Abstract
Bacterial Chondronecrosis with Osteomyelitis (BCO) is a specific cause of lameness in commercial fast-growing broiler (meat-type) chickens and represents significant economic, health, and wellbeing burdens. However, the molecular mechanisms underlying the pathogenesis remain poorly understood. This study represents the first comprehensive characterization of the proximal tibia proteome from healthy and BCO chickens. Among a total of 547 proteins identified, 222 were differentially expressed (DE) with 158 up- and 64 down-regulated proteins in tibia of BCO vs. normal chickens. Biological function analysis using Ingenuity Pathways showed that the DE proteins were associated with a variety of diseases including cell death, organismal injury, skeletal and muscular disorder, immunological and inflammatory diseases. Canonical pathway and protein–protein interaction network analysis indicated that these DE proteins were involved in stress response, unfolded protein response, ribosomal protein dysfunction, and actin cytoskeleton signaling. Further, we identified proteins involved in bone resorption (osteoclast-stimulating factor 1, OSFT1) and bone structural integrity (collagen alpha-2 (I) chain, COL2A1), as potential key proteins involved in bone attrition. These results provide new insights by identifying key protein candidates involved in BCO and will have significant impact in understanding BCO pathogenesis.
Similar content being viewed by others
Introduction
Bacterial Chondronecrosis with Osteomyelitis (BCO) is the leading cause of lameness affecting tens of billions of broiler chickens worldwide1,2. Addition to the food safety issue as well as animal health and welfare concerns, BCO causes heavy annual economic losses to the poultry industries2. While BCO likely arises from a complex interaction of metabolic, functional, genetic, nutritional, and environmental factors, there is a clear consensus and strong evidence of bacterial colonization and infection of the proximal femora and tibiae3,4.
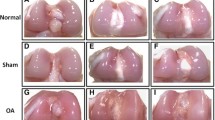
Multiple opportunistic organisms have been reported from BCO lesions, including Staphylococcus aureus, Staphylococcus agnetis, E. coli, and Enterococcus cecorum, often in mixed cultures with other bacteria3,5,6. Bacteria transmitted to chicks from breeder parents, contaminated eggshells or hatchery sources6,7, or that enter the chick's circulation via translocation through the integument, respiratory system or gastrointestinal tract8 spread hematogenously and exit the bloodstream through the capillaries supplying the growth plate. The translocated bacteria bind to the bone collagen and adhere directly to the cartilage matrix, colonize osteochondrotic clefts and form obstructive emboli in the metaphyseal vasculature9. Lytic substances released at sites of bacterial colonization promote generalized necrosis within the calcifying zone of the metaphysis, destroying the vasculature and eliminating struts of trabecular bone that normally provide structural support to prevent micro-fracturing of the epiphyseal and physeal cartilage10,11. Terminal BCO presents as necrotic degeneration and bacterial infection primarily within the proximal head of the femora and tibiae (Fig. 1), as well as in the growth plates of the 4th thoracic vertebrae.
Stages of proximal femoral degeneration (A–D) and proximal tibial degeneration (E–H) leading progressively to bacterial chondronecrosis with osteomyelitis: (A) Normal proximal femur with white cap of epiphyseal cartilage (e); (B) Femoral head separation (FHS: epiphyseolysis) with the epiphysis remaining in the socket when the femur was disarticulated, revealing the underlying surface of the growth plate or physis (p) and an early region of necrosis (n); (C) Fracturing of the growth plate (p) revealing a necrotic void (nv) within the metaphysis; (D) Terminal femoral head necrosis in which the femoral epiphysis, physis and most of the metaphysis remained attached to the acetabulum when the diaphysis weakened by widely dispersed necrosis was fractured during disarticulation, revealing copious fibrinonecrotic exudate (fe); (E) Normal proximal tibia showing the epiphysis (e) with a secondary center of ossification (*) and the physis/growth plate (p) fully supported by struts of trabecular bone in the metaphyseal zone (m); (F–H) Bacterial infiltration and sequestrae (s), necrotic voids (nv) and microfractures below the growth plate (arrows) provide macroscopic evidence of bone damage associated with osteomyelitis.
Despite seminal nutritional and managerial efforts to reduce BCO incidence in chickens, insufficient progress has been made due to limited understanding of the molecular mechanisms underlying BCO pathogenesis. We, therefore, undertook this study using high-throughput proteomics approach combined with advanced bioinformatics to identify molecular signatures of BCO disorder.
Materials and methods
Ethics statement
The present study was conducted in accordance with the recommendations in the guide for the care and use of laboratory animals of the National Institutes of Health. All procedures of animal care compiled with and were approved by the University of Arkansas Animal Care and Use Committee (IACUC) under protocol No 15043. All reported methods are in accordance with the Animal Research Reporting of in Vivo Experiments (ARRIVE).
Animals and samples preparation
The experiment was conducted at the Poultry Environmental Research Laboratory at the University of Arkansas Poultry Research Farm. One day-old male boiler chicks were obtained from a local commercial hatchery (Cobb-Vantress, Siloam Springs, AR) and randomly divided into two body weight-matched groups. The control group was reared on clean shaved wood litters at 50 birds/pen (6 pens/group, 300 birds/group) and the BCO group was maintained on wire floor model developed by Wideman Robert10. Pen conformation, bird densities, diet and water (ad libitum), light/dark cycle, and heating conditions were as previously described10,12. To minimize distress, birds were walked on a daily basis from day 15 to the end of the experiment (day 56). On day 56, birds were humanely euthanized, necropsied, and the right and left tibia were macroscopically scored for tibial head necrosis (THN) severity. Briefly, as described previously13, THN lesion severity was scored on a 0- to 3-scale with the following categories: 0- no abnormalities (Normal); 1- mild necrosis (THN); 2- severe tibial head necrosis (THNS); and 3- caseous THN (THNC).
Tibia samples and protein extraction
Chicken tibia samples were collected from healthy (score 0) and BCO (score 3) chickens (n = 6/group), snap frozen in liquid nitrogen and stored at −80 °C until use. Tibia proteins were extracted as previously described14. Briefly, samples were homogenized in lysis buffer (10 mM Tris base, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1 mM EGTA, 0.1% Triton X-100, 0.5% NP-40, protease, and phosphatase inhibitors), and proteins (100 μg) were run on 4–12% Novex Bis–Tris gels (ThermoFisher Scientific, Waltham, MA). The gel was then stained with Coomassie blue dye, and de-stained until the background was clear. Gel portions of each sample were excised and chopped into small pieces (< 1 mm2) and washed twice with 25 mM NH4HCO3 (Sigma Aldrich, St. Louis, MO). The gel pieces were de-stained with 25 mM NH4HCO3/50% acetonitrile (ACN), and dried with 100% ACN. Proteins were then reduced using 10 mM dithiothreitol (DTT, Sigma Aldrich, St. Louis, MO) in 25 mM NH4CO3 at 56° for 1 h. Subsequently, alkylation was conducted using 55 mM iodoacetamide (IAA, Sigma Aldrich, St. Louis, MO) in 25 mM NH4CO3, protected from light. The gel pieces were then washed with 25 mM NH4HCO3, dehydrated with 25 mM NH4HCO3/50% ACN, and completely dried via SpeedVac. Mass spectrometry grade Trypsin Gold (12.5 ng/μl in 25 mM NH4HCO3, Promega, Madison, WI) was added to cover dried gels, and incubated overnight at 37 °C. Peptides were extracted by 50% ACN/5% formic acid. The tryptic digests were desalted using Pierce C18 spin columns (150 × 0.3 mm, 3.5 µm particle size, 300 Å pore size, ThermoFisher Scientific, Waltham, MA) prior to LC–MS/MS at the State Wide Mass Spectrometry Facility, University of Arkansas at Fayetteville, Arkansas.
Shotgun proteomics
Individual extracted proteins were used in shotgun proteomics analysis with in-gel trypsin digestion followed by Liquid Chromatography with Tandem Mass Spectrometry (LC–MS/MS) conducted at the State Wide Mass Spectrometry Facility, University of Arkansas at Fayetteville. All LC–MS/MS samples were analyzed using Mascot ((Matrix Science, London, UK; version 2.2.1). Mascot was set up to search the UniProt_Gallus database assuming the digestion enzyme trypsin. Scaffold (version Sacffold_4.8.3, Proteome Software Inc., Portland, OR) was used to validate MS/MS based peptide and protein identifications. Peptide identifications were accepted if they could be established at greater than 95.0% probability by the Scaffold Local FDR algorithm. Protein identifications were accepted if they could be established at > 95.0% probability and contained at least 2 identified peptides. Protein probabilities were assigned by the Protein Prophet algorithm15,16. Proteins were annotated with the Gene Ontology Consortium (GO) terms from NCBI16. For quantitative changes, we set a P-value (t-test) < 0.05, set normal as the reference category for fold change and performed total spectra normalization (minimum value 0). Protein–protein interaction analysis was performance by String v10.8 (http://www.string-db.org/) at a 0.4 medium confidence value17.
Bioinformatics and ingenuity pathway analysis
Fold changes of the identified proteins were calculated by comparing the BCO conditions to healthy birds (control) and differentially expressed (DE) proteins (fold change and P-value < 0.05) as well as their ID (UniProt18) were submitted to QIAGEN Ingenuity Pathway Analysis (IPA, Ingenuity Systems, Redwood City, CA) for core analysis19. These proteins were mapped to the most significant canonical pathways, functional annotation, upstream regulators, as well as molecular discovery using the Ingenuity Knowledge Base as a reference set and a cut-off of FDR adjusted P-value < 0.05 and a fold-change between -1.5 and 1.5. Right-tailed Fisher’s exact test was used to determine the probability that biological functions and/or diseases were over-represented in the protein dataset. IPA also predicted potential upstream regulators and downstream effectors of the proteins in this study, which were assigned as inhibited or activated according to Z-score20.
