Abstract
To identify potential plasma biomarkers associated with microbial invasion of the amniotic cavity (MIAC) and/or intraamniotic inflammation (IAI) in women with preterm premature rupture of membranes (PPROM). This retrospective cohort study included 182 singleton pregnant women with PPROM (23–33 weeks) who underwent amniocentesis. Plasma samples; all subjects were chosen from these participants and were analyzed using label-free liquid chromatography-tandem mass spectrometry for proteome profiling using a nested case–control study design (cases with MIAC/IAI vs. non-MIAC/IAI controls [n = 9 each]). Three identified target molecules for MIAC/IAI were further verified by ELISA in the study cohort (n = 182). Shotgun proteomic analysis revealed 17 differentially expressed proteins (P < 0.05) in the plasma of MIAC/IAI cases. In particular, the levels of FCGR3A and haptoglobin, but not LRP1, were found to be increased in the plasma of patients with MIAC, IAI, and both MIAC/IAI compared with those without these conditions. Moreover, these differences remained significant after adjusting for gestational age at sampling. The area under the curves of plasma FCGR3A and haptoglobin ranged within 0.59–0.65 with respect to each of the three outcome measures. Plasma FCGR3A and haptoglobin were identified as potential independent biomarkers for less-invasively detecting MIAC/IAI in women with PPROM.
Similar content being viewed by others
Introduction
Preterm birth affects approximately 10% of all pregnancies and remains the leading cause of neonatal mortality and morbidity worldwide1. In particular, preterm premature rupture of membranes (PPROM), which occurs in 3–4% of all deliveries, is an important antecedent to approximately 30% of preterm births1,2,3. PPROM is often complicated by the subclinical presence of microorganisms in the amniotic fluid (AF) (i.e., microbial invasion of the amniotic cavity [MIAC]) and/or intraamniotic inflammation (IAI), with a frequency of approximately 50%4,5,6. A solid body of evidence suggests that the presence of MIAC/IAI is associated with additional risks of adverse pregnancies (i.e., delivery latency) and neonatal short- and long-term outcomes, which may be caused by fetal inflammation and injury to the immature organs7,8,9,10. Importantly, recent studies have shown that intravenous clarithromycin therapy may reduce the intensity of the intraamniotic inflammatory response in patients with PPROM with either MIAC or sterile IAI6,11. Thus, accurate and early identification of women at high risk for MIAC/IAI allows targeted use of novel therapeutics that can substantially reduce the incidence of complications in women with PPROM and their children, as well as improve the use of resources, such as transfer to a tertiary center, and use of corticosteroids and magnesium for neuroprotection.
AF analysis via amniocentesis for various interleukins (ILs) and matrix metalloproteinases remains the gold standard method for identifying MIAC/IAI complicated by PPROM2,4,8,12,13. However, this approach is clinically challenging owing to the need of invasive procedures and technical difficulty in some cases of PPROM with severe oligohydramnios secondary to ruptured membranes. In this context, a maternal blood sample, which can be obtained via a less invasive, easy-to-use, and inexpensive method, could be a preferable alternative to AF. Indeed, several studies have shown a significant elevation of various inflammatory proteins that occur concurrently in the AF and maternal blood compartments in the setting of MIAC/IAI and spontaneous preterm delivery (SPTD)12,14,15. However, to date, few studies have explored the predictive potential of these blood protein mediators for MIAC/IAI complicated by PPROM, particularly using high-throughput proteomic methodologies.
Mass spectrometry-based shotgun proteomics coupled with multidimensional chromatography separation has recently emerged as a high-throughput technique for characterizing proteomes with low abundance in complex biological samples (plasma and serum)16. This approach has shown promising results in the discovery of new protein markers for diseases with complex phenotypes, such as various forms of infection/inflammation17,18,19,20,21. The purpose of this study was (i) to identify potential plasma biomarkers associated with MIAC/IAI in women with PPROM using label-free shotgun proteomic analysis and (ii) to determine the top-ranked protein pathways activated under these conditions.
Materials and methods
Ethical approval
The study was approved by the local ethics committee of the Seoul National University Bundang Hospital, Seongnamsi, Korea (project number B-1105/128-102). All experiments were performed in accordance with the relevant guidelines and regulations of the ethics committee of the hospital. All participants provided written informed consent to collect and use biological samples and clinical information prior to the amniocentesis procedure.
Study population and research design
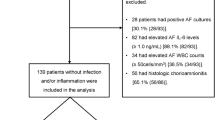
This retrospective study enrolled 182 singleton pregnant women admitted to the Department of Obstetrics and Gynecology at the Seoul National University Bundang Hospital, Seongnamsi, Korea, with a diagnosis of PPROM at 23 + 0 to 33 + 6 weeks of gestation between June 2004 and April 2019. The inclusion criteria were the following: (i) performance of transabdominal amniocentesis to assess possible subclinical intraamniotic infection and inflammation; (ii) delivery of a live fetus; and (iii) availability of a maternal plasma sample collected at the time of the amniocentesis. Participants were excluded if they had (i) active labor (defined as cervical dilation ≥ 4 cm in the presence of uterine contraction by sterile speculum examination), (ii) multiple pregnancies, (iii) a fetus with major congenital anomalies, and (iv) clinical chorioamnionitis at the time of admission. Gestational age was determined using the last menstrual period and ultrasound estimates based on first or second (≤ 20 weeks) trimester fetal biometry. PPROM was defined as clinically confirmed amniorrhexis occurring prior to labor onset and at < 37 weeks of gestation. This condition was diagnosed visually by sterile speculum examination to confirm the pooling of AF in the vagina (or AF leakage through the cervix) and a positive nitrazine test (and/or a positive AmniSure ROM test [Qiagen, Hilden, Germany]).
For the discovery phase of the study (Fig. 1A), a nested case–control approach was performed, comprising nine patients with MIAC (who also had IAI [case subjects]) and nine gestational age-matched patients without MIAC and IAI (control subjects). The case patients were randomly selected among the 52 patients with MIAC/IAI within the total study cohort of patients with PPROM using a random sequence generator. Each control patient was matched to the case patient in terms of gestational age at sampling, parity, years of admission, and maternal age.
