Abstract
Tomato (Solanum lycopersicum L.) is a major Solanaceae crop worldwide and is vulnerable to bacterial wilt (BW) caused by Ralstonia solanacearum during the production process. BW has become a growing concern that could enormously deplete the tomato yield from 50 to 100% and decrease the quality. Research on the molecular mechanism of tomato regulating BW resistance is still limited. In this study, two tomato inbred lines (Hm 2–2, resistant to BW; and BY 1–2, susceptible to BW) were used to explore the molecular mechanism of tomato in response to R. solanacearum infection by RNA-sequencing (RNA-seq) technology. We identified 1923 differentially expressed genes (DEGs) between Hm 2–2 and BY 1–2 after R. solanacearum inoculation. Among these DEGs, 828 were up-regulated while 1095 were down-regulated in R-3dpi (Hm 2–2 at 3 days post-inoculation with R. solanacearum) vs. R-mock (mock-inoculated Hm 2–2); 1087 and 2187 were up- and down-regulated, respectively, in S-3dpi (BY 1–2 at 3 days post-inoculation with R. solanacearum) vs. S-mock (mock-inoculated BY 1–2). Moreover, Gene Ontology (GO) enrichment analysis revealed that the largest amount of DEGs were annotated with the Biological Process terms, followed by Cellular Component and Molecular Function terms. A total of 114, 124, 85, and 89 regulated (or altered) pathways were identified in R-3dpi vs. R-mock, S-3dpi vs. S-mock, R-mock vs. S-mock, and R-3dpi vs. S-3dpi comparisons, respectively, by Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway analysis. These clarified the molecular function and resistance pathways of DEGs. Furthermore, quantitative RT-PCR (qRT-PCR) analysis confirmed the expression patterns of eight randomly selected DEGs, which suggested that the RNA-seq results were reliable. Subsequently, in order to further verify the reliability of the transcriptome data and the accuracy of qRT-PCR results, WRKY75, one of the eight DEGs was silenced by virus-induced gene silencing (VIGS) and the defense response of plants to R. solanacearum infection was analyzed. In conclusion, the findings of this study provide profound insight into the potential mechanism of tomato in response to R. solanacearum infection, which lays an important foundation for future studies on BW.
Similar content being viewed by others
Introduction
Tomato (Solanum lycopersicum L.) is one of the majorly consumed solanaceous vegetable crops, with a global annual yield of approximately 160 million tons1,2. It is cultivated worldwide for fresh vegetable consumption as well as for industrial processing3,4. Tomatoes are nutritionally significant that can provide vitamins, fibers, and minerals. They are essential sources of β-carotene and lycopene possessing antioxidant and have anti-cancer properties5. A large part of the tomato crop has been grown continuously for many years in the world. Continuous tomato cultivation in China has led to an outbreak of the devastating soil-borne disease "bacterial wilt" (BW), caused by virulent Ralstonia solanacearum. BW can enormously cause the production loss by 50–100% every year, and it has become one of the main diseases that seriously threaten the yield and quality of tomato6,7,8,9. BW is one of the most serious plant diseases in the world10,11. It is a typical vascular disease that harms the roots, stems, and leaves. BW has a rapid onset, withered leaves on the diseased side, wilted leaves, and striped diseased spots at the base of the stem. In severe cases, the roots would turn black and rot, and the pith would be hollow or honeycomb. When the diseased plant is cut open, the fibrous tube tissue inside will turn brown. When the transverse cut of the diseased plant base is pressed hard, the yellow-white bacterial mucus will flow out of the fracture, which is known as "bacteria pus"12,13. Bacterial wilt, known as plant cancer, seriously affects the yield and quality of crops. It has strong variability and soil-borne characteristics. The current traditional control methods, such as breeding resistant varieties, crop rotation and chemical control have some limitations14,15,16. Therefore, it is necessary to have a comprehensive and detailed understanding of plant–pathogen interactions during the progression of BW17.
The current research on tomato resistance to BW mainly focuses on the genetic mechanism of resistance18,19,20,21; identification20,22,23 and screening13,24,25,26,27,28,29 of molecular markers related to the disease resistance genes and other aspects. There are only a few studies available on the gene regulation of BW resistance30,31,32,33,34,35,36,37, and the molecular mechanism of tomato resistance-related genes regulating BW remains unclear. Thus, it necessitates performing the research on the response of tomato plant to BW. In recent years, with the application of transcriptome sequencing technology, several genes and miRNA functions have been identified. Transcriptome sequencing has been widely used in basic research, molecular breeding, pathogen-host interaction mechanism, comparison of resistance genes between susceptible and disease-resistant varieties, and biocontrol factors inducing plant disease-resistance mechanisms38,39,40. Transcriptomic technique has been used in many studies to identify the molecular mechanism of R. solanacearum resistance in a plethora of plant species, including Solanum dulcamara41, Arachis hypogaea42, Solanum commersonii43, Solanum melongena44, Capsicum annuum45, Solanum tuberosum46, and Nicotiana tabacum47. In tomato, the in-depth transcriptome data is available for its interaction with R. solanacearum15,17,48,49,50. French et al.48 analyzed the genome-wide transcriptional response of roots of resistant and susceptible tomato plants at multiple time points after inoculation with R. solanacearum and identified the molecular basis of this resistance. Furthermore, they determined the role of auxin signaling and transport pathways in resistance against R. solanacearum by functional analysis of an auxin transport tomato mutant. Jiang et al.49 investigated the root transcriptome profiles between silicon (Si)-treated (+ Si) and untreated (− Si) tomato plants at different days post-inoculation with R. solanacearum by using RNA-seq technology. They also determined the content of hormones including salicylic acid (SA), jasmonic acid (JA), and ethylene (ET). Finally, they suggested that Si enhanced BW resistance of tomato via several signaling pathol.ways. The molecular mechanism of tomato resistance-related genes regulating bacterial wilt resistance is still unclear. Therefore, it is necessary to screen out the genes related to tomato BW resistance through transcriptome analysis.
In this study, we used RNA-seq technology to analyze the transcriptome of two tomato inbred lines resistant to BW Hm 2–2 (R) and susceptible to BW BY 1–2 (S) before and after inoculation with R. solanacearum. The two tomato inbred lines are special and the inductive properties are stable. In addition, studies comfirmed that Ralstonia solanacearum can multiply in plant stems. In this study, tomato stems were collected and studied, while tomato roots were studied in most previous studies. This study aimed to determine the molecular mechanism involved in the tomato response towards R. solanacearum, and provide a theoretical foundation for future research in BW.
Results
Phenotypic characterization after inoculation with R. solanacearum
At 3 dpi (days post-inoculation), plants exhibited different phenotypic symptoms. As shown in Fig. 1, leaves of the resistant plants (Hm 2–2) and susceptible (BY 1–2) plants inoculated with sterile water showed no obvious symptoms. However, one or two leaves of the resistant plants (Hm 2–2) inoculated with pathogen showed wilting symptoms, but almost all the leaves of the susceptible (BY 1–2) plants inoculated with pathogen showed wilting symptoms. These results indicate that tomato Hm 2–2 and BY 1–2 plants respond differently to R. solanacearum infection.
Phenotypic symptoms of resistant (Hm 2–2) and susceptible (BY 1–2) tomato seedlings 3 days after Ralstonia solanacearum inoculation. R-mock represents mock-inoculated Hm 2–2 plants; R-3dpi represents 3 days post-pathogen-inoculated Hm 2–2 plants; S-mock represents mock-inoculated BY 1–2 plants; S-3dpi represents 3 days post-pathogen-inoculated BY 1–2 plants. The red arrows represent wilting symptoms of tomato leaves.
Analysis of RNA-seq data
To obtain a global overview of the transcriptome relevant to BW stress conditions in the resistant and susceptible tomato plants, the cDNA libraries from stem samples of Hm 2–2 and BY 1–2 plants with mock-inoculation and pathogen-inoculation were sequenced on the Illumina HiSeq2500 platform, separately. After removal of the adaptors and filtration of low-quality reads, an average of 3.3 × 107 clean reads were obtained for each sample, and clean reads from all twelve libraries were mapped to the tomato genome ITAG3.2. The coverage of overall mapped reads to all the samples ranged between 96–97% and the percentage of uniquely mapped reads ranged between 76 and 85%. Moreover, the percentage of bases with Q20 (high sequencing quality) was close to 98%. Detailed sequencing and mapping statistics are given in Supplementary Table S1. As shown in Supplementary Fig. S1, gene coverage ranged from 80 to 100%, accounting for approximately 75% of the total genes (Supplementary Fig. S1a). An experimental repeatability test was performed, the results of which indicated that the results between replicates were comparable (Supplementary Fig. S1b). These results showed that the transcriptome sequencing quality was appropriate for further analyses.