Western blot analysis
Proteins were extracted from proximal tibiae tissues as described above and the concentrations were determined using Synergy HT multi-mode microplate reader (BioTek, Winooski, VT) and a Bradford assay kit (Bio-Rad, Hercules, CA) with bovine serum albumin as a standard. Western blot was performed as previously described14,21. Briefly, total proteins (100 μg) were resolved on 4–12% Novex Bis–Tris gels (Life Technologies, grand Island, NY), and transferred to a polyvinylidene difluoride (PVDF) membrane. The transferred membranes were blocked in Tris-buffered saline (TBS) with 5% non-fat milk and Tween 20 (TBST) for 1 h at room temperature, and incubated overnight at 4 °C with primary antibodies diluted to 1:500–1:1000. The rabbit polyclonal antibodies used were as follows: anti-HSP90 (cat#PA5-17,610, ThermoFisher Scientific Waltham, MA), anti-OSTF1 (cat#A303-004A, Bethyl Laboratories, Inc, Waltham, MA), anti-ACLY (cat# LS-C290517, Lifespan Biosciences, Seattle, WA), anti-vinculin (VCL, cat#V4139, Sigma-Aldrich, Inc, St. Louis, MO), anti-STAT3 (cat# 4904, Cell Signaling Technology, Danvers, MA), anti-ACTN (cat#A1160, ABClonal Technology, Woburn, MA), and anti-β actin (cat#4967, Cell Signaling Technology, Danvers, MA). The membrane was washed twice with PBS-T and then incubated with anti-mouse or anti-rabbit horseradish peroxidase conjugated secondary antibodies (1:5000) for 1 h at room temperature. The signal was visualized by enhanced chemiluminescence (ECL plus) (GE Healthcare Bio-Sciences, Buckinghamshire, UK) and captured by FluorChem M MultiFluor System (Proteinsimple, Santa Clara, CA). Image Acquisition and Analysis were performed by AlphaView software (Version 3.4.0, 1993–2011, Proteinsimple, Santa Clara, CA).
Total RNA extraction and real-time quantitative PCR
Total RNA was extracted using Trizol reagent (Life Technologies, Carlsbad, CA) according to the manufacturer’s recommendations, and concentration and quality were determined with the Take3 microvolume plate and the Synergy HT multimode microplate reader (BioTek, Winooski, VT). RT and qPCR were performed as previously described22. Briefly, RNA was reverse transcribed using qScript cDNA Synthesis Supermix (Quanta Biosciences, Gaithersburg, MD), and amplified by qPCR (Applied Biosystems 7500 Real Time System) with Power-Up Sybr green master mix (Life Technologies, Carlsbad, CA). Relative expression of the target genes was determined using the 2-ΔΔCT method23, with normalization to ribosomal 18S expression and healthy bird as a calibrator (control). Oligonucleotide primer sequences specific for chicken are presented in Table 1.
Statistical analysis
Data were analyzed using Student “t” test and Graph Pad Prism software (version 7.03 for Windows, Graph Pad Software, La Jolla California, USA). Data are expressed as the mean ± SEM. Means were considered statistically significant at a P value ≤ 0.05.
Results
Protein expression profile in proximal tibiae of healthy- and BCO-affected broilers
To gain further insights into the molecular pathogenesis of BCO, we performed LC–MS/MS on protein isolated from the proximal end of the tibia from control (score 0) and BCO birds with severe THN (score 3). MASCOT and Scaffold analysis identified a total of 547 proteins that have been submitted to EMBL-EBI PRIDE database (https://doi.org/10.6019/PXD029085, accession# PXD029085PXD029085, reference #1-20211011-22240). Quantitative analysis identified 222 differentially expressed (DE) proteins (T-test, P \(<\) 0.05). Of those, there were 158 up- and 64 down-regulated proteins in tibia of BCO vs. normal chickens (Tables 2 and 3).
Metabolic pathway and network analysis
To gain biologically related molecular networks, the above identified DE proteins (222) were submitted into IPA knowledge-base. From these 222 proteins, 153 IDs were successfully mapped, while 69 IDs were unmapped to molecules in the Ingenuity Knowledge Base. The 153 mapped proteins were analyzed to outline the most enriched biological functions.
As shown in Table 4, the top five canonical pathways associated with Huntington’s disease, inhibition of ARE-mediated mRNA degradation, protein ubiquitination, hepatic fibrosis signaling, and glycolysis were enriched in BCO THN.
The top diseases and disorders enriched by the IPA core analysis for the DE proteins were ranked by P-value and summarized in Table 5. Cancer and organismal injury and abnormalities (including chronic bone and joint disease) were the top disorders enriched with 32 and 46 proteins, respectively (P = 4.83 × 10–2–3.05 × 10–3). Next, cardiovascular disease was embellished by IPA analysis and ranked the third with 15 proteins (P = 4.44 × 10–2-6.65 × 10–3), followed by connective tissue disorders and inflammatory disease with 10 molecules each (P = 0.01).
The top five molecular and cellular functions generated by IPA core analysis and rated by P- values are cellular development (P = 4.4 × 10–2–1.7 × 10–2, 17 molecules), cellular growth and proliferation (P = 4.4 × 10–2–1.7 × 10–2, 16 molecules), cell cycle (P = 3.2 × 10–2–3.2 × 10–2, 10 molecules), gene and protein expression (P = 4.8 × 10–2–3.2 × 10–2, 17 molecules), and cell-to-cell signaling and interaction (P = 4.4 × 10–2, 5 molecules) (Table 6).
The top physiological system development and functions enriched by IPA are summarized in Table 7 and are composed of organismal survival (P = 0.03, 26 molecules), connective tissue development and function (P = 0.04, 5 proteins), hematological system development and function (P = 0.04, 9 molecules), hematopoiesis (P = 0.04, 5 molecules), and immune cell trafficking (P = 0.04, 5 molecules).
Figure 2 provides a visual summary of the three IPA-predicted upstream regulators. The top predicted activated upstream transcription regulator in the BCO tibiae was the myelocytomatosis oncogene (MYC) with a computed activation z-score of 2.121, overlap P-value of 3.09 × 10–2, and with 15 of 23 proteins have measurement direction consistent with MYC activation. However, the progesterone receptor (PR) was significantly enriched by IPA in the THN and was predicted as an inhibitor upstream regulator (z-score −2.0, overlap P-value 4.44 × 10–2), with 4 of 5 proteins have measurement direction consistent with PR inhibition. The nuclear factor erythroid 2 like 1 (NRF1 or NFE2L1) was also predicted by IPA to be an upstream regulator (Overlap P-value 4.44 × 10–2), however its activation state was not determined. Validation studies on protein expression profile are more challenging as they rely on availability of antibodies that cross-react with chickens. Here, by using immunoblot, we were able to validate the upregulation of the osteoblast stimulating factor 1 (OSTF1, 4.52-fold change, P = 0.0002), heat-shock protein 90 (HSP90, 1.30-fold change, P < 0.0001), ATP citrate lyase (ACLY, 1.42-fold change, P < 0.0001), and vinculin (VCL, 1.44-fold change, P = 0.0005), and the down regulation of STAT3 (0.58-fold change, P < 0.0001) and ACTN4 (0.68-fold change, P < 0.05, Fig. 3), in BCO-affected compared to healthy-birds, which is in agreement with the LC–MS/MS data. As most of the DE markers are protein-encoding genes, we next determined the mRNA abundances of few selected genes. Real-time qPCR analysis confirmed the up regulation of HSP90 and the down regulation of Col2A1, ACTN4, NCAM1, and PR. Whilst qPCR determined the regulation direction (down regulation) of NRF1, it showed contrary to IPA prediction a down regulation of MYC gene expression (Figs. 2 and 3).
Interconnected proteins and predicted upstream regulators built with IPA program for DE protein data that was determined on BCO-affected tibiae. IPA analysis predicted MYC, PR (PGR), and NRF1 (or NFE2L1) as upstream regulators, which were assigned as inhibited or activated according to Z-score. ACTN4, α-actinin 4; ANXA5, annexin A5; APEX1, Apurinic/Apyrimidinic Endodeoxyribonuclease 1; Col2A1; Collagen Type II Alpha 1 Chain ; GCLM; Glutamate-Cysteine Ligase Modifier Subunit ; GLS; Glutaminase; HMOX1; Heme Oxygenase 1; HNRNPA1; Heterogeneous Nuclear Ribonucleoprotein A1; IPO7; Importin 7; LAMP1; Lysosomal Associated Membrane Protein 1; LUM; Lumican; MYC; Myelocytomatosis Oncogene; NFE2L1; NFE2 Like BZIP Transcription Factor 1; PCNA; Proliferating Cell Nuclear Antigen; PGAM1; Phosphoglycerate Mutase 1; PGR (PR); progesterone receptor; PKM; Pyruvate Kinase M1/2; PLS3; plastin 3; PSMA; Proteasome 20S Subunit Alpha 6; PSMD; Proteasome 26S Subunit, Non-ATPase 1; RPL; Ribosomal Protein L6; RPS; ribosomal protein S3A1; STAT; Signal Transducer And Activator Of Transcription; TF; Transferrin; VDAC2, voltage-dependent anion channel 2.
