Abstract
Target of rapamycin (TOR) is a conserved central growth regulator in eukaryotes that has a key role in maintaining cellular nutrient and energy status. Arbuscular mycorrhizal (AM) fungi are mutualistic symbionts that assist the plant in increasing nutrient absorption from the rhizosphere. However, the role of legume TOR in AM fungal symbiosis development has not been investigated. In this study, we examined the function of legume TOR in the development and formation of AM fungal symbiosis. RNA-interference-mediated knockdown of TOR transcripts in common bean (Phaseolus vulgaris) hairy roots notably suppressed AM fungus-induced lateral root formation by altering the expression of root meristem regulatory genes, i.e., UPB1, RGFs, and sulfur assimilation and S-phase genes. Mycorrhized PvTOR-knockdown roots had significantly more extraradical hyphae and hyphopodia than the control (empty vector) roots. Strong promoter activity of PvTOR was observed at the site of hyphal penetration and colonization. Colonization along the root length was affected in mycorrhized PvTOR-knockdown roots and the arbuscules were stunted. Furthermore, the expression of genes induced by AM symbiosis such as SWEET1, VPY, VAMP713, and STR was repressed under mycorrhized conditions in PvTOR-knockdown roots. Based on these observations, we conclude that PvTOR is a key player in regulating arbuscule development during AM symbiosis in P. vulgaris. These results provide insight into legume TOR as a potential regulatory factor influencing the symbiotic associations of P. vulgaris and other legumes.
Similar content being viewed by others
Introduction
Arbuscular mycorrhizal (AM) fungi, obligate biotrophs belonging to the phylum Mucoromycota and subphylum Glomeromycotina1, are the most ancient and perhaps the most important group of symbionts on earth. Fossil records and molecular clock dating indicate that AM fungal associations emerged 460 million years ago2, supporting the idea that the ability of plants to form these associations is one of the most remarkable and enduring adaptations to life on land. The vast majority of vascular plants can form symbioses with AM fungi3, which help the host plants take up and translocate macronutrients (phosphorus [P] and nitrogen [N]), micronutrients, and water from the soil4. In return, the host plant fulfills the organic carbon5 and lipid6,7 requirements of the AM fungi. AM fungi also protect against pathogenic fungi and diverse abiotic stresses8, making them of great interest for sustainable agriculture9.
The mutualistic relationship between plants and AM fungi begins in the rhizosphere with a molecular dialogue between the partners. In response to strigolactones in the host root exudate10, the germinating AM fungal spores synthesize and secrete Myc factors11. Chitooligosaccharides (COs) and lipo-chitooligosaccharides (LCOs), that promote symbiosis signaling by inducing oscillations in nuclear-associated calcium and inducing the expression of AM fungi-responsive plant genes that trigger phenotypic changes in the host roots11,12. The development of fungal hyphopodia establishes cell-to-cell contact between the host root and the AM fungus, thus initiate the common symbiotic signaling pathway (CSSP). Hyphal invasion and colonization of the root involves formation of an infection peg from the hyphopodium, which mediates hyphal growth into root epidermal cells, and then a prepenetration apparatus, which dictates the intracellular routes by which the fungus traverses the epidermis12. The fungal hyphae grow and ramify within the intra- and intercellular spaces of the root cortex and subsequently invade the inner cortical cells. Within these cells, the fungal hyphae develop finely branched structures called arbuscules. Each arbuscule remains separated from the plant cell cytoplasm by a plant-derived extension of the plasma membrane called the periarbuscular membrane. This membrane surrounds the hyphal branches and constitutes an interface that is specialized for the exchange of nutrients between the partners13. Given the importance of AM symbiosis, the signaling networks that underlie mycorrhizal interactions have been well studied. So far, several genes and transcription factors have been reported to be essential for AM symbiotic associations14 such as RAM1, RAD1, MIG1, ERF1, RAM2, STR1/2, PT4 and PT11 to name a few.
Target of rapamycin (TOR), a phosphatidylinositol kinase-related protein kinase, is functionally and structurally conserved from yeast to animals and plants15. TOR plays a key role in maintaining cellular nutrient and energy status, controlling the activity, localization, and stability of its target proteins by phosphorylation of serine and/or threonine residues16. In mammals and plants, TOR is encoded by a single gene, while fungi have two genes for TOR. TOR signaling pathway operates in two functionally and structurally distinct complexes, TORC1 and TORC217. TORC1 promotes translation, transcription of ribosomal protein genes, growth in response to sugar content and nitrogen availability but inhibits autophagy, while TORC2 governs cellular metabolism and reorganization of the cytoskeleton18.
In plants, several studies have deciphered TOR's crucial roles in developmental processes, including embryogenesis, seedling growth, root and shoot meristem activation, root hair elongation, chloroplast formation, photoautotrophic transition proliferation of leaf primordia, leaf expansion, flowering, and senescence to name few19. Inhibition of TOR causes quantitative changes in the phosphorylation status of hundreds of proteins in Chlamydomonas reinhardtii and Arabidopsis thaliana, indicating an indispensable role of this gene in regulating protein activity20,21. TOR activity is induced by glucose and sucrose by a complex mechanism19,22 and regulates meristem activity by inducing E2F and WUSCHEL transcription factors in root and shoot apical meristems, respectively23,24. Recent studies show TOR regulation of ribosomal protein gene activation and it also differentially regulates preinitiation complex formation at the promoters of the non-ribosomal protein genes thus, demonstrating a complex regulation of gene activation by TOR25. TOR overexpression lines of A. thaliana showed increased vulnerability to both bacterial and fungal pathogens, while plants with reduced TOR signaling exhibited increased resistance26,27. TOR antagonizes the actions of the classic defense hormones salicylic acid and jasmonic acid28. Abiotic and biotic stress affects plant growth and development by reprogramming transcriptional, translational, and metabolic pathways27,29. Previously, we have shown that TOR as an important regulatory factor for rhizobial infection and nodule development in P. vulgaris and hence essential for root nodule symbiosis30.
The nutritional status of the host plant is an important factor for successful AM fungal development31. Conditions that downregulate TOR expression could disrupt perception of the nutritional status of the plant and thus affect the symbiotic association. To study the function of legume TOR in AM symbiosis development and formation, we used RNA-interference to knockdown TOR transcript levels in transgenic hairy roots of common bean (Phaseolus vulgaris). We studied the symbiotic phenotype of the transgenic PvTOR-RNAi roots in detail and analyzed the spatiotemporal expression of PvTOR during AM symbiosis formation. Based on our results, we propose that PvTOR is required for fungal penetration, mycelial growth and arbuscule development in P. vulgaris.
