Abstract
Colletotrichum sublineola is a destructive fungal pathogen that causes anthracnose in sorghum. Senegalese sorghum germplasm is currently being considered as an option of sources for genetic resistance. In a recent study, Senegalese sorghum accessions were evaluated for response to a mixture of Texas isolates of C. sublineola at the 8-leaf stage in the greenhouse. As a comparison, 159 Senegalese sorghum accessions at the 1-leaf developmental stage were evaluated against a single Texas isolate of C. sublineola (FSP53) using an excised-leaf assay. A genome-wide association study (GWAS) was conducted based on the phenotypic data acquired to discover genetic variation associated with response to C. sublineola using 193,727 single nucleotide polymorphisms (SNPs) throughout the genome. Sorghum seedlings tended to be more resistant when compared with sorghum plants inoculated at the 8-leaf stage in the greenhouse in previous experiments. Based on the highest score evaluated in the 1-leaf developmental stage excised leaf assay for each accession, 16 accessions were labeled as susceptible. GWAS identified the SNP locus S01_72868925 that is associated with protein kinase domain // Leucine rich repeat N-terminal domain at a level of confidence that surpassed Bonferroni correction. Along with the SNP locus S01_72868925, other top SNP loci were also associated with genes that are known to play critical roles in plant defense or plant stress responses.
Similar content being viewed by others
Introduction
Sorghum [Sorghum bicolor (L.) Moench] is a multipurpose food crop which is ranked among the top five cereal crops in the world1. Like other important cereal crops, sorghum is consistently challenged by biotic and abiotic stresses that constrain overall productivity2. Sorghum anthracnose caused by Colletotrichum sublineola Henn. ex Sacc. & Trotter 1913 is an important disease of cultivated sorghum worldwide3. In recent years, genome-wide association studies (GWAS) have identified potentially important defense-related genes against C. sublineola in various sorghum germplasms4,5,6. As examples, genes that encode an F-box, a protein tyrosine kinase, a leucine rich repeat and a peroxidase were identified as top candidates for anthracnose resistance in the sorghum association panel (SAP) lines4. Similar studies identified genes whose products include motifs such as pentatricopeptide repeat and a leucine-rich repeat (LRR) as top candidates in the same SAP lines5.
In the sorghum mini core collection, genes that encode proteins with a zinc finger domain, an F-box domain, exodeoxyribonuclease VII, and ubiquitin-conjugating enzyme were listed as top candidate defense related genes6. Many germplasm lines from West and Central Africa where sorghum is cultivated in rainy and high humidity regions have been shown to be important sources of resistance genes to fungal diseases7. In a recent study, 163 Senegalese sorghum accessions were scored for responses to a mixture of eight Texas isolates (FSP2, FSP5, FSP7, FSP35, FSP36, FSP46, FSP50 and FSP53) of C. sublineola8. Plants of the Senegalese accessions were grown in a greenhouse, inoculated at the 8-leaf stage, and the subsequent GWAS analysis identified genes that encode leucine rich repeat // protein tyrosine kinase // leucine rich repeat N-terminal domain, selenium binding protein and zinc finger as genes highly associated with defense against Texas isolates of C. sublineola8. Sorghum seedlings contain the cyanogenic glycoside dhurrin, which may play a role in seedling protection9, and a phytoalexin pigment complex accumulates rapidly in sorghum seedlings when inoculated to C. sublineola (formerly known as C. graminicola (Ces.) G.W. Wils) 9,10. Therefore, compared to 8-leaf stage of sorghum, seedlings may show greater resistance against C. sublineola that can easily skew phenotypic data toward resistance. Previously however, comparisons between anthracnose inoculations made at juvenile and later growth stages of sorghum have not been studied in sorghum.
