Abstract
The intra-uterine components of labor, namely, myometrial contractility, cervical ripening, and decidua/membrane activation, have been extensively characterized and involve a local pro-inflammatory milieu of cellular and soluble immune mediators. Targeted profiling has demonstrated that such processes extend to the intra-amniotic space, yet unbiased analyses of the proteome of human amniotic fluid during labor are lacking. Herein, we utilized an aptamer-based platform to characterize 1,310 amniotic fluid proteins and found that the proteome undergoes substantial changes with term labor (251 proteins with differential abundance, q < 0.1, and fold change > 1.25). Proteins with increased abundance in labor are enriched for immune and inflammatory processes, consistent with prior reports of labor-associated changes in the intra-uterine space. By integrating the amniotic fluid proteome with previously generated placental-derived single-cell RNA-seq data, we demonstrated the labor-driven upregulation of signatures corresponding to stromal-3 and decidual cells. We also determined that changes in amniotic fluid protein abundance are reflected in the maternal plasma proteome. Collectively, these findings provide novel insights into the amniotic fluid proteome in term labor and support its potential use as a source of biomarkers to distinguish between true and false labor by using maternal blood samples.
Similar content being viewed by others
Introduction
Labor is a well-orchestrated process that includes physiological, biochemical, endocrinological, and immunological pathways in the mother and the fetus, culminating in a successful delivery. This complex process involves intra-uterine and extra-uterine components, with the former consisting of increased myometrial contractility, cervical ripening, and decidual/membrane activation1,2,3,4. A strong body of evidence indicates that the intra-uterine components include a local pro-inflammatory milieu characterized by increased concentration of soluble and number of cellular immune mediators in the myometrium5,6,7,8,9,10,11,12,13,14,15,16, cervix5,7,8,9,17,18,19,20,21,22,23,24,25, decidua7,8,26,27,28,29,30,31,32,33,34,35, and chorioamniotic membranes7,8,27,28,30,36,37,38,39,40. Such an intra-uterine inflammatory response is also reflected in the intra-amniotic space. Indeed, prior targeted studies have shown that the concentrations of several cytokines, including Interleukin-1β (IL-1β)41 and Interleukin-6 (IL-6)42, are elevated in the amniotic fluid of women with term labor compared to those without labor. Furthermore, cytomic approaches have shown that innate immune cells, such as neutrophils and monocytes/macrophages, are abundant in the amniotic fluid of women at term gestation43. In line with this concept, we recently profiled the amniotic fluid proteome from early to late pregnancy and found that immune-specific signatures were enriched in term gestation44. Notably, placental- and uterine-derived signatures were modulated in the amniotic fluid proteome44, showing that the fetal and maternal components are present in the intra-amniotic space. Taken together, these findings led us to hypothesize that amniotic fluid harbors cellular and soluble mediators at the end of gestation in preparation for the physiologic intra-amniotic inflammatory process of term parturition. Hence, the primary aim of the current study was to explore the human proteome of amniotic fluid during term parturition.
In addition to intra-uterine components, labor also includes extra-uterine components such as processes taking place in the maternal circulation1,2,3,4. Indeed, the detection of cell-free RNA and DNA derived from the fetus has been an invaluable clinical tool in evaluating fetal development as well as in screening for fetal abnormalities and obstetrical disease45,46,47,48,49,50,51,52,53,54,55,56,57,58,59. Moreover, the use of high-dimensional techniques such as transcriptomics, proteomics, and metabolomics has allowed for the exploration of the complex and dynamic processes that are modulated in the maternal circulation prior to and during labor60,61,62,63,64,65,66,67,68,69,70. Recently, we leveraged the maternal blood transcriptome to monitor single-cell signatures derived from placental tissue and demonstrated the increasing abundance of such transcripts with advancing gestation71 as well as in women with term71 or spontaneous preterm labor and birth66. Similarly, labor-specific transcriptomic changes in the maternal circulation can be correlated with those derived from the chorioamniotic membranes, cervix, or myometrium, suggesting that tissue signatures of labor are partly mirrored in this compartment70. By intersecting single-cell RNA-seq data of the laboring myometrium with maternal transcriptomic datasets, we also demonstrated that specific cell signatures were modulated throughout gestation and enriched with the process of labor72. Together, these prior observations demonstrate that labor-specific changes taking place in the gestational and reproductive tissues can be tracked in the maternal circulation. Yet, whether the processes occurring in the amniotic fluid during term labor are also reflected in the maternal circulation has not been investigated.
Herein, we utilized the aptamer-based SOMAscan proteomics platform73 to characterize changes in the concentrations of 1,310 amniotic fluid proteins during the normal process of labor at term. In addition to identifying amniotic fluid proteins dysregulated with labor, we tested in independent patient sets whether the amniotic fluid-derived signature can discriminate between women in labor and those not in labor based on blood protein profiles.
Results
Clinical characteristics of the study population
This study included amniotic fluid samples collected from women at term in labor (TIL, n = 24) or not in labor (TNL, n = 11) (Fig. 1, top panel). The clinical characteristics of the study participants are shown in Table 1. There were no significant differences in gestational age at amniocentesis between the TIL and TNL patients (median gestational age at amniocentesis: TIL 39 weeks vs. TNL 38 weeks, p = 0.25). There were no significant differences in maternal age, nulliparity, or birth weight between the two groups (Table 1). All women delivered a healthy neonate (Apgar score at 5 min > 8 for all cases) without major pregnancy or neonatal complications.
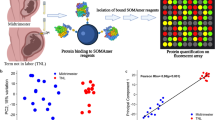
Study design and summary. The study included the determination of 1310 protein analytes in amniotic fluid samples collected at term from patients not in labor (TNL, n = 11) and from those in active labor (TIL, n = 24) (top panel). Maternal plasma concentrations of amniotic fluid proteins that significantly increased in abundance with labor were then used to discriminate labor from no labor groups in an independent set of pregnant women (TNL, n = 21 vs TIL, n = 59) (bottom panel). The figure was created with biorender.com155.
An additional set of plasma proteome data were available from a separate cohort of TIL (n = 59) and TNL (n = 21) patients (Fig. 1, bottom panel)74. The clinical characteristics of the study participants are shown in Supplementary Table S1. There was no significant difference in gestational age at sampling between the two groups (median gestational age at blood draw: TIL 39 weeks vs. TNL 38.7 weeks, p = 0.21).
Validation of the SOMAmer assay to evaluate amniotic fluid proteins
Although the SOMAscan v3 platform has been previously validated by using both targeted and unbiased proteomic platforms across a range of biological specimens, there is limited information about the cross-platform agreement when measuring amniotic fluid proteins75,76. Therefore, we utilized enzyme-linked immunoassay (ELISA)-based concentrations of C–X–C motif chemokine ligand 10 (CXCL10), neutrophil elastase (ELANE), interleukin (IL)-6, and secretory leukocyte protease inhibitor (SLPI) that were available for a subset of samples included in this study (Fig. 2). We found a significant positive correlation between the SOMAmer assay and the ELISA for each of these proteins [CXCL10: n = 19, Spearman's ρ = 0.76, p < 0.001; ELANE, n = 30, ρ = 0.57, p = 0.001; IL-6: n = 19, ρ = 0.65, p = 0.003; SLPI: n = 30, ρ = 0.61, p < 0.001], therefore indicating significant cross-platform agreement.
Cross-platform correlation of specific amniotic fluid proteins. Scatter plots of log2-transformed relative fluorescence units determined by using the SOMAscan assay (y-axis) and log2-transformed ELISA or RIA concentrations (x-axis) for (a) C–X–C motif chemokine ligand 10 (CXCL10), (b) neutrophil elastase (ELANE), (c) interleukin (IL)-6, and (d) secretory leukocyte protease inhibitor (SLPI). The number of sample pairs, Spearman's correlation coefficient, and the corresponding p-value are shown per analyte. The scatter plots were generated with the R package, ggplot2159.
The amniotic fluid proteome shows changes with labor at term
To visualize the relationship between the amniotic fluid proteome of TIL and TNL patients, principal component (PC) analysis was utilized. Despite some overlap, most amniotic fluid samples showed clear separation according to labor status (Supplementary Fig. S1). Furthermore, the meta-proteomes (PC1 and PC2, 22% and 16% of variance explained, respectively) were significantly associated with labor status (t-test, p < 0.05 for PC1 and for PC2).
Although there was no significant difference in gestational age at amniocentesis between the two study groups (Table 1), we and other investigators have shown that gestational age is a strong modulator of the amniotic fluid proteome and transcriptome, and even a small difference in gestational age at amniocentesis could potentially translate into changes in protein abundance44,77,78,79. However, Pearson correlation analysis showed no evidence of an association between gestational age at amniocentesis and the principal components of our current data [PC1: ρ = 0.01, p = 0.94, PC2: ρ = 0.03, p = 0.89)]. This finding is most likely due to the narrow range of gestational ages wherein amniocentesis was performed in the current study. Thus, we have not adjusted for gestational age at amniocentesis in our subsequent analyses.
