Abstract
Agricultural fertilization with organic amendments of animal origin often leads to antibiotic resistance dissemination. In this study, we evaluated the effect of different treatments (anaerobic digestion, biochar application, ozonation, zerovalent iron nanoparticle application, and spent mushroom substrate addition) on the resistome in dairy cow manure-derived amendments (slurry, manure, and compost). Anaerobic digestion and biochar application resulted in the highest reduction in antibiotic resistance gene (ARG) and mobile genetic element (MGE) gene abundance. These two treatments were applied to cow manure compost, which was then used to fertilize the soil for lettuce growth. After crop harvest, ARG and MGE gene absolute and relative abundances in the soil and lettuce samples were determined by droplet digital PCR and high-throughput qPCR, respectively. Prokaryotic diversity in cow manure-amended soils was determined using 16S rRNA metabarcoding. Compared to untreated compost, anaerobic digestion led to a 38% and 83% reduction in sul2 and intl1 absolute abundances in the soil, respectively, while biochar led to a 60% reduction in intl1 absolute abundance. No differences in lettuce gene abundances were observed among treatments. We conclude that amendment treatments can minimize the risk of antibiotic resistance in agroecosystems.
Similar content being viewed by others

Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Antibiotic resistance is an ancient phenomenon1; however, anthropogenic activities have increased the prevalence of antimicrobial-resistant microorganisms2,3, causing serious human health problems. The transfer of animal and environmental resistomes to humans is a matter of great concern for policymakers. Consequently, in 2006, the European Union banned the use of antibiotics for animal growth promotion4. Moreover, the EU Farm to Fork Strategy aims to reduce the overall EU sales of antimicrobials for farmed animals and aquaculture by 50% by 20305.
Although these regulations control the excessive use of antibiotics in livestock farming, none are related to the management and/or treatment of organic amendments of animal origin. Organic amendments of animal origin, which are commonly used for agricultural fertilization, are a source of antibiotic residues because a considerable percentage (30–90%) of antibiotics administered to livestock are discharged in the urine and feces6. The application of these organic amendments to soil as organic fertilizers can lead to the emergence and dissemination of antibiotic-resistant bacteria (ARB) harboring antibiotic resistance genes (ARGs) in soils and crops7. The principal mechanism underlying the spread of antibiotic resistance is horizontal gene transfer (HGT) via various mobile genetic elements (MGEs), such as conjugative plasmids, integrative conjugative elements, integrons, and transposons8.
The many benefits of organic amendments to agricultural soils and crops are undeniable9. Therefore, there is an urgent need to develop and implement management practices and treatments to reduce the abundance of ARGs and MGEs in animal-based organic amendments. Possible strategies for achieving this include sanitizing the amendments under certain environmental conditions (e.g., temperature, oxygenation) or with other substances, as well as seeking to immobilize ARB and ARGs by adding agents with a very high surface-to-volume ratio. Anaerobic digestion is the process of microbial decomposition of organic matter in the absence of oxygen10. Traditionally, it has been used to reduce the volume of solids from wastewater treatment, producing biogas and other organic compounds11. Nevertheless, its use to treat livestock manure is increasing; this process can reduce antibiotic residues, ARB, and ARGs in manure10,12. In addition, ozone disinfection destroys bacterial cell membranes13 and is widely used in wastewater treatment plants and the food industry. This disinfection process can also remove ARGs14.
On the other hand, biochar is a carbon rich material with a large surface area-to-volume ratio generated from the pyrolysis of biomass in the absence of oxygen15. It is used as a soil amendment to increase soil carbon sequestration, enhance soil fertility and productivity, reduce the bioavailability of organic compounds16, and remove antibiotic residues 17. Moreover, nanoscale zerovalent iron particles (nZVI) show high reactivity towards a broad range of pollutants and have a large surface-area-to-volume ratio18. For example, they are used for the removal of polycyclic aromatic hydrocarbons and metal(oid)s19, as well as bacteria and viruses20,21. They are mainly applied in groundwater remediation and, to a lesser extent, soil remediation22. Finally, five kilograms of spent mushroom substrate (SMS) are generated in the production of one kilogram of fresh mushrooms [for example, Spain produces around 166 thousand tons of mushrooms per year23]. Spent mushroom substrate is therefore generated in large quantities, and for many years, it has accumulated in landfills and has become an environmental problem. SMS has been used in the production of compost24 and animal feed25, as well as for enzyme extraction26, bioremediation27, and removal of antibiotic residues by combining their adsorption and biodegradation mechanisms28.
In the present study, we evaluated the potential of 11 organic amendment management processes to reduce the abundance of ARGs and MGE genes; the amendments used were cow slurry, manure, and compost. Subsequently, the most promising management strategies were applied to a compost that was used to fertilize soil and grow lettuce plants in a microcosm experiment. Thus, we quantified ARGs and MGE genes in (i) amendments, (ii) amended soils, and (iii) lettuce plants. In addition, we studied the structural diversity of soil bacterial communities. We hypothesized that both absolute and relative abundances of ARGs and MGE genes would be lower in the treated amendments than in the untreated controls, and consequently be lower in the amended soils and lettuce plants grown in these soils.
Results
Preliminary study with the three amendments
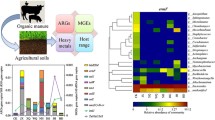
Eleven management processes were tested in a preliminary experiment, performed with slurry, manure and compost, that lasted 21 days. The ARG and MGE gene removal rates in each treatment for every amendment were evaluated using a droplet digital PCR (ddPCR; Fig. 1). Negative values indicated that the management process was successful in gene elimination; conversely, positive values indicated that compared to that in the control treatment, gene abundance increased from day 0 to day 21 in the studied treatment. To determine the success of the management processes in terms of the removal rate of the studied genes, we investigated both the number of genes reduced and the magnitude of their removal.
Removal rates of antibiotic resistance genes (ARGs) and mobile genetic element (MGE) genes under different treatments in three organic amendments (n = 1). Negative values indicate a reduction and positive values an increase in absolute gene abundances, compared to its respective control treatment. Treatments: AD: anaerobic digestion; nZVI: zerovalent iron nanoparticles; SMS: spent mushroom substrate. Amendments: SL: slurry; MAN: manure; COM: compost.
