Abstract
A comprehensive understanding of the extracellular matrix (ECM) is essential for developing biomimetic ECM scaffolds for tissue regeneration. As the periodontal ligament cell (PDLC)-derived ECM has shown potential for periodontal tissue regeneration, it is vital to gain a deeper understanding of its comprehensive profile. Although the PDLC-derived ECM exhibits extracellular environment similar to that of periodontal ligament (PDL) tissue, details of its molecular composition are lacking. Thus, using a multiomics approach, we systematically analyzed cultured mouse PDLC-derived ECM and compared it to mouse PDL tissue as a reference. Proteomic analysis revealed that, compared to PDL tissue, the cultured PDLC-derived ECM had a lower proportion of fibrillar collagens with increased levels of glycoprotein, corresponding to an immature ECM status. The gene expression signature was maintained in cultured PDLCs and was similar to that in cells from PDL tissues, with additional characteristics representative of naturally occurring progenitor cells. A combination of proteomic and transcriptomic analyses revealed that the cultured mouse PDLC-derived ECM has multiple advantages in tissue regeneration, providing an extracellular environment that closely mimics the environment in the native PDL tissue. These findings provide valuable insights for understanding PDLC-derived ECM and should contribute to the development of biomimetic ECM scaffolds for reliable periodontal tissue regeneration.
Similar content being viewed by others
Introduction
The periodontal ligament (PDL) is a highly specialized fibrous tissue that anchors the tooth to the alveolar bone. The PDL maintains its thin non-mineralized layer despite existing between two mineralized tissues (i.e., the tooth root cementum and alveolar bone). The PDL is important for oral function, including the dissipation of masticatory force, tooth eruption, and neuronal feedback1. Various pathological conditions, such as periodontitis and occlusal trauma, irreversibly damage the PDL, resulting in tooth loss. Therefore, regeneration of the PDL is essential to restore the structure and function of the periodontium and prevent tooth loss.
The extracellular matrix (ECM) is a principal tissue component that plays a pivotal role in tissue function. It primarily provides a structural framework for tissues whose mechanical properties rely on tissue-specific ECM composition and their secondary structures2,3. Another important function of the ECM is to provide mechanical and biochemical cues to cells as an extracellular microenvironment4. Because the ECM plays a key role in regulating cell behavior, and the composition and organization of the ECM vary depending on the tissue type, it is important to understand the tissue-specific ECM profile in the context of tissue regeneration.
The major ECM components of the PDL are fibrillar collagens (e.g., type I and III), minor collagens (e.g., types V, VI, and XII), proteoglycans, and ECM glycoproteins1. We recently reported the comprehensive ECM profile of mouse molar PDL using laser microdissection and mass spectrometry-based proteomic analysis with ECM-oriented data curation5. Our data revealed that type I collagen accounted for 86.5% and types III and XII collagens each accounted for 4.5% of the total collagen. The most abundant proteoglycan/ECM glycoprotein is periostin (POSTN), followed by lumican (LUM), asporin (ASPN), biglycan (BGN), and tenascin-N (TNN)5. Notably, the ECM proportion, together with the morphological characteristics of collagen fibers, was significantly changed by mechanical stress induced by an orthodontic appliance5. These data are in accordance with those of previous proteomic analyses showing dynamic changes in the ECM profile under different physiological environments6,7. Hence, altered ECM proportions in different physiological states may modulate cell behavior, playing essential roles in the homeostasis and regeneration of the periodontal tissue.
Recent advances in tissue engineering offer various approaches for the regeneration of the PDL8. One approach is the use of periodontal ligament cell (PDLC)-derived ECM, which is obtained through the decellularization of cultured PDLCs. The PDLC-derived ECM maintains its structural integrity, preserves growth factors, and supports the repopulation of allogeneic cells9. It was found that culturing PDLCs on PDLC-derived ECM maintained their stemness and osteogenic potential better than culturing on tissue culture polystyrene10. This method also induced osteoblastic differentiation of allogeneic MSCs11 and led to the differentiation of induced pluripotent cells into PDL stem cell-like cells12. Because of such cell instructive features, the PDLC-derived ECM is considered to preserve the original ECM composition of native PDL tissue to some extent. However the detailed profile of the PDLC-derived ECM remains elusive. Therefore, in the present study, we aimed to systematically analyze the PDLC-derived ECM and compare it with that in the native PDL tissue in mice. To gain a comprehensive and detailed understanding, we employed a multiomics approach, integrating data from different omics technologies (namely proteomics and transcriptomics).
Results
Proteomic characterization of cultured mouse PDLC-derived ECM
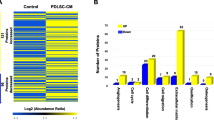
To elucidate the proteomic profile of the PDLC-derived ECM, we conducted an ECM-oriented proteomic analysis on cultured PDLCs (Fig. 1a). The ECM-enriched fraction of cultured PDLCs (Cell ECM) was prepared by decellularization and analyzed using liquid chromatography with tandem mass spectrometry. ECM-oriented data curation was performed using the Matrisome database13. Previously reported proteomic data of mouse PDL tissue dissected using laser microdissection5 were reanalyzed and used for the Tissue ECM. The number of detected proteins in Cell ECM was 459, while that in Tissue ECM was 655 (Fig. 1b). The number of matrisome proteins in Cell and Tissue ECM was 101 and 88, respectively. Forty-seven out of 142 matrisome proteins in total were detected in both Cell and Tissue ECM. Matrisome proteins accounted for 36.1% of the entire proteome in Cell ECM and for 56.1% of that in Tissue ECM (Fig. 1c). The relative proportions of the matrisome subclasses, collagens, proteoglycans, and ECM glycoproteins in Cell ECM were 43.8%, 12.4%, and 25.1%, respectively (Fig. 1d). Compared to Tissue ECM, the proportion of collagens was significantly lower in Cell ECM, while that of other matrisome subclasses (i.e., proteoglycans, ECM glycoproteins, ECM regulators, ECM-affiliated proteins, and secreted factors) was significantly higher (Fig. S1).
