Abstract
Mycoplasma gallisepticum (MG) is one of the most economically important pathogens worldwide. MG affects the respiratory system and impairs growth performance in poultry. In developing countries, the most widely used technique to identify MG is the conventional PCR assay. In this study, 24 MG isolates collected from Thailand farms with unvaccinated chickens during 2002–2020 were characterized by gene-targeted sequencing (GTS), followed by phylogenetic analysis using unweighted pair group method with arithmetic mean. These 24 Thai MG isolates differed from vaccine strains, including the F, ts-11 and 6/85 strains. One isolate showed 99.5–100% genetic similarity to the F strain with 4 partial gene analyses. This result may have been due to contamination from vaccinated flocks because the F strain is the most commonly used vaccine strain in Thailand. However, the GTS analysis using the partial MG genes in this study showed that the isolates could be grouped into different patterns based on individual gene sequences. The phylogenetic analysis of partial mgc2, gapA, pvpA and lp gene sequences classified the Thai MG isolates into 7, 11, 7 and 2 groups, respectively. In conclusion, at least 2 partial MG genes, especially partial gapA and mgc2 genes, are needed to differentiate MG isolates.
Similar content being viewed by others
Introduction
Mycoplasma gallisepticum (MG) remains one of the most important bacterial pathogens worldwide, causing a respiratory disease called chronic respiratory disease (CRD) in infected poultry flocks and resulting in monetary losses for treatment and control1. MG has both horizontal and vertical transmission. MG infection can cause a high feed conversion ratio, egg production loss, poor hatchability, and carcass degradation1. Stipkovits and Kempf2 investigated the economic loss from MG and found a 10–20% drop in egg production in infected layers and a body weight loss of 10–20% in infected broilers. In Thailand, approximately 25% of all laying hens in the poultry industry are infected with MG, leading to a loss of approximately 15 million U.S. dollars due to a decrease in egg production3.
Due to this widespread MG infection, vaccination is an important preventive strategy generally used in the Thailand poultry industry. Live vaccine strains, including the F, ts-11, and 6/85 strains, and inactivated MG vaccines have been used for years1. In particular, the F strain is one of the most effective vaccine strains and is widely used in Thailand. Therefore, a technique to differentiate between vaccine and field MG strains in flocks with suspected MG infection is needed. Several studies have investigated techniques for MG classification4,5,6,7,8,9. For example, gene targeted sequencing (GTS) analysis was developed by Ferguson et al.4 This technique has been used to determine the gene sequences of partial surface proteins of MG, including the gapA, mgc2, pvpA and MGA_0319 genes. The multilocus sequence typing scheme (MLST) is a technique that many studies have used and is regarded as the gold standard for bacterial typing6,10,11. This technique uses MG housekeeping genes for molecular identification, which is an effective way to determine the relationship between MG strains. Both GTS and MLST have been widely used to monitor and characterize MG strains6. Additionally, the whole genome sequence (WGS) can be used to analyse the entire genomic sequence of MG12,13. High-resolution melt (HRM) curve analysis is another new molecular technique that classifies MG strains by using the vlhA, pvpA, gapA, and mgc2 genes as well as the 16S-23S rRNA intergenic space region (IGSR) with conventional and real-time PCR8,9. The most commonly used technique in Thailand is random amplification of polymorphic DNA (RAPD). However, RAPD has low reproducibility, and results from different laboratories cannot be compared14,15. Sequencing is a potential technique for MG classification. MG strains can be differentiated with partial DNA sequences and compared among laboratories in different areas or countries4,16,17. In addition, gene targeted sequencing (GTS) is a cost-efficient and affordable method for use in developing countries, including Thailand, where advanced techniques are not generally feasible.
The important genes of MG, including gapA, mgc2, pvpA and MGA_0319 (lp), have been investigated in several epidemiological studies4,18,19. In Thailand, Limsatanun et al.20 classified MG strains with partial mgc2 gene sequences; thus, the partial mgc2 gene can be used to classify Thai MG strains from vaccine strains and various strains from different countries. However, partial mgc2 gene classification is not a reliable method for MG characterization4.
