Abstract
Honey DNA metabarcoding provides information of floral sources of honey and foraging plant preferences of honey bees. We evaluated the floral composition of honey from two different species of honey bees, Apis cerana honey (ACH) and A. mellifera honey (AMH) in a mixed apiary located in a semi-forest environment to understand the floral preference and level of interspecific competition on floral resource. Three honey samples were collected from different hives of each species in mid-August. In total, 56 plant taxa were identified across the honey samples and among them, 38 taxonomic units were found in ACH compared with a total of 33 in AMH. The number of major plants (> 1% of reads) in honey samples was 9 and 11 in ACH and AMH respectively indicating the higher diversity of plant taxa in AMH. 23 taxonomic units were found exclusively in ACH, 18 taxonomic units were found only in AMH and 15 taxonomic units were shared between ACH and AMH indicating that 73% of the taxonomic units were present only in honey originated from one of the honeybee species. Qualitative and quantitative analyses of the shared major plants revealed the division of floral resource between these co-existing honey bee species pointing to a low level of interspecific competition between these two important pollinators.
Similar content being viewed by others
Introduction
Among all insects, honey bees are considered to be one of the most important pollinators1. Animal pollination directly affects about three-quarters of essential crop types, including most fruits, seeds, and nuts2. Two species of honey bee exist in South Korea: Apis cerana (the Eastern honey bee), which is a native honey bee and A. mellifera (the European honey bee), which is an exotic bee introduced to Korea in the early 1900s. The native bees are usually kept in logs or smaller boxes with movable frames, while the European honey bees are mostly housed in the standard Langstroth hives3. Because of the greater honey production of A. mellifera, commercial interest in the European honey bee has risen in the last decades4 and resulted in a gradual decline of A. cerana colonies in Japan and South Korea3. On the other hand, A. cerana is more suitably adapted to East Asian environmental conditions with long periods of rainfall5,6 as well as being more resistant to parasitic mites7, native predators8 and local diseases9.
Previous studies on the foraging behavior of these honey bees showed that A. cerana is able to collect more nectar in resource-scarce areas. In addition, their ability to start foraging at lower temperatures makes them suitable to pollinate flowering plants in late winter and early spring. In summer, A. cerana workers start foraging earlier than A. mellifera in the morning and stop foraging later in the evening resulting in approximately one extra hour of foraging than A. mellifera5,10. However, there are some foraging traits that would allow A. mellifera to outperform A. cerana. A. mellifera performs better than A. cerana during an odor acquisition phase11. A. mellifera has a significantly longer maximum foraging distance (over 10 km)12, compared to A. cerana’s 1.4–2.5 km13,14. In an apple orchard and broccoli field, A. cerana visited a higher number of flowers than A. mellifera per minute due to the longer times that A. mellifera remains on each flower than the native A. cerana15. A contradictory study found that there was no significant difference between the two species for number of flowers visited per minute, however, it is generally believed that A. mellifera visits a higher number of flowers on each foraging trip since their foraging trips cover a greater distance10 and they are able to carry heavier pollen loads10,15.
Since the introduction of exotic pollinators can lead to negative consequences on survival, growth, reproduction, and foraging behavior of native pollinators16,17,18,19, resulting in declines and extinctions of native plants, possessing information on the effect of the introduction of A. mellifera as an introduced honey bee on foraging activities of local pollinators is of crucial importance. Considering the ability of honey bees to fly hundreds of meters, monitoring the foraging behavior and floral preference of several bees simultaneously is challenging over a large area20.
The presence of pollen grains inside honey not only indicates the floral composition of honey, but also reflects the foraging behavior of honey bees. Furthermore, it gives long-term information about foraging activities of honey bees throughout the season. The traditional method of recognition of floral composition of honey is based on melissopalynology, which is an identification of pollen grains inside honey using light microscopy21. This method is tedious and time-consuming requiring expert palynologist in different groups of plants. If the pollen grains from phylogenetically close species of plants are similar morphologically, the taxonomic resolution of plant identifications through this method to species or at least genus level is often impossible22,23. DNA barcoding allows for identification and classification of organisms based on a short nucleotide sequence. Mixed origin, environmental samples, such as pollen, are characterized by the presence of DNA from different organisms. Next‐generation sequencing (NGS) technologies now allow high‐throughput sequencing of complex DNA libraries including pollen24,25,26. DNA metabarcoding, which is combination of NGS and DNA barcoding, can be applied in identifications of mixed origin pollen loads and surpass in reliability the traditional microscopic identification methods employed by palynology or mellisopalynology by detecting a greater number of taxa and tracing infrequently detected species23,24.
Ideal DNA barcodes have significant interspecific genetic variation and are flanked by conserved regions for universal primer binding to allow easy amplification across a wide range of taxa27. There is still a debate on the optimal DNA barcode for plants, and different molecular marker candidates were used for plant recognition through DNA barcoding such as chloroplast ribulose-bisphosphate carboxylase (rbcL), maturase K (matK), tRNA gene (trnL), intergenic psbA-trnH spacer and ribosomal internal transcribed spacer 2 (ITS2)28,29. All of them were used in the recent pollen DNA metabarcoding studies by Keller et al.24, Richardson et al.25, Sickle et al.26. Among them the rbcL chloroplast gene has been suggested as reliable candidate genes for plant identification which its highly universal primers are available30. Furthermore, recent detailed studies indicated the reliability of quantitative analyses based on plastid markers such as rbcL, trnL and matK24,31,32,33.