Identification of DEGs
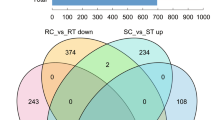
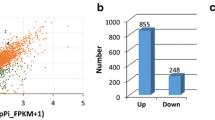
Differential expression analysis was performed by the DESeq2 package between two groups (treatments and control). The genes with FDR < 0.05 and |log2FC|≥ 1 were considered DEGs. The number of DEGs was different in each of the four comparisons (Fig. 2a). The most DEGs were found in S-3dpi vs. S-mock (1087 and 2187 significantly up- and down-regulated genes, respectively), followed by R-3dpi vs. R-mock (828 and 1095 significantly up- and down-regulated genes, respectively) and R-mock vs. S-mock (386 and 258 significantly up- and down-regulated genes, respectively), and the least DEGs were seen in R-3dpi vs. S-3dpi (337 and 263 significantly up- and down-regulated genes, respectively). In R-3dpi vs. R-mock and S-3dpi vs. S-mock, the total number of down-regulated genes was greater than that of up-regulated genes. While in R-mock vs. S-mock and R-3dpi vs. S-3dpi, the total number of up-regulated genes was greater than that of down-regulated genes. The volcano plots showed that the number of up- and down-regulated genes had a clear distribution pattern in all four comparisons (Fig. 2b). Moreover, the distribution pattern of down-regulated genes in S-3dpi vs. S-mock was much higher than that in R-mock vs. S-mock and R-3dpi vs. S-3dpi. Although the numbers of up- and down-regulated genes in R-mock vs. S-mock and R-3dpi vs. S-3dpi were lower compared with those of the other two comparisons, the distribution patterns were very similar. These results clearly showed the global gene expression patterns between different comparisons.
Differential expression analysis between treatments. (a) Comparison of the number of up- and down-regulated genes. Yellow and blue points represent up- and down-regulated genes, respectively. (b) Volcano plots between treatments and control. Red and blue points represent up- and down-regulated genes, respectively. Gray points represent no differential expression between genes. R-mock represents mock-inoculated Hm 2–2 plants; R-3dpi represents 3 days post-pathogen-inoculated Hm 2–2 plants; S-mock represents mock-inoculated BY 1–2 plants; S-3dpi represents 3 days post-pathogen-inoculated BY 1–2 plants.
The heat map analysis based on hierarchical clustering analysis of DEGs in four comparisons showed that DEGs in different comparisons showed different expression trends. Among the top 50 highly correlated genes across samples, 31/19 clustered together to reveal higher up-/down-regulating trends in R-3dpi vs. R-mock and S-3dpi vs. S-mock comparisons. Most of the top 50 highly correlated genes exhibited little difference between R-mock vs. S-mock and R-3dpi vs. S-3dpi (Fig. 3). The expression of the top 50 DEGs in different comparisons is listed in Supplementary Table S3.
Heat map showing a hierarchical cluster analysis of the top 50 highly expressed genes in four comparisons. The gradient scale represents expression levels with yellow indicating the highest expression and blue indicating the lowest expression. R-mock represents mock-inoculated Hm 2–2 plants; R-3dpi represents 3 days post-pathogen-inoculated Hm 2–2 plants; S-mock represents mock-inoculated BY 1–2 plants; S-3dpi represents 3 days post-pathogen-inoculated BY 1–2 plants.
SNP and InDel annotations
Variation detection based on SNPs and InDels of transcriptome sequencing was performed by the software GATKv3.4-46 (Fig. 4). Two types of nonsynonymous single nucleotide variation (SNV) and synonymous SNV represented dominant functional variations among the nine types of functional variations (Fig. 4a). The remaining types were frameshift deletion, stop gain, unknown, non-frameshift deletion stoploss, frameshift insertion, and non-frameshift insertion. Furthermore, these SNPs were distributed in different locations, with dominant distribution in intronic regions, followed by intergenic and exonic regions (Fig. 4b). We also identified all the possible mutations in this study. The two dominant types were transition and transversion (Fig. 4c), where transition accounted for 61.44%, while transversion accounted for 38.56%. These results suggest systematic, comprehensive, transcriptional regulation in tomato that orchestrates the response to BW.
GO enrichment analysis of DEGs
GO enrichment analyses were performed for all significant DEGs (Fig. 5). Different comparisons showed similar distribution patterns in terms of the number and type of enriched GO terms, which could be divided into three main functional groups, including 22 biological processes, 14 molecular functions, and 15 cellular components (Fig. 5a). The largest amount of DEGs were annotated with the biological process terms, with the cellular process, metabolic process, and single-organism process accounting for the largest number of DEGs. In molecular function terms, the majority of DEGs were associated with binding and catalytic activities. Other significantly abundant cellular component terms were cell, cell part, membrane, organelle, and membrane parts (Supplementary Table S4). However, the enrichment level (Q-value) for each functional comparison varied (Fig. 5b). Notably, other functional comparisons that were significantly enriched were involved in different cellular component pathways, such as extracellular region, cytoskeleton, microtubule cytoskeleton, external encapsulating structure, and cell periphery processes (Fig. 5b).
GO enrichment analyses of DEGs in four comparisons (R-3dpi vs. R-mock, R-mock vs. S-mock, R-3dpi vs. S-3dpi, and S-3dpi vs. S-mock). (a) Summary of the distribution and number of DEGs in three ontology classes, including biological process, molecular function, and cellular component. (b) Q-value heat map of the GO enrichment of the three main ontology classes. The color scale indicates the Q-value. Darker coloration indicates more significant enrichment. R-mock represents mock-inoculated Hm 2–2 plants; R-3dpi represents 3 days post-pathogen-inoculated Hm 2–2 plants; S-mock represents mock-inoculated BY 1–2 plants; S-3dpi represents 3 days post-pathogen-inoculated BY 1–2 plants.
KEGG pathway analysis of DEGs
It is known that genes do not perform independently in any organism. They often coordinate and interact with each other to jointly regulate various life activities of the organism. The KEGG enrichment analysis was used to investigate the major pathways that the DEGs participated in. A total of 114, 124, 85, and 89 regulated (or altered) pathways in R-3dpi vs. R-mock, S-3dpi vs. S-mock, R-mock vs. S-mock, and R-3dpi vs. S-3dpi were identified, respectively (Fig. 6a and Supplementary Table S5–S8). The largest numbers of DEGs belonging to the top 10 KEGG pathways are shown in Fig. 6b. For R-3dpi vs. R-mock, more DEGs were enriched in the “Metabolic pathways” (236 DEGs), “Biosynthesis of secondary metabolites” (162 DEGs), “Plant-pathogen interaction” (36 DEGs), “Phenylpropanoid biosynthesis” (32 DEGs) and “Plant hormone signal transduction” (31 DEGs) pathways. Out of the 114 pathways, 24 had p-values < 0.05 (Supplementary Table S5). For S-3dpi vs. S-mock, most of the DEGs were enriched in “Metabolic pathways” (387 DEGs), followed by the “Biosynthesis of secondary metabolites” (249 DEGs), “Plant hormone signal transduction” (57 DEGs), “Carbon metabolism” (55 DEGs), and “Plant-pathogen interaction” (55 DEGs) pathways. On the other hand, out of the 124 pathways, 42 had p-values < 0.05 (Supplementary Table S6). In R-mock vs. S-mock, “Metabolic pathways” (57 DEGs), “Biosynthesis of secondary metabolites” (39 DEGs), “Plant-pathogen interaction” (11 DEGs), “Carbon metabolism” (10 DEGs), and “Plant hormone signal transduction” (9 DEGs) pathways showed the greatest enrichment. However, only 11 pathways had p-values < 0.05 (Supplementary Table S7). In R-3dpi vs. S-3dpi, “Metabolic pathways” (52 DEGs), “Biosynthesis of secondary metabolites” (32 DEGs), “Carbon metabolism” (9 DEGs), “Plant-pathogen interaction” (9 DEGs), and “Plant hormone signal transduction” (7 DEGs) showed the greatest enrichment; nine pathways had p-values < 0.05 (Supplementary Table S8). In addition, the Q-value of the KEGG enrichment analysis indicated that the largest number of enriched DEGs were involved in the biosynthesis of secondary metabolites, although the degree of enrichment might not be the highest compared with the other top enriched pathways (Fig. 6c).
