Abstract
Circular RNA (circRNA) is a type of non-coding RNA known to affect cancer-related micro RNAs and various transcription factors. circRNA has promise as a cancer-related biomarker because its circular structure affords high stability. We found using high-throughput sequencing that seven candidate circRNAs (hsa_circ_0041150, hsa_circ_0025624, hsa_circ_0001020, hsa_circ_0028129, hsa_circ_0008558, hsa_circ_0036683, hsa_circ_0058087) were downregulated in HCC. The expression of these circRNAs was examined by quantitative PCR in 233 sets of HCC and matched background normal liver tissues, and correlations between candidate circRNA expression and prognosis were evaluated. The results of quantitative PCR showed that expression of hsa_circ_0041150, hsa_circ_0001020 and hsa_circ_0008558 was significantly lower in HCC than in background normal liver tissues. Kaplan–Meier analysis revealed that low expression of hsa_circ_0001020, hsa_circ_0036683, and hsa_circ_0058087 was associated with poor recurrence-free (RFS) and overall survival (OS) in HCC. Additionally, multivariate analysis revealed that low hsa_circ_0036683 expression was a significant prognostic factor, independent from other clinicopathological features, for inferior RFS and OS. There was no significant association between the expression of these circRNAs and hepatitis B/C status or cirrhosis. This study therefore identified circRNAs as potential prognostic markers for patients who undergo curative surgery for HCC and highlighted hsa_circ_0036683 as the most useful biomarker.
Similar content being viewed by others
Introduction
Recent advances in genome-wide analytical techniques have revealed that while almost all genomic lesions are transferred into RNA, as little as 2% of these encode proteins1. Thus, there is an abundance of non-coding RNA (ncRNA) in the eukaryotic cell. Numerous studies have uncovered important functional roles for ncRNA in regulating the interplay between DNA, RNA and protein expression2. Many different ncRNAs have been identified to date and they are predominantly categorized according to length3. Long non-coding RNAs (lncRNAs) are ncRNAs that exceed 200 base pairs and they represent a relatively abundant component of the mammalian transcriptome4. While lncRNAs form the biggest group of mammalian ncRNAs, circular RNA (circRNA) is a recently identified subtype of lncRNA that demonstrates greater stability than linear RNA owing to its covalently closed loop structure5. It is believed that circRNA regulates gene expression through various interactions with other RNA types (e.g., as sponging microRNAs) as well as through interactions with RNA-binding proteins. circRNAs may also regulate gene expression directly by influencing transcription and splicing. More recently, some circRNAs have been shown to encode proteins6. There is growing interest in the possible roles that circRNAs may play in the development of human diseases including malignant neoplasms. Recent studies have demonstrated abnormal circRNA expression in several types of malignancies6. However, the relationship between circRNA expression and cancer prognosis requires further study.
Liver cancer is predicted to be the sixth most commonly diagnosed cancer and the fourth leading cause of cancer-related death worldwide7. Hepatocellular carcinoma (HCC) accounts for 75–85% of primary liver cancers and is the second leading cause of cancer-related death in East Asia and sub-Saharan Africa, and the sixth most common cause in Western countries7,8. The main risk factors for HCC include lifestyle factors such as heavy alcohol intake, obesity, smoking, type 2 diabetes, and aflatoxin-contaminated foodstuffs, as well as background liver status including chronic hepatitis B (HBV) or hepatitis C (HCV) viral infection and associated cirrhosis7,9,10. Although there are several recommended treatment options for HCC, surgical resection remains the most effective therapy for prolonging patient survival11. However, because of complexities related to background liver status, the possibility of postoperative recurrence is higher in HCC, when compared with other gastrointestinal cancers. Consequently, the 5-year survival rate for HCC following surgery is approximately 30–57%12,13,14,15,16,17,18. Therefore, there is an urgent need to understand not only the molecular mechanisms of HCC itself, but also the molecular relationship between this disease and underlying background liver status.