Since research on comparisons of floral preference in different species of honey bees is limited and most of the information is based on visual records in the field, the aim of this research has been to study floral composition of honey from two different honey bees, namely A. cerana and A. mellifera using the rbcL metabarcoding approach. In addition, possessing some knowledge on foraging preferences could assist us in long-term conservation programs aimed at rare and endangered plant species which these honey bees are prone to collect pollen and nectar from them.
Results
Sequencing output
The total sample consisted of three ACH and three AMH and one negative control. In total 477,674 sequences obtained from six honey samples and initial quality control based on quality score, adaptor and length trimming resulted in 377,391 high-quality merged reads (Supplementary table 2). Sequences were compared against the NCBI database, with 99.4% being characterized to family, genus or species level. In total, 57 plant taxa were identified across the honey samples. Most (64.3%) were identified to genus level, 30.1% were identified to species and the remaining 5.6% to family level. Although significant numbers of the taxonomic units were recognizable to species level, there is a possibility to have false positives as reference sequences of several species from related genera are lacking, so that the data related to genus level alone are presented in this study (Table 1, Fig. 1).
(a) The Euclidean distance–based dendrogram (top) summarizes the differentiation among honey samples from Apis melifera (AM-n) and Apis cerena (AC-n). (b) Bar charts showing the taxonomic composition at genus level in the six honey samples. Abundances of taxa are reported with the percentage values of reads. Taxa accounting for < 1% of reads are grouped as “Other taxa”. (c) Boxplots of the taxa richness in honey samples from A. cerana and A. mellifera. (d) Boxplots of Shannon diversity index of A. cerana honey and A. mellifera honey. P values are based on Mann–Whitney U test.
Floral composition of honey of Apis cerana and A. mellifera
In total, 23 taxonomic units were found exclusively in ACH; 18 plant taxa were found only in AMH and 15 taxonomic units were shared between ACH and AMH. Although the number of taxonomic units was ranging from 17 to 26 (n = 3) in ACH samples and 12 to 23 in AMH, the difference in taxon richness between ACH and AMH was not significant (Mann–Whitney U test, W = 6.5, p = 0.506, N = 3) (Fig. 1c). In total, 38 taxonomic units were found in ACH in comparison to a total of 33 found in AMH. The Shannon diversity index ranged between 1.47 and 2.1 in AMH and 1.15–1.55 in ACH, but the difference between them was not significant (Mann–Whitney U test, W = 1, p = 0.190) (Figs. 1d and 2b).
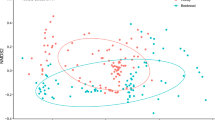
(a) Correspondence analysis (CA) representing the plant taxa and their abundances in the honey samples from Apis melifera (AM-n) and Apis cerena (AC-n). (b) A heat map representation of the Spearman correlation matrix of the six honey samples based on floral composition. The numbers indicate the p-value of the correlation.
At family level, a total of 37 plant families (S1 Dataset) was determined with 14 families shared between ACH and AMH; 14 families were found only in ACH and 8 families in AMH. We examined the proportion of the taxonomic units of the honey samples and in ACH, 9 (23.1%) major taxa (> 1% of reads) were detected whereas the number of major taxa in AMH was 11 (32.3%) (Supplementary table 3).
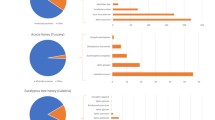
Castanea were found in a higher proportion in ACH including 44 percent of the reads, following by Actinidia and Anacardiaceae (Rhus/Toxicodendron) with relative abundance of 22% and 18%, respectively. Other major plant genera were Ricinus, Diospyros, Zanthoxylum, Brassica, Oryza and Flueggea found in a relatively high proportion (with relative abundance of 1.2–2.9%) (Table 1, Fig. 3). The results showed that in ACH, 95% of reads were represented by these taxonomic units and the remaining 29 taxonomic units comprised only 5% of the reads (Supplementary table 3). In AMH, Quercus is the dominant taxonomic unit with relative abundance of 38.5% of total reads followed by Diospyros, Amorpha, Chelidonium, Actinidia, Mazus, which are the other common taxonomic units with relative abundances of 14.9%, 12.7%, 7.3, 6.4%, 6% respectively, followed by Capsicum, Rosa and Flueggea and Robinia with relative abundances of 1.1–3.5% (Table 1, Fig. 3). In AMH, 97.3% of the reads represented 11 major plant taxa and the remaining 22 taxonomic units comprised only 2.7% of the reads (Supplementary table 3).
(a, b) Pie charts showing the taxonomic composition of major plants (> 1%) at genus level in A. cerana honey and A. mellifera honey. Abundances of taxa are reported with the percentage values of reads. Taxa accounting for < 1% of reads are grouped as “Other taxa”. (c) Venn diagram shows the number of unique and shared taxa identified at the genus level among honey samples.
