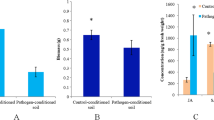
Abstract
Plants deposit photosynthetically-fixed carbon in the rhizosphere, the thin soil layer directly around the root, thereby creating a hospitable environment for microbes. To manage the inhabitants of this nutrient-rich environment, plant roots exude and dynamically adjust microbe-attracting and -repelling compounds to stimulate specific members of the microbiome. Previously, we demonstrated that foliar infection of Arabidopsis thaliana by the biotrophic downy mildew pathogen Hyaloperonospora arabidopsidis (Hpa) leads to a disease-induced modification of the rhizosphere microbiome. Soil conditioned with Hpa-infected plants provided enhanced protection against foliar downy mildew infection in a subsequent population of plants, a phenomenon dubbed the soil-borne legacy (SBL). Here, we show that for the creation of the SBL, plant-produced coumarins play a prominent role as coumarin-deficient myb72 and f6’h1 mutants were defective in creating a Hpa-induced SBL. Root exudation profiles changed significantly in Col-0 upon foliar Hpa infection, and this was accompanied by a compositional shift in the root microbiome that was significantly different from microbial shifts occurring on roots of Hpa-infected coumarin-deficient mutants. Our data further show that the Hpa-induced SBL primes Col-0 plants growing in SBL-conditioned soil for salicylic acid (SA)-dependent defenses. The SA-signaling mutants sid2 and npr1 were unresponsive to the Hpa-induced SBL, suggesting that the protective effect of the Hpa-induced shift in the root microbiome results from an induced systemic resistance that requires SA-signaling in the plant.
Similar content being viewed by others
Introduction
Microbial communities that associate with the plant root, together referred to as the root microbiome, are the focus of numerous studies that explore their potential to enhance plant health and growth1,2,3,4. Plants sculpt and sustain the root microbiome by depositing large amounts of photosynthetically-fixed carbon in the rhizosphere, creating nutrient-rich conditions for microbial growth and activity5,6. With a punishment and reward strategy, plants can modulate rhizosphere microbiome composition by the exudation of both microbe-stimulatory and -repellent metabolites7,8,9,10,11. By selectively adjusting the rhizosphere microbiome to favor beneficial microbes, plants can create conditions advantageous to their growth and health.
Previously, it was discovered that plants can dynamically recruit beneficial microbes in response to pathogen attack. Aboveground infection of the model plant Arabidopsis thaliana (hereafter: Arabidopsis) with Hyaloperonospora arabidopsidis12 (hereafter: Hpa) or Pseudomonas syringae pv. tomato13 led to specific changes in the root microbiome. A subsequent population of plants grown in soils thus conditioned with pathogen-infected plants was more resistant to the aboveground attacker. Moreover, the Arabidopsis root microbiome of Hpa-infected plants was enriched for a Xanthomonas, a Stenotrophomonas and a Microbacterium species. These bacteria were isolated and cultured, and only when all three were applied as a consortium they consistently induced systemic resistance (ISR) against Hpa12. Thus, we hypothesize that pathogen infection leads to changes in root exudation profiles that specifically promote beneficial microbes in their root microbiome and in turn trigger ISR to fend off subsequent pathogen attack. By increasing the abundance of these beneficial microbes in the rhizosphere, plants create a soil-borne legacy (SBL) that can protect a next generation of plants growing in that same soil14. The dynamic recruitment of beneficial microbes is suggested to be important for the buildup of disease suppressiveness in the field. This is exemplified by a study with sugar beet, in which attack by the soil-borne pathogen Rhizoctonia solani led to the activation of a disease-suppressive microbial consortium that protected the plant by antagonizing the pathogen15.
It is known that specific beneficial microbes can prime a plant’s immune system and bring about a state of ISR16,17. In this primed state of ISR, a plant activates defense responses upon pathogen detection much quicker and stronger, but without the plant growth penalty that is often associated with increased resistance18,19,20. Root colonization by Pseudomonas simiae WCS417 (hereafter: WCS417) can elicit ISR in Arabidopsis that is effective against a wide range of pathogens21,22,23. WCS417-mediated ISR is dependent on the transcription factor MYB72 that is expressed in the roots only, but is essential for the onset of ISR systemically21. MYB72 is also important in the plant’s iron deficiency response, as it regulates the biosynthesis and secretion of scopoletin8,24, a coumarin involved in mobilizing iron from the soil25. Moreover, scopoletin has specific antimicrobial activity that inhibits growth of soil-borne pathogens, such as Fusarium oxysporum and Verticillium dahlia, but to which beneficial MYB72-inducing WCS417 bacteria are tolerant8. MYB72 thus seems to serve a double role in signaling the onset of ISR as well as shaping the root microbiome. It was hypothesized that MYB72-dependent coumarin secretion is part of a positive feedback loop that stimulates colonization by MYB72-inducing and coumarin-tolerant beneficial microbes8,26,27,28, thus providing both enhanced pathogen protection and increased availability of iron in the soil environment.
In this study, we aimed to obtain mechanistic insight into how Hpa-infected plants create a SBL and how plants growing in SBL-conditioned soil become protected against Hpa infection. We found that the coumarin biosynthesis genes MYB72 and F6’H1 are essential for the creation of the SBL, as aboveground Hpa-infection in coumarin-deficient mutants myb72 and f6’h1 did not result in the creation of an effective SBL. We found that the secretion of a wide range of root metabolites is affected by foliar Hpa-infection and identified microbial taxa in the rhizosphere that differently respond to Hpa-infection on wild type and the coumarin-deficient mutant plants. Moreover, we found that plants growing on SBL-conditioned soil are primed for enhanced SA-dependent defenses, and that SA signaling mutants do not mount an ISR against Hpa on SBL-conditioned soil. Together these data highlight the role of coumarins in the build-up of a foliar pathogen-induced SBL and that this SBL is associated with the recruitment of a disease resistance-inducing microbiome that primes SA-dependent defenses in plants growing in the SBL-conditioned soil.
Materials and methods
SBL experiments
In this study, natural soil collected from the Reijerscamp nature reserve in the Netherlands was used12. The SBL setup was performed as previously described in detail (Methods S1)29. Briefly, approximately 30 Arabidopsis seeds were sown on 60-ml pots filled with soil and placed in a climate-controlled plant growth chamber at 21 °C, 70% relative humidity, 10-h light/14-h dark cycles. Two weeks after sowing, half of the pots were inoculated with a 50 spores/µl suspension of Hyaloperonospora arabidopsidis noco2 (Hpa) as previously described30. Briefly, an airbrush was used to spray spores suspension until tiny droplets started to form on the leaves of the plants, approximately 0.5 ml of spore suspension per pot. The other pots were sprayed with tap water as mock treatment. Hpa symptoms were allowed to develop for one week, after which all aboveground material of this so-called “conditioning population” of plants was cut off using a razor blade. A new population of plants was sown and grown as described above. After two weeks of growth of this second plant population, all plants were inoculated with Hpa and symptoms were allowed to develop for one week. Disease severity in this so-called SBL “responding population” of plants was determined based on Hpa spore densities normalized for plant fresh weight.