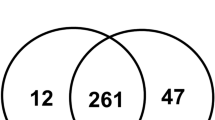
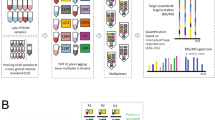
(A) Schematic workflow for the discovery (label-free LC–MS/MS) and verification (immunoassay) experiments. Three sets of pooled plasma samples from control (non-MIAC/IAI) and case (MIAC/IAI) groups were subjected to immunoaffinity depletion to remove the 14 most abundant proteins followed by tryptic digestion. Peptides were fractionated with high-pH reversed phase chromatography, after which were subjected to LC–MS/MS followed by label-free quantitative analysis based on peak intensities. Differentially expressed proteins were further investigated by IPA and the DEPs of interest were validated by ELISA. (B) Venn diagrams showing the distribution of common and uniquely peptides (above) and proteins (below) identified in the LC–MS/MS analyses. MIAC microbial invasion of the amniotic cavity, IAI intra-amniotic inflammation, LC liquid chromatography MS/MS tandem mass spectrometry, IPA Ingenuity pathway analysis, DEP differentially expressed proteins, ELISA enzyme-linked immunosorbent assaym, FCGR3A low affinity immunoglobulin gamma Fc region receptor III-A, LRP1 prolow-density lipoprotein receptor-related protein 1.
Diagnosis of MIAC and IAI using AF samples
Ultrasound-guided transabdominal amniocentesis was performed under aseptic conditions at the time of admission. AF samples were sent to the hospital laboratory for culturing aerobic and anaerobic bacteria, genital mycoplasma (Ureaplasma urealyticum and Mycoplasma hominis), and fungi, as well as for assessing the white blood cell count (WBC), as previously described22. The remaining AF was centrifuged for 10 min at 1500 × g, and the supernatant was divided into aliquots and stored at − 70 °C until further analysis. The managing physicians had access to the results of the AF analysis (AF culture results and WBC counts, but not IL-6 quantification). MIAC was defined as the presence of a positive AF culture for bacteria, genital mycoplasma (Ureaplasma spp. or M. hominis), and/or fungi.
AF IL-6 levels were assayed using the enzyme-linked immunosorbent assay (ELISA) human IL-6 DuoSet Kit (R&D Systems, Minneapolis, MN, USA) to define subclinical IAI. A detailed description of the measurements of IL-6 concentrations in AF is provided in Supplementary Methods. IAI was diagnosed when the AF IL-6 concentration was ≥ 2.6 ng/mL, as previously described23,24,25.
Collection and storage of plasma samples
On admission to a hospital, at the time of amniocentesis, prior to the administration of medications, maternal venous blood samples were collected into ethylenediaminetetraacetic acid tubes after measuring the C-reactive protein (CRP) concentration and WBC counts in patients diagnosed with PPROM, as part of the hospital protocol. Plasma was separated from the blood samples by centrifugation at 1500 × g at 4 °C for 10 min and stored in multiple aliquots at − 70 °C until further use.
Management of PPROM and clinical definitions of various factors
Management of PPROM has been previously described in detail26,27. Briefly, prophylactic antibiotics (macrolides plus ampicillin) were administered to all patients with PPROM. Tocolytic therapy (magnesium sulfate, ritodrine, or atosiban) and a course of corticosteroid treatment were administered to women with PPROM at 23–34 weeks of gestation at the discretion of the clinician. In patients with PPROM at < 34 weeks of gestation who proved to have positive AF cultures, the decision for delivery was not made only based on the positive AF cultures results; delivery was considered if the woman was diagnosed or suspected of chorioamnionitis or whose fetus was diagnosed or suspected of being in jeopardy. Delivery was considered for all women with PPROM ≥ 34 weeks. Acute histologic chorioamnionitis (HCA) was diagnosed based on the presence of acute inflammation (defined by neutrophil infiltration) in any placental tissue (chorionic plate, umbilical cord, or fetal membranes [chorion-decidua and amnion]) in accordance with previously published criteria28,29. Clinical chorioamnionitis was diagnosed using the criteria proposed by Gibbs et al.30; more detailed criteria are provided in Supplementary Methods.
Proteomic analysis (discovery phase)
Protein concentration was determined using a bicinchoninic acid protein assay kit (Thermo Fisher Scientific, Bremen, Germany) in 18 exploratory cohort samples (Fig. 1A). In each case (n = 9) and control (n = 9) groups, three sets of plasma samples were pooled in equal amounts per sample, thus creating three sets of pooled plasma samples for the case and control groups. These pooled plasma samples were subjected to immunoaffinity depletion to remove 14 high-abundance proteins, tryptic digestion, and high-pH reversed-phase peptide fractionation. The fractionated peptide samples were then analyzed in triplicate by liquid chromatography-tandem mass spectrometry (LC–MS/MS) using an Eksigent MDLC system (Eksigent Technologies, Dublin, CA, USA) interfaced to an LTQ XL-Orbitrap mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA).
For protein identification, a protein database search was performed against the SwissProt human database (20,329 entries; release date: July 2020) using MaxQuant (version 1.6.7.0)31,32. The false discovery rate for peptides and proteins was set at 1%, and all proteins were identified using two or more unique peptides. For quantification purposes, the MaxLFQ algorithm was employed for label-free quantification (LFQ) of the proteins. Proteins with fold change of LFQ intensities > 1.3 or < 0.77 and P < 0.05 were considered as differentially expressed proteins (DEPs). Perseus software (1.6.14.0; https://maxquant.net/perseus/) was used for the statistical analysis of the label-free quantitative datasets33,34. Detailed descriptions of the discovery proteomics experiments are provided in Supplementary Methods.
Ingenuity pathway analysis (IPA)
The proteomic dataset, which included UniProt accession numbers of the DEPs and their corresponding log2|Fold Change| of LFQ intensities, was submitted to IPA (data version 65,367,011; QIAGEN, Redwood City, CA) for functional analysis to identify the canonical pathways, diseases, and biological functions involved. The uploaded DEPs were mapped to the corresponding gene objects in the Ingenuity Pathways Knowledge Base as a reference set35,36. A right-tailed Fisher’s exact test was used to determine the likelihood of the mapped genes in each pathway (network) to be found together due to random chance.
