Abstract
Banana is an important fruit crop in the tropics and subtropics; however, limited information on biomarkers and signature volatiles is available for selecting commercial cultivars. Clonal fidelity is a major contributor to banana yield and aroma; however, there are no useful biomarkers available to validate clonal fidelity. In this study, we performed the molecular profiling of 20 banana cultivars consisting of diploid (AA or AB) and triploid (AAA or AAB or ABB) genomic groups. We screened 200 molecular markers, of which 34 markers (11 RAPD, 11 ISSR, and 12 SSR) yielded unequivocally scorable biomarker profiles. About 75, 69, and 24 allelic loci per marker were detected for RAPD, ISSR, and SSR markers, respectively. The statistical analysis of molecular variance (AMOVA) exhibited a high genetic difference of 77% with a significant FST value of 0.23 (p < 0.001). Interestingly, the UBC-858 and SSR CNMPF-13 markers were unique to Grand Nain and Ardhapuri cultivars, respectively, which could be used for clonal fidelity analysis. Furthermore, the analysis of banana fruit volatilome using headspace solid-phase microextraction-gas chromatography-tandem mass spectrometry (HS-SPME-GCMS) revealed a total of fifty-four volatile compounds in nine banana cultivars with 56% of the total volatile compounds belonging to the ester group as the significant contributor of aroma. The study assumes significance with informative biomarkers and signature volatiles which could be helpful in breeding and for the authentic identification of commercial banana cultivars.
Similar content being viewed by others
Introduction
Banana (Musa spp.) is one of the world’s major fruit crops belonging to the Musaceae family. It is native to the tropical region of Southeast Asia1. India ranks first in banana production, with 31.5 million tons accounting for 26.43% of global output2. Banana is rich in carbohydrates, vitamin B, and a good source of sodium, potassium, calcium, and magnesium3,4. It is a perennial with a faster relative growth rate than other fruit crops and produces fruits throughout the year. Banana cultivars are propagated asexually via micropropagation or suckers5.
The cultivated edible Musa spp. originated from two wild species, Musa acuminata Colla (A genome) and Musa balbisiana Colla (B genome); both belong to the section Eumusa. The banana cultivars possess good genetic diversity with genome groups of AA, AB, AAA, AAB, ABB, AABB, AAAB, or ABBB type resulting from inter or intra-specific hybridization of M. acuminata and M. balbisiana1,6. Morphological data have suggested that Musa spp. is diverse and showed some well-defined unifying characteristics, indicating the genomic constitution. Complex genome structure and phylogenetic relationship among the hybrid cultivars and wild types need further exploration. Due to its narrow genetic base, banana cultivation is prone to pests and diseases7. Further, this is exacerbated by devastating extremities of abiotic stresses induced by climate change and global warming8. Identifying genotypes with high potential is critical for an effective, economical, and environmentally friendly approach to boosting banana productivity. The wild germplasm (BB-rich genomes) possesses hardiness and could act as potential donors in developing high-performing cultivars through molecular breeding9,10. In this regard, the utility of molecular tools in selecting and developing biotic and abiotic stress-resistant genotypes is of paramount importance in banana breeding programs8,11,12.
Researchers and farmers essentially require authentic high-yielding cultivars to maximize the output. However, traditional methods based on the phenotypic selection are complex and irreproducible to identify authentic cultivars. Hence, it is necessary to search for and develop novel molecular biomarkers which are stably inherited and unperturbed by environmental deviations11. Such molecular markers can be valuable for assessing genetic diversity to complement traditional breeding methods. Moreover, the genetic content (molecular identity information) is highly stable and not influenced by environmental factors; hence differences among genotypes could be discerned at the DNA level. Molecular markers are being widely used in plants to detect the genetic basis of variation, genetic relationships within germplasm, identification of duplicate accessions, and genetic fidelity testing. Several marker-assisted traits have been used in banana breeding13. Restriction fragment length polymorphism (RFLP)1,7,14; random amplified polymorphic DNA (RAPD)9 amplified fragment length polymorphism (AFLP)15,16,17; and microsatellites or simple sequence repeats (SSRs) and inter-simple sequence repeats (ISSRs)18 have been employed in the characterization and evaluation of genetic diversity in Musa species. Amongst these markers, SSRs show a higher degree of polymorphism and codominant inheritance mode and are reproducible11,19.
Fruit volatile (the total volatile compounds in banana fruit) is the vital contributor to banana appeal and aroma characteristics which influence fruit delicacy, quality, and consumer preference. Banana fruits are rich in flavor and aroma with the occurrence of more than 250 volatile compounds of several chemical classes, including esters, ketones, terpenes, and aldehydes20. Despite the limited literature on banana fruit volatiles and their contribution to aroma21,22, the role of banana volatiles has been indicated in the resistance mechanism to some important fungal diseases like Sigatoka23 and insect pests. The study of volatile components from banana fruits is also very important to assess the effect of changing climatic conditions on fruit quality*1. In light of these, we sought to understand the characteristic volatilome of each banana cultivar. Profiling of volatile compounds representing characteristic aroma and flavor of banana germplasm by using headspace solid-phase microextraction (HS-SPME) followed by gas chromatography and mass spectrometry (GC–MS)24,25 provides an additional tool for characterization of banana germplasm which could be utilized for decoding the flavor of diverse banana cultivars and breeding for enhanced sensory quality20,22,25,26.
In the present study, we have used various molecular markers to evaluate the genetic diversity among commercially grown banana cultivars belonging to different genomic groups. This study revealed the identification of cultivars-specific biomarkers which may help to establish genetic relationships. Additionally, the HS-SPME and GC–MS were used to develop the volatile profile of the genotypes. Such markers will enable plant breeders to select banana cultivars with better aroma, flavor, and desirable characteristics, besides helping researchers and farmers verify commercially grown varieties’ genetic fidelity.