KEGG enrichment analysis of DEGs. (a) KEGG enrichment analyses in different comparisons. (b) The top ten KEGG pathways containing the largest number of DEGs in R-3dpi vs. R-mock, S-3dpi vs. S-mock, R-mock vs. S-mock, and R-3dpi vs. S-3dpi comparisons. (c) Q-value heat map of KEGG enrichment. R-mock represents mock-inoculated Hm 2–2 plants; R-3dpi represents 3 days post-pathogen-inoculated Hm 2–2 plants; S-mock represents mock-inoculated BY 1–2 plants; S-3dpi represents 3 days post-pathogen-inoculated BY 1–2 plants.
Plant–pathogen interaction pathways
Previous studies have concluded that increased Ca2+-dependent cyclic nucleotide-gated channels (CNGCs) play a key role in plant response to pathogens and pathogen-associated molecular pattern (PAMP) signals51. As shown in Supplementary Fig. S2a, the expression levels of CNGCs and HSP90 were up-regulated in R-mock vs. S-mock, but after R. solanacearum inoculation, their expression levels were significantly down-regulated (Supplementary Fig. S2b) in R-3dpi vs. S-3dpi. Furthermore, the CERK, MEKK1, Rboh, and PBS1 genes were down-regulated, but MYC2, PR1, and RIN4 were up-regulated in R-3dpi vs. S-3dpi. These results can help understand the resistance mechanism of tomato to R. solanacearum.
Plant hormone signal transduction pathways
Most of the plant hormone signal transduction pathways were represented by DEGs in both genotypes with only a few differences. Auxin responsive AUX/IAA and SAUR were found up-regulated in both R-mock vs. S-mock and R-3dpi vs. S-3dpi. Cytokine responsive B-ARR was found up-regulated in R-mock vs. S-mock but not significantly changed in R-3dpi vs. S-3dpi comparison. Moreover, AHP was found down-regulated in R-3dpi vs. S-3dpi but no significant change in AHP expression was found in R-mock vs. S-mock. The TGA transcription factor, a regulator of NPR1, was found up-regulated in both R-mock vs. S-mock and R-3dpi vs. S-3dpi. Furthermore, the downstream TGA and PR1 genes that are possibly responsible for disease resistance were up-regulated in both R-mock vs. S-mock and R-3dpi vs. S-3dpi (Supplementary Fig. S3).
MAPK signaling pathways
A total of seven spots were mapped on the mitogen-activated protein kinase (MAPK) signaling pathway (Supplementary Fig. S4). EBF1/2 genes in ethylene-responsive defense response were found down-regulated in both R-mock vs. S-mock and R-3dpi vs. S-3dpi comparisons, but interestingly, ChiB that was up-regulated in R-mock vs. S-mock showed down-regulation in R-3dpi vs. S-3dpi. Moreover, EIN3/EIL was found up-regulated in R-mock vs. S-mock but not significantly changed in R-3dpi vs. S-3dpi. Few plant-defense-related components, such as PR1 was found up-regulated, whereas EBF1/2 were commonly down-regulated in both R-mock vs. S-mock and R-3dpi vs. S-3dpi. Similarly, FLS2 was found down-regulated in both R-mock vs. S-mock and R-3dpi vs. S-3dpi. MPKKK17/18 were found up-regulated in R-3dpi vs. S-3dpi but not significantly changed in R-mock vs. R-mock. Furthermore, cell death-related RbohD was found down-regulated in R-3dpi vs. S-3dpi but not significantly changed in R-mock vs. S-mock (Supplementary Fig. S4).
Validation of RNA-seq data by qRT-PCR analysis
To verify the reliability of Illumina sequencing results, eight genes were randomly selected for qRT-PCR analysis with three biological replicates. The expression levels of these genes were normalized to the constitutively expressed Actin gene. The genes including 3-ketoacyl-CoA synthase 6 (KCS6, Solyc05g009270.3), calcium-dependent protein kinase CDPK1 (CDPK1, Solyc07g064610.3), and transcription factor MYB86-like (MYB86, Solyc06g071690.3) were found down-regulated after R. solanacearum inoculation. But cysteine protease RD19D (RD19D, Solyc11g008260.2) gene, PR1 protein precursor (PR1, Solyc01g106620.2) gene, pathogenesis-related leaf protein 6 precursor (PR6, Solyc00g174340.2) gene, WRKY transcription factor 75 (WRKY75, Solyc05g015850.3) gene, and cyclic nucleotide-gated ion channel 2 isoform X2 (CNGC2, Solyc02g088560.3) gene were up-regulated after R. solanacearum inoculation. The verification result is shown in Fig. 7, and the primers for qRT-PCR are given in Supplementary Table S2. Overall, the results obtained from qRT-PCR and RNA-seq showed the same up- and down-regulation trends, which suggested that the RNA-seq results were reliable.
Quantitative real-time PCR validation. (a) In the R-3dpi vs. R-mock comparison. (b) In the S-3dpi vs. S-mock comparison. The red column represents RNA-seq data, and the blue column represents qRT-PCR data. The experiments were performed in triplicate. R-mock represents mock-inoculated Hm 2–2 plants; R-3dpi represents 3 days post-pathogen-inoculated Hm 2–2 plants; S-mock represents mock-inoculated BY 1–2 plants; S-3dpi represents 3 days post-pathogen-inoculated BY 1–2 plants.
Silencing of WRKY75 in tomato leads to decreased resistance to R. solanacearum
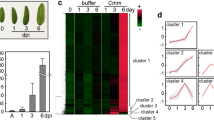
In order to further verify the reliability of the transcriptome data and the accuracy of qRT-PCR results, WRKY75, one of the eight DEGs was randomly selected for functional verification. Virus-induced gene silencing (VIGS) was used to verify the function of WRKY75 gene. At 7 dpi, the resistant Hm 2–2 tomato plants inoculated with TRV::WRKY75 showed wilting symptoms, while plants inoculated with TRV::empty showed little wilting symptoms in their lower leaves, as shown in Fig. 8a. Figure 8b shows the expression pattern of WRKY75 in VIGS plants. We evaluated the pathogenicity of the WRKY75-silencing plants in comparison with the wild-type Hm 2–2 plants (Fig. 8c). The result showed that the disease rating of WRKY75-silencing plants was greatly increased compared with the wild-type plants.
Resistance identification of tomato bacterial wilt after silencing WRKY75 gene. (a) Phenotypic symtoms after inoculation with Ralstonia solanacearum of wild-type Hm 2–2 (TRV::empty) and silencing WRKY75 (TRV::WRKY75) tomato plants; (b) qRT-PCR of WRKY75 gene; (c) Disease scoring after infection with R. solanacearum in wild-type Hm 2–2 (dark gray) and silencing WRKY75 (light gray) tomato plants. Results are averages ± s.e.m. (n = 20). *P < 0.05 using Student’s t-test. We repeated all experiments at least three times with similar results.
Discussion
RNA-seq technology, namely “whole transcriptome shotgun sequencing” technology, is an effective method for comprehensive analysis of the entire transcriptome through high-throughput sequencing52. Compared with the conventional hybridization technique, RNA-seq technology, without advanced probe design, can test all transcription information for any species at the single nucleotide level, and it has gradually become a new technology of genome and transcriptome research due to its advantages including high-throughput, easy to operate, can be quantitative, and low cost. It is widely used in differential gene detection, new gene identification, and gene function analysis. As a next generation sequencing technology, RNA-seq technology has also achieved a great breakthrough in the study of plant disease resistance mechanisms. Strau et al.53 identified Bs4C, a candidate gene responsible for Xanthomonas resistance from pepper by RNA-seq technology. Bs4C can regulate the recognition of Xanthomonas transcription activator like effectors (TALE) Avr Bs4. To assess the susceptible response of apple to the fire blight pathogen Erwinia amylovora, Kamber et al.54 found that 1080 transcripts were differentially expressed at 48 h post-inoculation with E. amylovora through analyzing RNA-seq data from challenged and mock-inoculated flowers. Li et al.55 found 1196 and 358 defense-related genes were DEGs in resistant and susceptible plants, respectively, through deep RNA-seq analysis. Dasgupta et al.56 performed RNA-seq analysis between two genotypes containing PMR-1 (resistant) and Pusa Vishal (susceptible) mungbeans in response to yellow mosaic virus. The resistance to mungbean yellow mosaic India virus (MYMIV) showed a very complicated gene network, starting from the production of general PAMPs, and activating various signaling cascades such as brassinosteroid (BR), jasmonic acid (JA), and kinases. Finally, the expression of specific genes (such as R-gene proteins, virus resistance, and PR-proteins) leads to disease resistance response. In this study, we identified 1923 and 3274 DEGs in R-3dpi vs. R-mock and S-3dpi vs. S-mock comparisons, respectively, by using RNA-seq technology. The obtained results revealed that these DEGs were involved in plant metabolism, signal transduction, synthesis of secondary metabolites, genetic information processing, responses to environmental stimuli, self-immunity, and other life activities. These results showed that the quality of the transcriptome sequencing output and assembly that we obtained meets the requirements of transcriptome analysis. The use of RNA-seq technology can yield very comprehensive and abundant transcription information, and that can provide us with abundant resources and paths for future research.