ELISA (validation phase)
Three selected candidate DEPs were validated in the study cohort, comprising 182 individual samples. The concentrations of haptoglobin (DuoSet ELISA, R&D Systems, Minneapolis, MN, USA), low-affinity immunoglobulin gamma Fc region receptor III-A (FCGR3A), and prolow-density lipoprotein receptor-related protein 1 (LRP1) (MyBioSource, San Diego, CA, USA) were assayed using commercial ELISA kits, according to the manufacturer’s instructions. Aliquots of the frozen plasma were thawed at room temperature (25 °C) for up to 1–2 h and vortexed thoroughly prior to analysis. The plasma dilutions used and the working range for each ESISA kit are described in detail in Supplementary Methods. The intra- and inter-assay coefficients of variation (CVs) were of < 10% for all analyzed proteins, except for the inter-assay CVs of FCGR3A (11.2%) and haptoglobin (11.4%). The above-mentioned three target molecules were selected for the validation study because: i) they revealed a high differential expression (i.e., fold change) or statistical significance; ii) little or no information was available regarding their expression change in the plasma in relation to MIAC/IAI; iii) they could have potential clinical relevance in the plasma with respect to inflammation/infection, considering their biological functions; and iv) ELISA kits for these proteins were readily available. Insulin-like growth factor II (DuoSet ELISA, R&D Systems, Minneapolis, MN, USA) and properdin (complement factor P; MyBioSourceSan Diego, CA, USA) were also assessed in the plasma using spike-and-recovery and linearity-of-dilution testing, but the results revealed poor assay performance; hence, these molecules were not further investigated.
Statistical analysis
The clinical data and plasma levels of the proteins were compared using Fisher’s exact test or the χ2-test for categorical data, Student’s t-test for normally distributed continuous data, and the Mann–Whitney U-test for non-normally distributed continuous data. Multivariate logistic regression was performed to estimate independent associations between plasma levels of each protein and the outcome measures after adjusting for gestational age at sampling and parity, which had a P-value < 0.1 in univariate analyses. Additionally, to determine the best blood-based multi-marker panels for MIAC, IAI, and microbial-associated IAI, multivariate analyses with a forward selection were performed on the two newly identified plasma biomarkers (FCGR3A, haptoglobin) along with conventional inflammatory marker in the blood (serum CRP), which were selected based on P < 0.1 in univariate analysis. The optimal cutoff and diagnostic value of each candidate protein were assessed through the analysis of their receiver operating characteristic (ROC) and area under the curve (AUCs). Thereafter, pairwise comparisons of the AUCs of each investigated plasma biomarker, serum CRP (as standard of inflammation biomarker), and multi-marker panel were performed using the method proposed by DeLong et al.37. The optimal cutoff value was determined using the maximal Youden's index (sum of sensitivity + specificity − 1). Potential interactions between three proteins profiled (FCGR3A, haptoglobin, and LRP1) were assessed using type-III tests. Spearman rank correlation was conducted to assess the linear correlations of each protein level. All probability values were two-sided and P < 0.05 were considered significant. Statistical analyses were performed using SPSS version 25.0 (IBM Inc., Armonk, NY, USA).
Results
Characteristics of the discovery cohort
The demographic and clinical characteristics of the exploratory cohort used for shotgun proteomics are shown in Table S1. Owing to the matched selection criteria, the cases and controls had similar gestational age at sampling, use of medications, maternal age, and parity. Nine patients with MIAC complicated by PPROM were positive for U. urealyticum (n = 9), M. hominis (n = 6), and Peptostreptococcus spp. (n = 1) as determined by AF culture, among whom 77.7% (7/9) of patients in the MIAC group had polymicrobial invasion.
Exploratory proteomic analysis
Overall, LC–MS/MS analyses of three plasma biological replicates revealed 4285 and 3964 unique peptides in the case and control groups, respectively, whereas 3566 peptides were identified in both groups (Fig. 1B). A total of 408 proteins were identified in the plasma of the control group and 396 in the case group, whereas 391 proteins were present in both groups (Fig. 1B and Table S2). Among these shared proteins, 7 (41.2%) were significantly upregulated and 10 (58.8%) were downregulated in MIAC/IAI cases as compared with cases without these conditions (Fig. 2 and Table S3). Hierarchical clustering analysis of these 17 DEPs further confirmed that their expression pattern in the MIAC/IAI cases was significantly different from that of the No-MIAC/Non-IAI controls (Fig. S1). Analysis of these 17 DEPs using IPA revealed five canonical pathways (Table S4): ‘acute phase response signaling (APR),’ ‘growth hormone signgling,’ ‘iron homeostasis signaling pathway,’ peroxisome proliferator-activated receptors (PPAR)/retinoid X receptor (RXR) activation,’ and ‘hypoxia-inducible factor 1 (HIF-1) signaling.’ IPA also identified ‘inflammatory response,’ ‘cancer,’ ‘organismal injury and abnormalities,’ and ‘metabolic disease’ as the top diseases and disorders associated with the presence of MIAC and IAI complicated by PPROM.
Label-free quantification proteomic data of DEPs in MIAC/IAI case vs. non-MIAC/IAI control. Representative protein identifiers in red indicate statistically significant DEPs with P < 0.05 and fold change of LFQ intensities > 1.3 or < 0.77. DEP differentially expressed proteins, MIAC microbial invasion of the amniotic cavity, IAI intra-amniotic inflammation, LFQ label-free quantification.
Verification of proteomic data in the total cohort
In the total study cohort (n = 182), the overall rates of MIAC and IAI were 34.6% (63/182) and 42.8% (78/182), respectively, and 28.5% (52/182) of women had both MIAC and IAI (microbial-associated IAI). MIAC and IAI alone were present in 6.0% (11/182) and 14.2% (26/182) of women, respectively, whereas 51% (93/182) of women exhibited neither MIAC nor IAI. Genital mycoplasmas (U. urealyticum [n = 49] and/or M. hominis [n = 33]) were the most common microbes found in the AF. Other microorganisms isolated from AF samples included Streptococcus agalactiae (n = 3), Peptostreptococcus spp. (n = 3), Streptococcus viridans (n = 2), Lactobacillus spp. (n = 2), Streptococcus mitis (n = 1), Haemophilus influenzae (n = 1), Escherichia coli (n = 1), gram-negative rods (n = 1), Candida glabrata (n = 1), and gram-positive cocci (n = 1). Polymicrobial findings were present in 55.5% (35/63) of the patients with MIAC.