To compare the response of sorghum plants inoculated at the 8-leaf stage, 159 Senegalese sorghum accessions were evaluated at the 1-leaf stage for response to a Texas isolate of C. sublineola through an excised-leaf assay11. All 159 accessions were the same as those included in the previous study for greenhouse survey at the 8-leaf stage8 (insufficient seed was available for the other 4 lines). It is hypothesized that Senegalese sorghum accessions would respond differently when inoculated at 1-leaf seedling stage compared to conventional 8-leaf stage. Moreover, it is hypothesized that an excised-leaf assay11, originally designed to be used in sorghum at the 8-leaf stage, can be practical for earlier screening of sorghum seedlings. By applying GWAS analysis to disease response based on an excised-leaf assay on sorghum seedlings, it was also hypothesized that it is possible to identify defense related genes that are particularly important at seedling stage against anthracnose. As in the previous study, the disease score results were combined with a GWAS analysis that includes 193,727 single-nucleotide polymorphic (SNP) loci from a publicly available genotype-by-sequencing (GBS) dataset for the Senegalese accessions8. GWAS analysis was conducted by using TASSEL12 version 5.2.80 based on the highest score for disease ratings in each accession; candidate genes that might contribute to resistance or susceptibility to anthracnose in Senegalese sorghum seedlings are reported.
Results
Phenotypic variation
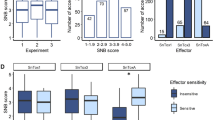
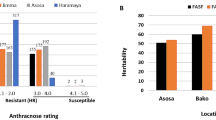
Senegalese seedlings overall showed strong resistance to FSP53; average scores based on all 9 1st leaves for all accessions were between 1 and 1.78 including resistant and susceptible checks. However, the accessions showed considerable variation based on the highest scores among 9 tested 1st leaves. Excluding resistant and susceptible checks, 82 accessions scored 1, and 61 accessions scored 2. Six accessions (PI514284, PI514298, PI514299, PI514372, PI514455 and PI514459) scored 3, and 3 accessions (PI514427, PI514449 and PI514474) scored 4. Lastly, 7 accessions (PI514293, PI514300, PI514448, PI514457, PI514467, PI514473 and PI514478) had at least one 1st leaf with a score of 5. The susceptible checks BTx623, PI609251, and TAM428 scored 3, 3, and 5 respectively. SC748-5, the resistant check, scored 2. As the majority of accessions using the excised leaf assay scored 1 or 2 at 1-leaf stage (seedlings exhibit only 1st leaf and flag leaf), juvenile plants of these Senegalese accessions can be assumed as generally resistant to the Texas isolate FSP53 of C. sublineola (Fig. 1a,b are representing score 1 & 2). Yet, 16 cultivars showed clear sign of infection (Fig. 1c).
Acervuli formation was confirmed on leaf segments of Sorghum seedlings response to FSP53 at 96 h post-inoculation. (a) Visual representation of score 1. No visible infection was shown on a leaf segment of SC748-5 (resistant check) (b) Visual representation of score 2. Germ tube formation was confirmed on a leaf segment of SC748-5 (c) Acervuli formation was confirmed on a leaf segment of PI514300. Although not prevalent, FSP53 successfully formed acervuli on Senegalese seedlings. Note: The images of (a) and (b) were used for visual representation of score 1 and 2. The images are not associated with this study as it captured responses of leaf segments of SC748-5 at 8-leaf stage to FSP53 at 96 h post-inoculation from a previous unpublished work.
In a previous study, the same sorghum accessions inoculated at the 8-leaf stage in a greenhouse rather than using excised-leaf assays were generally resistant to a mixture of Texas isolates8, but many different outcomes were shown when compared to the results from this study (Table 1).
Genome-wide association study
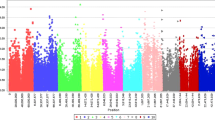
The SNP locus S01_72868925 is the only SNP that passed the Bonferroni threshold, but Table 2 listed 3 high confidence candidates close to the associated SNPs (Fig. 2 and Table 2).
The genome-wide association for response to anthracnose in Senegalese sorghum seedlings. Manhattan plot showing locations of SNP-detected QTLs associated with response to FSP53 on the ten chromosomes of Sorghum bicolor at 1-leaf stage. Bonferroni correction≈0.00000039 after filtering out SNPs with greater than 20% unknown alleles with minor allele frequency (MAF) below 5%. The red line indicates Bonferroni correction.
When mapped back to the published sorghum genome, top candidates SNPs that passed or nearly passed Bonferroni correction were nearest to genes that have previously reported roles in biotic or abiotic resistance/stress responses.
