Abstract
The single nucleotide polymorphisms (SNPs) of complement factor H (CFH) gene are well-known genetic risk factors for age-related macular degeneration (AMD). To identify whether the measurement of plasma protein concentrations of CFH variants using the multiple reaction monitoring (MRM) assay can determine the genotypes of CFH SNP rs1061170 and rs800292, 120 patients with AMD and 26 controls were included in this study. The number of cases were TT:TC:CC = 121:24:1 in CFH SNP Y402H and GG:AG:AA = 72:57:17 in CFH SNP I62V. Plasma concentrations of tryptic peptides were measured using the MRM assay, and tyrosine/histidine (Y/H) and valine/isoleucine (V/I) CFH variant protein ratios were obtained. To discriminate the genotypes by the plasma protein ratios, cut-off values were set for Y/H ratios (TT: > 4.428; TC: 1.00–4.428; CC: < 1.00) and V/I ratios (GG: > 1.09; AG: 0.0089–1.08; AA: < 0.0089). Correlation analysis revealed that the plasma CFH variant protein ratios and genotypes of CFH were exactly matched (100%) without overlap in the total patients and controls. The measurement of plasma protein CFH variants using the MRM assay can accurately identify the genotypes of CFH SNPs of Y402H and I62V.
Similar content being viewed by others
Introduction
Age-related macular degeneration (AMD) is one of the major causes of blindness among adults aged 65 years or above in developed countries. Ageing, genetic factors, and smoking are well-known risk factors for AMD1.
AMD is a multifactorial disease and Inflammation is known to be one of the major mechanisms for the development and progression of AMD2. The complement system plays an important role in the mediation of inflammatory response and is considered to be related to the pathophysiology of AMD3. Previous genome-wide association studies and validation studies revealed that the variants of complement factor H (CFH) are strongly associated with the risk of AMD: risk allele of CFH single nucleotide polymorphism (SNP) rs1061170 (risk allele: C, tyrosine 402 to histidine [Y402H]) and rs800292 (risk allele: G, isoleucine 62 to valine [I62V])4,5,6,7,8,9.
To determine the genotypes of CFH SNPs, deoxyriboNucleic Acid (DNA) purification and sequencing are required. If the genotype could be revealed by blood protein quantification, it can be a useful tool and can be integrated with other protein markers while alleviating the need for DNA analysis. However, quantifying highly homologous sequence variants of CFH using the currently available antibody and aptamer-based approaches is not feasible. Multiple reaction monitoring (MRM) assay has emerged as a useful method for precise and accurate quantification of proteins and their variants that cannot be detected using antibody-based methods10. Recently, the detection of multiple CFH protein variants using MRM assays has been reported, showing the possibility that the CFH genotypes can be identified using MRM assay11.
In this study, to determine the actual clinical utility of MRM assay for CFH SNP genotyping, we analyzed the correlation between the genotype of the major CFH SNPs rs1061170 (Y402H) and rs800292 (I62V) and the concentration ratios of plasma CFH variants measured using the MRM assay in a cohort of AMD patients and normal controls. Additionally, we also evaluated the possibility of replacing genetic testing with the MRM assay for the risk assessment of AMD via proteomic CFH genotyping.
Methods
This study protocol was reviewed and approved by the Institutional Review Board (IRB) of Seoul National University Bundang Hospital (IRB No: B-2105-682-105). In this study, a retrospective analysis was conducted on the collected clinical data and blood samples from the previous Bundang AMD Genetic Study9. The current study was conducted in accordance with the Declaration of Helsinki. A written informed consent was obtained from all participants in the secondary study.
Participants
Participants aged over 50 years who were newly diagnosed with intermediate AMD and nAMD between October 1, 2003, and December 31, 2016, were enrolled in the AMD group in our study. Participants aged over 50 years who were newly diagnosed with cataracts, without any retinal pathology between October 1, 2003, and December 31, 2016, were enrolled as the control group in our study. Intermediate AMD was defined as the presence of multiple large drusen (> 125 µm) on the macula with or without pigmentary changes, the absence of pre-existing central geographic atrophy (GA), and nAMD based on the international AMD classification12. nAMD was defined as the evidence of choroidal neovascularization (CNV) associated with non-drusenoid retinal pigment epithelium detachment, serous sensory retinal detachment, subretinal hemorrhage, and/or subretinal exudation. We collected basic demographic information, including age and sex, at the first visit. Smoking status was classified as non-smoker or former/current smoker.