To verify the proteomic data, the levels of three candidate DEPs, FCGR3A, haptoglobin, and LRP1, were determined in MIAC/IAI cases and then compared with those of controls without these conditions. The median plasma levels of FCGR3A and haptoglobin were significantly higher in women with MIAC, IAI, and microbial-associated IAI than in women without these conditions (P < 0.05, Table 1). However, univariate analysis showed that women with MIAC, IAI, and microbial-associated IAI had significantly lower gestational age at sampling and gestational age at delivery than those without these conditions (Table 2). Moreover, women with IAI and microbial-associated IAI were more or tended to be more parous than those without these conditions (Table 2). Additional multivariate analysis further demonstrated that high plasma levels of haptoglobin were significantly associated with MIAC, IAI, and microbial-associated IAI after adjusting for gestational age at sampling and parity (Table 3). Similar associations were observed between FCGR3A and MIAC and microbial-associated IAI, whereas no association of FCGR3A with IAI was observed. However, based on univariate analyses, no significant differences in plasma LRP1 levels were observed in relation to MIAC, IAI, and microbial-associated IAI (Table 1). Interactions between plasma FCGR3A, haptoglobin, and LRP1 levels were not found for the presence of MIAC, IAI, and microbial-associated IAI (Table S8).
The AUC values of plasma FCGR3A and haptoglobin for predicting MIAC were 0.65 and 0.59, and for IAI were of 0.60 and 0.63, respectively (Table 4 and Fig. 3A,B). Similarly, for the prediction of microbial-associated IAI, the AUC values of plasma FCGR3A and haptoglobin were of 0.64 and 0.62, respectively (Table 4 and Fig. 3C). Differences in the AUC values between plasma FCGR3A and haptoglobin were not statistically significant for predicting any of the three outcome measures (P = 0.370–0.758). Moreover, the AUC values of plasma FCGR3A and haptoglobin were similar to those of serum CRP with respect to each of the corresponding outcome measures (P = 0.299–0.879).
Repetition of the univariate and multivariate analyses after excluding data that had been included in the discovery study (n = 164), the results for each outcome measure were confirmed, being the same as those observed in the total cohort (n = 182) (Tables S5, S6, and S7). No correlations were found among the measured plasma proteins (FCGR3A, haptoglobin, and LRP1) (all variables, r = − 0.078 to 0.051, P > 0.2).
In the multi-marker panel for the diagnosis of MIAC, plasma FCGR3 and serum CRP levels were identified as the best combination (Table S9), with an AUC value of 0.68 (95% confidence interval [CI]: 0.59–0.76; P = 0.380 by Hosmer–Lemeshow test), which was not significantly higher than those for plasma FCGR3 and serum CRP (P = 0.373 and 0.107, respectively) (Table 4). Similarly, for predicting microbial-associated IAI, plasma FCGR3 levels along with serum CRP levels were identified as the best combination (Table S9), with an AUC value of 0.69 (95% CI: 0.61–0.78; P = 0.184 by Hosmer–Lemeshow test). The AUC value for this two-biomarker panel was similar to those of plasma FCGR3 and serum CRP (P = 0.141 and 0.156, respectively) (Table 4). However, in the IAI predictive model, plasma FCGR3A, plasma haptoglobin, and serum CRP levels were set in the logistic regression model as predictors, but only serum CRP level was selected for the best multi-marker panel; thus, a predictive model for IAI could not be generated.
Discussion
In the current study of women with PPROM, (i) 17 significant plasma DEPs related to MIAC/IAI and their potential biological pathways were identified using label-free shotgun proteomics approaches; and (ii) these proteomic findings were further validated and confirmed by ELISA. In particular, FCGR3A and haptoglobin were found to be significantly elevated in the plasma of women with MIAC, IAI, and both MIAC/IAI compared with those without these conditions. To the best of our knowledge, this is the first report demonstrating a comprehensive profile of the plasma proteome related to MIAC/IAI in patients with PPROM using a high-throughput shotgun label-free quantitation approach. The present study provides new insights for a better understanding of the biochemical mechanisms and molecular signals occurring in maternal circulation that are associated with inflammatory/infectious processes in the amniotic cavity.
In the last two decades, several investigations employed proteomic approaches to explore biomarkers for MIAC/IAI complicated by PPROM or PTL in AF and cervicovaginal fluid samples18,19,20,38,39,40. They reported several new proteins, including α1-acid glycoprotein, IGFBP-1, calgranulins, lipocalin-2, myeloperoxidase, neutrophil protein 1–3, as being associated with MIAC/IAI. However, to date, no study employed proteomic techniques in maternal blood to screen for potential biomarkers of MIAC/IAI. In our plasma proteomics and immunoassays, for the first time, we identified plasma FCGR3A and haptoglobin as novel biomarkers that may be used to differentiate MIAC/IAI from non-MIAC/IAI patients complicated by PPROM.
FCGR3A, which is also known as CD16A, is a transmembrane glycoprotein receptor that is member of the Fc gamma receptor family. It is one of the major receptors for IgG and is expressed on natural killer cells, monocytes/macrophages, trophoblasts, and dendritic cells41,42,43. FCGR3A is involved in antibody-dependent cell-mediated cytotoxicity, cytokine production, phagocytosis, and removal of antigen–antibody complexes44. Mutations and aberrant expression of FCGR3A have been reported to contribute to increased susceptibility to several inflammatory and immune diseases, including systemic lupus erythematosus, recurrent infectious diseases, and childhood chronic immune thrombocytopenic purpura45,46,47. Indeed, Presicce et al. showed that intraamniotic injection of lipopolysaccharide increases FCGR3 expression in chorio-decidua neutrophils in rhesus macaques48. Negishi et al. also found that the accumulation of FCGR3+ natural killer cells is increased in the decidua basalis of women who have experienced late preterm birth with acute chorioamnionitis49. The results of the above studies, along with the main biological characteristics of FCGR3A, support and in general agree with the plasma FCGR3 data in the present study, considering that significant associations have been reported between MIAC/IAI and acute HCA/SPTD risk in the PPROM context26,27,50.
Haptoglobin is an acute-phase protein that is produced mainly by the liver in response to proinflammatory stimuli, such as IL-1, IL-6, and tumor necrosis factor (TNF)-α51. Haptoglobin binds to free hemoglobin and matrix metalloproteinase-9 in circulation52, acts as an antioxidant, exhibits proangiogenic and antibacterial activities, and plays an important role in innate immunity51. In line with its known biological properties in inflammation, haptoglobin has been reported to be elevated in the plasma of patients with various inflammatory conditions, including inflammatory, infectious, and malignant diseases, and diabetes51. Particularly in the perinatal field, previous studies have shown that haptoglobin expression is upregulated in the cord blood or placenta of newborns with early onset neonatal sepsis and antenatal exposure to inflammation/infection (acute HCA, MIAC, and IAI) compared with those without these conditions53,54. Furthermore, a previous study by Oggé et al. showed that haptoglobin expression is upregulated in the AF of women with chronic HCA compared with controls55. However, haptoglobin expression has not yet been evaluated in maternal blood from women with PPROM with respect to MIAC/IAI. In the present study, we demonstrated for the first time that higher plasma haptoglobin levels are independently associated with the occurrence of MIAC/IAI in pregnancies complicated by PPROM.