In slurry, the anaerobic digestion (AD) 55 °C treatment showed the greatest success in removing all tested genes; the following most successful treatments were the biochar treatments for all genes except tetA. On the other hand, the ozonation treatment presented the opposite trend, showing a significant increase in all genes except tetA.
Regarding the manure, the nZVI 2% treatment had the greatest success in removing all tested genes, followed by the AD 55 °C, AD 75 °C, biochar 2%, biochar 5%, and SMS 1:0.5 treatments, which failed only with the tetA gene. In contrast, the ozonation and SMS 1:1 treatments increased the removal rates of five out of the six genes.
In compost, the AD 55 °C, AD 75 °C, and biochar 5% treatments showed the greatest success in removing ARGs and MGE genes. On the other hand, the nZVI 1%, ozonation, SMS 1:0.5, and SMS 1:1 treatments showed the opposite trend, increasing the removal rate of every gene.
Soil and lettuce antimicrobial resistance and mobile element genes
Of the three investigated amendments, we selected compost for the microcosm experiment because it is commonly used to fertilize lettuce. The compost management treatments AD 75 °C and biochar 5% were chosen because they were the most promising treatments in the preliminary study. In an additional treatment, a 5% biochar addition to the soil was tested in order to compare the application of biochar to the amendment with its direct application to the soil.
The absolute abundances of the genes were obtained from ddPCR for each treatment (Fig. 2). The abundance of sul1 was significantly lower in the mineral fertilization (NPK) and unamended treatments than that in the biochar soil treatment. Similarly, the abundance of sul2 was lower in the anaerobic digestion, NPK, and unamended treatments than in the untreated compost and biochar soil treatments. Moreover, untreated compost, NPK, and unamended treatments showed lower tetA abundance than those in anaerobic digestion, biochar compost, and biochar soil treatments (the difference was not statistically significant between untreated compost and biochar compost). Regarding the MGE genes, anaerobic digestion, biochar compost, NPK, and unamended treatments presented a lower intl1 gene abundance than that in the untreated compost. Finally, the abundance of tnpA-04 was lower in the unamended treatment than in the untreated compost treatment.
Absolute abundances of antibiotic resistance genes (ARGs) and mobile genetic element (MGE) genes in the soil samples according to the ddPCR analysis, expressed as gene copies per g of DW soil (n = 4). Different letters indicate statistically significant differences among treatments (p < 0.05) according to Tukey’s post hoc test. LoQ: limit of quantification.
The most abundant gene in the studied soils was tnpA-04 in all treatments, followed by sul1 in the biochar compost, biochar soil, and untreated compost treatments; sul2 in the NPK and unamended treatments; and tetA in the anaerobic digestion treatment. In contrast, the least abundant gene in almost all treatments was tetX, with values were under the limit of quantification, with the NPK treatment being an exception in which the least abundant gene was sul1.
With respect to lettuce plants, no statistically significant differences were found among the treatments in terms of ARG and MGE gene absolute abundances (Fig. 3). The most abundant gene in all treatments (expressed as gene copies 105 g−1 DW lettuce) was tnpA-04 (2.3 to 6.0, in untreated compost and biochar soil treatments, respectively), except in the biochar soil treatment, in which sul2 was most abundant (6.7). In contrast, the abundances of sul1, tetA, and tetX were below the quantification limit.
Absolute abundances of antibiotic resistance genes (ARGs) and mobile genetic element (MGE) genes in the lettuce samples according to the ddPCR analysis, expressed as gene copies per g of DW lettuce (n = 4). No statistically significant differences were found among the treatments. LoQ: limit of quantification.
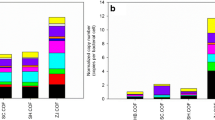
The results of HT-qPCR showed that out of the 76 ARGs and 17 MGE genes targeted in the soils, 69 and 17 genes were amplified, respectively. Out of the 16 ARGs and five MGE genes targeted in lettuce plants, eight and three genes were amplified, respectively. No statistically significant differences in gene families were detected between the treatments in any soil (Supplementary Fig. S1) or lettuce plant (Supplementary Fig. S2). Regarding individual genes, we just found a higher relative abundance of a gene conferring resistance to aminoglycosides (aph3-III) in soils amended with untreated compost than in the rest of the treatments. Moreover, we compared the relative abundance of individual genes in both compartments (i.e., soil and lettuce). We found significant differences in 16 of the 21 common genes studied between soils and lettuce (Supplementary Table S4). In all cases, the relative abundance of the genes was higher in soils than in lettuce.
Furthermore, the ordinary least squares (OLS) regression model showed that the relative abundance of MGE genes was linearly and positively correlated with the relative abundance of ARGs in the studied soils (R2 = 95.3, p < 0.001) (Fig. 4A) and lettuce plants (R2 = 37.2, p < 0.01) (Fig. 4B).
Results of the ordinary least squares (OLS) regression analysis of the relationship between the relative abundance of antibiotic resistance genes (ARGs) and mobile genetic element (MGE) genes in (A) soils and (B) lettuce plants. The line represents the best-fit curve. The shaded area represents the 95% confidence interval for the fitted OLS regression.
Soil bacterial community composition
Regarding the soil bacterial community composition, 99%, 93%, 84%, 47% and 6% of the reads were taxonomically classified at the class, order, family, genus, and species levels, respectively. The two most abundant classes were Alphaproteobacteria and Verrucomicrobiae, accounting for more than 60% of the relative abundance of bacteria; the exception was the anaerobic digestion treatment, in which the second most abundant class was Bacilli (Supplementary Fig. S3). The values of the Shannon’s diversity index (H’) ranged from 3.87 (in both the unamended and biochar compost treatments) to 4.12 (in the NPK treatment). No statistically significant differences were found among the treatments in terms of (i) taxa at class, order or genus levels; (ii) H’ index; or (iii) composition according to the ANOSIM test. Furthermore, no clustering of treatments was observed in the principal coordinates analysis (Supplementary Fig. S4).