A comparison of amniotic fluid protein abundance between the TIL and TNL groups resulted in 251 (19.2%) proteins with significant differences (q < 0.1 and fold change ≥ 1.25) (Supplementary Table S2). Of these, 100 (39.8%) proteins were more abundant, and 151 (60.2%) proteins were less abundant in TIL than in TNL. The summary results of this differential abundance analysis are displayed in Fig. 3, where the volcano plot indicates the magnitude of differences in protein abundance (log2 fold changes) against the statistical significance (q-values) of these differences (Fig. 3a). In addition, the heat map shown in Fig. 3b displays the log2 abundance in relative fluorescence units of the most significant (q < 0.05 and fold change ≥ 1.5) proteins in a color scale, indicating consistent patterns of protein abundance within each study group. These results illustrate the distinct amniotic fluid proteomic profile that characterizes labor at term.
Differential protein abundance in the amniotic fluid with and without labor at term. (a) Volcano plot showing log10-transformed false discovery rate adjusted p-values (q-values) against log2-transformed fold changes of the 1310 amniotic fluid proteins. (b) Heatmap based on proteins with significantly altered abundance (q<0.05 and fold change ≥ 1.5) between term not in labor (TNL, indicated by blue headers) and term in labor (TIL, indicated by red headers) samples. The R/Bioconductor packages, EnhancedVolcano157 and pheatmap158, were used to generate the volcano plot and heatmap, respectively.
Proteins differentially abundant with term labor are enriched for distinct signaling processes
Next, to aid in the interpretation of the proteomic dysregulation associated with term labor, we performed gene set enrichment analysis (GSEA) of the MSigDB C5 gene set collection and identified Gene Ontology (GO) terms (biological processes, molecular functions, and cellular components) associated with labor status. We identified 184 biological processes, 24 molecular functions, and 9 cellular components that were enriched in proteins with higher abundance in the TNL compared to TIL samples (q < 0.25) (Supplementary Table S3). Such biological processes included regulation of CD4 positive alpha–beta T-cell differentiation, lymphocyte activation, regulation of protein phosphorylation, cell–cell signaling, tyrosine phosphorylation of STAT protein, regulation of phospholipase activity, cardiac conduction system development, and transmission of nerve impulse (Supplementary Table S3). For proteins that were more abundant in the TIL group compared to the TNL group, we determined the enrichment of 65 biological processes, 28 molecular functions, and 22 cellular components (Supplementary Table S3). The significantly enriched biological processes included regulation of proteolysis, intracellular transport, macromolecule catabolic process, erythrocyte, and myeloid cell homeostasis (Supplementary Table S3).
We then conducted a GSEA of the MSigDB C2 gene set collection of canonical pathways to gain further insights into the physiology of normal labor as captured by the amniotic fluid proteome. GSEA identified 48 biological pathways that were significantly enriched among proteins more abundant in TNL compared to TIL samples (Supplementary Table S4), which included pathways related to cytokine signaling in T helper cells, e.g., cytokine-cytokine receptor interaction, JAK-STAT signaling, and selective expression of chemokine receptors during T-cell polarization as well as other signaling pathways (Fig. 4). By contrast, the 91 biological pathways enriched among proteins more abundant in TIL samples (Supplementary Table S4) were related to neutrophil degranulation, carbohydrate metabolism, degradation of the extracellular matrix, and myometrial relaxation and contraction pathways.
Functional enrichment of biological pathways in the amniotic fluid proteome with and without labor at term. Gene-concept network (cnetplot) of biological pathways that were significantly enriched (q-value < 0.1) before (blue) or after (red) the onset of labor. The size of the nodes, corresponding to enriched terms, represents the number of member genes that contributed to the enrichment (leading edge) of the term. The color of the nodes representing each leading-edge gene indicates the TIL/TNL fold change in the corresponding protein abundance. The relationships between the enriched terms are represented by the shared genes. The R/Bioconductor package, clusterProfiler162 was used to generate the figure.
Taken together, these findings indicate a distinct inflammatory proteomic signature associated with physiological labor at term compared to term no labor.
Integration of the amniotic fluid proteome with placental single-cell RNA-seq signatures reveals labor-specific enrichment of specific cell types
Given the crucial role of the placenta in cross-talk between the mother and the fetus during pregnancy and at labor onset, we further sought to interpret the labor-associated changes in the amniotic fluid proteome by intersecting these data with cell type signatures previously identified by using single-cell RNA-seq of placental tissues66. The aggregated proteomic signature of decidual cells and stromal-3 cells was significantly increased (q < 0.1) in amniotic fluid samples from TIL patients compared to TNL patients (Fig. 5, Supplementary Table S5). By contrast, there was a significant decrease in the signatures corresponding to extravillous trophoblasts, cytotrophoblasts, natural killer cells, endometrial cells, and fibroblasts with the onset of term labor (Fig. 5, Supplementary Table S5).
Placental single-cell RNA-seq signatures represented within the amniotic fluid proteome. For each placental cell type signature previously derived by single-cell RNA-seq analysis, the concentration of proteins coded by up to 20 most preferentially expressed genes was transformed into a Z-score and averaged. The Z-scores were compared between the term not in labor (TNL, blue bars) and term in labor (TIL, red bars) groups. The cell types that significantly (q-value < 0.1) changed in expression are shown. The box plots were generated with the R package, ggplot2159.
Changes in the amniotic fluid proteome are reflected in the maternal circulation and can distinguish term labor
We then sought to determine whether the labor-associated protein changes in amniotic fluid are also reflected in the maternal plasma proteome, given that signatures derived from the tissues surrounding this compartment can be monitored in the maternal circulation66,70,72. We first defined an amniotic fluid proteomic signature of term labor by selecting the top 20 proteins that were most increased in the TIL group compared to the TNL group. Among these 20 proteins, the abundance of 15 in the maternal plasma was also available in a dataset generated by SOMAScan v.2 in an independent set of uncomplicated pregnancies74. The aggregated proteomic signature (average of Z-scores) was significantly higher in maternal plasma samples collected at term labor compared to gestational age-matched samples collected from women not in labor (Fig. 6a), and the discrimination accuracy was substantial (area under the curve [AUC] = 0.76, 95% confidence interval [CI] 0.64–0.88) (Fig. 6b).
Changes in the amniotic fluid proteome are reflected in the maternal circulation and can distinguish term labor. (a) Boxplots showing changes in the amniotic fluid-derived labor signature in the maternal plasma proteomic profiles. The box plots were generated with the R package, ggpubr163. (b) Receiver operating characteristic curves for classifying maternal plasma samples into labor and no labor groups based on the plasma concentrations of proteins identified in the amniotic fluid. The black curve corresponds to the discrimination accuracy of the aggregated proteomic signature of the top 15 amniotic fluid proteins that were most increased in the TIL group compared to the TNL group. The red curve corresponds to the leave-one-out cross-validation (LOOCV)-based predictions of a random forest model trained with 10 proteins selected according to their importance from all amniotic fluid proteins increased in the TIL group compared to the TNL group. The ROC curves were plotted with the R package, pROC164.
Finally, we extended the pool of protein candidates to all proteins with significantly higher abundance in amniotic fluid during labor (Supplementary Table S2) and allowed a random forest classifier to select and assign different levels of importance to the different candidate proteins, as opposed to the simple average, as described above. Random forest models based on ten proteins selected according to their importance resulted in a leave-one-out cross-validation (LOOCV) AUC of 0.9 (95% CI 0.8–1) (Fig. 6b). Among the considered proteins, those most informative for distinguishing between labor and no labor in maternal plasma samples were ranked high to low: ERP29 (endoplasmic reticulum resident protein 29), SERPINE1 (plasminogen activator inhibitor 1), ICOSLG (ICOS ligand), SFTPD (pulmonary surfactant-associated protein D), HNRNPAB (heterogeneous nuclear ribonucleoprotein A/B), UFC1 (ubiquitin-fold modifier-conjugating enzyme 1), CTSV (cathepsin L2), PSME1 (proteasome activator complex subunit 1), IGFBP1 (insulin-like growth factor-binding protein 1), and PRDX1 (peroxiredoxin-1).
These data demonstrate that labor-associated changes in the amniotic fluid proteome can be monitored in the maternal circulation, providing a potential non-invasive means to distinguish labor.
Discussion
We utilized an aptamer-based proteomic platform to measure the abundance of 1,310 proteins in amniotic fluid samples collected from women with spontaneous term labor and from those at term without labor. First, we demonstrated that the amniotic fluid proteome undergoes substantial changes with spontaneous term labor that include the altered abundance of 251 proteins. Moreover, such proteins are enriched for immune and inflammatory processes, consistent with labor-associated changes occurring in other compartments. By integrating the amniotic fluid proteome with placenta-derived single-cell RNA-seq data, we demonstrated the labor-driven upregulation of signatures corresponding to decidual and stromal-3 cells. Importantly, we show that changes in amniotic fluid protein abundance are reflected in the maternal plasma proteome, providing the means to distinguish between the patients in labor from those not in labor by using minimally invasive samples. Such data may inform the development of blood tests to distinguish true labor from false labor at term and improve patient management.
Previous investigations have sought to describe the soluble proteome of the amniotic fluid in normal pregnancy by using MS-based approaches77,80,81,82,83, although such studies did not evaluate changes associated with spontaneous term labor. The proteomics platform utilized herein targets a predetermined number of proteins with specific aptamers, thus allowing for high throughput measurements comparable to immunoassays over a wide dynamic range75,76,84. While MS-based approaches identify more proteins, they tend to miss low abundant signaling proteins, e.g., cytokines85. The aptamer-based platform has been previously used to describe changes in the amniotic fluid44 and maternal plasma proteome86 with advancing gestation and pregnancy complications87,88,89,90; yet, the current study represents its first application to study physiological term labor.
