Abstract
Cutaneous melanoma (CM) is a highly malignant tumor originating from melanocytes, and its metastasis and recurrence are the major causes of death in CM patients. PANoptosis is a newly defined inflammatory programmed cell death that crosstalk pyroptosis, apoptosis, and necroptosis. PANoptosis participates in the regulation of tumor progression, especially the expression of PANoptosis related genes (PARGs). Although pyroptosis, apoptosis, and necroptosis have received attention in CM, respectively, the link between them remains elusive. Therefore, this study aimed to investigate the potential regulatory role of PANoptosis and PARGs in CM and the relationship among PANoptosis, PARGs and tumor immunity. We identified 3 PARGs associated with prognosis in CM patients by The Cancer Genome Atlas. Risk model and nomogram were established. Enrichment analysis of differentially expressed genes indicated that CM was immune-related. Subsequent analyses indicated that prognosis-related PARGs were associated with immune scores and infiltration of immune cells in CM patients. In addition, immunotherapy and drug sensitivity results indicated an association between prognosis-related PARGs and drug resistance in CM patients. In conclusion, PARGs play a key role in the progression of tumors in CM patients. PARGs can be used not only for risk assessment and OS prediction in CM patients, but also reflect the immune landscape of CM patients, which can provide a novel reference for individualized tumor treatment.
Similar content being viewed by others
Introduction
Among all skin tumors, cutaneous melanoma (CM) is the skin tumor with the highest mortality rate1. According to the GLOBOCAN 2020 cancer report prepared by the International Agency for Research on Cancer, there will be nearly more than 300,000 new cancer cases and 60,000 new cancer deaths in 2020 alone2. Notably, the mortality rate of CM is four times higher than that of non-melanoma skin tumors2. The 5-year survival rate of CM patients remains unsatisfactory due to the fact that CM is a highly metastatic malignancy3. Therefore, early detection and diagnosis of CM can be of great help in the treatment of CM4. Nowadays, research on the diagnosis and treatment of CM still worthwhile, especially the discovery of some reliable prognostic biomarkers will be crucial5.
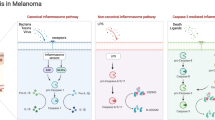
The imbalance between apoptosis and proliferation was the direct cause of tumorigenesis6. Notably, alterations in the apoptotic program can also affect tumor response to drugs7,8. Therefore, understanding the new programmed death pathways is essential for tumor prevention, diagnosis, and treatment. PANoptosis is a recently newly defined inflammatory programmed cell death that is based on an inflammatory vesicle complex (PANoptosome)9. It has been shown that PANoptosis, which contains molecules which associated with pyroptosis, apoptosis, and necroptosis, can crosstalk them and influence their relationship with each other10,11,12,13,14.
The key role of PANoptosis in many tumors has been reported, such as colorectal cancer15, gastric cancer16, and adrenocortical cancer17. It has also been shown that pyroptosis, apoptosis, and necroptosis are all closely related to CM18,19. However, current studies of PANoptosis in CM tend to focus on only one or two of them, and the joint roles of all three of them have not been reported. Therefore, the discovery of relevant biomarkers of PANoptosis in CM is crucial. In conclusion, the better understanding of the role of PANoptosis could provide new insights into the diagnosis and targeted therapy of CM.
In this study, PANoptosis-related genes (PARGs) associated with OS in CM patients were identified by analyzing datasets downloaded from the Cancer Genome Atlas database (TCGA). The risk model was constructed based on prognosis-related PARGs and the predictive power of the model was evaluated. Enrichment analysis of differentially expressed genes (DEGs) was used to further understand the differences in biological function between CM patients with different risks. The immune landscape of CM patients was assessed using a combination of immune algorithms. In addition, we analyzed the differences in immunotherapy response and drug sensitivity between patients with different risks. Overall, this study may provide new insights into the diagnosis and targeted therapy of CM.
Materials and methods
Data collection and preliminary processing
The transcriptome matrix and relevant clinical information of CM patients involved in this study were downloaded from the Cancer Genome Atlas database (TCGA) (https://portal.gdc.cancer.gov/). Transcriptomes without survival time information will not be considered and a total of 454 CM samples were finally available for subsequent analysis (Table 1 and Supplementary Table 1). The transcriptome of each CM sample was merged for annotation by perl script20. Age, sex, grade, stage, and TMN stage were obtained from the relevant clinical information downloaded from TCGA. The data involved in this study were downloaded from public databases and did not require ethics committee approval or written informed consent from patients.
Identification of prognosis-associated PARGs and construction of risk models
A total of 14 PARGs were extracted based on previous studies, and these genes are mainly components of the PANoptosome9. Univariate Cox regression analysis was used to assess the association of PARGs with CM. The least absolute shrinkage and selection operator (LASSO) algorithm was used to further screen PARGs associated with OS of CM. PARGs with independent prediction of CM prognosis were identified by multivariate Cox regression analysis, and risk models were constructed based on prognosis-related PARGs with the formula: Risk score = ∑i = Coefi ∗ (expression of PANRGi). The CM sample was divided into low-risk and high-risk groups according to the median risk score. Kaplan–Meier survival curves were used to assess the differences between OS of CM patients with different risks.