Discussion
Sorghum seedlings are known to be more resistant compared to later growth stages. It has been assumed this may be because they contain the preformed cyanogenic glycoside dhurrin, which may play a role in seedling protection9. Similarly, cultivars of Johnsongrass [Sorghum halepense (L.) Pers.], a wild relative of sorghum, were reported to be more resistant in young plants compared to fully grown plants when inoculated to a sorghum isolate of C. sublineola13. As Table 1 depicts, the results followed the previously reported patterns when compared to 8-leaf stage inoculated sorghum plants8. Majority of accessions were resistant in seedling stage, but they become more susceptible at the 8-leaf stage. In contrast, 11 cultivars showed completely opposite results; susceptible seedlings were detected, but acervuli formation was not observable in 8-leaf stage plants in the previous study (PI514298, PI514299, PI514300, PI514372, PI514427, PI514448, PI514449, PI514455, PI514459, PI514474 and PI514478)8. Both of these changes infer different resistance mechanisms operate in juvenile versus older plants. If dhurrin is inhibitory, these plants may simply be those with naturally low levels. The single isolate used in these studies was selected due to its pathogenicity on BTx635 at both stages of growth whereas a mix of isolates known to represent different pathotypes was used in in the previous tests. Nevertheless, a recent study showed that following inoculation with single or a mixture of two isolates of C. sublineola, had minimal to no effect, in that infection rates were more characteristic for the virulent isolate in mixed inoculum14. The use of the single isolate may contribute the decrease in number of high scores, but as FSP53 strain has a strong and broad virulence pattern to sorghum cultivars5,11, the effect is assumed to be minimal. The results of screening sorghum to C. sublineola can also be altered by environmental effects. For example, screening results in Texas and Puerto Rico differed in a study8. Similarly, SAP lines responded differently to anthracnose based on inoculum and field locations4. PI514284, PI514293 and PI514473 were susceptible at both seedling and 8-leaf stage8. It is speculated that change of phytoalexin levels may vary from seedling to 8-leaf stage in individual sorghum accessions.
The SNP locus S01_72868925 is located in protein kinase domain // Leucine rich repeat N-terminal domain (Sobic.001G451800). LRRs are a feature of nearly all cloned resistance genes and are widely known for roles in plant host defense8. Although not the same SNP, GWAS analysis based on identical accessions after inoculation at the 8-leaf stage showed the SNP locus S06_60609133 as the top candidate SNP, and the SNP locus S06_60609133 tagged leucine rich repeat/protein tyrosine kinase (Sobic.006G274866)8. In other studies, sorghum mini core collection and SAP lines, regarding anthracnose and head smut, leucine rich repeat containing proteins were commonly listed as top candidate genes4,5,6, but chromosomal locations differed in each study.
The SNP locus S08_7370058 is 16202 bp away from a poly (ADP-ribose) polymerase (PARP), catalytic domain. PARP domains have also been implicated as factors in stress responses of plants15. Transgenic plants with reduced PARP levels have broad-spectrum stress-resistant phenotypes16.
The SNP locus S09_51943886 is located within the gene coding for flavonoid 3'-monooxygenase. Chalcone synthase (CHS) is a key enzyme of the flavonoid/isoflavonoid biosynthesis pathway17. CHS expression causes accumulation of flavonoid and isoflavonoid phytoalexins and is involved in the salicylic acid defense pathway17.
As sorghum seedlings typically contain the preformed cyanogenic glycoside dhurrin9, it was expected to see majority of 1st leaves to be scored as 1. Sorghum pathologists often label an accession as susceptible when even a single sorghum plant among a number exposed to inoculum can be infected. An example is head smut caused by Sporisorium reilianum (Kühn) Langdon & Fullerton. Similarly, a GWAS analysis based on the highest score of the 1st leaves based on the excised-leaf assay in each accession identified candidate genes associated with SNP loci supported by strong statistical power. As sorghum responses differed at the 1-leaf stage versus the 8-leaf stage, it is not surprising to identify novel SNP loci, including the SNP locus Sobic.001G451800 that are associated with seedling defense, but it will be essential to explore the response more deeply in the future. It is quite possible that different SNPs are highly associated with sorghum defense in seedling stage only or throughout the whole growth stages. The candidate genes identified in this study can be tested for further analysis such as real-time quantitative reverse transcription PCR (Real-time qRT-PCR) or RNA sequencing analysis (RNA-Seq) to confirm that different genes are highly expressed in seedling stages. Gene editing technology can also be applied to verify the defensive roles of the candidate genes. Lastly, level of cyanogenic glycosides can be measured at seedling and 8-leaf stages to find associations with phenotypes and the identified genes.