Validation of selected protein-encoding genes. Protein expression was determined by Western blot (a, b) and mRNA abundances were measured by qPCR and 2-ΔΔCT method23. Data are mean ± SEM and * indicates a significant difference at P < 0.05. ACLY, ATP citrate lyase; ACTB, β actin; ACTN4, α-actinin 4; Col2A1, Collagen Type II Alpha 1 Chain; HSP90, heat shock protein 90; MYC, myelocytomatosis oncogene; NCAM1, Neural Cell Adhesion Molecule 1; NRF1/2, erythroid 2-related factor 1/2; OSTF1, Osteoclast-stimulating factor-1; STAT3, Signal Transducer And Activator Of Transcription 3; VCL, vinculin.
Discussion
Bacterial chondronecrosis with osteomyelitis is a significant health, welfare, and economic concern in the commercial broiler industry2, yet its underlying molecular mechanisms are not fully defined. In the present study, we used the well-established wire-flooring model that is reliable and reproducibly triggers a high incidence of BCO lesions that are similar to BCO-associated lesions observed in commercial flocks10. To gain large-scale in-depth knowledge, we used high throughput LC–MS/MS screening and comparative proteomics analysis, which provides a powerful tool to detect differentially or uniquely expressed proteins and their dynamic changes in a particular condition24,25. Previous studies have described differential plasma proteomic profile between healthy and spontaneously- or glucocorticoid-induced femoral head necrosis-affected birds26,27, and reported potential systemic (serum) biomarkers. In the current study, which constitutes the first to the best of our knowledge, we identified 222 locally (tibial) differentially expressed (DE) proteins, with 158 up- and 64 down-regulated proteins, in tibia of BCO compared to healthy chickens28.
The DE proteins were mapped to the reference pathway in IPA knowledge database (QIAGEN Inc., https://digitalinsights.qiagen.com/IPA)19 to derive biological insights and identify significantly enriched metabolic and/or signal transduction pathways. IPA identified significant top canonical pathways, including HD disease, ARE-mediated mRNA degradation, protein ubiquitination, hepatic fibrosis signaling, and glycolysis pathways, with several common and overlapping proteins. Based on the IPA core analysis, most of these proteins are involved in several disorders such as bone injury and abnormality, connective tissue disorders and inflammatory diseases. The top physiological functions depicted by IPA include organismal survival, connective tissue development, and immune cell trafficking, which are not surprising.
For instance, among these proteins, proteasomes (PSMA, PSMD, PSME), a multicatalytic proteinase complex, which are ubiquitously distributed in eukaryotic cells to cleave peptides in an ATP/ubiquitin-dependent (non-lysosomal) process29, are found in the first three top canonical pathways. They have been shown to be expressed in bone and involved in bone remodeling, resorption, and formation30,31,32. Moreover, infection of human endothelial (HUVEC) cells with Staphylococcus aureus (also the leading cause of BCO)3 has been shown to induce proteasome subunits at 16 h post-infection33,34. Immune and pro-inflammatory pathways have been found to be dependent on both proteasomal activity and ubiquitylation35, and our previous studies have shown that pro-inflammatory cytokines, such as IL-1β and TNFα, were induced in BCO-affected chickens at local and systemic levels36. This proteasome-induced inflammation is probably mediated through NF-κB and MAPK pathways that merit future investigations37. Additionally, proteasome inhibition by NIC-0102 has been reported to specifically prevent NLRP3 inflammasome activation, which has been shown to be induced in bone of BCO-affected chickens and in Staphylococcus-infected osteoblasts21.
The second group of biomarkers that proteomics analysis found with increased expression in the BCO group are molecular chaperones and stress proteins, including heat shock protein 90 (HSP 90), which also has been confirmed by qPCR and immunoblot analyses. Besides their classical roles as molecular chaperones and housekeepers (folding/unfolding, assembly/disassembly), HSPs are now understood to play a pivotal role in many cellular processes including transport and trafficking, protein degradation, and cell signaling38. Depending on the physiological context, these HSPs can be immune-stimulatory or immunosuppressive39,40. It has been shown that, after trauma or exposure to bacteria, cells express high levels of HSPs41, which in turn lead to cytokine transcription and release42,43. Indeed, Takahashi et al.44 have reported high levels of HSP70 and cytokines in articular cartilage and suggested a key role for HSPs in early stage of osteoarthritis in both rodents and humans. Moreover, recent studies proposed a role for HSP90 in chondrocyte biology and cartilage breakdown45,46. Tribelli et al.47, on the other hand, demonstrated that Staphylococcus aureus infection triggers human host primary keratinocyte and HaCaT cell line invasion through HSP90. Consistent with these findings and as mentioned above, the mRNA abundances of pro-inflammatory cytokine IL-1β and TNFα were also higher in the tibiae and femur of BCO compared to the healthy group. As a number of reports have shown that cytokines can stimulate HSP expression48, the cause-effect relationship between cytokines and HSP90 in BCO pathogenesis merit further in depth investigations.
The data from our proteomics, immunoblot and qPCR analyses agreed in denoting high levels of osteoclast-stimulating factor 1 (OSTF1), a protein known to activate osteoclasts and modulate trabecular bone remodeling49. OSTF1 was first described as an intracellular SH3-domain containing protein produced by osteoclasts that indirectly induces osteoclast formation and bone resorption49. Under normal physiological conditions, bone mass and structure homeostasis are maintained by constant bone remodeling with balanced bone formation by osteoblasts and bone resorption by osteoclasts50,51. Interestingly, Wideman group has previously found, in BCO model, a high osteoclastic activity52, which dissolve bone mineral by massive acid secretion and production of specialized proteinases that degrade the organic matrix, mainly type I collagen and erode the trabecular bone53. The increased levels of cathepsins here, which are responsible for the degradation of type I collagen in osteoclast-mediated bone resorption reinforce the aforesaid data54. Vermeren and co-workers55 reported that OSTF1 knockout mice suffer from a mild form of osteopetrosis, which caused by an increase in trabecular bone. In support of the abovementioned results, we reported here a reduced expression of Col2A1, at mRNA and protein levels, in BCO compared to control birds, which confirm a degradation status of collagen matrix and excessive bone resorption caused by exaggerated osteoclast activity-induced OSTF1 overexpression. Heretofore, molecular defects in the Col2A1 gene has been found to lead to low bone mass, bone deformity and fragility, and increased fracture incidence56, and thereby resulted in skeletal disorders such as skeletal dysplasia, achondrogenesis, stickler syndrome, and osteoarthritis57.
Parallel to Col2A1 down regulation, several other proteins (n = 20 from IPA top analysis-ready molecules) were found to be decreased (−2.3 < Expr Log Ratio) or increased (Expr Log Ratio > 2.4). Of particular interest, ovotransferrin (OTF), an 82-kDa glycoprotein and a member of transferrin family58, was significantly decreased in BCO. OTF and its receptor were found to be expressed in chicken bone and play key roles in bone formation59. In addition, mechanistically disruption of transferrin system altered iron uptake, heme biosynthesis, and bone homeostasis through glycolysis- and mitochondrial oxidative phosphorylation-dysmetabolisms60, both of which were delineated by proteomics and IPA analysis. In line with this, the ATPase H+ transporting V1 subunits (ATP6V1D/ATP6V1E1) that play critical roles in iron homeostasis and ATP synthesis were dysregulated. Similarly, the mitochondrial voltage dependent anion channels (VDAC2), glutaminase (GLS), and plastin 3 (also known as fimbrin, PLS3) expressions were dysregulated, indicating a mitochondrial dysfunction in bone of BCO birds, which has been previously reported by our group61. VDAC2 has been reported to play key roles in ADP-dependent mitochondrial bioenergetics62. PLS3, a member of actin-binding and bundling protein family, plays a pivotal role in actin cytoskeleton and in mitochondrial motility and function63. GLS, a key mitochondrial enzyme that catalyzes the deamidation of glutamine64, plays essential roles in oxidative phosphorylation, glutathione synthesis, and cellular redox homeostasis65. The decreased expression of glutathione S-transferase 1 (GSTT1), a multifunctional enzyme involved in oxidative stress, along with the antioxidant peroxiredoxin 6, in our experimental conditions supported the aforementioned data and indicated a potential accumulation of mitochondrial ROS in BCO-affected bone66,67.
One of the best-characterized pathways leading to cell death, a hallmark of BCO bone, involves mitochondria through outer membrane permeability, inner membrane potential changes, as well as elevated ROS production68. All the above DE mitochondrial markers (VDAC2, GLS, PLS3) have been reported to be involved in cell death69,70,71,72,73,74. Although further mechanistic studies are warranted, as bone resorption requires rapid cytoskeletal reorganization (sealing zone consisting of actin filament core surrounded by actin-binding proteins), we postulate that bacterial infection in BCO pathology dysregulates this actin cytoskeleton. This is supported, here, by dysregulation of PLS3, actin α1/γ1, and actin-binding proteins (ABPs) such as filamins (actin branching)75, tropomyosins (actin stabilizing)76, myosins (actin filament contraction and bundling)77, talin178, integrins (subunit αV, and β)79, actinins (ACTN1/4, actin cross-linking proteins)80, annexins81, fibronectin 182, hemopexin (heme scavenger)83, nebulin84, radixin85, stomatin86, vinculin87, vitronectin88, lamin89, scinderin (calcium-dependent actin filament-serving protein)90, and actin related protein 2/3 complex subunit 1B (ARPC1B). A number of studies have demonstrated a role for the actin cytoskeleton and several ABPs in triggering apoptosis upstream of caspases91,92, which has been shown by our group to be involved in bone attrition and osteoblast death21. This dysregulation of actin cytoskeleton complex alters the opening of VDAC, dysregulates the mitochondrial membrane permeabilization, depolarization, and integrity, and thereby leads to key apoptotic process via increased ROS production and oxidative stress93,94.