Internal validation of the risk model
To validate the predictive ability of the risk model, 454 CM samples were divided into a training cohort and a test cohort according to a 7:3 ratio and risk scores were calculated. There were 318 samples in the training cohort and 136 samples in the test cohort. Afterwards, the samples in each cohort were divided into low risk and high risk groups based on the median risk scores of each cohort. In each cohort, Kaplan–Meier survival curves were used to analyze the difference in OS between patients in the low- and high-risk groups.
Independent prognostic analysis of risk models and clinical characteristics
Prognostic independence of risk model and clinical characteristics was assessed by univariate and multivariate Cox regression analysis (performed by the R package “survival”). Kaplan–Meier survival curves were used to demonstrate differences in OS of CM patients with different clinical characteristics. The nomogram based on risk models and clinical characteristics were constructed by the R package "rms", and predictive accuracy was assessed by calibration plots and decision curves. The predicted survival probabilities of all variables were analyzed and visualized by the R package “pROC” and the R package “ggplot2”.
Screening and biological function enrichment for DEGs
The “limma” R package was used to evaluate and screen DEGs between different risk patient groups with a |Fold Change|≥ 2 and a p value < 0.05. Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis and Gene Ontology (GO) analysis were performed with the “clusterProfiler” R package to assess the enrichment of pathways and biological functions involved in DEGs21. Visualization of KEGG enrichment results by the “ggplot2” R package.
Immune landscape and drug sensitivity analysis
Multiple immune algorithms were used to assess the immune landscape of the samples in our study. The ESTIMATE algorithm can estimate stromal and immune cells and provide an ESTIMATE score. The CIBERSORT algorithm can assess the proportion of 22 immune cells in each sample. The single sample gene set enrichment analysis (ssGSEA) allows the assessment of 23 immune cells as well as immune function22. Tumor immune dysfunction and rejection (TIDE) scores for CM patients were calculated through the TIDE database (http://tide.dfci.harvard.edu/login/). Anti-tumor drug response prediction was performed in CM patients based on the Genomics of Drug Sensitivity in Cancer (GDSC) database, as reflected by half maximal inhibitory concentration (IC50)23. Differences in IC50 between different groups of CM patients were analyzed and visualized by the “ggplot2” R package.
Consensus clustering analysis
To further clarify the relationship between PARGs and prognosis, CM patients were distinguished into several subgroups based on prognosis-related PARGs. And a multifaceted comparative analysis of different subgroups will be performed, including Kaplan–Meier survival analysis, immune infiltration landscape, and immunotherapy response.
Statistical analysis
Statistical analyses involved in this study were performed using R software (version 4.1.2), and Wilcoxon rank sum test was used to analyze the differences between the two groups, with *p value < 0.05 as the threshold value to determine whether there was a statistical difference.
Results
Construction of a risk model for CM based on PARGs prognostic signature
To illustrate the relationship between PARGs and CM, a new risk model based on PARGs was constructed to predict the prognosis of CM. As shown in Fig. 1A, 8 PARGs were determined to be associated with OS of CM by univariate Cox regression analysis. Further analysis by the LASSO showed that 4 PARGs were able to predict the OS of CM (Fig. 1B,C). By multivariate Cox regression analysis, 3 PARGs with independent predictive power were finally identified for modeling. Patients with CM were ranked according to median risk score and divided into low- and high-risk groups. The scatter plot results showed that patients in the low-risk group tended to have higher OS (Fig. 1D,E). Kaplan–Meier survival curve analysis showed that patients in different risk groups had different OS (Fig. 1F). Notably, patients in the low-risk group had a higher OS than those in the high-risk group. The expression heat map of the 3 prognosis-related PARGs in the low-risk and high-risk group is shown in Fig. 1G. The results indicated that these 3 prognosis-related PARGs (ZBP1, MAP3K7, and RBCK1) were highly expressed in the low-risk group and less expressed in the high-risk group.
Construction of risk model based on prognosis-related PARGs and analysis of OS in CM patients. (A) Results of univariate cox regression analysis of PARGs. HR = 1: No impact; HR < 1: reduced risk, positively correlated with survival time; HR > 1: Increased risk, negatively correlated with survival time. Minimum λ (B) and optimal coefficients (C) of prognostic PARGs based on Lasso regression analysis. (D) Scatter plot of risk scores for CM patients. (E) Ranking of risk scores of CM patients. (F) Results of Kaplan–Meier survival curve analysis based on risk model. (G) Heat map of the expression of prognosis-related PARGs in CM patients. *p < 0.05; **p < 0.01; ***p < 0.001.
