Abstract
This study aimed to determine the presence and characteristics of locally circulating strains of Streptococcus suis, the most important streptococcal pathogen in swine. Oral swab samples were collected from pigs from 664 representative smallhold farms across nine provinces in the Philippines. Isolates were identified and characterized using PCR assays. The study revealed an isolation rate of 15.8% (105/664, 95% CI: 13.0–18.6) among the sampled farms. Two hundred sixty-nine (269) S. suis isolates were recovered from 119 unique samples. Serotype 31 was the most prevalent (50/269, 95% CI: 13.9–23.2) among the other serotypes identified: 5, 6, 8, 9, 10, 11, 15, 16, 17, 21, 27, 28, and 29. The detection of the three ‘classical’ S. suis virulence-associated genes showed that 90.7% (244/269, 95% CI: 87.2–94.2) were mrp-/epf-/sly-. Multilocus sequence typing (MLST) analysis further revealed 70 novel sequence types (STs). Notably, several local isolates belonging to these novel STs formed clonal complexes (CC) with S. suis strains recovered from Spain and USA, which are major pork-exporting countries to the Philippines. This study functionally marks the national baseline knowledge of S. suis in Philippines.
Similar content being viewed by others
Streptococcus suis, a Gram-positive bacterium, is the most important streptococcal pathogen affecting swine industries worldwide1. Pigs infected with virulent strains of this bacterium suffer from various clinical manifestations, including meningitis, arthritis, endocarditis, septicemia, pneumonia, and/or sudden death2. Despite the dense pig population in Southeast Asian countries and cultural practices involving the consumption of raw and undercooked pork in countries like Thailand, Vietnam and Laos, limited data on S. suis infections in pigs are available for countries such as Singapore, Philippines, Laos, and Cambodia3,4,5. This is of great concern as the number of human cases of S. suis infection has been increasing in this region3,4. Several major outbreaks of human infections were recorded in Thailand and China, resulting in deaths and significant economic loss to the affected provinces6,7.
In the Philippines, between 47 and 60% of the animal meat consumption of Filipinos comes from pork8. For years, the Philippines has been among the top pork producers worldwide, with an estimated value at around PHP 270 billion in 2021 and a pig population of around 10.18 million as of March 20238,9,10. Smallhold farms consistently account for most of the pork production in the Philippines, contributing between 62 and 78% of the total swine inventory annually, spanning from 2000 to 202310. However, this farm type is susceptible to significant losses due to limited practice of biosecurity measures such as vaccination, footbath provision, perimeter fencing, rodent control, and swill feeding management11.
In Huong et al.12, the Philippines, considering its wide and intensive hog raising practice, was identified as the Asian gap in information on S. suis. This is an important call that necessitates action particularly since multiple cases of human infections of S. suis have already been reportedly contracted in the country, including a number of cases diagnosed abroad3,13,14,15,16,17,18. This highlights years, if not decades, of gap in S. suis research data relevant to livestock health and disease monitoring, human health risks, and emergence and changes in antimicrobial resistance pattern in the country.
As a response, this study aimed to investigate the presence, distribution, and characteristics of S. suis in smallhold farms in the Philippines, specifically, the locally circulating serotypes, virulence gene profiles, and sequence types of the bacterium in the country.
Results
Isolation of Streptococcus suis
Out of the 664 farms surveyed, S. suis isolates were identified from 105 (15.8%, 95% CI: 13.0–18.6) farms using the S. suis-specific, recN-based PCR assay. None of the farms can be positively identified as affected by S. suis clinical infections. The observed proportion of farms with positive isolation of S. suis is 35.9% (28/78, 95% CI: 25.3–46.5) in Zamboanga del Norte, 23.6% (17/72, 95% CI: 13.8–33.4) in Bohol, 22.2% (16/72, 95% CI: 12.6–31.8) in Batangas, 21.9% (16/73, 95% CI: 12.4–31.4 ) in Marinduque, 12.3% (9/73, 95% CI: 4.8–19.9) in Misamis Oriental, 9.0% (7/78, 95% CI: 2.6–15.3) in Cebu, 8.2% (6/73, 95% CI: 1.9–14.5) in Albay, 6.9% (5/72, 95% CI: 1.1–12.8) in Misamis Occidental, and 1.4% (1/73, 95% CI: 0–4.0) in Iloilo. Zamboanga del Norte, which was estimated to have the highest positive isolation rate at provincial level, was significantly different from Misamis Oriental, Cebu, Albay, Misamis Occidental, and Iloilo. However, it was not significantly different from Bohol, Batangas, and Marinduque, as reflected by the overlapping interval estimates among these provinces. Alternatively, Iloilo, which posted the lowest positive isolation rate, was significantly different from Misamis Oriental, Marinduque, Batangas, Bohol, and Zamboanga del Norte but was not statistically different from Cebu, Albay, and Misamis Occidental (Fig. 1, Supplemental Fig. 1). Among the 105 S. suis positive farms, the proportion of sampled pigs positive for S. suis ranged from 13 to 100% per farm.
Geographical distribution of the recovered isolates of Streptococcus suis among nine provinces in the Philippines. The map was created using MapChart software (https://www.mapchart.net).
Comparing the proportions of S. suis-positive pigs by age group, isolation from piglets and sows outnumber isolation proportions from growers and boars. Results showed that 10.4% of piglets (45/433, 95% CI: 7.5–13.3), 8.1% of sows (36/447, 95% CI: 5.5–10.6), 5.6% of growers/finishers (37/660, 95% CI: 3.9–7.4), and 3.7% of boars (1/27, 95% CI: 0.0–10.8) tested positive for S. suis by isolation. However, the observed difference in the estimates could not support the idea that any age group is statistically more likely to be S. suis positive, as shown by the overlapping interval estimates (Supplemental Fig. 2).
In total, swab samples were individually collected from the oral cavities of 1,567 pigs, particularly from the surface of the palatine tonsils. Among these samples, 269 S. suis isolates were recovered from 119 unique samples (7.6%, 95% CI: 6.3–8.9). All isolates were recovered from asymptomatic pigs, except for two isolates obtained from a deceased piglet that exhibited fever symptoms before coincidentally dying on the day of sampling. No additional information was available for these two isolates that would confirm clinical S. suis infection. Therefore, these two isolates were characterized together with all other isolated S. suis strains. No isolates recovered from systemic or invasive clinical infections of S. suis were included.
To further validate the identity of the S. suis isolates, 11 representative isolates were subjected to 16 s rRNA gene sequencing. BLAST analysis revealed a 100% query cover and a 99 to 100% percentage identity (PIaD) to S. suis. All sequences were submitted to NCBI with GenBank accession numbers OR287118–OR287128. Figure 2 shows the phylogenetic tree constructed from the partial 16 s rRNA gene sequences of the recovered isolates, alongside several type and representative sequences from other streptococcal species.