Prior research into amniotic fluid proteins in term labor has primarily utilized targeted approaches to evaluate individual proteins or specific protein sets41,42,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123. Such studies support our current observations by indicating a general increase in inflammatory mediators in amniotic fluid during labor, as evidenced by the elevated concentrations of pro-inflammatory cytokines41,42,95,99,100,101,102,106,111, chemokines93,97,99,107,115,121, arachnoid acid metabolites124,125, and extracellular matrix-degrading proteases108,113,116,126. Herein, we confirmed the previously reported term labor-associated increase in concentrations of several such inflammatory proteins, including IL-642,99,100,101,102, C-X-C motif chemokine ligand 8 (CXCL8)93,97,99, colony stimulating factor-3 (CSF3)96,99, matrix metallopeptidase-8 (MMP8)110, matrix metallopeptidase-9 (MMP9)108,126, and heat shock protein family A (Hsp70) member 1A (HSPA1A)120. We also reproduced the previously reported decrease in amniotic fluid concentrations of tumor necrosis factor ligand superfamily member 6, soluble form (FASLG)127, intercellular adhesion molecule-1 (ICAM1)122, and lymphotoxin alpha (LTA)104 with labor. Yet, the application of an aptamer-based multiplex proteomics platform has enabled deeper exploration beyond markers of inflammation to describe labor-associated changes in proteins involved in other processes such as intracellular signaling, cell metabolism, intracellular transport, and cell–cell communications.
Herein, pathway analysis of the amniotic fluid protein changes in term labor identified innate immune responses and neutrophil degranulation as being enriched among upregulated proteins. This observation is in line with a previous report of amniotic fluid immunophenotyping, showing that neutrophils are the most abundant cell types in the amniotic fluid at term43. Moreover, our previous study comparing the amniotic fluid proteome between samples collected at term (without labor) and those collected during mid-trimester also identified neutrophil-mediated immunity as the most enriched functional term among proteins that increased in concentration at term44. Neutrophils produce matrix metalloproteinases that degrade the extracellular matrix, a process crucial for two key events of parturition: rupture of the fetal membranes and cervical ripening113,128,129,130. Indeed, GO and pathway analyses identified terms such as proteolysis, genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix, and extracellular matrix organization as enriched among proteins with increased abundance during labor.
Another set of functional terms that emerged from the functional profiling of proteins with increased abundance in term labor was related to carbohydrate metabolism. The top three upregulated proteins in labor were glycolytic enzymes: alpha-enolase (ENO1), fructose-bisphosphate aldolase A (ALDOA), and gamma-enolase (ENO2). Parturition requires diverse effector functions of multiple cell types across the maternal and fetal intrauterine tissues, making energy generation and consumption essential for accomplishing parturition without complications. Glucose metabolism meets a major portion of this increased energy demand131,132. Other studies of term and preterm pregnancies have also found an increase in the concentrations of proteins involved in carbohydrate metabolism with the spontaneous onset of labor in amniotic fluid and cervicovaginal fluid133,134,135,136. For example, analysis of paired cervicovaginal fluid samples collected a week before and a week after the onset of spontaneous term labor showed a significant increase in ENO1 with labor133. In addition, mass spectral proteomics analysis of amniotic fluid samples collected from women with spontaneous preterm labor has identified glycolysis/gluconeogenesis-associated proteins as differentially abundant in women who delivered preterm with134 and without134,135 intra-uterine infection or intra-uterine inflammation.
Studies in murine pregnancy have indicated that the fetal lung secretes surfactant components, particularly surfactant protein A (SP-A), into the amniotic fluid, where it can activate macrophages and induce inflammation and parturition137,138,139. However, it was observed that SP-A concentrations were lower in human amniotic fluid samples collected at term in labor compared to those at term without labor118. While SP-A was not measured in the current study, a functionally related protein, pulmonary surfactant-associated protein D (SFTPD), was among the most abundant in amniotic fluid samples collected after labor onset. Of note, while a moderate increase in the amniotic fluid concentration of SFTPD in amniotic fluid has been reported throughout the third trimester140, we did not previously observe a similar change in the amniotic fluid samples collected at term without labor compared to those collected during mid-pregnancy44. Thus, the observed changes in amniotic fluid concentrations of SFTPD could be specific to labor rather than to advanced gestational age. In addition, a genetic study has previously shown a significant association between SFTPD gene polymorphism and spontaneous preterm birth among women with recurrent preterm birth141.
Moreover, we showed that the meta-protein signature corresponding to decidual cells was increased in amniotic fluid samples collected during labor. Notably, we previously determined that the meta-RNA expression of decidual cells decreases with gestational age in amniotic fluid samples collected before the onset of labor79, suggesting that a labor-specific activation of the decidual cells occurs. Indeed, decidual activation is a crucial component of the common pathway of parturition4,142,143. Consistently, in addition to the increased meta-protein signature of decidual cells, we also observed that the biological pathway of myometrial contraction and relaxation was enriched among proteins that were more abundant in samples collected after labor. Given that myometrial contractility represents another component of the common pathway of parturition3,4,142, these findings further support the reflection of labor-specific changes in the intra-uterine tissues by the amniotic fluid proteome.
Amniocentesis is an invasive procedure that is no longer a part of routine maternal care144; thus, the development of diagnostic biomarkers that can be evaluated by using minimally invasive samples, such as maternal blood, is essential. To determine the potential of non-invasive biomarker discovery based on information derived from amniotic fluid, we followed a previously described analytic approach to define and track disease-specific69, single cell-specific79, and tissue-specific signatures44,145 in the maternal circulation and amniotic fluid. First, we defined a labor-specific signature as consisting of amniotic fluid proteins that increased the most in abundance with labor in a Hispanic population. Next, we showed that this meta-protein signature is significantly increased with term labor in the maternal plasma proteome of an independent set of predominantly African-American mothers74. The simple average of standardized plasma protein abundance distinguished between the groups, which is encouraging when considering that the proteins were identified in the amniotic fluid analysis. Moreover, when the list of candidate proteins was expanded from the top 15 to all amniotic fluid proteins significantly increased with labor, a plasma proteomic random forest model led to substantially improved accuracy. The set of proteins that most contributed to prediction accuracy included many previously known to be associated with labor in gestational tissues. For example, the expression of the gene coding for SERPINE1 was previously shown to be elevated during labor in the myometrium and placenta146,147,148,149. Moreover, mass spectrometry-based proteomic analysis of placental membranes collected after spontaneous labor (term or preterm) showed that ERP29 was detected only in term placentas150. The concentration of decidual IGFBP1 was shown to increase in vaginal secretions after fetal membrane rupture151, and an increased concentration of IGFBP1 in the cervicovaginal fluid has been proposed as a biomarker of PPROM152,153,154. These findings indicate that the amniotic fluid proteomic changes associated with term labor have biological plausibility and are translatable to the maternal circulation and across diverse cohorts of patients.
Strengths and limitations
The strengths of this study include the unbiased analysis of the proteome (1,310 proteins), although we recognize that this protein set represents a fraction of the human proteome. Another strength is the use of gestational age-matched samples to control for this variable, given that we have demonstrated its influence on the amniotic fluid proteome44. Finally, while amniotic fluid proteomic analysis was performed in a Hispanic population, the observed labor-specific changes were translatable to the maternal plasma proteome of a predominantly African-American population. The primary limitation of this study is the moderate sample size of the TNL group, which is to be expected since amniocenteses are not commonly performed in term deliveries. Yet, to overcome such a limitation, we utilized robust differential analysis and functional profiling methods with particular emphasis on controlling the false discovery rate. In addition, our study design (case–control, cross-sectional study) was not ideally suited to discriminate between events that cause labor and those that accompany labor since amniocentesis was performed either in the absence of labor or during labor. Nevertheless, by using robust functional profiling strategies, we provide corroborating evidence for several biological processes previously implicated in the onset of labor, including fetal lung maturity and decidual activation.
Whether the observed association between molecular changes in the amniotic fluid and maternal blood extends to other ethnicities and obstetrical syndromes, such as spontaneous preterm labor and birth, remains to be determined in future studies.
Conclusion
Collectively, the findings herein provide the first characterization of the amniotic fluid proteome during term parturition, thus demonstrating that such a process is reflected by the altered proteomic composition in this fetal compartment. Moreover, we validated our findings by corroborating the enrichment of specific immune and inflammatory processes associated with labor onset in other intrauterine compartments. Importantly, labor-driven perturbations of the amniotic fluid proteome can be observed in the maternal plasma proteome, thereby supporting its potential use as a biomarker to inform optimal patient management.
Methods
Ethics
The study protocol, collection of samples, and use of clinical data were approved by the Human Investigation Committee of Sotero del Rio Hospital, Santiago, Chile (amniotic fluid), and the Institutional Review Boards of Wayne State University and the Pregnancy Research Branch (formerly known as the Perinatology Research Branch), an intramural program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, U.S. Department of Health and Human Services (NICHD/NIH/DHHS), Detroit, MI, USA (maternal plasma). All patients provided written informed consent prior to sample collection. All experiments were performed in accordance with relevant guidelines and regulations.