In this study, we assessed the expression of circRNA in resected HCC and corresponding background normal liver tissues using a genome-wide approach. We also determined the level of circRNA in resected HCC tissues and paired normal liver tissues from patients who underwent curative HCC surgery. Expression data and clinical data were subsequently analysed to determine whether circRNA has utility as a prognostic marker in patients with HCC.
Results
Identification of a circRNA signature for HCC and background liver tissues using high throughput sequencing
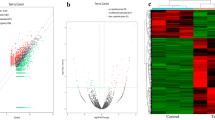
To develop a circRNA signature specific to HCC tumour and background normal liver tissue, we first interrogated high throughput sequencing (HTS)-based circRNA expression profiles for four cases of resected HCC. This genome-wide circRNA analysis identified 15 circRNAs that were differentially expressed between HCC and background normal liver tissue, omitting circRNAs with null expression in any of the four samples. A heatmap of the 15-circRNA signature is shown in Fig. 1, together with hierarchical clustering and principal component analysis. Eight of the 15 circRNAs corresponded to known hsa_circ IDs and we successfully generated primers for seven of these eight using CircPrimer software for the subsequent validation of the HTS data using qPCR19. The IDs for the seven circRNAs were hsa_circ_0041150, hsa_circ_0025624, hsa_circ_0001020, hsa_circ_0028129, hsa_circ_0008558, hsa_circ_0036683, and hsa_circ_0058087. According to the HTS data, these circRNAs were all expressed at a lower level in HCC tissues, when compared with paired background normal liver tissues (absolute log2 fold change > 2.0, p < 0.05, Student’s t-test).
Differential expression of circRNAs between HCC and background normal liver. The figure is prepared using R software (version 3.5.3, https://www.r-project.org/)43 and Gplots (version 3.1.1, https://github.com/talgalili/gplots).
qPCR-based validation of HTS data using HCC patient samples
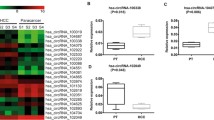
HCC and background normal liver tissues taken from 233 HCC patients who underwent liver resection, were analyzed by qPCR to determine the expression of the seven candidate circRNAs identified in the HTS analysis (hsa_circ_0041150, hsa_circ_0025624, hsa_circ_0001020, hsa_circ_0028129, hsa_circ_0008558, hsa_circ_0036683, hsa_circ_0058087). Expression profiles for these circRNAs in the collected paired samples are shown in Fig. 2. Among these seven circRNAs, the expression of hsa_circ_0041150, hsa_circ_0001020, and hsa_circ_0008558 was significantly lower in HCC tissues than in background normal liver tissues (p < 0.001 for all). These five circRNAs were therefore found to be expressed at a consistently lower level in HCC tissues, when compared with normal background tissues, in both the HTS and qPCR analyses. There was no correlation between the expression of these candidate cirRNAs in normal and tumour samples (Supplementary Fig. 1).
circRNA expression according to background liver status
Expression of the seven candidate circRNAs according to hepatitis virus or liver cirrhosis status is shown in Supplementary Fig. 2. Regarding background normal tissues, hsa_circ_0025624 was lower in HCV-positive patients than in HBV-positive patients (p = 0.047), hsa_circ_0008558 was lower in HCV-positive patients than in HBV-positive or HBV/HCV-negative patients (p = 0.012 and < 0.001, respectively), and hsa_circ_0036683 was lower in HCV-positive patients than in HBV/HCV-negative patients (p = 0.042). Regarding HCC tissues, hsa_circ_0041150 was lower in HBV/HCV-negative patients than in HCV-positive patients (p = 0.020), and hsa_circ_0058087 was higher in HBV/HCV-negative patients than in HBV or HCV-positive patients (p = 0.032 and 0.017, respectively). In HBV-positive patients, hsa_circ_0041150, hsa_circ_0001020, hsa_circ_0028129 and hsa_circ_0008558 were all lower in HCC tissues than in background normal tissues (p < 0.001 for all). In HCV-positive patients, hsa_circ_0041150, hsa_circ_0001020, hsa_circ_0028129, and hsa_circ_0008558 were lower in HCC tissues than in background normal tissues (p < 0.001 for all). In HBV/HCV-negative patients, hsa_circ_0041150, hsa_circ_0001020, hsa_circ_0028129, and hsa_circ_0008558 were all lower in HCC tissues than in background normal tissues (p < 0.001, p < 0.001, p = 0.001 and p < 0.001, respectively).