Discussion
Organic amendments of animal origin are important sources of antibiotic residues and microflora containing resistance genes (ARGs) in their genome29. Their dissemination in the environment and the associated risks to human and animal health are complex problems that require urgent action. Even though composting offers an environmentally friendly approach to biodegradation of antibiotic residues or ARGs to some extent30, some antibiotic residues and ARGs remain in the organic amendments. Therefore, it is essential to develop effective treatment methods for organic amendments of animal origin to reduce the risk.
The effectiveness of different management processes may differ depending on the type of organic amendment and whether it is in a solid, liquid, or semi-liquid state. In the present study, the best management processes were specific to each organic amendment. In general, the success was lower in compost than in the other two amendments, probably because it was more difficult for the treatment effect to spread well in this solid matrix.
Previous research suggested that anaerobic digestion may be able to remove pathogens and ARGs, with thermophilic digestion being particularly effective31,32. In a former study, the thermophilic digestion of dairy manure reduced the relative abundance of intl1, sul1, sul2, and tetX32. In contrast, an increase in sul1, sul233, and tetA34 was reported as a result of thermophilic digestion of cattle and pig manure, respectively. These inconsistent results may be attributable to the different characteristics of the raw materials, digestion parameters, and bacterial community composition, among others. The increase of tet gene abundances at AD higher temperatures could be associated with the specific groups of bacteria harboring them and their capacity to grow under these environmental conditions35. Also, some tet genes (i.e., tetX, tetW) were considered genes with high dissemination potential in previous studies due to their association with MGEs36,37, and under such conditions they could be transferred among a wide range of bacteria. Remarkably, the impact of anaerobic processes on ARG reduction could be gene-specific38. In the present study, anaerobic digestion was among the best treatments for the three amendments. Surprisingly, the success rate was lower at higher digestion temperatures.
Similarly, the application of biochar to the three amendments (slurry, manure, and compost) was a promising treatment. A previous study39 evaluated the effects of applying rice straw biochar and mushroom biochar during chicken manure composting on ARG removal (including sul1, sul2, tetA, and tetX genes). The removal rate of sul1, sul2, tetA and intl1 genes by adding biochar 5% was significantly higher than the control treatment during composting40. At the end of the composting process, the addition of biochar led to a decrease of the relative abundance of Bacteroidetes and Firmicutes40. Biochar application may potentially alter the resistome profile as a consequence of the changes in bacterial community composition, which is the primary determinant of ARG content41. Biochar can also adsorb heavy metals and reduce their bioavailability, thus reducing the selective pressure on ARB42.
Nanoparticles disrupt intracellular metabolism, damage DNA, and inhibit cell proliferation43. They can also enhance HGT via oxidative stress and increase the expression of mating pair formation44. In the present study, the application of nZVI 2% had the greatest success in removing all tested genes, but the removal rate was negative for almost every gene in the case of the compost. Another study also reported an increase in the relative abundance of intl1, sul2, and tetC in composted chicken manure 21 d after the application of 100 and 600 mg kg−1 of nZVI45. However, 42 d after the nZVI application, the relative abundance of the abovementioned genes decreased. This may have happened because nanoparticles reduced bacterial biomass and induced a succession of the bacterial community in the late stages of composting45.
In the present study, regarding ozonation, the abundances of ARGs and MGE genes increased in general compared to those in the control treatments. These results can be attributed to the consumption of ozone by organic constituents present in the amendments, such as humic substances, carbohydrates, and fatty acids46. Moreover, ozone damages the cell surface before releasing DNA, and intact ARGs may be transferred through HGT47. In another study, the inactivation of ARGs increased in municipal wastewater when the ozone concentration increased from 27 to 61 mg L−148.
Finally, the enrichment of ARGs and MGE genes observed in both manure and compost after SMS application (observed only with the 1:1 dose in manure) might be attributable to (i) the nutrients provided by the SMS, which led to the proliferation of bacterial hosts carrying ARGs and MGEs in the amendment or (ii) SMS-derived ARG and MGE survival and transfer to both manure and compost.
The results of the microcosm experiment with lettuce plants differed depending on the specific genes and the method used to quantify them. The risk tendency discrepancy detected between ddPCR and HT-qPCR may be attributable to the reaction volume (21 µL vs. 100 nL in ddPCR vs. HT-qPCR, respectively), primer sequences employed, annealing temperature (specific vs. general), and abundance units (absolute vs. relative). Droplet digital PCR generates up to 20,000 droplets and a PCR reaction occurs in each droplet. In terms of sensitivity, the lower limit of detection of ddPCR was 10 times more efficient than that of qPCR49. Therefore, we believe that HT-qPCR is useful for quantifying a wide range of genes simultaneously, but the results obtained should be confirmed with another method, such as ddPCR. In our case, we chose to study tetracyclines and sulfonamides in detail because they accounted for 31% and 8% of the total antimicrobial sales for use in food-producing animals in 31 European countries in 2018, respectively50. Although sulfonamides are used less frequently than tetracyclines, the main sulfonamide resistance genes (sul1 and sul2) are often located on transposable elements of self-transferable or mobilizable broad-host-range plasmids and occur in a wide range of bacterial species51. Furthermore, both antibiotics were categorized as highly important antimicrobials for human medicine by the World Health Organization52. Tetracycline and sulfonamide resistance genes have frequently been reported in manure and soil53,54.
In terms of the results obtained by ddPCR in the studied soils, anaerobic digestion led to a 38% and 83% reduction in sul2 and intl1 absolute abundance, respectively, compared to those in the untreated compost. In another study, dairy manure application resulted in higher abundances of intl1 and sul1 in soils fertilized with slurry digestate and untreated slurry than those in mineral fertilization and unamended soils55. The application of biochar to compost also led to a 60% reduction in intl1 gene abundance compared with that in the untreated compost. In another study, the application of 0.5% biochar to soil decreased the ARG abundance in unplanted soil, but failed to remove ARGs from soil planted with Brassica chinensis L.56. As expected, in general, we found that the application of compost increased the soil antimicrobial resistance and mobile element genes, as both mineral fertilization and unamended soil treatments presented the lowest ARG and MGE abundances.