Matrisome protein profiles of cultured mouse PDLCs and PDL tissue. (a) Experimental setting of the proteomic analysis. (b) Number of proteins detected in the ECM-rich fraction of cultured PDLCs (Cell) and native PDL tissue (Tissue). The number of detected proteins in the entire proteome and that in matrisome is shown separately. (c) Occupancy of matrisome proteins in the Cell and Tissue ECM within the entire proteome. (d) Matrisome protein profiles of the Cell and Tissue ECM. Occupancy was calculated based on the exponentially modified protein abundance index (emPAI) value of each matrisome subclass divided by the sum emPAI value of total matrisome proteins.
The detailed protein proportion in each matrisome subclass showed that the most abundant collagen in Cell ECM was type I collagen, occupying 31.2% of the matrisome, while that in Tissue ECM was 81.5% (Fig. 2a). The second major collagen in Cell ECM was type XII, followed by types VI, V, and III (Fig. S2). The major proteoglycans in Cell ECM were ASPN, LUM, BGN, and prolargin (PRELP), each accounting for 2%–3% of the matrisome. The most abundant ECM glycoprotein in Cell ECM was POSTN, which accounted for 12.5% of the matrisome and ranked second in abundance in Cell ECM matrisome.
Protein profile of matrisome subclasses in cultured mouse PDLCs and PDL tissue. Detailed profiling of the (a) core matrisome and (b) matrisome-associated proteins in the Cell and Tissue ECM. Occupancy was calculated based on the emPAI value of each matrisome protein divided by the sum emPAI value of total matrisome proteins. Protein order in each matrisome subclass corresponds to the descending order of emPAI value in the Cell ECM.
The matrisome-associated proteins were also more abundant in Cell ECM compared to Tissue ECM (Figs. 2b, S3). HSP47 (Seripnh1) is the major ECM regulator in both Cell and Tissue ECM. The high-temperature requirement (Htr) family of serine proteases (HTRA1 and HTRA3) and collagen-modifying enzymes (LOX, LOXL1, and LOXL2) were detected in Cell ECM, but not in Tissue ECM. A noticeable amount of galectin-1 (LGALS1) was detected in Cell ECM, accounting for 5.7% of the matrisome. Annexin-family proteins (ANXA1 and ANXA2), Ca2+ phospholipid-binding proteins, were detected in both Cell and Tissue ECM with similar occupancy. The S100 family (S100A4, S100A6, S100A13, and S100A11) of secreted factors was detected both in the Cell and Tissue ECM, with the Cell ECM showing a higher proportion of these factors.
Overrepresentation analysis using the differentially expressed proteins (DEPs) in cultured mouse PDLC-derived ECM
The number of collagens and proteoglycans in the detected proteins in Cell and Tissue ECM was similar, while the number of unique ECM glycoproteins was two-fold higher in Cell ECM than that in Tissue ECM (Fig. 3a). Many of the identified matrisome-associated proteins, namely ECM regulators, ECM-affiliated proteins, and secreted factors, were unique in either Cell or Tissue ECM.
Enrichment analysis of differentially expressed matrisome proteins in cultured mouse PDLCs and PDL tissue. (a) Number of detected proteins in matrisome subclasses in Cell and Tissue ECM. Matrisome proteins detected in both the Cell and Tissue ECM, and those only detected in either group are separately shown as Shared and Unique, respectively. Volcano plot of the differentially expressed proteins (DEPs) in the Cell ECM vs. those in the Tissue ECM. The matrisome proteins are shown in red dots. Top enrichment network of the overrepresentation analysis of (c) upregulated matrisome proteins and (d) downregulated matrisome proteins. FDR; false discovery rate, FC; fold change, pAdj; adjusted p value.
DEPs were identified based on the proportion of individual proteins within a defined threshold (Fig. 3b). Many of the upregulated DEPs were matrisome proteins, whereas downregulated DEPs contained the most abundant collagen types in Tissue ECM: type I and III collagens. Overrepresentation analysis showed that the assembly of collagen fibrils and other multimeric structures (R-MMU-2022090, pAdj = 2.8E−25) and ECM-receptor interaction (mmu04512, pAdj = 7.5E−21) were the most enriched terms for upregulated DEPs (Fig. 3c). Integrin cell–surface interactions (R-MMU-216083, pAdj = 3.3E−07) showed the highest enrichment score for downregulated DEPs (Fig. 3d).
Gene expression profiling of mouse PDLCs
Cells under culture are affected by an extracellular environment composed of their own secreted ECM in the same way that cells in tissues are affected by their native ECM as an extracellular environment. To this end, the gene expression of cultured cells reflects the effect of their secreted ECM on the cells. To investigate the gene expression profile of cultured mouse PDLCs, total RNA was isolated from the cells after 7D and 14D of culture. Total RNA directly prepared from the PDL tissue of the extracted teeth was used as a control. Gene expression in cultured PDLCs and in cells of the PDL tissue was analyzed using RNA sequencing (RNA-seq), and subsequently pathway and process enrichment analyses were performed. A schematic representation of the experimental workflow is shown in Fig. 4a. Principal component analysis showed that the samples were significantly different from one another, with the 7D and 14D PDLCs showing a close association (Fig. 4b). Using k-means clustering with the top 2000 significant genes, we showed that 571 genes were upregulated and 1404 were downregulated in cultured PDLCs compared with that in PDL tissue cells (Fig. S4a). Gene ontology (GO) analysis of differentially expressed genes (DEGs) showed that the upregulated genes in cultured PDLCs shared enrichment terms related to the ECM (Fig. S4b, d) and the extracellular space (Fig. S4c). On the contrary, the most enriched terms for the downregulated genes were keratinization (Fig. S4b) and ribosomal genes (Fig. S4c, d).