Study population
Amniotic fluid
Amniotic samples were collected from pregnant women seeking care at the Sotero del Rio Hospital, Santiago, Chile. A cross-sectional study was designed to include 35 women: 11 who underwent amniocentesis at term before the onset of labor and 24 who had an amniocentesis at term after the spontaneous onset of labor (Fig. 1, top panel). Labor was diagnosed by the presence of regular uterine contractions occurring at a frequency of four in 20 min for a minimum of 1 h associated with cervical dilatation and/or effacement changes. Women at term not in labor underwent amniocentesis for the assessment of fetal lung maturity prior to cesarean delivery, whereas those in labor underwent amniocentesis because of uncertain gestational age or for the diagnosis of intra-amniotic infection. Amniotic fluid samples were obtained by transabdominal amniocentesis under ultrasonographic guidance and cultured for the presence of microorganisms (aerobic and anaerobic bacteria and genital Mycoplasmas). Amniotic fluid not used in clinical assessment was centrifuged for 10 min at 4 °C, and the supernatant was aliquoted and stored at − 80 °C until analysis.
We included cases without evidence of medical or obstetrical complications such as preterm labor, preeclampsia, clinical chorioamnionitis, gestational or pregestational diabetes mellitus, positive amniotic fluid culture, meconium-stained amniotic fluid, multiple gestation, and pregnancy with fetal anomalies. The amniotic fluid samples had been previously used in targeted studies of amniotic fluid cytokines and arachidonic acid metabolites113,115,116,118.
Maternal plasma
Plasma samples were collected from women enrolled in a prospective longitudinal study at the Center for Advanced Obstetrical Care and Research of the NICHD's Pregnancy Research Branch, Detroit Medical Center, and Wayne State University74. Only the last sample collected prior to term delivery from women with spontaneous labor (n = 59) and the gestational age-matched samples from women who were not in labor (n = 21) were used (Fig. 1, bottom panel).
The study design is summarized in Fig. 1, which was created with Biorender.com155.
Proteomics
The abundance of 1310 proteins in amniotic fluid samples (75 µL aliquots) and 1125 proteins in maternal plasma samples was determined with the SOMAmer (Slow Off-rate Modified Aptamers) platforms v3 and v2, respectively (SomaLogic, Inc., Boulder, CO, USA), as previously described44,86. Briefly, the samples were incubated with SOMAmer mixes pre-immobilized onto streptavidin-coated beads and washed to remove non-specifically bound proteins. Proteins bound to their cognate SOMAmer reagents were tagged by using the NHS-biotin reagent. The beads were treated with an anionic competitor solution to prevent non-specific interactions. The beads were exposed to ultraviolet light to release pure cognate-SOMAmer complexes and unbound SOMAmer reagents, and the photo-cleavage eluate was incubated with a second streptavidin-coated bead to capture the biotinylated proteins. Unbound SOMAmer reagents were removed during subsequent washing, and the bound SOMAmer reagents were separated from their cognate proteins under denaturing conditions and hybridized to custom DNA microarrays. Protein abundance was measured in relative fluorescence units by detecting the Cyanine-3 signal from the fluorophores in SOMAmer reagents. The raw signal intensities were standardized by hybridization control normalization, median signal normalization, and interplate calibration84.
Data analysis
Demographic data analysis
Clinical characteristics and demographics of the study participants were summarized as the median and interquartile range for continuous variables and as proportions for categorical variables. To compare data between groups, Welch's t-test and Fisher's exact test were used for continuous and categorical variables, respectively. A p-value < 0.05 was considered statistically significant.
Differential protein abundance analysis and validation
The abundance of 1,310 proteins in the amniotic fluid was compared between samples collected from patients at term in labor and from those at term without labor by using moderated t-tests implemented in the limma R package156. A fold change ≥ 1.25 and a false discovery rate adjusted p-value (q-value) < 0.1 were used to determine statistical significance. The results of differential expression analysis were summarized and visualized with volcano plots and heatmaps, using the R packages EnhancedVolcano157 and pheatmap158, respectively.
Amniotic fluid concentrations of four proteins—CXCL10, ELANE, IL-6, and SLPI—were determined previously by specific immunoassays, ELISA, or radioimmunoassay, according to the manufacturer's instructions113. These data were used to assess cross-platform reproducibility via Spearman's correlation analysis. The correlations were visualized with scatter plots created with the R package ggplot2159.
Gene Ontology and biological pathway enrichment analysis
All proteins were mapped to Entrez gene identifiers per the manufacturer's provided annotation. We used GSEA160 to analyze the molecular signatures database (MSigDB)161 C5 sub-collection of gene sets corresponding to GO biological processes, molecular functions, and cellular components. We also performed the same analysis for the MSigDB C2 collection of canonical biological pathways curated from popular online databases, e.g., KEGG and Reactome. The analysis was restricted to gene sets with at least five genes having corresponding proteins measured on the SOMAScan platform. A q-value < 0.25 was considered statistically significant, as recommended by the authors of GSEA160. The enrichment of selected biological pathways was visualized as a gene-concept network created using the R/Bioconductor package, clusterProfiler162.
Placental single-cell-specific expression
We previously defined sets of genes as specific to a given population of cells identified by single-cell RNA-seq analyses of the placenta66. The log2 transformed relative fluorescence units reflecting the abundance of proteins encoded by these genes and measured herein with the SOMAmer assay were standardized by subtracting the mean and dividing by the standard deviation calculated from the reference (TNL) study group44,79,145. The standardized values referred to as Z-scores were then averaged and compared between groups with the Wilcoxon rank sum test. A q-value of less than 0.1 was considered significant. To obtain robust summaries of signatures, we considered only those cell types having at least five signature genes with corresponding proteins measured by the SOMAscan platform.
We implemented a previously described strategy66,69 to define an amniotic fluid labor signature consisting of the top 20 proteins most increased in abundance after the onset of labor. Of these 20 amniotic fluid proteins, 15 (HNRNPA2B1, MMP8, SFTPD, PRTN3, CSF3, PEBP1, ACY1, PPIA, TPT1, CTSV, ICOSLG, GDI2, CXCL8, MMP1, and AKR1A1) were also measured in maternal plasma by using the SOMAScan platform (v2) (Fig. 1)74. The average z-scores corresponding to these 15 proteins were then compared between the TIL and TNL groups with Wilcoxon tests and visualized in boxplots created with the R package, ggpubr 163.
Plasma proteomic models to classify labor and no labor groups
A random forest model was fit to discriminate between the labor and no labor groups by using protein data derived from the plasma samples. Only amniotic fluid proteins that significantly increased in abundance with labor were considered because there was no significant difference between the labor groups in the aggregated Z-score in the maternal plasma proteome of the 20 amniotic fluid proteins that decreased in abundance with labor. A LOOCV procedure was used to assess the model's generalizability to unseen data. Briefly, in each iteration of LOOCV, one patient profile was excluded from the training set used to fit the model, and the resulting model was applied to the data of the patient excluded from the training set. During training, an initial random forest classifier was fit, and the top 10 features, ranked according to metric importance, were selected to fit the final model. The procedure was repeated for all patients, and the model performance was evaluated by calculating the area under the receiver operating curve (AUROC). The receiver operating curve was plotted using the R package, pROC164, and the R packages, randomForest165, and ranger166, were used to fit the random forest models.
References
Norwitz, E. R., Robinson, J. N. & Challis, J. R. The control of labor. N. Engl. J. Med. 341, 660–666. https://doi.org/10.1056/NEJM199908263410906 (1999).
Romero, R. et al. The preterm parturition syndrome. BJOG 113(Suppl 3), 17–42. https://doi.org/10.1111/j.1471-0528.2006.01120.x (2006).
Smith, R. Parturition. N. Engl. J. Med. 356, 271–283. https://doi.org/10.1056/NEJMra061360 (2007).
Romero, R., Dey, S. K. & Fisher, S. J. Preterm labor: One syndrome, many causes. Science 345, 760–765. https://doi.org/10.1126/science.1251816 (2014).
Mackler, A. M., Iezza, G., Akin, M. R., McMillan, P. & Yellon, S. M. Macrophage trafficking in the uterus and cervix precedes parturition in the mouse. Biol. Reprod. 61, 879–883 (1999).
Thomson, A. J. et al. Leukocytes infiltrate the myometrium during human parturition: Further evidence that labour is an inflammatory process. Hum. Reprod. 14, 229–236 (1999).
Young, A. et al. Immunolocalization of proinflammatory cytokines in myometrium, cervix, and fetal membranes during human parturition at term. Biol. Reprod. 66, 445–449 (2002).
Osman, I. et al. Leukocyte density and pro-inflammatory cytokine expression in human fetal membranes, decidua, cervix and myometrium before and during labour at term. Mol. Hum. Reprod. 9, 41–45 (2003).
Yellon, S. M., Mackler, A. M. & Kirby, M. A. The role of leukocyte traffic and activation in parturition. J. Soc. Gynecol. Investig. 10, 323–338 (2003).
Shynlova, O., Tsui, P., Dorogin, A. & Lye, S. J. Monocyte chemoattractant protein-1 (CCL-2) integrates mechanical and endocrine signals that mediate term and preterm labor. J. Immunol. 181, 1470–1479 (2008).