Results
PvTOR is expressed during AM fungal symbiosis in wild-type P. vulgaris
To determine the role of PvTOR in AM fungal symbiosis in the legume, P. vulgaris, we examined the temporal and spatial expression of PvTOR in wild-type P. vulgaris. Quantitative RT-PCR (RT-qPCR) analysis revealed that PvTOR expression varies among organs and tissues (Fig. 1A). PvTOR transcript levels were higher in vegetative organs (hypocotyl, leaf, stem, and root) than in reproductive organs (flower and young pod). To analyze PvTOR expression during the AM fungal symbiosis, we inoculated 5-day-old wild-type seedlings with Rhizophagus irregularis under low Pi (10 µM phosphate (P)) conditions. Uninoculated plants grown under identical conditions were used as controls. Inoculated and uninoculated roots were harvested at various time points and a small portion of each sample was stained with trypan blue to measure the overall percentage of root length colonization (RLC)32 by AM fungi. All inoculated roots (1, 3, and 6 wpi [weeks post-inoculation] with R. irregularis) were colonized successfully, whereas the uninoculated control roots were free of such colonization (Fig. 1B).
Expression of PvTOR in various tissues and in mycorrhized wild-type P. vulgaris plants. (A) RT-qPCR analysis of PvTOR transcript levels in different vegetative and reproductive organs. (B) AM fungal colonized roots were stained and assessed for percent mycorrhizal root length colonization under a light microscope at different time points. The remaining root portions were used for RT-qPCR analysis to measure (C) PvTOR and (D) PT-4 expression in AM roots at different time points. Transcript accumulation was normalized to the expression of EIF4a and IDE, which were used as reference genes. For C and D, the statistical significance of differences between uninoculated and Ri-inoculated roots was determined using an unpaired two-tailed Student’s t-test (*P < 0.05; **P < 0.01; ***P < 0.001). For A–D, error bars refer to the SE of the mean of three biological replicates (n > 9). Ri, R. irregularis; dpi, days post inoculation; wpi, week(s) post inoculation.
We used the remaining portion of each root sample for RT-qPCR analysis of the expression levels of PvTOR and PvPT-4, a previously identified AM fungi-induced gene. The PvTOR mRNA levels were sharply increased at 3 dpi (days post inoculation) in inoculated roots and remained higher than in the uninoculated roots throughout the rest of the experiment (Fig. 1C). As expected, PvPT-4 expression was induced only in inoculated roots in all tested samples (Fig. 1D). Together, these results indicate that PvTOR is expressed in a majority of P. vulgaris tissues and is upregulated under AM fungi symbiotic conditions.
The tissue specificity of PvTOR expression changes in response to AM fungi
The spatial expression patterns of PvTOR during AM fungal symbiosis were investigated using a promoter activity assay in transgenic roots. To create the reporter construct, the 1-kb region upstream of the translation start codon of PvTOR was fused to the chimeric reporter β-glucuronidase (GUS)-enhanced green fluorescent protein (PvTORpro::GUS-GFP). The PvTORpro::GUS-GFP reporter construct was transfected into P. vulgaris by hairy root transformation and the plants were then inoculated with AM fungi. In uninoculated transgenic roots, GUS staining showed that PvTOR was expressed primarily in the root tip (Fig. 2A). After inoculation with AM fungi, the promoter activity was enhanced in the root tip and could also be detected in the elongation and maturation zones of the root (Fig. 2B).
Spatial expression of the PvTOR promoter in R. irregularis-inoculated transgenic P. vulgaris roots. (A–B) Histochemical GUS staining of (A) uninoculated and (B) R. irregularis-inoculated PvTORpro::GUS-GFP transgenic roots at 2 wpi. (C–E) Transmitted light and confocal images of PvTORpro::GUS-GFP transgenic roots at 2 dpi with R. irregularis showing (C) extraradical hyphae in contact with root epidermis, (D) induction of PvTORpro::GUS-GFP expression in the epidermal cells at the site of extraradical hyphal contact and (E) overlay image. (F–H) PvTORpro::GUS-GFP transgenic roots at 2 wpi with R. irregularis showing (F) extraradical hyphae on the root epidermis surface, (G) induction of PvTORpro::GUS-GFP expression in the root colonized by R. irregularis, and (H) overlay image. WGA Alexa Fluor 633 was used to stain (red) fungal cell walls. dpi, days post inoculation; wpi, week(s) post inoculation; e, epidermis; erh, extraradical hyphae.
To further evaluate PvTOR promoter activity during AM fungal invasion, we used confocal microscopy to examine PvTORpro::GUS-GFP transgenic roots. In this assay, R. irregularis was stained with wheat germ agglutinin (WGA) conjugated to Alexa Fluor 633, which produces far-red fluorescence. Once extraradical hyphae of the AM fungi came into contact with the root epidermis (Fig. 2C), PvTOR promoter activity (GFP fluorescence) was detected at the site of fungal penetration (Fig. 2D–E). PvTOR promoter activity was also observed along the mycorrhized root at 2 wpi (Fig. 2F–H). However, neither GUS staining nor GFP fluorescence was observed in uninoculated roots or in R. irregularis (data not shown). Taken together, these results show that the PvTOR promoter was active during the course of AM symbiosis.
PvTOR knockdown alters P. vulgaris root growth and the expression of root meristem regulatory genes
Myc-lipochitooligosaccharides produced by AM fungi during mycorrhizal colonization induce lateral root (LR) branching in M. truncatula33 and increase LR density in P. vulgaris34. In Arabidopsis, TOR specifically regulates the proliferation of primary root meristem cells23. To clarify the involvement of PvTOR in AM fungi-induced changes in LR development, we used RNAi to knockdown PvTOR expression in transgenic hairy roots of P. vulgaris. Quantitative RT-PCR assays confirmed that PvTOR expression was reduced at least sixfold in the PvTOR-RNAi roots relative to control roots transformed with empty pTdT-DC-RNAi vector (Fig. 3A).
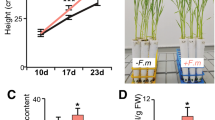
Knockdown of PvTOR inhibits AM fungus-stimulated lateral root production and downregulates the expression of root meristem regulatory genes in P. vulgaris. (A) Quantitative RT-PCR analysis of PvTOR transcript levels in control (empty vector) and PvTOR-RNAi roots at 10 days post emergence. Each bar represents three individual transformants (representative RNAi plant 1, 2, and 3). (B) Lateral root density in uninoculated and Ri-inoculated transgenic roots of control and PvTOR-RNAi at the indicated time points. (C) RT-qPCR analysis of the transcript levels of root meristem regulatory genes such as UPB1, RGF6, RGF9, GSS, APK2, SIR, APS1, APR3, ORC5, ETG1, MCM7, and CDC6 in control and PvTOR-RNAi transgenic roots at 6 dpi with Ri. Quantitative RT-PCR was performed on cDNA of root meristem RNA samples. Transcript accumulation was normalized to the expression of EIF4a and IDE, which were used as reference genes. For A, an unpaired two-tailed Student’s t-test was used to assess statistical significance (***P < 0.001). For B, the statistical significance of differences was determined using Tukey’s test followed by two-way ANOVA the results were statistically significant at p < 0.05 except for the Control (UI) vs. PvTOR-RNAi (UI), Control (UI) vs. PvTOR-RNAi (Ri—I), PvTOR-RNAi (UI) vs. PvTOR-RNAi (Ri—I) at 3 day samples and Control (UI) vs. PvTOR-RNAi (Ri—I) at 6 day samples. For D, Tukey’s test followed by two-way ANOVA was used to asses statistical significance and the mean values of each gene with unlike letters were significantly different (P < 0.05). Error bars refer to the SD of the mean of three biological replicates (n > 9 for A & D, n > 30 for B). Ri, R. irregularis.