Fortunately, we detected lower abundances of ARGs and MGE genes in lettuce plants than in soil using HT-qPCR. Moreover, sul1, tetA, and tetX gene abundances were below the quantification limit in all treatments of lettuce plants according to ddPCR. These results were consistent with the findings of previous studies indicating that the resistome and mobilome are more diverse and abundant in rhizosphere soil than in plants after the application of organic amendments36,56,57. Furthermore, ordinary least squares regression models pointed to the key role of MGEs in shaping the pattern of ARGs.
No differences were found between treatments in terms of soil bacterial diversity and composition. Therefore, we can argue that the changes in the antimicrobial resistance and mobile element genes were not a consequence of the modification of the bacterial communities.
In summary, anaerobic digestion and biochar addition could be beneficial for reducing the risk of antimicrobial resistance and mobile element genes in organic amendments of animal origin before they are applied to soils. However, further confirmatory studies are necessary before applying these methods on a large scale. Ideally, the treatment or a combination of treatments should be inexpensive and feasible for application on livestock farms.
Materials and methods
Preliminary experiment
The amendments used in this study were provided by a dairy cow farm located in Basque Country (Spain). Three types of amendment (slurry, manure, and compost) were used. The slurry was collected from the outlet of a slurry pond, manure was collected from cow beddings (made from feces, urine, and wheat straw), and compost was sampled from manure piles that had been stored for six months. Both the manure and compost samples were collected and placed in polyethylene bags, whereas the slurry sample was placed in a plastic barrel. The samples were immediately transferred to the laboratory and stored at 4 °C until further use. The physicochemical properties of the amendments are presented in Supplementary Table S1.
The following 11 management processes were tested in the preliminary experiment: anaerobic digestion at 55 °C (AD 55 °C), 75 °C (AD 75 °C), and 90 °C (AD 90 °C); biochar addition at the rate of 0.5% (biochar 0.5%), 2% (biochar 2%), and 5% dry weight (DW; biochar 5%); zerovalent iron nanoparticle addition (nZVI) at 1% (nZVI 1%) and 2% DW (nZVI 2%); ozonation at 2 ppm (ozonation), and spent mushroom substrate addition (SMS) at 1:0.5 (SMS 1:0.5) and 1:1 w:w (SMS 1:1).
For anaerobic digestion, 16, 23, and 80 g FW of compost, manure, and slurry, respectively, were placed in syringes, and distilled water was added to make a final volume of 80 mL. These samples were incubated at the corresponding temperatures in an incubator for 21 d. The rest of the management was carried out at room temperature in darkness. First, 100 g FW of each amendment were placed in plastic pots. Biochar was manually added to the amendments and thoroughly mixed. It had the following physicochemical properties: DW = 53%, pH = 7.5, organic matter (OM) = 69%, electrical conductivity (EC) = 4.1 mS cm−1, total N = 0.7%, K = 13 g kg−1, P = 2.2 g kg−1, and metal concentrations = 0.56, 18, 23, 14, and 103 mg kg−1 for Cd, Cr, Cu, Ni, and Zn, respectively. Zerovalent iron nanoparticles (Nanofer 25S, aqueous dispersion of Fe(0) nanoparticles, NANO IRON s.r.o., Czech Republici) were added to the amendments in the form of slurry and thoroughly mixed; nZVI were applied twice, one week apart. For ozonation, the amendments were placed on a tray and subjected to ozone exposure in an ozonation chamber (2 ppm) for three weeks. Finally, the SMS obtained from the Mushroom Research Technological Center of La Rioja (Spain) was mixed with the amendments. The SMS had the following physicochemical properties: pH = 4.65, OM = 71%, total N = 0.49%, K = 4.3 g kg−1, and P = 0.62 g kg−1. All the management processes had their respective untreated controls (n = 1), which were under the same experimental conditions. For every treatment, two DNA extractions were performed at the beginning (day 0) and at the end of the process (day 21).
Microcosm experiment
Before the beginning of the microcosm experiment, the following physicochemical properties were again determined in the compost, according to standard methods58: DW = 24%, pH = 9.1, C/N ratio = 14.6, EC = 0.54 mS cm−1, and metal concentrations = 0.49, 29, 229, 19, 9.4, 868 mg kg-1 DW for Cd, Cr, Cu, Ni, Pb, and Zn, respectively. The experimental soil was collected from the upper 30 cm layer of a semi-natural grassland field, which, to our knowledge, has never been amended. After collection, the soil was sieved to < 4 mm particles and was filled in 2 kg pots. The soil had the following physicochemical properties: clay loamy texture, OM = 5.4%, pH = 6.0, EC = 0.04 mS cm−1, total N = 0.22%, Olsen P = 2.3 mg kg−1 DW soil, and a K+ content of 97 mg kg−1 DW soil.
The following treatments were tested in the microcosm experiment: (i) untreated compost, (ii) compost subjected to anaerobic digestion at 75 °C for 21 d (anaerobic digestion), (iii) biochar 5% w/w to compost (biochar compost), (iv) biochar 5% w/w to soil (biochar soil), (v) mineral fertilization [NPK, N as NH4NO3 (33.5%), P as P2O5 (18%), and K as K2O (60%)], and (vi) unamended treatment. The compost dose was adjusted to provide an equivalent of 150 kg N ha−1 for lettuce plants. The compost was manually incorporated into the soil, thoroughly mixed (homogenized), and left to stabilize for two weeks. Lettuce seedlings were planted and bottom-watered every 2–3 days during the experimental period. The microcosm experiment was carried out in a growth chamber under the following controlled conditions: 14/10 h light/dark cycle, 20/16 °C day/night temperature, 70% relative humidity, and a photosynthetic photon flux density of 150 µmol photon m−2 s−1. Each treatment was replicated four times. After one month of growth, the plants were harvested and soil samples were collected.