In sum, this study explored sorghum seedling responses to C. sublineola with an excised-leaf assay and verified that the excised-leaf assay can be useful to study seedlings, but compared to 8-leaf stage inoculation methods, a greater number of seedlings should be screened to determine susceptibility in each accession as the results can easily be skewed.
However, the excised-leaf assay applied to 1-leaf stage seedlings is expected to promote studies associated with sorghum seedlings and C. sublineola interactions; it can be rapidly conducted within laboratory setting.
Methods
Sorghum lines
The 159 accessions (listed in Supplemental Information for the accessions and raw scores to C. sublineola) were obtained from the USDA-ARS Plant Genetic Resources Conservation Unit. BTx623, TAM428, and PI609251 (susceptible) and SC748-5 (resistant) were used as positive and negative controls8,18.
Excised-leaf assay and disease evaluations
Plug flats with 40 Square Cells (L x W x H≈ 5 cm × 5 cm × 7 cm for each cell) (The HC Companies, Twinsburg, OH) were filled with Metro Mix 200 (Sun Gro Horticulture, Agawam, MA), and sorghum accessions were grown at 23 °C with 65% humidity under LED light for approximately 12 h a day. When seedlings reached at l-leaf stage, an excised-leaf assay described by Prom et al.11 and Ahn et al.19 was applied. Prom et al.11 developed the assay to screen 8-leaf stage sorghum plants, but in this study, 1-leaf stage seedlings were used. In brief, a Texas C. sublineola isolate FSP53 was inoculated onto half strength Potato Dextrose Agar (PDA) plate and grown in an incubator for 2 weeks. FSP53 is one of the most virulent isolates based on the response of susceptible checks BTx623, TAM428, and RTx430 in field evaluation5, and the identical Senegalese sorghum accessions’ response to a mixture of Texas isolates including FSP53 were screened at the 8-leaf stage in a greenhouse recently8. Approximately 50 ml of sterile water was added to the plate, and a sterile spatula was used to scrape and remove colony growth, and the suspension mixture was screened through four layers of cheesecloth to obtain conidia19. Conidia concentrations were adjusted to a f 106 conidia/ml19. For the excised-leaf assay, whole leaves of each cultivar were placed, adaxial side up, on a half strength PDA plate, and 5 μl of the spore suspension was inoculated on the leaf blade19. In each trial, three 1st leaves were inoculated for each accession, and 3 trials of the excised-leaf assay were conducted (Total number of inoculated 1st leaves = 9 throughout 3 trials in each accession). Excised 1st leaves were inspected under an Olympus BX60 microscope (Olympus Co., Shinjuku, Tokyo, Japan) with 10 × magnification at 96 h post-inoculation, and scored for seedling responses to C. sublineola were recorded by using a 1–5 scale where 1 = fully resistant without visible fungal infection; 2 = fungal germ tube formed; 3 = fungal bed formed with some imperfectly formed acervuli; 4 = 1–5 acervuli perfectly formed and 5 = more than 5 acervuli perfectly formed19. Among 9 observed 1st leaves, the highest score was used to determine the symptom types to be categorized into two reaction types: Ratings 1 or 2 as resistant; ratings 3–5 as susceptible19. All raw scores are listed in Supplemental Information.
GWAS Analysis
SNP data from a recent study8 were used. Originally, SNP data were extracted from an integrated sorghum SNPs dataset based on sorghum reference genome version 3.1.1 and originally genotyped using GBS20,21,22,23, and missing data were imputed using Beagle 4.124.