Validation of PARGs prognostic signature in training and test cohorts
Internal validation was performed to assess the accuracy and independence of the PARGs prognostic signature in predicting the prognosis of CM patients.CM patients were randomly divided into a training cohort and a test cohort in a ratio of 7:3. Same as before, patients were divided into low- and high-risk groups according to the PARGs prognostic signature in both cohorts. As shown in Fig. 2A,B,D,E, scatter plots indicate that risk score was negatively correlated with survival time for patients in both the training and test cohorts. The Kaplan–Meier survival curve results indicate that in both cohorts, patients with low risk score had higher OS than patients with high risk score (Fig. 2C,F). The heat map showed that the expression of ZBP1, MAP3K7 and RBCK1 was significantly higher in the low-risk group (Fig. 2G,H). These results suggest that risk model construction based on the prognostic characteristics of PARGs can accurately assess the prognosis of CM patients.
Validation results of PARG prognostic signature in the training and test cohorts. (A) Scatter plot of risk scores for CM patients in training cohorts. (B) Ranking of risk scores of CM patients in training cohorts. (C) Results of Kaplan–Meier survival curve analysis based on risk model in training cohorts. (D) Scatter plot of risk scores for CM patients in test cohorts. (E) Ranking of risk scores of CM patients in test cohorts. (F) Results of Kaplan–Meier survival curve analysis based on risk model in test cohorts. Heat map of the expression of prognosis-related PARGs in CM patients in training (G) and test cohorts (H). *p < 0.05; **p < 0.01; ***p < 0.001.
Risk model based on the PARGs prognostic signature was an independent prognosis indicator
Univariate and multivariate Cox regression analyses were used to assess the independence of risk scores predicting prognosis based on the PARGs prognostic signature. Univariate Cox regression analysis showed that age (hazard ratio (HR) = 1.019, p = 0.004), stage (HR = 1.393, p = 0.003), T (HR = 1. 471, p < 0.001), N (HR = 1.365, p < 0.001) and risk score (HR = 1.589, p < 0.001) were associated with the OS rate of CM were strongly correlated (Fig. 3A). The results of multivariate Cox regression analysis showed that T (HR = 1.380, p = 0.001), N (HR = 1.671, p < 0.001) and risk score (HR = 1.503, p < 0.001) were independent prognostic indicators of CM (Fig. 3B). The ROC curve showed that the AUC of risk score was 0.670, indicating that the PARGs prognostic signature were satisfactory stability (Fig. 3C).
Results of independent prognostic analysis and clinical correlation analysis of risk models based on prognostic characteristics of PARGs. Results of (A) univariate Cox regression analysis and (B) multivariate Cox regression analysis based on risk score, clinical characteristics with OS. HR = 1: No impact; HR < 1: reduced risk, positively correlated with survival time; HR > 1: Increased risk, negatively correlated with survival time. (C) ROC results for predictive accuracy of risk score and clinical characteristics. Correlation analysis based on PARGs prognostic characteristics with different clinical characteristics: (D) Age ≤ 65; (E) Male; (F) Female; (G) M 0; (H) T I-II; (I) T III-IV; (J) N 0–1; (K) Stage I-II. *p < 0.05; **p < 0.01; ***p < 0.001.
To further assess the prognostic value of the PARGs prognostic signature under different clinical conditions, we grouped CM patients according to their clinical characteristics. Based on the PARGs prognostic signature, CM patients in the same clinical conditions were divided into low-risk and high-risk groups before Kaplan–Meier survival analysis. As shown in Fig. 3D–K, in some clinical conditions such as female, male, age ≤ 65 years, M0, T0-2, T3-4, N0-1 and Stage1-2 in the low-risk score of patients had significantly higher OS rates than those in the high-risk group. In contrast, OS rates were similar in some CM patients with clinical conditions of age > 65 years, M1, N2-3, and Stage3-4 (data not shown).
A novel nomogram model was subsequently developed to accurately predict the 1-, 3-, and 5-year OS rates of CM patients based on the PARGs prognostic signature and clinicopathological features. As shown in Fig. 4A, there are corresponding scoring criteria for each feature, by scoring each feature, the total score of the sample can be obtained, and the OS can be obtained according to the total score. The calibration curves showed that the 1-, 3-, and 5-year survival times predicted by nomogram showed satisfactory agreement with the actual OS rates of CM patients (Fig. 4B). In addition, the nomogram had a higher concordance index (C-index) and decision curve (Fig. 4C–E) compared to other clinical factors. The time-dependent ROC showed AUCs of 0.672, 0.655, and 0.685 for 1, 3, and 5 years, respectively (Fig. 4F). In conclusion, these results suggest that risk scores based on PARGs can accurately assess the prognosis of CM patients relative to clinicopathological characteristics.
Nomogram construction and prediction accuracy assessment. (A) Nomogram construction based on the PARGs prognostic signature and clinical characteristics. (B) Calibration curves comparing the accuracy of nomogram predicted OS with the actual OS. Concordance index of the PARGs prognostic signature and clinical characteristics (C) and decision curve plots (D). (E) ROC results for predictive accuracy of nomogram, risk score and clinical characteristics. (F) ROC curves based on nomogram predicted at 1, 3 and 5 years. *p < 0.05; **p < 0.01; ***p < 0.001.