DNA-based genotyping of CFH SNPs
Patient DNA was obtained from peripheral blood using a DNA extraction kit (QIAamp DNA Maxi kit, Qiagen Inc.). Multiplex polymerase chain reaction using a single base extension technique was performed for SNP genotyping using the iPLEX Gold kit and MassARRAY Typer software version 4.0 (Sequenom, San Diego, CA). Two SNPs known to be major risk alleles for AMD in Koreans, namely CFH rs1061170 and rs800292, were analyzed9.
Selection of target tryptic peptides
For proteomic genotyping, it is necessary to determine how the SNPs of the CFH gene are translated into proteins. Two SNPs of the CFH gene (rs1061170 and rs800292) were retrieved from NCBI dsSNP (http;// www.ncbi.nlm.nih.gov/snp) and the amino acid sequence of CFH was obtained from Uniport knowledgebase (Uniprot ID: P08603) (http;//www.uniprot.org/). CFH SNPs rs1061170 and rs800292 result in a change in codon assignment from tyrosine (Y) to histidine (H) at amino acid positions 402 and from isoleucine (I) to valine (V) at position 62. For the selection of target peptides in the MRM assay, tryptic peptides unique to each protein isoform were identified from the UniProt database using the BLAST search tool (https://www.uniprot.org/blast/) (Table 1). Stable isotope-labeled standard (SIS) peptides with the same amino acid sequence as the target peptides, but incorporating C-terminal [13C6, 15N4] arginine (> 95% purity) were purchased from JPT Technologies (Berlin, Germany). The SIS peptides are used as quantitative standards in a mass spectrometer as they are physicochemically identical to the target peptides and have only a difference in mass. For each peptide, amino acid analysis was performed, and the absolute amount of peptide was provided by the vendor. The SIS peptides were re-solubilized in 20% acetonitrile with 0.1% formic acid to prepare a 1 mg/mL stock solution. Two other dilution series (50 and 100 ng/mL) were prepared to optimize the MRM assay. All solutions were stored at − 80 °C until use.
Optimization of MRM assay
We experimentally optimized most of the empirical parameters of the MRM assay by directly infusing the SIS peptides into a 5500 Qtrap mass spectrometer equipped with a turbo v ion source (SCIEX, Foster City, CA). First, the intensities of the doubly and triply charged ions (initial Q1) were scanned, and the optimal declustering potential (DP) voltage was determined by ramping the DP, and the dominant precursor ions (Q1) were then measured with the optimal DP voltage (final Q1). Second, at the optimized Q1 and DP parameters, we initially measured 10 highly intense product ions of a precursor ion using the base collision energy (CE) and then ramped the CE voltages to find the best CE values. Subsequently, we obtained the optimal collision cell exit potential (CXP) values of the selected 10 product ions. All optimized data were collected and compared to the theoretical spectra, and three highly intense y-ions were used for subsequent MRM assays (Table 1 and Supplementary Fig. S1).
Sample preparation for MRM assay
The plasma (2 µL) sample was buffered with 50 µL of 8 M urea in 50 mM ammonium bicarbonate (ABC). The sample was treated using 7 µL of 100 mM tris (2-carboxyethyl) phosphine and 7 µL of 200 mM chloroacetamide at 37 °C for 1 h in the dark. Urea was diluted to 1 M with 50 mM ABC before trypsin (Promega, Madison, WI, USA) digestion at 1:50 enzyme: substrate ratio and incubated at 37 °C for 16 h with mixing on a shaker at 600 rpm. Formic acid was added at a final concentration of 0.5% to stop the digestion reaction. The mixture was desalted using a Sep-Pak tC18 96-well plate (Waters, Milford, MA, USA) and dried using a vacuum centrifuge (miVac Duo Concentrator; Genevac, Suffolk, UK). Dried samples were stored in a deep freezer at − 80 °C until use.