The intraspecific pairwise comparison of honey samples showed that there was a significant correlation in abundance of plant taxonomic units in ACH and AMH (p < 0.025 for all pairwise comparison) with Spearman’s Rho ranging from 0.309 to 0.544 in ACH and from 0.363 to 0.521 in AMH. The interspecific pairwise comparison of ACH and AMH was not significant (p > 0.025 for all pairwise comparison). The correspondence analysis (CA) result clearly showed the major plant species that the two different honey bee species visit, were different from each other (Fig. 2a). AMH showed relatively less intra-colony variability than ACH samples in floral compositions.
Actinidia and Rhus/Toxicodendron are the only taxonomic units found in all samples. Besides these taxonomic units, Castanea, Brassica and Ricinus are the major plants and Aralia, Robinia and Zea are the minor taxonomic units which are available in all ACH samples, whereas Quercus, Driospiros, Amorpha, Chelidonium and Capsicum are taxonomic units (all are among major taxa), which are present in all AMH samples (Table 1, Fig. 1b, Supplementary table 4).
Discussion
In this study, two species of honey bees, A. cerana and A. mellifera, had access to a wide range of agricultural and wild flowering plants in a semi-forested region. The study aimed to identify and understand the foraging preferences of these two important pollinators in a mixed apiary. In total 56 taxonomic units (genus level) were detected in the honey samples. According to the results, the number of taxonomic units which are exclusively presented in ACH and AMH was 23 and 15, respectively, showing that 73% of the taxonomic units were present only in one type of honey. In addition, both ACH and AMH shared Actinidia, Diospyros, Flueggea, and Anacardiaceae in the list of their major plants (Fig. 3c) which all together covered 43% and 25% of the total reads of ACH and AMH, respectively. Among them, Actinidia and Anacardiaceae included 40% of the total reads in ACH while representing only 7% of the reads in AMH. On the other hand, unlike AMH with Diospyros included to 15% of the total reads, it is only covered to about 2% of the reads in ACH. These differences clearly reveal the partitioning of foraging floral resources between A. cerana and A. mellifera (Table 1, Supplementary Fig. 3).
Pollen is a source of lipid, protein and micronutrients, necessary for growth and health of honey bees34 and its quality as well as other factors such as availability of plants within the landscape and quantity of nectar and morphology of flowers are crucial factors for bees to collect pollen from35,36. In addition, Pollen grains attach to honeybee body while visiting flowers during nectar collecting and study the pollen grains inside honey indicates the plants visited by honeybees37. One possible reason for the presence of different major plants in the two types of honey is the difference in nutritional requirements of these two species for healthy growth of their colonies; however, the absence of Castanea which is an important foraging target for A. mellifera among major plants of AMH is quite surprising. Another possible reason for visiting different flower resources by A. cerana and A. mellifera is due to the avoidance of interspecific competition. Previous studies have shown that the introduction of A. mellifera can reduce survival, growth, reproduction, and have a negative impact on feeding behavior of native pollinators17,18. Since A. cerana is a close relative of A. mellifera38, the introduction of A. mellifera in A. cerana distribution areas can cause devastating ecological effects such as cross-species mating with A. cerana and reductions of the native species’ fitness39, exchange of pests and diseases and more importantly interspecific competition on floral resource19,40. These negative effects have caused regional extinctions of the A. cerana in Japan41, India16 and China19. The negative effects of introducing an exotic pollinator may lead to behavioral modifications of endemic ones such as timing of flower visitations to avoid competition with introduced honey bees42. The introduction of the European honey bee into A. cerana’s territory had considerable negative effects on A. cerana’s population and the plants used to be pollinated by A. cerana. However, due to resource partitioning between the introduced and native honey bees, after about one hundred and twenty years of its first introduction, A. mellifera like A. cerana can both be considered to be important pollinators for Korean crops and wild flowering plants, showing that the preservation of both species must be taken seriously. Loss of a single pollinator can cause significant implications for plant communities due to the alteration in floral fidelity in the remaining pollinators and a reduction of plant reproduction43. Iwasaki et al.44 showed the resource partitioning between introduced bees, A. mellifera and Bombus terrestris, with New Zeeland native solitary bees indicated limited resource overlap between bee taxa being the result of low competition due to different nutritional requirements and the pollinators’ abilities to access floral resources. The presence of both honey bee species in the same apiary can lead honey bees to modify aspects of their behavioral repertoire and to avoid competition for floral resources and to provide pollination services for higher numbers of flowering plants in nature. Most previous research focused on the pollination network among a coexisting pollinator community44,45,46. However, there is still only limited information available about the impact of resource partitioning on colony growth and survival of pollinators which is worthwhile to be taken into consideration in future studies.
