Abstract
Highly pathogenic avian Influenza virus (HPAIV) has spread in an unprecedented extent globally in recent years. Despite the large reports of cases in Asia, Europe, and North America, little is known about its circulation in South America. Here, we describe the isolation, and whole genome characterization of HPAIV obtained from sampling 26 wild bird species in Peru, representing one of the largest studies in our region following the latest HPAIV introduction in South America. Out of 147 samples analyzed, 22 were positive for detection of avian influenza virus using a qRT-PCR-based assay. Following inoculation into embryonated chicken eggs, fourteen viral isolates were obtained from which nine isolates were selected for genome characterization, based on their host relevance. Our results identified the presence of HPAIV H5N1 subtype in a highly diverse wild bird species. Phylogenetic analysis revealed that these isolates correspond to the clade 2.3.4.4b, sharing a common ancestor with North American isolates and forming a monophyletic group along with isolates from Chile. Altogether, changes at the amino acid levels compared to their closest relatives indicates the virus is evolving locally, highlighting the need for constant genomic surveillance. This data evidence the chances for spillover events increases as the virus spreads into large populations of immunologically naïve avian species and adding conditions for cross species transmission.
Similar content being viewed by others
Introduction
Influenza A viruses are a major threat for animal and public health due to their evolutionary properties leading to the emergence of distinct novel viruses. The underlying mechanisms for evolution in Influenza viruses rely on both viral genetics and their animal hosts factors. In terms of viral genetics, its genome comprises of single-stranded, eight negative-sense viral RNAs segments1. Viral genome encodes for eleven proteins including some accessory proteins2. These eight segments are named and presented from the largest to shortest, as follows: PB2, PB1, PA, hemagglutinin (HA), nucleoprotein (NP), neuraminidase (NA), matrix (M), non-structural (NS)3,4. These proteins have different functions during the viral replication process. Thus, viral genome segmentation is the basis for reassortments, and recombination as evolutionary mechanisms commonly found in influenza viruses. Furthermore, influenza viruses are known to infect a wide host range5,6 which is the basis for spillover events and viral adaptation to novel species1. Altogether, influenza viruses are of the most rapidly evolving pathogens of major concern for animal and public health7.
Hemagglutinin (HA) and Neuraminidase (NA) are important viral proteins playing fundamental roles in viral replication8,9 and are used for Influenza virus classification. HA is a relevant protein for viral entry into target cells, therefore it is the main target for neutralizing antibodies and vaccine development. On the other hand, NA is a tetrameric protein with enzymatic activity that allow viral release from infected cells following replication10. In addition to their relevance in viral replication, these proteins are used for viral classification purposes. Hence, there are sixteen (16) HA and nine (9) NA described to date, in wild birds belonging to the Anseriformes and Charadriiformes orders11,12,13. In addition, there is a description of two HA and NA subtypes of influenza-like viruses found in bats from Peru14,15.
Furthermore, avian influenza is classified into low and highly pathogenic avian influenza viruses based on the HA cleavage site and the clinical severity of infections in chickens16. Thus, most avian influenza viruses harboured by wild birds correspond to low pathogenicity avian influenza viruses (LPAIVs) causing asymptomatic or mild infections in poultry4. On the other hand, some H5 and H7 lineages are known to produce a severe infection leading to massive death when domestic birds are infected and, specially H5 subtypes, causing death in their wild bird hosts. Thus, the co-circulation of Low- and High-pathogenicity AIVs in wild bird species is a contributing factor for the emergence of novel Influenza viruses with distinctive features.
Highly pathogenic avian influenza viruses (HPAIVs) have caught the attention from scientists due to its concern in animal and public health17. Since the emergence of the HPAIV H5N1 GS/Gd lineage in China in 1996, the virus has evolved into ten clades and multiple subclades18. More recently, one of these genetic clades, such as the 2.3.4.4b clade, represents the main influenza virus type causing the current outbreak around the world18,19,20,21. Despite this virus characterization in multiple geographical areas, little is known about the circulation of the HPAIV H5N1 in South America.