Enrichment analysis of DEGs
KEGG enrichment and GO enrichment were used to investigate the potential molecular mechanisms of DEGs in low and high risk groups. Volcano plot results of DEGs showed that most DEGs were downregulated in the high risk group (Fig. 5A). KEGG analysis as shown in Fig. 5B,D,E and the result showed that Cytokine-cytokine receptor interaction, Antigen processing and presentation, etc. were significantly enriched in DEGs (Fig. 5D). GO enrichment analysis demonstrated a high correlation between DEGs and immunity (Fig. 5C,F). Lymphocyte mediated immunity, positive regulation of leukocyte activation, positive regulation of lymphocyte activation, activation of immune response were enriched in the BP fraction. Immunoglobulin complex, T cell receptor complex, circulating immunoglobulin complex, and immunoglobulin receptor complex were enriched in the CC fraction. Immunoglobulin complex, circulating, immunoglobulin receptor binding, immune receptor activity, cytokine receptor activity were enriched in the MF fraction. These findings suggest that immune-related signaling pathways may mediate the role of PARGs in CM tumorigenesis.
Results of the enrichment analysis of DEGs. (A) Volcano plot of DEGs. Bar-plot of the results of KEGG enrichment analysis (B) and GO enrichment analysis (C). (D) Circle-plot of the results of KEGG enrichment analysis. Bubble-plot of the results of KEGG enrichment analysis (E) and GO enrichment analysis (F). *p < 0.05; **p < 0.01; ***p < 0.001.
Assessment and correlation analysis of the immune landscape based on PARGs prognostic signature
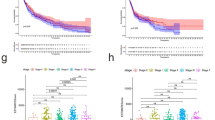
Multiple immune algorithms were used to assess the differences in immune infiltration landscape between patients in the low- and high-risk groups. Estimate results showed that patients in the low-risk group had lower tumor purity and higher stromal, immune, and estimate scores, while the opposite was true for patients in the high-risk group (Fig. 6A–D). CIBERSORT results showed that B cells naive, Plasma cells, T cells CD8, T cells CD4 memory activated , T cells follicular helper, T cells regulatory (Tregs), and Macrophages M1 were higher in the low-risk patient (Fig. 6E). In contrast, Macrophages M0, Macrophages M2, and Mast cells resting were higher in the high-risk patient population (Fig. 6E). The ssGSEA algorithm results showed that among all 23 immune cell species, except CD56 bright natural killer cellna did not differ in proportion in all patients, the other 21 immune cell were higher in the low-risk patient group than in the high-risk patient group (Fig. 6F). Correlation analysis was further performed to explore the association between prognostic related PARGs and immune infiltration landscape. The results showed a significant association between prognostic related PARGs and immune cells (Fig. 6G,H). As shown in Fig. 6G, T cells CD8, Plasma cells, T cells CD4 memory activated and T cells regulatory (Tregs) were positively correlated with ZBP1, while Macrophages M0, Macrophages M2 and NK cells activated were negatively correlated with ZBP1. Notably, for all 23 immune cells based on the ssGSEA algorithm, ZBP1 was positively correlated with most immune cells (Fig. 6H). The above results suggested significant differences in immune infiltration in CM patients, and we further evaluated patients’ immune function scores, the expression of key tumor genes, and the response to immunotherapy. The results showed that the low-risk patients exhibited higher immune function scores (Fig. 7A). The expression of key tumor genes was higher in all low-risk patients than in high-risk patients (Fig. 7B). Similarly, low-risk patients showed a better response regardless of the presence of CTAL4 or PD1 (Fig. 7C–F). And the TIDE results showed that patients in the low-risk group had higher TIDE scores, indicating a more pronounced response to immunotherapy in the low-risk group (Fig. 7G). In conclusion, these results suggest that a risk model based on the PARGs prognostic signature correlates with the immune infiltration status and immunotherapy response in CM patients.
The assessment and correlation analysis result of the immune landscape based on PARGs prognostic signature. (A–D) Tumor purity results as well as immune, stromal and ESTIMATE scores by ESTIMATE algorithm. (E) Proportion of immune cells of type 22 in CM patients of different risk groups by CIBERSORT. (F) Proportion of immune cells of type 23 in CM patients of different risk groups by ssGSEA. (G, H) Correlation results of prognosis-related PARGs with immune cells. *p < 0.05; **p < 0.01; ***p < 0.001.
The result of immune function and immunotherapy in CM patients with different risks. (A) Differential results of immune function in CM patients with different risks. (B) Results of immune checkpoint analysis in CM patients with different risks. (C–F) Results of immunotherapy response differences in CM patients with different risks. (G) TIDE score of different risk patients. *p < 0.05; **p < 0.01; ***p < 0.001.