Gene expression profiling of cultured mouse PDLCs using native mouse PDL tissue as reference. (a) Experimental workflow of the comprehensive gene expression analysis. (b) Principal component analysis showing that the groups were significantly different from one another. Volcano plot of the differentially expressed genes (DEGs) in the (c) Cell (14D) vs. Tissue, (d) Cell (7D) vs. Tissue, (e) Cell (14D) vs. Cell (7D). (f) Heatmap showing k-means clustering of differentially expressed matrisome genes. Expression values of each biological replicate are indicated according to a color gradient. Cluster 1 represents 166 upregulated matrisome genes on cultured PDLCs compared to native PDL tissue. Cluster 3 represents 102 downregulated matrisome genes on cultured PDLCs. Top enrichment network of the overrepresentation analysis on (g) upregulated matrisome genes and (h) downregulated matrisome genes. FDR; false discovery rate, FC; fold change, pAdj; adjusted p value.
Network enrichment analysis using differentially expressed matrisome genes of cultured mouse PDLCs
GO analysis strongly indicated that the major difference between cultured PDLCs and PDL tissue at the transcriptomic level was the ECM status; therefore, we focused on the ECM and its regulatory genes (matrisome genes)13 to gain a deeper understanding of these differences. Differentially expressed matrisome genes were identified between the groups and are shown in the volcano plot (Fig. 4c, d). Significant differences in gene expression patterns were observed between the 14D PDLCs and PDL tissue samples (Fig. 4c) and between the 7D PDLCs and PDL tissue samples (Fig. 4d), whereas the difference between the 7D and 14D PDLCs was minor (Fig. 4e). Among the matrisome genes identified (a total of 997 genes), upregulated genes (Cluster 1; 166 genes) and downregulated genes (Cluster 3; 102 genes) were identified using k-means clustering (Fig. 4f). Overrepresentation analysis revealed that the ECM–receptor interaction (mmu04512, pAdj = 1.0E−21) was most significantly enriched for upregulated matrisome genes (Fig. 4g). The ESC Pluripotency Pathways (WP339, pAdj = 8.2E−09), mainly controlled by the FGF and Wnt signaling pathways, were identified as the third most enriched pathway in upregulated matrisome genes. Neutrophil chemotaxis (GO:0030593, pAdj = 9.5E−17) was the most significantly enriched pathway in the downregulated genes (Fig. 4h). Notably, the IL-17 signaling pathway (mmu04657, pAdj = 1.0E−12) as well as cytokines and inflammatory response (WP222, pAdj = 1.6E−08), both related to the immunological response, did not exist under the culture conditions and were enriched in downregulated matrisome genes. In addition, the positive regulation of osteoblast differentiation (GO:0045669, pAdj = 6.8E−06) was enriched for downregulated genes.
Multiomic profiling of ECMs in cultured mouse PDLCs
To further analyze the details of ECMs and their regulators in cultured PDLCs, protein and gene expression changes were comparably analyzed (Figs. 5 and 6). The protein proportion of type I collagen significantly decreased (Fig. 5a), while its gene expression (Col1a1 and Col1a2) increased (Fig. 5b). Both the protein proportion and gene expression of type III collagen (Col3a1) decreased, while those of type XII collagen increased. The proportion of major proteoglycans and ECM glycoproteins significantly increased (Fig. 6a). In contrast, their gene expression levels showed an unstable pattern (Fig. 6b). The expression of Postn, the gene encoding the second most abundant protein in the Cell ECM, did not change significantly. The expression of Aspn and Dcn decreased, while that of Bgn increased. Correlation analysis showed a significant correlation between changes in gene expression and the protein proportion of the core matrisome (Fig. 6c). Discrepancies in gene expression and protein abundance were observed mainly in high-abundance proteins in the Cell and Tissue ECM, such as COL1A1, POSTN, TNN, ASPN, and LUM.
Protein and gene expression of collagens in cultured mouse PDLCs and PDL tissue. (a) Relative protein abundance of collagens in the Cell ECM compared with that in the Tissue ECM. (b) Relative gene expression of collagens in cultured PDLCs compared with that in cells from native PDL tissues. 7D, cultured PDLCs on day 7; 14D, cultured PDLCs on day 14. *p < 0.05, **p < 0.01.
Protein and gene expression of proteoglycans/ECM glycoproteins in cultured mouse PDLCs and PDL tissue. (a) Relative protein abundance of proteoglycans/ECM glycoproteins in the Cell ECM compared with that in the Tissue ECM. (b) Relative gene expression of proteoglycans/ECM glycoproteins in cultured PDLCs compared with that in cells from native PDL tissues. (c) Correlation analysis of gene and proteomic changes of the core matrisome in cultured PDLCs. 7D, cultured PDLCs on day 7; 14D, cultured PDLCs on day 14. *p < 0.05, **p < 0.01.
Post-translational modifications of collagen are essential for the integrity of the ECM of PDL tissue14. We observed an increased abundance of collagen-modifying enzymes (LOX, LOXL, and LOXL2) in Cell ECM compared with that in Tissue ECM (Fig. 3b). To gain further insights into the secretion and maturation of collagen in cultured PDLCs, we analyzed the gene expression of collagen-modifying enzymes known to play essential roles in collagen biosynthesis15 (Fig. S5). All Lox family proteins (encoded by Lox, Loxl1-4) showed significantly higher gene expression in cultured PDLCs than that in PDL tissue. Gene expression of all lysyl hydroxylases (encoded by Plod1-3) and essential binding partners for lysyl hydroxylase 2 (Plod2), HSP47 (Serpinh1) and Fkbp65 (Fkbp10), increased in the cultured PDLCs. Among the prolyl 4-hydroxylases (P4ha1-3, P4hb), the gene expression of P4ha1, P4ha3, and P4hb was increased, while that of P4ha2, the predominant form in chondrocytes, osteoblasts, and endothelial cells16, was unchanged.
