Abstract
The capsular polysaccharide (CPS) of Streptococcus suis defines various serotypes based on its composition and structure. Though serotype switching has been suggested to occur between S. suis strains, its impact on pathogenicity and virulence remains unknown. Herein, we experimentally generated S. suis serotype-switched mutants from a serotype 2 strain that express the serotype 3, 4, 7, 8, 9, or 14 CPS. The effects of serotype switching were then investigated with regards to classical properties conferred by presence of the serotype 2 CPS, including adhesion to/invasion of epithelial cells, resistance to phagocytosis by macrophages, killing by whole blood, dendritic cell-derived pro-inflammatory mediator production and virulence using mouse and porcine infection models. Results demonstrated that these properties on host cell interactions were differentially modulated depending on the switched serotypes, although some different mutations other than loci of CPS-related genes were found in each the serotype-switched mutant. Among the serotype-switched mutants, the mutant expressing the serotype 8 CPS was hyper-virulent, whereas mutants expressing the serotype 3 or 4 CPSs had reduced virulence. By contrast, switching to serotype 7, 9, or 14 CPSs had little to no effect. These findings suggest that serotype switching can drastically alter S. suis virulence and host cell interactions.
Similar content being viewed by others
Introduction
Streptococcus suis is an important porcine pathogen and zoonotic agent causing septicemia, meningitis and many other diseases1,2,3,4. This bacterium has evolutionarily adapted to pigs, with nearly 100% of carriage rate in the upper respiratory tract4,5. S. suis strains are serotyped based on structural differences in the capsular polysaccharide (CPS)2,4. So far, twenty-nine serotypes and twenty-seven additional novel CPS synthesis (cps) loci (NCL) were reported6,7,8,9. Among these serotypes, serotype 2 is responsible for the majority of human clinical cases and is the most frequently isolated from diseased pigs2. Serotypes 1/2, 3, 4, 7, 8, 9, and 14 are also frequently isolated from diseased pigs, although their distributions differ depending on the geographic location2. Multilocus sequence typing (MLST) for S. suis has demonstrated genetic diversity within this species, with more than 1,000 sequence types, and several clonal complexes (CCs) potentially associated with diseases in humans and pigs2,6. Accumulated serotyping and MLST data indicate the presence of different CCs in the population of serotype 2 strains (CC1, CC20, CC25, CC28, and CC104), and different serotypes in the respective CCs (e.g., CC1 include strains of serotypes 1/2, 1, 2, 8, 9, and 14 strains) [pubMLST: http://pubmlst.org/ssuis/]. Taken together, this suggests that serotype switching may occur between S. suis serotype 2 and different serotype isolates.
The S. suis CPS is produced by the repetition of a defined oligosaccharide unit formed by a unique arrangement of various sugars10. Indeed, unique CPS structures of serotypes 1, 2, 3, 7, 8, 9, 14, 18, and 1/2 have been previously determined11,12,13,14,15,16 (Supplementary Fig. S1). Furthermore, previous studies have shown that more than 10 genes related to S. suis CPS synthesis are clustered on a genomic locus7,8,9,10. Alongside, the cps gene clusters of serotypes 1 and 14 and serotypes 2 and 1/2 are almost identical10, with their CPS structure differing by the substitution of only a galactose (Gal) for a N-acetylgalactosamine (GalNAc)13 due to a single nucleotide polymorphism in the glycosyltransferase cpsK gene17. Except for these four serotypes, gene repertoires in the cps gene clusters greatly differ between serotypes7,8,9,10, indicating that up-take of genomic DNA of different serotypes and replacement of cps gene cluster by homologous recombination, using flanking sequences of the clusters, is usually required for serotype switching. In S. suis, some strains are naturally transformable, with the competent state induced by competence gene products18,19. Although serotype switching in S. suis has not yet been demonstrated, these findings suggest that replacement of the cps gene clusters may occur in strains in the competent state through up-take of genomic DNA of the other serotype strains from the environment.
Importantly, the serotype 2 CPS has been shown to play critical roles in protection against phagocytosis by innate immune cells and masking of bacterial surface proteins involved in host cell activation20. In addition, several studies have demonstrated non-virulence of isogenic non-encapsulated serotype 2 mutants in murine and porcine models of infection20. However, very little information is available regarding the CPS of other S. suis serotypes and is restricted to two studies on serotypes 9 and 1420,21. Furthermore, comparing the virulence of strains from different serotypes is impossible due to the high genotypic variation between strains. Accordingly, it remains unclear whether S. suis serotype switching (i.e., differences in CPS structure) can affect host cell interactions and strain virulence, even though serotype switching may occur among S. suis strains.
In the present study, serotype-switched S. suis mutants were experimentally generated to investigate the impacts of CPS type on the host cell interactions and virulence in vivo. The mutants were switched from serotype 2, which is the most important in this species, to serotypes 3, 4, 7, 8, 9, and 14, which are frequently isolated from diseased pigs and found in several CCs with serotype 2 human isolates (CC1, CC20, CC25, CC28, and CC104). Generated mutants have allowed us to study the modulation of the pathogenesis of S. suis caused by serotype switching. Preliminary information was discussed in a recent review22, although no data associated with the findings have been provided so far.
Results
Generated serotype-switched S. suis mutants contain few mutations other than the cps locus
Six different serotype-switched mutants (SS2to3, SS2to4, SS2to7, SS2to8, SS2to9, and SS2to14) and non-encapsulated mutant ΔCPS2, from which the cps locus was deleted, were generated from the serotype 2 strain P1/7 (hereafter SS2) (Table 1, generated as illustrated in Supplementary Figs. S2 and S3). Serotype-switched mutants were confirmed to belong to the correct serotype using classical serological techniques23.
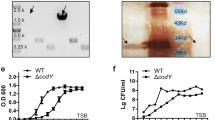
Serotype switching had little effect on bacterial growth in vitro (Supplementary Fig. S4). Well-encapsulation of the serotype-switched mutants was confirmed by surface hydrophobicity and transmission electron microscopy (TEM) (Fig. 1A,B). Since the CPS repeating unit composition and structure for serotypes 2, 3, 7, 8, 9, and 14 have been previously determined12,14,15,16 (Supplementary Fig. S1), CPS of the mutants SS2to3, SS2to7, SS2to8, SS2to9, and SS2to14 were purified to be analyzed by spectroscopy. Purified CPS yields of the mutants were comparable to those previously reported12,14,15,16 (Supplementary Table S1). Nuclear magnetic resonance (NMR) analyses confirmed the serotype identity for the serotype-switched mutants, except for SS2to9 (Supplementary Fig. S5)12,14,15,16. The CPS of SS2to9 slightly differed from that of serotype 9 strain 1,273,590 (used for CPS structure determination14) in that SS2to9 possessed a glucose instead of a galactose side chain (Supplementary Fig. S6a), suggesting that the donor strain and SS2to9 may be classified as a serotype 9 variant, which reacts with anti-serotype 9 serum (see Supplementary Notes for more detail). Taken together, these results confirm that the constructed serotype-switched mutants functionally possess and express the CPS of the donor serotype.
