Abstract
In the frame of the VITA mission of the Italian Space Agency (ASI), we addressed the problem of Space osteoporosis by using human blood-derived stem cells (BDSCs) as a suitable osteogenic differentiation model. In particular, we investigated proteomic and epigenetic changes in BDSCs during osteoblastic differentiation induced by rapamycin under microgravity conditions. A decrease in the expression of 4 embryonic markers (Sox2, Oct3/4, Nanog and E-cadherin) was found to occur to a larger extent on board the ISS than on Earth, along with an earlier activation of the differentiation process towards the osteogenic lineage. The changes in the expression of 4 transcription factors (Otx2, Snail, GATA4 and Sox17) engaged in osteogenesis supported these findings. We then ascertained whether osteogenic differentiation of BDSCs could depend on epigenetic regulation, and interrogated changes of histone H3 that is crucial in this type of gene control. Indeed, we found that H3K4me3, H3K27me2/3, H3K79me2/3 and H3K9me2/3 residues are engaged in cellular reprogramming that drives gene expression. Overall, we suggest that rapamycin induces transcriptional activation of BDSCs towards osteogenic differentiation, through increased GATA4 and Sox17 that modulate downstream transcription factors (like Runx2), critical for bone formation. Additional studies are warranted to ascertain the possible exploitation of these data to identify new biomarkers and therapeutic targets to treat osteoporosis, not only in Space but also on Earth.
Similar content being viewed by others
Introduction
Bone remodeling is an ongoing process, fundamental for adapting bone architecture to growth and repair processes, and for maintaining calcium homeostasis. Bone remodeling depends on the interaction between osteoblastic cells of mesenchymal origin and osteoclastic cells of hematopoietic origin.
Alterations of the equilibrium between bone resorption and bone formation can determine reduced bone density and increased fragility, hallmarks of osteoporosis1,2,3. On Earth, environmental elements (diet, physical activity, endocrine status, smoking, etc.) and genetic predisposition contribute to the onset of osteoporosis4,5,6.
In Space, however, astronauts lose on average more than 1% of bone mass per month and, while progression to osteoporosis is not an issue for short-term flights, it is indeed a handicap to long-term missions such as those on the International Space Station (ISS) and interplanetary flights. The cause of developing osteoporosis in Space is linked to low (micro- to zero-) gravity conditions, with possible contributions of cosmic ray radiation and circadian rhythm alterations7,8,9. In recent years, Space research has focused on osteoporosis in search for possible therapies able to counteract bone mass loss, crucial to prolong future manned Space missions.
Human blood-derived stem cells (BDSCs) are ideal candidates to study bone mass loss that affects astronauts, as these cells are autologous and pluripotent, and are able to differentiate into many different cell types. Of note, their osteogenic differentiation induced by rapamycin in the presence of suitable scaffolds has been widely studied10,11,12,13,14,15. Recently, rapamycin has also been approved for the treatment of bone pain in patients with bone metastases, due to its ability to counterbalance the effects of osteoclasts by promoting osteoblast activity16,17.
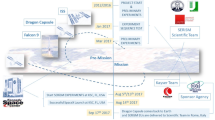
Recently, we performed the “SERiSM” (Role of the Endocannabinoid System in Reprogramming Human Pluripotent Stem Cells under Microgravity) project, that was selected by the Italian Space Agency (ASI) on a competitive basis, in order to: i) evaluate the osteogenic differentiation of BDSCs under real microgravity, and ii) interrogate the possible modulation of the so-called “endocannabinoid system” (i.e., a complex ensamble of bioactive lipids, their metabolic enzymes and binding receptors)18,19 by microgravity and its involvement in the osteogenic process. The SERiSM experiment was approved by the National Aeronautics and Space Administration (NASA) and was launched to the ISS on board the SpaceX Dragon Spacecraft CRS-12 from Cape Canaveral, Florida (USA), on August 14th, 2017, in the frame of the VITA mission of ASI (Fig. 1; for a detailed SERiSM mission profile see Supplementary Figure S1); then, it was incubated at 37 °C inside the ESA KUBIK incubator in the Columbus module onboard ISS. Preliminary data on the modulation of the endocannabinoid system under microgravity have been recently reported20. Here, we show that proteomic changes occur in BDSCs upon treatment with rapamycin, speaking in favour of an osteoblastic differentiation of these cells that shows a different timing under microgravity conditions compared to Earth.
It should be recalled that epigenetic regulation is deeply involved in bone homeostasis, and indeed histone modifications are considered the switches used by cells to control differentiation20,21,22,23,24. The expression of genes involved in bone remodeling, such as Runx2 and osteocalcin, is accompanied by the increase in methylation and acetylation of Runx2 promoter histones. Here, we also demonstrate that epigenetic modifications occur at H3 histone, and could support osteogenic differentiation of BDSCs on board the ISS.