To analyze the effects of PvTOR knockdown on root growth, we measured the primary root growth and LR density of PvTOR-RNAi plants with and without inoculation with R. irregularis. We observed that the primary roots of PvTOR-RNAi plants were shorter than those of control plants at all observed time points, regardless of the presence or absence of R. irregularis (Fig. S1). In control plants, the LR density was much higher for inoculated roots than for uninoculated roots at all time points. By contrast, PvTOR-RNAi roots showed only a marginal increase in LR density at the early time points after inoculation (i.e., 6 dpi; Fig. 3B). Quantitative measurements of the root mass confirmed these observations; a marginal increase of LR mass was observed in inoculated PvTOR-RNAi roots relative to uninoculated PvTOR-RNAi roots and, in general, LR mass was significantly decreased in PvTOR-RNAi roots relative to control roots, whether inoculated or not (Fig. S2).
Next, we quantified the transcription of key genes involved in the TOR-mediated regulation of root meristem activity23. The transcription factor gene UPB1, whose overexpression inhibits root meristem expansion through redox control35, was transcriptionally upregulated in PvTOR-RNAi relative to control inoculated root meristems (Fig. 3C). Other genes were upregulated in inoculated roots relative to uninoculated roots, to levels significantly higher in control roots but only marginally higher in PvTOR-RNAi roots. These genes included those encoding ROOT MERISTEM GROWTH FACTORS (RGF6 and RGF9), SULFUR ASSIMILATION GLUTATHIONE SYNTHETASE [GSS], ADENOSINE-5'-PHOSPHOSULFATE KINASE 2 [APK2], SULFITE REDUCTASE [SIR], ATP SULFURYLASE 1 [APS 1], and APS REDUCTASE 3 [APR 3]), and S-phase proteins E2F TARGET GENE 1 [ETG1], MINICHROMOSOME MAINTENANCE 7 [MCM7], and CELL DIVISION CONTROL 6 [CDC6], but not ORIGIN RECOGNITION COMPLEX PROTEIN 5 [ORC5]) (Fig. 3 C).
In summary, knockdown of PvTOR inhibited primary root growth and LR formation in both uninoculated and R. irregularis inoculated roots. Similarly, the transcripts of genes encoding RGFs, sulfur assimilation, and S-phase proteins are suppressed in uninoculated and R. irregularis inoculated PvTOR-RNAi roots.
Knockdown of PvTOR increases the length of AM extraradical hyphae
To examine the effect of PvTOR knockdown on the establishment of AM symbiosis, composite P. vulgaris plants (i.e., plants induced to form hairy roots by transformation with Agrobacterium rhizogenes) were inoculated with R. irregularis and monitored from 1 to 6 wpi. Light microscopy observations of inoculated root surfaces revealed that PvTOR-RNAi roots had more extensive extraradical hyphae (ERH) than control roots and that this difference increased over time (Fig. 4A,B).
Arbuscular mycorrhizal extraradical hyphae in P. vulgaris PvTOR-RNAi roots. (A–B) Transgenic root surface showing extraradical hyphae of mycorrhiza in control (A) and PvTOR-RNAi (B) plants at 1, 3, and 6 week(s) post inoculation (wpi) with R. irregularis. Arbuscule containing cells are indicated with an asterisk. erh, extraradical hyphae; v, vesicle. Bars = 2 mm. (C) Length of arbuscular mycorrhizal extraradical hyphae relative to the dry weight of transgenic roots at 1, 3, and 6 wpi with R. irregularis. n = 9 plants per time point for each line (AM control and AM PvTOR-RNAi). (D) RT-qPCR analysis of the transcript levels of sugar metabolism and sucrose transport genes in control and PvTOR-RNAi transgenic roots at 1 wpi with R. irregularis. Data are the fold-change expression relative to uninoculated transgenic roots. Transcript accumulation was normalized to the expression of EIF4a and IDE, which were used as reference genes. The statistical significance of differences between AM control and AM PvTOR-RNAi roots was determined using an unpaired two-tailed Student’s t-test (*P < 0.05; **P < 0.01). Error bars refer to the SD of the mean of three biological replicates.
To further confirm this observation, we measured the lengths of the ERH. The mean length of ERH on PvTOR-RNAi roots was 2.5-fold greater than that of ERH on control roots at 6 wpi (Fig. 4C). It is worth noting that in Medicago truncatula, knockdown of AM symbiosis-induced SUCROSE SYNTHASE 1 (SUCS1) impaired fungal colonization specifically, resulting in fewer radical hyphae and vesicles during AM symbiosis36. Further, MtSWEET1b is reported to be responsible for arbuscule maintenance by transporting sugar across the peri arbuscular membrane37. We identified the homologues of sugar transporter genes in P. vulgaris based on M. truncatula sequences. The phylogenetic alignment of the sequences showed that MtSWEET1b (Medtr3g089125), PvSWEET1 (Phvul.009G134300) and MtSWEET6 (Medtr3g080990), PvSWEET6 (Phvul.006G000600) were grouped together in the same clade (Fig. S3). With the exception of SWEET1, the expression levels of sugar metabolism and sugar transport genes—namely, SUCS1, SUCS2, CYTOSOLIC INVERTASE 1 (CINV1), and SWEET6-related— were found to be induced in AM fungi inoculated PvTOR-RNAi roots than in mycorrhized control roots (Fig. 4D). Subsequent observations on hyphopodia revealed that the longer ERH on PvTOR-RNAi roots were associated with a significantly higher number of hyphopodia compared to control roots (Fig. 5A, Fig. S4).
Quantification of hyphopodia and arbuscular mycorrhizal colonization in P. vulgaris PvTOR-RNAi roots. (A) Number of hyphopodia per transgenic root of control and PvTOR-RNAi bean plants at 6 wpi with R. irregularis. The statistical significance of differences between control and PvTOR-RNAi roots was determined using an unpaired two-tailed Student’s t-test (***P < 0.001). Error bars refer to the SD of the mean of three biological replicates (n > 30). (B) Total and arbuscular colonization levels in control and PvTOR-RNAi transgenic roots. The composite plants were harvested at 1, 3, and 6 wpi and mycorrhizal colonization parameters were quantified. Error bars refer to the SD of the mean of three biological replicates (n > 30). (C–D) Micrograph of R. irregularis-inoculated transgenic roots showing hyphopodia (arrowhead) and arbuscules in (C) empty vector (control) and (D) PvTOR-RNAi roots at 6 wpi. Insets show magnifications of an arbuscule containing cell (red-dashed regions). wpi, week(s) post inoculation; h, hyphopodia.