Effect of serotype switching on S. suis CPS expression. (a) Hydrophobicity of the different S. suis strains/mutants. Very low surface hydrophobicity is indicative of high encapsulation, which is demonstrated in the previous study27. Data are expressed as mean ± standard error of the mean (SEM) (n = 3). An asterisk denotes a significant difference with SS2 by Mann–Whitney rank sum test (p < 0.05). (b) Transmission electron micrographs showing CPS expression of the different S. suis strains/mutants. Scale bars = 0.5 µm.
To investigate potential mutations in the genomes of the serotype-switched mutants occurred following the transformation of whole genomic DNA, draft genome sequences of the mutants were compared with those of SS2 and the donors. The mutants had mutations in several genes besides the cps genes, which differed between mutants (Fig. 2, Supplementary Fig. S7, and Supplementary Table S2; see Supplementary Notes for more detail). However, no genes other than cps genes were gained in the genomes of the different mutants. Comparison of the genomes of mutants with those of the corresponding donor strains revealed that the regions of the mutants that were different from the recipient were highly similar to the corresponding region of the donor (> 99.7% of nucleotide identity) (Supplementary Table S3). Although it remains unclear whether these mutations might affect host–pathogen interactions and virulence, nonsense and frameshift mutations in genes, including reported virulence-associated genes20, did not occur (Supplementary Table S2). In addition, no mutations were found in reported small RNAs28. It should be noted that average nucleotide identity (ANI) between the recipient (strain SS2) and each the mutants was ≥ 99.9% and the alignment coverage was ≥ 97.8% (including the replaced cps locus), whereas ANI between the recipient and each the donor genomes was < 98.0% (with < 92.8% of the alignment coverage), except for the donor of SS2to14 (99.9% of ANI with 96.6% of the alignment coverage) (Supplementary Table S4). These data indicate that the mutants constructed in this study have almost identical genetic background to SS2 compared to the heterogenous genetic background of the different serotype strains, enabling more strict evaluation of the CPS effect hereafter.
Mutations present in the generated S. suis serotype-switched mutants. Each of the schematic representations illustrates the analysis data using Geneious Prime mapping of the draft genome sequence of each mutant (upper part) on the publicly available completed genome sequence of serotype 2 (accession no. AM946016) and the sequence alignment between two genomes (lower part). All gaps between the contigs of each mutant were due to multi-copy genes, such as rRNA genes, tRNA genes and IS elements, or repeated regions within genes. Gaps of the repeated regions within genes were found in the genes corresponding to the SS2 locus tags SSU0496, SSU1127, SSU1171, and SSU1172. Detailed data on mutated genes can be found in Supplementary Table S2. Below the bottom panel are displayed the descriptions for each color of the different drawings.
Switching from serotype 2 of S. suis can modulate host cell interactions
The serotype 2 CPS has been described to mask surface adhesins involved in the initial interactions with host cells, including adhesion to and invasion of epithelial cells21,29, to resist phagocytosis by macrophages and bactericidal killing by blood leukocytes to persist in the bloodstream and cause systemic dissemination20, and to mask subcapsular immunostimulatory components to interfere pro-inflammatory mediator production by dendritic cells (DCs)30,31.
First, using newborn pig trachea (NPTr) cells, the adhesion and invasion capacities were evaluated between SS2 and the mutants. While SS2, SS2to3, SS2to4, SS2to9, and SS2to14 similarly adhered to NPTr cells at 2 h, adhesion of SS2to7 and SS2to8 was significantly greater (P < 0.05), similar to that of ΔCPS2 used as a positive control (Fig. 3a). Unlike adhesion results, invasion of the different mutants was similar to that of SS2, with little invasion of NPTr cells overall, although ΔCPS2 showed high levels of invasion, as expected (Fig. 3b).
Impact of serotype switching on S. suis adhesion to and invasion of porcine tracheal epithelial cells, resistance to phagocytosis by macrophages, whole blood bacterial killing, and pro-inflammatory mediator production by dendritic cells. Adhesion (a) and invasion (b) of the different S. suis strains and mutants to NPTr porcine tracheal epithelial cells after 2 h of incubation. (c) Internalization of the different S. suis strains and mutants by J774A.1 murine macrophages after 2 h of incubation. (d) Killing of the different S. suis strains and mutants by murine whole blood after 4 h of incubation. (e) Growth capacity of the different S. suis strains and mutants in porcine whole blood after 4 h of incubation. (f) Pro-inflammatory mediator production by DCs at 16 h following infection with the different S. suis strains and mutants as measured by ELISA. Production of tumor necrosis factor (TNF), interleukin (IL)-6, IL-12p70, C-C motif chemokine ligand (CCL) 5, and C-X-C motif chemokine ligand (CXCL) 1, and CXCL9. C-denotes cells in medium alone. All the data represent the mean ± SEM (n = 4). An asterisk denotes a significant difference with SS2 by Mann–Whitney rank sum test (e) (p < 0.05).
Next, macrophage phagocytosis resistance was evaluated using the J774A.1 murine macrophage cell line. As expected, SS2 and ΔCPS2 were poorly and highly internalized by macrophages, respectively (Fig. 3c). No differences were observed in the internalization between SS2 and the serotype-switched mutants after 1 h incubation (data not shown); however, switching to serotype 4, 7 or 8 significantly increased phagocytosis after 2 h incubation (P < 0.05) (Fig. 3c). However, it should be noted that this increase was of approximately one log-fold, which is, though significant, relatively minor compared to the non-encapsulated mutant (4 log-fold increase).
The capacity to resist the bactericidal effect of leukocytes was then evaluated using murine and porcine whole blood. SS2 was completely resistant to killing by murine blood in contrast to ΔCPS2, which was efficiently killed (60% of killing) (Fig. 3d). While SS2to7, SS2to8, SS2to9, and SS2to14 were also resistant to killing by murine whole blood, SS2to3 and SS2to4 were significantly more killed, with 20% and 30% of killing, respectively (P < 0.05) (Fig. 3d). Using a porcine blood system, SS2 was not only able to persist, but also to some extent multiply, whereas ΔCPS2 was markedly cleared (P < 0.05) (Fig. 3e). Comparable to SS2, SS2to8 could significantly multiply, whereas all other mutants were cleared at different degrees (Fig. 3e). As with mouse blood, SS2to3 and SS2to4 showed the greatest impairment in their capacity to survive in porcine blood (Fig. 3e). It should be noted, however, that levels of cross-reactive antibodies against the different strains might affect the results observed with the swine blood and thus can be considered a confounding factor, although this fact also mimics the real situation in the field.
