Abstract
The order Macrodasyida (Gastrotricha) includes over 350 marine species, and only 3 freshwater species (Marinellina flagellata, Redudasys fornerise, R. neotemperatus). Herein we describe a new freshwater species of Macrodasyida, Redudasys brasiliensis sp. nov., from Brazil through an integrative taxonomic approach. The external morphology and internal anatomy were investigated using differential interference contrast microscopy, confocal microscopy, scanning and transmission electron microscopy. The systematization of the new taxon was inferred by nuclear (18S and 28S) and mitochondrial (COI) genes, and its intra-order relationships were assessed using data from most of available macrodasyids. Phylogenetic analyses yielded congruent trees, in which the new taxon is nested within the family Redudasyidae, but it was genetically distinct from the other species of the genus Redudasys. The new species shares the gross morphology and reproductive traits with other Redudasyidae and the presence of only 1 anterior adhesive tube per side with Redudasys neotemperatus, but it has a specific pattern of ventral ciliation and muscle organization. Results support the hypothesis that dispersion into fresh water habitats by Macrodasyida and Chaetonotida taxa occurred independently and that within Macrodasyida a single lineage invaded the freshwater environment only once. Furthermore, the Neotropical region seems to be peculiar for the evolution of the freshwater macrodasyid clade.
Similar content being viewed by others
Introduction
Gastrotricha is a group of aquatic, free-living microinvertebrates (80 µm to 3,000 µm in body length) divided into two orders: Macrodasyida and Chaetonotida1,2,3,4. The taxon Macrodasyida comprises over 350 worm-like species5, all interstitial in marine and estuarine habitats, except Marinellina flagellata Ruttner-Kolisko, 1955 collected from Austria, Redudasys fornerise Kisielewski, 1987 from Brazil and R. neotemperatus Kånneby & Kirk, 2017 from the U.S.A.6,7,8,9. However, at least two other unidentified freshwater macrodasyid species were collected from Brazil (Marinellina sp.10 and Redudasys sp.11).
Ruttner-Kolisko6 described for the first time a new freshwater species belonging to the order Macrodasyida, Marinellina flagellata, from the hyporheic region of an Austrian river, based on two likely immature specimens (the author reported that she could not see the reproductive organs). More than thirty years later Kisielewski7 described Redudasys fornerise, the first undoubted taxon of the order Macrodasyida recorded from a freshwater reservoir in Brazil. Todaro et al.8 redescribed the latter species few years ago. Recently, Garraffoni et al.11 found many specimens belonging to the genus Redudasys showing some minor differences from Redudasys fornerise, and Araújo et al.10 reported the occurrence of an undescribed species provisionally assigned to the genus Marinellina in South America for the first time. Furthermore, Kånneby & Wicksten12 collected one specimen of a putative new species of Redudasys from the Edwards Aquifer, Texas (U.S.A.): that was the first record of the genus from the Nearctic biogeographic region. Three years later, Kånneby & Kirk9 formally described the unnamed species reported by Kånneby & Wicksten12 as Redudasys neotemperatus based on specimens collected in Oregon (U.S.A.). An additional finding of a macrodasyid gastrotrich specimen much likely belonging to Marinellina flagellata was made by J.M. Schmidt-Araya in a different Austrian stream (Todaro et al.8).
Integrative taxonomy is a new approach for taxonomic and systematic studies that has spread in recent years13,14,15. The basic idea is to integrate information from different sources: ecological data, molecular data from nuclear and mitochondrial DNA, and morphological characters. In order to better understand the evolutionary history of the studied taxa, the data integration can be done by cumulation or congruence frameworks13. In the former case, the species identification is performed by a cumulative set of characters, without necessarily requiring concordance among all of them. Thus, a single set of characters considered good can be used to support the hypothesis about the new species. On the other hand, the framework of integration by congruence is based on a lineage divergence and the hypothesis under the assumption is based on the integration or coherent pattern of two or more independent sources of characters13. Considering that the advent of digital techniques in optical and electron microscopy have speed up the finding of new morphological characters and that new cybernetic infrastructures have been developed for analyzing and integrating data, the documentation and dissemination of data have greatly been improved16.
Herein, we formally describe a new species of freshwater Macrodasyida from Brazil through the approach of integrative taxonomy with the aim to investigate the relationships of the new taxon within the order. Morphological techniques (DIC, CLSM, SEM, TEM) and multigene molecular analyses (18S rRNA, 28S rRNA and COI mtDNA) were applied. We chose to integrate the data examined in this study by congruence: this framework has high confidence to promote taxonomic stability because the hypothesis regarding the description of the new species is supported by several character sets13. A study of additional specimens of Redudasys fornerise allowed a comparison within the genus.
Results
Taxonomic account
Phylum Gastrotricha Metschnikoff, 1865
Order Macrodasyida Remane, 1925 [Rao & Clausen, 1970]
Family Redudasyidae Todaro, Dal Zotto, Jondelius, Hochberg, Hummon, Kånneby & Rocha, 2012
Genus Redudasys Kisielewski, 1987
Emended diagnosis of the family Redudasyidae
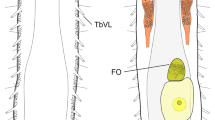
Macrodasyids from 302 to 400 µm in total length, with rounded head bearing several sensory cilia but without tentacles or ocelli. Lateral trunk margins even, without indentations or protrusions. Posterior end two-lobed, without a peduncle. Cuticular covering smooth, without scales or spines. Adhesive apparatus consisting of anterior (TbA) and posterior tubes (TbP); ventrolateral tubes (TbVL) may also be present (Anandrodasys). TbA, 1–3 per side: 1 tube per side arising from a lateral pouch, or 2–3 tubes of unequal length per side, borne from a common base and emerging from a ventrolateral furrow (Redudasys) or inserted in parallel (Anandrodasys), protruding obliquely to the rear. TbP, 4–12 in total, distributed symmetrically at the end of the two caudal lobes. TbVL if present at all, 5–6 per side, along the anterior intestinal region. Dorsal tubes (TbD) and lateral tubes (TbL) absent. Longitudinal muscles visibly cross-striated. Ventral ciliation of different patterns in the 2 genera: Redudasys: paired fields posterior to the mouth and along the pharyngeal region (U04–U36), and unpaired patches along the median line of the trunk region (U39–U50) or arranged in two large longitudinal bands close to each other, extending from the mouth ring to the anterior trunk (U48) and then merging into a single unpaired band to the caudal end; Anandrodasys: an unified field posterior to the mouth splits into 2 paired longitudinal bands, with the medial along the pharyngeal region, and the lateral extending up to the anus posterior to which is an isolated unpaired patch. Mouth, terminal or slightly subterminal; buccal cavity inconspicuous. Pharynx bearing pores at base, opening ventrolaterally. Intestine straight; anus ventral. Protonephridia present, 3 per side. Parthenogenetic; ovaries paired in hindgut region, with oocytes maturing anteriorly; male apparatus unknown; frontal and caudal organs unknown. Interstitial, marine or freshwater.