Wild-type plants of the Arabidopsis Col-0 accession (N1093; Nottingham Arabidopsis Stock Centre) were used in both the conditioning phase of the experiment (1st population of plants) and in the response phase (2nd population of plants). Depending on the experiment, Col-0 wild-type plants were replaced by mutant myb72-2, which is impaired in both rhizobacteria-mediated ISR21 and in the biosynthesis and secretion of coumarins8,24, the coumarin biosynthesis mutant f6’h1, which is impaired in the coumarin biosynthesis enzyme Feruloyl-CoA 6'-Hydroxylase131, or by SA signaling mutants npr1-132 or sid2-133. As controls, pots were left unplanted in either the conditioning phase or the response phase of the experiment. All experiments were performed at least twice with similar results.
Defense priming assay
To test for priming of plant defenses, 2-week-old plants in the response phase of an SBL experiment were dipped in a 1 mM SA solution supplemented with 0.015% Silwet L-77 or in a mock solution containing 0.015% Silwet L-77. Plants were harvested at 30 min, 4 h, and 6 h after dipping, immediately snap frozen in liquid nitrogen and stored at − 80 °C until further processing. RNA extraction was performed as described by Oñate-Sánchez34 with minor modifications. Quantitative real-time PCR (qPCR) analysis of the defense-related marker gene PATHOGENESIS-RELATED1 (PR1) was performed as described by Van Wees et al.35.
Preparation of amplicon sequencing library
For microbiome analysis, the roots from at least ten replicate pots were collected for each treatment at the end of the SBL conditioning phase. DNA was extracted using the DNeasy® Powerlyzer® Powersoil® kit (product #: 12855-100).The 16S rRNA gene amplicon libraries were constructed using the 16S metagenomic sequencing library protocol for the Illumina MiSeq system36 using 16S V3-V4 phasing primers adapted from De Muinck et al.37 (Table S1) and the barcoded PCR primers described by Baym and co-workers38. The pooled library was sent for sequencing on a 2 × 300 MiSeq at the USEQ sequencing facility (Utrecht University, the Netherlands). The fungal amplicon libraries were constructed in the same way but using primers targeting the internal transcribed spacer (ITS) region 2 of the ribosome encoding genes39.
Amplicon sequence data pre-processing
The 16S and ITS amplicon sequence data was analyzed using Qiime2 version 2019.740 as described in Methods S1. Reads annotated as chloroplast or mitochondria were removed from the data sets and only bacterial amplicon sequence variants (ASVs) with a total relative abundance ≥ 0.01% and detected in at least 10% of the samples were used for further analysis. Fungal ASVs with a total relative abundance of abundance ≥ 0.01% and detection in at least 10% of samples were used for downstream analysis.
Differential abundance analysis
Five statistical methods were used for differential abundance testing of ASVs: ANCOM-bc, DESeq2, Fisher’s exact test, Simper analysis and Spearman rank correlations41,42,43. ASVs were considered differentially abundant if the FDR-adjusted P ≤ 0.05 as generated by ANCOM-bc, DESeq2, and Spearman rank correlations. For Fisher’s exact test and Simper analysis we used unadjusted P ≤ 0.05 as the threshold for significance. Sparse ASVs denoted as structural zeroes by ANCOM-bc were ignored in downstream analysis unless also detected by at least one other statistical method.
Software used
All statistical analyses on the 16S amplicon sequencing data were performed in RStudio version 3.5.0 (data in Figs. 1 and 2) and version 3.6.1 (Figs. 3, 4 and 5). Figures were created in RStudio version 3.6.1 using phyloseq44, ggplot245, tidyheatmap (https://github.com/jbengler/tidyheatmap). PRISM8 Graphpad and Adobe Illustrator CC 2017 and 2022 were used to prepare graphs and figures.
Untargeted metabolomics
Sterile root exudates secreted by Arabidopsis roots in response to foliar Hpa inoculation were collected as described by Song et al. (summarized in Methods S1)46. Briefly, plants were sown on seed holders filled with agar-solidified 1/10 strength Hoagland (HG) solution. Roots of the germinating seedlings penetrated the agar to grow into vials with full strength HG solution and grown in controlled-climate conditions and kept sterile in Eco2boxes (Eco2 NV). After 19 days, leaves of half of the plants were carefully inoculated with an Hpa spore suspension. Five days post-inoculation, plants were removed from the HG medium and the medium was checked for sterility. Contaminated root exudates were discarded and sterile root exudates were lyophilized. Untargeted metabolomics was conducted on exudates using an ACQUITY ultra high performance liquid chromatography (UHPLC) coupled to a SYNAPT G2Si quadrupole time-of-flight (Q-TOF) mass spectrometer equipped with an electrospray (ESI) source (Waters) as previously described47 and summarized in Methods S1. Raw LCMS data were processed following the DIA MS2 deconvolution method using MS-DIAL software v. 4.748 and exploiting the online library MSMS-Public-Neg-VS15.msp (36,848 records). Putative annotation of differentially expressed metabolites resulted from MS-DIAL screening of the MS1 detected exact HR m/z and MS2 fragmentation patterns against multiple online databases (http://prime.psc.riken.jp/compms/msdial/main.html#MSP).48 After data-cleaning (blank check, SN > 10, CV QC < 30%), 923 metabolomic variables were retained for further chemometrics. The final dataset was normalized (median normalization, cube-root transformation and Pareto scaling) using MetaboAnalyst v 5.049 prior to multivariate statistical analyses. See Methods S1 for further details.
Collection of Hpa-responsive root exudates from soil
A population of Arabidopsis Col-0 plants was cultivated in the SBL experimental setup. Two weeks after sowing, half of the pots were inoculated with an Hpa spore suspension (50 spores/µl water) or with water. Seven days post-inoculation (dpi), approximately 65 ml of 5% ethanol was poured on top of the pots and allowed to flow through the soil. Following saturation of the soil, 50 ml of soil effluent was caught in 50-ml Falcon tubes and immediately frozen at − 20 °C. Samples were lyophilized for further processing.
Collection of Hpa-responsive root exudates from agar-solidified medium
Arabidopsis seeds were surface-sterilized and sown on agar-solidified Murashige and Skoog medium supplemented with 0.5% sucrose. After 4 days of stratification, plates were transferred to a climate chamber (21 °C, 70%, relative humidity, 10 h light/14 h dark, light intensity 100 μmol m−2 s−1). Twelve-day old seedlings were transplanted to agar-solidified HG medium with 5 seedlings per plate. On the same day, plants were inoculated with Hpa spores from a gnotobiotic laboratory-maintained Hpa noco2 culture (free of other microbes) by gently touching leaves from the infected plants against leaves of the healthy plants. Healthy plants that remained untreated and plates containing HG agar medium without any plants (empty control) were taken along as well. Plates were placed in sealed trays to ensure high humidity and stored in a climate chamber set at conditions favorable for infection (16 °C, 70% relative humidity, 9 h light/15 h dark, light intensity 100 μmol m−2 s−1). Seven days post-inoculation, rectangular pieces of agar medium were cut from the area occupied by the root elongation zone of plants that were visibly sporulating. Agar pieces were collected and pooled per plate as one biological replicate. For the empty control, agar samples of similar weight were taken from plates without plants. Samples were snap frozen in liquid nitrogen and stored at − 80 °C. Samples were lyophilized prior to further processing.