hsa_circ_0001020, hsa_circ_0008558 and hsa_circ_0036683 expression in background normal tissues was lower in patients with liver cirrhosis than in patients without cirrhosis (p = 0.010, p < 0.001 and p = 0.002, respectively), while expression in HCC tissues was independent of cirrhosis status. hsa_circ_0041150, hsa_circ_0025624, hsa_circ_0001020, hsa_circ_0028129, hsa_circ_0008558, and hsa_circ_0058087 were all lower in HCC tissues than in background normal tissues in patients with liver cirrhosis (p < 0.001, p = 0.005, p < 0.001, p < 0.001, p < 0.001 and p = 0.036, respectively). hsa_circ_0041150 hsa_circ_0001020, hsa_circ_0028129, and hsa_circ_0008558 were lower in HCC tissues than in background normal tissues in patients without liver cirrhosis (p < 0.001 for all).
circRNAs are predictive of HCC patient prognosis
Analysis of recurrence free-survival (RFS) and overall survival (OS) indicated that low expression of hsa_circ_0001020, hsa_circ_0036683, and hsa_circ_0058087 in HCC tissues was associated with a significantly inferior RFS (Fig. 3; median survival time [MST] 22 vs. 40 months, p = 0.008; MST 14 vs. 34, p < 0.001; MST 12 vs. 32, p = 0.006, respectively). Moreover, low expression of hsa_circ_0001020, hsa_circ_0036683, and hsa_circ_0058087 in HCC tissues was also associated with significantly inferior OS (Fig. 3; MST 67 vs. 116; p = 0.018, MST 33 vs. 92; p = 0.006, MST 55 vs. 94; p = 0.040, respectively). When the clinical features of the 233 HCC cases were stratified according to circRNA expression, hsa_circ_0001020 was found to be significantly associated with serum alpha fetoprotein (AFP) levels (p = 0.003), hsa_circ_0036683 (p = 0.006) and hsa_circ_0001020 (p = 0.004) with tumour size, hsa_circ_0001020 (p = 0.008) with portal vein or hepatic vein invasion, and hsa_circ_0001020 (p = 0.010) with pathological stage (Table 1).
Recurrence free and overall survival based on the expression of hsa_circ_0001020, hsa_circ_0036683 and hsa_circ_0058087. (a) Recurrence free survival. (b) Overall survival. The figure is prepared using R software (version 3.5.3, https://www.r-project.org/)43.
Univariate and multivariate analysis of prognostic factors associated with RFS and OS in HCC
Univariate analysis revealed that pathological stage (≥ III), hsa_circ_0001020 expression (low), hsa_circ_0036683 expression (low), and hsa_circ_0058087 expression (low) were all significant predictors of inferior RFS. When multivariate analysis was performed hsa_circ_0036683 expression (low) and hsa_circ_0058087 expression (low) were identified as independent predictive factors for inferior RFS (Table 2).
Regarding OS, univariate analysis revealed that AFP (≥ 20 ng/dl), pathological stage (≥ III), hsa_circ_0001020 expression (low), hsa_circ_0036683 expression (low), and hsa_circ_0058087 expression (low) were all significant predictors of worse OS. When multivariate analysis was performed on these predictors, AFP (≥ 20 ng/dl), and hsa_circ_0036683 expression (low) were identified as independent predictive factors for worse OS (Table 3).
Synergistic effect of hsa_circ_0036683 on alpha fetoprotein, as a biomarker
Supplementary Fig. 3 shows the results of ROC analysis of RFS and OS 2 years after surgery. Regarding RFS, the combination of hsa_circ_0036683 and AFP had a higher AUC (0.63) than hsa_circ_0036683 (0.57) or AFP alone (0.59). In addition, for OS, the combination of hsa_circ_0036683 and AFP had a higher AUC (0.69) than hsa_circ_0036683 (0.62) or AFP alone (0.62).