Since 2006, we have implemented a local system for monitoring avian influenza viruses in wild birds in Peru. This has led to the detection, isolation, and characterization of multiple AIV isolates that corresponded to those LPAIVs22,23,24. Nevertheless, the detection of HPAIV H5 was reported on November 14th in 2022, following an outbreak of high mortality in pelicans of Paita, Piura (North coast of Peru) by the national authorities. Since the arrival of this HPAIV, more than 55,000 wild birds have died due to these HPAIV H5 infections and 46 outbreaks in poultry farms have been controlled. More recently, spillover events from avian into marine wildlife have been reported by the National Service of Natural Protected areas (SERNANP). Thus, our objective was the identification, isolation, and whole genome characterization of this HPAIV detected in Peru, in wild birds from Peru. Our study represents the first complete genetic description of this virus in South America in a large group of local wild bird species.
Results
Avian influenza virus is detected in a large group of wild bird species with clinical presentation of highly pathogenic avian influenza (HPAIV) infection
Samples analysed were obtained from oropharyngeal, cloacal and fecal swabs from animals as part of our active surveillance for influenza viruses or those showing clinical sings of HPAIV infection in a collaboration with SERFOR (Servicio Nacional Forestal y de Fauna Silvestre). In total, we evaluated 147 samples that corresponded to 26 wild bird species ranging from migratory birds, waterfowl and some local birds that are part of the natural ecosystem in the pacific coast of Peru. A large proportion of the samples analyzed corresponded to Franklin’s gulls (Leucophaeus pipixcan), Peruvian pelicans (Pelecanus thagus), Sanderlings (Calidris alba), American oystercatchers (Haematopus palliates), mallard ducks (Anas platyrhynchos) among others. A list of the species and number of samples evaluated is presented in Table 1. Following sample collection, PCR showed that 22 out of 147 samples (14.9%) resulted positive for avian influenza. Cts values of these samples ranged from 12 to 30 cycles out of the 45 PCR cycles protocol. Positive samples were processed for inoculation into embryonated chicken eggs for viral isolation.
Avian influenza virus was successfully isolated in embryonated eggs from wild bird species
Embryonated chicken eggs have shown to be the ideal support for avian influenza virus isolation25. Here, we inoculated the 22 qPCR-positive samples in eggs selected previously. Out of those inoculated, 14 samples had evidence of viral replication through embryo death and hemagglutination assay. These samples were obtained from Peruvian pelican (3/6), Belcher’s gull (3/4), Harris’s hawk (1/3), American kestrel (1/2), Guanay cormorant (1/1), Humboldt penguin (1/1), Franklin’s gull (1/2), Peregrine falcon (1/1), Peruvian booby (1/1) and Western barn owl (1/1). We did not obtain the viral isolate from the Chesnut-eared aracari sample that was positive by PCR. These findings were detected in a single round of inoculation. A summary of the results is presented in Table 2. Following isolation, positive samples were selected based on the frequency of wild bird species reported by the National authorities and their migratory capacity, for whole genome characterization.
Highly pathogenic Avian influenza virus H5N1 clade 2.3.4.4b was identified and characterized in wild birds
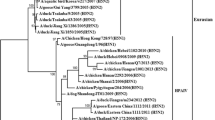
Genetic analysis allows the subtyping of influenza viruses based on the types of HA and NA proteins. Moreover, HA amino acid sequences show differential patterns of cleavage sites to differentiate highly and low pathogenic AIVs26. Here, we selected 09 isolates for subtyping and genomic reconstruction. Genetic analysis at the nucleotide and amino acid levels identified that our isolates corresponded to the H5N1 subtypes. Furthermore, cleavage sites at the HA regions identified sequences corresponding to those highly pathogenic avian influenza viruses. Hence, we found the multi basic motif (PLREKRRKR↓GLF) in all highly pathogenic avian influenza viruses. A geographical map showing the sample origin distribution is shown in Fig. 1. Furthermore, we took advantage of a Bayesian model to estimate the most recent common ancestor and define the phylogenetic relationship of the H5N1 viruses identified here compared with those described in America, Asia and Europe. Based on H5 phylogenetic reconstruction, our results showed that our isolates corresponded to highly pathogenic avian influenza clade 2.3.4.4b, sharing a common ancestor with isolates obtained from snow goose in North Dakota and Kansas, in the US, 2022. Furthermore, tree topology suggested that this virus is forming a monophyletic group along with those isolated from Chile, within the 2.3.4.4b clade, and appears to accumulate changes in this protein. Phylogenetic trees of the HA and NA proteins are shown in Fig. 2 and 3.