UHPLC-MS/MS analysis of coumarins
For detection of coumarins in root exudates, lyophilized samples were analyzed on a 1290 Infinity II UHPLC system equipped with an InfinityLab Poroshell 120 PFP, 2.1 × 100 (P/N: 695675-408) in combination with a Poroshell 120 PFP, with a 2.1 mm HPLC guard column and a 1.9 µm LC column (P/N: 821725-942; Agilent Technologies) coupled to an Agilent Technologies 6470A Tandem Quadrupole Mass Spectrometer (TQ-MS–MS). Coumarins were subsequently analyzed using MS with an ESI source in positive ion mode. Further details are described in Methods S1.
Ethics approval and consent to participate
The experimental research on plants in this study complies with relevant institutional, national, and international guidelines and legislation.
Results
MYB72 and coumarin biosynthesis are required for the creation of the soil-borne legacy mediated by Hpa-infected Arabidopsis
MYB72 regulates the production and secretion of coumarins, most notably scopoletin. Coumarins are secreted by the roots, have antimicrobial activity and can shape root microbiome assembly8,50. We therefore hypothesized that MYB72 and coumarin biosynthesis may play a role in the creation of the Hpa-induced SBL. This hypothesis was tested by preconditioning Reijerscamp soil with either wild-type Col-0, or the coumarin biosynthesis mutants myb72 or f6′h1 that were mock- or Hpa-inoculated. Moreover, we included non-planted, soil-filled pots that were sprayed with water or Hpa spores during the conditioning phase to investigate the possibility that the SBL was created via the Hpa inoculum. After removal of the first “conditioning population” of plants, a “responding population” of Col-0 plants was sown and grown on all pre-conditioned soils and inoculated with Hpa. One week post-inoculation, a significant reduction in Hpa spore production was observed for Col-0 plants growing on soil pre-conditioned by Hpa-infected Col-0 plants (Fig. 1), confirming previous findings (Berendsen et al.12). Interestingly, pre-conditioning of soil by the coumarin biosynthesis mutants myb72 and f6’h1 did not significantly affect Hpa resistance in the responding population of Col-0 plants (Fig. 1). Pre-conditioning of unplanted soil with Hpa inoculum had also no effect on Hpa resistance in Col-0 plants. We repeated this experiment with similar results (Fig. S1). Our results indicate that coumarin production is a pre-requisite for the creation of the SBL mediated by Hpa-infected plants.
Coumarin biosynthesis is essential for the creation of a Hpa-induced SBL that enhances resistance against downy mildew in a responding population of plants. Green bar graphs represent the disease severity of SBL-responding populations of Col-0 plants growing on natural Reijerscamp soil pre-conditioned with mock-treated or Hpa-inoculated Col-0, myb72, or f6’h1 seedlings (Experiment 1). Brown bar graphs represent the disease severity of responding populations of Col-0 plants growing in control pots that during the conditioning phase remained unplanted and were mock- or Hpa-inoculated directly on the soil. Bars depict the average Hpa spore production per gram of shoot fresh weight (n = 10). Error bars depict standard error of the mean. Asterisk denotes a significant difference (P = 0.01 in Students t-test).
Hpa infection results in significant changes in the root microbiome
To investigate the effect of Hpa infection on root microbiome assembly, we analyzed the composition of the root microbiomes of the healthy and Hpa-infected Col-0, myb72, and f6’h1 plants from the experiment described above (Experiment 1; Fig. 1). To this end, we sequenced 16S and ITS amplicons derived from roots of plants and of unplanted soil sampled at the moment the conditioning population of plants was removed. After filtering, the 16S data for this experiment comprised 67 samples with on average 97,108 reads per sample and a total of 1866 unique ASVs (Data S1). Principal coordinate analysis (PCoA) of these samples revealed that the root microbiome samples of Col-0, f6’h1 and myb72 plants clustered away from the unplanted soil samples, highlighting a strong rhizosphere effect for the bacterial communities (Fig. 2a, Table S2). Whereas the application of an Hpa spore suspension on unplanted soils did not result in changes in the soil microbiome, aboveground Hpa infection of plants led to a statistically significant shift (adjusted P < 0.05 in PERMANOVA) in the bacterial communities on the roots of Col-0 plants, as well as of the coumarin biosynthesis mutants myb72 and f6’h1 (Fig. 2b–d, Table S2). This shift constituted a change in the composition of the root bacterial communities, whereas the α-diversity of the communities was not affected (Fig. S2). Moreover, the bacterial communities on the roots of the mock-treated mutant plants were significantly different from those on the roots of mock-treated wild-type plants, whereas the bacterial communities on the roots of both mock-treated coumarin-deficient mutants did not differ from each other (Fig. 2e, Table S2). Also after Hpa-inoculation, the root bacterial communities on mutant plants were different from those on Col-0 plants (Fig. 2f), although the difference between infected myb72 and Col-0 plants was not significant at the P = 0.05 level (P = 0.081, Table S2).
Effect of foliar Hpa infection on root bacterial communities. (a) Principal coordinate analysis (PCoA) of bacterial communities in unplanted soil (filled triangle) and the rhizospheres of 3-week-old seedlings of Col-0 (filled circle), myb72 (⁎) and f6’h1 (filled rectangle). Samples were taken one week after either mock treatment (grey symbols) or inoculation with Hpa (blue-green symbols). (b) PCoA of only the Col-0 samples. (c) PCoA of only the myb72 samples. (d) PCoA of only the f6’h1 samples. (e) PCoA of root bacterial communities of mock-treated Col-0 (filled circle), myb72 (⁎) and f6’h1 (filled rectangle) plants. (f) PCoA of root bacterial communities of Hpa-infected Col-0 (filled circle), myb72 (⁎) and f6’h1 (filled rectangle) plants. Statistical tests on differences between microbial communities in the different genotypes and treatments is given in Table S2.
For the fungal ITS amplicon data, filtering resulted in a dataset of 74 samples with a total of 6,700,000 reads and 167 unique fungal ASVs. Also the fungal communities showed a clear rhizosphere effect as the roots of healthy and Hpa-inoculated Col-0, myb72 and f6’h1 plants were significantly different from the unplanted soil samples (adjusted P < 0.05 in PERMANOVA; Fig. S3, Table S2). In contrast to the bacterial microbiota (Table S2), root fungal microbiota were not significantly affected by foliar Hpa infection (Fig. S3, Table S3). This indicates that the root fungal microbiome is unlikely to play a role in the Hpa infection-mediated SBL and, for the further analyses in this study, we focused on the Hpa-induced changes in the bacterial root microbiome.