Quantification of antibiotic resistance genes and mobile genetic elements
Total DNA (i.e., both plasmid and genomic) was extracted from the amendments and soil samples (0.25 g DW) using the Qiagen DNeasy PowerSoil Pro Kit (Qiagen, Carlsbad, CA, USA) according to the manufacturer’s instructions, and from the plant samples using the innuPREP Plant DNA Kit (Analytik Jena, Jena, Germany). The quality of amendment, soil, and plant DNA was assessed using a NanoDrop™ One spectrophotometer (Thermo Scientific, Wilmington, DE, USA). The extracted DNA was stored at − 20 °C until further analysis.
Absolute quantification of ARGs and MGE genes in the preliminary and microcosm experiments was conducted using a droplet digital PCR (ddPCR, Bio-Rad Laboratories Inc., Hercules, CA, USA). Two sulfonamide (sul1 and sul2) and two tetracycline (tetA and tetX) resistance genes, as well as two MGE genes (intl1 and tnpA-04) and the 16S rRNA gene were analyzed (see primers and PCR conditions in Supplementary Table S2). Prior to generating droplets for ARGs and MGE genes, the DNA was digested with the XbaI restriction enzyme (Takara Bio, CA, USA) according to the manufacturer’s instructions. The reaction mixture (total volume of 25 µL) consisted of 12.5 µL of QX200 ddPCR EvaGreen Supermix (Bio-Rad), 0.50 µL each of forward and reverse primers (final concentration 10 nM each), 1.0 µL of the digested DNA extract (for 16S rRNA quantification, the DNA was diluted to 0.1 ng µL−1), and 10.5 Milli-Q water. The reaction mixture was added to a 96-well plate, sealed with foil, homogenized by vortexing, and centrifuged at 1000 g for 1 min. Aliquots of 21 µL ddPCR reaction mixture were dispensed into the sample well of the DG8 Droplet Generation Cartridge (Bio-Rad), and 70 µL of the QX200 Droplet Generation Oil for EvaGreen were added to the oil wells. Droplet generation was performed using a QX200 Droplet Generator (Bio-Rad). The generated droplets were transferred to a 96-well plate and incubated at 180 °C using a PX1 PCR plate sealer. The 96-well plate was then transferred to a C1000 Touch Thermal Cycler for PCR amplification. After thermal cycling, the plate was transferred to a QX200 droplet reader for data acquisition. Data analysis was performed using the QuantaSoft software (v.1.4., Bio-Rad). Target gene copies were quantified in duplicate, and positive and no-template controls were included in each ddPCR assay.
Furthermore, the relative abundance of ARGs and MGE genes in each sample of the microcosm experiment was analyzed using customized primer sets in a SmartChip qPCR system (TakaraBio) by Resistomap Oy (Helsinki, Finland). A total of 96 and 24 validated primer sets were used for soils and plants, respectively (see the list of primers in Supplementary Table S3); they were selected because they tested positive in a previous pre-screening analysis carried out with 384 genes. The quantified genes included (i) 76 and 16 primer sets (for soils and plants, respectively) targeting the ARGs conferring resistance against all major classes of antibiotics [aminoglycoside, β-lactam, FCA (phenicol, quinolone), MLSB (macrolide, lincosamide, streptogramin B), multidrug (i.e., those conferring resistance to more than one antibiotic), other (peptide, triclosan, mercury, hydrocarbon), sulfonamide, tetracycline, trimethoprim, and vancomycin]; (ii) 17 and 5 primer sets (for soils and plants, respectively) targeting MGE genes; and (iii) additional taxonomic reference genes (for Bacteroidetes, Firmicutes, and 16S rRNA).
PCR cycling conditions and initial data processing were the same as described previously59,60,61. A threshold cycle (CT) of 27 was used as the detection limit53. Detection of each ARG or MGE gene was considered positive when two out of three technical replicates for each sample were detected. The \({2}^{{-\mathrm{\Delta C}}_{T}}\) method was used to calculate the ARG and MGE gene relative abundances, normalized to the abundance of the 16S rRNA reference gene [where \({\mathrm{\Delta C}}_{T}= {C}_{T(target gene)}- {C}_{T(16S rRNA gene)}\)62].
Amplicon sequencing of soil bacterial communities
The prokaryotic 16S rRNA hypervariable region V4 was targeted using 519F (adapted from Ovreås et al.63) and 806R64 adapter-linked primer pairs as described by Lanzen et al.65. Although the sequencing of this single V-region is widely used for taxonomic classification purposes, the single target condition may lead to limitations in the estimation of diversity66. Pair-ended sequencing was performed using Illumina MiSeq at the Genomics Facility of SGIker (University of the Basque Country, Spain). Quality control of the reads was performed using FASTQC software67. PCR primers were removed from the sequences using Cutadapt68. The resulting FASTQ files were further analyzed via QIIME269 as follows: sequences were imported into QIIME2 as PairedEndSequencesWithQuality, after which both reads were joined by the qiime vsearch join-pairs plugin. Then, low-quality reads were filtered using the QIIME quality-filter Q-score-joined command with the default options. In the next step, deblur70 was used for denoising (qiime deblur denoise-16S), after which the resulting reads were classified using the qiime feature-classifier classify-sklearn and silva-132-99-nb-classifier.qza command as the reference model for assigning a taxonomical classification to the amplicon sequence variants (ASVs). The singletons were removed by the QIIME feature-table filter-features command, and contaminant and unclassified ASVs were also removed by the QIIME taxa filter-table. Finally, a table summarizing the ASVs was created using a qiime feature-table summarize command.
Statistical analysis
In the preliminary experiment, the removal rates were calculated following Yang et al.71 by first subtracting the abundances obtained on day 21 from those obtained on day 0 and then from those of the control treatment. Heatmaps were visualized with ggplot272.
In the microcosm experiment, statistically significant differences among the treatments in both the absolute and relative abundances of ARGs and MGE genes (p < 0.05) for soil and lettuce were assessed with ANOVA and Tukey’s post-hoc tests using the agricolae R package73, followed by Bonferroni’s multiple comparisons test on the relative abundances of individual genes from HT-qPCR results. This same procedure was followed with the ASVs obtained from the amplicon sequencing. Ordinary least squares (OLS) regression models were used to assess the relationships between ARG and MGE gene relative abundances obtained by HT-qPCR analysis using R, and the results were visualized using the ggplot2 package72. The analysis of similarities (ANOSIM) test was used to compared the mean of ranked dissimilarities between treatments to the mean of ranked dissimilarities within treatments. Principal coordinates analysis (PCoA) was used to visualize differences between treatments regarding soil microbial community composition with the ape package74. Bray–Curtis dissimilarity based on the abundance classes was calculated using the vegan package75.