The richness of the taxonomic units in the ACH was slightly higher than that of the AMH, resulting in a higher taxa diversity index of AMH indicating that the proportion of major plants was higher in AMH. However, the differences in taxa richness and diversity index between ACH and ADH were not significant which it is due to the low sample size. Lucek et al.47 also mentioned the requirement of high sample size to detect a statistical difference in taxa richness and diversity index between honey samples collected from urban and non-urban sites. Previous studies also demonstrated that A. cerana visited more flowering plants than A. mellifera48 and exhibited a higher performance while collecting pollen from scattered flowers of a variety of plant species, making more foraging trips per day and spending less time on each flower13,15,49. Being a native species of Asia, A. cerana commence foraging earlier in the day than A. mellifera and initiate foraging earlier in spring due to their ability to tolerate lower temperature (Supplementary Fig. 2). A. cerana also have access to a wider range of flowering plants to collect nectar and pollen from and the presence of Corylus among major plants of one of the A. cerana honey samples is indicative of the foraging activities of the recent species in early March (Table 1). Another plant taxon providing early season flowers is Brassica presented among major plants and identified from all ACH samples.
The list of taxonomic units of ACH and AMH contains several major plants followed by a long tail of minor plants. Previous melissopalynological or DNA metabarcoding studies on floral composition of honey also indicated that honey bees use a wide range of flowering plants with a relatively limited number of core plants22,36,50. Although the most abundant plants in the studied area were coniferous plants covering about 40% of landscape and about 60% of vegetation covered area (Supplementary table 1), Cerdus was the only taxonomic unit presented among minor plants which was visited exclusively by A. mellifera (Supplementary table 3).
The majority (92% of reads) of major plant taxa from ACH were the woody trees and shrubs including Castanea, Actinidia, Diospyros. Ricinus, Flueggea, Anacardiaceae, and Zanthoxylum. Likewise, in DNA extracts from AMH, apart from Capsicum, Mazus, and Chelidonium, the rest of the major plants (80% of total reads) were among trees and shrubs which included Quercus, Actinidia, Diospyros, Amorpha, Flueggea, Anacardiaceae and Rosa. Among horticultural plants, Brassica and Oryza were the major plants in ACH while Caspicum was present in AMH (Table 1, Supplementary table 3).
Both A. cerana and A. mellifera preferred to collect pollen from open type of corolla; the corolla type is quite diverse among major plants recognized from the honey produced by both species, however, Amorpha which have a tubular type of corolla found only in AMH. Most of the major taxa collected in this study by honey bees were among pollen-rich flowering plants except Robinia, which is an excellent source of nectar51,52. Moreover, Querqus and Ricinus which rarely become targets for honey bees as foraging plants, were among the major plants of A. mellifera and A. cerana, respectively. Previous studies on the foraging behavior of two species of stingless bees (Melipona bicolor and M. quadrifasciata) and A. mellifera showed that niche overlaps were more evident for nectar than for pollen53. Robinia, Amorpha and caspicus are introduced taxa, originating from Central and North America (Supplementary table 4), found as major plants of AMH, including 17% of the total reads whereas in ACH, Robinia, Ricinus and Oryza are the introduced major taxa with only about 5% of reads. Ricinus originated from Africa while Oryza was introduced from China (Supplementary table 4) where A. cerana is a native species14. En/Ex (Table 1) attributed to the major taxa containing at least one introduced and one endemic species in Korea and among them Anacardiaceae is one of the most abundant taxa in ACH with relative abundance of 17.7% while only covering 1.5% in AMH. The only introduced plant in Anacardiaceae, T. vernicifluum, is native to India and China (Supplementary table 4) where A. cerana is also endemic14. In total, compared with ACH, the abundance of introduced plant taxa in AMH is slightly higher than in ACH, which is congruent with previous studies in Japan where A. cerana visited native plant species more often than A. mellofera48. Nevertheless, some of the endemic taxa such as Quercus, Mazus, and Rosa are exclusively found in AMH.
Honey bees collect nectar and pollen from visiting flowers during their foraging trips to provide nutrients required for their colony development54. Nectar is processed to form honey. However, some pollen grains are also available in the honey allowing us to analyze the preferences that the bees have for certain species of foraging plants. In this study, the DNA was extracted from honey pellets after centrifuging. One possible reason for the fact that DNA metabarcoding is able to detect hidden and low abundant taxa in comparison with mellisopalynology22 is that the DNA can be also extracted from the plant cells which are available in the nectar through DNA metabarcoding. Recent studies indicated the presence of different groups of organisms such as Viruses, Bacteria, Arthropoda, Fungi, Plants and even vertebrates in the DNA extracted from pellets of the centrifuged honey55. Prosser and Hebert56 also documented the presence of different plant taxa inside DNA extract from the liquid part of the honey and the pellet as well. However, since different markers were used for liquid and pellet we were not able to discuss their taxon richness and diversity and it has to remain a task for future studies.
Methods
Honey samples
Three A. cerana cerana and three A. mellifera ligustica hives with healthy queens from a mixed stationary apiary, were selected randomly for sampling. A. mellifera colonies were kept in 10-framed langthroth hives while A. cerana colonies were kept in smaller size modern hives with 8 movable frames designed for Eastern honeybee. The apiary containing 10 A. cerana and 30 A. mellifera and located in a semi-forested hilly area with some agricultural fields of rice, potato, tomato, and pepper and small orchards of jujube, peach and cherry nearby, on the western coast of Andongho Lake, Andong, South Korea (reservoir, 36°40′00″ N, 128°49′34″ E, 177 m) (Supplementary Fig. 1). Five new frames were introduced into each colony of both honeybees in early March and Honey samples were harvested from introduced frames on the 15th August 2020 and 500 g honey from each blended sample were stored for forthcoming analyses.