A large cluster of ASVs is significantly enriched on roots of Hpa-infected Col-0 plants
To dig deeper in the Hpa infection-mediated changes in the root microbiome, we analyzed the differential abundance of individual ASVs between healthy and Hpa-infected plants at the end of the conditioning phase of the experiment presented in Fig. 1. Several methods of differential abundance testing are well-accepted in literature and each of these methods has strengths and weaknesses51,52. For example, DESeq243 can detect small but consistent differences in ASV abundance, but in our experience is unlikely to detect ASVs that are sparsely present in at least one of two treatments. To make optimal use of the available methods of differential abundance testing, we selected ANCOM-bc, DESeq2, Fisher’s exact test, Simper, and Spearman’s correlation as complementary methods to analyze our data41,42,43. We identified 176 differentially abundant ASVs between roots of mock- and Hpa-inoculated Col-0 plants that were detected by at least 1 of these 5 methods (Fig. 3a). Among these 176 ASVs, we found a cluster of ASVs (Fig. 3a; cluster I) that occur in most samples but are enriched on the roots of Hpa-infected plants, and a smaller cluster of ASVs (cluster II) with frequent occurrence that are depleted on the roots of Hpa-infected plants. Moreover, another two clusters of ASVs are sparse in one of the two treatments, but occur frequently and abundantly in the other treatment. Also among these more sparse ASVs we identify ASVs that are enriched on the roots of Hpa-infected plants (cluster III) as well as ASVs that are depleted on roots of Hpa-infected plants (cluster IV).
In the conditioning phase, only Hpa-infected Col-0 plants, not the coumarin-deficient mutants myb72 and f6’h1, created a SBL that increased Hpa resistance in the second, responding population of plants (Fig. 1). Thus, to identify microbiota that are associated with this increased resistance, we subsequently examined the response of rhizosphere-associated bacterial taxa to foliar Hpa infection on roots of myb72 and f6’h1 plants. Of the 176 ASVs responding to Hpa infection on roots of Col-0 plants, 14 ASVs responded significantly and were similarly enriched or depleted on at least one of the coumarin-deficient mutants (Fig. 3b). Hence, it is unlikely that the microbes corresponding to these ASVs are part of the disease-induced SBL on Col-0 plants that induces resistance against Hpa in a subsequent population of plants. The remaining 162 ASVs differentially abundant on Col-0 plants were either not significantly responsive to Hpa infection or inversely responsive on either myb72, f6’h1 or both mutant genotypes. From these 162 ASVs, 100 ASVs were enriched on the roots of Hpa-infected Col-0 plants (Fig. 3b), thus potentially responsible for the increased Hpa resistance in the response population of Col-0 plants. These 100 ASVs represented 12 different bacterial phyla, of which most belonged to the phylum Proteobacteria. Together, these data show that a group of phylogenetically diverse microbes increased in abundance on the roots of foliar-infected Col-0 plants, while their abundances remained stable or depleted on roots of Hpa-infected coumarin biosynthesis mutants.
Differentially abundant bacterial ASVs in the rhizosphere of Hpa- and mock-treated Col-0, myb72 and f6’h1 plants in Experiment 1. (a) Heatmap showing the log-transformed relative abundance of 176 ASVs that are differentially abundant between mock- and Hpa-treated Col-0 plants. Relative abundances were normalized by the average relative abundance of that ASV across all mock-treated plant samples. Colors above heatmap indicate phylum annotation and statistical method(s) by which each differentially abundant ASV was identified. I-IV indicate clusters of ASVs with similar differential abundance patterns between Hpa- and mock-treated plants. (b) Fold change of the 176 Hpa-responding ASVs between Col-0, myb72 and f6’h1 plants. Shrunk fold change in relative abundance between mock- and Hpa-treated plants is shown as calculated with DESeq2. Colors above the heatmap indicate phylum annotation and whether an ASV was also differentially abundant between mock- and Hpa-treated myb72 and f6’h1 plants. “Filtered” ASVs were differentially abundant and similarly enriched or depleted on Col-0 as well as on at least one plant mutant genotype. Enriched and depleted ASVs are highlighted as clusters with positive or negative log2 fold change in Col-0, respectively. In both heatmaps, M indicates Massilia ASV 09023.
Consistent enrichment of ASVs across two experiments
The SBL is reproducibly generated by Hpa-infected plants in independent experiments, and thus it was further investigated if there is consistency in the identity of microbes that are recruited by the roots of Hpa-infected plants. We analyzed the root microbiome of an independent experiment (Experiment 2) that included both Col-0 and myb72 plants in the conditioning phase. This experiment was carried out in soil from the same Reijerscamp field but was collected several months after collection of the soil for the first experiment (October 2018 vs April 2019). After quality control and filtering, the 16S data comprised 32 samples with on average 54,842 reads per sample comprising a total of 2003 unique ASVs over all samples. In this second experiment, again a significant shift in the bacterial root microbiome was detected in response to foliar Hpa infection in both Col-0 and myb72 plants (P = 0.013; Fig. S4, Table S4, Data S2). In this experiment, the bacterial root communities of healthy myb72 plants were not significantly different from those on healthy Col-0 plants (P = 0.098), whereas the differences in bacterial root communities between Hpa-infected Col-0 and myb72 plants were significant (P = 0.004).
In this second experiment, 120 ASVs were differentially abundant between the roots of Hpa-infected and healthy Col-0 plants (Fig. 4a). Of these, 110 ASVs did not respond similarly to Hpa infection in myb72 mutant plants. Thirteen of the 110 MYB72-dependent ASVs were significantly affected by Hpa infection in both the first and the second experiment. One of the 13 steadily affected ASVs on roots of Hpa-infected Col-0 plants was consistently enriched, while 7 ASVs were consistently depleted. The remaining 5 ASVs responded in varying directions between the two experiments (Fig. S5). The only ASV that was consistently enriched on roots of Hpa-infected Col-0 plants was annotated as a Massilia spp. This Massilia ASV 09023 was present in almost all root microbiome samples of the first experiment (Fig. 3a) and increased in abundance on Col-0 roots following Hpa infection, whereas it was depleted on the roots of Hpa-infected coumarin-deficient mutant plants (Figs. 3b, 4c). In the Experiment 2, this ASV was detected on the roots of most of the Hpa-infected Col-0 plants (5 out of 6) but virtually absent from all myb72 and mock-treated wild-type plants (Fig. 4a,d). Together, the data of this second experiment confirms that there is a phylogenetically diverse set of rhizosphere microbes that changes in abundance in response to foliar Hpa infection in a coumarin-dependent manner. Within the rhizosphere microbial community responding to foliar Hpa infection, a Massilia spp. was consistently enriched.