Data availability
All sequence data have been deposited in the European Nucleotide Archive under the study accession number PRJEB49515.
References
D’Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457–461. https://doi.org/10.1038/nature10388 (2011).
Cytryn, E. The soil resistome: The anthropogenic, the native, and the unknown. Soil Biol. Biochem. 63, 18–23. https://doi.org/10.1016/j.soilbio.2013.03.017 (2013).
Holmes, A. H. et al. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 387, 176–187. https://doi.org/10.1016/S0140-6736(15)00473-0 (2016).
Regulation (EC) No 1831/2003 of the European parliament and of the council of 22 September 2003 on additives for use in animal nutrition.
European Commission. Communication from the commission to the European parliament, the council, the European economic and social committee and the committee of the regions: A farm to fork strategy for a fair, healthy and environmentally-friendly food system COM/2020/381 Final, (2020).
Kumar, K. C., Gupta, S. C., Chander, Y. & Singh, A. K. Antibiotic use in agriculture and its impact on the terrestrial environment. Adv. Agron. 87, 1–54. https://doi.org/10.1016/S0065-2113(05)87001-4 (2005).
Chee-Sanford, J. C. et al. Fate and transport of antibiotic residues and antibiotic resistance genes following land application of manure waste. J. Environ. Qual. 38, 1086–1108. https://doi.org/10.2134/jeq2008.0128 (2009).
Heuer, H., Schmitt, H. & Smalla, K. Antibiotic resistance gene spread due to manure application on agricultural fields. Curr. Opin. Microbiol. 14, 236–243. https://doi.org/10.1016/j.mib.2011.04.009 (2011).
Epelde, L. et al. Characterization of composted organic amendments for agricultural use. Front. Sustain. Food Syst. 2, 44. https://doi.org/10.3389/fsufs.2018.00044 (2018).
Youngquist, C. P., Mitchell, S. M. & Cogger, C. G. Fate of antibiotics and antibiotic resistance during digestion and composting: A review. J. Environ. Qual. 45, 537–545. https://doi.org/10.2134/jeq2015.05.0256 (2016).
Ma, X., Xue, X., González-Mejía, A., Garland, J. & Cashdollar, J. Sustainable water systems for the city of tomorrow: A conceptual framework. Sustainability 7, 12071–12105. https://doi.org/10.3390/su70912071 (2015).
Wang, Y. et al. Degradation of antibiotic resistance genes and mobile gene elements in dairy manure anerobic digestion. PLoS ONE 16, e0254836. https://doi.org/10.1371/journal.pone.0254836 (2021).
Thanomsub, B. et al. Effects of ozone treatment on cell growth and ultrastructural changes in bacteria. J. Gen. Appl. Microbiol. 48, 193–199. https://doi.org/10.2323/jgam.48.193 (2002).
Sousa, J. M. et al. Ozonation and UV254nm radiation for the removal of microorganisms and antibiotic resistance genes from urban wastewater. J. Hazard. Mater. 323, 434–441. https://doi.org/10.1016/j.jhazmat.2016.03.096 (2017).
Park, J. H., Choppala, G. K., Bolan, N. S., Chung, J. W. & Chuasavathi, T. Biochar reduces the bioavailability and phytotoxicity of heavy metals. Plant Soil 348, 439–451. https://doi.org/10.1007/s11104-011-0948-y (2011).
Jeffery, S. et al. The way forward in biochar research: targeting trade-offs between the potential wins. GCB Bioenergy 7, 1–13. https://doi.org/10.1111/gcbb.12132 (2015).
Krasucka, P. et al. Engineered biochar: A sustainable solution for the removal of antibiotics from water. Chem. Eng. J. 405, 126926. https://doi.org/10.1016/j.cej.2020.126926 (2021).
Ken, D. S. & Sinha, A. Recent developments in surface modification of Nano zero-valent iron (nZVI): remediation, toxicity and environmental impacts. Environ. Nanotechnol. Monit. Manag. 14, 100344. https://doi.org/10.1016/j.enmm.2020.100344 (2020).
Zhao, X. et al. An overview of preparation and applications of stabilized zero-valent iron nanoparticles for soil and groundwater remediation. Water Res. 100, 245–266. https://doi.org/10.1016/j.watres.2016.05.019 (2016).
Diao, M. & Yao, M. Use of zero-valent iron nanoparticles in inactivating microbes. Water Res. 43, 5243–5251. https://doi.org/10.1016/j.watres.2009.08.051 (2009).
Shi, C. J., Wei, J., Jin, Y., Kniel, K. E. & Chiu, P. C. Removal of viruses and bacteriophages from drinking water using zero-valent iron. Sep. Purif. Technol. 84, 72–78. https://doi.org/10.1016/j.seppur.2011.06.036 (2012).
Anza, M., Salazar, O., Epelde, L., Alkorta, I. & Garbisu, C. The application of nanoscale zero-valent iron promotes soil remediation while negatively affecting soil microbial biomass and activity. Front. Environ. Sci. 7, 19. https://doi.org/10.3389/fenvs.2019.00019 (2019).
FAOSTAT. Mushrooms and truffles, production quantity (tons). https://www.tilasto.com/en/topic/geography-and-agriculture/crop/mushrooms-and-truffles/mushrooms-and-truffles-production-quantity/spain, (2020).
Polat, E., Uzun, H., Topçuo, B., Önal, K. & Onus, A. N. Effects of spent mushroom compost on quality and productivity of cucumber (Cucumis sativus L.) grown in greenhouses. Afr. J. Biotechnol. 8, 176–180 (2009).
Fazaeli, H. & Masoodi, A. R. T. Spent wheat straw compost of Agaricus bisporus mushroom as ruminant feed. Asian-Australas. J. Anim. Sci. 19, 845–851. https://doi.org/10.5713/ajas.2006.845 (2006).