LC-MRM-MS analysis
Peptide separation was performed using an ACQUITY UPLC M-class (Waters). Mobile phase A was 0.1% formic acid in the water, and mobile phase B was 0.1% formic acid in acetonitrile. Samples were reconstituted with 40 µL of SIS peptide mixture in mobile phase A, injected with a full sample loop injection of 5 µL, and separated in an ACQUITY UPLC peptide CSH C18 column (1 mm i.d., 10 cm length, pore size 130 Å, particle size 1.7 µm; Waters). A gradient with a flow rate of 20 µL/min and 5% B for 3 min, 5–25% B for 22 min, 25–60% for 1 min, 60–60% for 1 min, 60–5% B for 1 min, and 5% B for 5 min, followed by column washing with 80% B for 5 min, 80% to 5% for 6 min, 5–80% for 6 min, and re-equilibration with 5% B for 13 min. MRM analysis was performed using a 5500 Qtrap mass spectrometer. The MS detection was carried out in positive mode with the following parameters: ion spray voltage of 5500 V, curtain gas at 20 psi, nebulizer gas at 25 psi, heating gas at 20 psi, resolution at 0.7 Da (unit) for Q1/Q3, interface temperature at 400 °C, and scan mass range 300–1250 m/z QQQ mode. Quantification experiments were performed using the scheduled MRM mode. The MRM detection window time was 120 s, and the cycle time was 1.5 s. The mass spectrometer was operated with Analyst software (Version 1.6.2, SCIEX), which generated MRM-MS data with the file name *.wiff.
Quantitative MRM assay
The concentration of the endogenous target peptide can be calculated by comparing the peak areas of the peptide and the spiked SIS peptide resulting from the LC-MRM-MS analysis. Peak areas were integrated into the Analyst software by setting a three points Gaussian smooth width, 30 s RT window, 500 cps minimum peak height, and one point peak splitting factor. For an accurate quantification, analytical validation was performed using calibration curves, specificity, precision and accuracy13. The calibration curves were used to predict the concentration of targets and to determine the limit of quantification (LOQ), the lowest concentration within a coefficient of variation (CV) of ≤ 20% with a coefficient of determination (R2) of 0.99 (Supplementary Fig. S2). Specificity was calculated by comparing endogenous analytes and internal standards in double blank (DB), specificity analytes (S.A), blank and low QC samples. Under optimized MRM conditions, no interference was observed for each sample in the target peptide (Supplementary Table S1). To evaluate precision and accuracy, we prepared four types of quality control (QC) samples of different concentrations. For intra- and inter-day assays, each concentration level was tested for three consecutive days in five replicates. Both intra- and inter-day QC samples at each concentration for all four analytes were acceptable (Supplementary Table S2 and S3).
Statistical analysis
The statistical differences in age, sex, smoking status, and the genotype of CFH SNP rs1061170 and rs800292 between the normal controls and AMD groups were evaluated using the Chi-square test and Mann–Whitney test. The correlation between the plasma concentration of tyrosine and histidine and the genotypes of CFH SNP rs1061170 was evaluated using Kruskal–Wallis test. The correlation between the plasma concentration of valine and isoleucine and the genotypes of CFH SNP rs800292 was also evaluated using Kruskal–Wallis test. Scatter plots were drawn to visualize the correlation between the plasma tyrosine/histidine (Y/H) ratio and the genotypes of CFH SNP rs1061170 and between plasma valine/isoleucine (V/I) ratio and the genotypes of CFH SNP rs800292. Cut-off values of plasma CFH variant ratios were set to best delineate the CFH genotypes, and the correlation between MRM-based and DNA-based genotypes of CFH was evaluated. Statistical significance was defined as a P value of < 0.05. All statistical analyses were performed using SPSS (version 24.0; SPSS Inc., Chicago, USA).