The aim of this study was to determine a GTS technique for differentiating field and vaccine MG strains in commercial chicken flocks from different regions in Thailand. This is the first study to use 4 partial MG gene sequences for commercial MG classification in Thailand.
Results
PCR amplification
All twenty-four Thai MG isolates were detected by MG-specific PCR amplification following the Lauerman method21. To amplify partial mgc2 genes, which were 615 bp in size, 22 Thai MG isolates were successfully amplified and sequenced. According to the specific partial gapA PCR with 306 bp, 21 Thai MG isolates were positive and included in the phylogenetic analysis, while 20 samples of Thai MG isolates were successfully amplified using the pvpA and MGA_0319 (lp) primers with lengths of 456 and 495 bp, respectively. All nucleotide sequences from Thai MG isolates in this study were submitted to GenBank and given accession numbers (Table 1).
Phylogenetic analysis
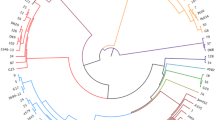
The phylogenetic tree based on the partial mgc2 gene demonstrated that 3 Thai MG isolates were closely related to the F strain. AHRU/2014/CU4508.1 was grouped together with the F strain, while AHRU/2020/CU0143.1 and AHRU/2020/0147.1 showed 97.6% genetic similarity to the F strain (Fig. 1). According to the phylogenetic tree based on the partial gapA gene, AHRU/2014/CU4508.1 showed 99.5% genetic similarity to the F strain. AHRU/2020/CU3704.1 was also grouped with the F strain (Fig. 2). The phylogenetic analysis of the partial pvpA gene placed all Thai MG isolates in the same cluster except the reference strain S6 (Fig. 3). Four Thai MG isolates showed 94.3% genetic similarity to the 6/85 strain. AHRU/2014/CU450 8.1 was grouped with the F strain with 100% similarity. The partial lp gene sequences of Thai MG were compared with reference strains. The 6/85 strain was grouped into different Clusters. AHRU/2014/CU4508.1 and AHRU/2020/CU0143.1 had 100% genetic similarity to the F strain and 99.2% genetic similarity to the ts-11 strain (Fig. 4). The genetic similarity of Thai MG strains and F strain is shown in Table 2. The lp gene showed the highest similarity of genetic sequences (98.2–100%) between the F strain and Thai MG strains. The phylogenetic trees with DNA sequence data are available in the Supplementary Information.
Discussion
Avian mycoplasmosis is an important disease-causing pathogen in the poultry industry with substantial economic impacts. Live, inactivated, and recombinant MG vaccines have been used in Thailand for a long time. Due to the increased use of MG vaccines, differentiation between field and vaccine strains is needed. Molecular characterization of MG has been investigated in many countries4,5,7,22,23. This study is the first to use the GTS technique on Thai MG strains with 4 partial MG gene sequences. The partial mgc gene has been used for MG characterization in many epidemiological studies6,20,24. It encodes the MGC2 protein, which coordinates with the gapA gene-encoded protein for cell attachment25 and is involved in MG immunogenicity12,26.
In a previous study, Armour et al.22 investigated MG isolates from South Africa using intergenic spacer regions (IGSRs), mgc2 and gapA genes. Thirty-six MG isolates were classified into 8 types by the mgc2 gene and 2 types by the gapA gene. Thus, the mgc2 gene had a higher discriminatory power than the gapA gene. Another study in Russia conducted an epidemiological investigation of MG7. The results showed that mgc2 gene had good discriminatory power, while gapA did not provide a good discriminatory index for MG classification. However, the use of only a single gene for classification could not determine the similarity between MG isolates. Additionally, some MG isolates were negative for the mgc2 gene according to the PCR assay, resulting in a failure to obtain mgc2 sequences5,6,19,22; thus, using only one partial gene sequence is insufficient to characterize MG.