Shynlova, O., Tsui, P., Jaffer, S. & Lye, S. J. Integration of endocrine and mechanical signals in the regulation of myometrial functions during pregnancy and labour. Eur. J. Obstet. Gynecol. Reprod. Biol. 144(Suppl 1), S2-10. https://doi.org/10.1016/j.ejogrb.2009.02.044 (2009).
Lombardi, A. et al. Expression of matrix metalloproteinases in the mouse uterus and human myometrium during pregnancy, labor, and preterm labor. Reprod. Sci. 25, 938–949. https://doi.org/10.1177/1933719117732158 (2018).
Ulrich, C. C. et al. Matrix metalloproteinases 2 and 9 are elevated in human preterm laboring uterine myometrium and exacerbate uterine contractility. Biol. Reprod. 100, 1597–1604. https://doi.org/10.1093/biolre/ioz054 (2019).
Leimert, K. B. et al. Maternal and fetal intrauterine tissue crosstalk promotes proinflammatory amplification and uterine transition. Biol. Reprod. 100, 783–797. https://doi.org/10.1093/biolre/ioy232 (2019).
Leimert, K. B. et al. Cooperative effects of sequential PGF2alpha and IL-1beta on IL-6 and COX-2 expression in human myometrial cells. Biol. Reprod. 100, 1370–1385. https://doi.org/10.1093/biolre/ioz029 (2019).
Wendremaire, M. et al. Macrophage-induced reactive oxygen species promote myometrial contraction and labor-associated mechanisms. Biol. Reprod. 102, 1326–1339. https://doi.org/10.1093/biolre/ioaa032 (2020).
Liggins, C. G. in Cervix in Pregnancy and Labour (eds D.A. Elwood & A.B.M. Andersson) 1 - 9 (Churchill Livingstone, 1981).
Bokström, H., Brännström, M., Alexandersson, M. & Norström, A. Leukocyte subpopulations in the human uterine cervical stroma at early and term pregnancy. Hum. Reprod. 12, 586–590. https://doi.org/10.1093/humrep/12.3.586 (1997).
Kelly, R. W. Inflammatory mediators and cervical ripening. J. Reprod. Immunol. 57, 217–224 (2002).
Sakamoto, Y., Moran, P., Bulmer, J. N., Searle, R. F. & Robson, S. C. Macrophages and not granulocytes are involved in cervical ripening. J. Reprod. Immunol. 66, 161–173. https://doi.org/10.1016/j.jri.2005.04.005 (2005).
Yellon, S. M., Ebner, C. A. & Sugimoto, Y. Parturition and recruitment of macrophages in cervix of mice lacking the prostaglandin F receptor. Biol. Reprod. 78, 438–444. https://doi.org/10.1095/biolreprod.107.063404 (2008).
Timmons, B. C., Fairhurst, A. M. & Mahendroo, M. S. Temporal changes in myeloid cells in the cervix during pregnancy and parturition. J. Immunol. 182, 2700–2707. https://doi.org/10.4049/jimmunol.0803138 (2009).
Yellon, S. M. et al. Remodeling of the cervix and parturition in mice lacking the progesterone receptor B isoform. Biol. Reprod. 85, 498–502. https://doi.org/10.1095/biolreprod.111.091983 (2011).
Payne, K. J., Clyde, L. A., Weldon, A. J., Milford, T. A. & Yellon, S. M. Residency and activation of myeloid cells during remodeling of the prepartum murine cervix. Biol. Reprod. 87, 106. https://doi.org/10.1095/biolreprod.112.101840 (2012).
Myers, D. A. The recruitment and activation of leukocytes into the immune cervix: Further support that cervical remodeling involves an immune and inflammatory mechanism. Biol. Reprod. 87, 107. https://doi.org/10.1095/biolreprod.112.105049 (2012).
Vince, G. S., Starkey, P. M., Jackson, M. C., Sargent, I. L. & Redman, C. W. Flow cytometric characterisation of cell populations in human pregnancy decidua and isolation of decidual macrophages. J. Immunol. Methods 132, 181–189 (1990).
Fidel, P. L. Jr. et al. Interleukin-1 receptor antagonist (IL-1ra) production by human amnion, chorion, and decidua. Am. J. Reprod. Immunol. 32, 1–7 (1994).
Keelan, J. A. et al. Cytokine abundance in placental tissues: Evidence of inflammatory activation in gestational membranes with term and preterm parturition. Am. J. Obstet. Gynecol. 181, 1530–1536 (1999).
Keski-Nisula, L., Aalto, M. L., Katila, M. L. & Kirkinen, P. Intrauterine inflammation at term: A histopathologic study. Hum. Pathol. 31, 841–846. https://doi.org/10.1053/hupa.2000.8449 (2000).
Gomez-Lopez, N., Estrada-Gutierrez, G., Jimenez-Zamudio, L., Vega-Sanchez, R. & Vadillo-Ortega, F. Fetal membranes exhibit selective leukocyte chemotaxic activity during human labor. J. Reprod. Immunol. 80, 122–131. https://doi.org/10.1016/j.jri.2009.01.002 (2009).
Gomez-Lopez, N., Guilbert, L. J. & Olson, D. M. Invasion of the leukocytes into the fetal-maternal interface during pregnancy. J. Leukoc. Biol. 88, 625–633. https://doi.org/10.1189/jlb.1209796 (2010).
Hamilton, S. et al. Macrophages infiltrate the human and rat decidua during term and preterm labor: Evidence that decidual inflammation precedes labor. Biol. Reprod. 86, 39. https://doi.org/10.1095/biolreprod.111.095505 (2012).
Gomez-Lopez, N. et al. Evidence for a role for the adaptive immune response in human term parturition. Am. J. Reprod. Immunol. 69, 212–230. https://doi.org/10.1111/aji.12074 (2013).
Hamilton, S. A., Tower, C. L. & Jones, R. L. Identification of chemokines associated with the recruitment of decidual leukocytes in human labour: potential novel targets for preterm labour. PLoS ONE 8, e56946. https://doi.org/10.1371/journal.pone.0056946 (2013).
Castillo-Castrejon, M. et al. Choriodecidual cells from term human pregnancies show distinctive functional properties related to the induction of labor. Am. J. Reprod. Immunol. 71, 86–93. https://doi.org/10.1111/aji.12179 (2014).
Lonergan, M. et al. Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL), TRAIL receptors, and the soluble receptor osteoprotegerin in human gestational membranes and amniotic fluid during pregnancy and labor at term and preterm. J. Clin. Endocrinol. Metab. 88, 3835–3844. https://doi.org/10.1210/jc.2002-021905 (2003).
Esplin, M. S. et al. Monocyte chemotactic protein-1 expression is increased in human gestational tissues during term and preterm labor. Placenta 26, 661–671. https://doi.org/10.1016/j.placenta.2004.09.012 (2005).
Gomez-Lopez, N. et al. Specific inflammatory microenvironments in the zones of the fetal membranes at term delivery. Am. J. Obstet. Gynecol. 205(235), e215-224. https://doi.org/10.1016/j.ajog.2011.04.019 (2011).
Gomez-Lopez, N., Vadillo-Perez, L., Nessim, S., Olson, D. M. & Vadillo-Ortega, F. Choriodecidua and amnion exhibit selective leukocyte chemotaxis during term human labor. Am. J. Obstet. Gynecol. 204(364), e369–e316. https://doi.org/10.1016/j.ajog.2010.11.010 (2011).
Lozovyy, V., Richardson, L., Saade, G. & Menon, R. Progesterone receptor membrane components: Key regulators of fetal membrane integrity. Biol. Reprod. 104, 445–456. https://doi.org/10.1093/biolre/ioaa192 (2021).
Romero, R. et al. Amniotic fluid interleukin-1 in spontaneous labor at term. J. Reprod. Med. 35, 235–238 (1990).
Romero, R., Avila, C., Santhanam, U. & Sehgal, P. B. Amniotic fluid interleukin 6 in preterm labor. Assoc. Infect. J Clin Invest. 85, 1392–1400. https://doi.org/10.1172/JCI114583 (1990).
Gomez-Lopez, N. et al. The immunophenotype of amniotic fluid leukocytes in normal and complicated pregnancies. Am. J. Reprod. Immunol. 79, e12827. https://doi.org/10.1111/aji.12827 (2018).
Bhatti, G. et al. The amniotic fluid proteome changes with gestational age in normal pregnancy: a cross-sectional study. Sci. Rep. 12, 601. https://doi.org/10.1038/s41598-021-04050-9 (2022).
Lee, T. et al. Down syndrome and cell-free fetal DNA in archived maternal serum. Am. J. Obstet. Gynecol. 187, 1217–1221. https://doi.org/10.1067/mob.2002.127462 (2002).
Wataganara, T. et al. Maternal serum cell-free fetal DNA levels are increased in cases of trisomy 13 but not trisomy 18. Hum. Genet. 112, 204–208. https://doi.org/10.1007/s00439-002-0853-9 (2003).
Angert, R. M. et al. Fetal cell-free plasma DNA concentrations in maternal blood are stable 24 hours after collection: analysis of first- and third-trimester samples. Clin. Chem. 49, 195–198. https://doi.org/10.1373/49.1.195 (2003).
Levine, R. J. et al. Two-stage elevation of cell-free fetal DNA in maternal sera before onset of preeclampsia. Am. J. Obstet. Gynecol. 190, 707–713. https://doi.org/10.1016/j.ajog.2003.12.019 (2004).
