Abstract
New information on the intensive care applications of new generation ‘high-density data clinical information systems’ (HDDCIS) is increasingly being published in the academic literature. HDDCIS avoid data loss from bedside equipment and some provide vital signs statistical calculations to promote quick and easy evaluation of patient information. Our objective was to study whether manual records of continuously monitored vital signs in the Paediatric Intensive Care Unit could be replaced by these statistical calculations. Here we conducted a prospective observational clinical study in paediatric patients with severe diabetic ketoacidosis, using a Medlinecare® HDDCIS, which collects information from bedside equipment (1 data point per parameter, every 3–5 s) and automatically provides hourly statistical calculations of the central trend and sample dispersion. These calculations were compared with manual hourly nursing records for patient heart and respiratory rates and oxygen saturation. The central tendency calculations showed identical or remarkably similar values and strong correlations with manual nursing records. The sample dispersion calculations differed from the manual references and showed weaker correlations. We concluded that vital signs calculations of central tendency can replace manual records, thereby reducing the bureaucratic burden of staff. The significant sample dispersion calculations variability revealed that automatic random measurements must be supervised by healthcare personnel, making them inefficient.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
There is a broad consensus that future healthcare will require the intensive use of information technology to acquire, store, process, analyse, and use the information extracted from medical data1,2. The Anaesthesia and Critical Care specialities are the most technical and data-driven medical environments and so the development of new approaches for the integration and use of the data generated in these fields is almost mandatory1,2,3,4,5,6,7. However, the basic approach to data collection and management has remained largely unchanged over the past 40 years2. Indeed, it is now becoming especially difficult to integrate and take advantage of the information provided by patient bedside monitoring and treatment devices2. In critical patients, this information is traditionally registered by nursing staff, who collect the most representative values from each period, usually once an hour8. This chart is used as the baseline by which patient evolution is then assessed9.
Given that these patient monitoring charts are costly to prepare, some centres have replaced this type of record keeping with automatic random data collection by different clinical information systems (CIS)2,8. These CIS usually collect one data point per parameter every 15, 30, or 60 min8,10. This means that more than 99% of the information generated at the bedside is lost with no possibility reaping the benefits that its exploitation could mean for patients and the healthcare system2,4,11,12. In addition, CIS do not provide any processing or analysis of the information obtained2. Thus, although some of these systems provide access to up to 1 piece of data per minute, the volume of data makes its routine manual evaluation by healthcare personnel difficult13 and most of the information is still lost. Because of all the above, Intensive Care Unit (ICU) teams continue to express frustration with the current patient data representation by CIS14.
Therefore, a new approach to the use of information generated at the bedside is required to prevent this data loss and to facilitate its clinical use4,8,11. The main objective of this new approach is to apply big-data principles (volume, velocity, variety, veracity, and data value) to the concept of personalised medicine. In this sense, information has already been published about the application of high-density data CIS (HDDCIS) in intensive care contexts, both in development studies2,3,13,15,16,17 and as original research4,8,12,18,19,20,21,22,23. Although the use of HDDCIS is not yet common in ICUs, in the future they will help revolutionise the monitoring of vital signs as we currently know it. In this context, Matam et al. analysed the technical difficulties and feasibility of adapting HDDCIS to their Paediatric Intensive Care Unit (PICU) and created a machine learning system that predicts cardiac arrest in children15,24. In turn, Brossier et al. argued for the importance of these systems being able to indefinitely store all the monitoring data for ‘perpetual patients’ as well as their advantages for the development of clinical decision support systems (CDSS)4,25,26.
Furthermore, the development of artificial intelligence (AI) is likely to strongly impact intensive care medicine. Indeed, one of the main lines of ongoing research for many groups working in this field involves the use of AI to give HDDCIS the ability to detect clinically important events. For example, in the field of mechanical ventilation, systems that predict accidental extubation27 or that protect the lungs by avoiding volutrauma28 are already available. The detection of events in the operating room during anaesthesia has also been analysed by leveraging the information contained in databases that integrate information from different clinical records such as electrocardiograms, oxygen saturation, heart rate, and bispectral index results29,30. Thus, the integration of vital signs records has already allowed the creation of machine learning systems capable of detecting, in real time, events such as acute hypotension31, responses to vasoactive drugs32, or the need for massive transfusion in the operating room33. In most cases, the AI technology used to detect events is based on the registration of vital signs data and leverages pre-defined scores. Among these indices, it is worth highlighting the Inadequate Oxygen Delivery Index (IDO2-Index) that warns of the risk of adverse events such as cardiorespiratory arrest, the development of enterocolitis, need for extracorporeal membrane oxygenation (ECMO), or renal replacement therapy in children undergoing cardiac surgery34.