Characterization of recovered Streptococcus suis isolates
Molecular serotyping
From the 119 unique swab samples gathered from the sampled animals, 269 S. suis isolates were recovered. The identification of the capsular polysaccharide (cps) types of S. suis was carried out using a four-reaction multiplex PCR assay19.
Among the recovered isolates, 131 of isolates tested were assigned to the 29 recognized serotypes, while the remaining 138 isolates could not be classified into any specific serotype and were categorized as nontypeable isolates. Nontypeable isolates were the most prevalent, followed by serotype 31. The proportion of the nontypeable strains (138/269, 51.3%, 95% CI: 45.3–57.3) was significantly higher than any of the typeable strains, as evidenced by their 95% confidence intervals. Additionally, among typeable strains the proportion of serotype 31 strains (50/269, 18.6%, 95% CI: 13.9–23.2) was significantly higher than all other detected typeable serotypes. Furthermore, the proportion of serotype 9 strains was significantly different from serotypes 31, 27, 5, 6, 29, 11, 17, 10, 28, and 15 but not significantly different from serotypes 21, 16, and 8. Subsequently, the proportion of serotype 27 strains was significantly different from serotypes 9 and 31 but not significantly different from the rest of the typeable strains (Supplemental Fig. 3).
Population snapshot of Streptococcus suis in the Philippines showing (A) minimal spanning tree representing the serotype distribution of STs and the newly identified clonal complexes, (B) the relationship of a novel ST with previously recognized clonal complexes (C) the newly identified clonal complexes, and (D) singletons.
Virulence gene profiling
Only 9.3% (95% CI: 5.8–12.8) of isolates carried at least one of the three classical virulence-associated genes (VAGs), namely mrp, epf, and sly. The predominant genotype detected is mrp-/epf-/sly- (90.7%, 95% CI: 87.2–94.2). Other observed genotypes include mrp-/epf-/sly+ (n = 11) and mrp-/epf+/sly- (n = 14). The sly-positive isolates were associated with serotypes 6, 11, and 15. Other sly-positive isolates were found among nontypeable strains. The virulence-associated gene epf was observed in serotype 8 (n = 11), serotype 27 (n = 2), and serotype 31 (n = 1). It could also be important to note that 52.0% (95% CI: 32.4–71.6) of isolates positive for VAGs were recovered from sows and piglets.
Multilocus sequence typing
From the 269 S. suis isolates, 145 were selected for multilocus sequence typing (MLST) using seven different housekeeping genes (aroA, cpn60, dpr, gki, mutS, recA, and thrA). The sequences were submitted to pubMLST database, and analyses revealed 101 novel alleles. Specifically, 14 new alleles were found in the aroA gene, 19 in the cpn60 gene, 10 in the dpr gene, 17 in the gki gene, 17 in the mutS gene, 13 in the recA gene, and 11 in the thrA gene.
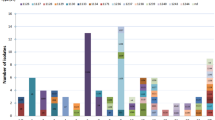
Interestingly, it was determined that all the 145 isolates could not be typed into any of the previously reported STs available in the database and were subsequently assigned to 70 novel STs. The most common ST observed was ST2157, which was observed in 11 isolates, followed by ST2162, ST2166, and ST2211, each with seven isolates. Meanwhile, ST2168 and ST2181 have six isolates each, and ST2170, ST2176 and ST2215 were represented in the collection by five isolates each. Further analysis of the association between serotypes and STs (Fig. 3.A.) showed that eight out of the 14 identified serotypes exhibited multiple STs. Serotype 31, the predominant serotype, exhibited 22 different STs. These include ST2152, ST2166, ST2167, ST2168, ST2175, ST2176, ST2189, ST2191, ST2192, ST2193, ST2194, ST2195, ST2197, ST2198, ST2199, ST2200, ST2201, ST2202, ST2270, ST2271, ST2272, and ST2274. The number of distinct STs within the other serotypes range from one to 10 (Fig. 3.A).
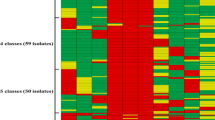
It was also determined that ST2154, ST2174, ST2196, ST2172, and ST2157 isolates tested positive for at least one VAG. These five STs were specifically linked to certain serotypes: ST2174 with serotype 15; ST2196 with serotype 31; ST2172 and ST2154 with serotype 6; and ST2157 with serotype 8. A clonal complex (CC) refers to a group of two or more independent isolates sharing identical genetic alleles at five or more loci20. Eleven (11) distinct CCs among the 145 isolates were identified using PHYLOViZ software goeBURST algorithm and illustration21: CC2217/2215, CC2159/2152, CC2179/2180, CC2211/2212, CC2189/2192, CC2170/2171, CC2272/2274, CC2198/2196, CC2270/2202, CC848/CC2161, and CC17 (Fig. 3A–C). ST2157 is related to CC17 via ST14. CC16 is related to CC87, whereas CC87 is closely related to CC89 (Fig. 3B). The remaining 50 STs were observed to be singleton isolates, which were not associated with any CCs (Fig. 3D).
Notably, two STs from the Philippines were connected to two other STs, one from Spain and another from USA. Specifically, ST2157 from Philippines belongs to CC17 via ST14 from Spain, while ST2161 from the Philippines formed a clonal complex with ST848 from USA. A total of 7.6% isolates (11/145) belonged to CC17 and 1.4% isolates (2/145) belonged to CC848/CC2161.
Discussion
Pigs and wild boars are considered natural reservoirs of S. suis, and colonization is generally regarded as present in almost all herds1,22. An individual pig, whether diseased or asymptomatic, may carry a mixture of multiple and diverse S. suis strains colonizing the upper respiratory tract, particularly the tonsils and nasal cavities, as well as the vaginal and possibly the alimentary tract1. The transition of S. suis from a colonizing pathobiont into an invasive pathogen is driven by a complex multifactorial process influenced by host health, environmental conditions, and strain-specific factors22,23.
In diseased pigs, examination of the clinical manifestation, the age of the affected individuals, and the characteristics of macroscopic lesions are common reference points for a presumptive diagnosis of S. suis clinical infections. This diagnosis is only confirmed when microscopic lesions typical of S. suis infection are observed, and isolation of S. suis is achieved from tissue samples, preferably from multiple organs or sites1. Meanwhile, in asymptomatic pigs, the recovery of S. suis strains from the nasal cavities or tonsils suggests a high load of this bacterium in these sites. This is because microbiological isolation is of low sensitivity for detection due to the presence of competing members of the microbiota. Isolation of S. suis strains from these sites may, therefore, indicate active transmission rather than a carrier state1,24,25. However, this distinction has not consistently been applied in the literature such that the proportion of pigs in a sampled population that is culturally positive for S. suis has also been referred to as the carriage rate, while the individual animals have also been called carrier pigs26,27,28,29.