Type genus: Redudasys Kisielewski, 1987; other genus: Anandrodasys Todaro, Dal Zotto, Jondelius, Hochberg, Hummon, Kånneby & Rocha, 2012.
Redudasys brasiliensis sp. nov. (Figs 1–4)
Redudasys brasiliensis sp. nov. Optical DIC micrographs. (a) Habitus. (b) Anterior body end. (c) Dorsal posterior region. (d) Posterior and caudal regions. Scanning electron micrographs. (e) Habitus in dorsolateral view and close-up of an anterior adhesive tube (inset). c = locomotory ventral ciliation, e = egg m = mouth, ph = pharynx, sb = sensory bristles, TbA = anterior adhesive tubes, TbP = posterior adhesive tubes.
Redudasys brasiliensis sp. nov. Transmission electron micrographs. (a) Longitudinal section of the anterior body region showing the mouth and the whole pharynx. Note the ventrolateral muscles running from the lateral pouch. (b) Longitudinal section of an anterior adhesive tube arising from a lateral pouch; the insertion of ventrolateral muscle is visible. (c) Longitudinal section of anterior body end, showing the terminal mouth, the buccal cavity lined with a thin cuticle (arrow) and the pharyngeal longitudinal muscles inserted at mouth ring. (d) Cross-section of the pharynx, showing the Y-inverted lumen and the insertion points of the TbA (arrows). (e) Longitudinal sections of a protonephridium, showing the nuclear and filter region of the terminal cell. (f) Longitudinal sections of a protonephridium, showing, anal cell with a single luminal cilium and a nephridiopore cell. bc = buccal cavity, c = cilium, cc = canal cell, fr = filter region, m = mouth, n = nucleus, nc = nephridiopore cell, plm = pharyngeal longitudinal muscles, TbA = anterior adhesive tubes, tcl = terminal cell, vlm = ventrolateral muscles.
Redudasys brasiliensis sp. nov. Reconstruction of the musculature from confocal laser scanning microscopy images. (a) Schematic drawing of musculature in dorsal view. (b) Schematic detail of ventral musculature in the pharyngeal and the caudal regions. (c) Volocity-rendered 3D view of muscles in lateral view. Confocal micrographs of phalloidin-stained specimens. (d) Helicoidal muscles in the pharynx (arrow). (e) Detail of the pharynx posterior end with pharyngeal pores. asm = anterior semicircular muscle, cm = circular muscles, dlm = dorsal longitudinal muscles, llm = dorsal longitudinal muscles, mr = mouth ring, plm = pharyngeal longitudinal muscles, pp = pharyngeal pore, psm = posterior semicircular muscle, vlm = ventrolateral muscles.
Diagnosis
Macrodasyid 302–376 µm in total body length. Body separated into head, trunk and caudal regions. Head rounded, bearing several sensory cilia, without tentacles or ocelli. Cylindrical trunk, posterior end two-lobed, without a peduncle. Cuticular covering smooth, with no ornamentation. Paired tactile bristles of equal length along the body sides. Adhesive tubes only at the anterior and posterior body ends. TbA, only one tube on each head side, arising from a ventrolateral pouch. TbP, two symmetrical groups, each composed of two tubes of unequal length arising from a common base. TbD and TbL absent. Ventral ciliation arranged in two large longitudinal bands close to each other, extending from the mouth ring to the anterior trunk then merging into a single band up to the caudal body end. Mouth terminal; buccal cavity inconspicuous, lined with thin cuticle. Cylindrical pharynx with pharyngeal pores at base. Circular muscles surround the entire pharynx and are in turn surrounded by helicoidal muscles. Ten to twelve longitudinal muscles are external to both circular and helicoidal muscles, and extend from the mouth ring to the caudal end. Two of these muscles, the ventrolateral bands, insert anteriorly on a transverse muscle at the region of the TbA, and caudally on a second transverse muscle. Muscles of the trunk include longitudinal bands along the intestine that are surrounded by somatic circular muscles; the paired ventrolateral longitudinal muscles run along their length externally to the somatic circular muscles. Protonephridia present. Intestine straight, narrowing posteriorly; anus ventral. Parthenogenetic; paired ovaries in posterior region of intestine, with oocytes maturing anteriorly; male system absent; caudal and frontal organs absent. Freshwater, interstitial.
Etymology
The specific name refers to the country where the species was found.
Species-specific characters
Ventral ciliation made of 2 large longitudinal bands close each other and merging into one at the anterior trunk region. Longitudinal muscles of the trunk up to posterior end: 2 dorsal from mid-pharynx and 2 lateral branching into 3 parallel bundles; thin anterior transverse muscle connecting the two ventral longitudinal muscles.
Material
Holotype. Adult, collected from sandy pebble-sediment on November 20, 2013, at 0.3 m depth in the Soberbo Stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil, mounted on glass slide, deposited at the Zoological Museum, State University of Campinas, Brazil, accession number ZUEC GCH 05.
Paratypes. Six adult specimens and one subadult specimen, collected from sandy pebble-sediment on November 19–20, 2013, at 0.3 m depth in the Soberbo Stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil, mounted on glass slide, deposited at the Zoological Museum, State University of Campinas, Brazil, accession numbers ZUEC GCH 06 (2 specimens), ZUEC GCH 07 (2 specimens) and ZUEC GCH 08 (2 specimens).
Repository
urn:lsid:zoobank.org:pub:B5C8B48B-FA7E-45DE-8160-DFA80204EA06.
Other material
Ten additional specimens collected on September 10, 2010 and on July 07, 2014 in the Água Limpa Stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil and two specimens collected on May 3, 2010 in an unnamed stream, São Francisco drainage basin, Sempre-Vivas Federal Park, Diamantina, State of Minas Gerais, Brazil were observed alive and are no longer available. Four were filmed and videos were deposited at the Zoological Museum, State University of Campinas, Brazil, accession numbers ZUEC GCH 51 to ZUEC GCH 54.