Differentially abundant bacterial ASVs in the rhizosphere of Hpa- and mock-treated Col-0 and myb72 plants in Experiment 2. (a) Heatmap showing the log-transformed relative abundance of 120 ASVs that are differentially abundant between mock- and Hpa-treated Col-0 plants in Experiment 2. Relative abundances were normalized by the average relative abundance of that ASV across all mock-treated plant samples. Colors above heatmap indicate phylum annotation and statistical method(s) by which each differentially abundant ASV was identified. (b) Fold change of the 120 Hpa-responding ASVs between Col-0 and myb72 plants in Experiment 2. Shrunk fold change in relative abundance between mock- and Hpa-treated plants is shown as calculated with DESeq2. Colors above the heatmap indicate phylum annotation and whether an ASV was also differentially abundant between mock- and Hpa-treated myb72 plants. Filtered ASVs were differentially abundant and similarly enriched or depleted on Col-0 and myb72 plants. Enriched and depleted ASVs are highlighted as clusters with positive or negative log2 fold change in Col-0, respectively. In both heatmaps, M indicates Massilia ASV 09023. (c) Relative abundance of Massilia ASV 09023 on plants in Experiment 1. (d) Relative abundance of Massilia ASV 09023 on plants in Experiment 2.
Root exudate composition and exuded coumarins are significantly affected by foliar Hpa-infection
In contrast to Col-0, mutant myb72 and f6’h1 plants cannot create an Hpa-induced SBL that increases resistance in a response population of Col-0 plants (Fig. 1), suggesting that root-secreted coumarins are required for the creation of an effective SBL. To examine how Hpa-infection affects root exudation, we first analyzed sterile root exudates of healthy and Hpa-infected Col-0 plants with an untargeted metabolomics approach. UPLC-QTOF-MS detected 923 metabolite features that were visualized by PCA, revealing that Hpa infection clearly impacted root exudate patterns (Fig. 5a). Clustering analysis of 459 significant metabolite features (ANOVA, P < 0.01, FDR) further confirmed that root exudates of Hpa-infected plants differ significantly from those of healthy plants (Fig. 5b). Based on accurate m/z and fragmentation patterns, we were able to putatively annotate metabolite features that were significantly enriched in the root exudates of diseased plants, in which especially compounds that belong to phenolic biosynthesis pathways (e.g. the shikimate pathway) and derivatives of amino and organic acids accumulated (Data S3). Conversely, phenolic glycosides, coumaric acids and azelaic acid were depleted in the exudates of infected plants. Using this untargeted approach, we were unable to detect the known MYB72- and F6’H1-dependent coumarins esculetin, fraxetin and scopoletin. Therefore, we re-analyzed the sterile root exudates by HPLC with a SB-C18 column using commercially available standards of esculetin, fraxetin and scopoletin to target our methodology. However, we again could not detect these specific coumarins. This could mean that they are not produced under these circumstances, or that this method is not sensitive enough for their detection of these coumarins.
Next we used a coumarin-targeted UHPLC-TQ-MS–MS method to detect whether soil-grown Col-0 plants produce coumarins in response to Hpa infection. To this end, Col-0 plants were cultivated in natural Reijerscamp soil and inoculated with Hpa similarly as for a SBL assay. Seven days post inoculation, the pots were flushed with 5% ethanol after which the root exudates were analyzed for coumarin accumulation. We found that scopoletin accumulated in the root exudates, but not fraxetin and esculetin. Remarkably, we detected significantly more scopoletin in the root exudates of mock-inoculated plant than in that of Hpa-infected plants (Fig. 5c). We hypothesized that scopoletin may be more rapidly metabolized by the microbiome that is recruited by plants in response to foliar Hpa infection. We therefore assessed the levels of scopoletin secreted by plants growing on agar-solidified medium that were inoculated with monoxenic Hpa spores (Fig. 5d). Again significantly lower levels of scopoletin were detected in the substrate surrounding the roots of Hpa-inoculated plants than in that of healthy control plants. These results suggest that foliar Hpa infection results in less secretion of scopoletin by the roots. Together our data show that plants respond to foliar infection by altering their root exudation profiles. This response affects a large number of metabolites and while putative phenolics and derivatives of amino and organic acids accumulate around the roots of infected plants they secrete significantly less of the coumarin scopoletin.
Root exudate profile of Col-0 is affected by foliar Hpa infection. (a) Principle component analysis (PCA) of 923 metabolic profiles (see Data S3) of sterile HG solution without plants (yellow symbols) or with root exudates of Col-0 plants that were either mock-inoculated (red symbols) or of which the leaves were inoculated with Hpa (blue symbols). (b) Clustering analysis with heatmap (Pearson’s correlation with Ward clustering) of 459 metabolite markers with significantly different detected levels between the treatments (ANOVA, P < 0.01, FDR). Heatmap depicts normalized intensities (blue: depleted, red: accumulated). (c) Quantification of scopoletin in root exudates of mock-inoculated or Hpa-infected plants growing in natural Reijerscamp soil. As a control, scopoletin levels were measured in unplanted soil. Bars represent the average of 5 samples. (d) Quantification of scopoletin in root exudates of mock-inoculated or Hpa-infected plants growing in gnotobiotic conditions on agar-solidified HG medium. As a control, scopoletin was measured in unplanted agar plates. Bars represent the average of 5 samples. Letters indicate significant differences (P < 0.05 in ANOVA with Tukey’s posthoc test) between treatments. Error bars show standard error.
Perception of the soil-borne legacy of Hpa-infected Arabidopsis depends on SA and primes foliar defenses
Arabidopsis plants growing on natural Reijerscamp soil that has been pre-conditioned by a population of Hpa-infected Arabidopsis plants are more resistant to Hpa infection than plants growing on soil pre-conditioned by healthy plants (Fig. 1, Fig. S1)12. We hypothesized that the perception of such a SBL is the result of systemic resistance induced by microbes that are recruited by the infected plants. To test this, we first pre-conditioned Reijerscamp soil by growing Col-0 plants that were either infected by Hpa or not. Subsequently, we cultivated Col-0 and specific mutants impaired in known defense signaling pathways in the pre-conditioned soil and examined their level of resistance against Hpa infection. In the experimental setup, we used the myb72 mutant that is blocked in its capacity to develop SA-independent ISR in response to various beneficial rhizobacteria and fungi16. Additionally, we used the SA-response mutants npr1 and sid2 because several beneficial rhizobacteria and fungi have been shown to activate an SA-dependent ISR response also known as systemic acquired resistance (SAR)16,20. Both Col-0 and myb72 plants were significantly more resistant to Hpa infection when growing on soil that was pre-conditioned by Hpa-infected Col-0 plants than when growing on soil pre-conditioned by healthy Col-0 plants (Fig. 6a). However, neither npr1 nor sid2 mutant plants developed resistance against Hpa when grown on soil pre-conditioned by Hpa-infected Col-0 plants (Fig. 6b). This suggests that perception of the Hpa-mediated SBL involves systemic defense signaling in the response population of plants that depends on SA, but does not require MYB72.
To further corroborate this finding we monitored the expression of the SA-responsive marker gene PR-1 before and after Hpa infection in Col-0 plants growing in SBL-conditioned and non-conditioned Reijerscamp soil. Col-0 plants transcribed significantly more of the SA-responsive marker gene PR-1 in response to SA treatment when growing on soil pre-conditioned by diseased plants than on soil pre-conditioned by healthy plants (Fig. 6c). This suggests that defense responses of plants perceiving an Hpa-induced SBL are primed for SA-dependent defenses.