We further examined the effect of PvTOR on AM fungal colonization by quantifying root length colonization (RLC) at various time points in control and PvTOR-RNAi roots. Total RLC, arbuscule RLC, and vesicle number were found to increase from 1 to 6 wpi in inoculated control roots. Inoculated PvTOR-RNAi roots showed significantly decreased total RLC (Fig. 5B) and significantly increased vesicle numbers relative to the control at all tested time points (Fig. S5). Furthermore, arbuscule RLC was more than eightfold lower in PvTOR-RNAi roots than in the control (Fig. 5B). Interestingly, the majority of arbuscules present in PvTOR-RNAi roots were abnormal (Fig. 5C–D). These results suggest that PvTOR knockdown causes an increase in ERH, hyphopodia, and vesicles but a decrease in total RLC and arbuscule RLC.
PvTOR is indispensable for arbuscule maturation
Given that PvTOR knockdown affected AM-symbiosis, we examined this phenotype in more detail. We inspected control and PvTOR-RNAi roots for fungal structures at 1 and 3 wpi with R. irregularis. Closer observation revealed that during the initial phases of symbiosis, such as hyphopodia formation and fungal entry into the cortex (intraradical hyphae; IRH), occurred normally in PvTOR-RNAi roots (Fig. 5C–D; Fig. S4). By contrast, although arbuscule development was initiated in PvTOR-RNAi roots, the arbuscules were stunted and clumped (Fig. 6B–C). Furthermore, the PvTOR-RNAi arbuscules were smaller and less densely branched (Fig. 7B, E) than those of the controls (Fig. 6A, 7A, D).
Arbuscular mycorrhizal phenotype of P. vulgaris PvTOR-RNAi roots colonized by R. irregularis. (A) The arbuscules in the cortex cells of the control transgenic roots are well developed. (B–C) Stunted arbuscules in the cortex cells of the PvTOR-RNAi transgenic roots at 1 wpi and 3 wpi, respectively. (D) Quantification of stunted arbuscules per transgenic control and PvTOR-RNAi roots at 3 wpi. The statistical significance of differences between control and PvTOR-RNAi roots was determined using an unpaired two-tailed Student’s t-test (***P < 0.001). Error bars refer to the SD of the mean of three biological replicates (n > 30 for D). a, arbuscule; ih, intercellular hyphae; sa, stunted arbuscule, iah, intracellular hyphae; wpi, week(s) post inoculation, Ri, R. irregularis.
Analysis of arbuscule size in P. vulgaris PvTOR-RNAi roots colonized by R. irregularis. (A-B) Confocal images of R. irregularis arbuscules stained (green) with wheatgerm agglutinin (WGA) conjugated with Alexa Fluor 488 in the cortex cells of control (A) and PvTOR-RNAi (B) bean plants. (C) Frequency of the fluorescence pixel intensity in arbuscule populations measured in control and PvTOR-RNAi transgenic roots at 3 weeks post inoculation. (D–E) Confocal images of arbuscules stained with WGA conjugated with Alexa Fluor 488 in the cortex cells of control (D) and PvTOR-RNAi (E) bean plants. Dotted lines mark the arbuscule area. (F) Frequency of the distribution of the arbuscule area in arbuscule populations measured in control and PvTOR-RNAi transgenic roots. For C and F, mean values of 25 arbuscules per composite plant and a total of three plants are shown. The statistical significance of differences between control and PvTOR-RNAi roots was determined using an unpaired two-tailed Student’s t-test (*P < 0.05; **P < 0.01). Error bars refer to the SE of the mean.
As another measure of arbuscule development, roots were stained with WGA conjugated with Alexa Fluor 488 to label fungal structures and the fluorescence intensity of arbuscule populations was measured using pixel intensity. The majority of arbuscules in PvTOR-RNAi transgenic roots showed fluorescence intensities ranging from 11–30 a.u. compared to intensities of 30–70 a.u. in controls (Fig. 7C). To precisely determine the arbuscule size range in control and PvTOR-RNAi roots, we measured the cross-sectional areas of arbuscule populations and sorted them into size categories. The majority of arbuscules in control roots had an area of 400–600 µm2, while in PvTOR-RNAi roots the area was significantly reduced, with most of arbuscules in the 100–200 µm2 range (Fig. 7F). Over 90% of arbuscules on PvTOR-RNAi roots were stunted and this phenotype persisted at all observed time points (Fig. 6D; Fig. S6). By contrast, the control arbuscules were well developed and highly branched, completely filling the arbuscule-containing cortical cells (Fig. 6A, 7A,D). Interestingly, the number of AM vesicles was significantly higher in the PvTOR-RNAi roots than in the control (Fig. S5).
To test whether PvTOR is required for the uptake of P by the AM fungi, we measured P concentrations in inoculated and uninoculated PvTOR-RNAi and control roots at 3 and 6 wpi. A portion of the root samples was stained to determine AM fungal colonization and the total % RLC was approximately the same as previously shown in Fig. 5B. However, in inoculated plants, the total P concentrations were significantly lower in PvTOR-RNAi roots i.e., 40.2% at 3wpi and 46.9% at 6wpi; and 53.2% at 3wpi and 61.7% at 6wpi in shoots relative to the controls. The same was true of uninoculated plants; P concentrations were lower in PvTOR-RNAi roots and shoots than in the corresponding control samples (Fig. S7), confirming that in the absence of PvTOR, P uptake and transport are disturbed. Therefore, we conclude that PvTOR is essential for mycorrhizal P uptake in P. vulgaris.
Activation of the common symbiosis pathway and expression of AM fungi-induced genes are altered in PvTOR-knockdown roots
In M. truncatula, induction of GRAS-type transcription factors specific to mycorrhizal signaling depends on activation of the common symbiosis signaling pathway (CSSP)38. To confirm that this is also the case for R. irregularis-inoculated transgenic roots of P. vulgaris, we used RT-qPCR to analyze transcript accumulation of the P. vulgaris CSSP genes, SYMRK (Phvul.002G143400), CCAMK (Phvul.011G186900), and IPD3 (Phvul.002G128600), and of genes encoding the AM symbiosis-specific GRAS-type TFs, NODULATION SIGNALING PATHWAY 2 (NSP2) (Phvul.008G165200) REDUCED ARBUSCULAR MYCORRHIZATION 1 (RAM1) (Phvul.001G089900) and RAM2 a GLYCEROL-3-PHOSPHATE ACYL TRANSFERASE (GPAT) (Phvul.007G233600). Transcripts for both CCAMK and IPD3 were less abundant in PvTOR-RNAi roots, while there was no difference in SYMRK expression relative to controls (Fig. 8A). Transcripts for RAM1 and RAM2 were significantly more abundant in PvTOR-RNAi roots than in the control. AM specific markers PvPT4 and P. vulgaris homologues Phvul.003G143400 and Phvul.010G050900 of Oryza sativa AM139 and M. truncatula H+ATPase, HA140 were found to be induced in PvTOR-RNAi roots confirming successful colonization of the symbiont (Fig. 8B).