Results
In our study, the number of cases was 26 in the control group and 120 in the AMD group. The average age of the subjects was 68.4 ± 9.0 years. Among the patients with AMD, intermediate AMD was observed in 55 patients (45.8%), and nAMD was observed in 65 patients (54.2%). There were no statistical differences between the control and AMD groups in age, sex, and smoking status (P = 0.092, 0.840, and 0.303, respectively).
The distribution of alleles in CFH SNP rs1061170 was 121 for the TT allele (82.9%), 24 for the TC allele (16.4%), and 1 for the CC allele (0.7%). The distribution of alleles in CFH SNP rs800292 was 72 for the GG allele (49.3%), 57 for the AG allele (39.0%), and 17 for the AA allele (11.6%) (Table 2).
The average plasma concentrations of tyrosine and histidine in the MRM assay according to CFH SNP rs1061170 genotypes were as follows: Tyrosine: 46.2 ± 18.4 ng/mL (TT allele), 27.0 ± 11.8 ng/mL (TC allele), 0.0001 ng/mL (CC allele); histidine, 0.861 ± 1.076 ng/mL (TT allele), 12.6 ± 4.5 (TC allele), and 12.7 ng/mL (CC allele) (Table 3, Fig. 1). There was a statistically significant difference between the plasma concentrations of CFH with tyrosine and histidine and the genotypes of CFH SNP rs1061170 (P < 0.001). The average plasma concentrations of valine and isoleucine in the MRM assay according to CFH SNP rs800292 were as follows: Valine: 423.1 ± 152.2 ng/mL (GG allele), 225.2 ± 93.8 (AG allele), 0.818 ± 1.017 (AA allele), isoleucine—3.1 ± 9.3 (GG allele), 341.3 ± 183.5 (AG allele), and 683.4 ± 354.5 (AA allele) (Table 4, Fig. 2). There was a statistically significant difference between the plasma concentrations of valine and isoleucine and the genotypes of CFH SNP rs800292 (P < 0.001).
The correlation between the plasma CFH tyrosine/histidine (Y/H) variant protein ratio and the genotypes of CFH SNP rs1061170 are depicted in Fig. 3. The average plasma CFH Y/H ratio according to the genotypes of CFH SNP rs1061170 was as follows: TT, 91.301 ± 68.915 (5.804–403,100.775); TC, 2.218 ± 0.882 (1.059–4.428); CC: 0.00000785453. We set the cut-off values as 4.428 and 1.00, to match the plasma Y/H ratio and the SNP genotypes. (TT: Y/H ratio > 4.428; TC: 1.00 ≤ Y/H ratio ≤ 4.428; CC: Y/H ratio < 1.00) The distribution of plasma CFH Y/H ratio by measuring MRM assay and genotype of CFH SNP rs1061170 was exactly matched (100%) (Table 3, Fig. 4).
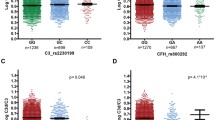
The distribution of logarithm plasma CFH tyrosine/histidine variant protein ratio according to the genotype of complement factor H single nucleotide polymorphism rs1061170 and the presence of age-related macular degeneration. C, control; A, age-related macular degeneration; CFH, complement factor H; SNP, single nucleotide polymorphism; Y, tyrosine; H, histidine; log, logarithm.
The correlation between the plasma CFH valine/isoleucine (V/I) variant protein ratio and the genotypes of CFH SNP rs800292 is depicted in Fig. 5. The average plasma CFH V/I ratio according to the genotypes of CFH SNP rs800292 was as follows: GG, 606.906 ± 641.983 (7.854–4654.144); GA, 0.711 ± 0.138 (0.519–1.090); AA, 0.001 ± 0.002 (0.0001–0.009). We set the cut-off values as 1.09 and 0.0089 to match the plasma V/I ratio and SNP genotypes (GG: V/I ratio > 1.09; AG: 0.0089 ≤ V/I ratio ≤ 1.08; AA: V/I ratio < 0.0089). The distribution of plasma V/I ratio by measuring MRM assay and genotype of CFH SNP rs800292 was exactly matched (100%) (Table 4, Fig. 6). The plasma concentrations of CFH protein variants could accurately predict the genotypes of CFH SNPs rs1061170 and rs800292.