Drug susceptibility analysis
Antineoplastic drugs remain one of the main clinical treatments for CM. Due to the heterogeneity of CM, the sensitivity of the same drug varies for different patients, so understanding the differences in the sensitivity of antineoplastic drugs in CM patients with different risks can provide new suggestions for personalized treatment of clinical. As shown in Fig. 8, the IC50 of most antineoplastic drugs in low-risk CM patients is lower than that in high-risk CM patients, such as Paclitaxel, Dasatinib, Imatinib and Lapatinib. Notably, the IC50 of Sorafenib was higher in low-risk CM patients than in high-risk CM patients, suggesting that Sorafenib may be more effective when used in high-risk CM patients. The above results suggest that CM patients with different risks have different sensitivity to antineoplastic drugs, and the prediction of drug sensitivity in CM patients with different risks can provide new thinking for personalized treatment of patients.
Differential analysis of drug sensitivity in CM patients with different risk groups. (A) Paclitaxel, (B) Bleomycin, (C) 5-Fluorouracil, (D) Sorafenib, (E) Sunitinib, (F) Lapatinib, (G) Ruxolitinib, (H) Saracatinib, (I) Pazopanib, (J) Imatinib, (K) Dasatinib and (L) Crizotinib. *p < 0.05; **p < 0.01; ***p < 0.001.
Consensus clustering analysis and immune evaluation
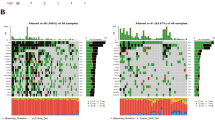
Consensus cluster analysis was used to classify CM patients into different subgroups according to the 3 prognostic related PARGs. The heat map showed that the best classification of CM patients was K = 2 with 125 samples in Cluster A and 329 samples in Cluster B (Fig. 9A). Kaplan–Meier survival analysis showed that Cluster B had a higher OS compared to Cluster A (Fig. 9B). ESTIMATE results showed that Cluster A had higher ESTIMATE, immune score, but lower tumor purity (Fig. 9C-F). Meanwhile, the CIBERSORT algorithm and ssGSEA were applied to each subgroup. As shown in Fig. 9G, T cells CD8, T cells CD4 memory activated, NK cells activated and Macrophages M1 were higher in Cluster A, while T cells CD4 memory resting, Macrophages M0 and Macrophages M2 were higher in Cluster B. The results of ssGSEA showed that the proportion of most immune cells was significantly higher in patients in Cluster A (Fig. 9H). In addition, the low-risk patients exhibited higher immune function scores (Fig. 10A). The TIDE results showed that patients in Cluster B had lower TIDE scores, which suggests that CM in Cluster B patients had a better potential immune therapy response (Fig. 10B). Similarly, low-risk patients showed a better response regardless of the presence of CTAL4 or PD1 (Fig. 10C–F). In conclusion, these results further suggest that PARGs are associated with prognosis and may indicate the immune response and immune infiltration status of CM.
Consensus clustering of CM patients and immune landscape analysis. (A) Consensus clustering heat map showing the best classification of CM patients with K = 2. (B) The result of Kaplan–Meier survival curve in different subgroups. (C–F) Tumor purity results as well as immune, stromal and ESTIMATE scores by ESTIMATE algorithm. (G) Proportion of immune cells of type 22 in CM patients of different subgroups by CIBERSORT. (H) Proportion of immune cells of type 23 in CM patients of different subgroups by ssGSEA. *p < 0.05; **p < 0.01; ***p < 0.001.
The result of immune function and immunotherapy in CM patients with different subgroups. (A) Differential results of immune function in CM patients with different subgroups. (B) TIDE score of different subgroups patients. (C–F) Results of immunotherapy response differences in CM patients with different subgroups. *p < 0.05; **p < 0.01; ***p < 0.001.
Discussion
As one of the most aggressive and metastatic malignancies, the OS of CM patients remains suboptimal24. PANoptosis is a recently defined inflammatory programmed cell death process involving multiple apoptotic pathways such as pyroptosis, apoptosis, and necroptosis10,11,12,13,14,25,26. A growing number of studies suggest that PANoptosis is associated with tumor development and metastasis8,27. However, the link between PANoptosis and CM is poorly studied, especially the exact mechanism of PARGs in CM is unclear. Therefore, we aimed to link PANoptosis to the prognosis of CM patients to reveal the role of PANoptosis and PARGs in CM. The study aims to investigate the link between PANoptosis and the prognosis of CM patients. This research could provide insights into the underlying mechanisms of CM progression and potentially lead to the development of new therapeutic strategies for CM.
In our study, ZBP1, MAP3K7 and RBCK1 were finally identified to be associated with OS in CM and could be used as reliable prognostic biomarkers for CM. A risk model was constructed based on these three prognosis-related PARGs, and the prediction accuracy of the model was better compared to other clinical characteristics. Notably, low-risk CM patients had better OS compared to high-risk CM patients. We subsequently constructed nomogram based on clinical characteristics as well as the risk model, which can be used for prediction of OS in CM patients.