Noteworthily, despite the fact that plasma FCGR3A and haptoglobin can discriminate MIAC/IAI from non-MIAC/IAI among a cohort of women with PPROM with similar diagnostic accuracy to serum CRP (a prototype marker of inflammation tested in the blood), their diagnostic performance is poor to fair (AUC: 0.59–0.65; Table 4). These observations are similar to those reported for other maternal blood biomarkers and may be attributed to the common inherent characteristics of non-specific inflammation biomarkers12,26,56. Consequently, the clinical utility of the aforementioned plasma biomarkers alone may be limited in the PPROM setting; thus, whether these newly identified plasma-based biomarkers could contribute to identifying PPROM-associated MIAC/IAI when used in combination with other currently available non-invasive tests (e.g., cervical length and inflammatory cytokines in the cervicovaginal fluid) needs to be addressed in future studies5,12,57,58. In addition, considering the significant association between the inflammatory response in circulation and SPTD59,60,61, it is highly likely that these two biomarkers may be useful for identifying patients at high risk for infection/inflammation-associated preterm delivery, which deserves to be the focus of future studies.
In the present study, IPA revealed the most significant canonical pathways associated with the 17 DEPs identified in maternal plasma potentially involved in MIAC/IAI. Overall, the five top signaling pathways identified are parallel to those associated with SPTD development, as reported in previous proteomics studies on patients at risk for preterm delivery62,63,64,65. APR is a complex systemic inflammatory reaction triggered by various factors, including local infection/inflammation, trauma, and tissue damage66. APR is mediated by proinflammatory cytokines (notably IL-1, IL-6, and TNF-α), leading to the release of acute-phase proteins from hepatocytes into the plasma67,68. Previous studies have shown that the levels of proinflammatory cytokines are significantly elevated in the AF and maternal plasma (or serum) from women with MIAC/IAI complicated by PPROM, and that simultaneously increased levels of various acute-phase proteins occur in the blood during this condition12,23,56. Thus, it is natural that the APR signaling pathway plays an important role in the link between local infection/inflammation in the amniotic cavity and systemic inflammatory responses. In growth hormone signaling, growth hormones play an important role in the regulation of growth and metabolism (glucose and lipid) during development, which is mediated through insulin-like growth factor (IGF) 1 and 269, as well as the immune system70. Furthermore, reduced growth hormone levels are associated with increased plasma secretion of proinflammatory cytokines (such as IL-6 and TNF-α) and increased proinflammatory function of monocytes/macrophages69,71. Iron homeostasis acts as an essential regulator of adaptive and innate immunity, and plays a crucial role in inflammatory processes and immune responses, especially during infection72. In particular, immune cells (including B cells, T cells, and macrophages) require sufficient amounts of iron to sustain their development and effector functions; thus, iron deficiency robustly impairs T and B cell activation, and the production and release of cytokines72. The PPAR forms a heterodimeric DNA-binding complex with RXR, which is called “PPAR/RXR activation,” that plays a critical role in energy balance, including lipid metabolism and glucose homeostasis, and is also involved in the inflammatory and vascular responses73. HIF-1 signaling pathway is activated mainly by hypoxia and is modulated by cytokines (e.g., ILs) and growth factors (e.g., IGFs), and has been implicated in the regulation of inflammatory response, tumor progression, and metastasis74,75.
Our proteomic study was performed using semi-pooled samples (three pools containing three, three, or three samples in each pool). Noteworthy, sample pooling strategy for proteomic analysis have some disadvantages: (i) may not represent the biological average of individual samples, and (ii) reduces the statistical value of the identified biomarkers (false identification of biomarkers and their missed detection than when using individual samples)76. However, compared with the individual sample strategy, it provides the following advantages: (i) reduced biological variation, especially for individual proteomes not associated with MIAC/IAI; (ii) requires a small amount of samples for analysis; and (iii) reduced experimental time and cost76,77. Thus, the semi-pooling strategy herein used may compensate for the drawbacks inherent to pooling all 9 samples from each group together, as well as provide three independent biological replicates for each group78,79.
The current study had several limitations. First, conventional microbial culture-based techniques were used only for microorganism detection in AF specimens, which may have led to false-negative MIAC results. A combination of culture and non-culture methods, including 16S rDNA polymerase chain reaction, provide an opportunity to precisely identify and characterize the microbes in AF80. Second, stored (at − 70 °C) plasma samples were used, which may have affected the results of proteomic and immunoassay analyses owing to protein degradation caused by long-term storage81,82,83. Third, the retrospective nature of our study had inherent drawbacks, including selection bias, and the validation study of candidate biomarkers and clinical validation of cutoff values were not conducted in a completely independent dataset. All these factors may limit the generalizability of the present findings, which may warrant further validation in other cohorts. Fourth, the label-free quantification strategy based on peak intensity herein used may result on considerably lower quantification accuracy and precision than label-based quantitation methods84. Nevertheless, the label-free approach has the key advantage of high proteome coverage and dynamic range, as well as no restrictions on sample type84. The major strengths of the present study are as follows: (i) a relatively large sample and cohort size; (ii) a relatively high rate for which proteins identified as DEPs in this proteomic study were reproduced in the results of the ELISA (2/3, 66.6%), which suggests that the shotgun proteomics experiments were properly conducted; and (iii) identification of MIAC/IAI-specific protein biomarkers in a less-invasive, but still complex biological sample (i.e., plasma), which is often challenging owing to the wide dynamic range of proteins, and high levels of salts and other interfering compounds in such samples85.
The discrepancy concerning the results of LRP1 between the shotgun proteomics and ELISA data can be generally found in other proteomics-based biomarker discovery and verification studies19,62,64,86,87,88, but this disagreement may be attributed to (i) sample types evaluated (pooled vs. individual samples), (ii) somewhat loose threshold used in the present study as selection criteria for DEPs (fold change of LFQ intensities > 1.3 or < 0.77), and (iii) inherent disadvantages of the ELISA method, including cross-reactivity of the antibodies with non-target antigens, high-dose hook effect, and that the target proteins might be denatured during ELISA processing and thus their epitopes cannot be detected by the secondary antibodies89,90.