In addition to the mitochondria, ribosomes, nucleus, and nucleolus are central hubs for stress sensors95. Proteomic analysis identified here several DE ribosomal proteins (RPs), including RPL6, RPL7a, RPL19, RPL70, RPS2, RPS3, RPS3A1, RPSA, and RRBP1. Although RPs are well established as the basic building blocks in the ribosome assembly and biogenesis, as well as protein translation and synthesis96, there is increasing evidence indicating that RPs play critical roles in normal cell physiology, cellular response to stress, insults, and diseases97,98,99. In fact, it has been demonstrated that RPs have extra-ribosomal functions, including DNA repair, cell- cycle arrest, and apoptosis100,101,102,103. Specifically, RPS2 and RPL7A have been shown to be targets for mir-320a and to be involved in cartilage degradation104 and osteoporosis105. RPL19 and RPS3A1 have been found to be targets for mir-16-5p and to be involved in osteoarthritis106, osteoclastogenesis107, and rheumatoid arthritis108. In addition of being a RP, RPS3 has been shown to be a DNA repair endonuclease that is involved in apoptosis109. Together our proteomic data unveiled for the first time a potential key watchguard role of RPs, ribosomal stress, and ribosomopathy in BCO pathogenesis, however more questions related to whether (1) these RPs are free or membrane-bound? (2) These RPs are nucleolar or mitochondrial? (3) The RP perturbation is a consequence or an associated feature of BCO? And the nature of downstream cascades of these RPs beg to be answered. It is possible that RPs are directly or indirectly involved in various downstream signaling pathways, including RP-MDM2-P53 signaling110, NF-κB-Gadd45β pathway111, and/or proteasome-ubiquitin pathways112,113, which has been pinpointed by LC–MS/MS analysis. For instance, it has been shown that the binding of RPL26 to MDM2 promote the ubiquitination and proteasomal degradation of RPL26, which inhibits the P53 protein synthesis through the disruption of PRL26-P53 mRNA association114. Furthermore, the RPS7 was found to be a substrate for MDM2-meditaed ubiquitination, and the RPS7-ubiquitin fusion protein selectively inhibits MDM2-mediated P53 degradation and induces apoptosis115.
As disruption of ribosome biogenesis leads to nucleolar stress, proteomics and IPA analyses identified several DE-nuclear proteins between BCO-affected and healthy birds. Signal transducer and activator of transcription (STAT1 and STAT3) were oppositely dysregulated in tibiae of BCO-affected compared to healthy birds. These proteins belong to JAK-STAT family that contains at least seven members encoded by distinct genes, which are both signal transducers and transcription factors116. It has been shown that the STAT proteins were differentially activated in a context-dependent manner in response to various stimuli117. The down regulation of STAT3 in our experimental conditions supports its role in apoptosis and bone attrition118. Davidson and colleagues119 have shown that loss of STAT3 has a detrimental effect on osteoclast and bone structure. Boone et al.120 have reported an association between STAT3 deficiency and child hip osteonecrosis. Zhou et al.121 demonstrated a critical role for STAT3 in skeletal development and bone homeostasis. Using hematopoietic cell-specific STAT3 knockout mice, Zhang et al.122 have reported an accelerated osteoporosis with increased osteoclastogenesis. STAT1, on the contrary, has been shown when it is overexpressed, to enhance apoptotic cell death in cardiac myocytes exposed to ischemia-reperfusion123, however overexpression of STAT3 reduced STAT1-induced cell death. Furthermore, increasing number of studies confirmed that STAT1 and STAT3 have opposing actions on apoptotic cell death in various cell types124, via antagonistic effects on promoters of genes encoding anti-apoptotic BCL-2 and BCL-X proteins125. Kim et al.126 showed that STAT1-/- mice exhibited excessive osteoclastogenesis. Moreover, Xiao et al.127 have reported that STAT1 control bone formation via FGF signaling (Supplementary Fig. S1).
Curiously, IPA predicted PR, NRF1, and MYC as potential upstream regulators involved in BCO pathogenesis. Although the opposite regulation was detected by qPCR, the role of MYC in the apoptotic pathways is confounding and not fully understood128,129,130. MYC is a proto-oncogene, which encodes for a nuclear phosphoprotein that plays a key role in cellular transformation and apoptosis129,131,132. The target genes for MYC approached 4000 in human, that are involved in various cellular processes, including cell cycle, survival, protein synthesis, cell adhesion, cytoskeleton, and metabolism133,134. It is worth mentioning that HSP90135, OSTF1136, VDAC2137, RPs138,139, mitochondrial genes140, Col2A1141, and cytoskeleton-associated proteins142,143,144, are all targets for MYC. Of particular interest, MYC was found to be expressed in bone and required for osteoclast differentiation145. Moreover, MYC dysregulation has been shown to affect collagen and induce apoptosis and cartilage degeneration141.
Although its function remains elusive, epidemiological, clinical, and experimental data indicated that progesterone is active in bone metabolism and that PRs are expressed in human osteoblast146. Turner’s group has reported a high bone mass phenotype in global-PR knockout mice, which appeared to result from a reduced bone resorption rate in male, and a greater bone formation rate in female147. Wang and co-workers demonstrated that progesterone suppressed murine osteoblast MC3T3-E1 apoptosis via activation of PR and inhibition of caspase 3 and 9 activities, as well as cytochrome c release148. This effect was reversed by PR antagonist which supports IPA-predicted and qPCR-confirmed data here, that down regulation of PR might induce bone attrition in BCO-affected tibiae. Although the upstream mechanism by which the Staphylococcus infection down regulates PR is not known, the down-stream pathways mediated by PR is likely involve MYC149, Bcl-2150, STAT1151, MAPK152, and/or mTOR153.mTOR complexes are well established to regulate, among other things, protein synthesis and cell survival pathways154,155. Recently, Manning’s group has uncovered a surprising new function of mTOR in increasing cellular proteasome via NRF1 induction156,157, which has been predicted here by IPA analysis and confirmed by qPCR. NRF1, which is also known as NFE2L1/LCRF1/TCF11, is a member of the CNC subfamily of basic-leucine zipper (bZIP) transcription factors158. There are two NRF1 isoforms; a 120 kDa isoform localized primarily in the ER as an integral membrane protein, whereas the 65 kDa isoform is nuclear159. Binding-site selection experiments have shown that NRF1 binds preferentially to a consensus sequence that is identical to the antioxidant response element (ARE)160, which regulates numerous oxidative stress-related genes161. In furtherance of our data, osteoblast-specific NRF1-knockout mice have reduced bone mineral content and bone area162.
In summary, this is the first study using high throughput analysis in combination with bioinformatics tools to evaluate tibia proteome in BCO-affected and healthy broilers. Several DE proteins, protein interaction networks, disease-and function-based networks, canonical pathways, and upstream regulator were identified. We validated a panel of protein/gene candidates that following further mechanistic and functional studies may be potential biomarkers for BCO pathogenesis.
Data availability
The proteomic datasets generated during the current study are available in PRIDE database (EMBL-EBI ProteomeXchange, PRIDE database, https://doi.org/10.6019/PXD029085, with the accession PXD029085). To access please use the following ID: dridi@uark.edu and the PW: DrR76fJD.
References
Knowles, T. G. et al. Leg disorders in broiler chickens: Prevalence, risk factors and prevention. PLoS ONE 3, e1545–e1616. https://doi.org/10.1371/journal.pone.0001545 (2008).
Wideman, R. F. Jr. Bacterial chondronecrosis with osteomyelitis and lameness in broilers: A review. Poult. Sci. 95, 325–344. https://doi.org/10.3382/ps/pev320 (2016).
Al-Rubaye, A. A. et al. Genome analysis of staphylococcus agnetis, an agent of lameness in broiler chickens. PLoS ONE 10, e0143336-e143416. https://doi.org/10.1371/journal.pone.0143336 (2015).
Jiang, T. et al. Molecular survey of bacterial communities associated with bacterial chondronecrosis with osteomyelitis (BCO) in broilers. PLoS ONE 10, e0124403. https://doi.org/10.1371/journal.pone.0124403 (2015).
Tate, C. R., Mitchell, W. C. & Miller, R. G. Staphylococcus hyicus associated with turkey stifle joint osteomyelitis. Avian Dis. 37, 905–907 (1993).
Stalker, M. J., Brash, M. L., Weisz, A., Ouckama, R. M. & Slavic, D. Arthritis and osteomyelitis associated with Enterococcus cecorum infection in broiler and broiler breeder chickens in Ontario, Canada. J. Vet. Diagn. Invest. 22, 643–645. https://doi.org/10.1177/104063871002200426 (2010).
Rodgers, J. D., McCullagh, J. J., McNamee, P. T., Smyth, J. A. & Ball, H. J. Comparison of Staphylococcus aureus recovered from personnel in a poultry hatchery and in broiler parent farms with those isolated from skeletal disease in broilers. Vet. Microbiol. 69, 189–198. https://doi.org/10.1016/s0378-1135(99)00112-1 (1999).