Three adult specimens, collected on July 07, 2014 in Água Limpa Stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil, were mounted for SEM, and kept at the State University of Campinas.
Four adult specimens collected on March 15, 2014 in Água Limpa stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil, were prepared for TEM and are no longer available.
Three adult specimens collected on August 20, 2015 in Água Limpa stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil, were prepared for confocal analysis and are no longer available.
Four adult specimens, collected in Veado Pool and Preto River, Jequitinhonha drainage basin, São Gonçalo do Rio Preto, State of Minas Gerais, Brazil (March 2013 and June 2015) and eight adult specimens collected in Água Limpa and Soberbo streams, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil (April 2015, February 2016 and December 2017), were used for DNA extraction and are no longer available (Supplementary Table S1).
Description
The description is based on both the holotype and 7 paratypes (Fig. 1a,b; Table 1). Worm-like body, flattened ventrally, vaulted dorsally, 302–376 μm in total length. Body well-delimited into head/trunk/caudal regions. Head width (U17) 44–65 μm, anterior trunk width (U40) 30–64 μm, midtrunk width (U60) 33–68 μm, and caudal base width (U93) 18–30 μm. Head rounded anteriorly, without tentacles or ocelli. Caudal end bearing two symmetrical groups each of two tubes of unequal length arising from a common base.
Cuticular covering smooth, without scales, spines or epidermal glands, (Figs 2a–e and 3a,b). Protonephridia composed of 3 cells: proximal terminal cell, canal cell and distal nephridiopore cell (Fig. 3e,f).
Ciliation
Sensory cilia distributed irregularly, scattered around the mouth (ca.10 μm in length) and along the anterolateral head margin (ca. 30 μm in length). Six paired dorsolateral sensory bristles of equal length (ca. 20 μm in length) along the body sides and one paired bristle on each caudal appendage (Fig. 2c,d). Ventral locomotory ciliation arranged in two large longitudinal bands close to each other, extending from the mouth ring to the anterior trunk (U48) and then merging into a single band to the caudal body end (Fig. 1b).
Adhesive tubes
Only TbA and TbP are present (Figs 1a,b, 2b,d and 3b). TbA, 1 per side, 15 μm long in the holotype, located at U16–18, and arising from a deep ventrolateral pouch (Fig. 3b,d). Lateral and dorsal tubes absent (Figs 1a,b and 2a,e). TbP, 2 per side, of unequal length, the outer 18–21 μm long and the inner 10–15 μm (Figs 1a,b and 2d,e).
Digestive tract
Mouth opening terminal and rounded, 10–29 μm in diameter (26 μm in the holotype). Buccal cavity inconspicuous, 14 µm long, lined with thin cuticle and supported by a strong musculature (Fig. 3a,c). Cylindrical pharynx, 65–147 μm long and 20–34 μm wide (Figs 2a and 3a). Triradial inverted-Y lumen (Fig. 3d). Poorly developed ventrolateral pores occur at the posterior end of the pharynx (U36) (Fig. 4e). Intestine slightly narrowing posteriorly, anus at ca. U90.
Reproductive system
Male reproductive organs not seen, species probably parthenogenetic. Two ovaries lateral to the posterior intestine (ca. U82); usually one fully-grown egg (120 μm long) is visible dorsally (Fig. 2a).
Muscular system
Thin circular muscles, very numerous, surround the pharynx for its whole length (U01 to U43). Circular muscles of the intestine (U44 to U82) are less numerous and more regularly spaced; muscles noticeably closer each to other at the caudal end (U83 to U93) (Fig. 4a,c). Along the pharynx, 10 to 12 large, longitudinal muscles are inserted at the mouth ring and extend to the PhJIn (U43): they get thinner posteriorly (Figs 3c and 4a–c) and are external to circular muscles. One pair of dorsal longitudinal muscles extends from the mid pharynx to the caudal end (U23-U95) (Fig. 4a,c). They are external to circular muscles, except in the posterior pharyngeal region (U31-U37) and in the caudal body region (U83-U93), where they lie beneath the circular muscles (Fig. 4a,c). One pair of lateral longitudinal muscles extend from the PhJIn (U43) and branch into 3 parallel, thin bundles on each side from U61 to the body posterior end (U93). These longitudinal muscles are surrounded by circular muscles for their whole length (Fig. 4a,c).
The largest muscles in the body are two longitudinal muscles that are inserted laterally to the pharynx in the region of the TbA (U14) and run ventrolaterally extending up to the caudal appendages (Figs 3a,b and 4a–c). These muscles are connected anteriorly (U16) by a thin ventral transverse muscle and posteriorly (U96) by another transverse semicircular muscle (Fig. 4b,c).
Helicoidal muscles were observed along the pharyngeal region, ending at the anterior intestinal region (U48) (Fig. 4d).
Ecology
Species relatively common in sediment of the Jequitinhonha drainage basin sites; rare in sediment of the São Francisco drainage basin sites.
Redudasys fornerise Kisielewski, 1987
Redudasys fornerise. Optical DIC micrographs. (a) Body anterior region. (b) Posterior and caudal regions. (c) Close-up of a pair of anterior adhesive tubes. Scanning electron micrographs. (d,e) Close-up of the anterior adhesive tubes. (f) Ventrolateral view of the posterior region. c = locomotory ciliation, m = mouth, ph = pharynx, sc = sensory cilia, TbA = anterior adhesive tubes, TbP = posterior adhesive tubes.
Redudasys fornerise. Reconstruction of the musculature from confocal laser scanning microscopy images. (a) Schematic drawing of musculature in ventral view. (b) Volocity-rendered 3D view of muscles in ventral view. Confocal micrograph of a phalloidin-stained specimen. (c) Detail of the pharynx posterior end with pharyngeal pores. asm = anterior semicircular muscle, cm = circular muscles, llm = lateral longitudinal muscles, mr = mouth ring, plm = pharyngeal longitudinal muscles, pp = pharyngeal pore, psm = posterior semicircular muscle, vlm = ventrolateral muscles.
Redudasys fornerise - Todaro et al.8.
Redudasys sp. – Garraffoni et al.11.