Overrepresentation analysis indicated diminished osteoblastic differentiation of PDLCs in culture (Fig. 4h). We analyzed proteomic and gene expression changes in osteoblast/cementoblast-associated ECM (Fig. 7). The abundance of osteopontin (OPN/Spp1) and bone sialoprotein (BSP/Ibsp) tended to decrease (Fig. 7a). The large SD values of OPN and BSP may be due to the contamination of bone/cementum tissue during the laser microdissection procedure. The expression of Spp1 and Ibsp was significantly decreased in cultured PDLCs (Fig. 7b), while that of other osteoblast/cementoblast differentiation markers, including Alpl, Dspp, and Bglap, was decreased.
Protein and gene expression of osteoblast/cementoblast differentiation-associated components in mouse PDLCs and PDL tissue. (a) Relative abundance of osteoblast/cementoblast differentiation-associated proteins in the Cell ECM compared with that in the Tissue ECM. (b) Relative expression of osteoblast/cementoblast differentiation-associated genes in cultured PDLCs compared with that in cells from native PDL tissues. 7D, cultured PDLCs on day 7; 14D, cultured PDLCs on day 14. *p < 0.05, **p < 0.01.
Gene expression profiling of cultured mouse PDLCs
Gene expression of transcription factors critical for osteoblast/cementoblast differentiation, osterix (Sp7) and Cbfa1/Runx2 (Runx2), and important for PDL homeostasis, scleraxis (Scx) and mohawk (Mkx), was analyzed (Fig. 8a). The gene expression of Sp7 and Runx2 significantly decreased, consistent with that of osteoblast/cementoblast markers (Fig. 7b). Gene expression of Scx increased in cultured PDLCs whereas that of Mkx decreased on day 7 but increased on day 14.
Gene expression of key transcriptional factors and stem cell markers in cultured mouse PDLCs and PDL tissue. Relative gene expression of (a) key transcriptional factors and (b) stem cell markers in cultured PDLCs compared with that in cells from native PDL tissue. 7D, cultured PDLCs on day 7; 14D, cultured PDLCs on day 14. *p < 0.05, **p < 0.01.
Overrepresentation analysis generated the enrichment term ESC Pluripotency Pathways (WP339, pAdj = 8.2E−09), indicating enhanced stem cell properties or an increased proportion of stem cells in cultured PDLCs (Fig. 4g). We selected stem cell marker genes, which are reported to have stem cell-like characteristics in PDL tissue (Fig. 8b). Gene expression of MSC markers CD90 (Thy1), αSMA (Acta2), and CD29 (Itgb1), and neural crest cell markers p75NTR (Ngfr) and Sox10, significantly increased in cultured PDLCs. In contrast, the gene expression of Gli1, a Wnt signal responsive gene, and PTHrP (Pthlh), confirmed to play crucial roles in tooth root development, was significantly decreased in cultured PDLCs.
Discussion
The ECM is a complex network of proteins that provides mechanical and biochemical signaling cues to guide cell behavior4. The composition and organization of the ECM vary depending on the tissue and environment; therefore, understanding these differences is important in the context of tissue regeneration. Here, we systemically analyzed the PDLC-derived ECM using a multiomics approach. The major change in the matrisome protein profile of the cultured PDLCs was the lower proportion of fibrillar collagens (type I and III collagens) compared to that in PDL tissue. Fibrillar collagens play a key role in tissue integrity and the tensile behavior of the PDL, whereas the rest of the ECM components, such as proteoglycans and ECM glycoproteins, are responsible for the viscoelastic response to compressive force17. Therefore, a lower proportion of fibrillar collagens with an increased proportion of glycoproteins results in significant changes in mechanical properties (e.g., tensile strength and elastic modulus). This also affects cell behavior, as the ECM provides an extracellular microenvironment to the surrounding cells18. We recently reported changes in the matrisome protein profile of the mouse PDL in response to mechanical stress5. Similar to the cultured PDLC-derived ECM, decreased levels of fibrillar collagens with increased glycoprotein levels have been observed during orthodontic tooth movement, in which mechanical stress-induced tissue remodeling actively takes place. Furthermore, the levels of matrisome proteins that are preferentially expressed in the PDL, such as POSTN19, TNN20, ASPN21, and S100A422, tended to increase in the ECM of cultured PDLCs. Our results indicate that the cultured PDLC-derived ECM preserves PDL-specific proteins and has characteristics similar to those of PDL tissue with active collagen fibrogenesis.
Previous studies have shown differences between the gene expression profiles of PDLCs in culture and in native PDL tissues. The cells under culture are influenced by their secreted ECM, and as a result, the gene expression in cultured PDLCs may partially represent the impact of their own-secreted ECM on the cells. In a recent study comparing the gene expression profiles of cultured PDLCs and cells in the PDL tissue of extracted human teeth using cap analysis of gene expression, the expression of many ECM genes, including Postn, Aspn, osteomodulin (Omd), Tnn, Spp1, and osteocalcin (Ocn), was remarkably decreased in cultured PDLCs23. Another study comparing human PDL tissue and cultured PDLCs using microarray reported that Alpl, Ibsp, Postn, Fmod, Spp1, and Omd were highly expressed in the PDL tissue, whereas gremlin (Grem1), follistatin (Fst), CTGF, and FGF5 were expressed at higher levels in cultured PDLCs24. Subtractive hybridization analysis revealed that type I and III collagen, Lum, Postn, Aspn, and follicular dendritic cell-secreted protein (Fdcsp) were highly expressed in the PDL tissue compared with that in cultured PDLCs25. Our gene expression data obtained using RNA-seq are consistent with those of previous studies showing that the expression of genes encoding ECM glycoproteins (Postn and Tnn), proteoglycans (Aspn and Lum), and osteoblast/cementoblast differentiation proteins (Alpl, Ibsp, Spp1, and Ocn) decreased in the cultured PDLCs compared with that in the PDL tissue. Moreover, our comprehensive gene expression analysis revealed that the major transcriptomic changes in cultured PDLCs were associated with the ECM. It remains unclear as to which cell population is responsible for the ECM turnover in the PDL tissue because the cells comprising the PDL tissue are heterogeneous and not well characterized1,26. The difference in the cell population between culture and in the native tissue also needs to be taken into consideration.