In the present study, the lp gene of Thai MG isolates was more conserved than the gapA, mgc2 and pvpA genes, as 18 out of 20 Thai MG isolates showed 100% genetic similarity on this gene. The use of partial gapA showed the highest genetic variation among Thai isolates. These results contradicted those of previous studies4,7,22, which indicated that MG isolates from the same area would have lower genetic diversity than MG isolates from different regions22. In the present study, Thai MG isolates were identified with 4 genes using the phylogenetic tree (UPMGA) method. AHRU/2014/CU4508.1 had the closest genetic relationship to the F strain. The UPMGA results showed that AHRU/2014/CU4508.1 was grouped with the F strain on all 4 partial gene analyses. Interestingly, all Thai MG isolates in this study were collected from farms with unvaccinated flocks. In Thailand, poultry breeders and layers are widely vaccinated with the F strain. Interestingly, the AHRU/2014/CU4508.1 isolate from these farms might have been contaminated from other farms with vaccinated flocks. The F strain from the live MG vaccine can be transmitted both horizontally and vertically27,28,29. Furthermore, several epidemiological studies have shown that the F strain can cause MG outbreaks if it spreads from vaccinated to nonvaccinated flocks5,24,30. Other Thai MG isolates in this study varied in genetic classification depending on the gene analysed. The results of gapA and mgc2 gene analysis showed that AHRU/2003/CU5113.2 and AHRU/2003/CU5808.2 were grouped with the S6 strain with 97% and 99.4% genetic similarity, respectively. In contrast, using the partial pvpA gene sequence indicated that the S6 strain was separated from all Thai MG isolates, including AHRU/2003/CU5113.2 and AHRU/2003/CU5808.2. These results indicated that AHRU/2003/CU5113.2 and AHRU/2003/CU5808.2 might be genetically related to the S6 strain. DNA sequences of all 4 virulence genes could not be obtained for some Thai MG isolates. For example, the Thai MG isolate AHRU/2009/CU3704 could only be classified by phylogenetic analysis of gapA and pvpA genes because it was negative for mgc2 and pvpA according to the PCR analysis. Plausibly, this lack of detection could be because of the poor quality of DNA due to the presence of multiple strains in the broth medium sample and/or genetic mutations between and within MG strains6,31,32.
In conclusion, the Thai MG isolates in this study could be differentiated with partial MG genes, including the gapA, mgc2, pvpA and MGA_0319 (lp) genes. All Thai MG isolates could be classified with at least 2 out of 4 partial gene sequences, especially the partial gapA and mgc2 genes, which had satisfactory discriminatory power for Thai MG characterization. Using partial DNA sequencing for MG characterization is an effective and reproducible method for establishing the genetic relationship between MG strains and differentiating between vaccine and field strains. In addition, this study was the first epidemiological study of Thai MG strains to use 4 partial MG gene sequences, demonstrating the genetic diversity of circulating MG strains in Thailand. In future studies, the GTS technique should be implemented along with other molecular techniques, including a multilocus sequence typing scheme, to provide more epidemiological and evolutionary data and improve the system for monitoring MG outbreaks in poultry farms in Thailand.
Materials and methods
MG isolates
Twenty-four Thai MG isolates were used in this study. All isolates were collected during 2003–2020 by Prof. Somsak Pakpinyo, Department of Veterinary Medicine, Faculty of Veterinary Science, Chulalongkorn University. All MG isolates were collected from choanal cleft of dead chicken and were propagated in FMS medium supplemented with 15% swine serum following previously reported methods33 and incubated at 37 °C until the broth colour changed from pink to orange. All isolates were confirmed as MG by polymerase chain reaction (PCR) assay21.