Material
Three specimens collected in July 07, 2014 in Água Limpa Stream, Jequitinhonha drainage basin, Diamantina, State of Minas Gerais, Brazil, were observed alive and filmed. The specimens are no longer available, but videos were deposited at the Zoological Museum, State University of Campinas, accession numbers ZUEC GCH 51, ZUEC GCH 55 and ZUEC GCH 56. One adult specimen collected in September 15, 2015 in Broa Reservoir, Tietê/Jacarei drainage basin, São Carlos, State of São Paulo, Brazil, prepared for confocal analysis and no longer extant. Two additional adult specimens collected in April 11, 2015 and an another two in September 15, 2015, in Broa Reservoir, São Carlos, State of São Paulo, Brazil, mounted for SEM and kept in the first authors’ collection at State University of Campinas.
Description
The external and internal morphology are in accordance with the original description7 and the subsequent redescription8 (Figs 5a–f and 6a–c). It is important to highlight that this species has 2 TbA per side, the inner tube shorter than the outer, with a common base that arises from a ventrolateral furrow (Fig. 5c–e). Here we only focus on the muscular system (U46 to U98) (Fig. 6a–c).
Thin circular muscles, very numerous, surround the pharynx for its whole length (U01 to U45), whereas along the intestinal region (U49 to U93) they are in low number and regularly spaced. Eight large longitudinal pharyngeal muscles are inserted at the mouth ring and extend to U45, externally to circular muscles. A group of 4 thick longitudinal muscles per side, internal to the circular musculature, extend ventrally from the PhJIn (U45) to the body posterior end (U96) (Fig. 6a,b).
Two very large longitudinal muscles arise laterally to the pharynx (U13), at TbA insertion (Fig. 6a,b), turning ventrolaterally at U20 and extending up to the posterior body end. They are connected ventrally by two thick transversal muscles at U13 and at U96 respectively (Fig. 6a,b).
Helicoidal muscles were observed along the pharynx, ending at around the PhIJ (U51), but their insertion point could not be observed.
Ovaries and testes not observed; caudal and frontal organs not observed.
Phylogeny
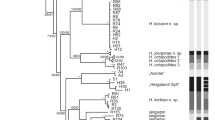
The final alignment of each dataset yielded 2149 positions in 18S rDNA, 4816 positions in 28S rDNA and 730 positions in COI mtDNA). Maximum Likehood (Supplementary Fig. S1) and Bayesian Analysis (Figs 7, 8) analyses yielded congruent topologies. For multigene analysis, the phylogeny showed monophyly of the families Redudasyidae, Thaumastodermatidae, Turbanellidae, Dactylopodolidae, Cephalodasyidae and Macrodasyidae supported in most cases with high bootstrap and Bayesian posterior probability values. Within Redudasyidae, Redudasys brasiliensis sp. nov. and Redudasys fornerise nested together and this clade formed the sister group of R. neotemperatus: all these relations are supported by high bootstrap and Bayesian posterior probability values (respectively, 100 and 1; Supplementary Fig. S1, Fig. 7). The clade with all specimens of Redudasys brasiliensis sp. nov. showed high bootstrap (99; Supplementary Fig. S1) but low Bayesian posterior probability (0.6; not shown in Fig. 7) values.
For the analysis of the COI mtDNA, the gene tree only showed the monophyly of the families Dactylopodolidae and Redudasyidae. Redudasys brasiliensis sp. nov. resulted as the sister-group of a clade composed by R. fornerise and R. neotemperatus, all supported by high bootstrap and Bayesian posterior probability values. All specimens of Redudasys brasiliensis sp. nov. were resolved as a monophyletic group, but with relative low bootstrap (61; not shown in Supplementary Fig. S1) and Bayesian posterior probability values (0.83; not shown in Fig. 8). The specimens of R. brasiliensis sp. nov. from São Gonçalo do Rio Preto and from Diamantina formed two distinct monophyletic clades highly supported by bootstrap (respectively, 98 and 97; Supplementary Fig. S1) and Bayesian posterior probability values (respectively, 1 and 1; Fig. 8).
The genetic distance (Table 2) of the haplogroups within each of the two Redudasys brasiliensis sp. nov. populations was very low (São Gonçalo do Rio Preto: p-distance = 0.3%; Diamantina: p-distance = 0.2%) as well as was between the two populations (4.2%). Comparing the values of the two populations with the two other species, the variation was higher: p-distances between the Diamantina clade of R. brasiliensis and R. fornerise ranged from 14.2% to 14.4%; p-distances between the São Gonçalo do Rio Preto clade of R. brasiliensis and R. fornerise ranged from 13.8% to 14.0%; p-distances between the Diamantina clade of R. brasiliensis and R. neotemperatus ranged from 15.6% to 15.8%; p-distances between the São Gonçalo do Rio Preto clade of R. neotemperatus and R. fornerise ranged from 14.6% to 14.8%. Thus, the relation of the maximum within-group genetic distance and the minimum between-group distances supported the identification of four candidate species (Redudasys brasiliensis sp. nov. population São Gonçalo do Rio Preto; R. brasiliensis sp. nov. population Diamantina; R. fornerise; R. neotemperatus).
The parsimony network representing 10 sequences detected a total of seven haplotypes of Redudasys brasiliensis sp. nov. (Supplementary Fig. S2). As in the COI mtDNA gene tree, the haplotypes from the two populations were divided into two lineages. The latter species is separated from R. fornerise and R. neotemperatus by high number of mutations in the network (Supplementary Fig. S2).
Discussion
In this section, we discuss some morphological and biogeographical patterns that may be associated with the diversification of freshwater macrodasyids.
Comparison among members of Redudasyidae
Members of family Redudasyidae show few external and internal morphological characters and hence the few characters concerning the adhesive system and locomotory cilia take on systematic significance12. Originally7,8, the main diagnostic characters of the genus Redudasys were the possession of 2 TbA per side and a ventral locomotory ciliation arranged into separate ciliary fields of unequal size (paired in the pharyngeal region and unpaired along the median trunk region). Anandrodasys, the only marine genus of the family, shows instead 3 TbA per side and ventral cilia arranged into an unified field posterior to the mouth and splitting into 2 paired longitudinal bands of different length.
Kånneby and Kirk9 described the first Redudasys species with the presence of a single pair of anterior adhesive tubes, as observed in R. brasiliensis sp. nov. Garraffoni et al.11. also reported an unnamed species of Redudasys with a single TbA per side from Brazil, but in this case that observation was a misinterpretation. During the present study we had the opportunity to collect fresh material and analyze Redudasys specimens from one of the sites previously investigated11. This allowed us to verify that these individuals in fact had 2 TbA of unequal size per side arising from a common base just like in R. fornerise. Thus, we considered that the Redudasys specimens found by Garraffoni et al.11. can be treated as R. fornerise.