It is crucial to note, however, that such colonizing isolates of S. suis from asymptomatic pigs should not be automatically considered avirulent or non-pathogenic strains. The classification of S. suis isolates as virulent or avirulent, pathogenic, opportunistic, or strictly commensal is a complex issue that has previously been problematized and discussed. In brief, there is no standard definition of what a virulent strain is. Asymptomatic pigs may carry virulent strains, particularly taking into account that most human cases of infections are acquired due to the consumption of contaminated pork products that are presumably from healthy pigs. Further complicating this problem is the absence of standardized assessment methods and models, either in vivo or in vitro, to assess strain virulence or evaluate the importance of suggested critical virulence factors, which would enable direct comparison of results30.
As such, for both clinical infections and asymptomatic colonization of S. suis, direct detection of S. suis by molecular detection from tonsils or nasal cavities offer limited practical utility1,31. On the other hand, while microbiologic isolation of S. suis strains may be less sensitive than molecular or serologic detection, isolation is a requirement for the confirmation of clinical infection and identification of potentially actively transmitted strains in asymptomatic pigs1. Furthermore, recovered isolates, regardless of the clinical condition of the source, offer opportunities for further genomic and phenotypic characterizations, including but not limited to serotyping, detection of virulence-associated genes, MLST, host challenge assays, and antimicrobial resistance, among others. These pieces of information are of utmost importance in the characterization of the epidemiology of S. suis and in the holistic evaluation of the importance of circulating S. suis strains. Microbiologic isolation of S. suis strains, therefore, has been utilized in numerous studies across various countries to describe the prevalence and characteristics of S. suis in both diseased and asymptomatic pigs28,32,33,34,35,36,37,38,39,40,41,42.
Almost a decade after Huong et al. (2014) singled out the Philippines as the S. suis knowledge gap in Asia, this study presents a representative cross-sectional survey in smallhold swine farms in the country12. No prior information on outbreaks, if they had occurred, or list of farms with a history of S. suis-related diseases was available to guide the selection of farms for sampling. Instead, smallhold farms were sampled from high pig population density provinces in the country.
The observed positive isolation rate among pigs in the current study (119/1567, 7.6%, 95% CI: 6.3–8.9) is comparable to an initial report of S. suis isolation in the Philippines in 2020, which achieved a 9.09% (4/44, 95% CI: 0.6–17.6) isolation rate from a single farm8. However, this rate is higher than an older report detailing zero recovered isolates from 220 nasal and tonsil swabs of slaughtered pigs from a province not included in the current study43. Similarly, reports on S. suis isolation from healthy pigs from farms or slaughterhouses in other countries, such as Northern Vietnam44, China (Jiangsu)45, Northern Thailand46, and Turkey43, showed isolation rates ranging from 0 to 6.0% . On the other hand, reports from China (Anhui)47, Thailand (Chiang Mai)48, and China (Xinjiang)47 ranged from 7.8 to 8.3%. Lastly, isolation rates reported from other provinces in China47, Korea49,50, India51, Spain52, Canada (Quebec)53, Thailand (Phayao)27,54, UK26, Northern and Central Thailand28,46, Southern Vietnam36, and Canada (Ontario)55 ranged from 11.2 to 73.1%, demonstrating a wide range of reported S. suis isolation rates in the literature (Supplemental Fig. 4).
Currently, 29 serotypes of S. suis are recognized based on the serological characteristics of the bacterial capsular polysaccharide56,57,58,59. Among these serotypes, serotype 2 is most associated with infections in both pigs and humans. It has been determined to be responsible for over 80% of human cases, putting emphasis on its importance as a global zoonotic threat60. As a result, a significant portion of the research and reports has primarily focused on S. suis serotype 2 samples obtained from diseased pigs and humans. Globally, the predominant S. suis serotypes in clinical pig cases are serotypes 2, 9, 3, 1/2, and 7. However, within North America, serotypes 2 and 3 were the predominant serotypes, followed by serotypes 1/2, 8, and 73,30,61. This can be attributed to the fact that the distribution of serotypes exhibits significant variations over time and in different geographical areas41.
In this study, no serotype 2 strains were recovered. Serotype 31 (38.17%; 50/131) was detected in 7 out of 9 provinces sampled, which may suggest that there is a need to put serotype 31 under surveillance to further understand its clinal and epidemiological importance, and identify potential control measures, if necessary. This is particularly important since other studies in Vietnam, Thailand, Spain, Canada, UK, and China only reported isolation rates of serotype 31 strains ranging from 0 to 7.2% in healthy or asymptomatic pigs28,26,62. On the other hand, there have also been several reports of serotype 31 associated with diseased pigs in Canada42 and China63 (Supplemental Fig. 5a). Notably, this serotype has also been reported to have caused a human infection in Thailand56.
Furthermore, over the last two decades, there has been an observed increase in the prevalence of serotype 9 isolates from diseased pigs in Netherlands33, Spain39, and Canada62,64, and it is also among the most commonly isolated serotypes in both clinically healthy and diseased pigs in commercial farms in China65. Reported proportion of serotype 9 isolates from diseased pigs ranged from 0–47.3% as reported from studies in Canada42,55,62, Spain39, Brazil66, Taiwan67, and the Netherlands68. Meanwhile, studies on healthy or asymptomatic pigs in Canada55,62, China26, Vietnam36, Korea49,50, Thailand27,28,54, UK26, and Spain52 ranged from 0 to 15.9% (Supplemental Fig. 5b).
Meanwhile, serotype 8 is reported to be isolated in diseased pigs from Spain, Brazil, Canada, Taiwan and Korea39,49,62,66,67,69. Serotype 8 ranked 5th as the most predominant and pathogenic serotype in diseased pigs worldwide from 2002 to 20133. Serotype 16, on the other hand, has also been reportedly isolated from diseased pigs in Germany and South Korea70. Lastly, serotype 21 is also recovered in clinically healthy pigs in North America but is associated with diseased pigs in Canada42,71,72. Variable detection rates of serotypes 8, 16, and 21 both from diseased and non-diseased samples have been reported in the literature27,28,36,39,42,49,50,52,53,54,55,26,62,66,67,68. Detection rates of serotype 8 strains ranged from 0 to 54.0%, while serotypes 16 and 21 were reportedly detected at 0–24.7% and 0–6.3%, respectively (Supplemental Fig. 5c–e).