Results
Proteomic changes
To study osteogenic differentiation of BDSCs under low gravity conditions (LG) on board the ISS and at ground gravity (GG) on Earth, expression levels of 15 proteins were analyzed as markers of pluripotency, commitment and stem cell differentiation (see Supplementary Table S1 for details). Because of the paucity of the biological material available, limited by the many constraints of Space research under true microgravity conditions, a “Proteome Profiler Array” (Human Pluripotent Stem Cell Array Kit) was used to analyze BDSCs at time zero (LG t0 and GG t0), and after 72 hours of differentiation (LG t72 and GG t72), as detailed in Materials and Methods. Thus, we were able to analyze 15 protein markers, some of which are important to determine stem cell phenotype, and others are necessary for stem cell commitment (Supplementary Table S1). The overall data obtained by proteome analysis are shown in Fig. 2a, and are described in detail below.
Proteome Profiler Array. (a) Human Pluripotent Stem Cell Array results on BDSCs controls (LG t0 and GG t0) and after 72 hours of differentiation (LG t72 and GG t72). (b) Statistical significance of proteomic profile experiments on BDSCs controls (LG t0 and GG t0). (c) Changes in protein expression measured as Δ% BDSCs LG t0/BDSCs GG t0. *p < 0.05, 0.0332 < **p < 0.0021, 0.0021 < ***p < 0.0001, ****p < 0.0001.
Effect of the space flight on BDSC controls
Figure 2b shows densitometric analysis of the data obtained from BDSCs LG t0 and GG t0 only. In both controls (LG and GG) BDSCs showed a pluripotent stem cell phenotype, as demonstrated by the expression of the transcriptional factors Oct3/4, Sox2 and Nanog. Figure 2c shows that almost all proteins analyzed increased on board the ISS, compared to Earth (BDSCs LG t0/GGt0). The difference in protein expression between the two controls (LG and GG) led us to normalize the data as Δ% for all timepoints, in order to compare all subsequent results.
Markers of pluripotency
Figure 3a shows protein expression changes (as Δ%) that occurred after 72 hours of differentiation on ground (GG BDSC t72/GG BDSC t0) and on board the ISS (LG BDSC t72/LG BDSC t0). The raw data of densitometric analysis are shown as insets of Fig. 3a, and relative p values are reported in Supplementary Table S2. Figure 3b shows the Δ% in LG BDSCs t72 and GG BDSCs t72 of the embryonic markers Sox2, Oct3/4, Nanog and E-cadherin, that are responsible for the maintenance of stem cell pluripotency. It was found that Sox2, Nanog and E-cadherin were less expressed in GG BDSCs than in LG BDSCs, while Oct3/4 decreased more markedly in LG BDSCs after 72 hours of osteogenic differentiation.
Proteome Profiler Array. (a) BDSCs GG and LG proteomic profile comparison after 72 hours of differentiation (Insets in panel a are the graphical statistical significance experiments on BDSCs GG t72/t0 and BDSCs LG t72/t0); (b–f) Changes in protein expression measured as Δ% BDSCs GG t72/t0 and BDSCs LG t72/t0 concerning: the decrease of the four pluripotent stem cell markers expression (b); the differences in expression of endoderm (c), trophoectoderm (d) and mesoendoderm (e) markers, and the differential expression of transcription factors (f). *p < 0.05, 0.0332 < **p < 0.0021, 0.0021 < ***p < 0.0001, ****p < 0.0001.
Markers of commitment
Figure 3c shows markers of endodermal commitment, such as PDX/IPF1 (insulin promoter factor 1), a transcription factor necessary for pancreatic development, and HNF-3b/FoxA2, that regulates the expression of hepatotrophic factor ALR in liver cells. Of note α-fetoprotein (AFP), a glycoprotein produced in early fetal life by the liver, was the only marker that did not change its expression in LG BDSCs t72 when compared to its LG BDSCs t0 control. In Fig. 3d, a marker of the trophoectoderm is shown: HCG (human chorionic gonadotropin), a glycoproteic hormone secreted during pregnancy by embryonic trophoblast cells. Also in this case, a difference between LG and GG BDSCs was observed after 72 hours of differentiation, along with a decreased expression on ground.
In Fig. 3e markers of mesoendodermal germinal layer are shown, namely: TP63/TP73L, a key player in embryonic urogenital development; Goosecoid (GCS), a homeobox protein expressed in cells that become pharyngeal endoderm, head mesoderm and notochord; and VEGF R2/KDR/Flk-1, that plays critical roles in skeletal development and directly controls differentiation and function of osteoblasts. Only VEGF R2/KDR/Flk-1 expression was found to increase in LG BDSCs t72, as expected during an osteogenic differentiation process.
Markers for transcriptional processes
Figure 3f highlights the changes in the expression of 4 transcription factors that are critical in determining whether the differentiation process can take place. Otx2 and Snail are negative transcriptional factors, necessary for the maintenance of the stemness phenotype, while GATA4 and SOX17 are considered fundamental in promoting differentiation, in particular towards osteogenesis. On board the ISS, BDSCs showed a remarkable increase of SOX17 and GATA4 after 72 hours, and a more pronounced expression of Otx2 and Snail when compared with the ground control cells.