Farina, A. et al. High levels of fetal cell-free DNA in maternal serum: A risk factor for spontaneous preterm delivery. Am. J. Obstet. Gynecol. 193, 421–425. https://doi.org/10.1016/j.ajog.2004.12.023 (2005).
Bianchi, D. W. et al. Fetal nucleic acids in maternal body fluids: An update. Ann. N Y Acad. Sci. 1075, 63–73. https://doi.org/10.1196/annals.1368.008 (2006).
Lo, Y. M. & Chiu, R. W. Noninvasive prenatal diagnosis of fetal chromosomal aneuploidies by maternal plasma nucleic acid analysis. Clin. Chem. 54, 461–466. https://doi.org/10.1373/clinchem.2007.100016 (2008).
Lun, F. M. et al. Noninvasive prenatal diagnosis of monogenic diseases by digital size selection and relative mutation dosage on DNA in maternal plasma. Proc. Natl. Acad. Sci. U S A 105, 19920–19925. https://doi.org/10.1073/pnas.0810373105 (2008).
Lo, Y. M. et al. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci. Transl. Med. 2, 61ra91. https://doi.org/10.1126/scitranslmed.3001720 (2010).
Tsui, N. B. et al. Noninvasive prenatal diagnosis of hemophilia by microfluidics digital PCR analysis of maternal plasma DNA. Blood 117, 3684–3691. https://doi.org/10.1182/blood-2010-10-310789 (2011).
Edlow, A. G. & Bianchi, D. W. Tracking fetal development through molecular analysis of maternal biofluids. Biochim. Biophys. Acta 1970–1980, 2012. https://doi.org/10.1016/j.bbadis.2012.04.005 (1822).
Bianchi, D. W. et al. Noninvasive prenatal testing and incidental detection of occult maternal malignancies. JAMA 314, 162–169. https://doi.org/10.1001/jama.2015.7120 (2015).
Bianchi, D. W. & Chiu, R. W. K. Sequencing of circulating cell-free DNA during pregnancy. N. Engl. J. Med. 379, 464–473. https://doi.org/10.1056/NEJMra1705345 (2018).
Sin, S. T. K. et al. Identification and characterization of extrachromosomal circular DNA in maternal plasma. Proc. Natl. Acad. Sci. U S A 117, 1658–1665. https://doi.org/10.1073/pnas.1914949117 (2020).
Ma, M. L. et al. Fetal mitochondrial DNA in maternal plasma in surrogate pregnancies: Detection and topology. Prenat. Diagn. 41, 368–375. https://doi.org/10.1002/pd.5860 (2021).
Koh, W. et al. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc. Natl. Acad. Sci. USA 111, 7361–7366. https://doi.org/10.1073/pnas.1405528111 (2014).
Tsui, N. B. et al. Maternal plasma RNA sequencing for genome-wide transcriptomic profiling and identification of pregnancy-associated transcripts. Clin. Chem. 60, 954–962. https://doi.org/10.1373/clinchem.2014.221648 (2014).
Fragiadakis, G. K. et al. Mapping the fetomaternal peripheral immune system at term pregnancy. J. Immunol. 197, 4482–4492. https://doi.org/10.4049/jimmunol.1601195 (2016).
Aghaeepour, N. et al. An immune clock of human pregnancy. Sci. Immunol. 2. https://doi.org/10.1126/sciimmunol.aan2946 (2017).
Aghaeepour, N. et al. A proteomic clock of human pregnancy. Am. J. Obstet. Gynecol. 218(347), e341-347.e314. https://doi.org/10.1016/j.ajog.2017.12.208 (2018).
Ghaemi, M. S. et al. Multiomics modeling of the immunome, transcriptome, microbiome, proteome and metabolome adaptations during human pregnancy. Bioinformatics 35, 95–103. https://doi.org/10.1093/bioinformatics/bty537 (2019).
Pique-Regi, R. et al. Single cell transcriptional signatures of the human placenta in term and preterm parturition. Elife 8. https://doi.org/10.7554/eLife.52004 (2019).
Liang, L. et al. Metabolic dynamics and prediction of gestational age and time to delivery in pregnant women. Cell 181, 1680–1692 e1615. https://doi.org/10.1016/j.cell.2020.05.002 (2020).
Stelzer, I. A. et al. Integrated trajectories of the maternal metabolome, proteome, and immunome predict labor onset. Sci. Transl. Med. 13. https://doi.org/10.1126/scitranslmed.abd9898 (2021).
Tarca, A. L. et al. Maternal whole blood mRNA signatures identify women at risk of early preeclampsia: A longitudinal study. J. Matern. Fetal. Neonatal. Med. 34, 3463–3474. https://doi.org/10.1080/14767058.2019.1685964 (2021).
Gomez-Lopez, N. et al. Transcriptome changes in maternal peripheral blood during term parturition mimic perturbations preceding spontaneous preterm birth. Biol. Reprod. 106, 185–199. https://doi.org/10.1093/biolre/ioab197 (2022).
Tarca, A. L. et al. Targeted expression profiling by RNA-Seq improves detection of cellular dynamics during pregnancy and identifies a role for T cells in term parturition. Sci. Rep. 9, 848. https://doi.org/10.1038/s41598-018-36649-w (2019).
Pique-Regi, R. et al. A single-cell atlas of the myometrium in human parturition. JCI Insight 7. https://doi.org/10.1172/jci.insight.153921 (2022).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One 5, e15004. https://doi.org/10.1371/journal.pone.0015004 (2010).
Tarca, A. L., Chaemsaithong, P., Chaiworapongsa, T., Hassan, S. S. & Romero, R. Kits and methods to distinguish false labor and true labor. United States patent 20220065867 (2022-03-03)
Kim, C. H. et al. Stability and reproducibility of proteomic profiles measured with an aptamer-based platform. Sci. Rep. 8, 8382. https://doi.org/10.1038/s41598-018-26640-w (2018).
Joshi, A. & Mayr, M. In aptamers they trust: the caveats of the somascan biomarker discovery platform from somalogic. Circulation 138, 2482–2485. https://doi.org/10.1161/CIRCULATIONAHA.118.036823 (2018).
Michaels, J. E. et al. Comprehensive proteomic analysis of the human amniotic fluid proteome: Gestational age-dependent changes. J. Proteome Res. 6, 1277–1285. https://doi.org/10.1021/pr060543t (2007).
Hui, L., Wick, H. C., Edlow, A. G., Cowan, J. M. & Bianchi, D. W. Global gene expression analysis of term amniotic fluid cell-free fetal RNA. Obstet Gynecol. 121, 1248–1254. https://doi.org/10.1097/AOG.0b013e318293d70b (2013).
Tarca, A. L. et al. Amniotic fluid cell-free transcriptome: A glimpse into fetal development and placental cellular dynamics during normal pregnancy. BMC Med. Genomics 13, 25. https://doi.org/10.1186/s12920-020-0690-5 (2020).
Nilsson, S., Ramstrom, M., Palmblad, M., Axelsson, O. & Bergquist, J. Explorative study of the protein composition of amniotic fluid by liquid chromatography electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry. J. Proteome Res. 3, 884–889. https://doi.org/10.1021/pr0499545 (2004).
Tsangaris, G. T. et al. The normal human amniotic fluid supernatant proteome. In Vivo 20, 479–490 (2006).
Cho, C. K., Shan, S. J., Winsor, E. J. & Diamandis, E. P. Proteomics analysis of human amniotic fluid. Mol. Cell Proteom. 6, 1406–1415. https://doi.org/10.1074/mcp.M700090-MCP200 (2007).
Liu, X., Song, Y., Guo, Z., Sun, W. & Liu, J. A comprehensive profile and inter-individual variations analysis of the human normal amniotic fluid proteome. J. Proteom. 192, 1–9. https://doi.org/10.1016/j.jprot.2018.04.023 (2019).
Candia, J. et al. Assessment of Variability in the SOMAscan Assay. Sci. Rep. 7, 14248. https://doi.org/10.1038/s41598-017-14755-5 (2017).
Billing, A. M. et al. Complementarity of SOMAscan to LC-MS/MS and RNA-seq for quantitative profiling of human embryonic and mesenchymal stem cells. J. Proteom. 150, 86–97. https://doi.org/10.1016/j.jprot.2016.08.023 (2017).
Romero, R. et al. The maternal plasma proteome changes as a function of gestational age in normal pregnancy: A longitudinal study. Am. J. Obstet. Gynecol. 217, 67 e61–67 e21. https://doi.org/10.1016/j.ajog.2017.02.037 (2017).
Erez, O. et al. The prediction of late-onset preeclampsia: Results from a longitudinal proteomics study. PLoS One 12, e0181468. https://doi.org/10.1371/journal.pone.0181468 (2017).
Tarca, A. L. et al. The prediction of early preeclampsia: Results from a longitudinal proteomics study. PLoS One 14, e0217273. https://doi.org/10.1371/journal.pone.0217273 (2019).
Shainker, S. A. et al. Placenta accreta spectrum: Biomarker discovery using plasma proteomics. Am. J. Obstet. Gynecol. 223, 433 e431–433 e414. https://doi.org/10.1016/j.ajog.2020.03.019 (2020).