Plant hormones are a class of organic substances produced by the plant itself. They play an important role in regulating various crucial activities including growth, development, senescence, and death at very low concentrations. Relevant studies have shown that SA, JA, and ET are the crucial signal molecules that induce plants to produce defense responses57,58,59. In addition, BRs, auxins, and cytokinins (CTKs) are also taking part in the defense response of plants to pathogens60,61,62. As a gaseous hormone, ET not only has a wide-ranging regulatory effect on plant growth and development but also participates in plant disease resistance and defense response63,64,65,66. Tezuka et al.67 reported that the rice ethylene response factor (ERF) OsERF83 positively regulates blast resistance by regulating the defense-related genes’ expression in rice. In this study, the expression of EBF1/2 was up-regulated in the S-3dpi vs. S-mock comparison (Supplementary Fig. S3b). SA is a type of phenolic hormone widely distributed in plants. It participates in various physiological processes of plants and plays a key role in forming defense responses against pathogenic bacteria68,69,70. It has been proved that after plants are infected by pathogenic bacteria, the SA that increases exponentially in the body can induce plants to produce hypersensitive response (HR) and systemic acquired resistance (SAR) so that plants exhibit disease resistance71. The SA-mediated disease resistance signal transduction pathway is regulated by multiple genes. Among them, NPR1 (nonexpressor of PR-1) is a key gene located downstream of the SA signal pathway. NPR1 interacts with the transcription factor TGA to activate the expression of a series of SAR genes, and finally confers improved resistance of plants to diseases71. Yang et al.72 reported that enhanced activation of SA biosynthesis in Arabidopsis thaliana hybrids may contribute to their increased resistance to a biotrophic bacterial pathogen. In this study, NPR1 was down-regulated in both R-3dpi vs. R-mock and S-3dpi vs. S-mock comparisons, but PR-1 was up-regulated only in the S-3dpi vs. S-mock (Supplementary Fig. S3). Moon et al.73 suggested that overexpression of OsTGA2 can improve the resistance of rice against bacterial leaf blight disease by directly regulating the expression of defense-related genes. Liu et al.74 found that overexpression of NtPR1a contributed to increasing resistance to R. solanacearum in tobacco Yunyan 87 via activating the defense-related genes. Aux/IAAs (Auxin/indole-3-acetic acid proteins) play important roles in auxin signaling pathways, Fan et al.75 identified some MeAux/IAAs from Manihot esculenta as novel members in plant disease resistance through gene profiling and functional analysis, and these observations provide important information for further utilization of MeAux/IAAs. Similarly, AUX1 was found down-regulated after inoculation with pathogen (Supplementary Fig. S3). The plant hormone abscisic acid (ABA) was found in various parts of the plant. ABA can convert the adversity stimulus in the environment into internal signals of plant cells, and further, make plants to produce physiological reactions and resistance towards the invasion of pathogens76,77. Earlier studies have shown that the protein phospholipase 2 (PP2C) and sucrose non-glycolytic protein kinase 2 (SnRK) genes in plants can eliminate the inhibitory function of phosphate groups when the content of ABA is low, and then the production and release of ABA will be increased. It further promotes the formation of the ABA-PYR/PYL-PP2C complex, which in turn activates SnRK2. SnRK2 further activates the expression of downstream components, stress-related transcription factors, secondary messengers, and related genes, and regulates stress-responsive gene expression, ultimately protecting plants from adverse stresses78,79,80,81. In our study, PP2C and SnRK2 were up-regulated in both R-3dpi vs. R-mock and S-3dpi vs. S-mock comparisons, but PYR/PYL was down- and up-regulated in R-3dpi vs. R-mock and S-3dpi vs. S-mock, respectively (Supplementary Fig. S3). In summary, several genes in the SA, auxin, ABA, and ET signaling pathways are involved in the defense response against R. solanacearum infection in tomato plants.
In the process of long-term interaction with pathogenic bacteria, plants have formed a set of the natural immune system, which includes two levels, namely PAMP-triggered immunity (PTI) and effector-triggered immunity (ETI). PTI promotes the recognition of PAMPs through cell surface receptors, such as bacterial flagellin and fungal chitin, thereby inducing host plants to produce a series of defense responses, including the formation of phytoalexins, the expression of disease-related proteins, and the deposition of the corpus callosum. ETI means that plants recognize the avirulence gene (Avr) of pathogens through the resistance gene (R), which triggers a series of specific defense responses82,83,84. In this study, Rd19 was up-regulated and CDPK, FLS2, and CERK were down-regulated in both R-3dpi vs. R-mock and S-3dpi vs. S-mock (Supplementary Fig. S2). Shi et al.85 reported that silencing of TaCP1 (an RD19-like cysteine protease) reduced wheat resistance to Puccinia striiformis f. sp. tritici. Fu et al.86 showed that rice OsCPK10 is an important regulatory factor in plant immune response, which may regulate disease resistance by activating SA- and JA-dependent defense responses.
In addition, the study also found that the MAPK signal transduction pathway-related genes that mediate plants in response to pathogen infection are up-/down-regulated, respectively, including genes of MAPK and PR-1 (Supplementary Fig. S4). MAPK is a type of serine/threonine-protein kinase, which is ubiquitous in plants. It can receive extracellular or intracellular signals, and further integrate and amplify the signals, ultimately causing cell physiological responses. PR-1 is a pathogenesis-related protein (PRP) that is closely related to disease resistance in plants. This study has also been shown that various stimuli such as pathogen infection, mechanical damage, or pathogen elicitor can activate MAPK, and the MAPK system can further activate related transcription factors. The domains of transcription factors can combine with the W box (T) TGACC (A/T) of PR-1 and can quickly activate the early defense response signal of plants87. In this study, genes that displayed up-/down-regulated expression indicate that the MAPK signal transduction pathway is involved in the response of tomato to R. solanacearum.
Conclusion
In this study, we analyzed the responses of BW-resistant and BW-susceptible tomato lines to R. solanacearum using RNA-seq. Gene expression changes following R. solanacearum-inoculation were identified, the expression levels and types of DEGs were investigated, and GO and KEGG enrichment analyses were performed to annotate the function of DEGs. Our results suggest complex and different responses of the two tomato inbred lines to R. solanacearum infection. This study provides an overall representation of the regulatory gene network for both resistant and susceptible tomato inbred lines responding to R. solanacearum infection. These R. solanacearum-responsive genes could serve as important candidates for future functional research and may be helpful in developing an effective method to resist this significant disease.
Materials and methods
Plant materials and growth conditions
Two S. lycopersicum inbred lines (selected and bred in our laboratory), Hm 2–2 (R, resistant to BW) and BY 1–2 (S, susceptible to BW), were used in this study. No approvals were required for this study, and we complied with all relevant regulations during the experiments. The collection, preservation, and use of plant materials involved in this study complied with relevant institutional, national, and international guidelines and legislation. S. lycopersicum seedlings (preserved in our laboratory) were planted in plastic trays (54 × 28 × 5 cm) filled with substrate (peat: perlite = 3:1, v/v).They were placed in an artificial climate chamber with a temperature of 28 ~ 30/15 ~ 17 °C (day/night), a photoperiod of 14/10 h (day/night), and relative humidity of 50% at Yichun University, Yichun, China. Three weeks later, 100 seedlings at the three-leaf stage were transplanted into black plastic pots (130 mm height × 150 mm diameter) filled with substrate (peat: perlite = 3:1, v/v), and plants were set 10–13 cm from each other. These plants were maintained in an artificial climate chamber with growth conditions as described above.