Discussion
RNA deep sequencing technology has revealed roles for ncRNAs in the progression of several diseases, including hepatitis, cirrhosis and liver cancer20. circRNA is a class of RNA that exhibits a unique single-stranded circular structure that is formed when the upstream 3′ splice site and downstream 5′ splice site join to form a covalently closed and stable continuous loop. circRNAs have been implicated in many important biological processes21. In this study, a genome-wide approach employing HTS was used to evaluate circRNA expression in HCC. Seven candidate circRNAs identified in the initial HTS analysis were subsequently examined in 233 sets of HCC samples to evaluate the relationship between expression and prognosis. The HTS data indicated that these seven circRNAs were all expressed at a lower level in HCC tissues, when compared with background normal tissues. Subsequent qPCR analysis using matched patient samples showed that hsa_circ_0041150, hsa_circ_0025624, hsa_circ_0001020, hsa_circ_0008558, and hsa_circ_0036683 were all expressed at a significantly lower level in HCC tissues, when compared with background normal tissues. Regarding prognosis, hsa_circ_0001020, hsa_circ_0036683, and hsa_circ_0058087 were all associated with worse RFS and OS. Moreover, multivariate analysis revealed that low hsa_circ_0036683 expression was an independent prognostic factor for worse RFS and OS.
Several circRNAs have been reported to be involved in HCC with some demonstrating elevated or reduced expression, suggesting that they may have utility as biomarkers in this disease22,23. It has been reported that circRNAs may contribute to the pathology of HCC through their role as micro RNA-sponges, protein-sponges, micro RNA transporters and as regulators of parental gene expression22,23,24. The altered expression of the seven circRNAs examined in this study has not previously been reported for HCC, although changes in hsa_circ_0041150, hsa_circ_0028129, and hsa_circ_0008558 expression have been reported in other carcinomas. An analysis of 20 sets of pancreatic ductal adenocarcinoma samples using microarray and qPCR revealed that hsa_circ_0041150 is downregulated in this disease25. Microarray data indicate that hsa_circ_0028129 is downregulated in colorectal cancer26, and that hsa_circ_0008558 is upregulated in both bladder cancer and oral mucosal melanoma27,28. Our study therefore demonstrates that these three circRNAs are implicated in the regulation of HCC in addition to these other carcinoma types.
Previous studies have examined the relationship between circRNA expression and background liver status29. Zhou et al. showed that expression profiles for hepatic circRNAs were significantly different between chronic hepatitis B (CHB) and normal hepatic tissues, with 99 dysregulated circRNAs identified in total29. Ji et al., reported the involvement of circ_0070963 in liver fibrosis30. Unexpectedly, this study did not reveal any significant association between circRNA expression and HBV, HCV, or cirrhosis status.
In addition to our study, others have also examined the relationship between circRNA expression and patient prognosis in HCC using qPCR. In a study of 70–200 HCC patients, high expression of circ_0021093, circ_0008450, circ_0128298, circ_0003998 and hsa_circ_0006916 was associated with unfavorable prognosis, as determined using Kaplan–Meier and multivariate analysis31,32,33,34,35. Low circ_0000567 and circ-ITCH expression was predictive of poor prognosis in a study of 134–288 HCC patients, as determined using Kaplan–Meier analysis36,37. In our study of 233 HCC patients we now demonstrate, using both Kaplan–Meier and multivariate analysis, that decreased expression of hsa_circ_0001020, hsa_circ_0036683, and hsa_circ_0058087 is predictive of poor disease prognosis.
Although we have identified several circRNAs as useful biomarkers and prognostic predictors for HCC, there are some limitations to this study. Firstly, the study cohort consisted of individuals from a single institution. Secondly, we did not investigate the underlying mechanisms of altered circRNA expression and how these changes may impact on prognosis by identifying potential gene or microRNA targets. This study simply evaluated circRNA expression in HCC and normal background liver tissues to examine its utility as a biomarker and prognostic factor in clinical practice.