The new species can be distinguished from Redudasys neotemperatus by: a different pattern of locomotory ciliation (i.e., two large longitudinal bands close to each other, extending from the mouth ring to the anterior trunk, then merging into a single band up to the caudal body end instead of regularly spaced paired tufts in R. neotemperatus), a greater body size (302–376 µm instead of 220–284 µm), a lower number of dorsolateral sensorial cilia (6 pairs instead of 13–14). Furthermore, Redudasys brasiliensis sp. nov. is also characterized by pharyngeal pores very poorly developed, unlike the other species of the family.
Redudasys brasiliensis sp. nov. and R. fornerise possess a quite similar muscular system, despite some specific muscles showing clearly distinct morphological patterns. The circular muscles of R. brasiliensis sp. nov. are external to the dorsal longitudinal muscles in the posterior pharyngeal region whereas in R. fornerise, they are internal to all longitudinal muscles. Moreover, the main longitudinal body muscles are ventrolateral in R. brasiliensis sp. nov. but clearly ventral for almost the whole length in R. fornerise. Finally, both species show two semicircular transverse muscles connecting the two ventrolateral muscles, but the anterior one is clearly thinner in R. brasiliensis sp. nov. than in R. fornerise.
The two populations of Redudasys brasiliensis sp. nov. were considered two separated species based on “4X rule” approach to delimiting species using DNA sequences15,17. However, this result can be biased due to limited number of individuals and candidate species (respectively, 12 and 3–4) and also because two of these 3–4 candidate species (R. fornerise and R. neotemperatus) had only 1 sequence each available in the GenBank (Supplementary Table S1). Thus, until more specimens of the three species can have their DNA sequenced, we still considered the individuals of both São Gonçalo do Rio Preto and Diamantina populations belonging to the same species, i.e., R. brasiliensis sp. nov.
Araújo et al.10. identified two freshwater macrodasyid specimens found in Brazil as belonging to genus Marinellina, basically because they had a single TbA per side. A subsequent study18 on the spatial and temporal distribution patterns of freshwater meiofaunal taxa in a lotic system in Brazil found additional macrodasyid specimens with a single TbA per side and reported them as an unidentified species of Redudasyidae. Considering the diagnostic character of the number of TbA, the metric data which fall in the measurement range of Redudasys brasiliensis sp. nov., as well as the geographic origin of these specimens, we consider the specimens found in both studies as belonging to the new species. Furthermore, the putative assignment of these specimens to the genus Marinellina10 cannot be supported with certainty due to several reasons: (a) insufficiency of the original description of the genus Marinellina1,6,7, (b) uncertainty about the exact position of TbA in Marinellina specimens2 (c) absence of pharyngeal pores in Marinellina specimens, (d) geographic distance between the sampling sites of european individuals and of the specimens found by Araújo et al.10, respectively.
The new species joins the few other species of macrodasyids that lack a male system including the frontal and caudal accessory organs, useful to store foreign spermatozoa and to act as a copulatory organ – sensu Ruppert and Shaw19 –, respectively. In these species, that are Anandrodasys agadasys, Redudasys fornerise, R. neotemperatus (Redudasyidae), Paradasys pacificus (Cephalodasyidae) and Urodasys viviparus, Thaidasys tongiorgii (Macrodasyidae)20,21, a probable parthenogenetic condition occurs. Only the latter species has a muscular organ of unknown function, which appears similar in position and structure to the caudal organ of other Macrodasyida20.
Colonization of freshwater habitats by Macrodasyida
Marine Macrodasyida are widely distributed along coastlines of the world and some level of endemism can be found in the North Hemisphere22. However, the levels of diversity and endemism of freshwater macrodasyids are considerably lower than in marine ones and, until now, no hypothesis or factor was presented to shed light on this great faunistic heterogeneity.
The Neotropical region seems to be a special area for the speciation of freshwater Macrodasyida. In Brazil, Kisielewski7 described the first undoubted macrodasyid from inland waters (Redudasys fornerise); subsequently Garraffoni et al.11 and Araújo et al.10 reported other freshwater specimens of Macrodasyida and in the present study we describe Redudasys brasiliensis sp. nov. These findings highlight the presence and the relatively wide distribution of Macrodasyida in the inland waters of this geographical area.
As pointed out by Ribeiro23 the hydrographic basins where populations of Redudasys fornerise and R. brasiliensis sp. nov. were found have an ancient biogeographic component. The populations of R. fornerise from State of São Paulo and the specimens of R. fornerise and R. brasiliensis sp. nov. from State of Minas Gerais are all very far from the Atlantic shore, respectively around 280 km and 420–450 km. However, these distances were only established during the last sea-level fluctuations in the Holocene transgression, about 5,600 years ago24,25. It is important to highlight that the South American continent has not always responded as a single rigid blocks during most of its geological history. The old history of South America shows that this large landmass has an intricate mosaic of amalgamation and break-up of several distinct continental fragments record in different periods23,26. Looking into the tectonic-sedimentary evolution of some provinces in the South American Platform, such as areas of São Paulo and Minas Gerais States, it is possible to observe specific orogenic cycles and depositional ages27,28,29. The sampling site in the State of São Paulo is located in the Bauru Basin represented by sandstones and reddish siltstones formed mainly in the Upper Cretaceous (99.6 to 65.5 Mya)27. On the other hand, the sampling sites from State of Minas Gerais lie in Lower Espinhaço Basin that is a mountain chain built up mainly of quartzitic rocks formed by a sequence of an intracontinental rift-sag basin system that developed around 1,500 Mya (Mesoproterozoic) to 600 Mya (Neoproterozoic)28,29.
These mosaics of smaller microblocks were surrounded by seaway and the current sampling sites could have been much closer to shallow marine environments than today. In this perspective, environment-specific diversification of freshwater macrodasyids within continental waters happened after the incursions of ancestral marine populations. The phylogenetic position of the monophyletic lineages composed by all Redudasys species is clearly nested into Macrodasyida clade, suggesting that the freshwater macrodasyids invaded inland environment only once. Furthermore, the phylogenetic hypothesis supports an independent colonization of freshwater habitats by Macrodasyida and Chaetonotida. This scenario is in agreement with the hypothesis advanced by Todaro et al.8, and in contrast with that proposed by Kieneke et al.4, implying a single colonization event of freshwater habitats in the stem lineage of Abursata ((Redudasys+Marinellina) + Paucitubulatina).