Lastly, the interactions with DCs were evaluated. Absence of CPS significantly increased production of all mediators tested (P < 0.05), with the exception of CCL2 (Fig. 3f), as previously reported21,29. SS2to3, SS2to7, SS2to9, or SS2to14, along with SS2, did not modulate pro-inflammatory mediator production (Fig. 3f). However, stimulation with SS2to8 significantly increased production of TNF, IL-6, IL-12p70, CCL5, CXCL1, and CXCL9, compared to SS2 (P < 0.05) (Fig. 3f). By contrast, SS2to4 induced significantly lower levels of TNF, IL-6, IL-12p70, and CXCL9 than SS2 (P < 0.05), but CCL5 or CXCL1. CCL2 production was not modulated regardless of the CPS type (Fig. 3f).
Serotype switching can differentially modulate S. suis virulence in a mouse model of systemic infection
The impact of switching from serotype 2 on S. suis virulence was evaluated using a well-established C57BL/6 mouse infection model for S. suis serotype 2 virulence studies32. Following intraperitoneal inoculation of SS2, 60% of mice died after developing clinical signs of systemic infection (Fig. 4a). By contrast, none of the ΔCPS2-inoculated mice died, presenting no or very mild clinical signs the first 24 h only (Fig. 4a). No significant differences in mortality were observed between SS2 and SS2to3, SS2to7, SS2to9 or SS2to14 (Fig. 4a). However, clinical signs of infection caused by SS2to3 were generally less severe than those by SS2. Unexpectedly, inoculation of SS2to8 significantly increased mouse mortality, with 100% of mice succumbing to septic shock within 24 h post-infection (P < 0.05) (Fig. 4a). By contrast, none of the SS2to4-infected mice died, presenting transient clinical signs within the first 48 h (P < 0.05) (Fig. 4a).
Impact of serotype switching on S. suis virulence and plasma pro-inflammatory mediator production in a mouse model of infection. (a) Survival of C57BL/6 mice following intraperitoneal inoculation of 1 × 107 CFU of the different S. suis strains and mutants. (b) Blood bacterial burden 24 h post-infection of C57BL/6 mice. A blood bacterial burden of 2 × 109 CFU/mL, corresponding to average burden upon euthanasia, was attributed to euthanized mice. (c) Plasma levels of IL-6, IL-12p70, IFN-γ, CCL2, CCL3, CCL4, CCL5, and CXCL2 in C57BL/6 mice at 12 h following intraperitoneal inoculation of 1 × 107 CFU of the different S. suis strains and mutants. Data represent survival curves (a) (n = 10–12), geometric mean (b) (n = 10–12) or mean ± SEM (C) (n = 8). An asterisk denotes a significant difference with SS2 by Log-rank (Mantel-Cox) test (c) and Mann–Whitney rank sum test (b and c) (p < 0.05).
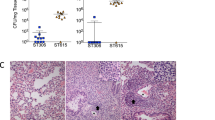
Blood bacterial burdens of infected mice were also determined to investigate the effect on persistent bacteremia. Twenty-four hours post-infection, bacterial burdens of SS2-infected mice averaged 3 × 107 colony-forming unit (CFU)/mL, whereas those in mice infected with ΔCPS2 were not detectable (< 1 × 102 CFU/mL) (Fig. 4b). Similar to mortality, no significant difference was observed between SS2 and SS2to3, SS2to7, SS2to9 or SS2to14 (Figs. 4b and S8). Meanwhile, blood bacterial burden of SS2to8-infected mice was significantly greater than that of SS2-infected mice (P < 0.05), averaging 2 × 109 CFU/mL (Fig. 4b). By contrast, blood bacterial burden was significantly reduced in SS2to4-infected mice compared to SS2 (P < 0.05), although blood burden remained detectable until at least 72 h post-infection, which differs from ΔCPS2-infected mice (Fig. 4b and Supplementary Fig. S8).
Furthermore, plasmatic levels of different pro-inflammatory mediators (12 h post-infection) were evaluated to investigate exacerbated systemic inflammation. The levels were elevated in SS2-infected mice, whereas they were undetectable in ΔCPS2-infected mice (Fig. 4c). Globally, no differences were observed in systemic inflammation between SS2-infected mice and those infected with SS2to7, SS2to9, or SS2to14 (Fig. 4c). However, a significant increase in the production of all the inflammatory mediators was observed in SS2to8-infected mice (P < 0.05), in accordance with the results on mortality observed above (Fig. 4a). Meanwhile, plasmatic levels of all mediators were significantly decreased in SS2to4-infected mice compared to SS2 (P < 0.05), although levels were detectable (Fig. 4c). Notably, infection with SS2to3 resulted in a significant reduction of most pro-inflammatory mediators compared to SS2, though reduction was not as great as with SS2to4 (Fig. 4c).
Serotype switching can differentially modulate S. suis virulence in piglets
Impact of serotype switching on S. suis virulence was subsequently evaluated in the natural host of this bacterium by an experimental intranasal infection model, representing the natural route of exposure to S. suis. The mutants were divided into two experiments (experiment I: SS2, ΔCPS2, SS2to4, or SS2to7; experiment II: SS2, SS2to3, SS2to8, or SS2to14) (Table 2). Virulence of the SS2to9 was not evaluated for ethical reasons, since no differences were observed in host cell interactions assays in vitro nor in the mouse infection model. In experiment I, none of the ΔCPS2-infected pigs developed any clinical signs of infection, while all SS2-infected pigs showed clinical signs of systemic and/or central nervous system infection, including lame and shivering (Supplementary Table S5). In fact, three out of four SS2-infected pigs were euthanized at 3 or 4 days post-infection (dpi) due to severity of clinical signs (Table 2 and Supplementary Table S5). The inoculated strain was recovered from the blood and several organs, including the joints and brain, in all SS2-infected pigs (Table 3 and Supplementary Table S6). Recovery of SS2 from the joints and brain was also confirmed in the animals presenting lameness or shivering (Table 3 and Supplementary Table S6). Meanwhile, recovery of the inoculum was not observed from any of the investigated sites in the ΔCPS2-infected pigs, except for the tonsils (two pigs) and the liver (one pig) (Table 3 and Supplementary Table S6). All SS2to4- and three of SS2to7-infected pigs presented no clinical signs of infection (Table 2 and Supplementary Table S5), which were, except for the tonsils and a single organ, negative for bacterial recovery (Table 3 and Supplementary Table S6). However, one of the SS2to7-infected pigs developed shivering, and bacteria were only recovered from the brain and tonsils (Supplementary Table S6).