The predominance of nontypeable strains in the current study aligns with findings from other publications reporting that nontypeable S. suis strains are commonly isolated in field conditions. These strains may be considered as potential novel serotypes or mutants of known serotypes19,73,74. Studies demonstrated that unencapsulated strains, which include nontypeable isolates, exhibit unique properties such as increased adherence to surfaces and cells, as well as the ability to form biofilms, which may contribute to their persistence and transmission75. Nontypeable isolates are also frequently recovered from both clinically ill and healthy pigs74. Although available information is limited, there have been reports of nontypeable S. suis isolates from pigs with meningitis in China (designated novel serotype variant Chz)76, and a nontypeable unencapsulated S. suis strain has also been reported in a human case in Thailand77. Hence, it is also important to consider potential risks associated with nontypeable isolates, considering that their role in disease transmission could not be disregarded.
The predominant S. suis VAG genotype (90.71%; 244/269) identified in this study is mrp−/epf−/sly−. This is consistent with other reports on S. suis isolated from healthy/carrier pigs from China78, Thailand28, and Germany35, forwarding the observation that isolates positive for epf, mrp, and sly genes were significantly less frequently detected in clinically healthy pigs35. It is important to note, however, that this determination does not automatically indicate avirulence, since most virulence-related studies that considered epf, mrp, and sly as the main VAGs primarily studied serotype 2 strains79. In fact, the mrp+/epf+/sly+ genotype is almost always reported in serotype 2 strains isolated from diseased pigs in Europe and Asia34,80. Moreover, it has been suggested that mrp, epf, and sly are coincidentally associated with virulence rather than being the actual determining factors of virulence, which means the absence of one or more of these proteins does not necessarily result in lack of virulence30,81. A more recent whole genome and pan-genome analyses suggested that ofs (encoding for serum opacity factor) and srtF (encoding for sortase F) are stronger predictors for differentiating pathogenicity compared to 71 other previously suggested VAGs in S. suis, including mrp, epf, and sly82.
Among the 145 S. suis isolates that were examined by MLST, no previously known STs were found. These isolates were submitted to pubMLST and were assigned as 70 novel STs. Generally, S. suis isolates from clinically healthy pigs are not subjected to MLST, which could partially account for the numerous novel STs in this study83. Of note, two specific STs were found to have connections to STs from other countries. ST2157 isolates, which were recovered from the dead piglet, was linked to ST14 from Spain, a serotype 3 strain isolated from a lung tissue showing pneumonia. The diseased piglet was not reported to exhibit manifestations of pneumonia but was only noted to have symptoms of fever before sudden death. On the other hand, ST2161 was connected to ST848 from the United States of America (USA), which was isolated from a pig in 2015. Considering these observed connections, it is tempting to hypothesize if importation and international trade could also be important factors to account for the diversity of S. suis in the Philippines, given that pork and pig products are also common sources of S. suis infections84. Based on the data obtained from the Foreign Agricultural Service (FAS) under the US Department of Agriculture (USDA), the Philippines has maintained a longstanding practice of importing live pigs from the United States, with a cumulative total of pigs imported at 14,689 heads from 1967 to 2022. The Philippine Statistics Authority (PSA) reported that a total of 2,703 live pigs were imported mainly from Canada, Spain, Netherlands, Brazil, and the USA 2018 to 2020. The problem, however, is that there is no information on the movement of these imported animals within the Philippines making it impossible to prove the connection of importation and the observed relationship among Philippine isolates sharing a clonal complex with isolates from other countries.
Pending the availability of information on the predominant serotypes of S. suis from clinical cases of infections, the current results, considering the observed predominant serotypes and the entirely novel set of MLST, suggest a potentially different epidemiology of S. suis in the Philippines34,41,49,38.
Smallhold farms contribute about 80% of the total hog production in the Philippines. Despite this, the scant information on important bacterial pathogens, especially those that pose zoonotic risks like S. suis, coupled with the limited implementation and understanding of biosecurity-related practices and measures, point to an increased danger of transmission of the pathogen among animals, and at the same time, from animals to humans.
Altogether, the current findings, which provided detailed molecular characterization of circulating S. suis strains in the Philippines for the first time, are highly relevant to both veterinary and human medicine, especially for individuals with direct contact with pigs and pork products, including pig farmers, abattoir workers, as well as the general consumers3,12,28,35,61. However, the current study is not without limitations, which must be given attention with further research studies. Particularly, subsequent studies may focus on clinical isolates of S. suis, including historical isolates (if present in regional or national diagnostic laboratories) and newer isolates recovered from more recent cases. Thus, this also emphasizes the importance of continuous surveillance of this pathogen among farms, and even among slaughterhouses and markets in the country, as performed in other countries before35,36,60. Also, in consideration to the growing number of studies indicating the propensity of S. suis for rapid development of multidrug resistance27,85,86,87,88,89,90,91,92,93,94,95, characterization of local isolates must also be extended to cover phenotypic and genotypic antimicrobial resistance characteristics to gain an understanding on the applicability and efficacy of currently implemented practices regarding antibiotic usage. Finally, whole genome sequencing and related bioinformatics studies on local isolates, particularly those that were identified as epidemiologically or clinically significant, would further bolster grounded local understanding of this important pathogen.
With plans to further intensify the swine raising industry in the Philippines, the current study provides a snapshot of the on-going conditions related to S. suis infection in key hog-raising provinces in the country. This study functionally marks the national baseline knowledge on S. suis in Philippines.
Methods
Ethics statement
Ethics approval for the research, specifically for the conduct of and the protocol for sample collection from farm animals, was obtained from the Institutional Animal Care and Use Committee (IACUC) of the University of the Philippines Los Baños, with the assigned protocol number BIOTECH-2021-001. Additionally, the collection of samples for this study was permitted by each farm owner prior to the sample collection from their pigs. All experiments conducted in this study are in accordance with the relevant regulations, guidelines, and protocols, as evaluated and approved by the UPLB-IACUC.
Sampling collection
The Philippines is generally divided into three major island groups (Luzon, Visayas, and Mindanao), further subdivided into regions and provinces. While the specific relationships between and among provinces within and among these island groups in relation to trade and transport of live and slaughtered pigs are unknown, it is assumed that a significant percentage of these movements are regional96.
Using a multistage stratified random sampling approach, it was determined that a minimum of 648 farms were to be sampled based on Cochran’s method (1977) with the following assumptions: assumed prevalence (P): 0.3, confidence level (1-α): 0.95, margin of error (α): 0.5, and design effect (DE): 2.097.
Based on the Philippine Statistics Authority’s Swine Situation Report as of January 1, 201998, provincial pig population densities were determined, and high hog production provinces were identified. A random sample of three high hog production provinces was selected from each of the three geographical island groups. The provinces randomly selected were Marinduque, Albay, and Batangas for Luzon, Iloilo, Bohol, and Cebu for Visayas, and Misamis Occidental, Zamboanga del Norte, and Misamis Oriental for Mindanao.