Molecular typing
The DNA from each Thai MG isolate was extracted with an equal volume of phosphate buffered saline (PBS) and then amplified by PCR. The primers in this study were designed by Ferguson4 (Table 3). A PCR assay was performed to detect the partial gapA, pvpA, MGA_0319 and mgc2 genes. The PCR mixture consisted of 500 mM KCl, 100 mM Tris–HCl (pH 8.3), 1.25 mM MgCl2, 1 mM dNTP (Thermo Scientific, Vilnius, Lithuania), 10 pmol each of primer (Qiagen®, Valencia, CA, USA), 1.25 µl of Taq polymerase (Promega, Madison, WI, USA) and 2.5 µl (125 ng) of the DNA template. The amplification reaction was performed in a DNA thermal cycler at 94 °C for 3 min, followed by 40 cycles of 94 °C for 20 s, 55–60 °C for 40 s, 72 °C for 60 s, and 72 °C for 5 min for the gapA, pvpA, MGA_0319 (lp) and mgc2 genes. The PCR products were 332, 702, 590 and 824 bp, respectively4.
Reference sequences
Four reference strains were used in this study. The F strain was the vaccine strain provided by a local distributor (MSD, Thailand). The S6 strain was obtained from ATCC (15302). The ts-11 and 6/85 strain sequences were obtained from Prof. Somsak Pakpinyo, Department of Veterinary Medicine, Faculty of Veterinary Science, Chulalongkorn University.
DNA sequence analysis
Amplified PCR products of MG-targeted gene-positive extracts were submitted to determine the sequence. Partial mgc2 gene sequences (Accession Numbers KX268616–KX268632) from 16 Thai MG isolates had been submitted to GenBank in a previous study20 (Table 3.) All sequences were analysed with the Editseq program (Lasergene, DNASTAR Inc., USA), and a consensus was constructed with the Seqman program (Lasergene, DNASTAR Inc., USA). Thai MG isolates and reference gene sequence data were aligned to construct a phylogenetic tree in Bionumeric version 7.6 software (Applied Maths, Sint-Martens-Latem, Belgium). Cluster analysis was performed with the UPGMA method. The similarity coefficients of Thai MG isolates and reference strains were determined from multiple sequence alignments.
Data availability
All data generated or analysed during this study are included in this published article [and its Supplementary Information files]. The sequence data are available at the NCBI Nucleotide (https://www.ncbi.nlm.nih.gov/nuccore); see Table 3 for sample accession numbers.
References
Armour, N. K. in Diseases of poultry (ed Swayne D. E. et al.) 911–923 (Wiley-Blackwell, 2020).
Stipkovits, L. & Kempf, I. Mycoplasmoses in poultry. Rev. Sci. Tech. 15, 1495–1525. https://doi.org/10.20506/rst.15.4.986 (1996).
Pakpinyo, S. in Important mycoplasmosis in Thailand (โรคติดเชื้อมัยโคพลาสมาที่สำคัญของสัตว์ปีก) (ed Pakpinyo S.) 1–11 (WorkDee Idea, 2017).
Ferguson, N. M. et al. Use of molecular diversity of Mycoplasma gallisepticum by gene-targeted sequencing (GTS) and random amplified polymorphic DNA (RAPD) analysis for epidemiological studies. Microbiology 151, 1883–1893. https://doi.org/10.1099/mic.0.27642-0 (2005).
Gharaibeh, S., Laibinis, V., Wooten, R., Stabler, L. & Ferguson-Noel, N. Molecular characterization of Mycoplasma gallisepticum isolates from Jordan. Avian Dis. 55, 212–216. https://doi.org/10.1637/9526-091510-Reg.1 (2011).
Matucci, A. et al. Molecular differentiation of Mycoplasma gallisepticum outbreaks: A last decade study on Italian farms using GTS and MLST. Vaccines-Basel https://doi.org/10.3390/vaccines8040665 (2020).
Sprygin, A. V. et al. Biological characterization of Russian Mycoplasma gallisepticum field isolates. Avian Pathol. 40, 213–219. https://doi.org/10.1080/03079457.2011.554795 (2011).