Unfortunately, none of the SS2-infected pigs developed clinical signs in experiment II, with recovery only from the tonsils and joints (Table 3 and Supplementary Table S6), although slight fever was observed 4 dpi (Table 2 and Supplementary Table S5). These difference in results of SS2 between experiments may be due to the pigs being used originated from different suppliers. Although most SS2to3-, SS2to8-, or SS2to14-infected pigs showed no clinical signs, one of the SS2to8-infected pigs developed clinical symptoms, including inactivity and clear incoordination (Table 2 and Supplementary Table S5). Nevertheless, SS2to14 was recovered from the blood and organs of one of the infected pigs. Excluding this individual, however, bacterial recovery was mostly negative for SS2to3- or SS2to14-infected pigs. Meanwhile, bacteria were recovered from multiple organs in all the SS2to8-infected pigs, though recovery from blood was recorded in only the individual presenting clinical symptoms (Table 3 and Supplementary Table S6).
Discussion
In the present study, serotype-switched S. suis mutants were generated from serotype 2 to serotypes 3, 4, 7, 8, 9 or 14 via induction of competence state using XIP, which is the first report experimentally demonstrating full cps locus exchange in S. suis. CPS sugar composition and structure of the mutants were identical to those of the same serotype strains previously described in the literatures11,12,13,14,15,16, except for the SS2to9 mutant, of which cps locus was identical to the donor serotype 9 strain used in this study but different from the strain previously used to determine the serotype 9 CPS structure14. In addition, our whole genome sequencing confirmed the deletion of the serotype 2 cps locus and the gain of the expected serotype cps locus of the respective donor strains, and revealed almost the same genetic background of each of the constructed mutant as SS2, enabling strict evaluation of the CPS effect alone; these findings suggest that replacement of the cps locus between the different serotypes alone was responsible for S. suis serotype switching. It should be noted, however, that it remains unclear whether several mutations, which were found in the genome other than cps locus of the respective mutants, affect the phenotype, including virulence and host cell interactions.
This study also provides the first evidence that serotype switch in S. suis can definitively modify the interactions with host cells and in vivo (Summarized in Table 4). CPS expression of S. suis serotypes 2, 9, and 14 plays critical roles on colonization and anti-phagocytic activity, important steps of the pathogenesis20,21,33. In this study, only switching to serotype 7 or 8 changed the adhesion pattern of SS2 to porcine tracheal epithelial cells. The possibility that differential exposure of cell wall components, particularly adhesins, might be the explanation for the increase of S. suis adhesion is worth testing in the future. Differences in thickness of expressed CPS may also responsible, although TEM results suggested similar thickness among mutants. It is also possible that a yet unknown host cell receptor might recognize certain motifs of specific S. suis CPSs. Moreover, results obtained in this study confirmed the previous speculation that the bacterial factors involved in S. suis adhesion and invasion probably differ and that the CPS itself is not involved in the latter21.
Regarding anti-phagocytic activity, no significant (serotype 3, 9 or 14) or minor difference (serotype 4, 7, or 8) was observed by serotype switching, suggesting that the switch of CPS expressed at the S. suis surface may, at least partially, affect the anti-phagocytic properties conferred. A previous study34 reported that serotype 4 and 7 wild-type strains were relatively more internalized by human monocyte-derived DCs than a serotype 2 strain, though the strains used have completely different genetic backgrounds. Here again, this study suggests the possibility that differences in phagocytosis is due to the exposure of differential cell wall component and/or activation of phagocytic receptors such as scavenger receptors and C-type lectins by specific CPS composition/structure. Indeed, C-type lectin receptors are involved in the uptake of Streptococcus pneumoniae35,36, and their potential involvement in S. suis recognition remains to be evaluated.
Results obtained for the two parameters described above (adhesion to epithelial cells and phagocytosis by macrophages) provided the first evidence that the CPS composition/structure can definitively modify S. suis interactions with host cells. However, a single cell-type system does not accurately represent the complexity of the bacterial interplay with its host. By further evaluation of the effects on serotype-switching using ex vivo (blood) and in vivo infection models (mouse and pig), only mutants switched to serotype 4 or 8 showed a marked and consistent impact on several bacterial virulence traits. The CPS4 conferred to S. suis a non-virulent phenotype characterized by increased susceptibility to killing by mouse and pig blood, reduced bacteremia in mice, diminished cytokine production (in vitro and in vivo), and low bacterial recovery from internal organs in pigs. In marked contrast, the CPS8 conferred to S. suis an hyper-virulent phenotype characterized by high capacity to multiply in pig blood, high bacteremia (mice) and organ dissemination (pigs), and increased capacity to induce a cytokine storm (in vitro and in vivo in the mouse model).
It should be noted that switching to serotype 14 or 9 (variant) had no major effects on S. suis virulence or its interactions with the host either in vitro or in vivo in the mouse model. The results on CPS9 in this study makes a striking contrast with the previous study comparing serotypes 2, 14 and 9 with their corresponding non-encapsulated mutants21. Indeed, the CPS9 observation is somehow unexpected; it has been shown that the serotype 9 strain 1135776 adhered more to porcine tracheal epithelial NPTr cells, was more internalized by macrophages, and induced much lower in vitro pro-inflammatory mediator production than the serotype 2 strain P1/7 and the serotype 14 strain 1373021. However, the serotype 9 strain 1135776 used in the study was genetically distinct from the serotype 2 strain P1/7, suggesting that combination of CPS and genetic background of other factors, such as cell wall components, are important for virulence. Meanwhile, serotype switch to CPS7 or CPS3 has restricted impact and affected few of the evaluated parameters. The SS2to7 mutant has slightly increased susceptibility to killing by pig blood and reduced virulence in the swine infection model, being mainly recovered from tonsils. Interestingly, one of the SS2to7-infected pigs developed shivering, and bacteria were only recovered from the brain. It should be noted that serotype 7 strains are isolated in a greater proportion from the central nervous system than from other organs in diseased pigs37. The SS2to3 mutant presented increased susceptibility to killing by mouse and pig blood, slightly reduced bacteremia in mice, and diminished capacity to induce cytokine production in vivo. Though serotype 3 CPS expression still caused S. suis-induced host death, clinical signs were less severe than those caused by SS2 in the mouse model. None of the pigs infected with SS2 developed clinical signs in experiment II, so a reduced virulence of SS2to3 mutant could not be definitively confirmed in the natural host. Overall, results obtained with the different mutants confirmed the delicate balance between bacterial burden, systemic dissemination, level of the inflammatory response, and clinical outcome32,38,39. Given that only different CPSs were expressed between mutants, these differences in effects depending on switched serotypes might be due to differential cell wall component exposure, including adhesins and immunostimulatory components, and/or recognition of certain motifs of specific S. suis CPSs by unknown host cell receptors.