Inoculation of R. solanacearum
The R. solanacearum strain belonging to race 1 biovar 3 was isolated from the BW-susceptible tomato line (BY 1–2)88. Bacteria were grown on triphenylterazolium chloride (TTC) medium (containing 3 g sucrose, 5–10 g tryptone, beef extract, and 7 g agar per liter) and incubated with 1% TTC for 2 days at 30 °C. Tomato plants at the six-leaf stage were inoculated with R. solanacearum by wounding the roots and incubating in the bacterial suspension (108 colony forming units (cfu)/ml) for 30 min, while sterile water was used for mock-inoculation. Later, all inoculated plants were planted in the plastic pots. Pathogen-inoculated and mock-inoculated Hm 2–2 plants were denoted as R-3dpi and R-mock, respectively, while pathogen-inoculated and mock-inoculated BY 1–2 plants were denoted as S-3dpi and S-mock, respectively. Stem tissue samples of seven tomato plants at 3 dpi were flash-frozen in liquid nitrogen and stored at – 80 °C for further analysis. Each experiment was performed in triplicate.
RNA extraction, RNA-seq library construction, and sequencing
Total RNA of the seven plant stem tissue samples from each of the four lines (R-mock, R-3dpi, S-mock, and S-3dpi) was extracted using a Trizol reagent kit (Invitrogen, Carlsbad, CA, USA) as per the manufacturer’s protocol. Each experiment was performed in triplicate. RNA quality was assessed on an Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA, USA) and checked using RNase-free agarose gel electrophoresis. After extracting the total RNA, eukaryotic mRNA was enriched by oligo (dT) beads, while prokaryotic mRNA was enriched by removing rRNA by Ribo-Zero Magnetic Kit (Epicentre, Madison, WI, USA). The enriched mRNA was fragmented into short fragments using fragmentation buffer and reverse transcribed into complementary DNA (cDNA) with random primers. Second-strand cDNA was synthesized by DNA polymerase I, RNase H, dNTP, and buffer. Then the cDNA fragments were purified with the QiaQuick PCR extraction kit (Qiagen, Venlo, the Netherlands), end-repaired, poly(A)-added, and ligated to Illumina sequencing adapters. The ligation products were size selected by agarose gel electrophoresis, PCR amplified, and sequenced using an Illumina HiSeq2500 platform by Gene Denovo Biotechnology Co. (Guangzhou, China).
RNA-seq data analysis
Raw reads obtained from the sequencer contain adapters or low-quality bases, which would affect the following assembly and analysis. Thus, to get high-quality clean reads, raw reads were filtered by fastp v0.18.089. Short reads alignment tool Bowtie2 v2.2.8 was used for mapping reads to the ribosome RNA (rRNA) database90. The rRNA-mapped reads were removed. The remaining clean reads were further used in assembly and gene abundance calculation. An index of the reference genome was built, and paired-end clean reads were mapped to the S. lycopersicum reference genome (ITAG3.2) using HISAT v2.2.4 with “-rna-strandness RF”, while other parameters were set as default91. The mapped reads of each sample were assembled by using a reference-based approach by StringTie v1.3.192,93. For each transcription region, an FPKM (fragment per kilobase of transcript per million mapped reads) value was calculated to quantify its expression abundance and variations, using StringTie v1.3.1 software.
Analysis of DEGs
DEGs were detected using the DESeq2 package94. The analysis mainly consisted of three steps: (1) normalization of the read count; (2) calculation of the probability of hypothesis testing (p-value) according to the model; (3) conduction of multiple hypothesis testing to obtain the false discovery rate (FDR) value. The genes with a p-value ≤ 0.05 and |log2FC|≥ 1 were considered DEGs.
GO enrichment analysis
Gene ontology (GO) is an international standardized gene functional classification system that offers a dynamic-updated controlled vocabulary and a strictly defined concept to a comprehensive description of gene properties and their products in any organism95. GO has three ontologies: molecular function, cellular component, and biological process. The basic unit of GO is GO term. Each GO term belongs to a type of ontology. GO enrichment analysis provides all GO terms that are significantly enriched in DEGs comparing to the genome background and filter the DEGs that correspond to biological functions. First, all DEGs were mapped to the GO database and assigned with GO terms. Then, gene numbers were calculated for each GO term. Finally, significantly enriched GO terms among DEGs comparing to the genome background were defined by a hypergeometric test. This analysis is able to recognize the main biological functions that DEGs participate in.
Pathway enrichment analysis
Genes usually interact with each other to play roles in certain biological processes. Pathway-based analysis helps to further understand gene biological functions. KEGG (Kyoto Encyclopedia of Genes and Genomes) is the major public pathway-related database96,97. The KEGG pathway was taken as the unit and a hypergeometric test was used to identify pathways that were significantly enriched among the DEGs compared to the entire genome background. Through significant enrichment of pathways, the most important biochemical metabolic pathways and signal transduction pathways involving DEGs can be determined.
Single-nucleotide polymorphism (SNP) analysis
Variation detection based on transcriptome sequencing mainly includes SNPs and insertions/deletions (InDels). SNP variation refers to the DNA sequence polymorphism caused by the change of a single nucleotide at the genome level, while InDel is the variation caused by the insertion or deletion of a nucleotide. The software GATK v3.4-6 was used to detect the variation of SNPs and InDels and finally conduct data statistics98.
Validation of RNA-seq data by qRT-PCR
To validate the RNA-seq results, quantitative real-time PCR (qRT-PCR) was performed in this study. RNA was extracted using the HiPure Total RNA Mini Kit (Magen, Guangzhou, China). Later, RNA was reverse transcribed to cDNA using the HiScript II 1st Stand cDNA Synthesis Kit (+ gDNA wiper; Vazyme, Nanjing, China). The qRT-PCR was performed in triplicate for each sample using the 2 × RealStar Green Fast Mixture (with ROX) as per the manufacturer’s instruction manual. The qRT-PCR amplification was performed using the StepOne Real-Time PCR Instrument (Applied Biosystems, Thermo Fisher, USA) and corresponding software (Applied Biosystems). The tomato Actin (Solyc03g078400) gene was used as the internal control99. The relative expression levels of the genes from three biological replicates were calculated using the 2−△△Ct method100. All primers for qRT-PCR are listed in Supplementary Table S1.
Vector construction and virus-induced gene silencing (VIGS) transformation
The VIGS vectors pTRV1 and pTRV2101 were stored in our laboratory (Fig. 9). A 199-bp WRKY75 (initially characterized by López-Galiano et al.102 and Chen et al.103) fragment was amplified from the stem tissue of resistant tomato plants (Hm 2–2) with the specific primers V-F and V-R (listed in Supplementary Table S2) by PCR. The WRKY75 fragment and the vector pTRV2 were then digested with EcoR I and BamH I (Takara, China), and the WRKY75 fragment was ligated into the vector using T4 ligase (Takara, China) to create the TPV::WRKY75 construct; pTRV2 with empty was created to TRV::empty and was used as control. The resulting vector was introduced into Agrobacterium tumefaciens GV3101 (WEIDI, China). The VIGS experiments were carried out as described previously104. Three-week-old newly emerged leaves of Hm 2–2 tomato plants were transformed with Agrobacterium containing TPV::WRKY75 (V(pTRV1):V(pTRV2::WRKY75) = 1:1) and TRV::empty (V(pTRV1):V(pTRV2::empty) = 1:1) vectors, respectively, using a 1-mL syringe. Each experiment included three biological replicates. One week later, the plants were inoculated with R. solanacearum by using the root-cutting and root-grafting method. Disease scoring was performed according to that previously described by Lacombe et al105.
Statistical analysis
Data were expressed as the mean ± standard deviation (SD). Statistical analyses were performed by SPSS (Statistical Product and Service Solutions) and Excel 2007. Differences were considered statistically significant at p < 0.05 and p < 0.01.
Data availability
RNA sequencing data were deposited at the National Center for Biotechnology Information (NCBI) in the Sequence Read Archive (SRA) under the PRJNA787007 Bioproject accession.
References
Lee, C. G. et al. Comparison of prokaryotic and eukaryotic communities in soil samples with and without tomato bacterial wilt collected from different fields. Microbes Eniviron. 32, 376–385. https://doi.org/10.1264/jsme2.ME17131 (2017).
Singh, V. K., Singh, A. K. & Kumar, A. Disease management of tomato through PGPB: Current trends and future perspective. 3 Biotech. 7(4), 255. https://doi.org/10.1007/s13205-017-0896-1 (2017).
Li, Y., Niu, W. Q., Dyck, M., Wang, J. W. & Zou, X. Y. Yields and nutritional of greenhouse tomato in response to different soilaeration volume at two depths of subsurface drip irrigation. Sci. Rep. 6, 39307. https://doi.org/10.1038/srep39307 (2016).
Zheng, X., Zhu, Y., Wang, J., Wang, Z. & Liu, B. Combined use of a microbial restoration substrate and a virulent Ralstonia solanacearum for the control of tomato bacterial wilt. Sci. Rep. 9(1), 20091. https://doi.org/10.1038/s41598-019-56572-y (2019).