Ghorashi, S. A., Bradbury, J. M., Ferguson-Noel, N. M. & Noormohammadi, A. H. Comparison of multiple genes and 16S–23S rRNA intergenic space region for their capacity in high resolution melt curve analysis to differentiate Mycoplasma gallisepticum vaccine strain ts-11 from field strains. Vet. Microbiol. 167, 440–447. https://doi.org/10.1016/j.vetmic.2013.09.032 (2013).
Ghorashi, S. A., Noormohammadi, A. H. & Markham, P. F. Differentiation of Mycoplasma gallisepticum strains using PCR and high-resolution melting curve analysis. Microbiology 156, 1019–1029. https://doi.org/10.1099/mic.0.031351-0 (2010).
Beko, K. et al. Genotyping Mycoplasma gallisepticum by multilocus sequence typing. Vet. Microbiol. 231, 191–196. https://doi.org/10.1016/j.vetmic.2019.03.016 (2019).
Ghanem, M. & El-Gazzar, M. Development of a multilocus sequence typing assay for Mycoplasma gallisepticum. Avian Dis. 63, 693–702. https://doi.org/10.1637/aviandiseases-D-19-00072 (2019).
Papazisi, L. et al. The complete genome sequence of the avian pathogen Mycoplasma gallisepticum strain R-low. Microbiol-Sgm 149, 2307–2316. https://doi.org/10.1099/mic.0.26427-0 (2003).
Song, Y. et al. Complete genome sequence of Mycoplasma gallisepticum strain KUVMG001, an isolate from South Korea. Microbiol. Resour. Announc. https://doi.org/10.1128/MRA.00331-21 (2021).
Tyler, K. D., Wang, G., Tyler, S. D. & Johnson, W. M. Factors affecting reliability and reproducibility of amplification-based DNA fingerprinting of representative bacterial pathogens. J. Clin. Microbiol. 35, 339–346. https://doi.org/10.1128/jcm.35.2.339-346.1997 (1997).
Mettifogo, E. et al. Molecular characterization of MG isolates using RAPD and PFGE isolated from chickens in Brazil. J. Vet. Med. B 53, 445–450. https://doi.org/10.1111/j.1439-0450.2006.00978.x (2006).
Raviv, Z. et al. The Mycoplasma gallisepticum 16S–23S rRNA intergenic spacer region sequence as a novel tool for epizootiological studies. Avian Dis. 51, 555–560. https://doi.org/10.1637/0005-2086(2007)51[555:Tmgsri]2.0.Co;2 (2007).
Ricketts, C. et al. Identification of strain-specific sequences that distinguish a Mycoplasma gallisepticum vaccine strain from field isolates. J. Clin. Microbiol. 55, 244–252. https://doi.org/10.1128/Jcm.00833-16 (2017).
Liu, T., Garcia, M., Levisohn, S., Yogev, D. & Kleven, S. H. Molecular variability of the adhesin-encoding gene pvpA among Mycoplasma gallisepticum strains and its application in diagnosis. J. Clin. Microbiol. 39, 1882–1888. https://doi.org/10.1128/Jcm.39.5.1882-1888.2001 (2001).
Felice, V. et al. Molecular detection and characterization of Mycoplasma gallisepticum and Mycoplasma synoviae strains in backyard poultry in Italy. Poultry Sci. 99, 719–724. https://doi.org/10.1016/j.psj.2019.12.020 (2020).
Limsatanun, A., Sasipreeyajan, J. & Pakpinyo, S. Comparison of GTS and RAPD assays to characterize Thai Mycoplasma gallisepticum strains. Thai J. Vet. Med. 48, 331–337 (2018).
Lauerman, L. H. in Nucleic acid amplification assays for diagnosis of animal disease (ed Lauerman L. H.) 41–42 (American Association of Veterinary Laboratory Diagnosticians, 1998).