This work also highlighted the complexity of S. suis host–pathogen interactions and the carefulness required when analyzing data from single cell type cultures vs. more complex biological systems (such as blood). For instance, neutrophils and monocytes are the main phagocytes in blood, with little to no macrophages being present. Therefore, results obtained with macrophages might not necessary reflect S. suis fitness in blood, but rather mimic the situation in tissues. Similarly, the interactions of S. suis with swine blood leukocytes are more complex than those evaluated when using mouse blood due to the presence of swine antibodies reacting against the bacteria. Thus, by using multiple in vitro and in vivo models, a more comprehensive analysis is obtained.
In Streptococcus pneumoniae, strict evaluations of the CPS effects using CPS switch mutants have already been performed, and several studies demonstrated that capsule type affected resistance to both complement C3b deposition and opsophagocytic uptake40, nonopsonic neutrophil-mediated killing41, and adhesion to the pharyngeal or lung epithelial cells42. Some of these studies also indicated the effect on virulence within the respiratory tract42, colonization41, survival in blood40, and brain injury43 by in vivo infection models. The structure and composition of CPS8 of S. suis is known to be identical to that of S. pneumoniae serotype 19F16, with serotype 19F pneumococcus mutant being shown to be the most resistant to non-opsonic killing by human neutrophils among the mutants41, suggesting that this structure of CPS provides the bacteria with high resistance to killing in blood. Previous studies using serotype-switched mutants41,44 also showed that CPS type affects the degree of encapsulation and growth phenotype due to the difference in metabolic costs for producing capsule between CPS types. In one of the study44, mix of the bacterial cells with thick capsule and thinner capsule was observed when the pneumococcus mutant switching CPS to serotype 19F was grown in the nutrient-limiting condition, unlike the other serotype-switched mutants (switching to serotypes 7F, 18C and 6B). These points should be evaluated in S. suis, especially serotype 8 CPS, in the future, because these may be one of the explanations of the difference in host interactions and virulence between SS2 and the serotype-switched mutants, in case different effect on the degree of encapsulation or mix of bacterial cells with thin and thick capsule, similar to the pneumococcus mutant switching CPS to serotype 19F, in the nutrient-limiting condition like in vivo.
In conclusion, these data demonstrate that serotype switching in S. suis serotype 2 can modulate host cell interactions and virulence. Among the tested serotypes, switch to serotype 8 increased the virulence. Although it remains unknown whether S. suis serotype switching affects virulence in humans, one serotype 8 strain having a genetic background similar to virulent serotype 2 clinical isolates has already been recovered (unknown source: pubMLST: http://pubmlst.org/ssuis/). Therefore, these results clearly demonstrate that more attention should be given to serotype switching in S. suis with regards to both commensal and pathogenic strains.
Methods
S. suis culturing
The S. suis strains used in this study are listed in Table 1. The serotype 2 strain P1/7 (SS2 in this study)24 was used as the parental strain for construction of the serotype-switched mutants. P1/7 belongs to CC1 and was shown to be induced to a competent state using XIP18. S. suis strains of serotypes 3, 4, 7, 8, 9, and 14 were used as donors to construct the serotype-switched mutants. All strains were cultured overnight on Todd-Hewitt (TH) agar (Becton Dickinson, Franklin Lakes, NJ, USA) at 37 °C with 5% CO2 unless indicated otherwise. Chloramphenicol was added to the medium at 5 μg/mL, when needed.
General molecular biology techniques
All PCRs were completed using the iProof HF Master Mix (BioRad Laboratories, Hercules, CA, USA) and QIAGEN Multiplex Master PCR Mix (Qiagen, Hilden, Germany) according to the manufacturers’ instructions. The PCR primers used in this study are listed in Supplementary Table S7. The amplified PCR products were purified using the QIAQuick PCR Purification Kit (Qiagen) and sequenced on a 3130xl Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) using a BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) where required. The sequence assembly of the PCR products was performed using SEQUENCHER 5.4 (Gene Codes Corp., Ann Arbor, MI, USA).
Construction of serotype-switched mutants and non-encapsulated mutant
An outline of the approach developed for the construction of the serotype-switched mutants is represented in Supplementary Fig. S1. First, a non-encapsulated mutant whose cps locus was replaced with a chloramphenicol resistance gene (ΔCPS2tocat) was generated from SS2. DNA fragments comprising the chloramphenicol cassette flanked by approximately 1 kbp of the upstream and downstream regions of the cps gene cluster were amplified by overlap-extension PCR. SS2 corresponding locus tags of genes deleted were SSU0515-SSU0538. The plasmid pSET145 was used as a template for the PCR to amplify the cat cassette. Five-microliters of the DNA fragment (approximately 500 ng) was then transformed into 100 µL of the SS2 culture [optical density 600 nm (OD600) of 0.035–0.045] by inducing competent state with 10 µL of 2.5 mM XIP as previously described18. After selection of transformants by culturing on TH plates with chloramphenicol, non-encapsulation was confirmed by co-agglutination with anti-serotype 2 serum (Supplementary Fig. S1, panel 1). Then, 100 µL of the ΔCPS2tocat (OD600 of 0.035–0.045) was transformed with 5 µL of whole genome of donor strains (approximately 2 µg of each genome DNA) and XIP (Fig. S1, panel 2). For screening the desired serotype-switched mutant candidates, bacterial cells were collected from the transformed culture by centrifugation at 2,600 × g for 20 min and washed once with 1 mL of 0.15 M NaCl. Percoll density gradient centrifugation was performed as previously described46. The washed cells were suspended with 10 µL of the undiluted Percoll PLUS (GE Healthcare UK Ltd., Buckinghamshire, UK), and the bacterial cell suspension was added to the bottom of a 2 mL microtube. Four hundred microliters of 20, 40, 60, and 80% stock isotonic Percoll (SIP) solution was gently layered onto the washed cell suspension to produce a step gradient with 80% SIP at the bottom of the tube and 20% SIP at the top. The tube was centrifuged at 2600 × g for 20 min to separate bacterial cells according to density. After centrifugation, 100 µL of the solution was collected from the interface between 20 and 40% SIP and between 40 and 60%, since most of the S. suis cells considered to be non-encapsulated are concentrated at the interface between 60 and 80% SIP and those considered to be encapsulated at the interface between 20 and 40% and between 40 and 60%. The collected solutions were spread and cultured on TH agar (Supplementary Fig. S1, panel 3). All of the colonies grown on the agars (100–200 colonies) were subcultured overnight at 37ºC in sterile U-bottom 96-well plates (Corning, NY, USA) with 100 µL of TH broth. The cultures that formed clear precipitates at the tip sections of the bottoms were selected and subcultured overnight at 37ºC with 5% CO2 on both TH agar plates with and without chloramphenicol (Supplementary Fig. S1, panel 4). The cultures that grew only on TH agar without CP were chosen, and the gain of cps gene cluster from the introduced genome DNA was verified by cps type-specific PCR as previously described25. Serotype switch was also confirmed using co-agglutination with antisera of the respective serotypes. For generation of the markerless non-encapsulated mutant, blue-white screening method using 5-bromo-4-chloro-3-indoxyl-α-L-fucopyranoside (X-α-L-fucopyranoside) was performed as represented in Supplementary Fig. S2 (See Supplementary Methods for more detail).