Conclusions
In summary, using proteomic approaches, 17 DEPs were identified as novel potential candidate biomarkers for MIAC/IAI in plasma samples collected from women with PPROM. In particular, plasma FCGR3A and haptoglobin were confirmed by ELISA as potential independent biomarkers for less-invasive identification of MIAC/IAI. Nevertheless, these biomarkers alone exhibited poor-to-fair diagnostic performance for PPROM-associated MIAC/IAI. The possible mechanistic roles of FCGR3A and haptoglobin in maternal circulation as pathophysiological links with inflammation/infection in the amniotic cavity warrant further investigation. Further studies are warranted to examine whether the alteration of these two proteins in other biological samples, such as cervicovaginal fluid, AF, saliva, or urine, could be used to identify MIAC/IAI risk.
Data availability
All relevant data are within the paper, and the authors can make available materials, data and associated protocols if requested.
References
Goldenberg, R. L., Culhane, J. F., Iams, J. D. & Romero, R. Epidemiology and causes of preterm birth. Lancet 371, 75–84 (2008).
Menon, R. & Richardson, L. S. Preterm prelabor rupture of the membranes: A disease of the fetal membranes. Semin. Perinatol. 41, 409–419 (2017).
Sae-Lin, P. & Wanitpongpan, P. Incidence and risk factors of preterm premature rupture of membranes in singleton pregnancies at Siriraj Hospital. J. Obstet. Gynaecol. Res. 45, 573–577 (2019).
Goncalves, L. F., Chaiworapongsa, T. & Romero, R. Intrauterine infection and prematurity. Ment. Retard. Dev. Disabil. Res. Rev. 8, 3–13 (2002).
Lee, S. M., Park, K. H., Jung, E. Y., Jang, J. A. & Yoo, H. N. Frequency and clinical significance of short cervix in patients with preterm premature rupture of membranes. PLoS ONE 12, e0174657 (2017).
Kacerovsky, M. et al. Antibiotic administration reduces the rate of intraamniotic inflammation in preterm prelabor rupture of the membranes. Am. J. Obstet. Gynecol. 223(114), e1-114.e20 (2020).
Romero, R., Chaiworapongsa, T. & Espinoza, J. Micronutrients and intrauterine infection, preterm birth and the fetal inflammatory response syndrome. J. Nutr. 133, 1668S-1673S (2003).
Shim, S. S. et al. Clinical significance of intra-amniotic inflammation in patients with preterm premature rupture of membranes. Am. J. Obstet. Gynecol. 191, 1339–1345 (2004).
Anblagan, D. et al. Association between preterm brain injury and exposure to chorioamnionitis during fetal life. Sci. Rep. 6, 37932 (2016).
Bierstone, D. et al. Association of histologic chorioamnionitis with perinatal brain injury and early childhood neurodevelopmental outcomes among preterm neonates. JAMA Pediatr. 172, 534–541 (2018).
Lee, J., Romero, R., Kim, S. M., Chaemsaithong, P. & Yoon, B. H. A new antibiotic regimen treats and prevents intra-amniotic inflammation/infection in patients with preterm PROM. J. Matern. Fetal. Neonatal. Med. 29, 2727–2737 (2016).
Lee, S. M. et al. Inflammatory proteins in maternal plasma, cervicovaginal and amniotic fluids as predictors of intra-amniotic infection in preterm premature rupture of membranes. PLoS ONE 13, e0200311 (2018).
Cobo, T. et al. Intra-amniotic inflammation predicts microbial invasion of the amniotic cavity but not spontaneous preterm delivery in preterm prelabor membrane rupture. Acta Obstet. Gynecol. Scand. 91, 930–935 (2012).
Calhoun, D. A. et al. Granulocyte colony-stimulating factor in preterm and term pregnancy, parturition, and intra-amniotic infection. Obstet. Gynecol. 97, 229–234 (2001).
Chow, S. S. et al. Differences in amniotic fluid and maternal serum cytokine levels in early midtrimester women without evidence of infection. Cytokine 44, 78–84 (2008).
Dowle, A. A., Wilson, J. & Thomas, J. R. Comparing the diagnostic classification accuracy of iTRAQ, peak-area, spectral-counting, and emPAI methods for relative quantification in expression proteomics. J. Proteome Res. 15, 3550–3562 (2016).
Park, J. M. et al. Predictive proteomic biomarkers for inflammatory bowel disease-associated cancer: Where are we now in the era of the next generation proteomics?. World J. Gastroenterol. 20, 13466–12476 (2014).
Vajrychova, M. et al. Comprehensive proteomic investigation of infectious and inflammatory changes in late preterm prelabour rupture of membranes. Sci. Rep. 10, 17696 (2020).
Hitti, J. et al. Noninvasive diagnosis of intraamniotic infection: Proteomic biomarkers in vaginal fluid. Am. J. Obstet. Gynecol. 203(32), e1-8 (2010).
Gravett, M. G. et al. Proteomic analysis of cervical-vaginal fluid: Identification of novel biomarkers for detection of intra-amniotic infection. J. Proteome Res. 6, 89–96 (2007).
Gravett, M. G. et al. Diagnosis of intra-amniotic infection by proteomic profiling and identification of novel biomarkers. JAMA 292, 462–469 (2004).
Lee, S. Y. et al. Intra-amniotic infection/inflammation as a risk factor for subsequent ruptured membranes after clinically indicated amniocentesis in preterm labor. J. Korean Med. Sci. 28, 1226–1232 (2013).
Park, K. H. et al. Noninvasive prediction of intra-amniotic infection and/or inflammation in preterm premature rupture of membranes. Reprod. Sci. 19, 658–665 (2012).
Chaemsaithong, P. et al. A point of care test for interleukin-6 in amniotic fluid in preterm prelabor rupture of membranes: A step toward the early treatment of acute intra-amniotic inflammation/infection. J. Matern. Fetal. Neonatal. Med. 29, 360–367 (2016).
Romero, R. et al. Sterile and microbial-associated intra-amniotic inflammation in preterm prelabor rupture of membranes. J. Matern. Fetal. Neonatal. Med. 28, 1394–1409 (2015).
Joo, E., Park, K. H., Kim, Y. M., Ahn, K. & Hong, S. Maternal plasma and amniotic fluid LBP, Pentraxin 3, Resistin, and IGFBP-3: Biomarkers of microbial invasion of amniotic cavity and/or intra-amniotic inflammation in women with preterm premature rupture of membranes. J. Korean Med. Sci. 36, e279 (2021).
Kim, H. J. et al. A protein microarray analysis of amniotic fluid proteins for the prediction of spontaneous preterm delivery in women with preterm premature rupture of membranes at 23 to 30 weeks of gestation. PLoS ONE 15, e0244720 (2020).