In conclusion, the circRNAs hsa_circ_0041150, hsa_circ_0025624, hsa_circ_0001020, hsa_circ_0028129, hsa_circ_0008558, hsa_circ_0036683, and hsa_circ_0058087 were all associated with prognosis in HCC. In particular, hsa_circ_0036683, which was found to be an independent and significant prognostic factor for HCC, has potential utility as a candidate biomarker for this disease. Moreover, we believe that further functional analysis of these circRNAs may identify novel therapeutic targets for the treatment of this disease.
Methods
Patients and samples
A total of 233 frozen tumour specimens and paired para-tumour normal background liver tissues were collected from patients with HCC who underwent surgery at Nagoya University Hospital between January 1998 and January 2012. All fresh tissues were immediately frozen in liquid nitrogen and stored at − 80 °C until required. Patient characteristics are summarized in Table 1. After surgery, blood examinations, ultrasonography (US), and computed tomography (CT) were used to monitor all patients. In cases of possible recurrence, additional examination including contrast US, positron emission tomography-CT, and/or angiography were performed for confirmation. After a median follow-up duration of 66.6 months (range 0.3–206.2 months), 182 (78.1%) recurrences and 135 (57.9%) deaths occurred in the 233 patients. The median follow-up duration of all cases was 66.6 months (range 0.3–206.2 months). This study and all procedures were approved by the Institutional Review Board at Nagoya University and all patients provided written informed consent. All clinical investigations were conducted in accordance with the principles of the Declaration of Helsinki38.
RNA isolation and genome-wide high-throughput sequencing
Total RNA was extracted from tissue samples using the Qiagen miRNeasy mini-kit (Qiagen, Hilden, Germany). Approximately 10 μg total RNA was then subject to ribosomal RNA depletion using the Ribo-Zero Gold Kit, as per the manufacturer’s instructions (Illumina, San Diego, CA, USA). The RNA fragments were then reverse-transcribed to create the final cDNA library using the ncRNA-Seq sample preparation kit (Illumina) according to the manufacturer’s recommended protocol. The prepared libraries were then sequenced on an Illumina Hiseq X ten platform (Illumina) and 2 × 100-bp paired-end reads (PE100) were generated according to the standard Illumina protocol. All procedures for circRNA sequencing were performed by BGI Genomics Services (Beijing, China). Sequencing reads containing low-quality, adaptor-polluted and a high content of unknown base (N) reads were removed using QCleaner v4.0.1 (Amelieff, Tokyo, Japan) before downstream analysis.
Annotation of human circRNAs
CIRI39 was used to predict circRNA in this project and to integrate results based on the start and end position of circRNA. Burrows-Wheeler Aligner software (BWA 0.7.17, http://bio-bwa.sourceforge.net/)40 was used to align discovered reads to the hg38 reference genome. circRNAs that had the same start and end position within 10 bases were assigned to the same class. If a circRNA was recorded in the circBase (http://www.circbase.org/cgi-bin/downloads.cgi), the corresponding ID code was provided. circRNA was considered novel if it did not overlap with any registered circRNA in circBase.
Estimation of tumour-specific circRNA and differential expression analysis
circRNAs detected in only HCC samples were considered to be tumour-specific. Tumour-specific circRNAs with a number of reads ≥ 6 were used in down-stream analyses. Differential expression analysis of circRNAs between HCC and background normal liver tissues was conducted using the limma package (3.38.3, https://bioconductor.org/packages/release/bioc/html/limma.html)41. The read count for circRNA was logarithmically transformed (log2 [count + 1]) and established linear models were assessed using the empirical Bayes method. The acquired p-value was adjusted by the Benjamini–Hochberg method.