In this line, our team has several years of clinical experience with the use of this technology35 and we have been able to verify the clinical usefulness of the hourly statistical calculations of vital signs provided by the HDDCIS6. However, the use of these calculations has not yet been validated. Thus, the objective of this current work was to study whether manual records of continuously monitored vital signs can be replaced by the statistical calculations of central tendency provided by an HDDCIS.
Materials and methods
This prospective observational clinical study was conducted in children with severe diabetic ketoacidosis consecutively admitted to the PICU at the University Clinical Hospital of Valencia (a 6-bed multivalent unit in a tertiary hospital), between 2017 and 2020. The inclusion criteria were (1) blood glucose exceeding 11.1 mmol/L; (2) ketonemia greater than 1 mmol/L; (3) bicarbonate less than 8 mEq/L; and (4) base excess exceeding − 20 mEq/L. We selected these patients for their special clinical characteristics. Upon admission, they presented a serious metabolic alteration that affected their vital signs but without associated respiratory, cardiocirculatory, or other pathologies or confounding factors such as the requirement for mechanical ventilation, administration of inotropic drugs, sedatives, or analgesics, among others9,22. Thus, with appropriate treatment, their vital signs returned to normal within hours, therefore making them good models to assess clinical evolutionary changes and to compare different study parameters.
This study was conducted in accordance with the amended Declaration of Helsinki and was approved by the Ethics Committee at the Biomedical Research Institute INCLIVA (grant number 2017/022). Informed consent was obtained from the families of the patients. We used the Medlinecare® HDDCIS (Medical Online Technology S.L., Valencia, Spain) which receives and transmits information from bedside monitoring and treatment equipment at a rate of 1 data point per monitored parameter every 3–5 s, (720–1200 data points per hour), including equipment from multiple manufacturers. This allowed us to continuously monitor the clinical status of our patients in real time. In addition, the HDDCIS stores data and automatically calculates hourly statistical indicators of the sample central trends (mean, mode, and median) and dispersion as the maximum (99th percentile), and minimum (1st percentile) values. These calculations were performed in the first few minutes of each new hour.
The data collected during patient admissions to the PICU are shown in Table 1. The nursing staff did not have access to the information provided by the HDDCIS or knowledge of the objective of this current study. We used pulse oximetry technology from Masimo Corp. (Irvine, CA) and collected oxygen saturation (SpO2) and pulse oximetry heart rate (pHR) data using disposable fingertip probes. We also collected electrocardiography heart rate (eHR) and respiratory rate data measured by impedance (iRR) using Infinity Delta XL multiparameter monitors from Dräger Medical (Lübeck, Germany). To uncover whether the clinical evolution of the patients could affect the correlation study of the parameters, the data series were divided into two groups according to the level of bicarbonate present in the blood of the patients: the severe acidosis group versus the improvement group, with the bicarbonate cut-off point being ≥ 10 meq/L.
Statistical analysis
SPSS software (v26.0 IBM Corp., Armonk, NY) was used to carry out the statistical analyses. The relationships between continuous variables were evaluated employing the Pearson or Spearman correlation coefficient, depending on the data distribution (after assessing the latter using the Kolmogorov–Smirnov test). We also used Student t-tests for related samples, Lin’s concordance correlation coefficient (CCC) to assess the data agreement36, and Bland–Altman plots for multiple measurements per patient, as calculated with MedCalc software37. Analysis of variance (ANOVA) was used to study the relationship between the continuous variables of each group. However, when statistically significant results failed to meet the ANOVA assumptions, we resorted to an alternative robust test (Welch’s test). The significance level threshold was set at an alpha of 0.05 in all cases.