S. suis growth measurements
Strains were streaked onto TH agar plates and incubated overnight at 37 °C with 5% CO2 and then subcultured in TH broth to OD600 of 0.6 using a spectrophotometer Ultrospec 2100 (Biochrom Ltd., Cambridge, UK). After adding 1/500 of the volume of each adjusted culture diluted 1,000 times by TH broth to TH broth, the cultures were incubated at 37 °C under air plus 5% CO2 conditions. The CFU (/mL) of each of the cultures was measured at 2, 4, 6, 8, 10, 12, and 14 h after incubation by plating serial dilutions on TH agar.
Confirmation of serotype switching
Serotyping, cell surface hydrophobicity test, TEM, measurement of CPS yields, NMR spectroscopy were performed to confirm well-encapsulation and serotype switching as previously described [22,47, serotyping and TEM; 27, hydrophobicity tests; 12,14,15,16, CPS purification and NMR] (see Supplementary Methods for more detail).
Whole genome sequence analyses
Whole genome draft sequences were determined using Illumina HiSeq X ten sequencing platform at the Beijing Genomics Institute (Shenzhen, China) or Illumina NovaSeq platform at Novogene Corporation (San Diego, CA, USA) (See Supplementary Methods for more detail). The final draft genome sequence of each of the mutants was then mapped and aligned with the publicly available complete genome sequence of strain P1/7 using Geneious Prime ver. 2019.1.1 (Tomy Digital Biology, Tokyo, Japan) with the default parameters. Calculations of ANI and a fraction shared between genome pairs were conducted using FastANI48.
In vitro assays for evaluation of impacts on serotype switching
Adhesion and invasion assays using the porcine tracheal epithelial NPTr cell line, phagocytosis assays using J774A.1 murine macrophages, murine whole blood bactericidal assay using blood collected from 6- to 10-week-old C57BL/6J mice and from a five-week-old piglet, and measurement of pro-inflammatory mediator production by DCs generated using the femur and tibia of C57BL/6J mice were performed as previously described21,32,49. (see Supplementary Methods for more detail).
In vivo assays for evaluation of impacts on serotype switching
Mouse infections were performed using 10–12 six-week-old male and female C57BL/6J mice per group via intraperitoneal inoculation (dose of 1 × 107 CFU/mouse) for survival and blood bacterial burden evaluation as previously described32. Plasma (systemic) pro-inflammatory mediators were measured using blood collected from eight mice intraperitoneally infected with 1 × 107 CFU 12 h post-infection as previously described32. Pig infections were performed for evaluation of appearance of symptoms and organ dissemination using 4–5 five-week-old crossbred male and female piglets per group purchased from Shokukanken Inc. (Gunma, Japan) or CIMCO Co. Ltd. (Tokyo, Japan). Infections were carried out via intranasal inoculation (dose of 2 × 109 CFU) for survival as previously described50 and divided into two experiments per four groups (Experiment I: SS2, ΔCPS2, SS2to4, and SS2to7; experiment II: SS2, SS2to3, SS2to8, and SS2to14) (see Supplementary Methods for more detail).
Statistical analyses
Normality of data distribution was verified using the Shapiro–Wilk test and Mann–Whitney rank sum tests were performed to evaluate statistical differences between groups. Data are presented as mean ± SEM or as geometric mean. Log-rank (Mantel-Cox) tests were used to compare survival between groups of mice. P < 0.05 was considered statistically significant.
Ethics statement
The animal experiments in this study were carried out in compliance with the ARRIVE guidelines and approved by the institutional committees for Ethics of Animal Experiments of the National Institute of Animal Health Japan (approval numbers 17-002, 17-010, and 17-085) and by the Animal Welfare Committee of the University of Montreal (approval number Rech-1570). Both committees formulated the guidelines and policies required to meet and adhere to the standards in the Guide for the Care and Use of Laboratory Animals.
Data availability
The sequence assembly data determined in this study and their raw data files were deposited in the DDBJ/ENA/GenBank databases under the accession numbers (P1/7, WABV00000000 and SRR13496243; ΔCPS2tocat, WABW00000000 and SRR13496636; SS2to3, WABX00000000 and SRR13488957; SS2to4, WABY00000000 and SRR13488797; SS2to7, WABZ00000000 and SRR13489086; SS2to8, WACA00000000 and SRR13489169; SS2to9, JABMDA000000000 and SRR13485874; SS2to14, WACB00000000 and SRR13489049; MO690, WACC00000000 and SRR13515771; MO691, WACD00000000 and SRR13516280; MO941, WACE00000000 and SRR13516281).
References
Staats, J. J., Feder, I., Okwumabua, O. & Chengappa, M. M. Streptococcus suis: past and present. Vet. Res. Commun. 21, 381–407. https://doi.org/10.1023/a:1005870317757 (1997).
Goyette-Desjardins, G., Auger, J. P., Xu, J., Segura, M. & Gottschalk, M. Streptococcus suis, an important pig pathogen and emerging zoonotic agent—an update on the worldwide distribution based on serotyping and sequence typing. Emerg. Microbes Infect. 3, e45. https://doi.org/10.1038/emi.2014.45 (2014).
Gottschalk, M., Xu, J., Calzas, C. & Segura, M. Streptococcus suis: a new emerging or an old neglected zoonotic pathogen?. Future Microbiol. 5, 371–391. https://doi.org/10.2217/fmb.10.2 (2010).
Gottschalk M. & Segura M. Streptococcosis in Diseases of Swine, 11th ed (ed. Zimmerman, J. J., Karriker, L. A., Ramirez, A., Schwartz, K. J. & Stevenson, G. W.) 934–950. (Wiley-Blackwell, 2019).
Vötsch, D., Willenborg, M., Weldearegay, Y. B. & Valentin-Weigand, P. Streptococcus suis—the “two faces” of a pathobiont in the porcine respiratory tract. Front. Microbiol. 9, 480. https://doi.org/10.3389/fmicb.2018.00480 (2018).
Okura, M. et al. Current taxonomical situation of Streptococcus suis. Pathogens 5, e45. https://doi.org/10.3390/pathogens5030045 (2016).