Validation of circRNA expression by qPCR
Total RNA was extracted from tissue samples using the Qiagen miRNeasy mini-kit and then converted to complementary DNA using M-MLV Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA) for subsequent use in qPCR assays. PCR was performed using SYBR Premix Ex Taq II (Takara Clontech, Kyoto, Japan) under the following conditions: denaturing at 95 °C for 10 s, 40 cycles of denaturing at 95 °C for 5 s, and annealing/extension at 60 °C for 30 s. The SYBR Green signal was detected in real-time using a StepOne Plus real-time PCR System (Life Technologies, Carlsbad, CA, USA). The relative quantification method was used where the expression level of each gene in a sample was determined after normalization to the housekeeping control glyceraldehyde-3-phosphate dehydrogenase (GAPDH). Relative gene expression levels were determined using the comparative threshold cycle (2-ΔΔCT) method. To design PCR primers for circRNA targets, templates were generated using CircPrimer (Ver.1.2, https://www.bioinf.com.cn/)19 and divergent primers were designed with primer3 (ver.0.4.0, https://bioinfo.ut.ee/primer3-0.4.0/)42. All qPCR experiments were performed in duplicate, including the template-omitted negative controls.
Statistical analysis
Continuous variables were expressed as the median and range, and gene expression comparisons were performed using the Mann–Whitney U, Wilcoxon singed rank, Kruskal–Wallis or Steel–Dwass tests. Categorical variables were compared using Fisher’s exact tests. The OS and RFS rate at each follow-up time point was estimated using the Kaplan–Meier method and comparisons were made using a log-rank test. The Cox proportional hazard model was used to perform univariate and multivariate analysis for RFS and OS. All statistical analyses were performed using R software (version 3.5.3, https://www.r-project.org/) 43. Statistical significance was set at p < 0.05, using two-tailed tests.
References
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563. https://doi.org/10.1126/science.1112014 (2005).
Mattick, J. S. & Makunin, I. V. Non-coding RNA. Hum. Mol. Genet. 15(Spec No 1), R17-29. https://doi.org/10.1093/hmg/ddl046 (2006).
Hombach, S. & Kretz, M. Non-coding RNAs: Classification, biology and functioning. Adv. Exp. Med. Biol. 937, 3–17. https://doi.org/10.1007/978-3-319-42059-2_1 (2016).
Mercer, T. R., Dinger, M. E. & Mattick, J. S. Long non-coding RNAs: insights into functions. Nat. Rev. Genet. 10, 155–159. https://doi.org/10.1038/nrg2521 (2009).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157. https://doi.org/10.1261/rna.035667.112 (2013).
Hsiao, K. Y., Sun, H. S. & Tsai, S. J. Circular RNA—New member of noncoding RNA with novel functions. Exp. Biol. Med. (Maywood) 242, 1136–1141. https://doi.org/10.1177/1535370217708978 (2017).
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424. https://doi.org/10.3322/caac.21492 (2018).
Choo, S. P., Tan, W. L., Goh, B. K. P., Tai, W. M. & Zhu, A. X. Comparison of hepatocellular carcinoma in Eastern versus Western populations. Cancer 122, 3430–3446. https://doi.org/10.1002/cncr.30237 (2016).
Rawla, P., Sunkara, T., Muralidharan, P. & Raj, J. P. Update in global trends and aetiology of hepatocellular carcinoma. Contemp. Oncol. (Pozn) 22, 141–150. https://doi.org/10.5114/wo.2018.78941 (2018).
London, W. T., Petrick, J. L. & McGlynn, K. A. Liver cancer. In Cancer Epidemiology and Prevention 4th edn (ed. Thun, M. J.) 635–660 (Oxford University Press, 2018).
Rahbari, N. N. et al. Hepatocellular carcinoma: Current management and perspectives for the future. Ann. Surg. 253, 453–469. https://doi.org/10.1097/SLA.0b013e31820d944f (2011).
Sun, H. C. et al. Postoperative interferon alpha treatment postponed recurrence and improved overall survival in patients after curative resection of HBV-related hepatocellular carcinoma: A randomized clinical trial. J. Cancer Res. Clin. Oncol. 132, 458–465. https://doi.org/10.1007/s00432-006-0091-y (2006).
Kubo, S. et al. Randomized clinical trial of long-term outcome after resection of hepatitis C virus-related hepatocellular carcinoma by postoperative interferon therapy. Br. J. Surg. 89, 418–422. https://doi.org/10.1046/j.0007-1323.2001.02054.x (2002).