Perception of the Hpa-induced SBL is SA dependent and leads to priming of SA-dependent defense responses. (a) Disease severity of Hpa-inoculated Col-0 and myb72 plants (Responding population) growing in the on soil pre-conditioned with healthy (Mock) or Hpa-infected Col-0 plants. Bars depict the average spore production per gram of shoot fresh weight of 9–10 replicates. Error bars depict standard error of the mean. Asterisk denotes a significant difference (P < 0.05 in Students t-test). (b) Disease severity of Hpa-inoculated Col-0, npr1 and sid2 plants (Responding population) growing on soil pre-conditioned by healthy (Mock) or Hpa-infected Col-0 plants. Bars depict the average spore production per gram of shoot fresh weight of 20 replicates with values based on 2 independent experiments. Error bars depict standard error of the mean. Asterisk indicates a significant difference (univariate GLM with interaction term, P = 0.029). (c) PR-1 expression in leaves of Col-0 plants growing on soil pre-conditioned by healthy or diseased Col-0 plants at 30 min, 4 h, and 6 h after dipping in 1 mM SA or tap water (control). PR-1 expression is expressed as fold-change in SA-treated plants relative to mock-treated control plants growing on soil conditioned by healthy plants. Asterisk denotes a significant difference as determined by Student’s t-test (P < 0.05; n = 3).
Discussion
Recent studies show that plants can create a SBL in response to both biotic and abiotic stresses, resulting in the modulation of microbial communities in the rhizosphere that can act favorably on plant growth and immunity12,13,53,54. Here, we investigated biological mechanisms involved in the creation and perception of the SBL created by downy mildew-infected Arabidopsis plants.
Creation of the SBL requires the coumarin biosynthesis genes MYB72 and F6’H1
In our study, conditioning of a natural soil with Hpa-infected plants resulted in a SBL that consistently reduced disease severity in a subsequent Hpa-inoculated population of plants growing on this SBL-conditioned soil, compared to soil conditioned with healthy plants. We show that the coumarin biosynthesis genes MYB72 and F6’H1 are required for the creation of the Hpa-mediated SBL (Fig. 1) and identified MYB72- and F6’H1-dependent shifts in the rhizosphere microbial communities of Hpa-infected plants (Figs. 2–4) that may be responsible for the enhanced resistance of plants growing in SBL-conditioned soil. Hpa infection on the leaves of Col-0 plants resulted in dramatic metabolomic changes in the root exudates (Fig. 5), with increased levels of metabolite features representing a diversity of secondary and primary metabolites. Surprisingly, the MYB72- and F6’H1-dependent coumarins scopoletin, fraxetin and esculetin were not among them. For the main coumarin of Arabidopsis, scopeletin, we showed that its levels even decreased in root exudates of Hpa-infected plants. We, thus, hypothesize that either (1) a drop in coumarin secretion is responsible for the creation of the Hpa-induced SBL, allowing coumarin-sensitive microbes to increase in abundance, or (2) the tested coumarins scopoletin, fraxetin and esculetin are themselves not responsible for the creation of the Hpa-induced SBL, but other so-far unidentified MYB72- and F6’H1-dependent compounds play a role. An argument against hypothesis 1 is that myb72 and f6’h1 plants do not create a SBL that reduces Hpa disease severity in a response population of Col-0 plants to the extent that Hpa-infected Col-0 plants do (Fig. 1). Future, more detailed metabolite profiling of Hpa-infected Col-0, myb72, and f6’h1-plants should shed light on this matter.
Foliar Hpa infection confers a shift in bacterial microbiota in the rhizosphere
Analysis of the root microbiome showed that the rhizosphere bacterial community is significantly affected by foliar Hpa infection. Moreover, the majority of bacteria that became more abundant on roots of Col-0 plants in response to Hpa infection were not promoted on the roots of myb72 and f6’h1 mutant plants, again underscoring that MYB72 and F6’H1 are essential for the creation of the Hpa-induced SBL. While the root bacterial community on Col-0 roots was significantly affected by foliar Hpa infection (Table S2), the root fungal community was not (Table S3). This suggests that bacteria and not fungi are responsible for the Hpa-triggered SBL. We identified a taxonomically diverse group of bacterial ASVs of which the abundance on the root was significantly affected by foliar Hpa infection in two independent experiments. The Xanthomonas, Stenotrophomonas and Microbacterium spp. rhizobacterial strains that were identified as being responsive to foliar Hpa infection in our previous study12 were not among them. Also in the present study, the overlap between the bacterial communities that significantly responded to foliar Hpa infection was not large (13 of the 162 MYB72- and/or F6’H1-dependent ASVs in Experiment 1 and 110 ASVs in Experiment 2 overlapped between the two experiments (Fig. S5). This suggests that the Hpa-induced SBL is not mediated by a select set of defined microbial species, but merely represents a shift in taxa representing microbes that either alone or in concerted action with other microbes in the microbial community are able to confer enhanced resistance against Hpa infection in a subsequent population of plants. In our previous study (Berendsen et al.18), the identified and selected Xanthomonas, Stenotrophomonas and Microbacterium spp. strains were demonstrated as a consortium to enhance biofilm formation and to consistently confer enhanced resistance against Hpa infection. This suggests that this small consortium of microbes can be responsible for a Hpa-induced SBL that effectively enhances resistance in a next generation of plants. However, the present study shows that depending on the microbial context in the soil, other microbial taxa may as well be capable of creating an effective SBL.
One of the candidate microbial taxa that consistently emerged in this study as being enriched on roots of Hpa-infected Col-0 plants and not on coumarin-deficient mutant plants represents a Massilia sp. (Massilia ASV 09023; Figs. 3, 4 and Fig. S5). Massilia is a genus of bacteria that is abundantly found in soils. Moreover, increased abundance of Massilia has previously been associated with the induction of SA signaling-dependent resistance of rice plants against bacterial pathogen Xanthomonas oryzae55. Future research will be focused on identifying the Massilia sp. strain(s) involved in the creation of the SBL and investigate the potential for their use as a bioinoculant in agriculture.
Perception of the SBL induces a SA-dependent disease resistance
Because the Hpa-induced SBL leads to enhanced protection against a foliar pathogen, it is likely that this is conferred via rhizobacteria-mediated ISR. Activation of ISR by the well-characterized beneficial rhizosphere bacterium P. simiae WCS417 and several other beneficial root-associated microbes requires the transcription factor MYB72 and functions independently of SA16,21. Other beneficial microbes, however, have been found to systemically induce disease resistance that is SA-dependent56,57,58. Our results show that the Hpa-mediated SBL induces a systemic resistance in the responding population of plants that does not require MYB72, but rather depends on SA-dependent signaling and involves priming of SA-responsive defenses. Thus, collectively, our results show that downy mildew infection of a first population of plants consistently results in a MYB72/F6’H1-dependent shift in the bacterial rhizosphere microbiome that is capable of inducing SA-dependent defense priming in a subsequent response population of plants, leading to increased resistance to the pathogen that initiated the SBL. From an ecological and agricultural perspective, this would mean that a diseased plant can provide protection to a next generation of plants growing in the conditioned soil by creating a microbial SBL that induces systemic immunity against the attacking pathogen. Interestingly, it has been demonstrated that soil, that was taken from fields in Washington state that were suppressive to either take-all or Rhizoctonia root rot of wheat, induced resistance in Arabidopsis plants to foliar Pseudomonas syringae infection. This shows that, also in the field, soils conditioned by leaf-infected plants can trigger ISR in a subsequent generation of plants59. Elucidating the mechanistic basis of pathogen-induced SBLs for both the involvement of the plant and the recruited microbiome will be highly instrumental for the development of more sustainable microbiome-assisted agricultural practices.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Moreover, the raw sequence data generated of this study are available at https://www.ncbi.nlm.nih.gov/bioproject/838813.