Epigenetic modifications
Figure 4a shows the results on 8 epigenetic modifications of H3 histone (expressed as fold changes) in BDSCs under GG and LG conditions at t0, t48 and t72. Due to the paucity of the biological material, we focused on the following lysine di/tri-methylations: H3K79me2/me3, H3K27me2/me3, H3K4me3, H3K9me2/me3 and HK36me3. Indeed, these epigenetic modifications are known to be involved in: heterochromatization of DNA and transcriptional silencing (H3K27me2, H3K27me3 and H3K4me3, shown in Fig. 4b); chromatin remodeling and transcriptional activation during differentiation (H3K79me2, H3K79me3, H3K9me2 and H3K9me3, shown in Fig. 4c); and more directly osteogenic differentiation (HK36me3, Fig. 4d). Similar changes were found at LG and at GG, yet to different extents. An increased trimethylation of lysine 4 (H3K4me3) and di/tri-methylation of lysine 27 (H3K27me2/3) were observed in LG BDSCs (both t48 and t72), compared to GG BDSCs at t72; at the same timepoints di/tri-methylation of lysine 79 (H3K79me2/3) was also increased. Additionally, we found very high levels of di/tri-methylation of H3K9 in both LG and GG cells, while in LG BDSCs (t48 and t72) we observed an increased trimethylation of H3K36 compared to GG BDSCs.
Epigenetic H3 modifications during differentiation in LG and GG conditions (expressed as fold changes). (a) Global histone modifications occurred, respect to controls, at 48 (LG) and 72 hours (GG, LG) of osteogenic differentiation (b) Histone modifications concerning the gene silencing occurred during the differentiation process. (c) Histone modifications concerning the gene opening and chromatin remodeling during differentiation. (d) Histone modification related to osteogenic differentiation.
Discussion
Proteomic and epigenetic changes occurring in BDSCs under true low gravity conditions on board the ISS were interrogated in a model of osteogenic differentiation that had been previously validated on Earth10. The advantage of an easy accessibility of blood-derived stem cells, compared to other (e.g., mesenchymal) stem cells, makes them a promising tool to identify biomarkers and/or therapeutic targets potentially useful for monitoring and/or treating osteoporosis in Space11,12,13,14,15. Of note, our preliminary data on Earth demonstrated that already after 72 hours of treatment with rapamycin and a suitable scaffold, BDSCs differentiated with a mesoendodermal commitment, secreting calcium phosphate and forming hydroxyapatite10. Therefore, we chose to perform proteomic analysis at t0 and t72, also including a t48 timepoint to interrogate possible early changes in epigenetic regulation of gene expression. Protein expression analysis demonstrated changes in LG t72 samples when compared to the GG t72 controls. Almost all proteins analyzed increased on board the ISS compared to Earth (LG BDSCs t0 compared to GG BDSCs t0), suggesting that the flight towards the ISS affected per se gene expression. Thus, we chose to normalize the data in order to properly compare ISS and Earth samples.
In LG and GG samples at t72, a decrease in the expression of the 4 embryonic markers Sox2, Oct3/4, Nanog and E-cadherin was observed (Fig. 3b), suggesting that BDSCs were losing their pluripotency both on board the ISS and on Earth. Yet, under microgravity the process was more pronounced. Differences between LG and GG samples were more evident when endodermal, trophoectodermal and, more notably, mesoendodermal markers were analyzed (Fig. 3c,d,e). In particular, after 72 hours only LG cells showed an increase of VEGFR-2, that is fundamental to recruit osteoprogenitor cells and thus form bones25. In keeping with these data, Dhaliwal and colleagues have recently demonstrated enhanced osteogenesis in bone marrow-derived human mesenchymal stem cells through induction of VEGFR-226.
Taken together, these data suggested an earlier activation of the differentiation processes in LG versus GG cells, an hypothesis that was confirmed by comparing the expression of the 4 transcription factors Otx2, Snail, GATA4 and Sox17. While Otx2 decreased in both LG and GG cells, Snail decreased in LG cells and increased in GG cells; both Sox17 and GATA4 increased in LG cells and showed little expression in GG cells at t72 compared to t0 (Fig. 3f). Also the decrease of Otx, which counteracts commitment and promotes pluripotency27,28, supported our findings. The data on Sox17 appeared of particular interest, because Sox17 is a transcriptional regulator that promotes differentiation of pluripotent cells; indeed, Sox17-deficient embryonic stem cells do not differentiate into extraembryonic cells and keep on expressing pluripotency-associated transcription factors like Oct4, Nanog and Sox2. Instead, forced expression of Sox17 downregulates embryonic stem cell-associated gene expression, and activates genes responsible for differentiation29. Thus, low expression of Sox17 in GG BDSCs after 72 h of induction suggests that these cells are still at the beginning of the differentiation process, while Sox17 increase in LG BDSCs suggests that these cells are at a later stage of differentiation.