Materials and methods
Plant genetic resources
The leaf samples of twenty different germplasm of banana comprising diploid and triploid genomes were collected from the Banana Research Station, Nanded affiliated to Vasantrao Naik Marathwada Krishi Vidyapeeth (VNMKV), Parbhani (Table 1). The study complies with relevant institutional biosafety regulations and guidelines. The genomic DNA was extracted using the modified CTAB method27 followed by RNase treatment. The qualitative and quantitative analysis of genomic DNA was done using a spectrophotometer and by resolving on 0.8% agarose gel. The Genomic DNA was diluted to 50 ng/µL and subsequently used for RAPD, ISSR, and SSR fingerprint analysis.
RAPD, ISSR, and SSR genotyping
A total of 200 primers were used for pre-screening and of which 11 RAPD, 11 ISSR, and 12 SSR primers were used for genotyping twenty banana cultivars (Table 2). Banana cultivars representing each genome type i.e. Namarai from diploid, Birbutia and Grand Nain from triploid, and NRCB-3 from tetraploid were selected for pre-screening. Among the 75 RAPD, 60 ISSR and 65 SSR primers; the primers showing amplification in the all above selected banana cultivars were shortlisted. PCR reactions for RAPD genotyping were carried out in 25.0 µL reaction volume containing 2.5 µL 10× PCR assay buffer with MgCl2(10×), 0.5 µL dNTP mix (10 mM), 0.33 µL Taq DNA polymerase (3 U/µL), 2 µL Primer (10 µm/µL) and 1 µL template DNA (50 ng/µL). ISSR PCR reaction of 25.0 µL comprised 2.5 µL 10× PCR assay buffer with MgCl2, 0.5 µL dNTP mix (10 mM), 0.4 µL Taq DNA polymerase (3 U/µL), 1 µL Primer (10 µm/µL) and 1 µL template DNA (50 ng/µL). The SSR genotyping were performed in 25.0 µL reaction with 2.5 µL 10× PCR assay buffer, 1.5 µL MgCl2(25 mM), 0.5 µL dNTP mix (10 mM), 0.5 µL Taq DNA polymerase (3 U/µL), 1.5 µL each forward and reverse primer (10 µm/µL) and 1.5 µL template DNA (50 ng/µL). The RAPD PCR was carried out in Eppendorf thermal cycler for 5 min at 94 °C, followed by 40 cycles of 45 s at 94 °C, 1 min at 36 °C, 1 min at 72 °C with a final extension time of 10 min at 72 °C. The thermal cycler conditions for ISSR genotyping were initial denaturation at 94 °C for 5 min followed by 40 cycles of 45 s at 94 °C, 1 min at 52 °C, 1 min at 72 °C with a final extension time of 10 min at 72 °C. A thermal cycler for SSR PCR was carried out at 94 °C for 4 min, 35 cycles of 94 °C for 40 s, 54 °C for 30 s, and 72 °C for 40 s followed by a final extension of 10 min at 72 °C. The RAPD and ISSR PCR amplified products were resolved on 1.5% agarose gel at 100 V. Whereas, SSR PCR products were resolved on 2.5% agarose gel at 100 V, followed by ethidium bromide staining. The images were captured by a gel documentation system (AlphaImager, TM-2200).
Data scoring and analysis
All visible and unambiguously scorable fragments amplified by the RAPD, ISSR, and SSR primers were scored as scorable fragments. Amplification profiles of the germplasm were compared, and bands of DNA fragments were scored as present (1) or absent (0). The DNA fingerprint data of the primers were utilized to estimate genetic similarity based on a number of shared amplification products28. The equation used was the number of shared amplification products = 2 × (number of typical bands between any two lanes)/(Total number of bands in the same two lanes). Genetic relationship among the germplasm was estimated with the dendrogram constructed through an unweighted pair group of arithmetic means using (UPGMA)29 and NTSYS pc 2.02 software. Further, the population genetic structure was analyzed based on SSR amplicons score as allele size (bp) using the Bayesian clustering method and STRUCTURE v.v 2.3.3 software30. The admixture model and correlated allele frequencies were applied for the estimation of ancestry fractions of each cluster. The value of K (range 1–10) was estimated using five independent runs and a burn-in period of 20,000 followed by 200,000 MCMC (Markov Chain Monte Carlo) repetitions. The web-based software STRUCTURE HARVESTER version 0.6.92 was used to determine the optimum K value using the log probability of data, LnP(D) based on the rate of change in LnP(D) between successive K31.
Volatile profiling of Banana cultivars using HS-SPME GCMS
The unripe banana fruits of available banana cultivars were collected from the Banana Research Station, Nanded, and kept for ripening at 20 °C.The pulp of fully ripe banana fruits was used for volatile analysis. A blended pulp juice from the three banana fruits of each banana cultivar was prepared separately, and 16 g of juice in a 30 ml glass vial tightly sealed with a Teflon septum with a plastic cap was used for volatile analysis. Optimized SPME parameters, i.e., sample equilibration for 15 min, adsorption for 60 min at 25 °C, and desorption time of 10 min, were used for GCMS sampling32. The 430 GC analyzed the volatile compounds, and 210 MS (Varian) gas chromatograph-mass spectrometer (GC–MS) equipped VF-5 MS column with helium as the carrier gas at a flow rate of 1.0 mL/min was used for detection of m/z ratio of fragmented ions and derived spectrum. The injection port was lined with a 0.75 mm, i.e., splitless glass inserter, and maintained at 200 °C. The oven temperature was programmed to rise from 50 to150 °C at 2 °C/min and the total GC run time was 55 min. MS transfer line was maintained at 290 °C, ionization energy was 70 eV, and the mass range was 50–550 m/z. The volatile compounds were identified by comparing the results obtained with the reference mass spectra from the NIST library (NIST98, version 2.0, Gaithersburg, USA) and retention index values obtained using a standard mixture of alkanes (C8–C20).