References
Arif, I., Batool, M. & Schenk, P. M. Plant microbiome engineering: Expected benefits for improved crop growth and resilience. Trends Biotechnol. 38, 1385–1396 (2020).
Berendsen, R. L., Pieterse, C. M. J. & Bakker, P. A. H. M. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486. https://doi.org/10.1016/j.tplants.2012.04.001 (2012).
Compant, S., Samad, A., Faist, H. & Sessitsch, A. A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J. Adv. Res. 19, 29–37 (2019).
Oyserman, B. O., Medema, M. H. & Raaijmakers, J. M. Road MAPs to engineer host microbiomes. Curr. Opin. Microbiol. 43, 46–54 (2018).
Drigo, B. et al. Shifting carbon flow from roots into associated microbial communities in response to elevated atmospheric CO2. Proc. Natl. Acad. Sci. USA 107, 10938–10942 (2010).
Griffiths, R. I. et al. 13CO2 pulse labelling of plants in tandem with stable isotope probing: Methodological considerations for examining microbial function in the rhizosphere. J. Microbiol. Methods 58, 119–129 (2004).
Lundberg, D. S. & Teixeira, P. Root-exuded coumarin shapes the root microbiome. Proc. Natl. Acad. Sci. USA 115, 5629–5631. https://doi.org/10.1073/pnas.1805944115 (2018).
Stringlis, I. A. et al. MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc. Natl. Acad. Sci. USA 115, E5213–E5222. https://doi.org/10.1073/pnas.1722335115 (2018).
Sasse, J., Martinoia, E. & Northen, T. Feed your friends: Do plant exudates shape the root microbiome?. Trends Plant Sci. 23, 25–41 (2018).
Pascale, A., Proietti, S., Pantelides, I. S. & Stringlis, I. A. Modulation of the root microbiome by plant molecules: The basis for targeted disease suppression and plant growth promotion. Front. Plant Sci. 10, 1741 (2020).
Jacoby, R. P., Koprivova, A. & Kopriva, S. Pinpointing secondary metabolites that shape the composition and function of the plant microbiome. J. Exp. Bot. 72, 57–69. https://doi.org/10.1093/jxb/eraa424 (2020).
Berendsen, R. L. et al. Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J. 12, 1496. https://doi.org/10.1038/s41396-018-0093-1 (2018).
Yuan, J. et al. Root exudates drive the soil-borne legacy of aboveground pathogen infection. Microbiome 6, 156. https://doi.org/10.1186/s40168-018-0537-x (2018).
Bakker, P. A. H. M., Pieterse, C. M. J., De Jonge, R. & Berendsen, R. L. The soil-borne legacy. Cell 172, 1178–1180 (2018).
Carrión, V. J. et al. Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366, 606–612 (2019).
Pieterse, C. M. J. et al. Induced systemic resistance by beneficial microbes. Annu. Rev. Phytopathol. 52, 347–375. https://doi.org/10.1146/annurev-phyto-082712-102340 (2014).
Van Wees, S. C. M., Van der Ent, S. & Pieterse, C. M. J. Plant immune responses triggered by beneficial microbes. Curr. Opin. Plant Biol. 11, 443–448. https://doi.org/10.1016/j.pbi.2008.05.005 (2008).
Van Hulten, M., Pelser, M., Van Loon, L. C., Pieterse, C. M. J. & Ton, J. Costs and benefits of priming for defense in Arabidopsis. Proc. Natl. Acad. Sci. USA 103, 5602–5607 (2006).
Martinez-Medina, A. et al. Recognizing plant defense priming. Trends Plant Sci. 21, 818–822 (2016).
De Kesel, J. et al. The induced resistance lexicon: Do’s and dont’s. Trends Plant Sci. 26, 685–691. https://doi.org/10.1016/j.tplants.2021.01.001 (2021).
Van der Ent, S. et al. MYB72 is required in early signaling steps of rhizobacteria-induced systemic resistance in Arabidopsis. Plant Physiol. 146, 1293–1304. https://doi.org/10.1104/pp.107.113829 (2008).
Pieterse, C. M. J. et al. A novel signaling pathway controlling induced systemic resistance in Arabidopsis. Plant Cell 10, 1571–1580 (1998).
Ton, J., Davison, S., Van Loon, L. C. & Pieterse, C. M. J. Heritability of rhizobacteria-mediated induced systemic resistance and basal resistance in Arabidopsis. Eur. J. Plant Pathol. 107, 63–68 (2001).
Zamioudis, C., Hanson, J. & Pieterse, C. M. J. Beta-Glucosidase BGLU42 is a MYB72-dependent key regulator of rhizobacteria-induced systemic resistance and modulates iron deficiency responses in Arabidopsis roots. New Phytol. 204, 368–379. https://doi.org/10.1111/nph.12980 (2014).
Palmer, C. M., Hindt, M. N., Schmidt, H., Clemens, S. & Guerinot, M. L. MYB10 and MYB72 are required for growth under iron-limiting conditions. PLoS Genet. 9, e1003953 (2013).
Voges, M. J. E. E. E., Bai, Y., Schulze-Lefert, P. & Sattely, E. S. Plant-derived coumarins shape the composition of an Arabidopsis synthetic root microbiome. Proc. Natl. Acad. Sci. USA 116, 12558–12565. https://doi.org/10.1101/485581 (2018).
Li, E. et al. Rapid evolution of bacterial mutualism in the plant rhizosphere. Nat. Commun. 12, 3829 (2021).
Stassen, M. J., Hsu, S.-H., Pieterse, C. M. & Stringlis, I. A. Coumarin communication along the microbiome–root–shoot axis. Trends Plant Sci. 26, 169–183 (2021).
Vismans, G., Spooren, J., Pieterse, C. M. J., Bakker, P. A. H. M. & Berendsen, R. L. In The Plant Microbiome: Methods and Protocols. Methods in Molecular Biology (eds Carvalhais, L. C. & Dennis, P. G.) 209–218 (Springer, 2021).
Asai, S., Shirasu, K. & Jones, J. D. G. Hyaloperonospora arabidopsidis (Downey mildew) infection assay in Arabidopsis. Bio-Protocols 5, e1627. https://doi.org/10.21769/BioProtoc.1627 (2015).