Mutalib, A., Riddell, C. & Osborne, A. D. Studies on the pathogenesis of staphylococcal osteomyelitis in chickens. II. Role of the respiratory tract as a route of infection. Avian Dis 27, 157–160 (1983).
Emslie, K. R., Fenner, L. M. & Nade, S. M. Acute haematogenous osteomyelitis: II. The effect of a metaphyseal abscess on the surrounding blood supply. J. Pathol. 142, 129–134. https://doi.org/10.1002/path.1711420203 (1984).
Wideman, R. F. Jr. et al. A wire-flooring model for inducing lameness in broilers: Evaluation of probiotics as a prophylactic treatment. Poult. Sci. 91, 870–883. https://doi.org/10.3382/ps.2011-01907 (2012).
Emslie, K. R. & Nade, S. Acute hematogenous staphylococcal osteomyelitis. A description of the natural history in an avian model. Am. J. Pathol. 110, 333–345 (1983).
Wideman, R. F. Jr. et al. Susceptibility of 4 commercial broiler crosses to lameness attributable to bacterial chondronecrosis with osteomyelitis. Poult. Sci. 92, 2311–2325. https://doi.org/10.3382/ps.2013-03150 (2013).
Weimer, S. L. et al. Impact of experimentally induced bacterial chondronecrosis with osteomyelitis (BCO) lameness on health, stress, and leg health parameters in broilers. Poult Sci. 100, 101457. https://doi.org/10.1016/j.psj.2021.101457 (2021).
Dridi, S. et al. ERK1/2 activation is a therapeutic target in age-related macular degeneration. Proc. Natl. Acad. Sci. USA 109, 13781–13786. https://doi.org/10.1073/pnas.1206494109 (2012).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Ashburner, M. et al. Gene ontology: Tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25, 25–29. https://doi.org/10.1038/75556 (2000).
Szklarczyk, D. et al. STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447-452. https://doi.org/10.1093/nar/gku1003 (2015).
UniProt, C. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515. https://doi.org/10.1093/nar/gky1049 (2019).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530. https://doi.org/10.1093/bioinformatics/btt703 (2014).
Sardiu, M. E., Florens, L. & Washburn, M. P. Evaluation of clustering algorithms for protein complex and protein interaction network assembly. J. Proteome Res. 8, 2944–2952. https://doi.org/10.1021/pr900073d (2009).
Greene, E. et al. Double-stranded RNA is a novel molecular target in osteomyelitis pathogenesis: A translational avian model for human bacterial chondronecrosis with osteomyelitis. Am. J. Pathol. 189, 2077–2089. https://doi.org/10.1016/j.ajpath.2019.06.013 (2019).
Lassiter, K. et al. Orexin system is expressed in avian muscle cells and regulates mitochondrial dynamics. Am. J. Physiol. Regul. Integr. Comp. Physiol. 308, R173-187. https://doi.org/10.1152/ajpregu.00394.2014 (2015).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108. https://doi.org/10.1038/nprot.2008.73 (2008).
Belbin, O., Lehmann, S., Sabido, E. & Hirtz, C. Editorial: Proteomics as a tool for biomarker and drug target discovery: Improving the diagnosis and treatment of neurodegenerative diseases. Front. Aging Neurosci. 12, 232. https://doi.org/10.3389/fnagi.2020.00232 (2020).
Zhou, W., Petricoin, E. F. 3rd. & Longo, C. Mass spectrometry-based biomarker discovery. Methods Mol. Biol. 1606, 297–311. https://doi.org/10.1007/978-1-4939-6990-6_19 (2017).
Packialakshmi, B., Liyanage, R., Lay, J. O. Jr., Okimoto, R. & Rath, N. C. Proteomic changes in the plasma of broiler chickens with femoral head necrosis. Biomark. Insights 11, 55–62. https://doi.org/10.4137/BMI.S38291 (2016).
Packialakshmi, B., Rath, N. C., Huff, W. E. & Huff, G. R. Poultry femoral head separation and necrosis: A review. Avian Dis. 59, 349–354. https://doi.org/10.1637/11082-040715-Review.1 (2015).
Kim, J. A. et al. Quantitative proteomics analysis for the identification of differential protein expression in calf muscles between young and old SD rats using mass spectrometry. ACS Omega 6, 7422–7433. https://doi.org/10.1021/acsomega.0c05821 (2021).
Rousseau, A. & Bertolotti, A. Regulation of proteasome assembly and activity in health and disease. Nat. Rev. Mol. Cell Biol. 19, 697–712. https://doi.org/10.1038/s41580-018-0040-z (2018).
Terpos, E., Sezer, O., Croucher, P. & Dimopoulos, M. A. Myeloma bone disease and proteasome inhibition therapies. Blood 110, 1098–1104. https://doi.org/10.1182/blood-2007-03-067710 (2007).
Garrett, I. R. et al. Selective inhibitors of the osteoblast proteasome stimulate bone formation in vivo and in vitro. J. Clin. Invest. 111, 1771–1782. https://doi.org/10.1172/JCI16198 (2003).
Zavrski, I. et al. Proteasome inhibitors abrogate osteoclast differentiation and osteoclast function. Biochem. Biophys. Res. Commun. 333, 200–205. https://doi.org/10.1016/j.bbrc.2005.05.098 (2005).
Gronnemose, R. B. et al. Bacteria-host transcriptional response during endothelial invasion by Staphylococcus aureus. Sci. Rep. 11, 6037. https://doi.org/10.1038/s41598-021-84050-x (2021).
Matussek, A. et al. Infection of human endothelial cells with Staphylococcus aureus induces transcription of genes encoding an innate immunity response. Scand. J. Immunol. 61, 536–544. https://doi.org/10.1111/j.1365-3083.2005.01597.x (2005).
Cetin, G., Klafack, S., Studencka-Turski, M., Kruger, E. & Ebstein, F. The ubiquitin-proteasome system in immune cells. Biomolecules https://doi.org/10.3390/biom11010060 (2021).
Ramser, A., Greene, E., Wideman, R. & Dridi, S. Local and systemic cytokine, chemokine, and FGF profile in bacterial chondronecrosis with osteomyelitis (BCO)-affected broilers. Cells https://doi.org/10.3390/cells10113174 (2021).
Alkalay, I. et al. Stimulation-dependent I kappa B alpha phosphorylation marks the NF-kappa B inhibitor for degradation via the ubiquitin-proteasome pathway. Proc. Natl. Acad. Sci. USA 92, 10599–10603. https://doi.org/10.1073/pnas.92.23.10599 (1995).
Calderwood, S. K., Mambula, S. S., Gray, P. J. Jr. & Theriault, J. R. Extracellular heat shock proteins in cell signaling. FEBS Lett. 581, 3689–3694. https://doi.org/10.1016/j.febslet.2007.04.044 (2007).
van Eden, W., van der Zee, R. & Prakken, B. Heat-shock proteins induce T-cell regulation of chronic inflammation. Nat. Rev. Immunol. 5, 318–330. https://doi.org/10.1038/nri1593 (2005).
Daniels, G. A. et al. A simple method to cure established tumors by inflammatory killing of normal cells. Nat. Biotechnol. 22, 1125–1132. https://doi.org/10.1038/nbt1007 (2004).
Hunter-Lavin, C. et al. Hsp70 release from peripheral blood mononuclear cells. Biochem. Biophys. Res. Commun. 324, 511–517. https://doi.org/10.1016/j.bbrc.2004.09.075 (2004).
Asea, A. et al. HSP70 stimulates cytokine production through a CD14-dependant pathway, demonstrating its dual role as a chaperone and cytokine. Nat. Med. 6, 435–442. https://doi.org/10.1038/74697 (2000).
Asea, A. et al. Novel signal transduction pathway utilized by extracellular HSP70: Role of toll-like receptor (TLR) 2 and TLR4. J. Biol. Chem. 277, 15028–15034. https://doi.org/10.1074/jbc.M200497200 (2002).
Takahashi, K. et al. Analysis of heat shock proteins and cytokines expressed during early stages of osteoarthritis in a mouse model. Osteoarthr. Cartil. 5, 321–329 (1997).
Boehm, A. K., Seth, M., Mayr, K. G. & Fortier, L. A. Hsp90 mediates insulin-like growth factor 1 and interleukin-1beta signaling in an age-dependent manner in equine articular chondrocytes. Arthritis Rheum. 56, 2335–2343. https://doi.org/10.1002/art.22664 (2007).
Terauchi, R. et al. Hsp70 prevents nitric oxide-induced apoptosis in articular chondrocytes. Arthritis Rheum. 48, 1562–1568. https://doi.org/10.1002/art.11040 (2003).
Tribelli, P. M. et al. Staphylococcus aureus Lpl protein triggers human host cell invasion via activation of Hsp90 receptor. Cell Microbiol. 22, e13111. https://doi.org/10.1111/cmi.13111 (2020).
Cruz, T. F., Kandel, R. A. & Brown, I. R. Interleukin 1 induces the expression of a heat-shock gene in chondrocytes. Biochem. J. 277(Pt 2), 327–330 (1991).
Reddy, S. et al. Isolation and characterization of a cDNA clone encoding a novel peptide (OSF) that enhances osteoclast formation and bone resorption. J. Cell Physiol. 177, 636–645. https://doi.org/10.1002/(SICI)1097-4652(199812)177:4%3c636::AID-JCP14%3e3.0.CO;2-H (1998).