Results
Various molecular markers have been used extensively to screen the diversity and develop biomarkers for clonal identity in crops, including banana9,17,33. Here we employed three different markers—RAPD, ISSR, and SSR to search the cultivar-specific biomarkers. The molecular marker provided genetic information and assisted in finding out biomarkers specific to Grand Nain and Ardhapuri cultivars. Additionally, we performed the volatile profiling of banana cultivars to characterize the compounds that contribute to the characteristic aroma of the banana fruits.
Screening of diploid and triploid cultivars of banana (Musa spp) predominantly cultivated in India
RAPD profiling of twenty banana cultivars produced polymorphic DNA fingerprint patterns and generated 725 polymorphic amplicons. Most RAPD primers produced 100% polymorphism (except OPA-13, OPA-18, OPD-15, and OPN-02, with an average polymorphism of 89.97%; Tables 3, 4, 5). Additionally, we recorded the average polymorphic bands per genotype ranging from 4.95 to 1.75 with a median value of 3.29. Moreover, the average number of loci per primer ranged from 5 to 10, with a median score of 6.81 loci per primer. The RAPD primer OPD-18 amplified the highest allelic loci, i.e., 10 followed by primers OPA-13, OPD-20, and OPN-02, each with 08 allelic loci. We evaluated the allele size of the RAPD profile, and it ranged from 0.1 to 1.5 kb. The polymorphic information content (PIC) value ranged from 0.43 for OPA-17 and OPA-10 to 0.25 for OPA-11 (Table 3). Furthermore, the average Shannon’s diversity index (H), expected heterozygosity (He), resolution power (RP), and marker index (MI) values for RAPD primers were 0.53, 0.35, 7.11, and 2.62, respectively. The RAPD primers OPD-18 and OPA-13 have recorded the highest resolution power and MI value. Overall, the RAPD marker-assisted molecular profiling produced highly polymorphic content.
Next, the ISSR primers were used to evaluate the molecular diversity of banana cultivars. The ISSR markers showed an average of 79.62% polymorphism. Among all ISSR markers used, five ISSR primers—ISSR-810, ISSR-818, ISSR-826, ISSR-857, and ISSR-858 generated 100% polymorphism (Table 4). The number of loci observed per primer varied from 3 to 9, with an average of 6.27 loci per primer. The PIC value for ISSR primer ranged from 0.12 to 0.42, with an average of 0.31. The parameters such as H, He, RP, and MI values measured using ISSR primers were: 0.44, 0.29, 6.39, and 2.26, respectively (Table 5). The ISSR primers UBC-810, ISSR-841, and ISSR-827 were found to provide significant molecular markers with the highest resolution power and marker index value. Hence, the ISSR yielded a highly informative marker profile that successfully profiled selected genotypes of Musa spp.
Additionally, SSR markers were also used to gain insights into selected genotypes of banana cultivars. The SSR marker produced polymorphic bands ranging from 1 to 5 loci per primer, and a total of 217 unique amplicons were recorded (Table 5). The SSR marker analysis yielded an average polymorphism of 71.86%. The indexes like H, He, RP, and MI values calculated using SSR primers were 0.19, 0.52, 0.34, 2.72, and 0.47, respectively. Seven locus-specific SSR markers, Ma3/103, CNMPF-8, CNMPF-10, CNMPF-21, CNMPF-16, Ma1/16, and Ma3/2, showed a 100% polymorphic pattern. The highest PIC value of 0.37 was reported by the SSR primer Ma1/16. The SSR primers CNMPF-13, CNMPF-19, and Ma1/16 were significantly powerful in characterizing banana genotypes with the highest RP and MI value.
Identification and assignment of cultivar-specific markers for clonal fidelity analysis
Molecular screening and data analysis provided several polymorphic markers for diploid and triploid genotypes of the banana crop. This study identified several cultivar-specific biomarkers using RAPD, ISSR, and microsatellite markers (Table 6, Fig. S1). We further analyzed the molecular data and assigned the biomarkers specific to the cultivar for clonal fidelity analysis in commercial agriculture. The RAPD primer OPC-08 has shown a particular fragment of 750 bp to Safedvelchi and Elavazhai cultivars (Fig. S1A). At the same time, the RAPD primer OPD-18 has shown a 1.4 kb specific fragment in cultivar Grand Nain, Ardhapuri, and Rasthali (Fig. S1B).