Kim, S. G. et al. Evaluation of resistance to Ralstonia solanacearum in tomato genetic resources at seedling stage. Plant Pathol. J. 32(1), 58–64. https://doi.org/10.5423/PPJ.NT06.2015.0121 (2016).
Shou, S. Y., Feng, Z. Z., Yin, Y. P., Tan, Y. & Miao, L. X. Biochemical and physiological differences between resistant and susceptible tomato cultivars infected by Ralstonia solanncearum Smith. J. Zhejiang Univ. Agric. Life Sci. 31, 550–554 (2005).
Zhou, X. G. & Wu, F. Z. Dynamics of the diversity of fungal and Fusarium communities during continuous cropping of cucumber in the green house. FEMS Microbiol. Ecol. 802(2), 469–478. https://doi.org/10.1111/j.1574-6941.2012.01312.x (2012).
Nion, Y. A. & Toyata, K. Recent trends in control methods for bacterial wilt diseases caused by Ralstonia solanacearum. Microbes Environ. 30(1), 1–11. https://doi.org/10.1264/jsme2.ME14144 (2015).
Ling, L. et al. A streptomyces sp. NEAU-HV9: Isolation, identification, and potential as a biocontrol agent against Ralstonia Solanacearum of tomato plants. Microorganisms. 8(3), 351. https://doi.org/10.3390/microorganisms8030351 (2020).
Yabuuchi, E., Kosako, Y., Yano, I., Hotta, H. & Nishiuchi, Y. Transfer of two burkholderia and an alcaligenes species to Ralstonia gen. Nov: Proposal of Ralstonia pickettii (Ralston, palleroni and doudoroff 1973) comb. Nov., Ralstonia solanacearum (Smith 1896) comb. Nov. and Ralstonia eutropha (Davis 1969) comb. Nov. Microbiol. Immunol. 39(11), 897–904. https://doi.org/10.1111/j.1348-0421.1995.tb03275.x (1995).
Aslam, M. N., Mukhtar, T., Ashfaq, M. & Hussain, M. A. Evaluation of chili germplasm for resistance to bacterial wilt caused by Ralstonia solanacearum. Aust. Plant Pathol. 46(3), 289–292. https://doi.org/10.1007/s13313-017-0491-2 (2017).
Genin, S. Molecular traits controlling host range and adaptation to plants in Ralstonia solanacearum. New Phytol. 187(4), 920–928. https://doi.org/10.1111/j.1469-8137.2010.03397.x (2010).
Wang, J. F. et al. Identification of major QTLs associated with stable resistance of tomato cultivar “Hawaii 7996” to Ralstonia solanacearum. Euphytica 190(2), 241–252. https://doi.org/10.1007/s10681-012-0830-x (2013).
Hayward, A. C. Biology and epidemiology of bacterial wilt caused by Pseudomonas solanacearum. Annu. Rev. Phytopathol. 29, 65–87. https://doi.org/10.1146/annurev.py.29.090191.000433 (1991).
Ishihara, T., Mitsuhara, I., Takahashi, H. & Nakaho, K. Transcriptome analysis of quantitative resistance-specific response upon Ralstonia solanacearum infection in tomato. PLoS ONE 7(10), e46763. https://doi.org/10.1371/journal.pone.0046763 (2012).
Peeters, N., Guidot, A., Vailleau, F. & Valls, M. Ralstonia solanacearum, a widespread bacterial plant pathogen in the post-genomic era. Mol. Plant Pathol. 14(7), 651–662. https://doi.org/10.1111/mpp.12038 (2013).
Wang, G. et al. Resistance against Ralstonia solanacearum in tomato depends on the methionine cycle and the γ-aminobutyric acid metabolic pathway. Plant J. 97(6), 1032–1047. https://doi.org/10.1111/tpj.14175 (2019).
González, W. G. & Summers, W. L. A comparison of Pseudomonas solanacearum-resistant tomato cultivars as hybrid parents. J. Am. Soc. Hortic. Sci. 120(6), 891–895 (1995).
Yue, S. J., Wu, D. H. & Liang, C. Y. Studies on resistance heredity of bacterial wilt of tomato. J. South China Agric. Univ. 16(4), 91–95 (1995).
Li, H. T., Zou, Q. D., Lv, S. W., Mu, X. & Xu, W. K. Studies on resistance heredity of bacterial wilt of tomato. Liaoning Agric. Sci. 5, 1–4 (2001).
Osiru, M. O., Rubaihayo, P. R. & Opio, A. F. Inheritance of resistance to tomato bacterial wilt and its implication for tomato improvement in Uganda. Afr. Crop Sci. J. 9(1), 9–16. https://doi.org/10.4314/acsj.v9i1.27619 (2001).
Li, N. J., Yuan, S. Q. & Li, Y. Identification of solanaceae crop resistance to bacterial wilt by hydroponics I. Hydroponics conditions and nutrient solution formula screening for tobacco, pepper and tomato. Guangdong Agric. Sci. 3, 36–40. https://doi.org/10.16768/j.issn.1004-874x.2000.03.016 (2001).
Wang, G. P., Xiong, Z. K. & Lin, M. B. A preliminary study on the evaluation of bacterial wilt resistance in tomato by a stem imprint method. Acta Agric. Univ. Jiangxiensis. 25(5), 780–782 (2003).
Mangin, B., Thoquet, P., Olivier, J. & Grimsley, N. H. Temporal and multiple quantitative trait loci analyses of resistance to bacterial wilt in tomato permit the resolution of linked loci. Genetics 151(3), 1165–1172. https://doi.org/10.1093/genetics/151.3.1165 (1999).
Ling, M. B., Wang, G. P., Lu, T., Liang, S. N. & Yue, S. J. Preliminary screening for bacterial wilt-resistance tomato and SSR marker linked with resistance. J. Anhui Agric. Sci. 36(9), 3538–3539. https://doi.org/10.13989/j.cnki.0517-6611.2008.09.206 (2008).
Miao, L. X. Identification of AFLP markers linked to bacterial wilt resistance in tomato and isolation of related genes. Doctoral thesis. Zhejiang University, China (2008).
Zhou, G. Z., Li, Z. M., Yang, Y. J., Xing, J. & Wang, R. Q. Genetic diversity of tomato germplasm resources resistant to bacterial wilt (Ralstonia solanacearum) revealed by AFLP. J. Zhejiang Univ. (Agric. Life Sci.) 35(4), 390–394. https://doi.org/10.3785/j.issn.1008-9209.2009.04.006 (2009).
Geethanjali, S., Chen, K. Y., Pastrana, D. V. & Wang, J. F. Development and characterization of tomato SSR markers from genomic sequences of anchored BAC clones on chromosome 6. Euphytica 173(1), 85–97. https://doi.org/10.1007/s10681-010-0125-z (2010).
Ercolano, M. R., Sanseverino, W., Carli, P., Ferriello, F. & Frusciante, L. Genetic and genomic approaches for R-gene mediated disease resistance in tomato: Retrospects and prospects. Plant Cell Rep. 31(6), 973–985. https://doi.org/10.1007/s00299-012-1234-z (2012).
Chen, S. C., Liu, A. R. & Zou, Z. R. Overexpression of glucanase gene and defensin gene in transgenic tomato enhances resistance to Ralstonia solanacearum. Russ. J. Plant Physiol. 53(5), 671–677. https://doi.org/10.1134/S1021443706050116 (2006).
Pan, I. C. et al. Ectopic expression of an EAR motif deletion mutant of SlERF3 enhances tolerance to salt stress and Ralstonia solanacearum in tomato. Planta 232(5), 1075–1086. https://doi.org/10.1007/s00425-010-1235-5 (2010).
Milling, A., Babujee, L. & Allen, C. Ralstonia solanacearum extracellular polysaccharide is a specific elicitor of defense responses in wilt-resistant tomato plants. PLoS ONE 6(1), e15853. https://doi.org/10.1371/journal.pone.0015853 (2011).
Wang, J. G. & Hsuliang, H. Induction of tomato jasmonate-resistant 1-like 1 gene expression can delay the colonization of Ralstonia solanacearum in transgenic tomato. Bot. Stud. 75(1), 75–84 (2012).
Wang, G. P. et al. Studies of two genes related to bacterial wilt resistance in tomato. Acta Hortic. Sin. 41(6), 1096–1104. https://doi.org/10.16420/j.issn.0513-353x.2014.06.006 (2014).