GATA4 is consistently expressed in pre-osteoblast cells and gradually down-regulated during osteoblast differentiation30. Guo and colleagues recently suggested a role for GATA4 in maintaining normal trabecular bone mass. Interestingly, both in vivo and in vitro reduction of GATA4 correlates with reduced Runx2 gene expression, along with reduced osteoblast mineralization31. It should be recalled that Runx2 is a transcription factor crucial for osteoblast differentiation32, as it is responsible for the synthesis of osteoblastic proteins like Osterix (Otx) and osteocalcin (Ocn)33,34,35. It has been suggested that GATA4 binds near the Runx2 promoter and enhancer, and helps maintaining open chromatin to regulate Runx2 expression that leads to bone mineralization31. In the same context, also Snail plays a crucial role in osteogenic differentiation by acting as a direct Runx2 repressor36. Thus, GATA4 and Snail have opposite roles in osteogenic differentiation by enhancing or repressing Runx2, respectively37,38. Taken together, in LG BDSCs the increase of GATA4 and Sox17 combined with the decrease of Snail and the increase of VEGFR-2 leads to activation of Runx2, and hence of osteogenesis.
In order to ascertain whether regulation of gene expression could depend on epigenetic mechanisms, we interrogated changes of histone H3 that is crucial in this type of gene control39. Indeed, during the differentiation process cells become more mature, lose pluripotency and undergo epigenetic rearrangements that shut off stemness genes, making cells resistant to returning to the undifferentiated state39. Figure 4a summarizes the overall modifications observed at t72 and at the earlier point t48. During stem cell differentiation, H3K4me3, H3K27me2/3, H3K79me2/3 and H3K9me2/3 residues are engaged in cellular reprogramming that controls gene expression39. We found that these changes did occur in LG cells (Fig. 4b and c, respectively).
The presence of di/tri-methylated H3K4 and H3K27 generally indicates inhibition of transcription and heterochromatinization, while during cellular differentiation their modulation is indicative of gene opening and closing39. Our results are consistent with literature data, reporting an increase in H3K27me3 and H3K4me3 at the beginning of differentiation, when chromatin is reassembled to silence genes responsible for stemness and to open lineage-specific genes only40.
The methylation of H3K79 (me2/3) seems also remarkable, because it is considered to impair cell reprogramming41. Indeed, it has been shown that a high level of H3K79 di/tri-methylation prevents reprogramming of an adult cell towards iPSC, while its reduction enhances the number of reprogrammed cells41,42.
Another epigenetic modification considered an obstacle to cellular reprogramming is the methylation of H3K9 (me2/3)43. H3K9me3 prevents initial binding of OSKM (Oct3/4, Sox2, Klf4 and c-Myc), thus hindering iPSC formation and reducing reprogramming efficiency44. We observed a high increase of H3K9me2/3, suggesting that BDSCs are leaving the pluripotent state to enter the differentiation program. Once again, it appears that the differentiation process builds up epigenetic barriers to drive cell fate, while stem cells do not show relevant histone modifications such as H3K79me2/3 and H3K9me2/345,46,47,48.
Our present results are in agreement with those by Dhaliwal and colleagues24, who demonstrated that epigenetic modifications like H3K9me3, H3K27me3 and H3K4me3 can be modulated to direct optimal osteoblastic differentiation within 72 hours, in the presence of osteogenic cues, and can regulate osteogenic differentiation of mesenchymal stem cells. An additional modification that we observed was trimethylation of H3K36 (Fig. 4d), that is known to increase during osteogenic differentiation49,50,51. H3K36me3 is required in promoters for Osterix, a specific transcription factor of osteoblasts52. Consistently, it increased both in LG and in GG BDSCs, confirming that osteogenic differentiation was activated. The different extent of epigenetic changes in LG versus GG BDSCs at t72 may be a corollary to the differences in protein expression observed at the same timepoint, indicating that these cells follow the same trends during their differentiation, but with an earlier start under LG conditions.
In conclusion, osteogenic differentiation of BDSCs induced by rapamycin follows a coordinated pattern in protein expression and epigenetic arrangement, as schematically depicted in Fig. 5. GG BDSCs at t72 are still in an osteogenic commitment phase, while the early expression of GATA-4 and Sox17 in LG BDSCs at t72 indicates that under microgravity conditions these cells are already in full osteogenic differentiation, rather than simply committed (Fig. 5a). Moreover, the results obtained in this investigation are in agreement with literature data showing how genes involved in embryonic stem cell maintenance exhibit high levels of H3K27me3 and H3K9me3, leading to a stable chromatin53. In addition, H3K36me3 must increase during osteogenic differentiation49, as we indeed observed in our cells (Fig. 5b). Overall, we suggest that rapamycin induces transcriptional activation of BDSCs through increased GATA4 and Sox17 that modulate Runx2 expression by means of VEGFR-2. In turn, Runx2 induces osteogenic differentiation by activating the transcription of Osterix (Osx) and Osteocalcin (Ocn) (Fig. 5c). Additional studies are warranted to ascertain the possible exploitation of these data to identify new biomarkers and therapeutic targets to treat osteoporosis, not only in Space but also on Earth.