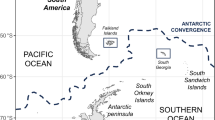
Geographical map of avian influenza isolates and species harbouring HPAIV H5N1 avian influenza virus clade 2.3.4.4b. Red dots represents the locations where isolates were obtained in Lima and La Libertad, two major cities in the coastal area in Perú (A). Pictures identifying the wild bird species source of the isolates characterized in our study are presented (B). (Geographical map was designed and prepared by Geog. José More-Bayona as requested).
Phylogenetic analysis of H5 gene shows the virus circulating in Peru corresponds to the clade 2.3.4.4b. We collected 588 H5 complete sequences from GenBank encompassing those from Asia, Europe, and America. A phylogenetic analysis was performed using the Markov Chain Monte Carlo (MCMC) algorithm in Mr. Bayes. A general time reverse (GTR) substitution model with gamma distribution was used. Our Peruvian sequences (red asterisks) grouped together with those isolates from Chile (blue) forming a monophyletic group with a common ancestor in the US.
Phylogenetic analysis of N1 gene from the Peruvian isolates has a common ancestor in the US. We collected 567 N1 complete sequences from Genbank encompassing those from Asia, Europe and America. A phylogenetic analysis was performed using the Markov Chain Monte Carlo (MCMC) algorithm in Mr. Bayes. A general time reverse (GTR) substitution model with gamma distribution was used. Our Peruvian sequences (red asterisks) grouped together with those isolates from Chile (blue) forming a monophyletic groups with a common ancestor in the US.
Genetic analysis of HA shows unique changes in the HA of HPAIV H5N1 from wild birds in Peru
Following phylogenetic reconstruction, we evaluated whether the HPAIV circulating in Peru is presenting some relevant changes compared to those members of the 2.3.4.4b. Hence, we identified some changes that were absent in those observed in most isolates from the US, while others were present in those closely related to our Peruvian isolates. Genetic distances in HA ranged from 99.08 to 99.5% in the samples sequenced. A summary of the genetic distances among all the genome segments with their closest relatives is presented in Table 3. Amino acid changes were identified in 7 out of 8 viral segments, in all nine isolates compared to their most recent ancestor. These changes suggest that our isolates along with those closely related from Chile share a common ancestor with a North American isolate, while depicting a subclade within the 2.3.4.4b phylogroup. Further studies are required to assess whether these changes are relevant for HA function or vaccine evasion. Amino acid changes observed in all nine isolates are presented in Table 4.
Discussion
Anseriformes and Charadriiformes are considered the natural reservoirs for avian influenza viruses, while migratory birds are responsible for viral spread in large geographical areas, playing a significant role in the emergence of novel viral forms. Currently, the emergence of HPAIV H5N1 clade 2.3.4.4b affecting multiple species of wild and domestic birds has been described in Asia, Europe, and America, becoming of major concern for animal and public health. Despite this, little is known about the viral circulation and genetic characterization in South America.