Results
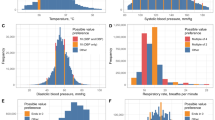
Thirty-two consecutive patients (21 boys and 11 girls) aged 9.1 ± 4.3 years were included in this study cohort. Around 2,761,000 measurements were used to obtain 1027 hourly statistical calculations for pHR, eHR, and SpO2, as well as 745 calculations of iRR. Simultaneous hourly nursing records (denoted with the ‘n’ prefix) were also collected for the heart rate (nHR = 1025), respiratory rate (nRR = 716), and oxygen saturation (nSpO2 = 980). In addition, 212 periodic determinations of glycaemia, ketonemia, lactate, and acid–base balance were also collected. Upon admission, the patients presented the following blood analytical test data: blood glucose = 25.8 ± 7.6 mmol/L, ketonemia = 5.3 ± 1.7 mmol/L, lactate = 2.2 ± 0.9 mmol/L, pH = 7.05 ± 0.1, PCO2 = 19.6 ± 7.5 mmHg, bicarbonate = 5.6 ± 2.9 mEq/L, and base excess = − 24.7 ± 4.7 mEq/L.
The hourly calculations of the central tendency for pHR and eHR were identical and their CCC was almost perfect (Fig. 1). These calculations showed identical values, very strong correlations, and a substantial CCC with the nHR data (Table 2). Indeed, both the nHR and the central tendency calculations for pHR and eHR showed the same moderately significant correlations with the evolution of the acid–base balance. For example, for pH, r = − 0.51 for nHR and r = − 0.5 for pHR, for PCO2, r = − 0.53 and − 0.55, respectively, for bicarbonate r = − 0.55 and − 0.55, respectively, and for base excess, r = − 0.56 and − 0.56, respectively.
Scatter diagram of the automatic hourly calculation of the ‘median heart rate’ obtained by pulse oximetry (pHR-median) and electrocardiography (eHR-median). Note how the line of equality and the trend line (blue) are identical (locally weighted scatterplot smoothing span = 0%; concordance correlation coefficient [CCC] = 0.9983; Pearson p for precision = 0.9984; bias correction factor for accuracy = 1 [95% CI 0.9982–0.9986]; p < 0.0001). Similar results were obtained for the central tendency calculations for the ‘mean heart rate’ and ‘modal heart rate’ (CCC > 0.99).
The central tendency calculations for the iRR and SpO2 behaved in a similar way, with identical or remarkably similar values and strong correlations with the nursing staff references (except for the mean iRR, which showed a moderate correlation). The maximum and minimum hourly calculations differed from their nursing references and from the central tendency calculations in all the variables by showing weaker correlations. These differences were clinically significant for heart rate (HR) and respiratory rate (RR). The concordance study results are shown in Tables 2 and 3 and in Fig. 2.
In the comparison by groups (severe acidosis versus improvement), blood glucose, ketonemia, lactate, and acid–base balance measurements all improved, reaching levels of clinically and statistically significant differences (p < 0.001). HR and RR were also significantly improved for all variables from both the clinical and statistical perspective, except for the maximum iRR (Tables 4 and 5). In contrast, SpO2 decreased with improving acidosis, although these differences did not reach the level of clinical significance (Table 4).
Discussion
The physiological basis and main characteristics of high-density data clinical information systems
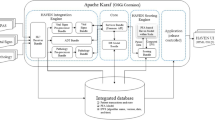
Drs. Horvat and Ogoe pointed out that “with the conventional approach, the patient data generated in the ICU are continually reduced to summary information, which has the risk of over-distilling the relevant and complex physiology of these patients”. Thus, these authors and others believe that the use of high-frequency data systems, “may further our understanding of intensive care physiology and eventually support the development of more individualised therapeutic regimens”5,9. As described in the introduction, this technology is already being applied in hospitals all over the world. More specifically, four basic properties characterise HDDCIS (Fig. 3) as follows:
-
1.
Sampling frequency. Although this can differ according to the parameters recorded, for vital signs and other fundamental parameters, this frequency must be less than 1 data point every 10 s. However, many current systems capture these values at intervals of minutes, which is insufficient to follow patient evolution in real time or to detect and evaluate clinical events.
-
2.
Multidevice capacity. Anaesthesia and critical care environments are complex by nature and are home to countless medical devices for patient monitoring and treatment. Thus, to include any parameter of clinical interest, it must be possible to capture data, at a high sampling frequency, from different pieces of medical equipment. To do this, the problem of device synchronisation must be solved.