Additionally, the RAPD profile of OPD-20 primer generated a unique band of 1.5 kb for the Neypoovan cultivar (Fig. S1C). The ISSR marker analysis of cultivars Grand Nain and Ardhapuri yielded a specific 0.9 kb fragment during fingerprinting using the ISSR-818 primer (Fig. S1D). The ISSR-818 primer was found to be located in the gene (KU977463.1) Musa acuminata AAA Group Constans-like 3b (COL3b) mRNA, complete CDS with E Value 0.062. Thus ISSR-818 was identified as a genome A specific genic biomarker. The cultivar Ankur yielded a 0.250 kb unique single fragment with an ISSR primer ISSR-827 and UBC-858 (Fig. S1E,G). The Grand Nain cultivar displayed a unique single specific amplicon of 0.7 kb with the primer ISSR-827 and a 0.2 kb amplicon for the primer UBC-858. The ISSR-827 marker yielded 250 bp and 700 bp unique alleles for the cultivars Ankur and Grand Nain, respectively. The banana cultivars Ankur with genome ABB representing M. balbisiana genetic background. Thus ISSR-827 microsatellite marker showed 100% similarity with M. balbisiana microsatellite Mbc08 sequence (0.281 kb; Acc. No. EF467424.1). The ISSR-827 was yielding 0.7 kb unique fragment in banana cultivar Grand Nain to confirm the presence M. acuminata calmodulin-like protein mRNA, partial CDS (765 bp) (AF414128.1) in M. acuminata Grand Nain (AAA) banana cultivar. Thus ISSR-827 yielding ⁓750 bp fragment was identified as a genic biomarker specific to the Grand Nain banana cultivar. The ISSR marker UBC-858 displaying 0.4 kb unique fragment in Ankur and 0.2 kb in Grand Nain is identified as a genic microsatellite biomarker as it was showing similarity with the M. acuminata subsp. malaccensis transcription factor bHLH112 (LOC103998874), transcript variant X2, mRNA, M. acuminata subsp. malaccensis abscisic acid 8'-hydroxylase 3-like (LOC103999378), transcript variant X1, mRNA, M. acuminata subsp. malaccensis indole-3-pyruvate monooxygenase YUCCA2-like (LOC103984381), mRNA etc., with E value 0.023 and confirmed its genic nature. Further, the ISSR-825 profile yielded a specific fragment of 1.0 kb (Fig. S1F) in the banana cultivar Lamby. The ISSR-825 represents the M. acuminata subsp. malaccensis pectinesterase 1-like (LOC103999580), mRNA (XM_009421366.2) (1370 bp), M. acuminata subsp. malaccensis uncharacterized LOC103999495 (LOC103999495), mRNA (1004 bp)( XM_018818029.1). The banana cultivar Ardhapuri produced a specific fragment of 0.5 kb in the DNA fingerprinting analysis using SSR primer CNMPF-8, and a 0.9 kb fragment was observed in the fingerprint of SSR CNMPF-9 (Fig. S1H,I). The CNMPF-8 and CNMPF-9 were identified as genetic markers showing similarity with M. acuminata subsp. burmannicoides var. Calcutta 4 mRNA containing microsatellite, clone MAC4-67K01-F, 595 bp linear mRNA (FM878688.1); and M. acuminata subsp. malaccensis uncharacterized LOC103989204 (LOC103989204), transcript variant X3, mRNA 925 bp linear mRNA (XM_009407999.2). The identified SSR and ISSR biomarkers were genic markers and represented transcripts of the Musa genome (Table 6). The detailed annotation of the SSR and ISSR biomarkers in the banana genome is listed in Table 6. This set of molecular markers could be further developed as the biomarker for the clonal fidelity testing of tissue culture-raised plantlets of cultivars of commercial significance.
Estimation of genetic diversity using molecular clustering
The genetic diversity of bananas has been carried out using different molecular markers, and significant success has been made for analyzing commercial cultivars and some wild germplasm34,35. However, not much is known about the local germplasm, which harbors important traits for adaptation to local climatic conditions and small-holdings. In this regard, genetic diversity analysis was carried out in the diverse banana germplasm. Molecular clustering analysis of 20 banana genotypes based on RAPD fingerprinting resulted in two significant subgroups comprising ten genotypes in each group with 55% similarity (Fig. 1A). Group-I comprised banana genotypes with AA, AB, AAB, ABB, and AAA genomes, including Namarai, Ankur, Karpura Chakrakeli, Rajeli, Ney Poovan, Nendran, Ardhapuri, Grand Nain, Rasthali, and Neyvazhai. Within Group-I, cultivars Karpura Chakrakeli and Rajeli were found to be closely related with 88% of similarity. Group-II consisted of nine banana cultivars carrying AB, BB, ABB, and AAB genomes, which consisted of Safed Velchi, Elavazhai, Bhurkel, Bluggoe, Karpuravalli, Birbutia, Karimbontha, NRCB-3, and Borkal Baista. The cultivars in Group-II were found to have at least 71% similarity level. Whereas the cultivar Lamby was observed to have 37% uniqueness from other cultivars in Group-II.
The phylogenetic analysis based on ISSR fingerprinting generated a dendrogram, which categorized 20 banana genotypes into two major groups in addition to the banana cultivars Namarai (AA) and Ankur (ABB) were clustered separately (Fig. 1A). Group I comprised banana cultivar of genome AB and BB type (Safed Velchi, Bhurkel, Karpuravalli, Borkal Baista, Bluggoe, NRCB-3, Elavazhai, Birbutia, Ney Poovan, Karimbontha and Lamby) with 70% of similarity level; while group II comprised of banana cultivars carrying AAA, AAB and ABB genomes, i.e., Karpura Chakrakeli, Rajeli, Nendran, Ardhapuri, Grand Nain, Rasthali, and Neyvazhai (Fig. 1B).
The clustering analysis helped to obtain a dendrogram based on the SSR fingerprinting data of twenty banana cultivars (Fig. 1C). The SSR analysis divided twenty banana cultivars into two major groups, cultivar like Ankur was grouped separately with 46% dissimilarity level. Group-I comprised of 13 banana cultivars, viz., Namarai, Karpuravalli, Birbutia, Nendran, Safed Velchi, Lamby, Rajeli, Karpura Chakrakeli, Grand Nain, Ney Poovan, Rasthali, NRCB-3, and Borkal Baista with 80% similarity; whereas group II comprised 6 cultivars, i.e., Bhurkel, Bluggoe, Karimbontha, Elavazhai, Neyvazhai, and Ardhapuri with 86% similarity index.
Consequently, we combined the RAPD, ISSR, and SSR fingerprinting data of twenty banana cultivars to construct a dendrogram (Fig. 1D). The combined molecular fingerprinting analysis revealed two major clusters, each comprised of 10 genotypes sharing 62% similarity. The first group contains Namarai, Ankur, Ardhapuri, Grand Nain, Ney Poovan, Karpura Chakrakeli, Rajeli, Neyvazhai, Nendran, and Rasthali. Among the genotypes in group I, Karpura Chakrakeli and Rajeli were genetically closer with 88% similarity, followed by genotypes Ardhapuri and Grand Nain with 81% similarity index. Whereas genotypes Neyvazhai, Namarai, and Ankur were grouped separately from other genotypes at a 47% dissimilarity level, Group-II comprised nine banana genotypes sharing 70% similarity. Lamby’s genotype was separated at a 35% dissimilarity level from other genotypes in group II. The dendrogram obtained after collective clustering described the precise grouping of all twenty banana genotypes than the individual marker-based grouping. The results confirmed the applicability of a three-way molecular marker system in genotyping with higher confidence in estimating cultivar diversity among banana diploid and triploid genotypes.