Provincial government authorities were requested for a list of villages or barangays that have at least 24 small hold farms. Within these provinces, a random sample of three villages or barangays was selected, and a random sample of 24 households/backyard farms was selected from each barangay. Up to three samples per age group of pigs were collected whenever possible. A sampled pig was classified as either a sow, boar, piglet, or grower by adapting local classification and farming practice. Young pigs housed with the sow and feeding on her milk are considered piglets. A pig housed separately from the sow (i.e., weaned) is considered a grower since distinction among weaners, growers, and finishers is rarely locally applied.
The sampled pigs were manually restrained, and a nose snare and a mouth gag were used to open the mouth and prevent biting. Oral swab samples were collected by rubbing a cotton swab against the oral cavities of the pigs, particularly on the surface of the palatine tonsils whenever safely possible, following the approved protocol identified above. The collection of the swab samples was performed under the guidance of provincial/municipal veterinarians or veterinary technicians. Samples were immediately placed in transport tubes containing 4.0 mL of Amies transport medium. All samples were stored and transported in insulated coolers and immediately processed for streptococcal isolation afterward.
Isolation and Identification
Serial dilutions of the transport medium were prepared, and 100 µL aliquots of varying dilutions were spread plated onto Columbia Agar plates with 5% defibrinated sheep blood. The plates were then incubated at 37 °C under 5% CO2 for 18–24 h. Up to eight α-hemolytic colonies were selected and purified from each sample25,26,27,28,29,36,37. All successfully purified isolates were identified using a conventional PCR assay utilizing S. suis-specific recN gene primers, as described elsewhere99.
Partial 16 s rRNA gene sequences of selected isolates were determined by Sanger sequencing following the amplification of the said gene using the F1/R13 primers, as described elsewhere100. Sequences were submitted to the NCBI Genbank database with accession numbers OR287118–OR287128. To better determine sequence similarity, the 16 s rRNA gene sequences of reference isolates of relevant Streptococcus species101 were also retrieved from the GenBank. Sequences obtained from this study and the retrieved representative sequences from NCBI were aligned using ClustalW method in Molecular Evolutionary Genetics Analysis (MEGA-X) software102. Neighbor-joining method103 with bootstrap analysis of 1000 replications was used to construct a phylogenetic tree.
Molecular characterization
Further characterization of the identified S. suis isolates was conducted using serotyping, virulence gene profiling, and multilocus sequence typing (MLST). The serotyping of all identified S. suis isolates was carried out using four multiplex PCR assays19, all targeting capsular polysaccharide synthesis (cps) genes located on a single locus in the S. suis chromosome. These assays can identify the currently recognized 29 S. suis serotypes.
All S. suis isolates were similarly profiled for the presence of virulence-associated genes using a multiplex PCR assay that targets the extracellular protein factor (epf), muramidase-release protein (mrp), and suilysin (sly)104, commonly referred to as virulence-associated genes (VAGs). The MLST was done using multiplex PCR protocol targeting seven housekeeping genes, including aroA (EPSP synthase), cpn60 (60-kDa chaperonin), dpr (peroxide resistance), gki (glucose kinase), mutS (DNA mismatch repair enzyme), recA (homologous recombination), and thrA (aspartokinase)20. The PCR products were subsequently submitted for sequencing. One hundred twenty-five serotypeable isolates were sequenced for MLST. The remaining six serotypeable isolates did not yield amplicons for one or more of the target genes and thus, were not typed by MLST. An additional 20 nontypeable isolates were selected in random and similarly sequenced. Recovered sequences were submitted to pubMLST (March–May 2023) for verification and typing assignment. The complete dataset pertaining to the isolates, typed by MLST, including provenance/relevant metadata and sequences, can be accessed through the pubMLST database under ID numbers 3539-3546, 3550-3606 and 3660-3739.
The entire S. suis database in pubMLST (available on August 13, 2023) was downloaded and used to visualize and identify groups of related genotypes and clonal complexes using the goeBURST algorithm in the PHYLOViZ software21.
Statistical analysis
Data were stored in an in-house Excel 365 (Microsoft) database. Descriptive statistics related to isolation rates per province, farm, and age group as well as the proportion of each serotype among the isolates, were calculated using the same software. Confidence interval estimates of the proportion were calculated with a confidence level of 95%. Meanwhile, in categories with zero observations of the character of concern, the rule of three states that the upper limit of the 95% confidence interval can be estimated by 3/n44,105.
Data availability
The datasets generated in the current study pertaining to the isolates typed by MLST, including provenance/relevant metadata and sequences, can be accessed through the pubMLST database under ID numbers 3539-3546, 3550-3606 and 3660-3739, while the generated and reported partial 16 s rRNA sequences can be accessed through the NCBI database with GenBank accession numbers OR287118-OR287128. All data presented in this study are included in this article.
References
Gottschalk, M. & Segura, M. Streptococcosis. in Diseases of Swine (eds. Zimmerman, J. J. et al.) 934–950 (Wiley Blackwell, 2019).
Vecht, U., Wisselink, H. J., Jellema, M. L. & Smith, H. E. Identification of two proteins associated with virulence of Streptococcus suis type 2. Infect. Immun. 59, 3156–3162 (1991).
Goyette-Desjardins, G., Auger, J. P., Xu, J., Segura, M. & Gottschalk, M. Streptococcus suis, an important pig pathogen and emerging zoonotic agent—An update on the worldwide distribution based on serotyping and sequence typing. Emerg. Microbes Infect. 3, 1–20 (2014).
Dutkiewicz, J. et al. Streptococcus suis: A re-emerging pathogen associated with occupational exposure to pigs or pork products. Part I—Epidemiology. Ann. Agric. Environ. Med. 24, 683–695 (2017).
Wertheim, H. F. L., Nghia, H. D. T., Taylor, W. & Schultsz, C. Streptococcus suis: An emerging human pathogen. Clin. Infect. Dis. 4, 617–625. https://doi.org/10.1086/596763 (2009).
Kerdsin, A. Human Streptococcus suis infections in Thailand: Epidemiology, clinical features, genotypes, and susceptibility. Trop. Med. Infect. Dis. 7, 359 (2022).
Ji, L. et al. Epidemiological and genomic analyses of human isolates of Streptococcus suis between 2005 and 2021 in Shenzhen, China. Front. Microbiol. 14, 1118056 (2023).
Sedano, S. A., Silva, B. B. I., Sangalang, A. G. M. & Merlyn, S. Streptococcus suis and S. parasuis in the Philippines: Biochemical, molecular, and antimicrobial resistance characterization of the first isolates from local swine. Philipp. Sci. Lett. 13, 107–119 (2020).
(PSA), P. S. A. Philippine Statistics Authority: Swine situation report. (2020).
(PSA), P. S. A. Philippine Statistics Authority. Swine situation report. vol. 81 (2023).
Bernardes, D. T. C. & Peña, S. T. Biosecurity and readiness of smallholder pig farmers against potential African Swine Fever outbreak and other pig diseases in Baybay City, Leyte, Philippines. Sci. Agropecu. 11, 611–620 (2020).