TASSEL12 version 5.2.80 was used to conduct a mixed linear model (MLM) association analysis based on the highest score for disease ratings in each accession. False associations were reduced by removing SNPs with greater than 20% unknown alleles. Furthermore, SNPs with minor allele frequency (MAF) below 5% were removed as well. SNPs contributing to seedling responses to C. sublineola were tracked to the specific chromosome location by using the sorghum genome sequence, version 3.1.1 available at the JGI Phytozome 13 website25. To verify statistical significance, the mean disease rating score for all Senegalese accessions with either of the two prevalent bases was determined and verified to differ significantly (P < 0.05) based on Student’s T-test by using JMP Pro 15 (SAS Institute, Cary, NC, USA)5. Any top candidate SNP with (P ≥ 0.05) were removed from the results reported in Table 2.
Data availability
The raw phenotypic data is available as a Supplemental Information. The SNP dataset used is originally from a study deposited to Dryad Data Repository (https://doi.org/10.5061/dryad.63h8fd4, database/repository name: An integrated genotyping-by-sequencing polymorphism map for over 10,000 sorghum genotypes)21.
References
Ananda, G. K. S. et al. Wild Sorghum as a promising resource for crop improvement. Front. Plant Sci. https://doi.org/10.3389/fpls.2020.01108 (2020).
AshokKumar, A. Sorghum hybrids development for important traits: Progress and way forward. In Sorghum: A state of the art and future perspectives agronomy monographs (eds Ciampitti, I. A. & Prasad, P. V. V.) 97–117 (American Society of Agronomy and Crop Science Society of America, USA, 2019).
Xavier, K. V., Mizubuti, E. S. G., Queiroz, M. V., Chopra, S. & Vaillancourt, L. Genotypic and pathogenic diversity of Colletotrichum sublineola Isolates from Sorghum (Sorghum bicolor) and Johnsongrass (S. halepense) in the Southeastern United States. Plant Dis. 102, 2341–2351. https://doi.org/10.1094/PDIS-04-18-0562-RE (2018).
Cuevas, H. E., Prom, L. K., Cooper, E. A., Knoll, J. E. & Ni, X. Genome-wide association mapping of Anthracnose (Colletotrichum sublineolum) resistance in the U.S. Sorghum association panel. The Plant Genome. https://doi.org/10.3835/plantgenome2017.11.0099 (2018).
Prom, L. K., Ahn, E., Isakeit, T. & Magill, C. GWAS analysis of sorghum association panel lines identifies SNPs associated with disease response to Texas isolates of Colletotrichum sublineola. Theor. Appl. Genet. 132, 1389–1396. https://doi.org/10.1007/s00122-019-03285-5 (2019).
Ahn, E. et al. Genome wide association analysis of sorghum mini core lines regarding anthracnose, downy mildew, and head smut. PLoS ONE 14, e0216671. https://doi.org/10.1371/journal.pone.0216671 (2019).
Cuevas, H. E., Prom, L. K. & Rosa-Valentin, G. Population structure of the NPGS Senegalese sorghum collection and its evaluation to identify new disease resistant genes. PLoS ONE 13, e0191877. https://doi.org/10.1371/journal.pone.0191877 (2018).
Ahn, E., Prom, L. K., Hu, Z., Odvody, G. & Magill, C. Genome-wide association analysis for response of Senegalese sorghum accessions to Texas isolates of anthracnose. The Plant Genome. https://doi.org/10.1002/tpg2.20097 (2021).
Nicholson, R. L., Jamil, F. F., Snyder, B. A., Lue, W. L. & Hipskind, J. Phytoalexin synthesis in the juvenile sorghum leaf. Physiol. Mol. Plant Pathol. 33, 271–278. https://doi.org/10.1016/0885-5765(88)90027-6 (1988).
Tenkouano, A., Miller, F. R., Hart, G. E., Frederiksen, R. A. & Nicholson, R. L. Phytoalexin assay in juvenile sorghum: An aid to breeding for anthracnose resistance. Crop Sci. 33, 243 (1993).
Prom, L., Cuevas, H., Isakeit, T. & Droleskey, R. Excised Leaf Method for High Volume Evaluation of Sorghum Germplasm for Resistance Against Colletotrichum sublineolum. Vol. 15 (2016).
Bradbury, P. J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635. https://doi.org/10.1093/bioinformatics/btm308 (2007).