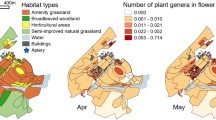
Landscape analysis was conducted over a 2.5 km radius around the apiary to identify landcover types. The radius was decided based on foraging distances of both A. mellifera and A. cerana12,13,57. We used landcover map data with 1 m spatial resolution from the Korea Ministry of Environment (KME) website (http://eng.me.go.kr/). A circle of 2.5 km radius was given to the base map and the landcover types were extracted in ArcGIS pro58. In the given circle, potential foraging habitats for nectar of 14.3 km2 were identified. Among the forest types, coniferous, broad-leaf and mixed forests were dominant, followed by agricultural fields and grassland (Supplementary table 1).
DNA extraction
Total DNA was extracted using a modified version of protocol of DNeasy Plant Mini Kit (Qiagen) for DNA isolation from honey used by de Vere et al.36 with modification on preparing samples. Briefly, 10 g of each honey were used for DNA extraction. DNA from each of the honey samples extracted four times and mixed together before library preparation. The honey sample was placed into a sterile 50 ml centrifuge tube and 30 ml pure water was added. Samples were incubated at 65 °C for 30 min with shaking in every five minutes, and then centrifuged for 30 min at 15,000 g. The supernatant was discarded and each pellet was resuspended in 10 ml ultrapure water and after homogenizing centrifuged again for 30 min at 15,000 g. The pellet resuspended in 1 ml water and transferred to 1.5 ml eppendorf tube and centrifuged for 10 min at 15000 g and the sediment were used for DNA isolation following the protocol by de Vere et al.36. An isolation negative control was also included using 40 ml of ultrapure water. The extracted DNA stored at − 20 °C prior to library preparation.
Library preparation and sequencing
Two rounds of PCR were applied for library preparation of rbcL DNA barcode marker region. The first PCR was conducted to amplify the rbcL region using the universal primers rbcL2f60 and rbcLaR61 to which adaptor ‘tails’ had been added and the second PCR was applied to attach unique tags for separation of the sequences into samples after sequencing. PCR was conducted in a final volume of 25 μl. A total of 2.5 μl of template DNA was combined with 12.5 μl of 2 × KAPA HiFi HotStart ReadyMix (KAPABIOSYSTEMS) and 5 μl of each primer (1 μM). The reaction was performed following program: initial denaturing at 95 °C for 3 min, followed by 30 cycles of 95 °C for 30 s, 50 °C for 30 s, 72 °C for 30 s with a final extension at 72 °C for 5 min. Products from the first PCR were purified using AMPureXP purification kit. The second PCR was conducted in a final volume of 50 μl using 5 μl of the first PCR product as a template, mixed with 25 μl of KAPA HiFi HotStart ReadyMix, 5 μl of Nextra XT Index Primer 1 (N7xx), 5 μl of Nextra XT Index Primer 2 (S5xx) and 10 μl of PCR Grade water. The PCR reaction was repeated as for the first PCR but with 8 cycles. The resulting products were purified using a AMPureXP purification kit. The pooled mixture was sequenced in a single flow cell on a 600-cycle run of the Illumina MiSeq instrument (Illumina, USA) at Macrogen (South Korea). All sequence information has been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (accession code PRJNA768940).
Bioinformatic analysis
Raw FASTQ files were demultiplexed by the sequencing company. Reads were trimmed for presence of adapter sequence using Trimmomatic62. Paired-end reads were quality-controlled considering the phred scale quality score trimming threshold, which was set to 25 and only the paired-end reads more than 180 bp after trimming were kept. Forward and reverse reads were merged using VSEARCH 2.14.163 and the merged reads lesser than 450 bp and quality score lower than 20 were discarded using SICKLE version 1.3364. Length trimmed sequences were converted to FASTA format and dereplicated using VSEARCH. An rbcL reference database was assembled as described below using OBITools365. All available relevant sequences were downloaded from EMBL database (2021.04.17), dereplicated and trimmed based on same primer sets used in this study and a formatted taxon list was created using only taxa with complete binomial species names. The assembled reference database was converted to FASTA format and used to check potential chimeras using uchime-denovo command in VSEARCH. Chimera free sequences were clustered at 97% threshold using VSEARCH. The BLASTN algorithm (blast-2.11.0)66 was used to align representative sequences with the GenBank derived sequence library (20201.06.21) using the following settings: E-value cutoff 1e−125, number of alignments 5, output format 0, number of descriptions 10, percent identity threshold 95%. The species name assigned to the sequence when the top bit score is matched to a single species. If the top bit score be the same for different species belonging to the same genus, then the sequence identified to that genus. A family level assigned to the sequence when the top bit score matched to multiple genera within the same family. The sequences not identified in family level were discarded. The scientific name of OTUs matched with the list of Korean plants (http://koreanplant.info/wordpress/index.php/plantlist-eng/) and the ones with species name not available in the list recognized in genus level. To reduce sequencing errors, OTUs with lesser than 10 reads were discarded. In addition, the negative control was used to set threshold values for sequence removals and any taxonomic classifications recorded from fewer reads than the read number we obtained from a negative control were removed from further analysis. The relative abundances of each taxon in percentage were used as a semi-quantitative measurement to demonstrate the honey bees’ foraging behavior.