Schmid, N. B. et al. Feruloyl-CoA 6′-Hydroxylase1-dependent coumarins mediate iron acquisition from alkaline substrates in Arabidopsis. Plant Physiol. 164, 160–172 (2014).
Cao, H., Glazebrook, J., Clarke, J. D., Volko, S. & Dong, X. The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell 88, 57–63 (1997).
Wildermuth, M. C., Dewdney, J., Wu, G. & Ausubel, F. M. Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414, 562–565 (2001).
Oñate-Sánchez, L. & Vicente-Carbajosa, J. DNA-free RNA isolation protocols for Arabidopsis thaliana, including seeds and siliques. BMC Res. Notes 1, 1–7 (2008).
Van Wees, S. C. M., Van Pelt, J. A., Bakker, P. A. H. M. & Pieterse, C. M. J. Bioassays for assessing jasmonate-dependent defenses triggered by pathogens, herbivorous insects, or beneficial rhizobacteria. Methods Mol. Biol. 1011, 35–49 (2013).
Illumina. 16S Metagenomic Sequencing Library Preparation, https://support.illumina.com/ (2013).
De Muinck, E. J., Trosvik, P., Gilfillan, G. D., Hov, J. R. & Sundaram, A. Y. M. A novel ultra high-throughput 16S rRNA gene amplicon sequencing library preparation method for the Illumina HiSeq platform. Microbiome 5, 68. https://doi.org/10.1186/s40168-017-0279-1 (2017).
Baym, M. et al. Inexpensive multiplexed library preparation for megabase-sized genomes. PLoS ONE 10, e0128036. https://doi.org/10.1371/journal.pone.0128036 (2015).
Ihrmark, K. et al. New primers to amplify the fungal ITS2 region—evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 82, 666–677. https://doi.org/10.1111/j.1574-6941.2012.01437.x (2012).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857. https://doi.org/10.1038/s41587-019-0209-9 (2019).
Clarke, K. R. Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 18, 117–143. https://doi.org/10.1111/j.1442-9993.1993.tb00438.x (1993).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514. https://doi.org/10.1038/s41467-020-17041-7 (2020).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217. https://doi.org/10.1371/journal.pone.0061217 (2013).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis 2nd edn. (Springer, 2016).
Song, Y., Pieterse, C. M. J., Bakker, P. A. H. M. & Berendsen, R. L. In The Plant Microbiome: Methods and Protocols (eds Carvalhais, L. C. & Dennis, P. G.) 305–317 (Springer, 2021).
Pétriacq, P. et al. Metabolite profiling of non-sterile rhizosphere soil. Plant J. 92, 147–162. https://doi.org/10.1111/tpj.13639 (2017).
Tsugawa, H. et al. MS-DIAL: Data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526. https://doi.org/10.1038/nmeth.3393 (2015).
Pang, Z. et al. MetaboAnalyst 5.0: Narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 49, W388–W396. https://doi.org/10.1093/nar/gkab382 (2021).
Stringlis, I. A., de Jonge, R. & Pieterse, C. M. J. The age of coumarins in plant-microbe interactions. Plant Cell Physiol. 60, 1405–1419. https://doi.org/10.1093/pcp/pcz076 (2019).
Weiss, S. et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5, 27. https://doi.org/10.1186/s40168-017-0237-y (2017).
Nearing, J. T. et al. Microbiome differential abundance methods produce different results across 38 datasets. Nat. Commun. 13, 342. https://doi.org/10.1038/s41467-022-28034-z (2022).
Li, H., La, S., Zhang, X., Gao, L. & Tian, Y. Salt-induced recruitment of specific root-associated bacterial consortium capable of enhancing plant adaptability to salt stress. ISME J. 15, 2865–2882. https://doi.org/10.1038/s41396-021-00974-2 (2021).
Santos-Medellín, C. et al. Prolonged drought imparts lasting compositional changes to the rice root microbiome. Nat. Plants 7, 1065–1077. https://doi.org/10.1038/s41477-021-00967-1 (2021).
Chen, S. et al. Diclofop-methyl affects microbial rhizosphere community and induces systemic acquired resistance in rice. J. Environ. Sci. 51, 352–360. https://doi.org/10.1016/j.jes.2016.06.027 (2017).
Van de Mortel, J. E. et al. Metabolic and transcriptomic changes induced in Arabidopsis by the rhizobacterium Pseudomonas fluorescens SS101. Plant Physiol. 160, 2173–2188 (2012).
De Vleesschauwer, D. & Höfte, M. Rhizobacteria-induced systemic resistance. Adv. Bot. Res. 51, 223–281 (2009).
Pieterse, C. M. J. et al. Pseudomonas simiae WCS417: Star track of a model beneficial rhizobacterium. Plant Soil 461, 245–263 (2021).
Weller, D. M., van Pelt, J. A., Pieterse, C. M. J. & Bakker, P. A. H. M. Disease-suppressive soils induce systemic resistance in Arabidopsis thaliana against Pseudomonas syringae pv. tomato. Phytopathology 105, 147 (2015).
Acknowledgements
We would like to thank Annemiek van Andel for propagation of Hpa in planta, and Natuurmonumenten for permitting the sampling of soil in the Reijerscamp nature reserve.
Funding
This work was supported by the Dutch Research Council (NWO) through Perspective program “Back2Roots” (grant no. 14219), the Industrial Doctorates program (project no. NWA.ID.17.040), the Gravitation program MiCRop (grant no. 024.004.014), and XL program “Unwiring beneficial functions and regulatory networks in the plant endosphere” (grant no. OCENW.GROOT.2019.063), by Foundation TKI Horticulture (grant no. 1605–106), and by the China Scholarship Council.
Author information
Authors and Affiliations
Contributions
G.V., Sv.B., J.S., Y.S., C.M.J.P., P.A.H.M.B. and R.L.B. designed the experiments. G.V., Sv.B., J.S., M.S., Y.S. performed the experiments. G.V., Sv.B., P.G., B.L.S. analyzed the microbiome data. J.V., B.T., L.D., H.J.B. and P.P. performed and supervised the chemical analyses. G.V., Sv.B., J.S., Y.S., C.M.J.P., P.A.H.M.B. and R.L.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vismans, G., van Bentum, S., Spooren, J. et al. Coumarin biosynthesis genes are required after foliar pathogen infection for the creation of a microbial soil-borne legacy that primes plants for SA-dependent defenses. Sci Rep 12, 22473 (2022). https://doi.org/10.1038/s41598-022-26551-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26551-x
- Springer Nature Limited
This article is cited by
-
Combined metabolome and transcriptome reveal HmF6’H1 regulating simple coumarin accumulation against powdery mildew infection in Heracleum moellendorffii Hance
BMC Plant Biology (2024)
-
Nonpathogenic Pseudomonas syringae derivatives and its metabolites trigger the plant “cry for help” response to assemble disease suppressing and growth promoting rhizomicrobiome
Nature Communications (2024)
-
MAPK Cascades in Plant Microbiota Structure and Functioning
Journal of Microbiology (2024)
-
Obligate biotroph downy mildew consistently induces near-identical protective microbiomes in Arabidopsis thaliana
Nature Microbiology (2023)