Some questions about the biogeographical and evolutionary history of the freshwater Macrodasyida remains unanswered: (a) how did Redudasys lineages achieve their current distributions in independent freshwater systems of Nearctic and Neotropics?; (b) why appear Redudasys populations found in freshwater habitats of the Austral hemisphere to be more widespread than those in Boreal hemisphere?; (c) how can the awareness of the existence of Marinellina, a taxon incertae sedis, affect our current biogeographic and evolutionary understanding of Redudasyidae?
Faunistic observations
Until now, 76 freshwater species of Gastrotricha from Brazil have been reported7,8,10,11,30,31,32,33. As highlighted by Garraffoni et al.30, due to the low number of sampled sites and wide variety of Brazilian freshwater habitats (with distinct geological and abiotic conditions) the number of new species will increase considerably in the next years. As an example for this statement, both the recently described new genus and species Cephalionotus kisielewskii30 and Redudasys brasiliensis sp. nov. are psammic and from the same region, Diamantina Plateau in the Espinhaço Range. The geological structure of this area is composed of high altitude formations (more than 900 m), has a very specific vegetation known as rocky fields (“campos rupestres”), and shows special conditions of climate, soil, and water causing high rates of endemisms, especially of plant species34.
Methods
Samples of upper stream sediment were taken from 7 distinct stations (Supplementary Fig. S3): State of Minas Gerais: Jequitinhonha drainage basin - 1. Água Limpa Stream, Biribiri State Park, City of Diamantina (sandy substrate): 18°12′S – 43°37′W; 2. Soberbo Stream, City of Diamantina (sandy/rocky substrate): 18°11′S – 43°33′W; 3. Preto River, Rio Preto State Park, City of São Gonçalo do Rio Preto (sandy/rocky substrate): 18°06′S – 43°20′W; 4. Veado Pool, Rio Preto State Park, City of São Gonçalo do Rio Preto (sandy substrate): 18°06′S – 43°20′W; São Francisco drainage basin - 5. unnamed stream 1 in Sempre-Vivas Federal Park, City of Diamantina (sandy/rocky substrate): 17°50′S – 43°45′W; 6. unnamed stream 2 in Sempre-Vivas Federal Park, city of Diamantina (sandy/rocky substrate): 17°55′S – 43°48′W; State of São Paulo: 7. Broa reservoir, City of São Carlos (sandy substrate): 22°20′S – 47°48′W. The sampling sites 5 and 6 were the same reported by Araújo et al.10.
Living individuals were located by sorting the sediment poured into Petri dishes under a stereomicroscope Leica EZ4; they were mounted singly on glass slides, observed in vivo and anaesthetized with 2% MgCl2 under a PrimoStar Zeiss light microscope, and photographed and filmed using a Leica DM2500 microscope equipped with differential interference contrast optics (DIC) and a camera Moticam 2300 of 3.0 megapixel. The videos are available upon request from the first author.
The positions of morphological characters along the body were measured using the percentage units (U)35.
Scanning Electron Microscopy
For Scanning Electronic Microscopy (SEM) analysis, the specimens were fixed in 4% formalin, dehydrated through a graded series of ethanol, treated with HMDS (hexamethyldisilazane)36, mounted onto aluminum stub and coated with gold-palladium using sputter coating. Observations were carried out under a SEM JSM 5800LV, at the State University of Campinas (Brazil), and a SEM Philips 515, at the University of Urbino (Italy).
Transmission Electron Microscopy
For Transmission Electronic Microscopy (TEM) analysis, the specimens were fixed in 2% glutaraldehyde in a 0.1 M sodium cacodylate buffer (pH 7.4), post-fixed in 1% osmium tetroxide buffered with 0.1 M sodium cacodylate, washed in a clean 0.1 M sodium cacodylate buffer, dehydrated in a graded ethanol series and embedded in Araldite. Semi-thick and ultra-thin sections were cut with a LKB Ultrotome 2088 V microtome and contrasted with toluidine blue and lead citrate, respectively. The semi-thin sections were observed in transmission light under a VANOX AHBT3 Olympus optical microscope, whereas the ultra-thin sections (70–80 nm) were observed under a TEM Philips CM10 at the University of Urbino (Italy).
Confocal Laser Scanning Microscopy
For Confocal Laser Scanning Microscopy (CLSM) the specimens were fixed in 4% paraformaldehyde in 0.1 M phosphate buffer saline (pH 7.2) for at least one week. Specimens were then rinsed in PBS and stained with Alexa Fluor 488 phalloidin (Life Technologies) to document the musculature. Stained specimens were briefly rinsed in PBS before mounting in Fluoromount G (Southern Biotechnology Associates, Birmingham, AL) on glass slides. An Olympus FV 300 confocal laser-scanning microscope was used to observe the specimens at the State University of Campinas (Brazil). An Argon laser (488 nm) was used to excite the samples, and Olympus software was used to capture the images. Confocal z-stacks were collected and processed as TIF files and MOV video files. Files were further processed with the software Volocity (Perkin Elmer) to generate z-projections.
DNA extraction and PCR amplification
Genomic and mitochondrial DNA was extracted from entire individuals of Redudasys brasiliensis sp. nov. using a column-based method QIAamp DNA Micro Kit (Qiagen), following the manufacturer’s instructions. DNA samples were subjected to PCR amplifications in a reaction volume of 25 µL containing 12.5 µL of 2x Taq PCR Master Mix (Qiagen), 3 µL of DNA, 8.7 µL of water, and 0.4 µL (4 pmol) of specific primers. The primer sequences and PCR conditions are indicated in Supplementary Table S2. The amplification products were electrophoresed in 1% agarose gels containing SYBR Green (Life Technologies). The bands with expected sizes were excised and then purified using a QIAquick Gel Extraction Kit (Qiagen) or the PCR product was directly purified using Illustra ExoProStar 1-Step (GE Healthcare). The DNA fragments were sequenced using BigDye Terminator reactions in a 3730XL DNA Analyzer (Applied Biosystems) at the facility of the LaCTAD laboratory (Campinas, Brazil). The 18S rDNA, 28S rDNA and COI mtDNA partial DNA sequences of Redudasys brasiliensis sp. nov. were deposited in GenBank (Supplementary Table S2).