-
3.
Information processing. This is important both in terms of real-time care functionality, as well as in the evaluation of clinical evolution based on historical data6. As Sun et al. stated, “we must develop a data acquisition system that facilitates the access and review of historical data for medical personnel. Furthermore, acquired data should be […] presented to clinical staff in such a manner that supports clinical decision making”13. Thus, improving this information processing will directly enhance care provider wellbeing, patient outcomes, and quality of care14. Moreover, HDDCIS are of great educational utility and can also help improve quality of care because they can fully detect clinical episodes as they happen, at high resolution, and can also reproduce them for later analysis (see Supplementary Figs. S1–S3).
-
4.
Ability to exploit information. These systems will provide medical staff with a powerful research tool which, by combining their clinical observations with supervised and unsupervised machine learning, can be used to develop and test CDSS and other AI functions4,38. However, the loss of information from the current CIS prevents the development of these functions. Moreover, it should be noted that, after the COVID-19 global pandemic, the development and use of AI has exponentially increased39,40 thanks to its freer availability and ability to integrate large amounts of information about an unknown disease and present it in a simple way to clinicians.
Our previous experience and justification of this study
In a previous study in ventilator-dependent patients hospitalised at home, we observed the usefulness of the HDDCIS for telemedical real-time patient assessment. Furthermore, the statistical indicators it provides are of great clinical use because they allow the quick and complete analysis of all the information during the morning telemedical rounds6,35. However, we were previously unable to validate its use given that specialised healthcare personnel were not available to perform monitoring in domestic settings. In our experience, the application of this technology in the fields of anaesthesiology and intensive care is proving to be extremely useful, especially in the most serious clinical cases, because the resulting monitoring of these patients is more complete and produces fewer artifacts caused by movements, disconnections, or sensor misplacements.
A possible limitation of this study was related to the gold standard used for comparison, given that inaccuracy in the manual recording of vital signs in hospitalised patients has been previously reported41,42. However, unlike the random automatic vital sign data-point collections carried out by the current CIS or collected during patient visits by nursing staff in conventional hospitalisation wards41, the anaesthesiologist in the operating room and nursing staff in ICUs continuously follow-up and assess patients. Thus, this information is of immense value as it is the most representative of the period in question. Therefore, these registries have formed the basis used to assess the evolution of patients over the last 40 years, thereby facilitating clinical communication and working well in countless clinical studies9.
Discussion of our results
In this current study the perfect correlation we found between the central tendency indicators of both heart rate measurements supports the use of this technology for the purposes set out in this study (Fig. 1). Thus, the values we obtained for manual nursing heart rate measurements (nHR) and the automatic calculations of central tendency (pHR and eHR) were identical and showed very strong correlations and concordances (Tables 2, 3, 4). In addition, the identical correlation of these parameters with the evolution of the acid–base balance—the main pathophysiological alteration present in these patients—indicates that all the clinical value of the nursing records was also captured by the automatic calculations of central tendency. These results indicate that these calculations could replace manual records, thus reducing the bureaucratic burden this task places on staff. Moreover, they allow reliable monitoring measurements to be obtained in settings where staff are not available, as we have already seen in our previous work in home-based settings6,35.
In a similar vein, the hourly central tendency calculations for iRR and SpO2 were analogous to those for HR, with good (but in this case, not exceptional) correlations with their nursing references. This meant that when the more demanding CCC test (comprising two metrics, one for precision and the other of accuracy) was done, poor values (< 0.9) were obtained and, for reasons intrinsic to the test, these could not exceed their own measure of precision (the Pearson r coefficient). These results can be explained in a different way for each parameter. For the iRR, it was related to the movement artifacts that affect this measurement, as discussed below. For SpO2, it was related to the low variability of this parameter (especially in these patients without oxygenation problems), which determined the low Pearson correlation coefficient values, regardless of the accuracy and overall concordance of these measurements43. Thus, the almost perfect values of its accuracy component (bias correction factor) indicate adequate concordance with the nursing reference values (Table 2).