Huong, V. T. L. et al. Epidemiology, clinical manifestations, and outcomes of Streptococcus suis infection in humans. Emerg. Infect. Dis. 20, 1105–1114 (2014).
Lee, G. T. et al. Streptococcus suis meningitis, United States. Emerg. Infect. Dis. 14, 183–185 (2008).
Wongjittraporn, S., Teerasukjinda, O., Yee, M. & Chung, H. H. Streptococcus suis meningoencephalitis with seizure from raw pork ingestion: a case report. Hawaii J. Med. Public Health 73, 13–14 (2014).
Coner-Nobleza, M. J., Ramos-Jason, R., Lerios, J. K. G. & Gregorio, K. M. R. Streptococcus suis Septicemia: A case series from the Philippines.
Domado, A. M. & Itable, J. Streptococcus suis: Bacteremia presenting with fever, rashes, arthritis and neurologic deficits. Phillippine J. Intern. Med. 56, 27–33 (2018).
Quieta, R. Bacterial meningitis Streptococcus suis: Causes, treatment, and prevention. https://www.gmanetwork.com/entertainment/celebritylife/health/72831/bacterial-meningitis-streptococcus-suis-causes-treatment-and-prevention/story#:~:text=Asia’s Nightingale Lani Misalucha revealed,deaf in her right ear (2020).
Kerdsin, A., Segura, M., Fittipaldi, N. & Gottschalk, M. Sociocultural factors influencing human Streptococcus suis disease in Southeast Asia. Foods 11, 1–13 (2022).
Kerdsin, A. et al. Streptococcus suis serotyping by a new multiplex PCR. J. Med. Microbiol. 63, 824–830 (2014).
King, S. J. et al. Development of a multilocus sequence typing scheme for the pig pathogen Streptococcus suis: Identification of virulent clones and potential capsular serotype exchange. J. Clin. Microbiol. 40, 3671–3680 (2002).
Francisco, A. P. et al. PHYLOViZ: Phylogenetic inference and data visualization for sequence based typing methods. BMC Bioinform. 13, 1–10 (2012).
Vötsch, D., Willenborg, M., Weldearegay, Y. B. & Valentin-Weigand, P. Streptococcus suis—The ‘two faces’ of a pathobiont in the porcine respiratory tract. Front. Microbiol. 9, 1–10 (2018).
Obradovic, M. R., Segura, M., Segalés, J. & Gottschalk, M. Review of the speculative role of co-infections in Streptococcus suis-associated diseases in pigs. Vet. Res. 52, 1–22 (2021).
Cloutier, G. et al. Epidemiology of Streptococcus suis serotype 5 infection in a pig herd with and without clinical disease. Vet. Microbiol. 97, 135–151 (2003).
Marois, C., Le Devendec, L., Gottschalk, M. & Kobisch, M. Detection and molecular typing of Streptococcus suis in tonsils from live pigs in France. Can. J. Vet. Res. 71, 14–22 (2007).
Zou, G. et al. Effects of environmental and management-associated factors on prevalence and diversity of Streptococcus suis in clinically healthy pig herds in China and the United Kingdom. Appl. Environ. Microbiol. 84, 1–15 (2018).
Kerdsin, A. et al. Genotypic comparison between Streptococcus suis isolated from pigs and humans in Thailand. Pathogens 9, 4–11 (2020).
Meekhanon, N., Kaewmongkol, S., Phimpraphai, W., Okura, M. & Osaki, M. Potentially hazardous Streptococcus suis strains latent in asymptomatic pigs in a major swine production area of Thailand. J. Med. Microbiol. 66, 662–669 (2017).
Werinder, A. et al. Streptococcus suis in Swedish grower pigs: Occurrence, serotypes, and antimicrobial susceptibility. Acta Vet. Scand. 62, 1–9 (2020).
Segura, M., Fittipaldi, N., Calzas, C. & Gottschalk, M. Critical Streptococcus suis virulence factors: Are they all really critical?. Trends Microbiol. 25, 585–599 (2017).
Wileman, T. M. et al. Pathotyping the zoonotic pathogen Streptococcus suis: Novel genetic markers to differentiate invasive disease-associated isolates from non-disease-associated isolates from England and Wales. J. Clin. Microbiol. 57, 10–1128 (2019).
Kittiwan, N. et al. Genetic diversity and variation in antimicrobial-resistance determinants of non-serotype 2 Streptococcus suis isolates from healthy pigs. Microb. Genomics 8, 1–17 (2022).
Willemse, N. et al. Clonal expansion of a virulent Streptococcus suis serotype 9 lineage distinguishable from carriage subpopulations. Sci. Rep. 9, 1–11 (2019).
Wei, Z. et al. Characterization of Streptococcus suis isolates from the diseased pigs in China between 2003 and 2007. Vet. Microbiol. 137, 196–201 (2009).
Louise Prüfer, T. et al. Molecular typing of Streptococcus suis strains isolated from diseased and healthy pigs between 1996–2016. PLoS One 14, 1–17 (2019).
Ngo, T. H. et al. Slaughterhouse pigs are a major reservoir of Streptococcus suis serotype 2 capable of causing human infection in Southern Vietnam. PLoS One 6, 1–7 (2011).
Zhang, C. et al. Capsular serotypes, antimicrobial susceptibility, and the presence of transferable oxazolidinone resistance genes in Streptococcus suis isolated from healthy pigs in China. Vet. Microbiol. 247, 108750 (2020).
Besung, I. N. K. et al. Isolation and identification of Streptococcus suis from sick pigs in Bali, Indonesia. BMC Res. Notes 12, 1–6 (2019).
Tarradas, C. et al. Distribution of serotypes of Streptococcus suis isolated from diseased pigs in Spain. Vet. Rec. 154, 665–666 (2004).
Vela, A. I. et al. Analysis of genetic diversity of Streptococcus suis clinical isolates from pigs in Spain by pulsed-field gel electrophoresis. J. Clin. Microbiol. 41, 2498–2502 (2003).
Lunha, K. et al. Antimicrobial susceptibility of Streptococcus suis isolated from diseased pigs in Thailand, 2018–2020. Antibiotics 11, 2018–2020 (2022).
Gottschalk, M. et al. Characterization of Streptococcus suis isolates recovered between 2008 and 2011 from diseased pigs in Québec, Canada. Vet. Microbiol. 162, 819–825 (2013).
Baǧcigil, A. F. et al. Isolation of streptococcus species from the tonsils of slaughtered pigs. Turkish J. Vet. Anim. Sci. 37, 94–96 (2013).
Nguyen, N. T. T. et al. An epidemiological study of Streptococcus suis prevalence among swine at industrial swine farms in Northern Vietnam. One Heal. 13, 1–5 (2021).