Statistical analysis
Statistical analyses involved R 4.1.0. Spearman’s Rank Correlations for multiple testing were used to assess whether each colony of honey bees used the same plants in similar proportions. Differences between taxa richness and Shannon diversity index from the samples of each group (A. cerana and A. mellifera honey samples) were compared using Mann–Whitney U test. Spearman’s Rank Correlations and Mann–Whitney U tests were used as the data did not meet the assumption of normality required for parametric tests.
We conducted correspondence analysis (CA) to visualize the relationships between plant taxa and their abundances in the honey samples from A. melifera and A. cerena. CA was conducted with “FactoMineR” package in R 4.1.067. A hierarchical clustering analysis (HCA) was also conducted based on plant species information to identify whether combination of floral sources can specify foraging characteristics of two honey bee species, A. melifera and A. cerena. The ward algorithm with minimum variance method was used for the clustering with “cluster” package in R 4.1.067. The clustering result was visualized in a dendrogram plot.
Data availability
All sequence information has been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (accession code PRJNA768940).
References
Ballantyne, G., Baldock, K. C. R., Rendell, L. & Willmer, P. G. Pollinator importance networks illustrate the crucial value of bees in a highly speciose plant community. Sci. Rep. 7, 8389 (2017).
Smith, M. R., Singh, G. M., Mozaffarian, D. & Myers, S. S. Effect of decreases of animal pollinators on human nutrition and global health: A modeling analysis. Lancet 386, 1964–1972 (2015).
Jung, C. & Cho, S. Relationship between honey bee population and honey production in Korea: A historical trend analysis. J. Apic. 30(1), 7–12 (2015).
Abrol, D. P. Asiatic Honey Bee Apis cerana: Biodiversity Conservation and Agricultural Production (Springer, 2013).
Chandel, Y. S., Kumar, A. & Srivastva, S. Comparative performance of Apis mellifera L. vis a vis Apis cerana Fab. on toria (Brassica campestris var Toria) in mid-hill zone of Himachal Pradesh, India. Indian J. Agric. Res. 34, 264–267 (2000).
Feng, M., Ramadan, H., Han, B., Yu, F. & Li, J. Hemolymph proteome changes during worker brood development match the biological divergences between western honey bees (Apis mellifera) and eastern honey bees (Apis cerana). BMC Genomics 15, 563–576 (2014).
Peng, Y. S., Fang, Y., Xu, S. & Ge, L. The resistance mechanism of the Asian honey bee, Apis cerana Fabr., to an ectoparasitic mite, Varroa jacobsoni Oudemans. J. Invertebr. Pathol. 49, 54–60 (1987).
McClenaghan, et al. Behavioral responses of honey bees, Apis cerana and Apis mellifera, to Vespa mandarinia marking and alarm pheromones. J. Apic. Res. 58(1), 141–148 (2018).
Lin, Z. et al. Go east for better honey bee health: Apis cerana is faster at hygienic behavior than A. mellifera. PLoS ONE 11(9), e0162647 (2016).
Verma, L. R. & Dutta, P. C. Foraging behaviour of Apis cerana indicia and Apis mellifera in pollinating apple flowers. J. Apic. Res. 25, 197–201 (1986).
Wang, Z. & Tan, K. Comparative analysis of olfactory learning of Apis cerana and Apis mellifera. Apidologie 45(1), 45–52 (2014).
Beekman, M. & Ratnieks, F. L. W. Long-range foraging by the honey-bee Apis mellifera L.. Funct. Ecol. 14, 490–496 (2000).
Dyer, F. C. & Seeley, T. D. Dance dialects and foraging range in three Asian honey bee species. Behav. Ecol. Sociobiol. 28, 227–233 (1991).
Koetz, A. H. Ecology, behaviour and control of Apis cerana with a focus on relevance to the Australian incursion. Insects 4(4), 558–592 (2013).
Joshi, N. C. & Joshi, P. C. Foraging behavior of Apis spp. On apple flowers in a subtropical environment. N. Y. Sci. J. 3(3), 71–76 (2010).
Atwal, A. S. & Sharma, O. P. The dominance of Apis mellifera over Apis indica. Am. Bee J. 111, 343 (1971).
Kato, M., Shibata, A., Yasui, T. & Nagamasu, H. Impact of introduced honey bees, Apis mellifera, upon native bee communities in the Bonin (Ogasawara) Islands. Res. Popul. Ecol. 41, 217–228 (1999).
Thorp, D. W., Wenner, A. M. & Barthell, J. F. Pollen and nectar resource overlap among bees on Santa Cruz Island. MBC Appl. Environ. Sci. 2020, 261–267 (2000).
Yang, G. Harm of introducing the western honey bee Apis mellifera L. to the Chinese honey bee Apis cerana F. and its ecological impact. Acta Entomol. Sin. 48, 401–406 (2005) ((in Chinese)).