The clinical evolution by group was as expected, with statistically significant data indicating patient progression towards clinical normality (Tables 4 and 5). Moreover, it was interesting to observe how the decrease in respiratory rate also conditioned a statistically significant decrease in SpO2, although this did not translate into clinical significance (Table 4). The logic of this observation is rooted in patient physiology; however, the accuracy of this evolution, reflected both by the nursing records and the hourly statistical calculations, was of far more relevance. Hence, taken together, these current findings also support the use of HDDCIS calculations.
The evolution of the RR was also interesting (Table 5). Movements in these patients, which were initially infrequent while they rested but later increased as they started to recover, likely caused artefacts to appear in the iRR measurements. This could explain the fact that the central tendency calculations initially showed strong Pearson correlations with the nursing reference data, which then became moderate correlations in the improvement group. Thus, these inaccuracies were attributable to the measurement method, not to the data processing. Moreover, this processing ensured that the mode and median still maintained a certain level of clinical utility. Nonetheless, given the importance of this variable, as with HR, we can simultaneously monitor RR using capnography and spirometry as well, thereby allowing us to isolate any discrepancies of this type. Thus, for example, increases only in RR measured by impedance are usually related to the patient's movements.
Finally, we can say that at their highest degree of accuracy, the manual records should match the automatic calculations of central tendency. Therefore, in the absence of major measurement artifacts, we consider that these automatic central tendency calculations should be used as the gold standard for assessing patient evolution.
Like the central tendency parameters, the clinical evolutionary assessment of the minimum (1st percentile) and maximum (99th percentile) hourly calculations was simple for the care staff and was based on the patient age and their clinical situation. Interestingly, despite the selection of these percentiles and the haemodynamic and respiratory stability of these patients, there were significant clinical differences in the HR and RR minimum and maximum hourly calculations with their corresponding nursing references (Fig. 2 and Tables 2, 3, 4, 5). Of note, similar variations were also described for the HR8. These findings indicate that automatic random sampling to assess the evolution of these important parameters can differ significantly according to when the sampling is conducted. This could explain, in part, the so-called ‘smoothing effect’. In other words, the trend toward normal or average physiology in the nursing and anaesthesia records41,42. Hence, in the operating room and ICU, this smoothing phenomenon could indicate just the opposite: the inaccuracy of random samples and their impaired ability to reflect the true clinical situation of patients. This therefore highlights the need to monitor and modify automatic random registrations4,8,9. Thus, given all the above, and in line with these authors, we also believe that random vital sign measurements require supervision, thereby making them inefficient.
Conclusions
The current CIS discard most of the information generated at the bedside and so the benefits that its storage, processing and exploitation could entail are lost. Furthermore, the random automatic data collection they perform must be supervised by healthcare personnel, making them inefficient. However, a new generation of HDDCIS now being used in anaesthesia and critical care medicine could overcome these limitations. These systems avoid data loss and improve data processing and integration to support the development of more personalised therapeutic regimens. Although more research is still needed to validate this potential for individualising therapeutics, our findings indicate that automatic hourly vital signs calculations of central tendency could replace manual anaesthesia and critical care records, thereby freeing up highly qualified staff for other more demanding tasks.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Vellido, A. et al. Machine learning for critical care: State-of-the-art and a sepsis case study. BioMed. Eng. OnLine. 135, 101–118 (2018).
De Georgia, M. A., Kaffashi, F., Jacono, F. J. & Loparo, K. A. Information technology in critical care: Review of monitoring and data acquisition systems for patient care and research. Sci. World J. 2015, 727694. https://doi.org/10.1155/2015/727694 (2015).
Goldstein, B. et al. Physiologic data acquisition system and database for the study of disease dynamics in the intensive care unit. Crit. Care Med. 31, 433–441 (2003).
Brossier, D. et al. Creating a high-frequency electronic database in the PICU: The perpetual patient. Pediatr. Crit. Care Med. 19, e189-198 (2018).
Vincent, J. L. The future of critical care medicine: Integration and personalization. Crit. Care Med. 44, 386–389 (2016).
Muñoz-Bonet, J. I. et al. Medical complications in a telemedicine home care programme for paediatric ventilated patients. J. Telemed. Telecare. 26, 462–473 (2020).
Mathews, S. C. & Pronovost, P. J. The need for systems integration in health care. JAMA 305, 934–935 (2011).
Eytan, D. et al. Temporal variability in the sampling of vital sign data limits the accuracy of patient state estimation. Pediatr. Crit. Care Med. 20, e333-341 (2019).