Siddiqui, J. A. & Partridge, N. C. Physiological bone remodeling: Systemic regulation and growth factor involvement. Physiol. (Bethesda) 31, 233–245. https://doi.org/10.1152/physiol.00061.2014 (2016).
Zaidi, M. Skeletal remodeling in health and disease. Nat. Med. 13, 791–801. https://doi.org/10.1038/nm1593 (2007).
Wideman, R. F. & Prisby, R. D. Bone circulatory disturbances in the development of spontaneous bacterial chondronecrosis with osteomyelitis: A translational model for the pathogenesis of femoral head necrosis. Front. Endocrinol. 3, 183. https://doi.org/10.3389/fendo.2012.00183 (2012).
Blair, H. C. How the osteoclast degrades bone. BioEssays News Rev. Mol., Cellular Dev. Biol. 20, 837–846. https://doi.org/10.1002/(SICI)1521-1878(199810)20:10%3c837::AID-BIES9%3e3.0.CO;2-D (1998).
Wilson, S. R., Peters, C., Saftig, P. & Bromme, D. Cathepsin K activity-dependent regulation of osteoclast actin ring formation and bone resorption. J. Biol. Chem. 284, 2584–2592. https://doi.org/10.1074/jbc.M805280200 (2009).
Vermeren, M. et al. Osteoclast stimulation factor 1 (Ostf1) KNOCKOUT increases trabecular bone mass in mice. Mamm. Genome 28, 498–514. https://doi.org/10.1007/s00335-017-9718-3 (2017).
Marini, J. C. & Blissett, A. R. New genes in bone development: What’s new in osteogenesis imperfecta. J. Clin. Endocrinol. Metab. 98, 3095–3103. https://doi.org/10.1210/jc.2013-1505 (2013).
Richards, A. J. et al. High efficiency of mutation detection in type 1 stickler syndrome using a two-stage approach: vitreoretinal assessment coupled with exon sequencing for screening COL2A1. Hum. Mutat. 27, 696–704. https://doi.org/10.1002/humu.20347 (2006).
Gentili, C. et al. Cell proliferation, extracellular matrix mineralization, and ovotransferrin transient expression during in vitro differentiation of chick hypertrophic chondrocytes into osteoblast-like cells. J. Cell Biol. 122, 703–712. https://doi.org/10.1083/jcb.122.3.703 (1993).
Gentili, C. et al. Ovotransferrin and ovotransferrin receptor expression during chondrogenesis and endochondral bone formation in developing chick embryo. J. Cell Biol. 124, 579–588. https://doi.org/10.1083/jcb.124.4.579 (1994).
Das, B. K. et al. Transferrin receptor 1-mediated iron uptake regulates bone mass in mice via osteoclast mitochondria and cytoskeleton. Elife https://doi.org/10.7554/eLife.73539 (2022).
Ferver, A., Greene, E., Wideman, R. & Dridi, S. Evidence of mitochondrial dysfunction in bacterial chondronecrosis with osteomyelitis-affected broilers. Front. Vet. Sci. 8, 640901. https://doi.org/10.3389/fvets.2021.640901 (2021).
Rostovtseva, T. K. et al. Tubulin binding blocks mitochondrial voltage-dependent anion channel and regulates respiration. Proc. Natl. Acad. Sci. USA 105, 18746–18751. https://doi.org/10.1073/pnas.0806303105 (2008).
Ligon, L. A. & Steward, O. Role of microtubules and actin filaments in the movement of mitochondria in the axons and dendrites of cultured hippocampal neurons. J. Comp. Neurol. 427, 351–361. https://doi.org/10.1002/1096-9861(20001120)427:3%3c351::aid-cne3%3e3.0.co;2-r (2000).
Hensley, C. T., Wasti, A. T. & DeBerardinis, R. J. Glutamine and cancer: Cell biology, physiology, and clinical opportunities. J. Clin. Invest. 123, 3678–3684. https://doi.org/10.1172/JCI69600 (2013).
Amores-Sanchez, M. I. & Medina, M. A. Glutamine, as a precursor of glutathione, and oxidative stress. Mol. Genet. Metab. 67, 100–105. https://doi.org/10.1006/mgme.1999.2857 (1999).
Leiers, B. et al. A stress-responsive glutathione S-transferase confers resistance to oxidative stress in Caenorhabditis elegans. Free Radic. Biol. Med. 34, 1405–1415. https://doi.org/10.1016/s0891-5849(03)00102-3 (2003).
Dancy, B. M. et al. Glutathione S-transferase mediates an ageing response to mitochondrial dysfunction. Mech. Ageing Dev. 153, 14–21. https://doi.org/10.1016/j.mad.2015.12.001 (2016).
Danial, N. N. & Korsmeyer, S. J. Cell death: Critical control points. Cell 116, 205–219. https://doi.org/10.1016/s0092-8674(04)00046-7 (2004).
Shoshan-Barmatz, V. & Gincel, D. The voltage-dependent anion channel: Characterization, modulation, and role in mitochondrial function in cell life and death. Cell Biochem. Biophys. 39, 279–292. https://doi.org/10.1385/CBB:39:3:279 (2003).
Godbole, A., Varghese, J., Sarin, A. & Mathew, M. K. VDAC is a conserved element of death pathways in plant and animal systems. Biochim. Biophys. Acta 1642, 87–96. https://doi.org/10.1016/s0167-4889(03)00102-2 (2003).
Tsujimoto, Y. & Shimizu, S. The voltage-dependent anion channel: An essential player in apoptosis. Biochimie 84, 187–193. https://doi.org/10.1016/s0300-9084(02)01370-6 (2002).
Zhang, J. et al. Inhibition of GLS suppresses proliferation and promotes apoptosis in prostate cancer. Biosci. Rep. https://doi.org/10.1042/BSR20181826 (2019).
Tong, Y. et al. SUCLA2-coupled regulation of GLS succinylation and activity counteracts oxidative stress in tumor cells. Mol. Cell 81, 2303–2316. https://doi.org/10.1016/j.molcel.2021.04.002 (2021).
Desouza, M., Gunning, P. W. & Stehn, J. R. The actin cytoskeleton as a sensor and mediator of apoptosis. BioArchitecture 2, 75–87. https://doi.org/10.4161/bioa.20975 (2012).
Iwamoto, D. V. et al. Structural basis of the filamin A actin-binding domain interaction with F-actin. Nat. Struct. Mol. Biol. 25, 918–927. https://doi.org/10.1038/s41594-018-0128-3 (2018).
Vilfan, A. The binding dynamics of tropomyosin on actin. Biophys. J. 81, 3146–3155. https://doi.org/10.1016/S0006-3495(01)75951-6 (2001).
Bekyarova, T. I. et al. Reverse actin sliding triggers strong myosin binding that moves tropomyosin. Proc. Natl. Acad. Sci. USA 105, 10372–10377. https://doi.org/10.1073/pnas.0709877105 (2008).
Owen, L. M., Bax, N. A., Weis, W. I. & Dunn, A. R. The C-terminal actin-binding domain of talin forms an asymmetric catch bond with F-actin. Proc. Natl. Acad. Sci. USA 119, e2109329119. https://doi.org/10.1073/pnas.2109329119 (2022).
Vicente-Manzanares, M., Choi, C. K. & Horwitz, A. R. Integrins in cell migration–the actin connection. J. Cell Sci. 122, 199–206. https://doi.org/10.1242/jcs.018564 (2009).
Hampton, C. M., Taylor, D. W. & Taylor, K. A. Novel structures for alpha-actinin:F-actin interactions and their implications for actin-membrane attachment and tension sensing in the cytoskeleton. J. Mol. Biol. 368, 92–104. https://doi.org/10.1016/j.jmb.2007.01.071 (2007).
Hayes, M. J., Rescher, U., Gerke, V. & Moss, S. E. Annexin-actin interactions. Traffic 5, 571–576. https://doi.org/10.1111/j.1600-0854.2004.00210.x (2004).
Bloom, L., Ingham, K. C. & Hynes, R. O. Fibronectin regulates assembly of actin filaments and focal contacts in cultured cells via the heparin-binding site in repeat III13. Mol. Biol. Cell 10, 1521–1536. https://doi.org/10.1091/mbc.10.5.1521 (1999).
Lennon, R. et al. Hemopexin induces nephrin-dependent reorganization of the actin cytoskeleton in podocytes. J. Am. Soc. Nephrol. 19, 2140–2149. https://doi.org/10.1681/ASN.2007080940 (2008).
Pappas, C. T., Krieg, P. A. & Gregorio, C. C. Nebulin regulates actin filament lengths by a stabilization mechanism. J. Cell Biol. 189, 859–870. https://doi.org/10.1083/jcb.201001043 (2010).
Loebrich, S., Bahring, R., Katsuno, T., Tsukita, S. & Kneussel, M. Activated radixin is essential for GABAA receptor alpha5 subunit anchoring at the actin cytoskeleton. EMBO J. 25, 987–999. https://doi.org/10.1038/sj.emboj.7600995 (2006).
Chen, J. C. et al. Up-regulation of stomatin expression by hypoxia and glucocorticoid stabilizes membrane-associated actin in alveolar epithelial cells. J. Cell Mol. Med. 17, 863–872. https://doi.org/10.1111/jcmm.12069 (2013).