From stem cells to osteoblasts: proteomic and epigenetic pathways. (a) Timeline of protein expression during osteoblastic differentiation. (b) Proposed epigenetic changes occurring during stem cell differentiation. (c) Proposed role of transcription factors and VEGFR2 during the first steps of osteogenesis differentiation.
Materials and Methods
Preparation of BDSCs
Human BDSCs were purified from blood samples, as previously reported12. Briefly, the nucleated blood cell fraction was isolated by ammonium chloride incubation (dilution 1: 3 in NH4Cl 1 M), centrifuged at 1023 g for 20 minutes and washed several times with phosphate-buffered saline (PBS), pH7.2 (Base Catalog BR0014G, Oxoid, Hampshire, England), to remove the majority of erythrocytes. Cells were then resuspended in 5 ml PBS and incubated for 72 h at 37 °C in the presence of 50 nM macrophage colony-stimulating factor (Base catalog M9170, Sigma-Aldrich, St. Louis, MO), and 5 μM gentamicin sulphate (Base catalog L0012, BioWest, Nuaillé, France). For the VITA mission experiments, 16 samples were prepared and distributed as follows: 6 BDSC controls (LG t0), 4 samples for the 48 hours BDSCs (LG t48), 6 samples for the 72 hours BDSCs (LG t72). In the proteomic and epigenetic experiments 3 LG t0, 2 LG t48 and 4 LG t72 were used. Healthy blood donors gave their written informed consent to the study according to the Legislative Decree n. 196/2003. The study was approved by the Ethical Committee of Campus Bio-Medico University of Rome and was conducted according to the ethical principles arising from Helsinki Declaration. Participation of the subjects was voluntary and unpaid.
SERiSM Hardware
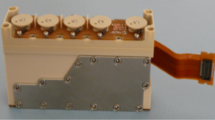
Cell growth and differentiation of BDSCs occurred in the hardware developed on purpose by Kayser Italia Srl, shown in Supplementary Figure S2.
The SERiSM Experiment Unit (KEU-RO) is a device capable of performing automatic cell culture of non–adherent cells in microgravity. It is equipped with reservoirs for chemicals and a culture chamber allowing cell growth in suspension. The scientific protocol is led by the KEU-RO electronics following a predefined timeline. Each KEU-RO Experiment Unit (EU) is made of a semi-crystalline thermoplastic polymer with excellent mechanical and chemical resistance properties, biologically inert. Cross contamination among the fluids chambers are avoided due to proper sealing gaskets. The EU itself provides one Level of Containment (LoC) that is increased to two by using KIC-SL container class. The experiment is fully autonomous; all the actions are electrically controlled by a predefined timeline uploaded into the on-board microcontroller. Housekeeping data are recorded during the mission and downloaded at re-entry.
The typical fluidic concept carries out the KEU-RO experimental protocol which relies on three main steps, i.e. Activation, Incubation, Fixation. On the whole, the actions performed by the fluidic system are led by preloaded spring actuators activated by the control electronics. Such mechanism pushes the pistons inward displacing the fluids (Activator or Fixative) contained into the chemicals reservoirs (Activator or Fixative reservoir) towards the Culture Chamber (CC). Short channels connect independently the reservoirs to the CCs so that cells are activated or fixed. The SERiSM Experiment Unit (KEU-RO) assembled with control electronics was integrated inside the KIC-SL containers (Kayser Italia Containers- Single Level), and then placed inside the BIOKON transportation Container for the upload onboard launcher. Onboard ISS the SERiSM Experiment Hardware was inserted inside the ESA KUBIK incubator in the ISS Columbus module, set at 37 °C.
Osteogenic differentiation
To promote in vitro osteogenic differentiation, BDSCs were grown in DMEM-F12 (Lonza, Belgium, BE12-719F) supplemented by 10% FBS (not-USA origin, Sigma, USA, F9665), 1% penicillin (100 units/ml)/streptomycin (100 mg/ml) (Lonza, Belgium, DE17-602E), rapamycin 10 nM as osteogenic inductor (Sigma-Aldrich, St. Louis, MO, R0395) and Bio-Oss scaffold (Geistlich, Switzerland, 30643.3/500079), as previously reported10,15.