Horvat, C. M. & Ogoe, H. Variability in vital sign documentation as a barrier to modeling patient state: Why patient records may need more complexity!. Pediatr. Crit. Care Med. 20, 690–691 (2019).
Sauer, C. M. et al. Systematic review and comparison of publicly available ICU data sets—A decision guide for clinicians and data scientists. Crit. Care Med. 50, e581–e588 (2022).
Celi, L. A., Marshall, J. D., Lai, Y. & Stone, D. J. Disrupting electronic health records systems: The next generation. JMIR Med. Inform. 3, e4192. https://doi.org/10.2196/medinform.4192 (2015).
Malunjkar, S., Weber, S. & Datta, S. A highly scalable repository of waveform and vital signs data from bedside monitoring devices. ArXiv. 2021, 1–12 (2021).
Sun, Y. et al. INSMA: An integrated system for multimodal data acquisition and analysis in the intensive care unit. J. Biomed. Inform. 106, 103434. https://doi.org/10.1016/j.jbi.2020.103434 (2020).
Jalilian, L. & Khairat, S. The next-generation electronic health record in the ICU: A focus on user-technology interface to optimize patient safety and quality. Perspect. Health Inf. Manag. 19, 1 (2022).
Matam, B. R. & Duncan, H. Technical challenges related to implementation of a formula one real time data acquisition and analysis system in a paediatric intensive care unit. J. Clin. Monit. Comput. 32, 559–569 (2018).
Meyer, M. A. et al. A computerized perioperative data integration and display system. Int. J. Comput. Assist. Radiol. Surg. 2, 191–202 (2007).
Goodwin, A. J. et al. A practical approach to storage and retrieval of high-frequency physiological signals. Physiol. Meas. 41, 035008. https://doi.org/10.1088/1361-6579/ab7cb5 (2020).
Smielewski, P. et al. ICM+, a flexible platform for investigations of cerebrospinal dynamics in clinical practice. Acta Neurochir. Suppl. 102, 145–151 (2008).
Vinecore, K. et al. Design and implementation of a portable physiologic data acquisition system. Pediatr. Crit. Care Med. 8, 563–569 (2007).
Eytan, D. et al. Revisiting oxygen dissociation curves and bedside measured arterial saturation in critically ill children. Intensive Care Med. 45, 1832–1834 (2019).
Walsh, B. et al. Categorization in mechanically ventilated pediatric subjects: A proposed method to improve quality. Respir. Care. 61, 1168–1178 (2016).
Eytan, D., Goodwin, A. J., Greer, R., Guerguerian, A. M. & Laussen, P. C. Heart rate and blood pressure centile curves and distributions by age of hospitalized critically ill children. Front. Pediatr. 5(52), 1–8 (2017).
Walsh, B. K. et al. Daily goals formulation and enhanced visualization of mechanical ventilation variance improves mechanical ventilation score. Respir. Care. 62, 268–278 (2017).
Matam, B. R., Duncan, H. & Lowe, D. Machine learning based framework to predict cardiac arrests in a paediatric intensive care unit: Prediction of cardiac arrests. J. Clin. Monit. Comput. 33, 713–724 (2019).
Brossier, D. et al. Qualitative subjective assessment of a high-resolution database in a paediatric intensive care unit—Elaborating the perpetual patient’s ID card. J. Eval. Clin. Pract. 26, 86–91 (2020).
Fartoumi, S., Emeriaud, G., Roumeliotis, N., Brossier, D. & Sawan, M. Computerized decision support system for traumatic brain injury management. J. Pediatr. Intensive Care. 05, 101–107 (2015).
Hur, S. et al. Development and validation of unplanned extubation prediction models using intensive care unit data: Retrospective, comparative, machine learning study. J. Med. Internet. Res. 23, e23508. https://doi.org/10.2196/23508 (2021).
Hagan, R., Gillan, C. J., Spence, I., McAuley, D. & Shyamsundar, M. Comparing regression and neural network techniques for personalized predictive analytics to promote lung protective ventilation in Intensive Care Units. Comput. Biol. Med. 126, 104030 (2020).
Lee, H. C. et al. VitalDB, a high-fidelity multi-parameter vital signs database in surgical patients. Sci. Data. 9, 279. https://doi.org/10.1038/s41597-022-01411-5 (2022).