Jung, E. Y. et al. Relation between amniotic fluid infection or cytokine levels and hearing screen failure in infants at 32 wk gestation or less. Pediatr. Res. 81, 349–355 (2017).
Kim, C. J. et al. Acute chorioamnionitis and funisitis: Definition, pathologic features, and clinical significance. Am. J. Obstet. Gynecol. 213, S29-52 (2015).
Gibbs, R. S., Blanco, J. D., St Clair, P. J. & Castaneda, Y. S. Quantitative bacteriology of amniotic fluid from women with clinical intraamniotic infection at term. J. Infect. Dis. 145, 1–8 (1982).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Cox, J. et al. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Calvano, S. E. et al. A network-based analysis of systemic inflammation in humans. Nature 437, 1032–1037 (2005).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: A nonparametric approach. Biometrics 44, 837–845 (1988).
Buhimschi, I. A., Christner, R. & Buhimschi, C. S. Proteomic biomarker analysis of amniotic fluid for identification of intra-amniotic inflammation. BJOG 112, 173–181 (2005).
Ruetschi, U. et al. Proteomic analysis using protein chips to detect biomarkers in cervical and amniotic fluid in women with intra-amniotic inflammation. J. Proteome Res. 4, 2236–2242 (2005).
Romero, R. et al. Proteomic analysis of amniotic fluid to identify women with preterm labor and intra-amniotic inflammation/infection: The use of a novel computational method to analyze mass spectrometric profiling. J. Matern. Fetal. Neonatal. Med. 21, 367–388 (2008).
Ravetch, J. V. & Kinet, J. P. Fc receptors. Annu. Rev. Immunol. 9, 457–492 (1991).
Mancardi, D. & Daëron, M. Fc Receptors in Immune Responses. Reference Module in Biomedical Sciences B978-0-12-801238-3.00119-7 (2014).
Atyeo, C. et al. Compromised SARS-CoV-2-specific placental antibody transfer. Cell 184, 628–642 (2021).
Wang, Y. & Jonsson, F. Expression, role, and regulation of neutrophil fcgamma receptors. Front Immunol. 10, 1958 (2019).
Ozturk, C., Aksu, G., Berdeli, A. & Kutukculer, N. Fc gamma RIIa, IIIa and IIIb polymorphisms in Turkish children susceptible to recurrent infectious diseases. Clin. Exp. Med. 6, 27–32 (2006).
Ye, D. et al. A novel single-nucleotide polymorphism of the Fcgamma receptor IIIa gene is associated with genetic susceptibility to systemic lupus erythematosus in Chinese populations: A family-based association study. Clin. Exp. Dermatol. 31, 553–557 (2006).
Foster, C. B. et al. Polymorphisms in inflammatory cytokines and Fcgamma receptors in childhood chronic immune thrombocytopenic purpura: A pilot study. Br. J. Haematol. 113, 596–599 (2001).
Presicce, P. et al. IL-1 signaling mediates intrauterine inflammation and chorio-decidua neutrophil recruitment and activation. JCI Insight 3, e98306 (2018).
Negishi, Y., Shima, Y., Takeshita, T. & Takahashi, H. Distribution of invariant natural killer T cells and dendritic cells in late pre-term birth without acute chorioamnionitis. Am. J. Reprod. Immunol. 77, e12658 (2017).
Palmsten, K. et al. Subclinical and clinical chorioamnionitis, fetal vasculitis, and risk for preterm birth: A cohort study. Placenta 67, 54–60 (2018).
di Masi, A. et al. Haptoglobin: From hemoglobin scavenging to human health. Mol. Aspects Med. 73, 100851 (2020).
Hanthorn, C. J. et al. Serum concentrations of haptoglobin and haptoglobin-matrix metalloproteinase 9 (Hp-MMP 9) complexes of bovine calves in a bacterial respiratory challenge model. BMC Vet. Res. 10, 285 (2014).
Buhimschi, C. S. et al. Proteomics mapping of cord blood identifies haptoglobin “switch-on” pattern as biomarker of early-onset neonatal sepsis in preterm newborns. PLoS ONE 6, e26111 (2011).
McCarthy, M. E. et al. Identification of haptoglobin switch-on status in archived placental specimens indicates antenatal exposure to inflammation and potential participation of the fetus in triggering preterm birth. Placenta 62, 50–57 (2018).
Oggé, G. et al. Chronic chorioamnionitis displays distinct alterations of the amniotic fluid proteome. J. Pathol. 223, 553–565 (2011).
Dulay, A. T. et al. Compartmentalization of acute phase reactants Interleukin-6, C-Reactive Protein and Procalcitonin as biomarkers of intra-amniotic infection and chorioamnionitis. Cytokine 76, 236–243 (2015).
Holmstrom, E. et al. Cervical and amniotic fluid matrix metalloproteinase-8 and interleukin-6 concentrations in preterm pregnancies with or without preterm premature rupture of membranes. Fetal. Diagn. Ther. 46, 103–1107 (2019).
Kacerovsky, M. et al. Cervical fluid IL-6 and IL-8 levels in pregnancies complicated by preterm prelabor rupture of membranes. J. Matern. Fetal. Neonatal. Med. 28, 134–140 (2015).
Hong, S. et al. A protein microarray analysis of plasma proteins for the prediction of spontaneous preterm delivery in women with preterm labor. Reprod. Sci. 27, 1187–1196 (2020).
Pereira, L. et al. Insights into the multifactorial nature of preterm birth: Proteomic profiling of the maternal serum glycoproteome and maternal serum peptidome among women in preterm labor. Am. J. Obstet. Gynecol. 202(555), e1-10 (2010).
Park, H. et al. Plasma inflammatory and immune proteins as predictors of intra-amniotic infection and spontaneous preterm delivery in women with preterm labor: A retrospective study. BMC Preg. Childbirth 18, 146 (2018).
Lee, J. E. et al. Proteomic identification of novel plasma biomarkers associated with spontaneous preterm birth in women with preterm labor without infection/inflammation. PLoS ONE 16, e0259265 (2021).
Lynch, A. M. et al. The relationship of circulating proteins in early pregnancy with preterm birth. Am. J. Obstet. Gynecol. 214, 517 (2016).
Hong, S. et al. Identifying potential biomarkers related to pre-term delivery by proteomic analysis of amniotic fluid. Sci. Rep. 10, 19648 (2020).