Proteome profiler
Changes in protein expression were evaluated by a Proteome Profiler Array (Human Pluripotent Stem Cell Array Kit, R&D Systems, Abingdon, UK, ARY010), according to the manufacturer’s instructions. For the proteomic experiments, two samples were used for each timepoint, each in duplicate (BDSCs LG t0, BDSCs LG t72, BDSCs GG t0, BDSCs GG t72). Briefly, cells were solubilized in 100 μl lysis Buffer supplemented with protease inhibitor cocktail (Calbiochem, EMD Millipore, Billerica, MA USA, 535140-1 ML), centrifugated a 4 °C at 14000 g for 5 minutes and the supernatant collected. Protein concentrations were assessed by Bradford assay. For each sample 200 μg of proteins were used to perform the array. After development of autoradiographies, pixel density of each marker was evaluated using ImageJ analysis software and Microsoft Office and the results expressed in Relative-delta percentage as suggested by manufacter’s protocol:
Total histone purification
Histones were isolated using EpiQuik Total Histone Extraction Kit (Epigentek, Farmingdale, NY, #OP-0006) according to the manufacturer’s instructions. For epigenetic experiments one sample of BDSCs GG t0, two samples of BDSCs GG t72, one sample of BDSCs LG t0, two samples of BDSCs LGt48, two samples of BDSCs LG t72 were used. Briefly, the cellular pellet was resuspended in 1 ml Diluted Pre-Lysis Buffer (1X) and was placed on ice for 10 minutes under stirring to remove the plasmatic membranes. The lysate was centrifuged at 9391 g for 1 minute at 4 °C and the supernatant removed. The pellet was resuspended in 50 μl of the Lysis Buffer and incubated on ice for 30 minutes. The lysate was centrifuged at 13523 g for 5 minutes at 4 °C to remove the nuclear membranes and the supernatant transferred to a new tube. For each sample, 0.3 volumes of Balance Buffer were immediately added to the supernatant without DTT since its presence causes scaffold precipitates that hindered histones extraction. Histones concentration was assessed by Bradford assay.
Histone H3 modification
To evaluate H3 modifications the EpiQuik Histone H3 Modification Multiplex Assay Kit (Epigentek, Farmingdale, NY, #P-3100) was used according to the manufacturer’s instructions. Due to the limited amount of material available, the following histone modifications were considered for each sample: H3K4me3, H3K9me2, H3K9me3, H3K27me2, H3K27me3, H3K36me3, H3K79me2 and H3K79me3. The amount of each histone modification was compared to the own total histone and then expressed as a percentage, using the following formulas, as described in the manufacturer’s protocol.
To better highlight the results obtained, we reported each H3 modification as fold change, using the following formula:
Statistical analysis
Statistical analysis of proteomics data was performed with GraphPad Prism3 by multiple T test (one per line). Statistical significance was determined by using the Holm-Sidak method, with alpha = 0.05. Each row was analyzed individually. The test number was equal to 15. p < 0.05 (*), 0.0332 < p < 0.0021 (**), 0.0021 < p < 0.0001 (***), and p < 0.0001 (****), as reported in Supplementary Table S2.
References
Feng, X. & McDonald, J. M. Disorders of bone remodeling. Annu. Rev. Pathol. 6, 121–145 (2011).
Demontiero, O., Vidal, C. & Duque, G. Aging and bone loss: new insights for the clinician. The. Adv. Musculoskeletal Dis. 4, 61–76 (2012).
Rouhi, G. Biomechanics of Osteoporosis: The Importance of Bone Resorption and Remodeling Processes. In Tech. 3 (2012)
Sözen, T., Özışık, L. & Başaran, N. Ç. An overview and management of osteoporosis. Eur. J. Rheumatol. 4, 46–56 (2016).
Bartolozzi, E. The natural approach to osteoporosis. Clin. Cases Miner. Bone Metab. 12, 111–115 (2015).
Wood, C. L., Stenson, C. & Embleton, N. The Developmental Origins of Osteoporosis. Curr. Genomics 16, 411–418 (2015).
Demontis, G. C. et al. Human Pathophysiological Adaptations to the Space Environment. Front. Physiol. 8, 547 (2017).
Willey, J. S., Lloyd, S. A., Nelson, G. A. & Bateman, T. A. Space Radiation and Bone Loss. Gravit. Space Biol. Bull. 25, 14–21 (2011).
Tian, Y. et al. The Impact of Oxidative Stress on the Bone System in Response to the Space Special Environment. Int. J. Mol. Sci. 18, 2132 (2017).
Carpentieri, A. et al. Rapid Rapamycin-Only Induced Osteogenic Differentiation of Blood-Derived Stem Cells and Their Adhesion to Natural and Artificial Scaffolds. Stem Cells Int. 2017, 1–13 (2017).
Spaas, J. H. et al. Purification and expansion of stem cells from equine peripheral blood, with clinical applications. Vlaams Diergeneeskd. Tijdschr. 80, 129–135 (2011).
Alaimo, G. et al. Blood-Derived Stem Cells (BDSCs) Plasticity: In Vitro Hepatic Differentiation. J. Cell. Physiol. 228, 1249–1254 (2013).
Marfe, G. et al. A new clinical approach: use of blood-derived stem cells (BDSCs) for superficial digital flexor tendon injuries in horses. Life Sci. 90, 825–830 (2012).