Ahn, E., Odvody, G., Prom, L. K. & Magill, C. Late growth stages of johnsongrass can act as an alternate host of Colletotrichum sublineola. Plant Health Progress. https://doi.org/10.1094/PHP-11-19-0084-RS (2020).
Ahn, E., Fan, F., Prom, L. K., Odvody, G. & Magill, C. The response of sorghum cultivars to mixtures of compatible and incompatible isolates and different conidia concentrations of Colletotrichum sublineola. Physiol. Mol. Plant Pathol. https://doi.org/10.1016/j.pmpp.2021.101690 (2021).
Rissel, D. & Peiter, E. Poly(ADP-Ribose) polymerases in plants and their human counterparts: Parallels and peculiarities. Int. J. Mol. Sci. 20, 1638. https://doi.org/10.3390/ijms20071638 (2019).
Vanderauwera, S. et al. Silencing of poly(ADP-ribose) polymerase in plants alters abiotic stress signal transduction. Proc. Natl. Acad. Sci. 104, 15150–15155. https://doi.org/10.1073/pnas.0706668104 (2007).
Dao, T. T. H., Linthorst, H. J. M. & Verpoorte, R. Chalcone synthase and its functions in plant resistance. Phytochem. Rev. 10, 397. https://doi.org/10.1007/s11101-011-9211-7 (2011).
Prom, L., Cuevas, H., Isakeit, T. & Droleskey, R. Excised leaf method for high volume evaluation of sorghum germplasm for resistance against Colletotrichum sublineolum. Plant Pathol. J. 15, 11–16. https://doi.org/10.3923/ppj.2016.11.16 (2016).
Ahn, E., Prom, L. K., Odvody, G. & Magill, C. Defense responses against the sorghum anthracnose pathogen in leaf blade and midrib tissue of johnsongrass and sorghum. Physiol. Mol. Plant Pathol. 106, 81–86. https://doi.org/10.1016/j.pmpp.2018.12.008 (2019).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379. https://doi.org/10.1371/journal.pone.0019379 (2011).
Hu, Z., Olatoye, M. O., Marla, S. & Morris, G. P. An integrated genotyping-by-sequencing polymorphism map for over 10,000 Sorghum Genotypes. The Plant Genome 12, 180044. https://doi.org/10.3835/plantgenome2018.06.0044 (2019).
Upadhyaya, H. D. et al. Developing a mini core collection of sorghum for diversified utilization of Germplasm. Crop Sci. 49, 1769–1780. https://doi.org/10.2135/cropsci2009.01.0014 (2009).
Wang, Y.-H. et al. Genetic structure and linkage disequilibrium in a diverse, representative collection of the C4 model plant, Sorghum bicolor. G3 Bethesda, Md 3, 783–793. https://doi.org/10.1534/g3.112.004861 (2013).
Browning, B. L. & Browning, S. R. Genotype imputation with millions of reference samples. Am. J. Human Genet`. 98, 116–126. https://doi.org/10.1016/j.ajhg.2015.11.020 (2016).
McCormick, R. F. et al. The Sorghum bicolor reference genome: Improved assembly, gene annotations, a transcriptome atlas, and signatures of genome organization. Plant J. 93, 338–354. https://doi.org/10.1111/tpj.13781 (2018).
Acknowledgements
This study was funded by AFRI, NIFA, USDA (grant number 20156800423492) & by The Feed the Future Innovation Lab for Collaborative Research on Sorghum and Millet, a United States Agency for International Development Cooperative (agreement number AID-OAA-A-13-00047 with title name: Enabling marker assisted selection for sorghum disease resistance in Senegal and Niger). We thank Dr. Zhenbin Hu for aiding GWAS analysis.
Author information
Authors and Affiliations
Contributions
E.A. designed experiments, investigated, analyzed, validated data, curated data, and wrote original draft of manuscript. C.F. investigated and validated data. L.P.K. provided resources and wrote and revised manuscript. C.M. acquired funding for the experiments, administrated project, wrote and revised manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahn, E., Fall, C., Prom, L.K. et al. Genome-wide association study of Senegalese sorghum seedlings responding to a Texas isolate of Colletotrichum sublineola. Sci Rep 12, 13025 (2022). https://doi.org/10.1038/s41598-022-16844-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16844-6
- Springer Nature Limited