Huan, H., Jiang, L., Tang, L., Wang, Y. & Guo, S. Isolation and characterization of Streptococcus suis strains from swine in Jiangsu province, China. J. Appl. Microbiol. 128, 1606–1612 (2020).
Lakkitjaroen, N. et al. Prevalence and antimicrobial susceptibility of Streptococcus suis isolated from slaughter pigs in Northern Thailand. Kasetsart J. - Nat. Sci. 45, 78–83 (2011).
Liu, P., Zhang, Y., Tang, H., Wang, Y. & Sun, X. Prevalence of Streptococcus suis in pigs in China during 2000–2021: A systematic review and meta-analysis. One Heal. 16, 100513 (2023).
Padungtod, P. et al. Incidence and presence of virulence factors of streptococcus suis infection in slaughtered pigs from Chiang Mai, Thailand. Southeast Asian J. Trop. Med. Public Health 41, 1454–1461 (2010).
Oh, S. I. et al. Capsular serotypes, virulence-associated genes and antimicrobial susceptibility of Streptococcus suis isolates from pigs in Korea. J. Vet. Med. Sci. 79, 780–787 (2017).
Han, D. U. et al. Prevalence, capsular type and antimicrobial susceptibility of Streptococcus suis isolated from slaughter pigs in Korea. Can. J. Vet. Res. 65, 151–155 (2001).
Rajkhowa, S. & Rajesh, J. B. Virulence associated gene profiling and antimicrobial resistance pattern of Streptococcus suis isolated from clinically healthy pigs from North East India. Lett. Appl. Microbiol. 73, 392–397 (2021).
Luque, I. et al. Genetic analysis of Streptococcus suis isolates recovered from diseased and healthy carrier pigs at different stages of production on a pig farm. Vet. J. 186, 396–398 (2010).
Brisebois, L. M., Charlebois, R., Higgins, R. & Nadeau, M. Prevalence of Streptococcus suis in four to eight week old clinically healthy piglets. Can. J. Vet. Res. 54, 174–177 (1990).
Thongkamkoon, P., Kiatyingangsulee, T. & Gottschalk, M. Serotypes of Streptococcus suis isolated from healthy pigs in Phayao Province, Thailand. BMC Res. Notes 10, 1–7 (2017).
Denich, L. C. et al. A case-control study to investigate the serotypes of S. suis isolates by multiplex PCR in nursery pigs in Ontario, Canada. Pathogens 9, 44 (2020).
Hatrongjit, R. et al. First human case report of sepsis due to infection with Streptococcus suis serotype 31 in Thailand. BMC Infect. Dis. 15, 1–7 (2015).
Guo, G. et al. Genomic characterization of Streptococcus parasuis, a close relative of Streptococcus suis and also a potential opportunistic zoonotic pathogen. BMC Genomics 23, 1–16 (2022).
Okura, M. et al. Current taxonomical situation of Streptococcus suis. Pathogens 5, 45 (2016).
Athey, T. B. T. et al. Population structure and antimicrobial resistance profiles of streptococcus suis serotype 2 sequence type 25 strains. PLoS One 11, e0150908 (2016).
Scherrer, S. et al. Population structure, genetic diversity and pathotypes of Streptococcus suis isolated during the last 13 years from diseased pigs in Switzerland. Vet. Res. 51, 1–16 (2020).
Segura, M. et al. Update on Streptococcus suis research and prevention in the era of antimicrobial restriction: 4th international workshop on S. suis. Pathogens 9, 1–39 (2020).
Arndt, E., Farzan, A., Slavićc, D., Maclnnes, J. & Friendship, R. An epidemiological study of Streptococcus suis serotypes of pigs in Ontario determined by a multiplex polymerase chain reaction. Can. Vet. J. 59, 997–1000 (2018).
Wang, J. et al. Investigation of the genomic and pathogenic features of the potentially zoonotic Streptococcus parasuis. Pathogens 10, 1–18 (2021).
Zheng, H. et al. Genomic comparisons of Streptococcus suis serotype 9 strains recovered from diseased pigs in Spain and Canada. Vet. Res. 49, 1–13 (2018).
Dong, C. L. et al. New characterization of multi-drug resistance of Streptococcus suis and biofilm formation from swine in Heilongjiang Province of China. Antibiotics 12, 1–14 (2023).
Costa, A. T. R. et al. Serotyping and evaluation of the virulence in mice of Streptococcus suis strains isolated from diseased pigs. Rev. Inst. Med. Trop. Sao Paulo 47, 113–115 (2005).
Wu, C. F. et al. Serotype and multilocus sequence typing of Streptococcus suis from diseased pigs in Taiwan. Sci. Rep. 13, 8263 (2023).
Schultsz, C. et al. Differences in the population structure of invasive Streptococcus suis strains isolated from pigs and from humans in The Netherlands. PLoS One 7, 5–10 (2012).
Gottschalk, M. & Lacouture, S. Distribution of Streptococcus suis (from 2012 to 2014) and Actinobacillus pleuropneumoniae (from 2011 to 2014) serotypes isolated from diseased pigs. Can. Vet. J. 58, 802–804 (2015).
Nghia, H. D. T. et al. Human case of Streptococcus suis serotype 16 infection. Emerg. Infect. Dis. 14, 155–157 (2008).
Estrada, A. A. et al. Serotype and genotype (multilocus sequence type) of Streptococcus suis isolates from the United States serve as predictors of pathotype. J. Clin. Microbiol. 57, 10–1128 (2019).
Lacouture, S., Olivera, Y. R., Mariela, S. & Gottschalk, M. Distribution and characterization of Streptococcus suis serotypes isolated from January 2015 to June 2020 from diseased pigs in Québec, Canada. Can. J. Vet. Res. 86, 78–82 (2022).
Gottschalk, M. et al. Streptococcus suis infections in humans: What is the prognosis for Western countries? (Part II). Clin. Microbiol. Newsl. 32, 97–102 (2010).
Denich, L. et al. Study of the relationship between untypable and typable isolates of Streptococcus suis recovered from clinically ill and healthy nursery pigs. Vet. Microbiol. 257, 1–6 (2021).
Bonifait, L., Gottschalk, M. & Grenier, D. Cell surface characteristics of nontypeable isolates of Streptococcus suis. FEMS Microbiol. Lett. 311, 160–166 (2010).
Pan, Z. et al. Novel variant serotype of Streptococcus suis isolated from piglets with meningitis. Appl. Environ. Microbiol. 81, 976–985 (2015).
Kerdsin, A. et al. A human case of Streptococcus suis infection caused by an unencapsulated strain. JMM Case Rep. 1, 10–13 (2014).
Wang, K. et al. Characterization of Streptococcus suis isolates from slaughter swine. Curr. Microbiol. 66, 344–349 (2013).