Dubois, T., Pasquaretta, C., Barron, A. B., Gautrais, J. & Lihoreau, M. A model of resource partitioning between foraging bees based on learning. PLoS Comput. Biol. 17(7), e1009260 (2021).
Louveaux, J., Maurizio, A. & Vorwohl, G. Methods of melissopalynology. Bee World 59, 139–153 (1978).
Hawkins, J., de Vere, N., Griffith, A. & Ford, C. R. Using DNA metabarcoding to Identify the floral composition of honey: A new tool for investigating honey bee foraging preferences. PLoS ONE 10(8), e0134735 (2015).
Smart, M. D. et al. A Comparison of honey bee-collected pollen from working agricultural lands using light microscopy and its metabarcoding. Environ. Entomol. 46(1), 38–49 (2016).
Keller, A. et al. Evaluating multiplexed next-generation sequencing as a method in palynology for mixed pollen samples. Plant Biol. 17, 558–566 (2015).
Richardson, R. T. et al. Rank-based characterization of pollen assemblages collected by honey bees using a multi-locus metabarcoding approach. Appl. Plant Sci. 3, 1500043 (2015).
Sickel, W. et al. Increased efficiency in identifying mixed pollen samples by meta-barcoding with a dual-indexing approach. BMC Ecol. 15, 20 (2015).
Kress, W. J. & Erickson, D. L. DNA barcodes: Genes, genomics, and bioinformatics. PNAS 105, 2761–2762 (2008).
Hollingsworth, P. M., Graham, S. W. & Little, D. P. Choosing and using a plant DNA barcode. PLoS ONE 6, e19254 (2011).
Chen, S. et al. Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 5, e8613 (2010).
CBOL Plant Working Group. A DNA barcode for land plants. PNAS 106, 12794–12797 (2009).
Pornon, A. et al. Using metabarcoding to reveal and quantify plant-pollinator interactions. Sci. Rep. 6, 27282 (2016).
Bell, K. L. et al. Quantitative and qualitative assessment of pollen DNA metabarcoding using constructed species mixtures. Mol. Ecol. 28(2), 431–455 (2018).
Baksay, S. et al. Experimental quantificarion of pollen with DNA metabarcoding using ITS1 and trnL. Sci. Rep. 10, 4202 (2020).
Ghosh, S. & Jung, C. Nutritional value of bee-collected pollens of hardy kiwi, Actinidia arguta (Actinidiaceae) and oak, Quercus sp. (Fagaceae). J. Asia Pac. Entomol. 20(1), 245–251 (2017).
Brunet, J., Thairu, M. W., Henss, J. M., Link, R. I. & Kluevert, J. A. The effects of flower, floral display, and reward sizes on bumblebee foraging behaviour when pollen is the reward and plants are dichogamous. Int. J. Plant Sci. 176(9), 811–819 (2015).
de Vere, N. et al. Using DNA metabarcoding to investigate honey bee foraging reveals limited flower use despite high floral availability. Sci. Rep. 7(1), 42838 (2017).
Barth, O. M. In O polen no mel brasileiro. Rio de Janeiro, Luxor 151 (1989).
Lo, T., Gloag, R. S., Anderson, D. L. & Oldroyd, B. P. A molecular phylogeny of the genus Apis suggests that the giant honey bee of southern India, A. indica Fabricius, are valid species. Syst. Entomol. 35, 226–223 (2010).
Pirk, C. W. W., Sole, C. L. & Crewe, R. M. Pheromones. In Honey Bees of Asia (eds Hepburn, H. R. & Radloff, S. E.) 207–214 (Springer, Berlin, 2011).
Theisen-Jones, H. & Bienefeld, K. The Asian honey bee (Apis cerana) is significantly in decline. Bee World 93, 90–97 (2016).
Sakagami, S. F. Some interspecific relations between Japanese and European honey bees. J. Anim. Ecol. 28, 51–68 (1959).
Thomson, D. Competitive interactions between the invasive European honey bee and native bumble bees. Ecology 85, 458–470 (2004).
Brosi, B. J. & Briggs, H. M. Single pollinator species losses reduce floral fidelity and plant reproductive function. PNAS 110(32), 13044–13048 (2013).
Iwasaki, J. M. et al. Floral usage partitioning and competition between social (Apis mellifera, Bombus terrestris) and solitary bees in New Zealand: Niche partitioning via floral preferences?. Austral Ecol. 43(8), 937–948 (2018).
Rodrigues, C. S., Ferasso, D. C., Mossi, A. J. & Coelho, G. C. Pollen resources partitioning of stingless bees (Hymenoptera: Apidae) from the southern Atlantic forest Acta Scientiarum. Biol. Sci. 42, e48714 (2020).
Lucas, A. et al. Floral resource partitioning by individuals within generalised hoverfly pollination networks revealed by DNA metabarcoding. Sci. Rep. 8(1), 5133 (2018).
Lucek, K. et al. Metabarcoding of honey to assess differences in plant-pollinator interactions between urban and non-urban sites. Apidologie 50, 317–329 (2019).
Tatsuno, M. & Osawa, N. Flower visitation patterns of the coexisting honey bees Apis cerana japonica and Apis mellifera (Hymentoptera: Apidae). Entomol. Sci. https://doi.org/10.1111/ens.12206 (2016).