Vistisen, S. T., Pollard, T. J., Enevoldsen, J. & Scheeren, T. W. L. VitalDB: Fostering collaboration in anaesthesia research. Br. J. Anaesth. 127, 184–187 (2021).
Wijnberge, M. et al. Effect of a machine learning-derived early warning system for intraoperative hypotension vs standard care on depth and duration of intraoperative hypotension during elective noncardiac surgery: The HYPE randomized clinical trial. JAMA 323, 1052–1060 (2020).
Ross, C. E. et al. Physiologic response to pre-arrest bolus dilute epinephrine in the pediatric intensive care unit. Resuscitation 126, 137–142 (2018).
Lee, S. M. et al. Development and validation of a prediction model for need for massive transfusion during surgery using intraoperative hemodynamic monitoring data. JAMA Netw. Open. 5, e2246637. https://doi.org/10.1001/jamanetworkopen.2022.46637 (2022).
Rogers, L. et al. The inadequate oxygen delivery index and low cardiac output syndrome score as predictors of adverse events associated with low cardiac output syndrome early after cardiac bypass*. Pediatr. Crit. Care Med. 20, 737–743 (2019).
Muñoz-Bonet, J. I. et al. Usefulness of telemedicine for home ventilator-dependent children. J. Telemed. Telecare. 26, 207–215 (2020).
Lin, L. I. A concordance correlation coefficient to evaluate reproducibility. Biometrics 45, 255–268 (1989).
MedCalc Software Ltd, Ostend, Belgium; https://www.medcalc.org; 2022. MedCalc® Statistical Software version 20.109.
Neill, D. B. Using artificial intelligence to improve hospital inpatient care. IEEE Intell. Syst. 28, 92–95 (2013).
Wang, L. et al. Artificial intelligence for COVID-19: A systematic review. Front. Med. 8, 704256. https://doi.org/10.3389/fmed.2021.704256 (2021).
Ghaderzadeh, M. et al. Deep convolutional neural network-based computer-aided detection system for COVID-19 using multiple lung scans: Design and implementation study. J. Med. Internet Res. 23, e27468 (2021).
Taenzer, A. H., Pyke, J., Herrick, M. D., Dodds, T. M. & McGrath, S. P. A comparison of oxygen saturation data in inpatients with low oxygen saturation using automated continuous monitoring and intermittent manual data charting. Anesth. Analg. 118, 326–331 (2014).
Buell, K. G. et al. Big data for clinical trials: Automated collection of SpO2 for a trial of oxygen targets during mechanical ventilation. J. Med. Syst. 44, 153 (2020).
Cortés-Reyes, E., Rubio-Romero, J. A. & Gaitán-Duarte, H. Métodos estadísticos de evaluación de la concordancia y la reproducibilidad de pruebas diagnósticas. Rev. Colomb Obstet. Ginecol. 61, 247–255 (2010).
Acknowledgements
We would like to express our gratitude to Dr. Francisco Ruza and Dr. Ignacio Ibarra for critically reviewing this manuscript. We also thank Dr. Maria Ledran for linguistic services.
Author information
Authors and Affiliations
Contributions
J.I.M. and J.B. conception of the study. J.I.M, V.P. and L.G. design of the study. J.I.M., V.P., L.G. and J.S. literature review. V.P. and L.G. acquisition of the data. All the authors contributed to the analysis and interpretation of the data. J.I.M., V.P. and L.G. drafted the submitted article. J.S., J.L.V., A.C. and J.B. provided substantial revision of the intellectual content of the article. All the authors approved the final version to be published and have agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding author
Ethics declarations
Competing interests
J.I.M. declares, as a researcher at the University of Valencia and the INCLIVA Biomedical Research Institute, that he collaborates in the development of telemedicine solutions at the Medical Online Technology S.L. company. None of the other authors have any conflicts of interest to declare.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Muñoz-Bonet, J.I., Posadas-Blázquez, V., González-Galindo, L. et al. Exploring the clinical relevance of vital signs statistical calculations from a new-generation clinical information system. Sci Rep 13, 15068 (2023). https://doi.org/10.1038/s41598-023-40769-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-40769-3
- Springer Nature Limited