In Peru, the national service for animal health (SENASA) reported the first detection of a HPAIV infection in November 2022. This case was reported in a coastal city of the Northern Peru, following the occurrence of Peruvian pelicans found death in the Paita beach in Piura. Our initial findings suggested that a viral infection was responsible for causing severe disease in the animals evaluated. Our results allowed the identification of avian influenza viruses with a highly variable viral load based on the Ct values observed. Although Ct values can be used as approximation method to assess the viral load for other viral infections27, we did not find evidence of correlation in terms of severity and viral levels based on Ct values (data not shown). Thus, clinical presentation and the PCR results indicate that influenza virus was likely responsible for the severe cases. Further genetic characterization was required to establish the presence of a highly pathogenic avian influenza virus.
Genomic sequencing and genotyping of these viruses detected the presence of HPAIV H5N1 clade 2.3.4.4b, affecting a broad range of species. We speculate that these species play a differential role into the viral spread since some of them appeared to be more affected than others. Moreover, movement dynamics of these species might add contributing factors in viral dissemination into local areas. Despite differences in the host source, our isolates showed a high degree of genetic conservation. Our isolates belonging to a monophyletic group share a common ancestor with isolates from Chile. However, some genetic changes have been detected compared to those isolates. Hence, genetic surveillance needs to be reinforced in the upcoming months to closely track the viral evolution in our region. Evolutionary analysis shows that these viruses have been transmitted to the US from Europe and Asia and are co-circulating with other viral lineages in the Americas. It is well known, the H5N1 viruses circulating are reassortants of viruses with different origins28 and these events might continue in South America.
Avian influenza viruses in Peru have been tracked in wild birds since 2006 when we established a viral surveillance program that allowed the detection and characterization of only LPAIVs prior to the current outbreak. Thus, we identified that these LPAI viruses arose from common ancestors of viruses isolated in North America during the 2019 and 2022 period22. These findings indicate that migratory routes of wild birds play a major role into the viral entry in the South American region and are the potential route for those highly pathogenic avian influenza viruses as these birds arrive to this region in the summer period in the south hemisphere6,29. As expected, our current results confirmed this since these HPAIVs shared a common ancestor with those in the US.
Furthermore, although this current outbreak represents the first introduction of the HPAIV clade 2.3.4.4b virus in the Peruvian ecosystem, we speculate that multiple introductions might occur in the upcoming migration periods. Further studies are required to define other multiple introductions of the virus and whether there are alternative routes of viral entry30. These conditions are the fundamental basis for the emergence of viruses with novel characteristics that might infect other species including mammals. Although no human-to-human transmission has been reported, national authorities have reported the occurrence of infections in marine mammals in Peru. Even though this is not under the scope of this paper, we emphasize that genetic surveillance of these spillover events must be monitored closely since the viral adaptation to these species might reveal the capacity for infection in humans and the human-to-human transmission.
Peru is one of the most biodiverse countries around the world. Thus, the entry and spread of HPAIVs, such as H5N1 clade 2.3.4.4b, into immunologically naïve wildlife populations is rapidly diminishing the local species, with major impact in those in an endangered situation. Our study shows that multiple local avian species are affected by this virus, and some of those studied here especially Peruvian pelicans (Pelecanus thagus), and Guanay cormorants (Phalacrocorax bougainvillii) and others have been placed into an even more endangered condition. We call the authorities to pay close attention to these species of ecosystem importance. In addition, to its intrinsic importance for the local ecosystem, they do play a major economic role. The guano produced by these species is commercialized as a natural fertilizer of great value. Thus, we urge the national and regional authorities to take actions in a coordinated manner to avoid the impact on these species.
In summary, our study represents the first report of isolation and genome characterization of HPAIV H5N1clade 2.3.4.4b from the largest wild bird species pool affected of severe disease in South America. Furthermore, this report shows evidence that despite the close genetic relationship of our isolates with North American strains, there are some nucleotide and amino acid changes detected in the virus circulating in Peru, indicating the potential for spillover events into other species.