Expression of P. vulgaris AM regulatory genes in R. irregularis-inoculated transgenic roots. (A) RT-qPCR analysis of the transcript levels of CSSP genes and AM fungi induced TFs genes in control and PvTOR-RNAi transgenic roots at 1 wpi. (B) RT-qPCR analysis of the transcript levels of mycorrhizal-induced genes in control and PvTOR-RNAi transgenic roots at 1 wpi. (A & B) Quantitative RT-PCR was performed on cDNA of root RNA samples. The statistical significance of differences between mycorrhized roots of control and PvTOR-RNAi plants was determined using an unpaired two-tailed Student’s t-test (*P < 0.05; **P < 0.01). Error bars refer to the SE of the mean of three biological replicates (n > 9). CSSP, common symbiotic signaling pathway; AM, arbuscular mycorrhiza; TFs, transcription factors; wpi, week post inoculation.
Other genes induced by AM symbiosis include VAPYRIN (VPY)41 and VAMP (SNARE)42 in M. truncatula and STR1 and STR2 in Oryza sativa43,44. Mutation of these genes limits arbuscule growth of the AM fungi, resulting in a small and stunted arbuscule phenotype43,44. We measured the expression of the P. vulgaris homologs of these genes in inoculated roots and found that—with the exception of STR2—their transcript levels were significantly lower in PvTOR-RNAi roots than in control roots (Fig. 8B). Together, these results suggest that knockdown of PvTOR disrupts the expression of genes involved in arbuscule development but not those encoding AM symbiosis-specific GRAS-type TFs.
Discussion
In plants, TOR plays a central regulatory role in modulating multiple cellular activities, including embryogenesis, meristem activation, root and leaf growth, flowering, and life span determination, as well as controlling photosynthesis, autophagy, and senescence23,45,46. Information about the role of this important protein in the regulation of symbiotic interactions is quite fragmentary. We previously showed that in the model legume P. vulgaris, TOR is involved in regulating rhizobial symbiosis, including infection thread progression and nodule organogenesis30. The focus of our current study was to improve our understanding of the role of PvTOR in regulating interactions with another important endosymbiont, AM fungi. To this end, we analyzed the expression pattern of PvTOR in P. vulgaris, in the presence and absence of AM fungi, and studied the effect of PvTOR knockdown on AM symbiosis.
In terms of PvTOR expression, our key finding was that PvTOR promoter activity, assayed in P. vulgaris hairy roots transformed with a PvTORpro::GUS-GFP reporter construct, increased in response to AM inoculation. The promoter activity in root cortical cells increased in coordination with the ramifying AM fungal mycelia, from the hyphopodium to the ERH to the IRH, an indication of mycorrhiza specific PvTOR promoter expression.
Transformation of P. vulgaris hairy roots with a pTdT-PvTOR RNAi construct resulted in an almost six-fold reduction in PvTOR transcript abundance. This PvTOR knockdown inhibited the proliferation of lateral roots that is normally caused by AM inoculation47. The TOR signaling pathway is also known to regulate the expression of genes involved in the proliferation of progenitor cells for root meristem activation and growth in Arabidopsis23,45. An analysis of the expression of root meristem regulatory genes (RGF, sulfur assimilation, and S-phase genes) in uninoculated PvTOR-RNAi hairy roots revealed significantly lower transcript levels relative to PvTOR-RNAi hairy roots inoculated with AM fungi at 6 dpi. We also observed increased expression of the transcription factor gene UPB1, whose overexpression is known to inhibit root meristem expansion through redox control35. We propose that UPB1 is one of the genes regulated by PvTOR, though further experimental studies are required to verify how TOR and redox regulatory signaling execute the cell proliferation through UPB1 transcription factor.
The ERH perform two main functions: they increase the surface area for mineral and water absorption from the soil and transport to the host via the arbuscule-cortical cell interface and they provide structures capable of colonizing new roots via hyphopodia48. Our observation shows that ERH became more extensive when the arbuscules were truncated by PvTOR knockdown.
AM fungal development in the host is governed by the nutritional status of the host plant. In response to AM colonization, the sink strength of host plant roots increases, allowing more sucrose to be unloaded from the phloem and exported toward the arbuscular cells. This increase in sink strength involves the activity of sucrose-cleaving enzyme invertases (INV) and SUCROSE SYNTHASES (SUCS) and the tight regulation of sucrose transporters49,50. AM-inoculated PvTOR-RNAi roots showed high expression levels of the sucrose synthase genes SUCS1 and SUCS2 and the CYTOSOLIC INVERTASE gene CINV1. The most important finding in these tissues was the transcript downregulation of the sugar transporter PvSWEET1, a homologue of MtSWEET1b. In M. truncatula, SWEET1b is localized to the periarbuscular membrane and is responsible for providing the arbuscule containing cells with the required carbon source for the growth and differentiation of arbuscules. However, the gene encoding the sugar transporter PvSWEET6 was induced in AM-inoculated PvTOR-RNAi roots by two-fold compared to AM-inoculated controls. Hence, PvSWEET6 could be the potential sugar transporter involved in the AM-specific source-to-sink sucrose transport in PvTOR-RNAi plants. Due to the stunted nature of arbuscules in PvTOR-RNAi roots, the available carbon source might be taken up by intraradicular hyphae via an unknown mechanism, as previously proposed by Bago and colleagues51. The increase in vesicle numbers could be to store the extra carbon received from the host.
An increase in ERH was also found to increase the number of hyphopodia. The GRAS-type transcription factors RAM152 and RAM253 function in hyphopodia formation. Similarly, in P. vulgaris, there was an increase in transcript accumulation of NSP2, RAM1, and RAM2 in mycorrhized PvTOR-RNAi roots, providing molecular confirmation for the increased hyphopodia phenotype.
Analyses of the stunted arbuscule phenotype of PvTOR-RNAi roots revealed that the expression levels of VPY41,54, VAMP55, and STR43,44 were supressed. STR and STR2 function as a heterodimer in the periarbuscular membrane that may possibly export a nutrient signal molecule essential for arbuscule development, as in M. truncatula43. In PvTOR-RNAi roots, STR expression was significantly reduced whereas STR2 expression increased during AM symbiosis, implying that in the absence of STR, STR/STR2 heterodimer complex formation is affected and could account for the stunted arbuscules in PvTOR-RNAi roots.
Taken together, our study suggests that PvTOR permits arbuscule maturation during AM symbiosis in P. vulgaris by modulating AM-specific sugar transporters and arbuscule-specific genes. Though there was a surge in mycelial growth and hypopodial numbers, the P content of the PvTOR-RNAi plants did not differ from that of the controls. To improve our understanding of the involvement of PvTOR in the common symbiotic signaling pathway of legumes, future studies should examine the downstream proteins, subcellular localization of TOR, and physiological mechanism of legume TOR regulation during symbiotic interactions.