Type I collagen in the PDL undergoes distinct post-translational modifications that markedly affect the formation of intermolecular cross-links and determine the structural and mechanical properties of the tissue14,15. In the present study, collagen-modifying enzymes, such as LOX, LOXL1, and LOXL2 were detected only in the proteome datasets of the Cell ECM but not in the Tissue ECM (Fig. S3). In addition, the gene expression of collagen-modifying enzymes tended to increase in cultured PDLCs compared with that in the PDL tissue, which is indicative of activated collagen fibrogenesis (Fig. S5). We previously reported that mechanical stress accelerates the maturation of collagen cross-linking through post-translational modifications of type I collagen in human PDLCs27. Because cultured PDLCs are free from occlusal loading, some biomechanical cues that are crucial for ECM organization and maturation may be lacking. Although collagen-modifying enzymes are increased in cultured PDLCs, fibrillar collagens are most likely less cross-linked in the cultured PDLCs than in the PDL tissue, owing to the absence of occlusal loading. Furthermore, an enrichment pathway of ECM–receptor interactions in upregulated DEPs indicated accelerated ECM turnover because cell–ECM interactions generally stimulate ECM degradation28. Increased expression of MMP-2, a gelatinase with a wide substrate range, was observed in cultured PDLCs at both the mRNA and protein levels. In accordance with the matrisome profiles, the ECM of cultured PDLCs has an immature status with increased turnover.
Among the well-characterized transcriptional factors in PDL, the gene expression of Cbfa1/Runx229, Scx30, and Mkx31 was well maintained, while that of Sp732,33 was significantly decreased in cultured PDLCs. It has been shown that the deletion of Sp7 in Col1- or Ocn-expressing cells in mice results in a short root with thin root dentin33, while the conditional deletion of Cbfa1/Runx2 in Gli1-lineage cells completely inhibits tooth root formation29. Despite a slight difference in the Cre-driver system, deletion of Cbfa1/Runx2 resulted in a more severe phenotype compared to Sp7, indicating that Cbfa1/Runx2 is essential for tooth root development and also most likely for its maintenance. Therefore, the decreased expression of Sp7 in the cultured PDLCs may not significantly affect PDLC characteristics. Notably, the gene expression of Mkx and Scx showed the opposite pattern in 7D PDLCs; the expression of Scx increased, while that of Mkx decreased. This observation is consistent with a recent study showing the mutually exclusive role of Scx and Mkx in PDL, as revealed using the single-cell RNA-seq of Mkx-KO mice34.
As the PDL is abundant in tissue stem cells, and they contribute substantially to periodontal tissue regeneration, the identification and characterization of the tissue stem cells in the PDL has attracted considerable research interest35. Cultured PDLCs express several stem cell markers, including CD29/Itgb1, CD90/Thy1, and Stro-136. In addition, lineage-tracing studies have shown that αSMA/Acta2- 37, Axin2- 38,39, Prrx1- 40,41, Gli1- 42, PTHrP/Pthlh43, and CD90/Thy1-expressing cells39 contribute to PDL tissue maintenance. Furthermore, cultured PDLCs had approximately 50-fold higher Stro-1 protein levels than cells in PDL tissue24, and all the expanded cells isolated from the molar surface were Gli1-lineage cells42. Therefore, cultured PDLCs represent a naïve cell population; possibly, they are naturally occurring progenitor cells24. In our experimental settings, the cultured PDLCs were affected by the ECM produced by them. Consistent with the previous notion, our pathway enrichment analysis indicated that the ESC pluripotency pathway (WP339), mainly controlled by FGF and Wnt signaling, was enhanced in cultured PDLCs. Gene expression data also indicated a significant increase in MSC markers (CD90/Thy1, aSMA/Acta2, CD29/Itgb1, and Prrx1) and neural crest cell markers (p75NTR/Ngfr and Sox10). In contrast, the expression of genes expressed by well-characterized stem cell-like populations in PDL, Gli1, and PTHrP/Pthlh, significantly decreased. Our results partially support the notion that cultured PDLCs represent naturally occurring progenitor cells24, capable of accelerating tissue regeneration. Collectively, the PDLC-derived ECM possibly guides the cells toward a tissue progenitor cell state.
Currently, label-free proteomics is a versatile protein-quantification method in bottom-up proteomics44. Label-free spectral counting-based protein quantification relies on the abundance of tryptic peptides in each sample. Because larger proteins generate more peptides in theory, the spectral counts of these proteins tend to be overrepresented. In the present study, we used exponentially modified protein abundance index (emPAI), a spectral counting-based relative protein quantification technique, in which the number of observed peptides was divided by the number of theoretically observable peptides per protein45. We previously reported the matrisome profile of mouse PDL tissue using another spectral counting-based quantification technique, normalized spectral abundance factor (NSAF), which normalized the spectral counts with respect to the protein length46. Despite the same proteome data set being used, the collagen proportion of the Tissue ECM matrisome was 43.8% in the current study, but it was 55 to 58% in the previous report using NSAF5. Although it is difficult to ascertain the algorithm that is most suitable for evaluating the matrisome profile, it should be noted that the difference in the normalization algorithm affects relative protein quantification in spectral counting-based protein quantification.