Sequences, alignments and data analyses
18S rDNA, 28S rDNA and COI mtDNA sequences were aligned separately with Mafft v.7.215 using the L-INS-I approach37. The best-fit substitution model was determined with jModelTest 2.1.438. As the number of species analyzed with three genes and deposited in the GenBank is low, we run two distinct analyses (combined dataset and only COI) under maximum likelihood (ML) and Bayesian inference methods (BA) frameworks. ML analysis using RAxML39 was run with a GTRCAT model with 1000 bootstrap replicates. BA analysis was done using MrBAYES v.3.2.340 using two different runs with four chains each for a maximum of 20 million generations (sampled every 500 generations). Best-fit evolutionary model was selected using Akaike Information Criterion. The analysis was stopped when the two runs reached convergence (average standard deviation of split frequencies lower than 0.01). Convergence and estimated sample size (ESS) were verified using TRACER v.1.5, and 10% of each run was discarded as burn-in. Both ML and BA analyses were performed using the CIPRES Science Gateway, San Diego Supercomputer Center41.
For combined dataset, sequences of 37 species were retrieved from GenBank (2 Cephalodasyidae; 3 Dactylopodolidae; 2 Macrodasyidae; 1 Planodasyidae; 1 Lepidodasyidae; 18 Thaumastodermatidae; 6 Turbanellidae; 4 Redudasyidae) and for COI mtDNA, sequences of 27 species deposited in GenBank were included in the analysis (2 Cephalodasyidae; 3 Dactylopodolidae; 1 Macrodasyidae; 1 Lepidodasyidae; 13 Thaumastodermatidae; 4 Turbanellidae; 3 Redudasyidae) (Supplementary Table S2). Three species belonging to the order Chaetonotida, family Xenotrichulidae: Draculiciteria tesselata (Renaud Mornant, 1968), Xenotrichula intermedia Remane, 1934 and X. velox Remane, 1927 were used as outgroups (Supplementary Table S2). These species were selected as outgroups because members of Xenotrichulidae appeared as the sister group of all other Chaetonotida (e.g. Kånneby et al.42).
Genetic distances
MEGA7 software43 was used to calculate average uncorrected pairwise distances (p-distances) within and between species and haplogroups; alignment gaps were not considered. Moreover, the reliability of the new species diversification was tested applying the K/ϴ model (formerly known as 4X rule, Birky17, Fontaneto et al.15).
Statistical parsimony networks
To visualize possible intraspecific relationships between the COI haplotypes for Redudasys fornerise, R. neotemperatus and Redudasys brasilensis sp. nov., statistical parsimony networks were calculated using the program PopART44. The raw networks produced by PopART were redrawn using software Adobe Photoshop CS6 (Supplementary Fig. S2).
References
Balsamo, M., d’Hondt, J. L., Kisielewski, J. & Pierboni, L. Global diversity of gastrotrichs (Gastrotricha) in fresh waters. Hydrobiologia. 595, 85–91 (2008).
Todaro, M. A. & Hummon, W. D. An overview and a dichotomous key to genera of the phylum Gastrotricha. Meiofauna Marina. 16, 3–20 (2008).
Hummon, W. D. & Todaro, M. A. Analytic taxonomy and notes on marine, brackish water and estuarine Gastrotricha. Zootaxa. 2392, 1–32 (2010).
Kieneke, A., Riemann, O. & Ahlrichs, W. H. Novel implications for the basal internal relationships of Gastrotricha revealed by an analysis of morphological characters. Zool. Scr. 37, 429–460 (2008).
Todaro, M. A. Marine Gastrotricha. In Gastrotricha World Portal http://www.gastrotricha.unimore.it/marine.htm (2017). Accessed 01 Nov 2017
Ruttner-Kolisko, A. Rheomorpha neiswestnovae und Marinellina flagellata, zwei phylogenetisch interessante Würmtypen aus dem Siisswasserpsammon. Öst. Zool. Zeitschr. 6, 55–69 (1955).
Kisielewski, J. Two new interesting genera of Gastrotricha (Macrodasyida and Chaetonotida) from the Brazilian freshwater psammon. Hydrobiologia. 153, 23–30 (1987).
Todaro, M. A. et al. Marine Sister for a Freshwater Puzzle. PLoS One. 7, e31740 (2012).
Kånneby, T. & Kirk, J. J. A new species of Redudasys (Gastrotricha: Macrodasyida: Redudasyidae) from the United States. Proc. Biol. Soc. Wash. 130, 128–139 (2017).
Araújo, T. Q., Alcantara, F. C. & Garraffoni, A. R. S. New records of Gastrotricha from Minas Gerais, Brazil. Stud. Neotrop. Fauna Environ. 48, 68–75 (2013).
Garraffoni, A. R. S., Araújo, T. Q., Lourenco, A. P. & Balsamo, M. New data on freshwater psammic Gastrotricha from Brazil. Zookeys. 60, 1–12 (2010).
Kånneby, T. & Wicksten, M. K. First record of the enigmatic genus Redudasys Kisielewski, 1987 (Gastrotricha: Macrodasyida) from the Northern hemisphere. Zoosystema. 36, 723–733 (2014).
Padial, P. J., Miralles, A., De la Riva, I. & Vences, M. The integrative future of taxonomy. Front. Zool. 7, 16 (2010).
Goldstein, P. Z. & De Salle, R. Integrating DNA barcode data and taxonomic practice: determination, discovery, and description. BioEssays. 33, 135–147 (2011).
Fontaneto, D., Flot, J. F. & Tang, C. Q. Guidelines for DNA taxonomy, with a focus on the meiofauna. Mar. Biodivers. 45, 433–451 (2015).
Ramírez, M. J. et al. Linking of digital images to phylogenetic data matrices using a morphological ontology. Syst. Biol. 56, 283–294 (2007).
Birky, C. W. Species detection and identification in sexual organisms using population genetic theory and DNA sequences. PLoS One. 8, e52544 (2013).
Araújo, T. Q., Checon, H. H. & Garraffoni, A. R. S. Alpha and beta diversity of freshwater meiofauna at different spatial scales in a Neotropical lotic system. Mar. Freshwater Res. 68, 538–548 (2017).
Ruppert, E. & Shaw, K. The reproductive system of gastrotrichs. 1. Introduction with morphological data for two new Dolichodasys species. Zool. Scr. 6, 185–295 (1977).
Todaro, M. A., Dal Zotto, M. & Leasi, F. An integrated morphological and molecular approach to the description and systematisation of a novel genus and species of Macrodasyida (Gastrotricha). PLoS One 10, e0130278 (2015).