Methods
Plant materials, inoculation, and growth conditions
Seeds of Phaseolus vulgaris L. cv. Negro Jamapa were obtained from Instituto de Biotecnología, UNAM, Mexico. The seeds were surface-sterilized56, germinated in the dark on wet filter paper for two days at 28 °C, transferred to sterile vermiculite, and grown under a 16-h photoperiod at 28 ± 1 °C. All the experiments involving plants were carried out in accordance with appropriate guidelines. Five-day-old plants were inoculated with Rhizophagus irregularis (800 spores/plant) [Symplanta, Darmstadt, Germany] and irrigated twice weekly with half-strength B&D solution57 containing a low concentration of potassium phosphate (10 µM, K2HPO4) to promote AM colonization13. At different time points, a portion of root sample (50% of total root volume) was excised from each plant, immediately frozen in liquid nitrogen, and stored at –80 °C for RNA extraction. The remaining portion of the root samples was stained to determine the percent root length colonization of AM fungi32. A set of plants grown separately under identical conditions but without R. irregularis inoculation served as controls.
Plasmid construction and generation of composite plants
To develop the RNAi construct of PvTOR, a fragment corresponding to the non-conserved region of the C terminus and 3′-UTR of PvTOR (Phvul.002G049900) was amplified from the cDNA isolated from the root tips of 2-day-old germinated P. vulgaris, using specific oligonucleotides (Table S1). The PCR product was recombined with pTdT-DC-RNAi vector using the Gateway system (Invitrogen, Carlsbad, California, USA). The correct orientation of the clone was confirmed by sequencing the insert of the plasmid. Empty pTdT-DC-RNAi vector was used as the control.
Upstream of the TOR translation start site, a 1-kb promoter fragment was amplified from P. vulgaris genomic DNA using specific primers (Table S1) and cloned into the pENTR/SD/D-TOPO vector (Invitrogen, Carlsbad, California, USA). The Gateway LR reaction was performed between the entry vector pENTR/SD/D-TOPO-PvTOR and the destination vector pBGWSF7.0 according to the manufacturer’s instructions (Invitrogen). The Agrobacterium rhizogenes/K599 strain carrying the corresponding constructs was used to initiate hairy root formation on P. vulgaris tissues and form composite plants after transformation56. Transgenic hairy roots expressing the PvTOR-RNAi vector or PvTORpro::GFP-GUS were selected under an epifluorescence stereomicroscope based on red fluorescent protein (RFP) and green fluorescent protein (GFP) expression, respectively. RFP fluorescence was excited at 561 nm by a solid-state laser and emission was filtered using a band pass filter of 640/50 nm. GFP fluorescence was excited with a blue argon ion laser (488 nm) and emitted fluorescence was collected from 510 to 540 nm.
PvTOR promoter analysis
The PvTORpro::GFP-GUS promoter construct was transfected into common bean cv. Negro Jamapa by hairy root transformation, and the resulting transgenic composite plants were inoculated with R. irregularis (~ 800 spores per plant). The roots were harvested at 5–14 dpi and stained either for promoter fusion GUS activity according to Jefferson58 or WGA (Wheat Germ Agglutinin) conjugated to Alexa Fluor 633 (Invitrogen, Carlsbad, California, USA) to visualize fungal structures59 in RED fluorescence, using a ZEISS LSM-510 confocal laser-scanning microscope. The PvTOR promoter activity was monitored in transgenic roots expressing PvTORpro::GUS-GFP. WGA-Alexa Fluor 633 (red channel) was excited with an argon ion laser (633 nm), and emitted fluorescence was collected from 652 to 752 nm.
RNA extraction and quantitative real-time PCR analysis
Total RNA was isolated from P. vulgaris roots using TRIzol reagent, according to the manufacturer’s recommendations (Thermo Scientific, Waltham, USA). Genomic DNA contamination from RNA samples was eliminated by incubating the samples with RNase-free DNase (1 U µl–1) at 37 °C for 15 min and then at 65 °C for 10 min. RNA integrity and concentration were determined by electrophoresis and NanoDrop ND-2000 (Thermo Scientific, Wilmington, USA) spectrophotometry, respectively.
Quantitative real-time PCR was performed using an iScript One-step RT-PCR Kit with SYBR Green (Bio-Rad, Hercules, California, USA), following the manufacturer’s instructions, in an iQ5 Multicolor Real-time PCR Detection System (Bio-Rad, Hercules, California, USA). Each reaction was set up using 40 ng of RNA as template. A control sample, which lacked reverse transcriptase (RT), was included to confirm the absence of contaminant DNA. Relative gene expression levels were calculated using the formula 2–ΔCT, where cycle threshold value (ΔCT) is the CT of the gene of interest minus the CT of the reference gene. P. vulgaris EIF4a (Phvul.010G136300) and IDE were used as control genes, as previously described23,60. The relative expression values, normalized with two reference genes, were calculated according to Vandesompele and colleagues61. The data are averages of three biological replicates and each sample was assessed in triplicate. The expression of genes listed in Table S1 was quantified using gene-specific oligonucleotides.
Root growth parameters
Composite plants grown in pots of vermiculite and irrigated with B&D medium were used to determine root growth parameters and superoxide accumulation in transgenic roots. Transgenic roots expressing RFP were selected at various intervals from both uninoculated and R. irregularis-inoculated plants and root growth parameters were recorded. Lateral root density was calculated using the formula: D = LR/L´, where D = density of lateral roots; LR = number of lateral roots; and L´ = length of the main root between the first and last lateral root62.
Quantification of mycorrhizal colonization and microscopy analysis
The AM fungi-inoculated roots were sampled at 1, 3, and 6 wpi and stained with trypan blue using the modified histochemical staining method32 or WGA-Alexa Fluor 48859 to measure the mycorrhizal colonization. Using a light microscope (DMLB Bright-field Microscope; Leica, Wetzlar, Germany), trypan blue-stained root samples were analyzed to visualize fungal structures (extraradical hyphae, hyphopodia, intraradical hyphae, vesicles, and arbuscules) and assess the root length colonization (percent RLC) as per McGonigle and colleagues32. Lengths of extra-radical hyphae (ERH) were determined according to a published protocol63 with some modifications64. Arbuscule size was measured using images obtained using a ZEISS LSM-510 confocal laser scanning microscope (ZEISS, Oberkochen, Germany). Z-stacks of Alexa Fluor 488-stained mycorrhized roots were generated from 12–18 serial images taken at increments of 1.25 µM, and analyzed using the LSM 5 tool. Alexa Fluor 488 (green channel) was excited with an argon ion laser (488 nm) and emitted fluorescence was collected from 510 to 540 nm.
Quantification of phosphorus in leaves
To estimate total phosphorus levels in the leaves of composite plants, the dried leaf powder was wet digested using the nitric-perchloric acid method65 and the digested samples were dissolved at 1:20 (w/v) in distilled water and quantified spectrophotometrically (optical density 470 nm). The standard curve was prepared65 using the phosphorous containing salt, KH2PO4.