Tarca, A. L. et al. Crowdsourcing assessment of maternal blood multi-omics for predicting gestational age and preterm birth. Cell Rep. Med. 2, 100323. https://doi.org/10.1016/j.xcrm.2021.100323 (2021).
Kofinas, G. D., Kofinas, A. D., Pyrgerou, M. & Reyes, F. I. Amniotic fluid beta-endorphin levels and labor. Obstet. Gynecol. 69, 945–947 (1987).
Kubota, T., Tsuzuki, H. & Saito, M. Determination of prolactin, growth hormone, beta-endorphin, and cortisol in both maternal plasma and amniotic fluid during human gestation. Acta Endocrinol. (Copenh) 121, 297–303. https://doi.org/10.1530/acta.0.1210297 (1989).
Romero, R. et al. Neutrophil attractant/activating peptide-1/interleukin-8 in term and preterm parturition. Am. J. Obstet. Gynecol. 165, 813–820. https://doi.org/10.1016/0002-9378(91)90422-n (1991).
Romero, R. et al. Tumor necrosis factor in preterm and term labor. Am. J. Obstet. Gynecol. 166, 1576–1587. https://doi.org/10.1016/0002-9378(92)91636-o (1992).
Romero, R. et al. Interleukin-1 alpha and interleukin-1 beta in preterm and term human parturition. Am. J. Reprod. Immunol. 27, 117–123. https://doi.org/10.1111/j.1600-0897.1992.tb00737.x (1992).
Saito, S., Kato, Y., Ishihara, Y. & Ichijo, M. Amniotic fluid granulocyte colony-stimulating factor in preterm and term labor. Clin. Chim. Acta 208, 105–109. https://doi.org/10.1016/0009-8981(92)90027-n (1992).
Laham, N., Rice, G. E., Bishop, G. J., Ransome, C. & Brennecke, S. P. Interleukin 8 concentrations in amniotic fluid and peripheral venous plasma during human pregnancy and parturition. Acta Endocrinol. (Copenh) 129, 220–224. https://doi.org/10.1530/acta.0.1290220 (1993).
Casey, M. L., Brown, C. E., Peters, M. & MacDonald, P. C. Endothelin levels in human amniotic fluid at mid-trimester and at term before and during spontaneous labor. J. Clin. Endocrinol. Metab. 76, 1647–1650. https://doi.org/10.1210/jcem.76.6.8501173 (1993).
Saito, S., Kasahara, T., Kato, Y., Ishihara, Y. & Ichijo, M. Elevation of amniotic fluid interleukin 6 (IL-6), IL-8 and granulocyte colony stimulating factor (G-CSF) in term and preterm parturition. Cytokine 5, 81–88. https://doi.org/10.1016/1043-4666(93)90027-3 (1993).
Opsjln, S. L. et al. Tumor necrosis factor, interleukin-1, and interleukin-6 in normal human pregnancy. Am. J. Obstet. Gynecol. 169, 397–404. https://doi.org/10.1016/0002-9378(93)90096-2 (1993).
Gunn, L., Hardiman, P., Tharmaratnam, S., Lowe, D. & Chard, T. Measurement of interleukin-1 alpha and interleukin-6 in pregnancy-associated tissues. Reprod. Fertil. Dev. 8, 1069–1073. https://doi.org/10.1071/rd9961069 (1996).
Olah, K. S., Vince, G. S., Neilson, J. P., Deniz, G. & Johnson, P. M. Interleukin-6, interferon-gamma, interleukin-8, and granulocyte-macrophage colony stimulating factor levels in human amniotic fluid at term. J. Reprod. Immunol. 32, 89–98. https://doi.org/10.1016/s0165-0378(96)00990-4 (1996).
Mitchell, M. D., Hunter, C., Dudley, D. J. & Varner, M. W. Significant decrease in parathyroid hormone-related protein concentrations in amniotic fluid with labour at term but not preterm. Reprod. Fertil. Dev. 8, 231–234. https://doi.org/10.1071/rd9960231 (1996).
Laham, N. et al. Tumor necrosis factor-beta in human pregnancy and labor. J. Reprod. Immunol. 33, 53–69. https://doi.org/10.1016/s0165-0378(97)01012-7 (1997).
Rice, G. E., Reimert, C. M. & Bendtzen, K. Eosinophil cationic protein and eosinophil protein X: Human amniotic fluid concentrations and gestational tissue content at term. Placenta 19, 181–185. https://doi.org/10.1016/s0143-4004(98)90007-8 (1998).
Maymon, E. et al. The tumor necrosis factor alpha and its soluble receptor profile in term and preterm parturition. Am. J. Obstet. Gynecol. 181, 1142–1148. https://doi.org/10.1016/s0002-9378(99)70097-9 (1999).
Athayde, N. et al. A role for the novel cytokine RANTES in pregnancy and parturition. Am. J. Obstet. Gynecol. 181, 989–994. https://doi.org/10.1016/s0002-9378(99)70337-6 (1999).
Athayde, N. et al. Matrix metalloproteinases-9 in preterm and term human parturition. J. Matern. Fetal Med. 8, 213–219. https://doi.org/10.1002/(SICI)1520-6661(199909/10)8:5%3c213::AID-MFM3%3e3.0.CO;2-R (1999).
Denison, F. C., Kelly, R. W., Calder, A. A. & Riley, S. C. Secretory leukocyte protease inhibitor concentration increases in amniotic fluid with the onset of labour in women: Characterization of sites of release within the uterus. J. Endocrinol. 161, 299–306. https://doi.org/10.1677/joe.0.1610299 (1999).
Maymon, E. et al. Human neutrophil collagenase (matrix metalloproteinase 8) in parturition, premature rupture of the membranes, and intrauterine infection. Am. J. Obstet. Gynecol. 183, 94–99. https://doi.org/10.1067/mob.2000.105344 (2000).
Ietta, F. et al. Macrophage migration inhibitory factor in human pregnancy and labor. Am. J. Reprod. Immunol. 48, 404–409. https://doi.org/10.1034/j.1600-0897.2002.01152.x (2002).
Florio, P. et al. High levels of human chromogranin A in umbilical cord plasma and amniotic fluid at parturition. J. Soc. Gynecol. Investig. 9, 32–36. https://doi.org/10.1016/s1071-5576(01)00149-6 (2002).
Helmig, B. R. et al. Neutrophil elastase and secretory leukocyte protease inhibitor in prelabor rupture of membranes, parturition and intra-amniotic infection. J. Matern. Fetal. Neonatal. Med. 12, 237–246. https://doi.org/10.1080/jmf.12.4.237.246 (2002).
Espinoza, J. et al. Antimicrobial peptides in amniotic fluid: defensins, calprotectin and bacterial/permeability-increasing protein in patients with microbial invasion of the amniotic cavity, intra-amniotic inflammation, preterm labor and premature rupture of membranes. J. Matern. Fetal. Neonatal. Med. 13, 2–21. https://doi.org/10.1080/jmf.13.1.2.21 (2003).
Esplin, M. S. et al. Amniotic fluid levels of immunoreactive monocyte chemotactic protein-1 increase during term parturition. J. Matern. Fetal. Neonatal. Med. 14, 51–56. https://doi.org/10.1080/jmf.14.1.51.56 (2003).
Park, K. H. et al. Matrix metalloproteinase 3 in parturition, premature rupture of the membranes, and microbial invasion of the amniotic cavity. J. Perinat. Med. 31, 12–22. https://doi.org/10.1515/JPM.2003.002 (2003).
Gotsch, F. et al. Evidence of the involvement of caspase-1 under physiologic and pathologic cellular stress during human pregnancy: A link between the inflammasome and parturition. J. Matern. Fetal. Neonatal. Med. 21, 605–616. https://doi.org/10.1080/14767050802212109 (2008).
Chaiworapongsa, T. et al. The concentration of surfactant protein-A in amniotic fluid decreases in spontaneous human parturition at term. J. Matern. Fetal. Neonatal. Med. 21, 652–659. https://doi.org/10.1080/14767050802215193 (2008).
Vaisbuch, E. et al. Total hemoglobin concentration in amniotic fluid is increased in intraamniotic infection/inflammation. Am. J. Obstet. Gynecol. 199(426), e421-427. https://doi.org/10.1016/j.ajog.2008.06.075 (2008).
Chaiworapongsa, T. et al. Amniotic fluid heat shock protein 70 concentration in histologic chorioamnionitis, term and preterm parturition. J. Matern. Fetal. Neonatal. Med. 21, 449–461. https://doi.org/10.1080/14767050802054550 (2008).
Hamill, N. et al. Exodus-1 (CCL20): Evidence for the participation of this chemokine in spontaneous labor at term, preterm labor, and intrauterine infection. J. Perinat. Med. 36, 217–227. https://doi.org/10.1515/JPM.2008.034 (2008).
Kemp, B., von Rango, U., Brehme, A. & Rath, W. Amniotic fluid adhesion molecules during parturition at term. J. Perinat. Med. 37, 28–31. https://doi.org/10.1515/JPM.2009.002 (2009).
Panaitescu, B. et al. In vivo evidence of inflammasome activation during spontaneous labor at term. J. Matern. Fetal. Neonatal. Med. 32, 1978–1991. https://doi.org/10.1080/14767058.2017.1422714 (2019).