Kuang, B. Y. & Kuang, H. O. Biology of the Honey bee (Yunnan Science and Technology Press, 2002) ([In Chines]).
Ghorab, A. et al. Sensorial, melissopalynological and physico-chemical characteristics of honey from Babors Kabylia’s region (Algeria). Foods 10, 225 (2021).
Sasaki, M. In Bee’s eye view of flowering plants: Nectar- and pollen-source plants and related honey bee products. Kaiyusha, Tokyo, Japan (2010).
Simpson, M. G. Diversity and classification of flowering plant: Eudicots. Plant Syst. 2010, 275–448 (2010).
Wilms, W. & Wiechers, B. Floral resource partitioning between native Melipona bees and the introduced Africanized honey bee in the Brazilian Atlantic rain forest. Apidologie 28, 339–355 (1997).
Klein, S. et al. Honey bees increase their foraging performance and frequency of pollen trips through experience. Sci. Rep. 9, 6778 (2019).
Bovo, S. et al. Shotgun metagenomics of honey DNA: Evaluation of a methodological approach to describe a multikingdom honey bee derived environmental DNA signature. PLoS ONE 13(10), e0205575 (2018).
Prosser, S. W. J. & Hebert, P. D. N. Rapid identification of the botanical and entomological sources of honey using DNA metabarcoding. Food Chem. 214, 183–191 (2017).
Dhaliwai, H. S. & Sharma, P. L. Foraging range of the Indian honey bee. J. Apic. Res. 13, 137–141 (1974).
ESRI. ArcGis Pro (Version 10.6); ESRI Inc.: Redlands, CA, USA (2020).
Palmieri, L., Bozza, E. & Giongo, L. Soft fruit traceability in food matrices using real-time PCR. Nutrients 1, 316–328 (2009).
Kress, W. J. et al. Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot in Panama. PNAS 106, 18621–18626 (2009).
Kress, J. W. & Erickson, L. D. A two-locus global DNA barcode for land plants: The coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE 2(6), 1–10 (2007).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30(15), 2114–2120 (2014).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Joshi, N. A. & Fass, J. N. Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33). https://github.com/najoshi/sickle (2011).
Boyer, F. et al. OBITools: A Unix-inspired software package for DNA metabarcoding. Mol. Ecol. Res. 16, 176–182 (2016).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
R Core Team. R: A language and environment for statistical computing. https://www.gbif.org/tool/81287/r-a-language-andenvironment-for-statistical-computing (2022).
Bosch, J., Retana, J. & Cerdá, X. Flowering phenology, floral traits and pollinator composition in a herbaceous Mediterranean plant community. Oecologia 109, 583–591 (1997).
Endress, P. K. Flower structure and trends of evolution in eudicots and their major subclades. Ann. Missouri Bot. Gard. 97(4), 541–583 (2010).
Gómez, J. M., Torices, R., Lorite, J., Klingenberg, C. P. & Perfectti, F. The role of pollinators in the evolution of corolla shape variation, disparity and integration in a highly diversified plant family with a conserved floral bauplan. Ann. Bot. 117, 899–904 (2016).
Watts, S., Dormann, C. F., González, M. M. & Ollerton, J. The influence of floral traits on specialization and modularity of plant-pollinator networks in a biodiversity hotspot in the Peruvian Andes. Ann. Bot. 118, 415–429 (2016).
Lee, C. B. Coloured Flora of Korea. Hyangmunsa, Seoul, Republic of Korea (2003).
Acknowledgements
We are grateful to Dr. YD Lim to facilitate study in his apiary. We also thank Céline Mercier (Laboratoire de microbiologie des environnements extrêmophiles) for technical support with OBITools software. This study was partly supported by the BSRP through the National Research Foundation of Korea (NRF), Ministry of Education (NRF-2018R1A6A1A03024862).
Author information
Authors and Affiliations
Contributions
S.M.N. and C.J. conceived the study, collected samples and wrote the manuscript. S.M.N. designed and carried out the laboratory part of the study as well as bioinformatics analysis and statistical analysis. M.S. gathered plant related data and assessed plant determination and taxonomy as well as landscape analysis. M.K. conducted the clustering and correspondence analysis, climate data collection and analysis and completed the landscape analysis. C.J. provided the financial support of the study and supervised this research. All authors edited and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mohamadzade Namin, S., Kim, MJ., Son, M. et al. Honey DNA metabarcoding revealed foraging resource partitioning between Korean native and introduced honey bees (Hymenoptera: Apidae). Sci Rep 12, 14394 (2022). https://doi.org/10.1038/s41598-022-18465-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18465-5
- Springer Nature Limited
This article is cited by
-
The buzz about honey-based biosurveys
npj Biodiversity (2024)
-
Uncovering floral composition of paper wasp nests (Hymenoptera: Vespidae: Polistes) through DNA metabarcoding
Scientific Reports (2024)
-
The value of twinned pollinator-pollen metabarcoding: bumblebee pollination service is weakly partitioned within a UK grassland community
Scientific Reports (2023)
-
A preliminary metabarcoding analysis of Portuguese raw honeys
Archives of Microbiology (2023)