Zhang, Y. et al. A putative LysR-type transcriptional regulator PrhO positively regulates the type III secretion system and contributes to the virulence of Ralstonia solanacearum. Mol. Plant Pathol. 19(8), 1808–1819. https://doi.org/10.1111/mpp.12660 (2018).
Wang, J. et al. Transcriptomic and genetic approaches reveal an essential role of the NAC transcription factor SlNAP1 in the growth and defense response of tomato. Hortic. Res. 7(1), 209. https://doi.org/10.1038/s41438-020-00442-6 (2020).
Pandey, A. et al. Ralstonia solanacearum type III effector RipJ triggers bacterial wilt resistance in Solanum pimpinellifolium. Mol. Plant Microbe Interact. 34(8), 962–972. https://doi.org/10.1094/MPMI-09-20-0256-R (2021).
Gyetvai, G. et al. The transcriptome of compatible and incompatible interactions of potato (Solanum tuberosum) with Phytophthora infestans revealed by DeepSAGE analysis. PLoS ONE 7(2), e31526. https://doi.org/10.1371/journal.pone.0031526 (2012).
Bai, T. T. et al. Transcriptome and expression profile analysis of highly resistant and susceptible banana roots challenged with Fusarium oxysporum f. sp. cubense tropical race 4. PLoS ONE 8(9), e73945. https://doi.org/10.1371/journal.pone.0073945 (2013).
Zhang, Y. et al. Transcriptome profiling of Gossypium barbadense inoculated with Verticillium dahliae provides a resource for cotton improvement. BMC Genom. 14, 637. https://doi.org/10.1186/1471-2164-14-637 (2013).
Jacobs, J. M., Babujee, L., Meng, F., Milling, A. & Allen, C. The in planta transcriptome of Ralstonia solanacearum: Conserved physiological and virulence strategies during bacterial wilt of tomato. MBio 3(4), e00114-e212. https://doi.org/10.1128/mBio.00114-12 (2012).
Tan, X. et al. Complete genome sequence analysis of Ralstonia solanacearum strain PeaFJ1 provides insights into its strong virulence in peanut plants. Front. Microbiol. 13, 830900. https://doi.org/10.3389/fmicb.2022.830900 (2022).
Zuluaga, A. P. et al. Transcriptome responses to Ralstonia solanacearum infection in the roots of the wild potato Solanum commersonii. BMC Genom. 16(1), 246. https://doi.org/10.1186/s12864-015-1460-1 (2015).
Chen, N. et al. RNA-Seq-derived identification of differential transcription in the eggplant (Solanum melongena) following inoculation with bacterial wilt. Gene 644, 137–147. https://doi.org/10.1016/j.gene.2017.11.003 (2018).
Du, H. et al. Dual RNA-seq reveals the global transcriptome dynamics of Ralstonia solanacearum and pepper (Capsicum annuum) hypocotyls during bacterial wilt pathogenesis. Phytopathology https://doi.org/10.1094/PHYTO-01-21-0032-R (2021).
Tian, T. et al. A genome-wide analysis of StTGA genes reveals the rritical role in enhanced bacterial wilt tolerance in potato during Ralstonia solanacearum infection. Front. Genet. 13, 894844. https://doi.org/10.3389/fgene.2022.894844 (2022).
Li, Y. et al. Digital gene expression analysis of the response to Ralstonia solanacearum between resistant and susceptible tobacco varieties. Sci. Rep. 11(1), 3887. https://doi.org/10.1038/s41598-021-82576-8 (2021).
French, E., Kim, B. S., Rivera-Zuluaga, K. & Iyer-Pascuzzi, A. S. Whole root transcriptomic analysis suggests a role for auxin pathways in resistance to Ralstonia solanacearum in tomato. Mol. Plant Microbe Interact. 31(4), 432–444. https://doi.org/10.1094/MPMI-08-17-0209-R (2018).
Jiang, N., Fan, X., Lin, W., Wang, G. & Cai, K. Transcriptome analysis reveals new insights into the bacterial wilt resistance mechanism mediated by silicon in tomato. Int. J. Mol. Sci. 20(3), 761. https://doi.org/10.3390/ijms20030761 (2019).
Gopalan-Nair, R. et al. Convergent rewiring of the virulence regulatory network promotes adaptation of Ralstonia solanacearum on resistant tomato. Mol. Biol. Evol. 38(5), 1792–1808. https://doi.org/10.1093/molbev/msaa320 (2021).
Ma, W. Roles of Ca2+ and cyclic nucleotide gated channel in plant innate immunity. Plant Sci. 181(4), 342–346. https://doi.org/10.1016/j.plantsci.2011.06.002 (2011).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 10(1), 57–63. https://doi.org/10.1038/nrg2484 (2009).
Strauss, T. et al. RNA-seq pinpoints a Xanthomonas TAL-effector activated resistance gene in a large-crop genome. Proc. Natl. Acad. Sci. U. S. A. 109(47), 19480–19485. https://doi.org/10.1073/pnas.1212415109 (2012).
Kamber, T. et al. Fire blight disease reactome: RNA-seq transcriptional profile of apple host plant defense responses to Erwinia amylovora pathogen infection. Sci. Rep. 6, 21600. https://doi.org/10.1038/srep21600 (2016).
Li, W. M. et al. Deep RNA-seq analysis reveals key responding aspects of wild banana relative resistance to Fusarium oxysporum f. sp. cubense tropical race 4. Funct. Integr. Genom. 20(4), 551–562. https://doi.org/10.1007/s10142-020-00734-z (2020).
Dasgupta, U. et al. Comparative RNA-Seq analysis unfolds a complex regulatory network imparting yellow mosaic disease resistance in mungbean [Vigna radiata(L.) R. Wilczek]. PLoS ONE 16(1), e0244593. https://doi.org/10.1371/journal.pone.0244593 (2021).
Pauwels, L., Inzé, D. & Goossens, A. Jasmonate-inducible gene: What does it mean?. Trends Plant Sci. 14, 87–91. https://doi.org/10.1016/j.tplants.2008.11.005 (2009).
Vlot, A. C., Dempsey, D. A. & Klessig, D. F. Salicylic Acid, a multifaceted hormone to combat disease. Annu. Rev. Phytopathol. 47, 177–206. https://doi.org/10.1146/annurev.phyto.050908.135202 (2009).
Mersmann, S., Bourdais, G., Rietz, S. & Robatzek, S. Ethylene signaling regulates accumulation of the FLS2 receptor and is required for the oxidative burst contributing to plant immunity. Plant Physiol. 154(1), 391–400. https://doi.org/10.1104/pp.110.154567 (2010).
Kazan, K. & Manners, J. M. Linking development to defense: Auxin in plant-pathogen interactions. Trends Plant Sci. 14(7), 373–382. https://doi.org/10.1016/j.tplants.2009.04.005 (2009).
Choi, J. et al. The cytokinin-activated transcription factor ARR2 promotes plant immunity via TGA3/NPR1-dependent salicylic acid signaling in Arabidopsis. Dev. Cell. 19(2), 284–295. https://doi.org/10.1016/j.devcel.2010.07.011 (2010).
Oh, M. H. et al. Autophosphorylation of Tyr-610 in the receptor kinase BAK1 plays a role in brassinosteroid signaling and basal defense gene expression. Proc. Natl. Acad. Sci. U. S. A. 107(41), 17827–17832. https://doi.org/10.1073/pnas.0915064107 (2010).
Yang, C. et al. Activation of ethylene signaling pathways enhances disease resistance by regulating ROS and phytoalexin production in rice. Plant J. 89(2), 338–353. https://doi.org/10.1111/tpj.13388 (2017).
Dubois, M., Van den Broeck, L. & Inzé, D. The pivotal role of ethylene in plant growth. Trends Plant Sci. 23(4), 311–323. https://doi.org/10.1016/j.tplants.2018.01.003 (2018).
Debbarma, J. et al. Ethylene Response Factor (ERF) family proteins in abiotic stresses and CRISPR-Cas9 genome editing of ERFs for multiple abiotic stress tolerance in crop plants: A review. Mol. Biotechnol. 61(2), 153–172. https://doi.org/10.1007/s12033-018-0144-x (2019).
Lyu, J. et al. Proteomic analysis reveals key proteins involved in ethylene-induced adventitious root development in cucumber (Cucumis sativus L.). PeerJ 9, e10887. https://doi.org/10.7717/peerj.10887 (2021).
Tezuka, D. et al. The rice ethylene response factor OsERF83 positively regulates disease resistance to Magnaporthe oryzae. Plant Physiol. Biochem. 135, 263–271. https://doi.org/10.1016/j.plaphy.2018.12.017 (2019).