The distribution of logarithm plasma CFH valine/isoleucine variant protein ratio according to the genotype of complement factor H single nucleotide polymorphism rs800292 and the presence of age-related macular degeneration. C, control; A, age-related macular degeneration; CFH, complement factor H; SNP, single nucleotide polymorphism; V, valine; I, isoleucine; log, logarithm.
Discussion
In our study, we measured the plasma concentrations of CFH variants using MRM analysis and correlated the CFH variant ratios with the genotypes of CFH SNPs rs1061170 and rs800292. We identified an exact match between the genotypes of CFH SNPs rs1061170 and rs800292 and the ratios of plasma levels of CFH variants. Thus, the quantification of plasma CFH variants using the MRM assay may be a useful alternative to DNA-based genotyping of the two CFH SNPs.
The SNPs in the CFH and age-related maculopathy susceptibility 2 (ARMS2) genes have shown the greatest impact on risk assessment and the pathogenic mechanism of AMD6,14,15,16. CFH is present at relatively high concentrations in the blood. CFH is produced primarily in the liver and circulates throughout the body including the retina through the blood17. The abundant concentration of CFH in the blood enabled the measurement of CFH variants and the correlation with the corresponding genotypes. However, in the case of ARMS2, no protein has been found in the blood; thus, proteomic analysis is not feasible. Among other genes related to AMD, genes that produce proteins present in the blood can be analyzed using the MRM assay in the same way. However, the major risk SNPs of AMD are CFH and ARMS2 (odds ratio: ARMS2 SNP rs10490924, 3.13; CFH SNP rs1061170, 2.74; CFH SNP rs800292, 2.43), while the other known SNPs related to AMD have insufficient clinical significance (e.g. odds ratio: 2.82 for CFB rs4151667; 2.31 for CFB rs438999; 1.58 for SYN3/TIMP3 rs9621532; 1.52 for C3 rs2230199)18. In the previous studies, in Caucasians, Y402H (CFH SNP rs1061170) was reported to be highly related to AMD, while in Asians, I62V (CFH SNP rs800292) was reported to be highly related to AMD9,19,20,21,22,23,24,25. Thus, proteomic identification of CFH SNPs Y402H and I62V can be applied with value to both ethnicities, Asians and Caucasians.
A recent study by Zhang et al. quantified plasma CFH variants (Y402H, I62V) and CFH-related proteins (complement factor-related protein 1 to 5) using the multiplexed targeted mass spectrometry assay11. They analyzed the blood samples of participants in the Age, Gene/Environment Susceptibility-Reykjavik study and showed plasma CFH variant concentrations by inferred genotypes. Thus, they did not correlate plasma CFH protein concentrations with the true genotypes of the CFH gene SNPs. Additionally, the study was a population-based study that did not define patients with AMD and controls; thus, the actual clinical application of the MRM assay was not tested before.
In our study, we measured the plasma levels of CFH variants using MRM analysis and analyzed the genotypes of CFH SNPs rs1061170 and rs800292. Importantly, we proposed cut-off values for protein variant ratios for the discrimination of SNPs and identified an exact match between DNA-based genotypes and MRM-based genotypes of CFH SNPs rs1061170 and rs800292. The further advantages of proteomic genotyping can be suggested as follows: (1) It can be combined with other protein markers up to approximately 100 in a single MRM assay. Thus, additional proteomic analysis of plasma proteins can be performed simultaneously with minimal additional costs; (2) In the future, the risk SNPs of various genes that encode proteins expressed in the blood can be added for AMD and other diseases. This will shift the paradigm of genotyping from DNA-based genetic assays to protein-based multi-omics.
Our study has several limitations. First, our study was a retrospective, observational, single-center design. Second, the relatively small sample size of one ethnic group may be another limitation. Future validation including larger samples of different ethnicities is needed. However, our study showed an exact match between the proteomic genotypes defined by the concentration of plasma CFH variants and the DNA-based genotypes of CFH SNPs. To the best of our knowledge, this is the first study to suggest a clinically applicable method to identify the genotypes of CFH using proteomic MRM assay in a cohort of patients with AMD and controls.