Armour, N. K., Laibinis, V. A., Collett, S. R. & Ferguson-Noel, N. The development and application of a Mycoplasma gallisepticum sequence database. Avian Pathol. 42, 408–415. https://doi.org/10.1080/03079457.2013.819486 (2013).
Gerchman, I., Levisohn, S., Mikula, I., Manso-Silvan, L. & Lysnyansky, I. Characterization of in vivo-acquired resistance to macrolides of Mycoplasma gallisepticum strains isolated from poultry. Vet. Res. 42, 90. https://doi.org/10.1186/1297-9716-42-90 (2011).
Khalifa, R., Eissa, S., El-Hariri, M. & Refai, M. Sequencing analysis of Mycoplasma gallisepticum wild strains in vaccinated chicken breeder flocks. J. Mol. Microb. Biotech. 24, 98–104. https://doi.org/10.1159/000357733 (2014).
Boguslavsky, S. et al. Molecular characterization of the Mycoplasma gallisepticum pvpA gene which encodes a putative variable cytadhesin protein. Infect. Immun. 68, 3956–3964. https://doi.org/10.1128/IAI.68.7.3956-3964.2000 (2000).
Hnatow, L. L., Keeler, C. L. Jr., Tessmer, L. L., Czymmek, K. & Dohms, J. E. Characterization of MGC2, a Mycoplasma gallisepticum cytadhesin with homology to the Mycoplasma pneumoniae 30-kilodalton protein P30 and Mycoplasma genitalium P32. Infect. Immun. 66, 3436–3442. https://doi.org/10.1128/IAI.66.7.3436-3442.1998 (1998).
Kleven, S. H. Transmissibility of the F strain of Mycoplasma gallisepticum in leghorn chickens. Avian Dis. 25, 1005–1018 (1981).
Lin, M. Y. & Kleven, S. H. Egg transmission of two strains of Mycoplasma gallisepticum in chickens. Avian Dis. 26, 487–495 (1982).
Glisson, J. R. & Kleven, S. H. Mycoplasma gallisepticum vaccination: Effects on egg transmission and egg production. Avian Dis. 28, 406–415 (1984).
Ley, D. H., Avakian, A. P. & Berkhoff, J. E. Clinical Mycoplasma gallisepticum infection in multiplier breeder and meat turkeys caused by F strain: identification by sodium dodecyl sulfate-polyacrylamide gel electrophoresis, restriction endonuclease analysis, and the polymerase chain reaction. Avian Dis. 37, 854–862 (1993).
Delaney, N. F. et al. Ultrafast evolution and loss of CRISPRs following a host shift in a novel wildlife pathogen Mycoplasma gallisepticum. PLoS Genet 8, e1002511. https://doi.org/10.1371/journal.pgen.1002511 (2012).
Szczepanek, S. M. et al. Comparative genomic analyses of attenuated strains of Mycoplasma gallisepticum. Infect. Immun. 78, 1760–1771. https://doi.org/10.1128/IAI.01172-09 (2010).
Kleven, S. H. in A laboratory manual for the isolation and identification of avian pathogens (ed Swayne D. E. et al.) 74–80 (American Association of Avian Pathologists, 1998).
Acknowledgements
This research was financially supported by the Office of the Permanent Secretary, Ministry of Higher Education, Science, Research and Innovation Grant No. RGNS 63-227.
Author information
Authors and Affiliations
Contributions
A.L. conceptualized and designed the study, performed the experiment and carried out all analyses, interpreted the results and drafted the manuscript; T.P. constructed the phylogenetic trees; K.L. collected swab sample data; and S.P. supervised the study and reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Limsatanun, A., Pakpinyo, S., Limpavithayakul, K. et al. Targeted sequencing analysis of Mycoplasma gallisepticum isolates in chicken layer and breeder flocks in Thailand. Sci Rep 12, 9900 (2022). https://doi.org/10.1038/s41598-022-14066-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-14066-4
- Springer Nature Limited