The immune environment is an important factor in the interaction between tumor cells and immunity, and immune escape of tumor cells is often directly related to the effect of immunotherapy. In this study by enrichment analysis of DEGs in CM patients with different risk, we found that most immune-related pathways and biological functions were enriched, indicating that CM is very relevant to immunity. We found that the low-risk group had higher immune scores and a higher degree of immune cell infiltration by assessing the immune landscape of CM patients with different risk. The activation of the interferon signaling pathway has been shown to induce the expression of chemokines that attract T cells and NK cells, as well as the upregulation of major histocompatibility complex (MHC) molecules that present tumor antigens to T cells28. Therefore, the upregulation of interferon signaling pathway genes in the low-risk group may contribute to the higher infiltration of T cells CD8 and NK cells observed in these patients. In addition, the activation of the Wnt/β-catenin pathway has been associated with an immunosuppressive microenvironment characterized by the recruitment of myeloid-derived suppressor cells (MDSCs) and the inhibition of T cell activation29. Therefore, the upregulation of Wnt/β-catenin pathway genes in the high-risk group may contribute to the higher infiltration of Macrophages M0 and M2 observed in these patients. Notably, the prognosis-related PARGs we identified were significantly associated with immune infiltration. High expression of ZBP1, MAP3K7 and RBCK1 was accompanied by a higher degree of immune infiltration, suggesting that PANoptosis plays a key role in the immune landscape of CM.
Z-DNA binding protein 1 (ZBP1) was identified for its role in inducing tumor-associated proteins30. However, recent studies have shown that ZBP1 can function as a central regulator of multiple programmed death pathways31,32. Rajendra Karki showed that ZBP1 can mediate NLRP3 inflammatory vesicle activation and PANoptosis33. Yuanqin Yang found that ZBP1 can control anti-tumor immune responses through mixed-spectrum kinase structural domain-like pseudokinase (MLKL)34. Caspases regulate cell death, immune responses, and homeostasis, and Min Zheng found that caspase-6 can act by promoting ZBP1-mediated inflammatory vesicle activation and cell death10. In conjunction with our study, it is clear that ZBP1 has a very important association with immunity.
Mitogen-activated protein kinase kinase kinase 7 (MAP3K7) encodes transforming growth factor β-activated kinase 1 (TAK1) and can regulate the immune response, cell death and carcinogenesis35. Inokuchi S found that TAK1 deficiency can lead to cellular injury, inflammation, fibrosis and carcinogenesis36. More notably, TAK1 has a key role in differentiation and function of macrophage37. In conjunction with our study, the absence of MAP3K7 in the high-risk group may count for the worse immune environment.
Gamma interferon (IFN-γ) is a key driver of immune activation and immunosuppression38. RBCK1, a gene related to IFN-γ signaling, is associated with tumor immune infiltration as well as immunotherapy39,40. Martin Krenn found that N-terminal mutations in RBCK1 may lead to immune dysfunction41. RBCK1 is also able to promote NFKB signaling in the immune response through the linear ubiquitin assembly complex (LUBAC) to facilitate NFKB signaling in the immune response42. In our study, RBCK1 was highly associated with infiltration of many immune cells.The effect of RBCK1 on immune function may be responsible for the differences in sensitivity to immunotherapy and drug treatment in CM patients with different risks.
Overall, our study shows that PANoptosis plays a crucial role in CM. The risk models based on prognosis-related PARGs not only can effectively predict the CM patients prognosis but also indicate the immune landscape of CM patients. Understanding the role of PANoptosis and PARGs in CM could provide new avenues for targeted therapy and personalized treatment options for CM patients. This research could have a significant impact on improving the prognosis and survival rates of CM patients.
Data availability
The datasets involved in this study were obtained from public databases (TCGA, https://portal.gdc.cancer.gov/). And the raw data presented in the study are included in the article or supplementary material, further inquiries can be addressed to the respective authors.
References
Leonardi, G. C. et al. Cutaneous melanoma: From pathogenesis to therapy (review). Int. J. Oncol. 52, 1071–1080. https://doi.org/10.3892/ijo.2018.4287 (2018).
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Arheden, A. et al. Real-world data on PD-1 inhibitor therapy in metastatic melanoma. Acta Oncol. 58, 962–966. https://doi.org/10.1080/0284186X.2019.1620966 (2019).
Mar, V. J. et al. Diagnosis and management of cutaneous melanoma. Aust. J. Gen. Pract. 49, 733–739. https://doi.org/10.31128/AJGP-02-20-5238 (2020).
Lee, J. J. & Lian, C. G. Molecular testing for cutaneous melanoma: An update and review. Arch. Pathol. Lab. Med. 143, 811–820. https://doi.org/10.5858/arpa.2018-0038-RA (2019).
Mohammad, R. M. et al. Broad targeting of resistance to apoptosis in cancer. Semin. Cancer Biol. 35(Suppl), S78–S103. https://doi.org/10.1016/j.semcancer.2015.03.001 (2015).