In conclusion, the measurement of plasma CFH protein variants using the MRM assay can accurately predict the genotypes of CFH SNPs of Y402H and I62V. Proteomic genotyping of CFH using the MRM assays can be used without DNA-based genetic tests for the risk assessment of AMD.
Data availability
The datasets generated during and/or analysed during the current study are available in the Harvard Dataverse repository, [https://dataverse.harvard.edu/api/access/datafile/6413273].
References
Armstrong, R. A. & Mousavi, M. Overview of risk factors for age-related macular degeneration (AMD). J. Stem Cells 10, 171–191 (2015).
Ozaki, E. et al. Inflammation in age-related macular degeneration. Adv. Exp. Med. Biol. 801, 229–235. https://doi.org/10.1007/978-1-4614-3209-8_30 (2014).
Wu, J. & Sun, X. Complement system and age-related macular degeneration: drugs and challenges. Drug Des. Dev. Ther. 13, 2413–2425. https://doi.org/10.2147/dddt.s206355 (2019).
Joo, K., Mun, Y. S., Park, S. J., Park, K. H. & Woo, S. J. Ten-year progression from intermediate to exudative age-related macular degeneration and risk factors: Bundang AMD cohort study report 1. Am. J. Ophthalmol. 224, 228–237. https://doi.org/10.1016/j.ajo.2020.11.012 (2020).
Hu, J. et al. Age-related macular degeneration-susceptibility single nucleotide polymorphisms in a Han Chinese control population. Ophthalmic Epidemiol. 18, 137–142. https://doi.org/10.3109/09286586.2011.574335 (2011).
Tian, J. et al. Association of genetic polymorphisms and age-related macular degeneration in Chinese population. Investig. Ophthalmol. Vis. Sci. 53, 4262–4269. https://doi.org/10.1167/iovs.11-8542 (2012).
Babanejad, M. et al. Investigating the CFH gene polymorphisms as a risk factor for age-related macular degeneration in an Iranian population. Ophthalmic Genet. 37, 144–149. https://doi.org/10.3109/13816810.2014.955585 (2016).
García, M. et al. CFH polymorphisms in a Northern Spanish population with neovascular and dry forms of age-related macular degeneration. Acta Ophthalmol. 93, e658-666. https://doi.org/10.1111/aos.12790 (2015).
Woo, S. J. et al. Analysis of genetic and environmental risk factors and their interactions in Korean patients with age-related macular degeneration. PLoS ONE 10, e0132771. https://doi.org/10.1371/journal.pone.0132771 (2015).
MacLean, B. et al. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics (Oxford, England) 26, 966–968. https://doi.org/10.1093/bioinformatics/btq054 (2010).
Zhang, P. et al. A novel, multiplexed targeted mass spectrometry assay for quantification of complement factor H (CFH) variants and CFH-related proteins 1–5 in human plasma. Proteomics 17, 1600237. https://doi.org/10.1002/pmic.201600237 (2017).
Ferris, F. L. 3rd. et al. Clinical classification of age-related macular degeneration. Ophthalmology 120, 844–851. https://doi.org/10.1016/j.ophtha.2012.10.036 (2013).
Carr, S. A. et al. Targeted peptide measurements in biology and medicine: best practices for mass spectrometry-based assay development using a fit-for-purpose approach. Mol. Cell. Proteomics MCP 13, 907–917. https://doi.org/10.1074/mcp.M113.036095 (2014).
Kim, Y. H., Kim, H. S., Mok, J. W. & Joo, C. K. Gene-gene interactions of CFH and LOC387715/ARMS2 with Korean exudative age-related macular degeneration patients. Ophthalmic Genet. 34, 151–159. https://doi.org/10.3109/13816810.2012.749287 (2013).
Vavvas, D. G. et al. CFH and ARMS2 genetic risk determines progression to neovascular age-related macular degeneration after antioxidant and zinc supplementation. Proc. Natl. Acad. Sci. USA 115, E696-e704. https://doi.org/10.1073/pnas.1718059115 (2018).