Huang, J. et al. Identification of six novel capsular polysaccharide loci (NCL) from Streptococcus suis multidrug resistant non-typeable strains and the pathogenic characteristic of strains carrying new NCLs. Transbound. Emerg. Dis. 66, 995–1003. https://doi.org/10.1111/tbed.13123 (2019).
Pan, Z. et al. Novel variant serotype of Streptococcus suis isolated from piglets with meningitis. Appl. Environ. Microbiol. 81, 976–985. https://doi.org/10.1128/AEM.02962-14 (2015).
Zheng, H. et al. Genotyping and investigating capsular polysaccharide synthesis gene loci of non-serotypeable Streptococcus suis isolated from diseased pigs in Canada. Vet. Res. 48, 10. https://doi.org/10.1186/s13567-017-0417-6 (2017).
Okura, M. et al. Genetic analysis of capsular polysaccharide synthesis gene clusters from all serotypes of Streptococcus suis: potential mechanisms for generation of capsular variation. Appl. Environ. Microbiol. 79, 2796–2806. https://doi.org/10.1128/AEM.03742-12 (2013).
Van Calsteren, M. R., Gagnon, F., Lacouture, S., Fittipaldi, N. & Gottschalk, M. Structure determination of Streptococcus suis serotype 2 capsular polysaccharide. Biochem. Cell. Biol. 88, 513–525. https://doi.org/10.1139/O09-170 (2010).
Van Calsteren, M. R. et al. Structure determination of Streptococcus suis serotype 14 capsular polysaccharide. Biochem. Cell. Biol. 91, 49–58. https://doi.org/10.1139/bcb-2012-0036 (2013).
Van Calsteren, M. R. et al. Explaining the serological characteristics of Streptococcus suis serotypes 1 and 1/2 from their capsular polysaccharide structure and biosynthesis. J. Biol. Chem. 291, 8387–8398. https://doi.org/10.1074/jbc.M115.700716 (2016).
Vinogradov, E. et al. Structure determination of Streptococcus suis serotype 9 capsular polysaccharide and assignment of functions of the cps locus genes involved in its biosynthesis. Carbohydr. Res. 433, 25–30. https://doi.org/10.1016/j.carres.2016.07.005 (2016).
Goyette-Desjardins, G. et al. Streptococcus suis serotype 3 and serotype 18 capsular polysaccharides contain di-N-acetyl-bacillosamine. Carbohydr. Res. 466, 18–29. https://doi.org/10.1016/j.carres.2018.07.003 (2018).
Goyette-Desjardins, G. et al. Structure determination of Streptococcus suis serotypes 7 and 8 capsular polysaccharides and assignment of functions of the cps locus genes involved in their biosynthesis. Carbohydr. Res. 473, 36–45. https://doi.org/10.1016/j.carres.2018.12.009 (2019).
Roy, D. et al. A single amino acid polymorphism in the glycosyltransferase CpsK defines four Streptococcus suis serotypes. Sci. Rep. 7, 4066. https://doi.org/10.1038/s41598-017-04403-3 (2017).
Zaccaria, E. et al. Control of competence for DNA transformation in Streptococcus suis by genetically transferable pherotypes. PLoS ONE 9, e99394. https://doi.org/10.1371/journal.pone.0099394 (2014).
Okura, M. et al. A locus encoding variable defense systems against invading DNA identified in Streptococcus suis. Genome Biol. Evol. 9, 1000–1012. https://doi.org/10.1093/gbe/evx062 (2017).
Segura, M., Fittipaldi, N., Calzas, C. & Gottschalk, M. Critical Streptococcus suis virulence factors: are they all really critical?. Trends Microbiol. 25, 585–599. https://doi.org/10.1016/j.tim.2017.02.005 (2017).
Auger, J. P. et al. Interactions of Streptococcus suis serotype 9 with host cells and role of the capsular polysaccharide: comparison with serotypes 2 and 14. PLoS ONE 14, e0223864. https://doi.org/10.1371/journal.pone.0223864 (2019).
Segura, M. et al. Update on Streptococcus suis research and prevention in the era of antimicrobial restriction: 4th International Workshop on S. suis. Pathogens 9, 374. https://doi.org/10.3390/pathogens9050374 (2020).
Gottschalk, M., Higgins, R. & Boudreau, M. Use of polyvalent coagglutination reagents for serotyping of Streptococcus suis. J. Clin. Microbiol. 31, 2192–2194. https://doi.org/10.1128/JCM.31.8.2192-2194.1993 (1993).
Holden, M. T. et al. Rapid evolution of virulence and drug resistance in the emerging zoonotic pathogen Streptococcus suis. PLoS ONE 4, e6072. https://doi.org/10.1371/journal.pone.0006072 (2009).
Okura, M. et al. Development of a two-step multiplex PCR assay for typing of capsular polysaccharide synthesis gene clusters of Streptococcus suis. J. Clin. Microbiol. 52, 1714–1719. https://doi.org/10.1128/JCM.03411-13 (2014).
Zheng, H. et al. Genomic comparisons of Streptococcus suis serotype 9 strains recovered from diseased pigs in Spain and Canada. Vet. Res. 50, 62. https://doi.org/10.1186/s13567-019-0680-9 (2019).
Bonifait, L., Gottschalk, M. & Grenier, D. Cell surface characteristics of nontypeable isolates of Streptococcus suis. FEMS Microbiol. Lett. 311, 160–166. https://doi.org/10.1111/j.1574-6968.2010.02086.x (2010).
Wu, Z. et al. The Streptococcus suis transcriptional landscape reveals adaptation mechanisms in pig blood and cerebrospinal fluid. RNA 20, 882–898. https://doi.org/10.1261/rna.041822.113 (2014).
Fittipaldi, N., Segura, M., Grenier, D. & Gottschalk, M. Virulence factors involved in the pathogenesis of the infection caused by the swine pathogen and zoonotic agent Streptococcus suis. Future Microbiol. 7, 259–279. https://doi.org/10.2217/fmb.11.149 (2012).
Auger, J. P., Dolbec, D., Roy, D., Segura, M. & Gottschalk, M. Role of the Streptococcus suis serotype 2 capsular polysaccharide in the interactions with dendritic cells is strain-dependent but remains critical for virulence. PLoS ONE 13, e0200453. https://doi.org/10.1371/journal.pone.0200453 (2018).
Lecours, M. P. et al. Critical role for Streptococcus suis cell wall modifications and suilysin in resistance to complement-dependent killing by dendritic cells. J. Infect. Dis. 204, 919–929. https://doi.org/10.1093/infdis/jir415 (2011).
Auger, J. P., Fittipaldi, N., Benoit-Biancamano, M. O., Segura, M. & Gottschalk, M. Virulence studies of different sequence types and geographical origins of Streptococcus suis serotype 2 in a mouse model of infection. Pathogens 5, E48. https://doi.org/10.3390/pathogens5030048 (2016).