Principle coordinate (PCA) and population structure analysis (PSA) revealed genetic distance among banana cultivars
The two-and three-dimensional PCA based on genetic variance was performed, which helped to categorize banana germplasm into two distinct groups similar to the groups or clusters obtained during dendrogram analysis of the combined molecular marker system (Fig. 2A,B). PCA showed that the first three Eigenvalues explained 34% of the cumulative variation plotted to reveal diversity among banana germplasm. PCA corroborated cluster analysis results based on RAPD, ISSR, and SSR DNA fingerprinting. The cultivar Lamby was out-grouped and found genetically most distant.
The population structure of 20 banana genotypes was estimated using SSR data. The structure analysis results based on a K-value of 1–10 revealed a peak of dk at K = 4 (Fig. 3A), proposing four major populations (Fig. 3B). The probability of membership threshold (C = 0.44), ten banana genotypes were allocated to SP3 (as 7 pure and 3 admixture), seven genotypes into SP2 (as 5 pure and 3 admixture), two genotypes into SP4, and one genotype into SP1. The pairwise genetic distance among the population was estimated (see Tables 7, 8). The expected heterozygosity between members of the same population was observed as 0.1675 (SP1), 0.0674 (SP2), 0.1367 (SP3), and 0.1505 (SP4). The pairwise genetic differentiation among the four subpopulations was compared based on FST values; the pairwise FST values ranged from 0.171 to 0.555. A high genetic differentiation was observed between SP3 and SP4 with FST value of 0.171. The moderate differentiation was observed between other sub-populations i.e., FST = 0.172 for SP2 and SP3, FST = 0.171 for SP3 and SP4, FST = 0.2342 for SP2 and SP4, FST value 0.3748 for SP2 and SP1, and FST value 0.4987 for SP1 and SP4 (Table 8). Additionally, we performed AMOVA for all 20 banana genotypes considering four sub-populations to support this evidence. A highly significant FST value of 0.23 (p > 0.001) was obtained, suggesting the presence of overall high genetic differentiation among and within populations. Further, it was observed that 23% of genetic variation separated the four sub-populations, and 77% of the total variation existed within the sub-populations (Table 9).
Volatile profiling of banana cultivars
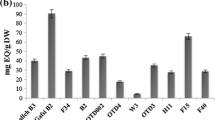
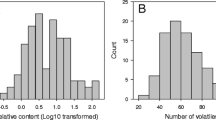
Banana fruits are characterized by specific aroma profiles (volatilome), and considerable effort is needed to investigate aroma profiles during the ripening and post-ripening stages or storage of banana cultivars26,36. In the present study, we performed the volatile compounds profiling to understand the genotypic variation for volatilome of selected banana cultivars of the different genomic constitutions. The volatile compounds identified in banana fruits of nine different cultivars are depicted in Table 10. Maximum volatile compounds were identified in Borkal Baista i.e. 29, followed by 23 each in Rajeli, Ankur, and Karpuravalli, 20 in Bhurkel,16 in Birbutia, 10 in Ardhapuri, and 9 in Grand Nain. A total of fifty-four volatile compounds were identified from fruit samples of nine cultivars of banana and classified into nine unique chemical groups (Fig. 4, Table 10). Almost 56% of the total identified volatile compounds belonged to the ester group, while a minimum of about 2% belonged to carboxylic acid (Fig. 5). The ester compounds were predominantly observed in all the studied banana cultivars (Fig. 5). Among the nine banana cultivars, the highest number of volatile components, i.e., more than 20 i.e., approximately 37%, were recorded in fruits of the cultivars of Borkal Baista, Karpuravalli, Rajeli, Ankur, and Bhurkel. Therefore, these cultivars are suggested to be as rich in aroma and can be utilized for flavor enhancement of banana fruit by selective breeding. The highest number of ester compounds was recorded in Borkal Baista (18), followed by Ankur (17), Karapurvalli (17), Rajeli (12), Bhurkel (10), and Ardhapuri and Grand Nine (7 each) (Table 10). In commercially grown cultivars, Ardhapuri and Grand Nine ester compounds contributed 70–78% of the volatilome suggesting that ester compounds are a major source of banana flavor and aroma and enhance the aesthetic value of the fruit. Another factor that contributed to the aroma was the alcohol groups. We found the distribution of alcohols in Birbutia (5, 31%), Bhurkel (5, 25%), Bluggoe (3, 34%) Rajeli (4, 18%), and Borkal Baista (3, 10%). The alcoholic compounds are also an important component of flavor in the cultivars desired for wine production. Additionally, aldehyde compounds were observed in Bluggoe, Borkal Baista, Bhurkel, Rajeli, and Ankur. Aldehyde compounds have an essential role in aroma37. Moreover, the ketone compounds have an important role in the banana aroma. In the present study, ketones were detected in five banana cultivars (Rajeli, Bhurkel, Karpuravalli, and Birbutia), contributing to the flavor of these cultivars. Although several compounds such as hydrocarbons, carboxylic acid, chlorine-containing ethers, and diverse functional groups were detected in the volatiome analysis, previous studies suggested that these compounds do not contribute to banana aroma26.