Boujemaa-Paterski, R. et al. Talin-activated vinculin interacts with branched actin networks to initiate bundles. Elife https://doi.org/10.7554/eLife.53990 (2020).
Furutani, Y. et al. Vitronectin induces phosphorylation of ezrin/radixin/moesin actin-binding proteins through binding to its novel neuronal receptor telencephalin. J. Biol. Chem. 287, 39041–39049. https://doi.org/10.1074/jbc.M112.383851 (2012).
Simon, D. N., Zastrow, M. S. & Wilson, K. L. Direct actin binding to A- and B-type lamin tails and actin filament bundling by the lamin A tail. Nucleus 1, 264–272. https://doi.org/10.4161/nucl.1.3.11799 (2010).
Tchakarov, L., Vitale, M. L., Jeyapragasan, M., Rodriguez Del Castillo, A. & Trifaro, J. M. Expression of scinderin, an actin filament-severing protein, in different tissues. FEBS Lett. 268, 209–212. https://doi.org/10.1016/0014-5793(90)81010-l (1990).
Parlato, S. et al. CD95 (APO-1/Fas) linkage to the actin cytoskeleton through ezrin in human T lymphocytes: A novel regulatory mechanism of the CD95 apoptotic pathway. EMBO J. 19, 5123–5134. https://doi.org/10.1093/emboj/19.19.5123 (2000).
Franklin-Tong, V. E. & Gourlay, C. W. A role for actin in regulating apoptosis/programmed cell death: Evidence spanning yeast, plants and animals. Biochem. J. 413, 389–404. https://doi.org/10.1042/BJ20080320 (2008).
Xu, X., Forbes, J. G. & Colombini, M. Actin modulates the gating of Neurospora crassa VDAC. J. Membr. Biol. 180, 73–81. https://doi.org/10.1007/s002320010060 (2001).
Rosenblatt, J., Raff, M. C. & Cramer, L. P. An epithelial cell destined for apoptosis signals its neighbors to extrude it by an actin- and myosin-dependent mechanism. Curr. Biol. 11, 1847–1857. https://doi.org/10.1016/s0960-9822(01)00587-5 (2001).
Boulon, S., Westman, B. J., Hutten, S., Boisvert, F. M. & Lamond, A. I. The nucleolus under stress. Mol. Cell 40, 216–227. https://doi.org/10.1016/j.molcel.2010.09.024 (2010).
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948. https://doi.org/10.1126/science.1211204 (2011).
Narla, A. & Ebert, B. L. Ribosomopathies: Human disorders of ribosome dysfunction. Blood 115, 3196–3205. https://doi.org/10.1182/blood-2009-10-178129 (2010).
Ebert, B. L. et al. Identification of RPS14 as a 5q- syndrome gene by RNA interference screen. Nature 451, 335–339. https://doi.org/10.1038/nature06494 (2008).
Xue, S. & Barna, M. Specialized ribosomes: A new frontier in gene regulation and organismal biology. Nat. Rev. Mol. Cell Biol. 13, 355–369. https://doi.org/10.1038/nrm3359 (2012).
He, H. & Sun, Y. Ribosomal protein S27L is a direct p53 target that regulates apoptosis. Oncogene 26, 2707–2716. https://doi.org/10.1038/sj.onc.1210073 (2007).
Kim, J. et al. Implication of mammalian ribosomal protein S3 in the processing of DNA damage. J. Biol. Chem. 270, 13620–13629. https://doi.org/10.1074/jbc.270.23.13620 (1995).
Pestov, D. G., Strezoska, Z. & Lau, L. F. Evidence of p53-dependent cross-talk between ribosome biogenesis and the cell cycle: Effects of nucleolar protein Bop1 on G(1)/S transition. Mol. Cell Biol. 21, 4246–4255. https://doi.org/10.1128/MCB.21.13.4246-4255.2001 (2001).
Fumagalli, S. et al. Absence of nucleolar disruption after impairment of 40S ribosome biogenesis reveals an rpL11-translation-dependent mechanism of p53 induction. Nat. Cell Biol. 11, 501–508. https://doi.org/10.1038/ncb1858 (2009).
Jin, Y. et al. The role of miR-320a and IL-1beta in human chondrocyte degradation. Bone Joint Res. 6, 196–203. https://doi.org/10.1302/2046-3758.64.BJR-2016-0224.R1 (2017).
De-Ugarte, L. et al. Pro-osteoporotic miR-320a impairs osteoblast function and induces oxidative stress. PLoS ONE 13, e0208131-e208216. https://doi.org/10.1371/journal.pone.0208131 (2018).
Li, L. et al. MicroRNA-16-5p controls development of osteoarthritis by targeting SMAD3 in chondrocytes. Curr. Pharm. Des. 21, 5160–5167. https://doi.org/10.2174/1381612821666150909094712 (2015).
Sang, S., Zhang, Z., Qin, S., Li, C. & Dong, Y. MicroRNA-16-5p inhibits osteoclastogenesis in giant cell tumor of bone. Biomed. Res. Int. 2017, 3173547. https://doi.org/10.1155/2017/3173547 (2017).
Dunaeva, M., Blom, J., Thurlings, R. & Pruijn, G. J. M. Circulating serum miR-223-3p and miR-16-5p as possible biomarkers of early rheumatoid arthritis. Clin. Exp. Immunol. 193, 376–385. https://doi.org/10.1111/cei.13156 (2018).
Jang, C. Y., Lee, J. Y. & Kim, J. RpS3, a DNA repair endonuclease and ribosomal protein, is involved in apoptosis. FEBS Lett. 560, 81–85. https://doi.org/10.1016/S0014-5793(04)00074-2 (2004).
Liu, Y., Deisenroth, C. & Zhang, Y. RP-MDM2-p53 pathway: Linking ribosomal biogenesis and tumor surveillance. Trends Cancer 2, 191–204. https://doi.org/10.1016/j.trecan.2016.03.002 (2016).
Wan, F. et al. IKKbeta phosphorylation regulates RPS3 nuclear translocation and NF-kappaB function during infection with Escherichia coli strain O157:H7. Nat. Immunol. 12, 335–343. https://doi.org/10.1038/ni.2007 (2011).
Wang, A. et al. Ribosomal protein RPL41 induces rapid degradation of ATF4, a transcription factor critical for tumour cell survival in stress. J. Pathol. 225, 285–292. https://doi.org/10.1002/path.2918 (2011).
Gao, M. et al. Ribosomal protein S7 regulates arsenite-induced GADD45alpha expression by attenuating MDM2-mediated GADD45alpha ubiquitination and degradation. Nucleic Acids Res. 41, 5210–5222. https://doi.org/10.1093/nar/gkt223 (2013).
Ofir-Rosenfeld, Y., Boggs, K., Michael, D., Kastan, M. B. & Oren, M. Mdm2 regulates p53 mRNA translation through inhibitory interactions with ribosomal protein L26. Mol. Cell 32, 180–189. https://doi.org/10.1016/j.molcel.2008.08.031 (2008).
Zhu, Y. et al. Ribosomal protein S7 is both a regulator and a substrate of MDM2. Mol Cell 35, 316–326. https://doi.org/10.1016/j.molcel.2009.07.014 (2009).
Darnell, J. E. Jr. STATs and gene regulation. Science 277, 1630–1635. https://doi.org/10.1126/science.277.5332.1630 (1997).
Bromberg, J. F. Activation of STAT proteins and growth control. BioEssays 23, 161–169. https://doi.org/10.1002/1521-1878(200102)23:2%3c161::AID-BIES1023%3e3.0.CO;2-0 (2001).
Aoki, Y., Feldman, G. M. & Tosato, G. Inhibition of STAT3 signaling induces apoptosis and decreases survivin expression in primary effusion lymphoma. Blood 101, 1535–1542. https://doi.org/10.1182/blood-2002-07-2130 (2003).
Davidson, R. K. et al. The loss of STAT3 in mature osteoclasts has detrimental effects on bone structure. PLoS ONE 15, e0236891. https://doi.org/10.1371/journal.pone.0236891 (2020).
Staines Boone, A. T. et al. Zoledronate as effective treatment for minimal trauma fractures in a child with STAT3 deficiency and osteonecrosis of the hip. Pediatr. Blood Cancer 63, 2054–2057. https://doi.org/10.1002/pbc.26119 (2016).
Zhou, S. et al. STAT3 is critical for skeletal development and bone homeostasis by regulating osteogenesis. Nat. Commun. 12, 6891. https://doi.org/10.1038/s41467-021-27273-w (2021).
Zhang, Z. et al. Osteoporosis with increased osteoclastogenesis in hematopoietic cell-specific STAT3-deficient mice. Biochem. Biophys. Res. Commun. 328, 800–807. https://doi.org/10.1016/j.bbrc.2005.01.019 (2005).
Stephanou, A. et al. Ischemia-induced STAT-1 expression and activation play a critical role in cardiomyocyte apoptosis. J. Biol. Chem. 275, 10002–10008. https://doi.org/10.1074/jbc.275.14.10002 (2000).
Battle, T. E. & Frank, D. A. The role of STATs in apoptosis. Curr. Mol. Med. 2, 381–392. https://doi.org/10.2174/1566524023362456 (2002).
Stephanou, A., Brar, B. K., Knight, R. A. & Latchman, D. S. Opposing actions of STAT-1 and STAT-3 on the Bcl-2 and Bcl-x promoters. Cell Death Differ. 7, 329–330. https://doi.org/10.1038/sj.cdd.4400656 (2000).