Marfe, G. et al. Blood derived stem cells: An ameliorative therapy in veterinary ophthalmology. J. Cell. Physiol. 227, 1250–1256 (2012).
Carpentieri, A. et al. Differentiation of human neuroblastoma cells towards the osteogenic lineage by mTOR inhibitor. Cell Death Dis. 6, e1974 (2016).
Jie, D. & Zhu, B. Rapamycin: A promising agent to treat cancer pain? Int. J. Cancer 139, 1670–1671 (2016).
Abdelaziz, D. M., Stone, L. S. & Komarova, S. V. Osteolysis and pain due to experimental bone metastases are improved by treatment with rapamycin. Breast Cancer Res. Treat. 143, 227–237 (2014).
Maccarrone, M., Guzmán, M., Mackie, K., Doherty, P. & Harkany, T. Programming of neural cells by (endo) cannabinoids: from physiological rules to emerging therapies. Nat. Rev. Neurosci. 15, 786–801 (2014).
Maccarrone, M. et al. Endocannabinoid signaling at the periphery: 50 years after THC. Trends Pharmacol. Sci. 36, 277–296 (2015).
Bari, M. et al. The SERiSM project: preliminary data on human stem cell reprogramming in microgravity. Front. Physiol. Conference Abstract: 39th ISGP Meeting & ESA Life Sciences Meeting (2018).
Delgado-Calle, J., Garmilla, P. & Riancho, J. A. Do epigenetic marks govern bone mass and homeostasis? Curr. genomics 13, 252–263 (2012).
Ghayor, C. & Weber, F. E. Epigenetic Regulation of Bone Remodeling and Its Impacts in Osteoporosis. Int. J. Mol. Sci 17, 1446 (2016).
Kim, K., Shin, Y., Kim, J., Ulmer, T. S. & An, W. H3K27me1 is essential for MMP-9-dependent H3N-terminal tail proteolysis during osteoclastogenesis. Epigenetics Chromatin 11, 23 (2018).
Dhaliwal, A., Pelka, S., Gray, D. S. & Moghe, P. V. Engineering Lineage Potency and Plasticity of Stem Cells using Epigenetic Molecules. Sci. Rep. 8, 16289 (2018).
Berendsen, A. D. & Olsen, B. R. How vascular endothelial growth factor-A (VEGF) regulates differentiation of mesenchymal stem cells. J. Histochem. Cytochem. 62, 103–108 (2014).
Berendsen, A. D. & Olsen, B. R. Regulation of adipogenesis and osteogenesis in mesenchymal stem cells by vascular endothelial growth factor A. J. Intern. Med. 277, 674–680 (2015).
Acampora, D., Di Giovannantonio, L. G. & Simeone, A. Otx2 is an intrinsic determinant of the embryonic stem cell state and is required for transition to a stable epiblast stem cell condition. Development 140, 43–55 (2013).
Harikumar, A. & Meshorer, E. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep. 16, 1609–1619 (2015).
Niakan, K. K. et al. Sox17 promotes differentiation in mouse embryonic stem cells by directly regulating extraembryonic gene expression and indirectly antagonizing self-renewal. Genes Dev. 24, 312–326 (2010).
Song, I. et al. GATA4 negatively regulates osteoblast differentiation by downregulation of Runx2. BMB Rep. 47, 463–468 (2014).
Guo, S. et al. Role of GATA binding protein 4 (GATA4) in the regulation of tooth development via GNAI3. Sci. Rep. 7, 1534 (2017).
Wysokinski, D., Pawlowska, E. & Blasiak, J. RUNX2: A Master Bone Growth Regulator That May Be Involved in the DNA Damage Response. DNA Cell. Biol. 34, 305–315 (2015).
Nakashima, K. et al. The Novel Zinc Finger-Containing Transcription Factor Osterix Is Required for Osteoblast Differentiation and Bone Formation. Cell. 108, 17–29 (2002).
Gersbach, C. A., Le Doux, J. M., Guldberg, R. E. & García, A. J. Inducible regulation of Runx2-stimulated osteogenesis. Gene Ther. 13, 873–882 (2006).
Zhang, X. et al. Runx2 overexpression enhances osteoblastic differentiation and mineralization in adipose-derived stem cells in vitro and in vivo. Calcif. Tissue Int. 79, 169–178 (2006).
Tang, Y. & Weiss, S. J. Snail/Slug-YAP/TAZ complexes cooperatively regulate mesenchymal stem cell function and bone formation. Cell Cycle 16, 399–405 (2017).
Khalid, A. B. et al. GATA4 Directly Regulates Runx2 Expression and Osteoblast. Differentiation. JBMR Plus 2, 81–91 (2018).
Park, S. J. et al. The transcription factor snail regulates osteogenic differentiation by repressing Runx2 expression. Bone 46, 1498–1507 (2010).