Phan, C. W. & Sabaratnam, V. Potential uses of spent mushroom substrate and its associated lignocellulosic enzymes. Appl. Microbiol. Biotechnol. 96, 863–873. https://doi.org/10.1007/s00253-012-4446-9 (2012).
Lau, K. L., Tsang, Y. Y. & Chiu, S. W. Use of spent mushroom compost to bioremediate PAH-contaminated samples. Chemosphere 52, 1539–1546. https://doi.org/10.1016/S0045-6535(03)00493-4 (2003).
Mayans, B. et al. An assessment of Pleurotus ostreatus to remove sulfonamides, and its role as a biofilter based on its own spent mushroom substrate. Environ. Sci. Pollut. Res. Int. 28, 7032–7042. https://doi.org/10.1007/s11356-020-11078-3 (2021).
Congilosi, J. L. & Aga, D. S. Review on the fate of antimicrobials, antimicrobial resistance genes, and other micropollutants in manure during enhanced anaerobic digestion and composting. J. Hazard. Mater. 405, 123634. https://doi.org/10.1016/j.jhazmat.2020.123634 (2021).
Oliver, J. P. et al. Invited review: fate of antibiotic residues, antibiotic-resistant bacteria, and antibiotic resistance genes in US dairy manure management systems. J. Dairy Sci. 103, 1051–1071. https://doi.org/10.3168/jds.2019-16778 (2020).
Beneragama, N. et al. Survival of multidrug-resistant bacteria in thermophilic and mesophilic anaerobic co-digestion of dairy manure and waste milk. Anim. Sci. J. 84, 426–433. https://doi.org/10.1111/asj.12017 (2013).
Sun, W., Qian, X., Gu, J., Wang, X. J. & Duan, M. L. Mechanism and effect of temperature on variations in antibiotic resistance genes during anaerobic digestion of dairy manure. Sci. Rep. 6, 30237. https://doi.org/10.1038/srep30237 (2016).
Sun, W., Gu, J., Wang, X., Qian, X. & Peng, H. Solid-state anaerobic digestion facilitates the removal of antibiotic resistance genes and mobile genetic elements from cattle manure. Bioresour. Technol. 274, 287–295. https://doi.org/10.1016/j.biortech.2018.09.013 (2019).
Zou, Y., Xiao, Y., Wang, H., Fang, T. & Dong, P. New insight into fates of sulfonamide and tetracycline resistance genes and resistant bacteria during anaerobic digestion of manure at thermophilic and mesophilic temperatures. J. Hazard. Mater. 384, 121433. https://doi.org/10.1016/j.jhazmat.2019.121433 (2020).
Agga, G. E., Kasumba, J., Loughrin, J. H. & Conte, E. D. Anaerobic digestion of tetracycline spiked livestock manure and poultry litter increased the abundances of antibiotic and heavy metal resistance genes. Front Microbiol. 11, 614424. https://doi.org/10.3389/fmicb.2020.614424 (2020).
Jauregi, L., Epelde, L., González, A., Lavín, J. L. & Garbisu, C. Reduction of the resistome risk from cow slurry and manure microbiomes to soil and vegetable microbiomes. Environ. Microbiol. 23, 7643–7660. https://doi.org/10.1111/1462-2920.15842 (2021).
Zhang, Z. et al. Assessment of global health risk of antibiotic resistance genes. Nat Commun 13, 1553. https://doi.org/10.1038/s41467-022-29283-8 (2022).
He, Y. et al. Antibiotic resistance genes from livestock waste: occurrence, dissemination, and treatment. npj Clean Water 3, 4. https://doi.org/10.1038/s41545-020-0051-0 (2020).
Cui, E., Wu, Y., Zuo, Y. & Chen, H. Effect of different biochars on antibiotic resistance genes and bacterial community during chicken manure composting. Bioresour. Technol. 203, 11–17. https://doi.org/10.1016/j.biortech.2015.12.030 (2016).
Fu, Y., Zhang, A., Guo, T., Zhu, Y. & Shao, Y. Biochar and hyperthermophiles as additives accelerate the removal of antibiotic resistance genes and mobile genetic elements during composting. Materials (Basel) 14, 5428. https://doi.org/10.3390/ma14185428 (2021).
Forsberg, K. J. et al. Bacterial phylogeny structures soil resistomes across habitats. Nature 509, 612–616. https://doi.org/10.1038/nature13377 (2014).
Li, H. et al. Effects of bamboo charcoal on antibiotic resistance genes during chicken manure composting. Ecotoxicol. Environ. Saf. 140, 1–6. https://doi.org/10.1016/j.ecoenv.2017.01.007 (2017).
Bondarenko, O., Ivask, A., Käkinen, A. & Kahru, A. Sub-toxic effects of CuO nanoparticles on bacteria: Kinetics, role of Cu ions and possible mechanisms of action. Environ. Pollut. 169, 81–89. https://doi.org/10.1016/j.envpol.2012.05.009 (2012).
Wang, X. et al. Bacterial exposure to ZnO nanoparticles facilitates horizontal transfer of antibiotic resistance genes. NanoImpact 10, 61–67. https://doi.org/10.1016/j.impact.2017.11.006 (2018).
Qiu, X., Zhou, G. & Wang, H. Nanoscale zero-valent iron inhibits the horizontal gene transfer of antibiotic resistance genes in chicken manure compost. J. Hazard. Mater. 422, 126883. https://doi.org/10.1016/j.jhazmat.2021.126883 (2022).
Zeng, T., Wilson, C. J. & Mitch, W. A. Effect of chemical oxidation on the sorption tendency of dissolved organic matter to a model hydrophobic surface. Environ. Sci. Technol. 48, 5118–5126. https://doi.org/10.1021/es405257b (2014).
Pak, G. et al. Comparison of antibiotic resistance removal efficiencies using ozone disinfection under different pH and suspended solids and humic substance concentrations. Environ. Sci. Technol. 50, 7590–7600. https://doi.org/10.1021/acs.est.6b01340 (2016).