Capece, A., Vasieva, O., Meher, S., Alfirevic, Z. & Alfirevic, A. Pathway analysis of genetic factors associated with spontaneous preterm birth and pre-labor preterm rupture of membranes. PLoS ONE 9, e108578 (2014).
Baumann, H. & Gauldie, J. The acute phase response. Immunol. Today 15, 74–80 (1994).
Gabay, C. & Kushner, I. Acute-phase proteins and other systemic responses to inflammation. N. Engl. J. Med. 340, 448–454 (1999).
Venteclef, N., Jakobsson, T., Steffensen, K. R. & Treuter, E. Metabolic nuclear receptor signaling and the inflammatory acute phase response. Trends Endocrinol. Metab. 22, 333–343 (2011).
de Leeuw, A. J. M., Oude Luttikhuis, M. A. M., Wellen, A. C., Muller, C. & Calkhoven, C. F. Obesity and its impact on COVID-19. J. Mol. Med. (Berl) 99, 899–915 (2021).
Bodart, G. et al. The somatotrope growth hormone-releasing hormone/growth hormone/insulin-like growth factor-1 axis in immunoregulation and immunosenescence. Front Horm. Res. 48, 147–159 (2017).
Serri, O., St-Jacques, P., Sartippour, M. & Renier, G. Alterations of monocyte function in patients with growth hormone (GH) deficiency: Effect of substitutive GH therapy. J. Clin. Endocrinol. Metab. 84, 58–63 (1999).
Ross, A. C. Impact of chronic and acute inflammation on extra- and intracellular iron homeostasis. Am. J. Clin. Nutr. 106, 1581S-1587S (2017).
Plutzky, J. The PPAR-RXR transcriptional complex in the vasculature: Energy in the balance. Circ. Res. 108, 1002–1016 (2011).
Balamurugan, K. HIF-1 at the crossroads of hypoxia, inflammation, and cancer. Int. J. Cancer 138, 1058–1066 (2016).
Dehne, N., Fuhrmann, D. & Brune, B. Hypoxia-inducible factor (HIF) in hormone signaling during health and disease. Cardiovasc. Hematol. Agents Med. Chem. 11, 125–135 (2013).
Molinari, N. et al. Sample pooling and inflammation linked to the false selection of biomarkers for neurodegenerative diseases in top-down proteomics: A pilot study. Front Mol. Neurosci. 11, 477 (2018).
Martins-de-Souza, D. et al. Alterations in oligodendrocyte proteins, calcium homeostasis and new potential markers in schizophrenia anterior temporal lobe are revealed by shotgun proteome analysis. J. Neural Transm. (Vienna) 116, 275–289 (2009).
Kusonmano, K. et al. Effects of pooling samples on the performance of classification algorithms: A comparative study. ScientificWorldJournal 2012, 278352 (2012).
Telaar, A., Nurnberg, G. & Repsilber, D. Finding biomarker signatures in pooled sample designs: A simulation framework for methodological comparisons. Adv. Bioinform. 2010, 318573 (2010).
Combs, C. A. et al. Amniotic fluid infection, inflammation, and colonization in preterm labor with intact membranes. Am. J. Obstet Gynecol. 210(125), e121-125.e115 (2014).
Mitchell, B. L., Yasui, Y., Li, C. I., Fitzpatrick, A. L. & Lampe, P. D. Impact of freeze-thaw cycles and storage time on plasma samples used in mass spectrometry based biomarker discovery projects. Cancer Inform. 1, 98–104 (2005).
Keustermans, G. C., Hoeks, S. B., Meerding, J. M., Prakken, B. J. & de Jager, W. Cytokine assays: An assessment of the preparation and treatment of blood and tissue samples. Methods 61, 10–17 (2013).
Spong, C. Y., Ghidini, A., Ossandon, M., Walker, C. N. & Pezzullo, J. C. Are the cytokines interleukin-6 and angiogenin stable in frozen amniotic fluid?. Am. J. Obstet. Gynecol. 178, 783–786 (1998).
Megger, D. et al. Comparison of label-free and label-based strategies for proteome analysis of hepatoma cell lines. Biochim. Biophys. Acta. 1844, 967–976 (2013).
Ray, S. et al. Proteomic technologies for the identification of disease biomarkers in serum: Advances and challenges ahead. Proteomics 11, 2139–2161 (2011).
Parry, S. et al. Maternal serum serpin B7 is associated with early spontaneous preterm birth. Am. J. Obstet. Gynecol. 211(678), e671–e612 (2014).
Parry, S. et al. Cervicovaginal fluid proteomic analysis to identify potential biomarkers for preterm birth. Am. J. Obstet. Gynecol. 222, 493 (2020).
Lee, J. et al. Proteomic analysis of amniotic fluid proteins for predicting the outcome of emergency cerclage in women with cervical insufficiency. Reprod. Sci. 27, 1318–1329 (2020).
Kessler, E., Matthes, H. F., Schein, E. & Wendt, M. Detection of antibodies in sera of weaned pigs after contact infection with Sarcoptes scabiei var. suis and after treatment with an antiparasitic agent by three different indirect ELISAs. Vet. Parasitol. 114, 63–73 (2003).
Hoofnagle, A. N. & Wener, M. H. The fundamental flaws of immunoassays and potential solutions using tandem mass spectrometry. J. Immunol. Methods 347, 3–11 (2009).
Acknowledgements
This research was supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute funded by the Ministry of Health & Welfare (HI20C0013), and by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT; No. 2020R1F1A1048362). The funders had no any role in the design of the study, data collection, analysis, and interpretation of data and in writing the manuscript.
Author information
Authors and Affiliations
Contributions
B.J.H., L.J.E., P.K.H. Design of the study; K.S.Y., G.M.B., K.H.J., L.J.E., P.K.H. Conduct of the study; K.S.Y., K.H.J., L.K.N., P.K.H. Collection and management of data; B.J.H., G.M.B., L.K.N., L.J.E., P.K.H. Analysis and interpretation of data; B.J.H., K.S.Y., K.H.J., L.J.E., P.K.H. Preparation of manuscript; B.J.H., K.S.Y., G.M.B., K.H.J., L.K.N., L.J.E., P.K.H. Review or approval of manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Back, J.H., Kim, S.Y., Gu, M.B. et al. Proteomic analysis of plasma to identify novel biomarkers for intra-amniotic infection and/or inflammation in preterm premature rupture of membranes. Sci Rep 13, 5658 (2023). https://doi.org/10.1038/s41598-023-32884-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-32884-y
- Springer Nature Limited