Wang, J. H. et al. The efficacy of treatment schedules according to Barcelona Clinic Liver Cancer staging for hepatocellular carcinoma—Survival analysis of 3892 patients. Eur. J. Cancer 44, 1000–1006. https://doi.org/10.1016/j.ejca.2008.02.018 (2008).
Lin, C. T. et al. Comparing hepatic resection and transarterial chemoembolization for Barcelona Clinic Liver Cancer (BCLC) stage B hepatocellular carcinoma: Change for treatment of choice?. World J. Surg. 34, 2155–2161. https://doi.org/10.1007/s00268-010-0598-x (2010).
Chang, W. T. et al. Hepatic resection can provide long-term survival of patients with non-early-stage hepatocellular carcinoma: extending the indication for resection?. Surgery 152, 809–820. https://doi.org/10.1016/j.surg.2012.03.024 (2012).
Torzilli, G. et al. A snapshot of the effective indications and results of surgery for hepatocellular carcinoma in tertiary referral centers: is it adherent to the EASL/AASLD recommendations? An observational study of the HCC East-West study group. Ann. Surg. 257, 929–937. https://doi.org/10.1097/SLA.0b013e31828329b8 (2013).
Zhong, J. H. et al. Hepatic resection associated with good survival for selected patients with intermediate and advanced-stage hepatocellular carcinoma. Ann. Surg. 260, 329–340. https://doi.org/10.1097/SLA.0000000000000236 (2014).
Zhong, S., Wang, J., Zhang, Q., Xu, H. & Feng, J. CircPrimer: A software for annotating circRNAs and determining the specificity of circRNA primers. BMC Bioinform. 19, 292. https://doi.org/10.1186/s12859-018-2304-1 (2018).
Roy, S., Trautwein, C., Luedde, T. & Roderburg, C. A General overview on non-coding RNA-based diagnostic and therapeutic approaches for liver diseases. Front. Pharmacol. 9, 805. https://doi.org/10.3389/fphar.2018.00805 (2018).
Chen, L. L. & Yang, L. Regulation of circRNA biogenesis. RNA Biol. 12, 381–388. https://doi.org/10.1080/15476286.2015.1020271 (2015).
Hu, J. et al. Progress and prospects of circular RNAs in Hepatocellular carcinoma: Novel insights into their function. J. Cell Physiol. 233, 4408–4422. https://doi.org/10.1002/jcp.26154 (2018).
Fu, L., Jiang, Z., Li, T., Hu, Y. & Guo, J. Circular RNAs in hepatocellular carcinoma: Functions and implications. Cancer Med. https://doi.org/10.1002/cam4.1574 (2018).
Yao, T., Chen, Q., Fu, L. & Guo, J. Circular RNAs: Biogenesis, properties, roles, and their relationships with liver diseases. Hepatol. Res. 47, 497–504. https://doi.org/10.1111/hepr.12871 (2017).
Li, H. et al. Circular RNA expression profile of pancreatic ductal adenocarcinoma revealed by microarray. Cell Physiol. Biochem. 40, 1334–1344. https://doi.org/10.1159/000453186 (2016).
Tian, J. et al. CircRNA hsa_circ_0004585 as a potential biomarker for colorectal cancer. Cancer Manag. Res. 11, 5413–5423. https://doi.org/10.2147/CMAR.S199436 (2019).
Zhong, Z., Lv, M. & Chen, J. Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci. Rep. 6, 30919. https://doi.org/10.1038/srep30919 (2016).
Ju, H. et al. Altered expression pattern of circular RNAs in metastatic oral mucosal melanoma. Am. J. Cancer Res. 8, 1788–1800 (2018).
Zhou, T. C. et al. Differential expression profile of hepatic circular RNAs in chronic hepatitis B. J. Viral. Hepat. 25, 1341–1351. https://doi.org/10.1111/jvh.12944 (2018).
Ji, D. et al. Hsa_circ_0070963 inhibits liver fibrosis via regulation of miR-223-3p and LEMD3. Aging (Albany NY) 12, 1643–1655. https://doi.org/10.18632/aging.102705 (2020).