Statistical analysis
Statistical analysis was performed using Prism Mac 6 (GraphPad Software, California, USA). An unpaired two-tailed Student’s t-test or two-way ANOVA was used to determine the statistical significance of differences between different groups. Single, double, and triple asterisks indicate differences that are statistically significant (P < 0.05 or P < 0.01) or highly significant (P < 0.001), respectively.
References
Spatafora, J. W. et al. A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108, 1028–1046 (2016).
Simon, L., Bosquet, J., Lévesque, C. R. & Lalonde, M. Origin and diversification of endomycorrhizal fungi and coincidence with vascular land plants. Nature 363, 67–68 (1993).
Wang, B. & Qiu, Y. L. Phylogenetic distribution and evolution of mycorrhizas in land plants. Mycorrhiza 16, 299–363 (2006).
Govindarajulu, et al. Nitrogen transfer in the arbuscular mycorrhizal symbiosis. Nature 435, 819–823 (2005).
Smith, S. E. & Smith, F. A. Roles of arbuscular mycorrhizas in plant nutrition and growth: new paradigms from cellular to ecosystems scales. Annu. Rev. Plant Biol. 63, 227–250 (2011).
Jiang, Y. et al. Plants transfer lipids to sustain colonization by mutualistic mycorrhizal and parasitic fungi. Science 356, 1172–1175 (2017).
Brands, M., Wewer, V., Keymer, A., Gutjahr, C. & Doermann, P. The Lotus japonicus acyl-acyl carrier protein thioesterase FatM is required for mycorrhiza formation and lipid accumulation of Rhizophagus irregularis. Plant J. 95, 219–232 (2018).
Lenoir, I., Fontaine, J. & Lounèshadj, S. A. Arbuscular mycorrhizal fungal responses to abiotic stresses: a review. Phytochemistry 123, 4–15 (2016).
Gianinazzi, S. et al. Agroecology: the key role of arbuscular mycorrhizas in ecosystem services. Mycorrhiza 20, 519–530 (2010).
Besserer, A. et al. Strigolactones stimulate arbuscular mycorrhizal fungi by activating mitochondria. PLoS Biol. 4, 226 (2006).
Maillet, F. et al. Fungal lipochitooligosaccharide symbiotic signals in arbuscular mycorrhiza. Nature 469, 58–63 (2011).
Genre, A., Chabaud, M., Faccio, A., Barker, D. G. & Bonfante, P. Prepenetration apparatus assembly precedes and predicts the colonization patterns of arbuscular mycorrhizal fungi within the root cortex of both Medicago truncatula and Daucus carota. Plant Cell 20, 1407–1420 (2008).
Smith, S. E. & Read, D. J. Mycorrhizal Symbiosis 3rd edn. (Academic Press, London, 2008).
Choi, J., Summers, W. & Paszkowski, U. Mechanisms underlying establishment of arbuscular mycorrhizal symbioses. Annu. Rev. Phytopathol. 56, 135–160 (2018).
Wullschleger, S., Loewith, R. & Hall, M. N. TOR signaling in growth and metabolism. Cell 124, 471–484 (2006).
Hay, N. & Sonenberg, N. Upstream and downstream of mTOR. Genes Dev. 18, 1926–1945 (2004).
Shimobayashi, M. & Hall, M. N. Making new contacts: the mTOR network in metabolism and signalling crosstalk. Nat. Rev. Mol. Cell Biol. 15, 155–162 (2014).
Oh, W. J. & Jacinto, E. mTOR complex 2 signaling and functions. Cell Cycle 10, 2305–2316 (2011).
Shi, L., Wu, Y. & Sheen, J. TOR signaling in plants: conservation and innovation. Development 145, dev160887 (2018).
Roustan, V. & Weckwerth, W. Quantitative phosphoproteomic and system-level analysis of TOR inhibition unravel distinct organellar acclimation in Chlamydomonas reinhardtii. Front. Plant Sci. 9, 1590 (2018).
Van Leene, J. et al. Capturing the phosphorylation and protein interaction landscape of the plant TOR kinase. Nat. Plants 5, 316–327 (2019).
Dobrenel, T. et al. TOR signaling and nutrient sensing. Annu. Rev. Plant Biol. 67, 261–285 (2016).
Xiong, Y. et al. Glucose-TOR signalling reprograms the transcriptome and activates meristems. Nature 496, 181–186 (2013).
Pfeiffer, A. et al. Integration of light and metabolic signals for stem cell activation at the shoot apical meristem. Elife 5, e17023 (2016).
Uprety, B., Kaja, A. & Bhaumik, S. R. TOR facilitates the targeting of the 19S proteasome subcomplex to enhance transcription complex assembly at the promoters of the ribosomal protein genes. Mol. Cell Biol. 38, e00469-e517 (2018).
Soprano, A. S., Smetana, J. H. C. & Benedetti, C. E. Regulation of tRNA biogenesis in plants and its link to plant growth and response to pathogens. BBA Gene Regul. Mech. 1861, 344–353 (2018).
Margalha, L., Confraria, A. & Baena, E. SnRK1 and TOR: modulating growth–defense trade-offs in plant stress responses. J. Exp. Bot. 70, 2261–2274 (2019).
De Vleesschauwer, D. et al. Target of rapamycin signaling orchestrates growth-defense trade-offs in plants. New Phytol. 217, 305–319 (2018).
Bechtold, U. & Field, B. Molecular mechanisms controlling plant growth during abiotic stress. J Exp Bot. 69, 2753–2758 (2018).
Nanjareddy, K. et al. A Legume TOR protein kinase regulates Rhizobium symbiosis and is essential for infection and nodule development. J. Plant Physiol. 172, 2002–2020 (2016).
Muller, A., Ngwene, B., Peiter, E. & George, E. Quantity and distribution of arbuscular mycorrhizal fungal storage organs within dead roots. Mycorrhiza 27, 201–210 (2017).
McGonigle, T. P., Millers, M. H., Evans, D. G., Fairchild, G. L. & Swan, J. A. A new method which gives an objective measure of colonization of roots by vesicular-arbuscular mycorrhizal fungi. New Phytol. 115, 495–501 (1990).
Oláh, B., Brière, C., Bécard, G., Dénarié, J. & Gough, C. Nod factors and a diffusible factor from arbuscular mycorrhizal fungi stimulate lateral root formation in Medicago truncatula via the DMI1/DMI2 signalling pathway. Plant J. 44, 195–207 (2005).
Arthikala, M. K. et al. PvRbohB negatively regulates Rhizophagus irregularis colonization in Phaseolus vulgaris. Plant Cell Physiol. 54, 1391–1402 (2013).
Tsukagoshi, H., Busch, W. & Benfey, P. N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 143, 606–616 (2010).
Baier, M. C. et al. Knockdown of the symbiotic sucrose synthase MtSucS1 affects arbuscule maturation and maintenance in mycorrhizal roots of Medicago truncatula. Plant Physiol. 152, 1000–1014 (2010).
An, J. et al. A Medicago truncatula SWEET transporter implicated in arbuscule maintenance during arbuscular mycorrhizal symbiosis. New Phytol. 224, 396–408 (2019).