Materials and methods
Samples
One hundred and forty-seven samples encompassing oropharyngeal, cloacal, and fecal swabs were collected and submitted to the Laboratory of Avian Pathology at the Universidad Nacional Mayor de San Marcos in Peru, between November 2022 and January 2023. These samples included those taken as part of our active surveillance of avian influenza in Peru, along with those obtained as a research collaboration with the Servicio Nacional Forestal y de Fauna Silvestre (SERFOR). All samples were taken in a universal transport media (VTM) tube and transported to the lab during the same collection day at 4 °C and processed immediately. In brief, all swabs were resuspended in 500 µl PBS with antibiotics, filtered through 0.22-micron syringe filters to remove bacterial contamination and centrifuged for clarification (8000×g for 10 min). Supernatants were saved for further molecular test by a specific PCR-based assay for detection of Influenza A viruses and isolation into embryonated chicken eggs. In some instances, samples were pooled into 4–5 samples according to geographical location, species, date, etc. Some live birds samples obtained from the National Forest and wildlife Service (SERFOR), presented neurological and respiratory clinical signs suggestive of influenza infections. A summary of the samples submitted is presented in Table 1 and video recordings of representative species affected by the infection is presented in the supplementary information (Sup S8 and S9). Protocols and methods were carried out in accordance and approved by the Ethics and animal welfare committee (CEBA) by the Faculty of Veterinary Medicine in the Universidad Nacional Mayor de San Marcos (CEBA202133). We confirm that the current study adheres to the ARRIVE guidelines31.
Viral detection
One hundred and forty (140) µL of samples processed were used for RNA extraction. For this purpose, we used the QIAamp Viral RNA Mini Kit (Qiagen, US), according to manufacturer’s instructions. For viral gene detection, we used the RealPCR Influenza A RNA mix for avian influenza A virus detection following the manufacturer’s instructions (IDEXX, US). The protocol, based on 45 amplification cycles, allow the identification of avian Influenza A viruses. The positive status was used as a primary criteria of sample selection for further isolation and sequencing procedures.
Viral isolation
Viral isolation was performed by inoculation of 0.2 mL of samples into the allantoid cavity of 9–11-days-old embryonated chicken eggs in five replicates25. Following inoculation, eggs were incubated at 37.5 °C for 4 days and checked daily to assess embryo survival. A round of inoculation was performed to increase the viral load for improving the metagenomic sequencing of all eight genomic segments. Following inoculation, an hemagglutination test was used as screening method to evaluate the presence of an hemagglutinating agent. In brief, allantoid fluid (25 uL) was serially diluted with two-fold volume of PBS (pH 7,4). This mix was incubated into an equal proportion of 1% chicken red blood cells (RBCs). After 20 min, the bottom formation or red blood cells agglutination was recorded. Following clarification (20,000×g for 10 min), positive samples were selected based on wild bird species host and viral load for genome sequencing.
Next generation sequencing
Positive samples for PCR against Influenza A virus and hemagglutination test following inoculation into embryonated eggs, were selected for genomic characterization using a metagenomic approach. First-strand cDNA was performed using the SuperScript™ III First-Strand Synthesis kit (Invitrogen™, US) with the FR20RV primer (5′-GCCGGAGCTCTGCAGATATC-3′) prior to library preparation. Library preparation was performed using Nextera XT method and sequenced on a MiSeq platform using paired 150 base read chemistry (Cambridge Technologies, Worthington, MN, USA).
Genome assembly and annotation
Raw data (fastq files) was evaluated based on Phred score using FastQC v. 0.11.9 software32, while adapters were removed using Trimmomatic version 0.3233. The genome was assembled using SPAdes Genome Assembler version 3.13.134 and annotated using Prokka genome annotation version 1.14.635, along with the use of the Bacterial and Viral Bioinformatics Research Center, version 3.28.2236. Gene annotation was performed using the tools of the Influenza virus genome database to address all eight genome segments. Where low coverage detected the sequences were deposited as partial CD and submitted to Genbank under the accession numbers: OQ747758-OQ747763 (HA), OQ747765-OQ747770 (NA), OQ747863-OQ747868 (PB2), OQ747878-OQ747883 (PB1), OQ747873-OQ747877 (PA), OQ747895-OQ747899 (MP), OQ747884-OQ747888 (NS), OQ747889-OQ747894 (NP).