Malireddi, R. K. S., Tweedell, R. E. & Kanneganti, T. D. PANoptosis components, regulation, and implications. Aging 12, 11163–11164. https://doi.org/10.18632/aging.103528 (2020).
Jiang, M. et al. Caspase-8: A key protein of cross-talk signal way in “PANoptosis” in cancer. Int. J. Cancer 149, 1408–1420. https://doi.org/10.1002/ijc.33698 (2021).
Samir, P., Malireddi, R. K. S. & Kanneganti, T. D. The PANoptosome: A deadly protein complex driving pyroptosis, apoptosis, and necroptosis (PANoptosis). Front. Cell. Infect. Microbiol. 10, 238. https://doi.org/10.3389/fcimb.2020.00238 (2020).
Zheng, M., Karki, R., Vogel, P. & Kanneganti, T. D. Caspase-6 is a key regulator of innate immunity, inflammasome activation, and host defense. Cell 181, 674–687. https://doi.org/10.1016/j.cell.2020.03.040 (2020).
Malireddi, R. K. S. et al. Innate immune priming in the absence of TAK1 drives RIPK1 kinase activity-independent pyroptosis, apoptosis, necroptosis, and inflammatory disease. J. Exp. Med. https://doi.org/10.1084/jem.20191644 (2020).
Lalaoui, N. et al. Mutations that prevent caspase cleavage of RIPK1 cause autoinflammatory disease. Nature 577, 103–108. https://doi.org/10.1038/s41586-019-1828-5 (2020).
Rickard, J. A. et al. RIPK1 regulates RIPK3-MLKL-driven systemic inflammation and emergency hematopoiesis. Cell 157, 1175–1188. https://doi.org/10.1016/j.cell.2014.04.019 (2014).
Dondelinger, Y. et al. Serine 25 phosphorylation inhibits RIPK1 kinase-dependent cell death in models of infection and inflammation. Nat. Commun. 10, 1729. https://doi.org/10.1038/s41467-019-09690-0 (2019).
Huang, J. et al. Analysis of PANoptosis-related LncRNA-miRNA-mRNA network reveals LncRNA SNHG7 involved in chemo-resistance in colon adenocarcinoma. Front. Oncol. 12, 888105. https://doi.org/10.3389/fonc.2022.888105 (2022).
Pan, H., Pan, J., Li, P. & Gao, J. Characterization of PANoptosis patterns predicts survival and immunotherapy response in gastric cancer. Clin. Immunol. 238, 109019. https://doi.org/10.1016/j.clim.2022.109019 (2022).
Ren, L. et al. CDK1 serves as a therapeutic target of adrenocortical carcinoma via regulating epithelial-mesenchymal transition, G2/M phase transition, and PANoptosis. J. Transl. Med. 20, 444. https://doi.org/10.1186/s12967-022-03641-y (2022).
Hussein, M. R., Haemel, A. K. & Wood, G. S. Apoptosis and melanoma: Molecular mechanisms. J. Pathol. 199, 275–288. https://doi.org/10.1002/path.1300 (2003).
Erb, P., Ji, J., Kump, E., Mielgo, A. & Wernli, M. Apoptosis and pathogenesis of melanoma and nonmelanoma skin cancer. Adv. Exp. Med. Biol. 624, 283–295. https://doi.org/10.1007/978-0-387-77574-6_22 (2008).
Liu, Y. et al. AC010973.2 promotes cell proliferation and is one of six stemness-related genes that predict overall survival of renal clear cell carcinoma. Sci. Rep. 12, 4272. https://doi.org/10.1038/s41598-022-07070-1 (2022).
Yu, L. et al. Multi-omics analysis reveals the interaction between the complement system and the coagulation cascade in the development of endometriosis. Sci. Rep. 11, 11926. https://doi.org/10.1038/s41598-021-90112-x (2021).
Wu, D. et al. Identification of novel autophagy-related lncRNAs associated with a poor prognosis of colon adenocarcinoma through bioinformatics analysis. Sci. Rep. 11, 8069. https://doi.org/10.1038/s41598-021-87540-0 (2021).
Zhang, D. et al. Prognostic role of DNA damage response genes mutations and their association with the sensitivity of olaparib in prostate cancer patients. Cancer Control J. Moffitt Cancer Center 29, 10732748221129452. https://doi.org/10.1177/10732748221129451 (2022).
Eggermont, A. M., Spatz, A. & Robert, C. Cutaneous melanoma. Lancet 383, 816–827. https://doi.org/10.1016/S0140-6736(13)60802-8 (2014).
Lee, S. et al. AIM2 forms a complex with pyrin and ZBP1 to drive PANoptosis and host defence. Nature 597, 415–419. https://doi.org/10.1038/s41586-021-03875-8 (2021).
Gullett, J. M., Tweedell, R. E. & Kanneganti, T. D. It’s all in the PAN: Crosstalk, plasticity, redundancies, switches, and interconnectedness encompassed by PANoptosis underlying the totality of cell death-associated biological effects. Cells. https://doi.org/10.3390/cells11091495 (2022).