Khan, M. I., Fatma, M., Per, T. S., Anjum, N. A. & Khan, N. A. Salicylic acid-induced abiotic stress tolerance and underlying mechanisms in plants. Front. Plant Sci. 6, 462. https://doi.org/10.3389/fpls.2015.00462 (2015).
Zhang, Y. & Li, X. Salicylic acid: Biosynthesis, perception, and contributions to plant immunity. Curr. Opin. Plant Biol. 50, 29–36. https://doi.org/10.1016/j.pbi.2019.02.004 (2019).
Van Butselaar, T. & Van den Ackerveken, G. Salicylic acid steers the growth-immunity tradeoff. Trends Plant Sci. 25(6), 566–576. https://doi.org/10.1016/j.tplants.2020.02.002 (2020).
Mou, Z., Fan, W. & Dong, X. Inducers of plant systemic acquired resistance regulateNPR1 function through redox changes. Cell 113(7), 935–944. https://doi.org/10.1016/s0092-8674(03)00429-x (2003).
Yang, L. et al. Salicylic acid biosynthesis is enhanced and contributes to increased biotrophic pathogen resistance in Arabidopsis hybrids. Nat. Commun. 6, 7309. https://doi.org/10.1038/ncomms8309 (2015).
Moon, S. J. et al. OsTGA2 confers disease resistance to rice against leaf blight by regulating expression levels of disease related genes via interaction with NH1. PLoS ONE 13(11), e0206910. https://doi.org/10.1371/journal.pone.0206910 (2018).
Liu, Y., Liu, Q., Tang, Y. & Ding, W. NtPR1a regulates resistance to Ralstonia solanacearum in Nicotiana tabacum via activating the defense-related genes. Biochem. Biophys. Res. Commun. 508(3), 940–945. https://doi.org/10.1016/j.bbrc.2018.12.017 (2019).
Fan, S. et al. Molecular functional analysis of auxin/indole-3-acetic acid proteins (Aux/IAAs) in plant disease resistance in cassava. Physiol. Plant. 168(1), 88–97. https://doi.org/10.1111/ppl.12970 (2020).
Mauch-Mani, B. & Mauch, F. The role of abscisic acid in plant–pathogen interactions. Curr. Opin. Plant Biol. 8(4), 409–414. https://doi.org/10.1016/j.pbi.2005.05.015 (2005).
Adie, B. A. et al. ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 19(5), 1665–1681. https://doi.org/10.1105/tpc.106.048041 (2007).
Ma, Y. et al. Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324(5930), 1064–1068. https://doi.org/10.1126/science.1172408 (2009).
Park, S. Y. et al. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324(5930), 1068–1071. https://doi.org/10.1126/science.1173041 (2009).
Umezawa, T. et al. Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc. Natl. Acad. Sci. U. S. A. 106(41), 17588–17593. https://doi.org/10.1073/pnas.0907095106 (2009).
Raghavendra, A. S., Gonugunta, V. K., Christmann, A. & Grill, E. ABA perception and signalling. Trends Plant Sci. 15(7), 395–401. https://doi.org/10.1016/j.tplants.2010.04.006 (2010).
Zipfel, C. et al. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 428(6984), 764–767. https://doi.org/10.1038/nature02485 (2004).
Jones, J. D. G. & Dangl, J. L. The plant immune system. Nature 444(7117), 323–329. https://doi.org/10.1038/nature05286 (2006).
Göhre, V. et al. Plant pattern-recognition receptor FLS2 is directed for degradation by the bacterial ubiquitin ligase AvrPtoB. Curr. Biol. 18(23), 1824–1832. https://doi.org/10.1016/j.cub.2008.10.063 (2008).
Shi, B. et al. Wheat Thioredoxin (TaTrxh1) associates with RD19-like cysteine protease TaCP1 to defend against stripe rust fungus through modulation of programmed cell death. Mol. Plant Microbe Interact. 34(4), 426–438. https://doi.org/10.1094/MPMI-11-20-0304-R (2021).
Fu, L., Yu, X. & An, C. Overexpression of constitutively active OsCPK10 increases Arabidopsis resistance against Pseudomonas syringae pv. tomato and rice resistance against Magnaporthe grisea. Plant Physiol. Biochem. 73, 202–210. https://doi.org/10.1016/j.plaphy.2013.10.004 (2013).
Yu, D., Chen, C. & Chen, Z. Evidence for an important role of WRKY DNA binding proteins in the regulation of NPR1 gene expression. Plant Cell 13(7), 1527–1540. https://doi.org/10.1105/tpc.010115 (2001).
Wang, L., Chen, Y. T., Cai, K. Z. & Wang, G. P. Effects of exogenous silicon supply on the activity of antioxidant enzymes of tomato leaves infected by Ralstonia solanacearum. J. South China Agric. Univ. 35(3), 74–78. https://doi.org/10.7671/j.issn.1001-411X (2014).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17), i884–i890. https://doi.org/10.1093/bioinformatics/bty560 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9(4), 357–359. https://doi.org/10.1038/nmeth.1923 (2012).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods. 12(4), 357–360. https://doi.org/10.1038/nmeth.3317 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33(3), 290–295. https://doi.org/10.1038/nbt.3122 (2015).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11(9), 1650–1667. https://doi.org/10.1038/nprot.2016.095 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15(12), 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Ashburner, M. et al. Gene ontology: Tool for the unification of biology. Gene Ontol. Consort. Nat. Genet. 25(1), 25–29. https://doi.org/10.1038/75556 (2000).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 43(1110), 11101–111033. https://doi.org/10.1002/0471250953.bi1110s43 (2013).
Yan, S. et al. Anthocyanin Fruit encodes an R2R3-MYB transcription factor, SlAN2-like, activating the transcription of SlMYBATV to fine-tune anthocyanin content in tomato fruit. New Phytol. 225(5), 2048–2063. https://doi.org/10.1111/nph.16272 (2020).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt Method. Methods 25(4), 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Liu, Y., Schiff, M. & Dinesh-Kumar, S. P. Virus-induced gene silencing in tomato. Plant J. 31(6), 777–786. https://doi.org/10.1046/j.1365-313x.2002.01394.x (2002).
López-Galiano, M. J. et al. Epigenetic regulation of the expression of WRKY75 transcription factor in response to biotic and abiotic stresses in Solanaceae plants. Plant Cell Rep. 37(1), 167–176. https://doi.org/10.1007/s00299-017-2219-8 (2018).
Chen, L. et al. The transcription factor WRKY75 positively regulates jasmonate-mediated plant defense to necrotrophic fungal pathogens. J. Exp. Bot. 72(4), 1473–1489. https://doi.org/10.1093/jxb/eraa529 (2021).
Chen, N. The expression characteristic and functional identification analysis of SmNAC transcription factor in the eggplant (Solanum melongena). Master’s thesis. South China Agricultural University, China (2016).
Lacombe, S. et al. Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat. Biotechnol. 28(4), 365–369. https://doi.org/10.1038/nbt.1613 (2010).
Acknowledgements
We would like to thank TopEdit (www.topeditsci.com) for the English language editing of this manuscript. We are also grateful to Guangzhou Genedenovo Biotechnology Co., Ltd for assisting in sequencing.
Funding
This work was financially supported by the National Natural Science Foundation of China (Grant No. 32260776), the Program for Science and Technology of the Education Department of Jiangxi Province (Grant No. GJJ190863), the Science and Technology Project of Yichun City (Grant No. YC2020082500022).
Author information
Authors and Affiliations
Contributions
N.C. designed the study and wrote the manuscript. N.C. and Q.S. performed experiments and analyzed the data. Q.L. supervised the study. Q.S. and X.L. performed the tests with R. solanacearum. N.C. and Y.G. performed the bioinformatics analysis. N.C. and Q.S. conducted the qPCR validation analysis. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, N., Shao, Q., Lu, Q. et al. Transcriptome analysis reveals differential transcription in tomato (Solanum lycopersicum) following inoculation with Ralstonia solanacearum. Sci Rep 12, 22137 (2022). https://doi.org/10.1038/s41598-022-26693-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26693-y
- Springer Nature Limited
This article is cited by
-
Transcriptome sequencing and expression analysis in peanut reveal the potential mechanism response to Ralstonia solanacearum infection
BMC Plant Biology (2024)
-
Dendrobine alleviates oleic acid-induced lipid accumulation by inhibiting FOS/METTL14 pathway
Journal of Molecular Histology (2024)
-
Transcriptomics: illuminating the molecular landscape of vegetable crops: a review
Journal of Plant Biochemistry and Biotechnology (2024)