Segura, M., Calzas, C., Grenier, D. & Gottschalk, M. Initial steps of the pathogenesis of the infection caused by Streptococcus suis: fighting against nonspecific defenses. FEBS Lett. 590, 3772–3799. https://doi.org/10.1002/1873-3468.12364 (2016).
Meijerink, M. et al. Immunomodulatory effects of Streptococcus suis capsule type on human dendritic cell responses, phagocytosis and intracellular survival. PLoS ONE 7, e35849. https://doi.org/10.1371/journal.pone.0035849 (2012).
Park, J. Y. et al. The C-type lectin CD209b is expressed on microglia and it mediates the uptake of capsular polysaccharides of Streptococcus pneumoniae. Neurosci. Lett. 450, 246–251. https://doi.org/10.1016/j.neulet.2008.11.070 (2009).
Rabes, A. et al. The C-type lectin receptor Mincle binds to Streptococcus pneumoniae but plays a limited role in the anti-pneumococcal innate immune response. PLoS ONE 10, e0117022. https://doi.org/10.1371/journal.pone.0117022 (2015).
Prüfer, T. L. et al. Molecular typing of Streptococcus suis strains isolated from diseased and healthy pigs between 1996–2016. PLoS ONE 14, e0210801. https://doi.org/10.1371/journal.pone.0210801 (2019).
Auger, J. P., Benoit-Biancamano, M. O., Bédard, C., Segura, M. & Gottschalk, M. Differential role of MyD88 signaling in Streptococcus suis serotype 2-induced systemic and central nervous system diseases. Int. Immunol. 31, 697–714. https://doi.org/10.1093/intimm/dxz033 (2019).
Lavagna, A. et al. Interleukin-1 signaling induced by Streptococcus suis serotype 2 is strain-dependent and contributes to bacterial clearance and inflammation during systemic disease in a mouse model of infection. Vet. Res. 50, 52. https://doi.org/10.1186/s13567-019-0670-y (2019).
Hyams, C. et al. Streptococcus pneumoniae resistance to complement-mediated immunity is dependent on the capsular serotype. Infect. Immun. 78, 716–725. https://doi.org/10.1128/IAI.01056-09 (2010).
Weinberger, D. M. et al. Pneumococcal capsular polysaccharide structure predicts serotype prevalence. PLoS Pathog. 5, e1000476. https://doi.org/10.1371/journal.ppat.1000476 (2009).
Sanchez, C. J. et al. Changes in capsular serotype alter the surface exposure of pneumococcal adhesins and impact virulence. PLoS ONE 6, e26587. https://doi.org/10.1371/journal.pone.0026587 (2011).
Hathaway, L. J., Grandgirard, D., Valente, L. G., Täuber, M. G. & Leib, S. L. Streptococcus pneumoniae capsule determines disease severity in experimental pneumococcal meningitis. Open Biol. 6, 150269. https://doi.org/10.1098/rsob.150269 (2016).
Hathaway, L. J. et al. Capsule type of Streptococcus pneumoniae determines growth phenotype. PLoS Pathog. 8, e1002574. https://doi.org/10.1371/journal.ppat.1002574 (2012).
Takamatsu, D., Osaki, M. & Sekizaki, T. Construction and characterization of Streptococcus suis-Escherichia coli shuttle cloning vectors. Plasmid 45, 101–113. https://doi.org/10.1006/plas.2000.1510 (2001).
Auger, J. P. et al. Streptococcus suis serotype 2 capsule in vivo. Emerg. Infect. Dis. 22, 1793–1796. https://doi.org/10.3201/eid2210.151640 (2016).
Lakkitjaroen, N. et al. Loss of capsule among Streptococcus suis isolates from porcine endocarditis and its biological significance. J. Med. Microbiol. 60, 1669–1676. https://doi.org/10.1099/jmm.0.034686-0 (2011).
Jain, C., Rodriguez, R. L., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114. https://doi.org/10.1038/s41467-018-07641-9 (2018).
Wang, Y. et al. Capsular sialic acid of Streptococcus suis serotype 2 binds to swine influenza virus and enhances bacterial interactions with virus-infected tracheal epithelial cells. Infect. Immun. 81, 4498–4508. https://doi.org/10.1128/IAI.00818-13 (2013).
Pallarés, F. J. et al. Comparison of experimental models for Streptococcus suis infection of conventional pigs. Can. J. Vet. Res. 67, 225–228 (2003).
Acknowledgements
This work was funded by the JSPS KAKENHI grants #18H02658 (MO and TS) and #26870840 (MO), as well as by the Natural Sciences and Engineering Research Council of Canada (NSERC) grants #04435 (MG) and #342150 (MS). JPA and GGD are recipients of an Alexander Graham Bell Graduate Scholarship – Doctoral Program from NSERC. MS is the holder of a Canada Research Chair – Tier 1. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication. The authors would like to thank Sonia Lacouture for technical help and advice, Mariane Grzebyk and Claudia Duquette for technical assistance with the production and purification of the CPSs, Kaori Tosaki, Kennosuke Sugie, Koujiro Yoshizaki, Yusuke Abeto, and Hirotaka Itoh for TEM analysis, and Han Zheng for providing information on genome sequence of serotype 9 strains. Computational resources were partly provided by the Data Integration and Analysis Facility, National Institute for Basic Biology, Japan.
Author information
Authors and Affiliations
Contributions
M.Okura wrote and revised the manuscript, conceived, designed and conducted the experiments, analyzed that data. J.P.A wrote and revised the manuscript, designed and conducted the experiments, analyzed that data. T.S., G.G.D, M.R.V.C., F.M., and M.K. revised the manuscript, designed and conducted the experiments, analyzed that data. M.Osaki revised the manuscript and helped to analyze the data. M.S., M.G., and D.T. revised the manuscript and conceived the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Okura, M., Auger, JP., Shibahara, T. et al. Capsular polysaccharide switching in Streptococcus suis modulates host cell interactions and virulence. Sci Rep 11, 6513 (2021). https://doi.org/10.1038/s41598-021-85882-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-85882-3
- Springer Nature Limited
This article is cited by
-
Comparative analysis of the interactions of different Streptococcus suis strains with monocytes, granulocytes and the complement system in porcine blood
Veterinary Research (2024)
-
Characterization of pig tonsils as niches for the generation of Streptococcus suis diversity
Veterinary Research (2024)
-
A critical review on experimental Streptococcus suis infection in pigs with a focus on clinical monitoring and refinement strategies
BMC Veterinary Research (2023)
-
Genomic comparison of two Streptococcus suis serotype 1 strains recovered from porcine and human disease cases
Scientific Reports (2023)