Romero, R. et al. Increased concentrations of arachidonic acid lipoxygenase metabolites in amniotic fluid during parturition. Obstet. Gynecol. 70, 849–851 (1987).
Lee, S. E. et al. Amniotic fluid prostaglandin concentrations increase before the onset of spontaneous labor at term. J. Matern. Fetal. Neonatal. Med. 21, 89–94. https://doi.org/10.1080/14767050701830514 (2008).
Maymon, E. et al. Evidence of in vivo differential bioavailability of the active forms of matrix metalloproteinases 9 and 2 in parturition, spontaneous rupture of membranes, and intra-amniotic infection. Am. J. Obstet. Gynecol. 183, 887–894. https://doi.org/10.1067/mob.2000.108878 (2000).
Behnia, F. et al. Chorioamniotic membrane senescence: a signal for parturition?. Am. J. Obstet. Gynecol. 213(359), e351–e316. https://doi.org/10.1016/j.ajog.2015.05.041 (2015).
Osmers, R. et al. Origin of cervical collagenase during parturition. Am. J. Obstet. Gynecol. 166, 1455–1460. https://doi.org/10.1016/0002-9378(92)91619-l (1992).
Winkler, M. et al. Parturition at term: parallel increases in interleukin-8 and proteinase concentrations and neutrophil count in the lower uterine segment. Hum. Reprod. 14, 1096–1100. https://doi.org/10.1093/humrep/14.4.1096 (1999).
Gomez-Lopez, N., StLouis, D., Lehr, M. A., Sanchez-Rodriguez, E. N. & Arenas-Hernandez, M. Immune cells in term and preterm labor. Cell. Mol. Immunol. 11, 571–581. https://doi.org/10.1038/cmi.2014.46 (2014).
Maheux, P. C. et al. Glucose homeostasis during spontaneous labor in normal human pregnancy. J. Clin. Endocrinol. Metab. 81, 209–215. https://doi.org/10.1210/jcem.81.1.8550753 (1996).
Scheepers, H. C., de Jong, P. A., Essed, G. G. & Kanhai, H. H. Fetal and maternal energy metabolism during labor in relation to the available caloric substrate. J. Perinat. Med. 29, 457–464. https://doi.org/10.1515/JPM.2001.064 (2001).
Dellios, N. L. et al. Increased expression of alpha-enolase in cervico-vaginal fluid during labour. Eur. J. Obstet. Gynecol. Reprod. Biol. 153, 16–22. https://doi.org/10.1016/j.ejogrb.2010.06.014 (2010).
Romero, R. et al. Isobaric labeling and tandem mass spectrometry: a novel approach for profiling and quantifying proteins differentially expressed in amniotic fluid in preterm labor with and without intra-amniotic infection/inflammation. J. Matern. Fetal Neonatal Med. 23, 261–280. https://doi.org/10.3109/14767050903067386 (2010).
Hong, S. et al. Identifying potential biomarkers related to pre-term delivery by proteomic analysis of amniotic fluid. Sci. Rep. 10, 19648. https://doi.org/10.1038/s41598-020-76748-1 (2020).
Gerson, K. D. et al. Second trimester short cervix is associated with decreased abundance of cervicovaginal lipid metabolites. Am. J. Obstet. Gynecol. https://doi.org/10.1016/j.ajog.2022.04.031 (2022).
Condon, J. C., Jeyasuria, P., Faust, J. M. & Mendelson, C. R. Surfactant protein secreted by the maturing mouse fetal lung acts as a hormone that signals the initiation of parturition. Proc. Natl. Acad. Sci. U S A 101, 4978–4983. https://doi.org/10.1073/pnas.0401124101 (2004).
Montalbano, A. P., Hawgood, S. & Mendelson, C. R. Mice deficient in surfactant protein A (SP-A) and SP-D or in TLR2 manifest delayed parturition and decreased expression of inflammatory and contractile genes. Endocrinology 154, 483–498. https://doi.org/10.1210/en.2012-1797 (2013).
Mendelson, C. R., Montalbano, A. P. & Gao, L. Fetal-to-maternal signaling in the timing of birth. J. Steroid. Biochem. Mol. Biol. 170, 19–27. https://doi.org/10.1016/j.jsbmb.2016.09.006 (2017).
Miyamura, K. et al. Surfactant proteins A (SP-A) and D (SP-D): levels in human amniotic fluid and localization in the fetal membranes. Biochim. Biophys. Acta 1210, 303–307. https://doi.org/10.1016/0005-2760(94)90233-x (1994).
Karjalainen, M. K. et al. A study of collectin genes in spontaneous preterm birth reveals an association with a common surfactant protein D gene polymorphism. Pediatr. Res. 71, 93–99. https://doi.org/10.1038/pr.2011.2 (2012).
Norwitz, E. R. et al. Molecular regulation of parturition: The role of the decidual clock. Cold Spring Harb. Perspect. Med. 5. https://doi.org/10.1101/cshperspect.a023143 (2015).
Menon, R. Fetal inflammatory response at the fetomaternal interface: A requirement for labor at term and preterm. Immunol. Rev. 308, 149–167. https://doi.org/10.1111/imr.13075 (2022).
Jindal, A., Sharma, M. & Chaudhary, C. in StatPearls (2022).
Bhatti, G. et al. The amniotic fluid cell-free transcriptome in spontaneous preterm labor. Sci. Rep. 11, 13481. https://doi.org/10.1038/s41598-021-92439-x (2021).
Esplin, M. S. et al. The use of cDNA microarray to identify differentially expressed labor-associated genes within the human myometrium during labor. Am. J. Obstet. Gynecol. 193, 404–413. https://doi.org/10.1016/j.ajog.2004.12.021 (2005).
Bukowski, R., Hankins, G. D., Saade, G. R., Anderson, G. D. & Thornton, S. Labor-associated gene expression in the human uterine fundus, lower segment, and cervix. PLoS Med 3, e169. https://doi.org/10.1371/journal.pmed.0030169 (2006).
Peng, H. H. et al. The effects of labor on differential gene expression in parturient women, placentas, and fetuses at term pregnancy. Kaohsiung J. Med. Sci. 27, 494–502. https://doi.org/10.1016/j.kjms.2011.06.012 (2011).
Chan, Y. W., van den Berg, H. A., Moore, J. D., Quenby, S. & Blanks, A. M. Assessment of myometrial transcriptome changes associated with spontaneous human labour by high-throughput RNA-seq. Exp. Physiol. 99, 510–524. https://doi.org/10.1113/expphysiol.2013.072868 (2014).
Butt, R. H. et al. An initial proteomic analysis of human preterm labor: Placental membranes. J. Proteome Res. 5, 3161–3172. https://doi.org/10.1021/pr060282n (2006).
Lockwood, C. J. et al. Fetal membrane rupture is associated with the presence of insulin-like growth factor-binding protein-1 in vaginal secretions. Am. J. Obstet. Gynecol. 171, 146–150. https://doi.org/10.1016/0002-9378(94)90461-8 (1994).
Akercan, F. et al. The value of the insulin-like growth factor binding protein-1 in the cervical-vaginal secretion detected by immunochromatographic dipstick test in the prediction of delivery in women with clinically unconfirmed preterm premature rupture of membranes. Eur. J. Obstet. Gynecol. Reprod. Biol. 121, 159–163. https://doi.org/10.1016/j.ejogrb.2004.12.006 (2005).
Heng, Y. J. et al. Human cervicovaginal fluid biomarkers to predict term and preterm labor. Front. Physiol. 6, 151. https://doi.org/10.3389/fphys.2015.00151 (2015).
Galletta, M. A. K., Bittar, R. E., Rodrigues, A. S., Francisco, R. P. V. & Zugaib, M. Comparative analysis of Insulin-like growth factor binding protein-1, placental alpha-microglobulin-1, phenol and pH for the diagnosis of preterm premature rupture of membranes between 20 and 36 weeks. J. Obstet. Gynaecol. Res. 45, 1448–1457. https://doi.org/10.1111/jog.13991 (2019).
Aoki, S., Shteyn, K. & Marian, R. Biorender: Create professional science figures in minutes (2022).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43, e47. https://doi.org/10.1093/nar/gkv007 (2015).
EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling v. 1.12.0 (2021).
Pheatmap: pretty heatmaps v. 1.0.12 (2019).
Wickham, H. in ggplot2 189–201 (Springer, 2016).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740. https://doi.org/10.1093/bioinformatics/btr260 (2011).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Kassambara, A. ggpubr:“ggplot2” based publication ready plots. (2020).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77. https://doi.org/10.1186/1471-2105-12-77 (2011).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R news 2, 18–22 (2002).
Wright, M. N. & Ziegler, A. ranger: A fast implementation of random forests for high dimensional data in C++ and R. arXiv preprint arXiv:1508.04409 (2015).
Author information
Authors and Affiliations
Contributions
A.L.T., G.B., N.G.L., and R.R.conceived and performed the research. G.B., A.L.T., and N.G.L. drafted the manuscript. T.C., N.G.T., K.R.T., J.G., F.G., R.P.R., S.M.B., and M.K. reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bhatti, G., Romero, R., Gomez-Lopez, N. et al. The amniotic fluid proteome changes with term labor and informs biomarker discovery in maternal plasma. Sci Rep 13, 3136 (2023). https://doi.org/10.1038/s41598-023-28157-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-28157-3
- Springer Nature Limited