Calo, E. & Wysocka, J. Modification of Enhancer Chromatin: What, How, and Why? Mol. Cell 49, 825–837 (2013).
Harikumar, A. & Meshorer, E. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep. 16, 1609–1619 (2015).
Banito, A. et al. Senescence impairs successful reprogramming to pluripotent stem cells. Genes Dev. 23, 2134–2139 (2009).
Onder, T. T. et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 483, 598–602 (2012).
Becker, J. S., Nicetto, D. & Zaret, K. S. H3K9me3-Dependent Heterochromatin: Barrier to Cell Fate Changes. Trends Genet. 32, 29–41 (2015).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and Impediments of the Pluripotency Reprogramming Factors’ Initial Engagement with the Genome. Cell 151, 994–1004 (2012).
Sridharan, R. et al. Proteomic and genomic approaches reveal critical functions of H3K9 methylation and Heterochromatin Protein-1γ in reprogramming to pluripotency. Nat. Cell Biol. 15, 872–882 (2013).
Herranz, M. & Esteller, M. DNA methylation and histone modifications in patients with cancer: potential prognostic and therapeutic targets. Methods Mol. Biol. 361, 25–62 (2007).
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell stem Cell 6, 479–91 (2010).
Meshorer, E. et al. Hyperdynamic Plasticity of Chromatin Proteins in Pluripotent Embryonic Stem Cells. Dev. Cell 10, 105–116 (2006).
Wu, H. et al. Chromatin dynamics regulate mesenchymal stem cell lineage specification and differentiation to osteogenesis. Biochim. Biophys. Acta. Gene Regul. Mech. 1860, 438–449 (2017).
Wang, L. et al. H3K36 trimethylation mediated by SETD2 regulates the fate of bone marrow mesenchymal stem cells. PLoS Biol. 16, e2006522 (2018).
Huang, B., Li, G. & Jiang, X. H. Fate determination in mesenchymal stem cells: a perspective from histone-modifying enzymes. Stem Cell Res. Ther. 6, 35 (2015).
Sinha, K. M., Yasuda, H., Coombes, M. M., Dent, S. Y. & de Crombrugghe, B. Regulation of the osteoblast-specific transcription factor Osterix by NO66, a Jumonji family histone demethylase. EMBO J. 29, 68–79 (2009).
Zhou, Y., Kim, J., Yuan, X. & Braun, T. Epigenetic modifications of stem cells: A paradigm for the control of cardiac progenitor cells. Circ. Res. 109, 1067–1081 (2011).
Acknowledgements
We wish to thank Professor Alessandro Finazzi Agrò (Campus Bio-Medico University of Rome) for critically reading the manuscript, Drs Derek Duflo and Kamber Scott (Space Station Processing Facility Support Team, Kennedy Space Center, Florida) for their valuable support in SERiSM preparation at Kennedy Space Center, and Mrs. Donna Hartley (Occupational Health Facility, Kennedy Space Center, Florida) for her kind help with blood withdrawal. Technical and logistic support by the Kayser Italia S.r.l. team is also gratefully acknowledged. The SERiSM project was made possible by the coordination and financial support of the Italian Space Agency (ASI), under the contract N. 2016-5-U.0 to M.M.
Author information
Authors and Affiliations
Contributions
A.G., M.B. and M.M. conceived the SERiSM project. A.G. and M.M. monitored project progression, data analysis and interpretation. M.G. performed the proteomic and epigenetic experiments and designed the figures, G.R. and G.D. aided in the experiments and statistical analysis, M.B. and B.N. prepared the space cell cultures, and M.Balsamo was responsible for Kayser Italia hardware. S.P., G.V. and G.M. supervised and coordinated the project for ASI. A.G. drafted the manuscript, that was revised and finalized by M.M. and approved by all authors.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gambacurta, A., Merlini, G., Ruggiero, C. et al. Human osteogenic differentiation in Space: proteomic and epigenetic clues to better understand osteoporosis. Sci Rep 9, 8343 (2019). https://doi.org/10.1038/s41598-019-44593-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44593-6
- Springer Nature Limited
This article is cited by
-
Conserved mechanisms of self-renewal and pluripotency in mouse and human ESCs regulated by simulated microgravity using a 3D clinostat
Cell Death Discovery (2024)
-
Chromosomal positioning and epigenetic architecture influence DNA methylation patterns triggered by galactic cosmic radiation
Scientific Reports (2024)
-
Hindlimb unloading, a physiological model of microgravity, modifies the murine bone marrow IgM repertoire in a similar manner as aging but less strongly
Immunity & Ageing (2023)
-
GATA4-activated lncRNA MALAT1 promotes osteogenic differentiation through inhibiting NEDD4-mediated RUNX1 degradation
Cell Death Discovery (2023)
-
Histone methyltransferase SETDB1 promotes osteogenic differentiation in osteoporosis by activating OTX2-mediated BMP-Smad and Wnt/β-catenin pathways
Human Cell (2023)