Discussion
Genetic diversity and the development of genotype-specific markers are crucial for successful breeding and crop improvement. Banana breeding is a time-consuming and laborious process considering the requirements of time for crop cultivation, field size, and human resources. Therefore, the inclusion of marker-assisted breeding (MAB) can speed up the process of germplasm improvement and make it appealing to develop better performing commercial varieties4,38. Further, it can reduce the number of years consummated from pollination to pre-assessment of first fruit setting and yield, assessing several pollination cycles prior to screening for other biotic and abiotic stresses. Another advantage of MAB is the reduced cost of genetic screening and the wide range of sensing technology that can easily complement phenotyping12.
Chronically several limitations are faced by banana and other horticultural crop breeders as well as farmers from the tropics or subtropics, for example, with the widespread occurrence of misnomers16,17. However, misnomers are frequently grown as commercially cultivars, mislabelled with different names in local languages. To avoid such mislabeling and inappropriate branding, the characterization of misnomers (or synonyms) using molecular markers is essentially required. Thus a quick, simple, reproducible, and cost-effective method for cultivar identification and validation is required, which can be further developed as a certification method for cultured tissue plantlets of bananas. Moreover, it could also help in germplasm preservation and documentation of novel genetic resources39. Therefore, molecular markers hold promise and might help to encourage banana cultivation and smooth trade of authentic genetic material among banana-growing regions like the Marathwada region of Maharashtra, India. The present study offers unique information on molecular markers and volatile profiles for commercially grown cultivars from the Marathwada region, a significant banana cultivation belt within India. This particular region has a growing concern with several misnomers. Hence, a desired molecular marker, a short unique genomic DNA sequence or fragment that carries a conserved inter-specific motif, would be beneficial for appropriate varietal identification. Moreover, such DNA markers possess minimum intra-specific separation to enable the classification of several cultivars and species40; this approach is exploited in different species and cultivars of horticultural crops, including grapes16,17, pineapple41, and figs42. Our results have demonstrated that ISSR primer UBC-858 was found to be unique to Grand Nain, whereas the SSR maker CNMPF-13 was able to confirm the specific loci from the Ardhapuri cultivar, which could be further developed as biomarkers for DNA barcoding the plantlets produced through tissue culture for use in commercial cultivation or breeding programs. Since these DNA markers possessed strong discriminatory power, it could be helpful in the validation of uniformity among the plant population. The study showed that all the phenotypic characteristics in cultivated Musa sp. are geographical location and climate-specific. An earlier analysis of 68 banana accessions performed using RFLP of ITS fragments (digested with RsaI), produced steady, reliable, and distinctive patterns of polymorphic DNA in Musa sp. cultivars43,44. Likewise, ITS1–5.8S-ITS2 sequences45 have also been successfully exploited in molecular diversity analysis of Musaceae.
The aesthetic value of fruits is primarily defined by the constituent volatiles (volatilome), which contribute to the unique aroma and flavor. Moreover, the aroma or flavor of fruits generates significant appeal driving the urge for consumer acceptance, consumption and usage25. The distinctive chemical nature of aroma and flavor are formed by combination of several volatile small molecules. In the case of banana, its agreeable and classic aromatic smell has been studied relatively recently, and there have been over 150 volatiles found to bequeath a unique aroma32.
In the present study, we report over 33% of the volatiles from fruits of nine banana cultivars. This volatiles belonged to several classes of small molecules like esters, ketones, terpenes, and aldehydes. Previously, isoamyl and isobutyl esters and ketones have been reported to be dominant in banana26. In our study, we observed that over 56% contribution to banana flavor is contributed by esters and in cultivar, Grand Nain esters contributed to 78% of the volatiles. Similarly, cultivars namely Ardhapuri, Karpurvalli, Ankur, and Borkee Baista represented significant portion of volatiles belonging to ester compounds. Suggesting dominance of the ester molecules in contribution towards characteristics aroma in these cultivars. Additionally, alcohols and carboxylic compounds were also significant constituents of banana aroma. The SPME method used to extract volatiles is simple, quick, robust, and solvent-free24. The method was efficient for the recovery of volatiles and could be optimized quickly for any fruit volatilome analysis26,32,46. Since some of the volatile compounds are reported to impart resistance against plant pathogens47, additional research is warranted on extensive spatial and temporal volatilome analysis from banana crops to establish the underlying mechanism of volatile biosynthesis and genetic control of resistance against certain pathogens47. In the future, novel technologies such as CRISPR-Cas47,48 and nanotechnological applications49,49,50,52 could help to enhance banana crop improvement.
Conclusions
Since banana is a polyploid and vegetatively propagated horticultural plant, genetic diversity and the development of genotype-specific markers are crucial for successful breeding and crop cultivation. The present study revealed a high polymorphism across the 20 banana genotypes evaluated with 11 RAPD, 11 ISSR, and 12 SSR primers. Phylogenetic and PCA clustered 20 banana genotypes into two significant clusters at 62% similarity level and 34% cumulative variation. AMOVA distributed 23% variation among the populations and 77% within the sub-populations with a significant FST value of 0.23, representing a high level of genetic differentiation. Interestingly, ISSR marker UBC-858 and SSR primer CNMPF-13 yielded a unique fingerprint for Grand Naine and Ardhapuri cultivars, respectively, and could be utilized as cultivar-specific biomarkers for clonal fidelity testing in the tissue culture industry. The volatile profiling revealed the presence of several volatiles in banana cultivars and highlighted the role of ester compounds in the characteristic aroma. Thus, the food processing industry could exploit the contribution of ester-like compounds in banana flavor.
Data availability
The accessions analyzed during the current study are available in the ICAR-NRC Banana repository (https://data.gov.in/resources/icar-publication-repository-date-nrc-banana-collection). The data generated and analyzed are presented in the figures or table of this submission.
References
D’Hont, A. et al. The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488, 213–217 (2012).
Knoema. Bananas production quantity. https://knoema.com/atlas/topics/Agriculture/Crops-Production-Quantity-tonnes/Bananas-production (2022).