Zhuang, Y. et al. Inactivation of antibiotic resistance genes in municipal wastewater by chlorination, ultraviolet, and ozonation disinfection. Environ. Sci. Pollut. Res. Int. 22, 7037–7044. https://doi.org/10.1007/s11356-014-3919-z (2015).
Park, S., Rana, A., Sung, W. & Munir, M. Competitiveness of quantitative polymerase chain reaction (qPCR) and droplet digital polymerase chain reaction (ddPCR) technologies, with a particular focus on detection of antibiotic resistance genes (ARGs). Appl. Microbiol. 1, 426–444. https://doi.org/10.3390/applmicrobiol1030028 (2021).
European Medicines Agency. European surveillance of veterinary antimicrobial consumption, (2020). Sales of Veterinary Antimicrobial Agents in 31 European Countries in 2018 (EMA/24309/2020).
Heuer, H. et al. The complete sequences of plasmids pB2 and pB3 provide evidence for a recent ancestor of the IncP-1β group without any accessory genes. Microbiology (Reading) 150, 3591–3599. https://doi.org/10.1099/mic.0.27304-0 (2004).
World Health Organization. Critically Important Antimicrobials for Human Medicine, 6th Revision (WHO, Geneva, Switzerland, 2019).
Zhu, Y. G. et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc. Natl Acad. Sci. U. S. A. 110, 3435–3440. https://doi.org/10.1073/pnas.1222743110 (2013).
Guo, T. et al. Increased occurrence of heavy metals, antibiotics and resistance genes in surface soil after long-term application of manure. Sci. Total Environ. 635, 995–1003. https://doi.org/10.1016/j.scitotenv.2018.04.194 (2018).
Nõlvak, H. et al. Inorganic and organic fertilizers impact the abundance and proportion of antibiotic resistance and integron-integrase genes in agricultural grassland soil. Sci. Total Environ. 562, 678–689. https://doi.org/10.1016/j.scitotenv.2016.04.035 (2016).
Chen, Q. L. et al. Effect of biochar amendment on the alleviation of antibiotic resistance in soil and phyllosphere of Brassica chinensis L.. Soil Biol. Biochem. 119, 74–82. https://doi.org/10.1016/j.soilbio.2018.01.015 (2018).
Zhu, B., Chen, Q., Chen, S. & Zhu, Y. G. Does organically produced lettuce harbor higher abundance of antibiotic resistance genes than conventionally produced?. Environ. Int. 98, 152–159. https://doi.org/10.1016/j.envint.2016.11.001 (2017) .
Métodos, M. A. P. A. Oficiales de análisis de suelos y Aguas Para riego. Minist. Agric. Pesca Aliment. Métodos Oficiales Anal. III (1994).
Muziasari, W. I. et al. Aquaculture changes the profile of antibiotic resistance and mobile genetic element associated genes in Baltic Sea sediments. FEMS Microbiol. Ecol. 92, fiw052. https://doi.org/10.1093/femsec/fiw052 (2016).
Muurinen, J. et al. Influence of manure application on the environmental resistome under Finnish agricultural practice with restricted antibiotic use. Environ. Sci. Technol. 51, 5989–5999. https://doi.org/10.1021/acs.est.7b00551 (2017).
Muziasari, W. I. et al. The resistome of farmed fish feces contributes to the enrichment of antibiotic resistance genes in sediments below Baltic Sea fish farms. Front. Microbiol. 7, 2137. https://doi.org/10.3389/fmicb.2016.02137 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Ovreås, L., Forney, L., Daae, F. L. & Torsvik, V. Distribution of bacterioplankton in meromictic Lake Saelenvannet, as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA. Appl. Environ. Microbiol. 63, 3367–3373. https://doi.org/10.1128/aem.63.9.3367-3373.1997 (1997) .
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624. https://doi.org/10.1038/ismej.2012.8 (2012).
Lanzén, A. et al. Multi-targeted metagenetic analysis of the influence of climate and environmental parameters on soil microbial communities along an elevational gradient. Sci. Rep. 6, 28257. https://doi.org/10.1038/srep28257 (2016).
Pinna, N. K., Dutta, A., Monzoorul, H. M. & Mande, S. S. Can targeting non-contiguous V-regions with paired-end sequencing improve 16S rRNA-based taxonomic resolution of microbiomes?: An in silico evaluation. Front. Genet. 10, 653. https://doi.org/10.3389/fgene.2019.00653 (2019).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12. https://doi.org/10.14806/ej.17.1.200 (2011).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857. https://doi.org/10.1038/s41587-019-0209-9 (2019).
Amir, A. et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems 2, e00191-e216. https://doi.org/10.1128/mSystems.00191-16 (2017).
Yang, Y., Li, B., Zou, S., Fang, H. H. P. & Zhang, T. Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Res. 62, 97–106. https://doi.org/10.1016/j.watres.2014.05.019 (2014).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
de Mendiburu, F. Agricolae: Statistical procedures for agricultural research. R package version 1.3-3. https://CRAN.R-project.org/package=agricolae (2020).
Paradis, E. & Schliep, K. ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528. https://doi.org/10.1093/bioinformatics/bty633 (2019).
Oksanen, J. et al. Vegan: Community ecology package. R Package Version 2.3-1. (2015).
Acknowledgements
The authors thank support from Euskampus – JRL Environmental Antibiotic Resistance.
Funding
This work was financially supported by the Basque Government (projects: URAGAN 18-00044 and KONTRAE-Elkartek-KK-2020-00007) and the grant PRADA PID2019-110055RB-C21 funded by MCIN/AEI/https://doi.org/10.13039/501100011033. LJ was the recipient of a predoctoral fellowship from the Department of Economic Development and Infrastructure of the Basque Government.
Author information
Authors and Affiliations
Contributions
L.J., C.G., and L.E. conceived the study and designed the project. L.J. and A.G. performed the experiments. L.J. performed the data analysis. L.J. and L.E. interpreted the data and wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jauregi, L., González, A., Garbisu, C. et al. Organic amendment treatments for antimicrobial resistance and mobile element genes risk reduction in soil-crop systems. Sci Rep 13, 863 (2023). https://doi.org/10.1038/s41598-023-27840-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-27840-9
- Springer Nature Limited