To the best of our knowledge, our study is the first to show a comprehensive profile of cultured mouse PDLC-derived ECM in reference to the mouse PDL tissue. A combination of proteomic and transcriptomic analyses revealed that the mouse PDLC-derived ECM displayed an immature status with active turnover. Moreover, in cultured PDLCs, the gene expression signature is maintained and is similar to that in cells from PDL tissues, with additional characteristics representative of tissue progenitor cells. These observations strongly suggest that PDLC-derived ECM has multiple advantages in periodontal tissue regeneration because it provides an extracellular environment that closely mimics the environment in the native PDL tissue with active ECM turnover, promoting tissue regeneration. Furthermore, cell culture is an indispensable tool for analyzing biological systems in a well-controlled environment. Therefore, understanding the differences between the in vitro and in vivo ECM statuses is crucial. Because our data clearly show the difference between the ECM of cultured PDLCs and that of the native PDL tissue, we can manipulate the ECM of cultured PDLCs to the tissue PDL end by adjusting culture conditions, for example, by supplementing growth factors, applying mechanical force, changing oxygen tension, or using a combination of these. The matrisome profiling that we performed in this study can also be used to evaluate the quality of cultured PDLC-derived ECM for optimal tissue regeneration. Although further studies are required to validate our findings, the multiomics characterization of the PDLC-derived ECM presented in this study provides further insights into developing tissue-specific biomimetic ECM scaffolds for periodontal tissue regeneration.
Methods
Ethics statement
The Niigata University Animal Experiment Ethics Committee reviewed and approved the animal experiment protocol (Approved number: SA00532). All animal handling and experiments strictly followed the ARRIVE guidelines for animal research and reporting for in vivo experiments. All methods were performed in accordance with the relevant guidelines and regulations.
Cell culture
C57BL/6 J mice (male, 5 weeks old, n = 8) were euthanized by carbon dioxide inhalation followed by cervical dislocation. The first and second molars of each quadrant were extracted, and the PDL tissue was dissociated with Liberase DL (Roche, Basel, Switzerland) for 60 min at 37 °C on a horizontal shaker at 100 rpm. The released PDLCs were collected by centrifugation (1000 rpm, 5 min) and maintained in alpha-MEM (Invitrogen, Waltham, MA, USA) containing 10% fetal bovine serum, 1% antibiotics, and antimycotics (Invitrogen). The PDLCs were then seeded onto a 35 mm dish (4 × 105 cells/dish) and cultured for 1 week. The culture medium was supplemented with ascorbic acid (50 µg/mL; Sigma-Aldrich, St. Louis, MO, USA).
Transcriptome analysis
Total RNA from cultured cells was isolated after seven and 14 days (7D and 14D, respectively, n = 4 each) of culture using the TRIzol reagent (Invitrogen). To isolate total RNA from PDL tissue cells, the molars were harvested as described earlier and treated with TRIzol reagent (n = 14). Next-generation sequencing library was prepared using the MGIEasy RNA Directional Library Prep Set (MGI Tech, Shenzhen, China). Poly(A) mRNA was isolated using the NEBNext Poly(A) mRNA Magnetic Isolation Module (New England Biolabs, Ipswich, MA, USA). Libraries with different indices were multiplexed and loaded onto an MGI DNBSEQ G-400 instrument (MGI Tech) with a 2 × 150-bp paired-end configuration. The sequences were processed and analyzed by Azenta Life Sciences (Tokyo, Japan).
Laser microdissection
Previously reported proteomic data of the mouse PDL tissue dissected using laser microdissection5 were reanalyzed and used in this study. In brief, the mesial and distal sides of the distal root PDL of the upper first molar were dissected from toluidine blue-stained cryosections using an LMD 7000 microdissection system (Leica, Wetzlar, Germany). Approximately 20 cryosections were collected from each tooth (n = 8) for a total of ~ 160 slices to obtain enough protein for proteomic analysis. Each proteomic experiment comprised five biological replicates. Samples were kept at − 80 °C for proteome analysis.
Proteome analysis
Mouse PDLCs were cultured for 2 weeks in the presence of ascorbic acid (n = 5). The cultures were collected using RIPA buffer (Wako, Osaka, Japan), and the ECM-rich fraction was prepared by centrifugation at 12,000 rpm for 30 min. The precipitate was thoroughly washed with phosphate-buffered saline and double distilled water, lysed with lysis buffer (100 mM Tris–HCl, pH 8.8, 7 M urea, 2% sodium dodecyl sulfate), sonicated, and the protein amount was quantified using the Pierce™ BCA Protein Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). After reduction with TCEP-HCl (Thermo Fisher Scientific) and alkylation with iodoacetamide (Sigma-Aldrich), tryptic peptides were generated via trypsin digestion.
Trypsin-digested peptides were dissolved in 0.3% formic acid and filtered through a 0.45-μm Ultrafree MC membrane filter (Merck-Millipore, Billerica, MA, USA). Samples containing 0.13 μg peptide were analyzed using direct-injection mode on an Eksigent NanoLC 415 nano-flow liquid chromatography system (Sciex, Framingham, MA, USA) using a 75 µm × 150 mm C18 spray-tip column (3 μm, 120 Å; Nikkyo Technos, Tokyo, Japan) coupled with a TripleTOF 5600 + tandem mass spectrometer (Sciex). Separation involved a 60 min gradient elution using 98% A, 2% B to 68% A, and 32% B at 300 nL/min (A and B refer to mobile phases comprising 0.1% aqueous formic acid and 0.1% formic acid in acetonitrile, respectively). The MS spectrum (250 ms) and the following 10 MS/MS spectra (100 ms each) were acquired in data-dependent mode. The dataset for the mouse PDL tissue obtained using laser microdissection (JPST001418/PXD030430, https://repository.jpostdb.org/entry/JPST001418) was previously published5.
Raw data generated using Analysis TF 1.6 (Sciex) were converted to Mascot generic files using the MS Data Converter (Sciex) and searched against a mouse reference protein sequence database (UniProt, Mus musculus) using the Mascot search engine (v.2.6; Matrix Science, Boston, MA, USA). The peptide and MS/MS tolerances were set at ± 20 ppm and ± 0.1 Da, respectively. A maximum of one missed cleavage was allowed. The search settings involved cysteine carbamidomethylation as a fixed modification and the following variable modifications: deamidation of asparagine and/or glutamine, N-terminal glutamine to pyroglutamate modification, and oxidation of methionine, proline, and lysine. The target false discovery rate (FDR) was set to < 1%. Proteins detected in at least two out of five samples were considered identified.