Ssebuliba, R. et al. Reproductive efficiency and breeding potential of East African highland (Musa AAA-EA) bananas. Field Crop. Res. 95, 250–255 (2006).
Hippolyte, I. et al. Foundation characteristics of edible Musa triploids revealed from allelic distribution of SSR markers. Ann. Bot. 109, 937–951 (2012).
Mahadev, S. R., Kathithachalam, A. & Marimuthu, M. Efficient protocol for large-scale plantlet production from male floral meristems of Musa spp. cultivars Virupakshi and Sirumalai. In vitro Cell Dev. Biol. Plant 47, 611–617 (2011).
Čížková, J. et al. Molecular analysis and genomic organization of major DNA satellites in banana (Musa spp.). PLoS ONE 8, e54808 (2013).
Sardos, J. et al. DArT whole genome profiling provides insights on the evolution and taxonomy of edible Banana (Musa spp.). Ann. Bot. 118, 1269–1278 (2016).
Lorenzen, J., Hearne, S., Mbanjo, G., Nyine, M. & Close, T. Use of molecular markers in banana and plantain improvement. Acta Hortic. 897, 231–236 (2011).
Ruangsuttapha, S. et al. Molecular phylogeny of banana cultivars from Thailand based on HAT-RAPD markers. Genet. Resour. Crop Evol. 54, 1565–1572 (2007).
Nwakanma, D. C., Pillay, M., Okoli, B. E. & Tenkouano, A. PCR-RFLP of the ribosomal DNA internal transcribed spacers (ITS) provides markers for the A and B genomes in Musa L. Theor. Appl. Genet. 108, 154–159 (2003).
Čížková, J. et al. Molecular and cytogenetic characterization of wild Musa species. PLoS ONE 10, 1–19 (2015).
Perrier, X. et al. Multidisciplinary perspectives on banana (Musa spp.) domestication. Proc. Natl. Acad. Sci. 108, 11311–11318 (2011).
Christelová, P. et al. Molecular and cytological characterization of the global Musa germplasm collection provides insights into the treasure of banana diversity. Biodivers. Conserv. 26, 801–824 (2017).
Bakry, F., Carreel, F., Jenny, C. & Horry, J.-P. Genetic improvement of banana BT—Breeding plantation tree crops: Tropical species. in (eds. Jain, S. M. & Priyadarshan, P. M.) 3–50 (Springer, New York, 2009). https://doi.org/10.1007/978-0-387-71201-7_1.
Ude, G., Pillay, M., Ogundiwin, E. & Tenkouano, A. Genetic diversity in an African plantain core collection using AFLP and RAPD markers. Theor. Appl. Genet. 107, 248–255 (2003).
Upadhyay, A., Kadam, U. S., Chacko, P. M., Aher, L. & Karibasappa, G. Microsatellite analysis to differentiate clones of Thompson seedless grapevine. Indian J. Hortic. 67, 260–263 (2010).
Upadhyay, A., Kadam, U. S., Chacko, P. & Karibasappa, G. Microsatellite and RAPD analysis of grape (Vitis spp.) accessions and identification of duplicates/misnomers in germplasm collection. Indian J. Hortic. 67, 8–15 (2010).
Drapal, M. et al. Metabolite profiling characterises chemotypes of Musa diploids and triploids at juvenile and pre-flowering growth stages. Sci. Rep. 9, 1–10 (2019).
Hřibová, E. et al. The ITS1–5.8S-ITS2 sequence region in the musaceae: Structure, diversity and use in molecular phylogeny. PLoS ONE 6, e17863 (2011).
Pino, J. A. & Febles, Y. Odour-active compounds in banana fruit cv. Giant Cavendish. Food Chem. 141, 795–801 (2013).
Liu, A.-Z., Kress, W. J. & Li, D.-Z. Phylogenetic analyses of the banana family (Musaceae) based on nuclear ribosomal (ITS) and chloroplast (trnL-F) evidence. Taxon 59, 20–28 (2010).
Pino, J. A., Castro-Benítez, M. & Winterhalter, P. Analysis of volatile compounds in baby banana peel and pulp (musa Acuminata AA Simmonds Cv. Bocadillo) in relation to the hyperpigmentation phenomenon. Int. J. Fruit Sci. 20, 403–413 (2020).
Deng, G. et al. Transcriptome and metabolome profiling provide insights into molecular mechanism of pseudostem elongation in banana. BMC Plant Biol. 21, 1–14 (2021).
Majcher, M. & Jeleń, H. H. Comparison of suitability of SPME, SAFE and SDE methods for isolation of flavor compounds from extruded potato snacks. J. Food Compos. Anal. 22, 606–612 (2009).
Thaiphanit, S. & Anprung, P. Physicochemical and flavor changes of fragrant banana (Musa acuminata aaa group “Gross Michel”) during ripening. J. Food Process. Preserv. 34, 366–382 (2010).
Zhu, X. et al. Comparative study of volatile compounds in the fruit of two banana cultivars at different ripening stages. Molecules 23, 2456 (2018).
Gawel, N. J. & Jarret, R. L. A modified CTAB DNA extraction procedure for Musa andIpomoea. Plant Mol. Biol. Rep. 9, 262–266 (1991).
Nei, M. Analysis of gene diversity in subdivided populations. Proc. Natl. Acad. Sci. 70, 3321–3323 (1973).
Sneath, P. H. A. & Sokal, R. R. Numerical Taxonomy. The Principles and Practice of Numerical Classification. (1973).
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332 (2009).
Earl, D. & vonHoldt, B. M. Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361 (2011).
Facundo, H. V. D. V., Garruti, D. S., Cordenunsi, B. R. & Lajolo, F. M. Isolation of volatiles compounds in banana by HS-SPME: optimization for the whole fruit and pulp. Int. J. Biosci. Biochem. Bioinform. 3, 110–115 (2013).