Todaro, M. A., Leasi, F. & Hochberg, R. A new species, genus and family of marine Gastrotricha from Jamaica, with a phylogenetic analysis of Macrodasyida based on molecular data. Syst. Biodivers. 12, 473–488 (2014).
Garraffoni, A. R. & Balsamo, M. Is the ubiquitous distribution real for marine gastrotrichs? Detection of areas of endemism using Parsimony Analysis of Endemicity (PAE). Proc. Biol. Soc. Wash. 130, 198–211 (2017).
Ribeiro, A. C. Tectonic history and the biogeography of the freshwater fishes from the coastal drainages of eastern Brazil: an example of faunal evolution associated with a divergent continental margin. Neotrop. Ichthyol. 4, 225–246 (2006).
Suguio, K. et al. Flutuações do nível relativo do mar durante o Quaternário Superior ao longo do litoral brasileiro e suas implicações na sedimentação costeira. Braz. J. Geol. 15, 273–286 (1985).
Dominguez, J. M. L. The Coastal Zone of Brazil: an Overview. J. Coast. Res. 39, 16–20 (2006).
De Wit, M. J., Stankiewicz, J. & Reeves, C. Restoring Pan-African–Brasiliano connections: more Gondwana control, less trans-Atlantic corruption. In West Gondwana: Pre-Cenozoic Correlations Across the South Atlantic Region (eds. Pankhurst, R.J., Trouw, R.A.J, Brito-Neves, B.B. & de Wit, M.J.) 399–412 (Spec Publ Geol Soc London, 2008).
Garcia, M. D. G. M. et al. The inventory of geological heritage of the state of São Paulo, Brazil, methodological basis, results and perspectives. Geoheritage. 1–20; https://doi.org/10.1007/s12371-016-0215-y (2018).
Abreu, P. A. A. O Supergrupo Espinhaço da Serra do Espinhaço Meridional (Minas Gerais): o rifte, a bacia e o orógeno. Geonomos 3, 1–18 (1995).
Chemale, Jr et al. Unravelling a Proterozoic basin history through detrital zircon geochronology: the case of the Espinhaço Supergroup, Minas Gerais, Brazil. Gondwana Res. 22, 200–206 (2012).
Garraffoni, A. R. S., Araújo, T. Q., Lourenço, A. P., Guidi, L. & Balsamo, M. New genus and new species of freshwater Chaetonotidae (Gastrotricha: Chaetonotida) from Brazil with phylogenetic position inferred from nuclear and mitochondrial DNA sequences. Syst. Biodivers. 15, 49–62 (2017).
Kisielewski, J. Inland-water Gastrotricha from Brazil. Ann. Zool. 43, 1–168 (1991).
Garraffoni, A. R. S. & Melchior, M. P. New species and new records of freshwater Heterolepidoderma (Gastrotricha: Chaetonotidae) from Brazil with an identification key to the genus. Zootaxa. 4057, 551–568 (2015).
Minowa, A. K. & Garraffoni, A. R. S. A new species of Haltidytes Remane, 1936 (Gastrotricha: Chaetonotida: Dasydytidae) from an urban lagoon of Brazil with a phylogenetic reconstruction of the genus based on morphological data. Zool. Anz. 269, 100–109 (2017).
Giulietti, A. M. & Pirani, J. R. Patterns of geographic distribution of some plant species from the Espinhaço Range, Minas Gerais and Bahia, Brazil. In Proceedings of a Workshop on Neotropical Distribution Patterns (eds Heyer, W. R. & Vanzolini, P. E.) 39–69 (An. Acad. Bras. Cienc., 1988).
Hummon, W. D., Balsamo, M. & Todaro, M. Antonio. Italian marine Gastrotricha: I. Six new and one redescribed species of Chaetonotida. Ital. J. Zool. 59, 499–516 (1992).
Hochberg, R. & Litvaitis, M. K. Hexamethyldisilazane for Scanning Electron Microscopy of Gastrotricha. Biotech. Histochem. 75, 41–44 (2000).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods. 9, 772–772 (2012).
Stamatakis, A., Hoover, P. & Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 57, 758–771 (2008).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 19, 1572–1574 (2003).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees in Proceedings of the Gateway Computing Environments Worksho p (GCE) 1–8 (IEEE, New Orleans, LA, USA) (2010).
Kånneby, T., Todaro, M. A. & Jondelius, U. Phylogeny of Chaetonotidae and other Paucitubulatina (Gastrotricha: Chaetonotida) and the colonization of aquatic ecosystems. Zool. Scr. 42, 88–105 (2013).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Leigh, J. & Bryant, D. POPART: full-feature software for haplotype network construction. Methods Ecol. Evol. 6, 1110 (2015).
Acknowledgements
We express our gratitude to the Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG: ETC-00017-13), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq: 478825/2013), Rede ComCerrado Program (50.6121/2008-09), and Biota Program/ Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP: 2014/23856-0) for financial support; to Rick Hochberg for his help with the confocal analysis; to William Pinheiro da Costa and Diego Fontaneto for their help with the genetic distance analysis. We are grateful to anonymous referees for greatly improving the first version of the manuscript. We also thank Alexander Kieneke for his constructive criticism.
Author information
Authors and Affiliations
Contributions
A.R.S.G. conceived the ideas, and T.Q.A., A.P.L., L.G. and M.B. contributed to the original data. A.P.L. generated molecular data and A.R.S.G. performed most of the initial research and molecular analysis. T.Q.A. performed the confocal analysis. L.G. and M.B. performed TEM and SEM analyses. A.R.S.G., A.P.L., L.G. and M.B. drafted the original manuscript; the final version was written by all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Garraffoni, A.R.S., Araújo, T.Q., Lourenço, A.P. et al. Integrative taxonomy of a new Redudasys species (Gastrotricha: Macrodasyida) sheds light on the invasion of fresh water habitats by macrodasyids. Sci Rep 9, 2067 (2019). https://doi.org/10.1038/s41598-018-38033-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-38033-0
- Springer Nature Limited
This article is cited by
-
Trends on Gastrotricha research: a bibliometric analysis
Biologia (2024)
-
An Information-theoretic approach to dimensionality reduction in data science
International Journal of Data Science and Analytics (2021)
-
The curious and neglected soft-bodied meiofauna: Rouphozoa (Gastrotricha and Platyhelminthes)
Hydrobiologia (2020)
-
Non-destructive genome skimming for aquatic copepods
Conservation Genetics Resources (2020)
-
Biodiversity analyses in freshwater meiofauna through DNA sequence data
Hydrobiologia (2020)