Liu, L. et al. Overexpression of circ_0021093 circular RNA forecasts an unfavorable prognosis and facilitates cell progression by targeting the miR-766-3p/MTA3 pathway in hepatocellular carcinoma. Gene 714, 143992. https://doi.org/10.1016/j.gene.2019.143992 (2019).
Zhang, J., Chang, Y., Xu, L. & Qin, L. Elevated expression of circular RNA circ_0008450 predicts dismal prognosis in hepatocellular carcinoma and regulates cell proliferation, apoptosis, and invasion via sponging miR-548p. J. Cell Biochem. 120, 9487–9494. https://doi.org/10.1002/jcb.28224 (2019).
Chen, D., Zhang, C., Lin, J., Song, X. & Wang, H. Screening differential circular RNA expression profiles reveal that hsa_circ_0128298 is a biomarker in the diagnosis and prognosis of hepatocellular carcinoma. Cancer Manag. Res. 10, 1275–1283. https://doi.org/10.2147/CMAR.S166740 (2018).
Qiao, G. L. et al. Hsa_circ_0003998 may be used as a new biomarker for the diagnosis and prognosis of hepatocellular carcinoma. Onco Targets Ther. 12, 5849–5860. https://doi.org/10.2147/OTT.S210363 (2019).
Zhao, M., Dong, G., Meng, Q., Lin, S. & Li, X. Circ-HOMER1 enhances the inhibition of miR-1322 on CXCL6 to regulate the growth and aggressiveness of hepatocellular carcinoma cells. J. Cell Biochem. https://doi.org/10.1002/jcb.29672 (2020).
Xu, L. et al. CircSETD3 (Hsa_circ_0000567) acts as a sponge for microRNA-421 inhibiting hepatocellular carcinoma growth. J. Exp. Clin. Cancer Res. 38, 98. https://doi.org/10.1186/s13046-019-1041-2 (2019).
Guo, W. et al. Polymorphisms and expression pattern of circular RNA circ-ITCH contributes to the carcinogenesis of hepatocellular carcinoma. Oncotarget 8, 48169–48177. https://doi.org/10.18632/oncotarget.18327 (2017).
Association, W. M. World Medical Association Declaration of Helsinki: Ethical principles for medical research involving human subjects. JAMA 310, 2191–2194. https://doi.org/10.1001/jama.2013.281053 (2013).
Gao, Y., Wang, J. & Zhao, F. CIRI: An efficient and unbiased algorithm for de novo circular RNA identification. Genome Biol. 16, 4. https://doi.org/10.1186/s13059-014-0571-3 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47. https://doi.org/10.1093/nar/gkv007 (2015).
Untergasser, A. et al. Primer3—New capabilities and interfaces. Nucleic Acids Res. 40, e115. https://doi.org/10.1093/nar/gks596 (2012).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Acknowledgements
We thank M. Kubokawa (Amelieff) for bioinformatics support. This work was partly supported by a Japan Society for the Promotion of Science, KAKENHI Grant-in-Aid for Scientific Research (C) number 18H0617 awarded to Hideki Takami. We thank James Monypenny, PhD, from Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
Conception and design: Y.S., S.Y., F.S.; financial support: Y.K., H.T.; administrative support: Y.K., S.Y.; provision of study materials and patients: Y.S., S.Y., F.S., K.K., N.T., Y.S., Y.I., H.T., M.H., M.K., C.T., G.N., M.K.; collection and assembly of data: Y.S., S.Y., F.S., K.K., N.T., Y.S., Y.I., H.T., M.H.; qPCR data analysis and interpretation: Y.S., F.S.; manuscript writing: Y.S., S.Y., F.S., Y.K.; final approval of manuscript: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sunagawa, Y., Yamada, S., Sonohara, F. et al. Genome-wide identification and characterization of circular RNA in resected hepatocellular carcinoma and background liver tissue. Sci Rep 11, 6016 (2021). https://doi.org/10.1038/s41598-021-85237-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-85237-y
- Springer Nature Limited