Hickey, L. T. et al. Breeding crops to feed 10 billion. Nat. Biotechnol. 37, 744–754 (2019).
Borborah, K. et al. Comparative analysis of genetic diversity in some non-commercial cultivars of Musa L. from Assam, India, using morphometric and ISSR markers. Int. J. Fruit Sci. 20, 1814–1828 (2020).
Mertens, A. et al. Genetic diversity and structure of Musa balbisiana populations in Vietnam and its implications for the conservation of banana crop wild relatives. PLoS ONE 16, e0253255 (2021).
Esteras, C. et al. Fruit flesh volatile and carotenoid profile analysis within the Cucumis melo L. species reveals unexploited variability for future genetic breeding. J. Sci. Food Agric. 98, 3915–3925 (2018).
Beekwilder, J. et al. Functional characterization of enzymes forming volatile esters from strawberry and banana. Plant Physiol. 135(4), 1865–1878. https://doi.org/10.1104/pp.104.042580 (2004). PMID: 15326278; PMCID: PMC520758.
Christelová, P. et al. A platform for efficient genotyping in Musa using microsatellite markers. AoB Plants 2011, plr024 (2011).
Uma S, Sathiamoorthy S, D. S. Banana Indian Genetic Resources and Catalogue. (National Research Center for Banana (NRCB) India (2005).
Němečková, A. et al. Molecular and cytogenetic study of east african highland banana. Front. Plant Sci. 9, 1–13 (2018).
Hidayat, T., Abdullah, F. I., Kuppusamy, C., Samad, A. A. & Wagiran, A. Molecular identification of malaysian pineapple cultivar based on internal transcribed spacer region. APCBEE Proc. 4, 146–151 (2012).
Castro, C., Hernandez, A., Alvarado, L. & Flores, D. DNA barcodes in fig cultivars (Ficus carica L.) using ITS regions of ribosomal DNA, the psbA-trnH spacer and the matK coding sequence. Am. J. Plant Sci. 06, 95–102 (2015).
Javed, M. A., Chai, M. A. K. & Othman, R. Y. Morphological characterization of malaysian wild banana Musa acuminata. Biotropia (Bogor) 18, 21–37 (2002).
Bhat, K. V., Jarret, R. L. & Rana, R. S. DNA profiling of banana and plantain cultivars using random amplified polymorphic DNA (RAPD) and restriction fragment length polymorphism (RFLP) markers. Electrophoresis 16, 1736–1745 (1995).
Hapsari, L. & Ayu Lestari, D. Fruit characteristic and nutrient values of four Indonesian banana cultivars (Musa spp.) at different genomic groups. Agrivita 38, 303–311 (2016).
Zhang, H. et al. Characterization of volatile profiles and marker substances by HS-SPME/GC-MS during the concentration of coconut jam. Foods 9, 347 (2020).
Hinge, V. R., Chavhan, R. L., Kale, S. P., Suprasanna, P. & Kadam, U. S. Engineering resistance against viruses in field crops using CRISPRCas9. Curr. Genomics 22, 214–231 (2021).
Kadam, U. S., Shelake, R. M., Chavhan, R. L. & Suprasanna, P. Concerns regarding ‘off-target’ activity of genome editing endonucleases. Plant Physiol. Biochem. 131, 22–30 (2018).
Kadam, U. S., Lossie, A. C., Schulz, B. & Irudayaraj, J. Gene expression analysis using conventional and imaging methods. DNA and RNA Nanobiotechnologies in Medicine: Diagnosis and Treatment of Diseases. 141–162 (Springer, Berlin, 2013). https://doi.org/10.1007/978-3-642-36853-0_6.
Kadam, U. S., Schulz, B. & Irudayaraj, J. M. K. K. Multiplex single-cell quantification of rare RNA transcripts from protoplasts in a model plant system. Plant J. 90, 1187–1195 (2017).
Kadam, U. S., Chavhan, R. L., Schulz, B. & Irudayaraj, J. Single molecule Raman spectroscopic assay to detect transgene from GM plants. Anal. Biochem. 532, 60–63 (2017).
Kadam, U., Moeller, C. A., Irudayaraj, J. & Schulz, B. Effect of T-DNA insertions on mRNA transcript copy numbers upstream and downstream of the insertion site in arabidopsis thaliana explored by surface enhanced raman spectroscopy. Plant Biotechnol. J. 12, 568–577 (2014).
Author information
Authors and Affiliations
Contributions
V.R.H., R.L.C., and U.S.K.: conceived and designed the experiments; V.R.H., I.M.S., and A.S.D.: performed all experiments; V.R.H., R.L.C., P.S., and U.S.K.: provided scientific advice and resources. V.R.H., R.L.C., R.M.S., S.A.G., A.M.D., P.S., U.S.K.: wrote the manuscript and prepared figures. All authors reviewed and read the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hinge, V.R., Shaikh, I.M., Chavhan, R.L. et al. Assessment of genetic diversity and volatile content of commercially grown banana (Musa spp.) cultivars. Sci Rep 12, 7979 (2022). https://doi.org/10.1038/s41598-022-11992-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-11992-1
- Springer Nature Limited
This article is cited by
-
Assessment of genetic diversity and population structure of common walnut (Juglans regia) germplasm with simple sequence repeat (SSR) markers
Genetic Resources and Crop Evolution (2024)
-
Correlation of released gases and quality classification of bananas during storage
Journal of Food Measurement and Characterization (2024)
-
Genetic diversity, population structure and selection signatures in Enset (Ensete ventricosum, (Welw.) Cheesman), an underutilized and key food security crop in Ethiopia
Genetic Resources and Crop Evolution (2023)
-
An efficient embryogenic cell suspension culture system through secondary somatic embryogenesis and regeneration of true-to-type plants in banana cv. Sabri (silk subgroup AAB)
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)