Karki, R. et al. Interferon regulatory factor 1 regulates PANoptosis to prevent colorectal cancer. JCI Insight. https://doi.org/10.1172/jci.insight.136720 (2020).
Glassman, C. R. et al. Structural basis for IL-12 and IL-23 receptor sharing reveals a gateway for shaping actions on T versus NK cells. Cell 184, 983–999. https://doi.org/10.1016/j.cell.2021.01.018 (2021).
Deng, H. et al. Translocation of Helicobacter hepaticus synergizes with myeloid-derived suppressor cells and contributes to breast carcinogenesis. Oncoimmunology 11, 2057399. https://doi.org/10.1080/2162402X.2022.2057399 (2022).
Fu, Y. et al. Cloning of DLM-1, a novel gene that is up-regulated in activated macrophages, using RNA differential display. Gene 240, 157–163. https://doi.org/10.1016/s0378-1119(99)00419-9 (1999).
Thapa, R. J. et al. DAI senses influenza A virus genomic RNA and activates RIPK3-dependent cell death. Cell Host Microbe 20, 674–681. https://doi.org/10.1016/j.chom.2016.09.014 (2016).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aag2045 (2016).
Karki, R. et al. ADAR1 restricts ZBP1-mediated immune response and PANoptosis to promote tumorigenesis. Cell Rep. 37, 109858. https://doi.org/10.1016/j.celrep.2021.109858 (2021).
Yang, Y. et al. ZBP1-MLKL necroptotic signaling potentiates radiation-induced antitumor immunity via intratumoral STING pathway activation. Sci. Adv. 7, eabf6290. https://doi.org/10.1126/sciadv.abf6290 (2021).
Tan, S. et al. Hepatocyte-specific TAK1 deficiency drives RIPK1 kinase-dependent inflammation to promote liver fibrosis and hepatocellular carcinoma. Proc. Natl. Acad. Sci. U.S.A. 117, 14231–14242. https://doi.org/10.1073/pnas.2005353117 (2020).
Inokuchi, S. et al. Disruption of TAK1 in hepatocytes causes hepatic injury, inflammation, fibrosis, and carcinogenesis. Proc. Natl. Acad. Sci. U.S.A. 107, 844–849. https://doi.org/10.1073/pnas.0909781107 (2010).
Richter, E., Ventz, K., Harms, M., Mostertz, J. & Hochgrafe, F. Induction of macrophage function in human THP-1 cells is associated with rewiring of MAPK signaling and activation of MAP3K7 (TAK1) protein kinase. Front. Cell Dev. Biol. 4, 21. https://doi.org/10.3389/fcell.2016.00021 (2016).
Kim, E. Y. et al. Post-sepsis immunosuppression depends on NKT cell regulation of mTOR/IFN-gamma in NK cells. J. Clin. Investig. 130, 3238–3252. https://doi.org/10.1172/JCI128075 (2020).
Xu, W. et al. Comprehensive multi-omics identification of interferon-gamma response characteristics reveals that RBCK1 regulates the immunosuppressive microenvironment of renal cell carcinoma. Front. Immunol. 12, 734646. https://doi.org/10.3389/fimmu.2021.734646 (2021).
Yu, S. et al. RBCK1 promotes p53 degradation via ubiquitination in renal cell carcinoma. Cell Death Dis. 10, 254. https://doi.org/10.1038/s41419-019-1488-2 (2019).
Krenn, M. et al. Mutations outside the N-terminal part of RBCK1 may cause polyglucosan body myopathy with immunological dysfunction: Expanding the genotype-phenotype spectrum. J. Neurol. 265, 394–401. https://doi.org/10.1007/s00415-017-8710-x (2018).
Li, Z. et al. RBCK1 is an endogenous inhibitor for triple negative breast cancer via hippo/YAP axis. Cell Commun. Signal 20, 164. https://doi.org/10.1186/s12964-022-00963-8 (2022).
Funding
This work was supported by the Zhejiang Provincial Natural Science Foundation of China (LYY21H300001), Zhejiang Provincial Health Science and Technology Program (2021KY906) and Construction Fund of Medical Key Disciplines of Hangzhou.
Author information
Authors and Affiliations
Contributions
W.W. and X.X. designed the study. W.W. and Z.Q. collected the data which involved in this study. W.W. and L.L. analyzed the data and visualized the results. W.W. and Z.Q. wrote this manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, W., Zhou, Q., Lan, L. et al. PANoptosis-related prognostic signature predicts overall survival of cutaneous melanoma and provides insights into immune infiltration landscape. Sci Rep 13, 8449 (2023). https://doi.org/10.1038/s41598-023-35462-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-35462-4
- Springer Nature Limited
This article is cited by
-
Latest advances in hepatocellular carcinoma management and prevention through advanced technologies
Egyptian Liver Journal (2024)