Oldroyd, G. E. Speak, friend, and enter: Signalling systems that promote beneficial symbiotic associations in plants. Nat Rev Microbiol. 11, 252–263 (2013).
Gutjahr, C. et al. Arbuscular mycorrhiza-specific signaling in rice transcends the common symbiosis signaling pathway. Plant Cell 20, 2989–3005 (2008).
Krajinski, F. et al. The H+-ATPase HA1 of Medicago truncatula is essential for phosphate transport and plant growth during arbuscular mycorrhizal symbiosis. Plant Cell 26, 1808–1817 (2014).
Pumplin, N. et al. Medicago truncatula Vapyrin is a novel protein required for arbuscular mycorrhizal symbiosis. Plant J. 61, 482–494 (2010).
Sogawa, A. et al. SNARE Proteins LjVAMP72a and LjVAMP72b are required for root symbiosis and root hair formation in Lotus japonicus. Front. Plant Sci. 9, 1992 (2019).
Zhang, Q., Blaylock, L. A. & Harrison, M. J. Two Medicago truncatula half-ABC transporters are essential for arbuscule development in arbuscular mycorrhizal symbiosis. Plant Cell 22, 1483–1497 (2010).
Gutjahr, C. et al. The half-size ABC transporters STR1 and STR2 are indispensable for mycorrhizal arbuscule formation in rice. Plant J. 69, 906–920 (2012).
Ren, M. et al. Target of rapamycin signaling regulates metabolism, growth, and life span in Arabidopsis. Plant Cell 24, 4850–4874 (2012).
Li, L. et al. TOR-inhibitor insensitive-1 (TRIN1) regulates cotyledons greening in Arabidopsis. Front Plant Sci. 6, 861 (2015).
Gutjahr, C., Casieri, L. & Paszkowski, U. Glomus intraradices induces changes in root system architecture of rice independently of common symbiosis signaling. New Phytol. 182, 829–837 (2009).
Friese, C. F. & Allen, M. F. The spread of VA mycorrhizal fungal hyphae in the soil: inoculum types and external hyphal architecture. Mycologia 83, 409–418 (1990).
Schaarschmidt, S., Roitsch, T. & Hause, B. Arbuscular mycorrhiza induces gene expression of the apoplastic invertase LIN6 in tomato (Lycopersicon esculentum) roots. J. Exp. Bot. 57, 4015–4023 (2006).
Boldt, K. et al. Photochemical processes, carbon assimilation and RNA accumulation of sugar transporter genes in tomato arbuscular mycorrhiza. J Plant Physiol. 168, 1256–1263 (2011).
Bago, B. et al. Carbon export from arbuscular mycorrhizal roots involves the translocation of carbohydrate as well as lipid. Plant Physiol. 131, 1496–1507 (2003).
Gobbato, E. et al. A GRAS-type transcription factor with a specific function in mycorrhizal signaling. Curr. Biol. 22, 2236–2241 (2012).
Wang, E. et al. A common signaling process that promotes mycorrhizal and oomycete colonization of plants. Curr Biol. 22, 2242–2246 (2012).
Feddermann, N. et al. The PAM1 gene of petunia, required for intracellular accommodation and morphogenesis of arbuscular mycorrhizal fungi, encodes a homologue of VAPYRIN. Plant J. 64, 470–481 (2010).
Ivanov, S. et al. Rhizobium-legume symbiosis shares an exocytotic pathway required for arbuscule formation. Proc. Natl. Acad. Sci. USA 109, 8316–8321 (2012).
Nanjareddy, K., Arthikala, M. K., Aguirre, A. L., Gómez, B. M. & Lara, M. Plant Promoter Analysis: Identification and Characterization of Root Nodule Specific Promoter in the Common Bean. J. Vis. Exp. 130, 56140 (2017).
Broughton, W. J. & Dilworth, M. J. Control of leghemoglobin synthesis in snake beans. Biochem J. 125, 1075–1080 (1971).
Jefferson, R. A. Assaying chimeric genes in plants, the GUS gene fusion system. Plant Mol. Biol. Rep. 5, 387–405 (1987).
Javot, H., Penmetsa, R. V., Terzaghi, N., Cook, D. R. & Harrison, M. J. A Medicago truncatula phosphate transporter indispensable for the arbuscular mycorrhizal symbiosis. Proc. Natl. Acad. Sci. USA 104, 1720–1725 (2007).
Borges, A., Tsai, S. M. & Caldas, D. G. Validation of reference genes for RT-qPCR normalization in common bean during biotic and abiotic stresses. Plant Cell Rep. 31, 827–838 (2012).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal reference genes. Genome Biol. 3, research0034.1-0034.11 (2002).
Dubrovsky, J. G., Gambetta, G. A., Hernandez-Barrera, A., Shishkova, S. & Gonzalez, I. Lateral root initiation in Arabidopsis: developmental window, spatial patterning, density and predictability. Ann. Bot. 97, 903–915 (2006).
Jakobsen, I., Abbott, L. K. & Robson, A. D. External hyphae of vesicular-arbuscular mycorrhizal fungi associated with Trifolium subterraneum L. 1. Spread of hyphae and phosphorus inflow into roots. New Phytol. 120, 371–380 (1992).
Schweiger, R., Baier, M. C., Persicke, M. & Müller, C. High specificity in plant leaf metabolic responses to arbuscular mycorrhiza. Nat. Communi. 5, 3886 (2014).
Miller, R. O. Nitric-Perchloric Acid Wet Digestion in an Open Vessel. In Handbook of methods for plant analysis. 57–61 (CRC Press, Taylor & Francis Group, London, 1998).
Acknowledgements
We thank Brenda-Mariana Gómez, ENES-Leon, UNAM, for technical assistance and for maintaining the plants in the growth chambers. Salma-Sarai González, ENES-Leon, UNAM for formatting the text.
Funding
The authors wish to acknowledge Dirección General de Asuntos del Personal Académico (DGAPA/PAPIIT-UNAM) for partially funding this research (Grant no. IA2O7219 and IN213221 to MKA, IN211218 to KN and CONACyT 240614 to ML).
Author information
Authors and Affiliations
Contributions
M.L. and M.-K.A. conceived the research and designed the experiments; K.N. and M.-K.A. performed all the experiments. LB maintained the plants and mycorrhizal cultures and X.A.-A. performed the confocal microscopy experiments. M.-K.A. and K.N. analyzed the data and wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arthikala, MK., Nanjareddy, K., Blanco, L. et al. Target of rapamycin, PvTOR, is a key regulator of arbuscule development during mycorrhizal symbiosis in Phaseolus. Sci Rep 11, 11319 (2021). https://doi.org/10.1038/s41598-021-90288-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90288-2
- Springer Nature Limited
This article is cited by
-
Genome-wide identification and comparative analysis of the Amino Acid Transporter (AAT) gene family and their roles during Phaseolus vulgaris symbioses
Functional & Integrative Genomics (2024)