Bioinformatics
For bioinformatics analysis, matrisome genes/proteins were selected from all identified proteins using the Matrisome Annotator (http://matrisomeproject.mit.edu/analytical-tools/matrisome-annotator/)47. The occupancy of each matrisome protein was calculated based on the emPAI value45 divided by the sum emPAI value of the total matrisome proteins. The matrisome profile was determined as the mean of five biological replicates. Statistical significance was calculated using the Benjamini–Hochberg procedure48 with the following thresholds: genes, log (fold change) >|0.2| and log (FDR) > 2; proteins, log (fold change) >|0.08| and log (FDR) > 2. Statistically significant genes/proteins were considered differentially expressed. Overrepresentation analysis was performed using eVITTA v1.3.149 based on the following ontology sources: KEGG Pathway, GO Biological Processes, GO Cellular Components, GO Molecular Functions, Reactome Gene Sets, and WikiPathways. k-Means clustering was conducted using iDEP 1.050. The top 2000 statistically significant genes were clustered (n = 3) based on their expression patterns across all samples and visualized as a heatmap. Overrepresentation analysis was performed in each cluster using eVITTA v1.3.1. The relative abundance of protein was calculated based on the occupancy of each matrisome protein, as described earlier. The occupancy of each protein in PDLCs in culture was divided by that in the PDL tissue. To calculate the relative gene expression, the transcripts per million (TPM) values of RNA-seq data were used. The TPM value for PDLCs in culture was divided by that for the PDL tissue to obtain the relative gene expression.
Statistical analysis
The results are expressed as the means ± standard deviations (SDs) from four or five independent experiments. Statistical analysis for differences between groups was performed with a two-tailed unpaired t-test with Welch’s correction using Prism 9 (GraphPad Software, San Diego, CA, USA), and a p value of < 0.05 was considered statistically significant.
Data availability
The datasets generated and/or analyzed during the current study are available in the Japan ProteOme STandard Repository/Database with the data set identifier JPST001418/ PXD030430 (https://repository.jpostdb.org/entry/JPST001418), or from the corresponding author on reasonable request.
References
Beertsen, W., McCulloch, C. A. & Sodek, J. The periodontal ligament: A unique, multifunctional connective tissue. Periodontology 2000(13), 20–40 (1997).
Frantz, C., Stewart, K. M. & Weaver, V. M. The extracellular matrix at a glance. J. Cell Sci. 123, 4195–4200 (2010).
Mouw, J. K., Ou, G. & Weaver, V. M. Extracellular matrix assembly: A multiscale deconstruction. Nat. Rev. Mol. Cell Biol. 15, 771–785 (2014).
Kumar, A., Placone, J. K. & Engler, A. J. Understanding the extracellular forces that determine cell fate and maintenance. Development 144, 4261–4270 (2017).
Thant, L. et al. Extracellular matrix-oriented proteomic analysis of periodontal ligament under mechanical stress. Front. Physiol. 13, 899699 (2022).
Denes, B. J., Ait-Lounis, A., Wehrle-Haller, B. & Kiliaridis, S. Core matrisome protein signature during periodontal ligament maturation from pre-occlusal eruption to occlusal function. Front. Physiol. 11, 174 (2020).
Connizzo, B. K. et al. Nonuniformity in periodontal ligament: Mechanics and matrix composition. J. Dent. Res. 100, 179–186 (2021).
Wang, X., Chen, J. & Tian, W. Strategies of cell and cell-free therapies for periodontal regeneration: The state of the art. Stem Cell. Res. Ther. 13, 536 (2022).
Farag, A. et al. Decellularized periodontal ligament cell sheets with recellularization potential. J. Dent. Res. 93, 1313–1319 (2014).
Zhang, J. C., Song, Z. C., Xia, Y. R. & Shu, R. Extracellular matrix derived from periodontal ligament cells maintains their stemness and enhances redifferentiation via the wnt pathway. J. Biomed. Mater. Res. A 106, 272–284 (2018).
Farag, A. et al. The effect of decellularized tissue engineered constructs on periodontal regeneration. J. Clin. Periodontol. 45, 586–596 (2018).
Hamano, S. et al. Extracellular matrix from periodontal ligament cells could induce the differentiation of induced pluripotent stem cells to periodontal ligament stem cell-like cells. Stem Cells Dev. 27, 100–111 (2018).
Naba, A. et al. The matrisome: In silico definition and in vivo characterization by proteomics of normal and tumor extracellular matrices. Mol. Cell Proteomics 11, M111 014647 (2012).
Hudson, D. M., Garibov, M., Dixon, D. R., Popowics, T. & Eyre, D. R. Distinct post-translational features of type I collagen are conserved in mouse and human periodontal ligament. J. Periodontal Res. 52, 1042–1049 (2017).
Kaku, M. & Yamauchi, M. Mechano-regulation of collagen biosynthesis in periodontal ligament. J. Prosthodont. Res. 58, 193–207 (2014).
Gjaltema, R. A. & Bank, R. A. Molecular insights into prolyl and lysyl hydroxylation of fibrillar collagens in health and disease. Crit. Rev. Biochem. Mol. Biol. 52, 74–95 (2017).
Ortun-Terrazas, J., Cegonino, J. & Perez Del Palomar, A. In silico study of cuspid’ periodontal ligament damage under parafunctional and traumatic conditions of whole-mouth occlusions. A patient-specific evaluation. Comput. Methods Programs Biomed. 184, 105 (2020).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Horiuchi, K. et al. Identification and characterization of a novel protein, periostin, with restricted expression to periosteum and periodontal ligament and increased expression by transforming growth factor beta. J. Bone Miner. Res. 14, 1239–1249 (1999).
Nishida, E. et al. Transcriptome database KK-Periome for periodontal ligament development: expression profiles of the extracellular matrix genes. Gene 404, 70–79 (2007).
Yamada, S. et al. PLAP-1/asporin, a novel negative regulator of periodontal ligament mineralization. J. Biol. Chem. 282, 23070–23080 (2007).