Ryoo, N. K. et al. Thickness of retina and choroid in the elderly population and its association with Complement Factor H polymorphism: KLoSHA Eye study. PLoS ONE 13, e0209276. https://doi.org/10.1371/journal.pone.0209276 (2018).
Khandhadia, S. et al. Age-related macular degeneration and modification of systemic complement factor H production through liver transplantation. Ophthalmology 120, 1612–1618. https://doi.org/10.1016/j.ophtha.2013.01.004 (2013).
Grassmann, F., Fritsche, L. G., Keilhauer, C. N., Heid, I. M. & Weber, B. H. Modelling the genetic risk in age-related macular degeneration. PLoS ONE 7, e37979. https://doi.org/10.1371/journal.pone.0037979 (2012).
Gotoh, N. et al. No association between complement factor H gene polymorphism and exudative age-related macular degeneration in Japanese. Hum. Genet. 120, 139–143. https://doi.org/10.1007/s00439-006-0187-0 (2006).
Lau, L. I. et al. Association of the Y402H polymorphism in complement factor H gene and neovascular age-related macular degeneration in Chinese patients. Investig. Ophthalmol. Vis. Sci. 47, 3242–3246. https://doi.org/10.1167/iovs.05-1532 (2006).
Uka, J. et al. No association of complement factor H gene polymorphism and age-related macular degeneration in the Japanese population. Retina (Philadelphia, Pa.) 26, 985–987. https://doi.org/10.1097/01.iae.0000244068.18520.3e (2006).
Klein, R. et al. Inflammation, complement factor h, and age-related macular degeneration: The Multi-ethnic Study of Atherosclerosis. Ophthalmology 115, 1742–1749. https://doi.org/10.1016/j.ophtha.2008.03.021 (2008).
Liu, X. et al. Association study of complement factor H, C2, CFB, and C3 and age-related macular degeneration in a Han Chinese population. Retina (Philadelphia, Pa.) 30, 1177–1184. https://doi.org/10.1097/IAE.0b013e3181cea676 (2010).
Arakawa, S. et al. Genome-wide association study identifies two susceptibility loci for exudative age-related macular degeneration in the Japanese population. Nat. Genet. 43, 1001–1004. https://doi.org/10.1038/ng.938 (2011).
Quan, Y. L., Zhou, A. Y. & Feng, Z. H. Association between complementary factor H Y402H polymorphisms and age-related macular degeneration in Chinese: Systematic review and meta-analysis. Int. J. Ophthalmol. 5, 242–246. https://doi.org/10.3980/j.issn.2222-3959.2012.02.25 (2012).
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (Nos. 2020R1F1A1072795, 2017M3C9A5031595, 2017M3A9F9030559). The funding organization had no role in the design or conduct of this study.
Author information
Authors and Affiliations
Contributions
C.L., S.J.W. and S.-J.P. designed the experiment. H.K. and Y.L. conducted the experiment. K.L.K., S.-J.P., C.L. and S.J.W. Analyzed and interpreted data. K.J., S.J.P., K.H.P. and S.J.W. Provided materials. K.L.K. and S.J.W. wrote the article. And, S.J.W., S.-J.P. and C.L. proofed and revised the article.
Corresponding authors
Ethics declarations
Competing interests
Se Joon Woo: Samsung Bioepis (C, S), Curacle (C, S), Alteogen (C, S), Novelty Nobility (C, S), Panolos Bioscience (C, O), Novartis (C, L, S), Janssen (C), Bayer (L), Allergan (C, L), Abbvie (L, S), Alcon (L), SCAI Therapeutics (L), Philophos (C), Retimark (O) (C: consultant; L: Lecture fee; O: Equity Owner; S: grant support. All other authors have no competing/conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, K.L., Kim, H., Lee, Y. et al. Proteomic genotyping of SNP of Complement Factor H (CFH) Y402H and I62V using multiple reaction monitoring (MRM) assays. Sci Rep 12, 19587 (2022). https://doi.org/10.1038/s41598-022-20936-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-20936-8
- Springer Nature Limited