Kim, S. et al. Stat1 functions as a cytoplasmic attenuator of Runx2 in the transcriptional program of osteoblast differentiation. Genes. Dev. 17, 1979–1991. https://doi.org/10.1101/gad.1119303 (2003).
Xiao, L. et al. Stat1 controls postnatal bone formation by regulating fibroblast growth factor signaling in osteoblasts. J. Biol. Chem. 279, 27743–27752. https://doi.org/10.1074/jbc.M314323200 (2004).
Pelengaris, S., Khan, M. & Evan, G. c-MYC: More than just a matter of life and death. Nat. Rev Cancer 2, 764–776. https://doi.org/10.1038/nrc904 (2002).
Evan, G. I. et al. Induction of apoptosis in fibroblasts by c-myc protein. Cell 69, 119–128. https://doi.org/10.1016/0092-8674(92)90123-t (1992).
Ayala-Torres, S., Zhou, F. & Thompson, E. B. Apoptosis induced by oxysterol in CEM cells is associated with negative regulation of c-myc. Exp. Cell Res. 246, 193–202. https://doi.org/10.1006/excr.1998.4308 (1999).
Hoffman, B. & Liebermann, D. A. Apoptotic signaling by c-MYC. Oncogene 27, 6462–6472. https://doi.org/10.1038/onc.2008.312 (2008).
Shi, Y. et al. Role for c-myc in activation-induced apoptotic cell death in T cell hybridomas. Science 257, 212–214. https://doi.org/10.1126/science.1378649 (1992).
Dang, C. V. et al. The c-Myc target gene network. Semin. Cancer Biol. 16, 253–264. https://doi.org/10.1016/j.semcancer.2006.07.014 (2006).
Elkon, R. et al. In silico identification of transcriptional regulators associated with c-Myc. Nucleic Acids Res. 32, 4955–4961. https://doi.org/10.1093/nar/gkh816 (2004).
Teng, S. C. et al. Direct activation of HSP90A transcription by c-Myc contributes to c-Myc-induced transformation. J. Biol. Chem. 279, 14649–14655. https://doi.org/10.1074/jbc.M308842200 (2004).
Lyraki, R. et al. Characterization of a novel RP2-OSTF1 interaction and its implication for actin remodelling. J. Cell Sci. https://doi.org/10.1242/jcs.211748 (2018).
Yang, Y. et al. Nedd4 ubiquitylates VDAC2/3 to suppress erastin-induced ferroptosis in melanoma. Nat. Commun. 11, 433. https://doi.org/10.1038/s41467-020-14324-x (2020).
van Riggelen, J., Yetil, A. & Felsher, D. W. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat. Rev. Cancer 10, 301–309. https://doi.org/10.1038/nrc2819 (2010).
Arabi, A. et al. c-Myc associates with ribosomal DNA and activates RNA polymerase I transcription. Nat. Cell Biol. 7, 303–310. https://doi.org/10.1038/ncb1225 (2005).
Popay, T. M. et al. MYC regulates ribosome biogenesis and mitochondrial gene expression programs through its interaction with host cell factor-1. Elife https://doi.org/10.7554/eLife.60191 (2021).
Melnik, S. et al. Impact of c-MYC expression on proliferation, differentiation, and risk of neoplastic transformation of human mesenchymal stromal cells. Stem. Cell Res. Ther. 10, 73. https://doi.org/10.1186/s13287-019-1187-z (2019).
Ying, X. et al. ANXA1 (Annexin A1) regulated by MYC (MYC proto-oncogene) promotes the growth of papillary thyroid carcinoma. Bioengineered 12, 9251–9265. https://doi.org/10.1080/21655979.2021.1996511 (2021).
Boudjadi, S., Carrier, J. C., Groulx, J. F. & Beaulieu, J. F. Integrin alpha1beta1 expression is controlled by c-MYC in colorectal cancer cells. Oncogene 35, 1671–1678. https://doi.org/10.1038/onc.2015.231 (2016).
Sauzeau, V., Berenjeno, I. M., Citterio, C. & Bustelo, X. R. A transcriptional cross-talk between RhoA and c-Myc inhibits the RhoA/Rock-dependent cytoskeleton. Oncogene 29, 3781–3792. https://doi.org/10.1038/onc.2010.134 (2010).
Battaglino, R. et al. c-myc is required for osteoclast differentiation. J. Bone Miner. Res. 17, 763–773. https://doi.org/10.1359/jbmr.2002.17.5.763 (2002).
MacNamara, P., O’Shaughnessy, C., Manduca, P. & Loughrey, H. C. Progesterone receptors are expressed in human osteoblast-like cell lines and in primary human osteoblast cultures. Calcif. Tissue Int. 57, 436–441. https://doi.org/10.1007/BF00301947 (1995).
Yao, W. et al. Inhibition of the progesterone nuclear receptor during the bone linear growth phase increases peak bone mass in female mice. PLoS ONE 5, e11410. https://doi.org/10.1371/journal.pone.0011410 (2010).
Wang, Q. P. et al. Effect of progesterone on apoptosis of murine MC3T3-E1 osteoblastic cells. Amino Acids 36, 57–63. https://doi.org/10.1007/s00726-008-0028-7 (2009).
Kavlashvili, T. et al. Inverse relationship between progesterone receptor and myc in endometrial cancer. PLoS ONE 11, e0148912. https://doi.org/10.1371/journal.pone.0148912 (2016).
Yin, P. et al. Progesterone receptor regulates Bcl-2 gene expression through direct binding to its promoter region in uterine leiomyoma cells. J. Clin. Endocrinol. Metab. 92, 4459–4466. https://doi.org/10.1210/jc.2007-0725 (2007).
Goodman, M. L. et al. Progesterone receptor attenuates STAT1-mediated IFN signaling in breast cancer. J. Immunol. 202, 3076–3086. https://doi.org/10.4049/jimmunol.1801152 (2019).
Skildum, A., Faivre, E. & Lange, C. A. Progesterone receptors induce cell cycle progression via activation of mitogen-activated protein kinases. Mol. Endocrinol. 19, 327–339. https://doi.org/10.1210/me.2004-0306 (2005).
Pedroza, D. A. et al. Progesterone receptor membrane component 1 promotes the growth of breast cancers by altering the phosphoproteome and augmenting EGFR/PI3K/AKT signalling. Br. J. Cancer 123, 1326–1335. https://doi.org/10.1038/s41416-020-0992-6 (2020).
Dibble, C. C. & Manning, B. D. Signal integration by mTORC1 coordinates nutrient input with biosynthetic output. Nat. Cell Biol. 15, 555–564. https://doi.org/10.1038/ncb2763 (2013).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141. https://doi.org/10.1038/ncb2152 (2011).
Zhang, Y. et al. Coordinated regulation of protein synthesis and degradation by mTORC1. Nature 513, 440–443. https://doi.org/10.1038/nature13492 (2014).
Zhang, Y. & Manning, B. D. mTORC1 signaling activates NRF1 to increase cellular proteasome levels. Cell Cycle 14, 2011–2017. https://doi.org/10.1080/15384101.2015.1044188 (2015).
Chan, J. Y., Han, X. L. & Kan, Y. W. Cloning of Nrf1, an NF-E2-related transcription factor, by genetic selection in yeast. Proc. Natl. Acad. Sci. U.S.A. 90, 11371–11375. https://doi.org/10.1073/pnas.90.23.11371 (1993).
Ohtsuji, M. et al. Nrf1 and Nrf2 play distinct roles in activation of antioxidant response element-dependent genes. J. Biol. Chem. 283, 33554–33562. https://doi.org/10.1074/jbc.M804597200 (2008).
Johnsen, O., Murphy, P., Prydz, H. & Kolsto, A. B. Interaction of the CNC-bZIP factor TCF11/LCR-F1/Nrf1 with MafG: Binding-site selection and regulation of transcription. Nucleic. Acids Res. 26, 512–520. https://doi.org/10.1093/nar/26.2.512 (1998).
Biswas, M. & Chan, J. Y. Role of Nrf1 in antioxidant response element-mediated gene expression and beyond. Toxicol. Appl. Pharmacol. 244, 16–20. https://doi.org/10.1016/j.taap.2009.07.034 (2010).
Xing, W. et al. Nuclear factor-E2-related factor-1 mediates ascorbic acid induction of osterix expression via interaction with antioxidant-responsive element in bone cells. J. Biol. Chem. 282, 22052–22061. https://doi.org/10.1074/jbc.M702614200 (2007).
Funding
This study was supported by a grant from the University of Arkansas Chancellor’s Innovation Funds (003226-00001A to SD).
Author information
Authors and Affiliations
Contributions
S.D. conceived and designed the study. S.D. provided the reagents. E.S.G., R.W. and S.D. conducted the experiments and analyzed the data. R.W. scored the birds. R.L. performed the LS-MS/MS. A.R. and E.S.G performed the immunoblot analysis. GM performed the qPCR analysis. J.S.D. created the Fig. 2. J.C. wrote the first draft and S.D. wrote the final paper with a critical review by E.S.G., A.R., G.M., J.S.D., R.L., and R.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cook, J., Greene, E.S., Ramser, A. et al. Comparative- and network-based proteomic analysis of bacterial chondronecrosis with osteomyelitis lesions in broiler’s proximal tibiae identifies new molecular signatures of lameness. Sci Rep 13, 5947 (2023). https://doi.org/10.1038/s41598-023-33060-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33060-y
- Springer Nature Limited