Duarte, W. R. et al. cDNA cloning of S100 calcium-binding proteins from bovine periodontal ligament and their expression in oral tissues. J. Dent. Res. 77, 1694–1699 (1998).
Hosiriluck, N., Kashio, H., Takada, A., Mizuguchi, I. & Arakawa, T. The profiling and analysis of gene expression in human periodontal ligament tissue and fibroblasts. Clin. Exp. Dent. Res. 8, 658–672 (2022).
Lallier, T. E. & Spencer, A. Use of microarrays to find novel regulators of periodontal ligament fibroblast differentiation. Cell Tissue Res. 327, 93–109 (2007).
Nakamura, S. et al. Identification of genes preferentially expressed in periodontal ligament: specific expression of a novel secreted protein, FDC-SP. Biochem. Biophys. Res. Commun. 338, 1197–1203 (2005).
Mizukoshi, M. et al. In vivo cell proliferation analysis and cell-tracing reveal the global cellular dynamics of periodontal ligament cells under mechanical-loading. Sci. Rep. 11, 9813 (2021).
Kaku, M. et al. Mechanical loading stimulates expression of collagen cross-linking associated enzymes in periodontal ligament. J. Cell. Physiol. 231, 926–933 (2016).
Lu, P., Takai, K., Weaver, V. M. & Werb, Z. Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb. Perspect. Biol. 3, e005058 (2011).
Wen, Q. et al. Runx2 regulates mouse tooth root development via activation of WNT inhibitor NOTUM. J. Bone Miner. Res. 35, 2252–2264 (2020).
Takimoto, A. et al. Scleraxis and osterix antagonistically regulate tensile force-responsive remodeling of the periodontal ligament and alveolar bone. Development 142, 787–796 (2015).
Koda, N. et al. The transcription factor mohawk homeobox regulates homeostasis of the periodontal ligament. Development 144, 313–320 (2017).
Takahashi, A., Ono, N. & Ono, W. The fate of Osterix-expressing mesenchymal cells in dental root formation and maintenance. Orthod. Craniofac. Res. 20(Suppl 1), 39–43 (2017).
Kim, T. H. et al. Osterix regulates tooth root formation in a site-specific manner. J. Dent. Res. 94, 430–438 (2015).
Takada, K. et al. Single cell RNA sequencing reveals critical functions of Mkx in periodontal ligament homeostasis. Front. Cell Dev. Biol. 10, 795441 (2022).
Bartold, P. M. & Gronthos, S. Standardization of criteria defining periodontal ligament stem cells. J. Dent. Res. 96, 487–490 (2017).
Tomokiyo, A. et al. Detection, characterization, and clinical application of mesenchymal stem cells in periodontal ligament tissue. Stem Cells Int. 2018, 5450768 (2018).
Roguljic, H. et al. In vivo identification of periodontal progenitor cells. J. Dent. Res. 92, 709–715 (2013).
Xie, X. et al. Axin2(+)-mesenchymal PDL cells, instead of K14(+) epithelial cells, play a key role in rapid cementum growth. J. Dent. Res. 98, 1262–1270 (2019).
Zhao, J., Faure, L., Adameyko, I. & Sharpe, P. T. Stem cell contributions to cementoblast differentiation in healthy periodontal ligament and periodontitis. Stem Cells 39, 92–102 (2021).
Bassir, S. H. et al. Prx1 Expressing cells are required for periodontal regeneration of the mouse incisor. Front. Physiol. 10, 591 (2019).
Gong, X. et al. Tracing PRX1(+) cells during molar formation and periodontal ligament reconstruction. Int. J. Oral. Sci. 14, 5 (2022).
Men, Y. et al. Gli1+ periodontium stem cells are regulated by osteocytes and occlusal force. Dev. Cell 54, 639-654 e636 (2020).
Takahashi, A. et al. Autocrine regulation of mesenchymal progenitor cell fates orchestrates tooth eruption. Proc. Natl. Acad. Sci. USA 116, 575–580 (2019).
Arike, L. & Peil, L. Spectral counting label-free proteomics. Methods Mol. Biol. 1156, 213–222 (2014).
Ishihama, Y. et al. Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein. Mol. Cell Proteomics 4, 1265–1272 (2005).
Zybailov, B. et al. Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J. Proteome Res. 5, 2339–2347 (2006).
Naba, A. et al. The extracellular matrix: Tools and insights for the “omics” era. Matrix Biol. 49, 10–24 (2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Cheng, X., Yan, J., Liu, Y., Wang, J. & Taubert, S. eVITTA: A web-based visualization and inference toolbox for transcriptome analysis. Nucleic Acids Res. 49, W207–W215 (2021).
Ge, S. X., Son, E. W. & Yao, R. iDEP: An integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinform. 19, 534 (2018).
Acknowledgements
We are grateful to D. Kobayashi (Center for Research Promotion, School of Medicine, Niigata University) for technical assistance. This work was conducted using the research equipment shared in the MEXT Project for promoting public utilization of advanced research infrastructure (program for supporting the introduction of the new sharing system) at Niigata University. Parts of the figures were created using BioRender.com. The RNA sequencing was processed and analyzed by Azenta Life Sciences (Tokyo, Japan).
Author information
Authors and Affiliations
Contributions
Conceptualization, M.Ka., and L.T.; Investigation, L.T., A.D., Y.O., M.Ki, M.Mi, M.A., H.I., and K.K.; Formal Analysis, M.Ka.; Writing—Original Draft, M.Ka., and L.T.; Writing—Review and Editing, M.Ma, I.S., and K.U.; Funding Acquisition, M.Ka.; Resources, M.Ma, and Y.K.; Supervision, M.Ma, I.S., and K.U.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kaku, M., Thant, L., Dobashi, A. et al. Multiomics analysis of cultured mouse periodontal ligament cell-derived extracellular matrix. Sci Rep 14, 354 (2024). https://doi.org/10.1038/s41598-023-51054-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-51054-8
- Springer Nature Limited