Staats, J. J., Feder, I., Okwumabua, O. & Chengappa, M. M. Streptococcus suis: Past and present. Vet. Res. Commun. 21, 381–407 (1997).
Wisselink, H. J., Smith, H. E., Stockhofe-Zurwieden, N., Peperkamp, K. & Vecht, U. Distribution of capsular types and production of muramidase-released protein (MRP) and extracellular factor (EF) of Streptococcus suis strains isolated from diseased pigs in seven European countries. Vet. Microbiol. 74, 237–248 (2000).
Fittipaldi, N., Segura, M., Grenier, D. & Gottschalk, M. Virulence factors involved in the pathogenesis of the infection caused by the swine pathogen and zoonotic agent Streptococcus suis. Future Microbiol. 7, 259–279 (2012).
Estrada, A. A. et al. Proposed virulence-associated genes of Streptococcus suis isolates from the United States serve as predictors of pathogenicity. Porc. Heal. Manag. 7, 1–16 (2021).
Gebhart, C., Estrada, A. A., Gottschalk, M., Rendahl, A. & Marthaler, D. G. Characterisation of Streptococcus suis isolates from the United States by serotyping, MLST, and virulence-associated gene profiling. In Pathogens vol. 9 (2020).
Ho, D. T. N. et al. Risk factors of Streptococcus suis infection in Vietnam. A case control study. PLoS One 6, e17604 (2011).
Hoa, N. T. et al. The antimicrobial resistance patterns and associated determinants in Streptococcus suis isolated from humans in southern Vietnam, 1997–2008. BMC Infect. Dis. 11, 1–8 (2011).
Ye, C. et al. Clinical, experimental, and genomic differences between intermediately pathogenic, highly pathogenic, and epidemic Streptococcus suis. J. Infect. Dis. 199, 97–107 (2009).
Zhu, Y. et al. ICESsuHN105, a novel multiple antibiotic resistant ICE in Streptococcus suis serotype 5 strain HN105. Front. Microbiol. 10, 1–13 (2019).
Lekagul, A., Tangcharoensathien, V. & Yeung, S. Patterns of antibiotic use in global pig production: A systematic review. Vet. Anim. Sci. 7, 100058 (2019).
Chen, L. et al. Antimicrobial susceptibility, tetracycline and erythromycin resistance genes, and multilocus sequence typing of Streptococcus suis isolates from diseased pigs in China. J. Vet. Med. Sci. 75, 583–587 (2013).
Uruén, C., García, C., Fraile, L., Tommassen, J. & Arenas, J. How Streptococcus suis escapes antibiotic treatments. Vet. Res. 53, 91 (2022).
Chen, C. et al. A glimpse of streptococcal toxic shock syndrome from comparative genomics of S. suis 2 Chinese isolates. PLoS One 2, e315 (2007).
Du, F., Lv, X., Duan, D., Wang, L. & Huang, J. Characterization of a linezolid- and vancomycin-resistant Streptococcus suis isolate that harbors optrA and vanG operons. Front. Microbiol. 10, 1–12 (2019).
Li, M. et al. GI-type T4SS-mediated horizontal transfer of the 89K pathogenicity island in epidemic Streptococcus suis serotype 2. Mol. Microbiol. 79, 1670–1683 (2011).
Hu, P. et al. Comparative genomics study of multi-drug-resistance mechanisms in the antibiotic-resistant Streptococcus suis R61 strain. PLoS One 6, e24988 (2011).
Palmieri, C., Varaldo, P. E. & Facinelli, B. Streptococcus suis, an emerging drug-resistant animal and human pathogen. Front. Microbiol. 2, 1–6 (2011).
Abao, L. N. B. et al. PRRS outbreaks and farm hygiene management in the Philippines. Agric. Manag. Res. 52, 131–136 (2014).
Cochran, W. G. Sampling Techniques (John Wiley & Sons Inc., 1977).
(PSA), P. S. A. Philippine Statistics Authority: Swine situation report. vol. 8600 (2019).
Ishida, S. et al. Development of an appropriate PCR system for the reclassification of Streptococcus suis. J. Microbiol. Methods 107, 66–70 (2014).
Dorsch, M. & Stackebrandt, E. Some modifications in the procedure of direct sequencing of PCR amplified 16S rDNA. J. Microbiol. Methods 16, 271–279 (1992).
Yamada, R. et al. Development of PCR for identifying Streptococcus parasuis, a close relative of Streptococcus suis. J. Vet. Med. Sci. 80, 1101–1107 (2018).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Saitou, N. & Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Silva, L. M. G. et al. Virulence-associated gene profiling of Streptococcus suis isolates by PCR. Vet. Microbiol. 115, 117–127 (2006).
Hanley, J. A. If nothing goes wrong, Is everything all right?. J. Am. Med. Assoc. 249, 1743 (1983).
Acknowledgements
The study was funded by the Department of Agriculture–Philippine Agriculture and Fisheries Biotechnology Program (DA-BIOTECH) under the Project Code No: DABIOTECH-R2002 and was supported by the University of the Philippines Los Baños (UPLB) and National Institute of Molecular Biology and Biotechnology (BIOTECH). The authors extend their gratitude to the local government units and personnel of the provinces of Albay, Batangas, Bohol, Cebu, Iloilo, Marinduque, Misamis Occidental, Misamis Oriental, and Zamboanga del Norte for their active assistance during the sampling process. Acknowledgement is also due to Abigail D. Perez, Mark Angelo D. Ipac, and Zyrah Marie M. Zapanta for their assistance in the processing and finalization of sequences for MLST. Special appreciation is also extended to Dr. Leila Fazeldehkordi and Mr. Mark Lorenz P. Mangosing for their expert assistance in data curation and production of visual materials relevant to this study. Finally, the authors would also like to acknowledge Dr. Merlyn S. Mendioro for their insightful comments and suggestions, which greatly contributed to the improvement of this manuscript.
Author information
Authors and Affiliations
Contributions
S.A.S., B.B.I.S., N.A.T., and M.A.E conceptualized and developed designs of the methodology, S.A.S and B.B.I.S. supervised the experiments, M.G.C.T.C., C.G.A.R.L., A.M.E.S.D., and J.T.C. conducted the experiments, M.G.C.T.C., C.G.A.R.L., A.M.E.S.D.G., N.A.T., and B.B.I.S. analyzed the results. S.A.S., M.A.E., M.G.C.T.C., A.M.E.S.D.G., N.A.T., B.B.I.S., and A.K. edited the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sedano, S.A., Cantalejo, M.G.C.T., Lapitan, C.G.A.R. et al. Epidemiology and genetic diversity of Streptococcus suis in smallhold swine farms in the Philippines. Sci Rep 13, 21178 (2023). https://doi.org/10.1038/s41598-023-48406-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48406-9
- Springer Nature Limited