Avian Influenza subtyping and phylogenetic analysis
Following genome assembly, we focused on the HA and NA subtyping. For this purpose, we collected the coding region of HA and NA genes from AIV viruses submitted to the National Center for bioinformatics information (NCBI), Influenza virus genome database (https://www.ncbi.nlm.nih.gov/genomes/FLU/Database/nph-select.cgi?go=genomeset). We only included in the analysis the coding complete sequences submitted from 2017 to 2023, isolates from avian species in Asia, Europe and America. The alignment was performed in MEGA X37, with posterior manual editing and files were saved as nexus files and used for phylogenetic reconstruction. We used the Markov Chain Monte Carlo (MCMC) algorithm in Mr. Bayes to estimate a Bayesian inference for phylogenetic reconstruction38. A general time reverse (GTR) substitution model with gamma distribution was used. Markov Chain Monte Carlo chains were run for 2 million iterations and sampled every 100 steps to allow all parameters converge with effective sample size values greater than 200, and with a burn-in of 0.25 to discard the 25% of samples at the beginning of the analysis. The consensus tree was summarized and further visualized using Figtree39. For the other six genome segments, sequences were retrieved from the NCBI database and selecting sequences collected from 2017 to date, for phylogenetic reconstruction, including sequences from America, Asia and Europe. A schematic representation of our methodology for detection, isolation and genome characterization is presented in Fig. 4.
Schematic representation of the general procedure for isolation and genome characterization of HPAIV H5N1 clade 2.3.4.4b. National authorities reported the outbreak of avian influenza virus during mid November 2022. Following this report, we performed a viral surveillance along a research collaboration with SERFOR to identify the virus involved in the current outbreak. Viral isolation and detection of avian influenza virus started on December 2022 and the genome characterization was performed with those positives for the hemagglutination and qRT-PCR assays. Currently, we are continuing with the surveillance program for analysis of the HPAIV evolution in Peru.
Data availability
Nucleotide sequences produced in the current study have been submitted to GenBank. Here are the accession numbers for each gene segment: HA: https://www.ncbi.nlm.nih.gov/nuccore/OQ747758, https://www.ncbi.nlm.nih.gov/nuccore/OQ747759, https://www.ncbi.nlm.nih.gov/nuccore/OQ747760, https://www.ncbi.nlm.nih.gov/nuccore/OQ747761, https://www.ncbi.nlm.nih.gov/nuccore/OQ747762, https://www.ncbi.nlm.nih.gov/nuccore/OQ747763, https://www.ncbi.nlm.nih.gov/nuccore/OR591549, https://www.ncbi.nlm.nih.gov/nuccore/OR591550, https://www.ncbi.nlm.nih.gov/nuccore/OR591551. NA: https://www.ncbi.nlm.nih.gov/nuccore/OQ747765, https://www.ncbi.nlm.nih.gov/nuccore/OQ747766, https://www.ncbi.nlm.nih.gov/nuccore/OQ747767, https://www.ncbi.nlm.nih.gov/nuccore/OQ747768, https://www.ncbi.nlm.nih.gov/nuccore/OQ747769, https://www.ncbi.nlm.nih.gov/nuccore/OQ747770, https://www.ncbi.nlm.nih.gov/nuccore/OR591758, https://www.ncbi.nlm.nih.gov/nuccore/OR591759, https://www.ncbi.nlm.nih.gov/nuccore/OR591760. PB2: https://www.ncbi.nlm.nih.gov/nuccore/OQ747863, https://www.ncbi.nlm.nih.gov/nuccore/OQ747864, https://www.ncbi.nlm.nih.gov/nuccore/OQ747865, https://www.ncbi.nlm.nih.gov/nuccore/OQ747866, https://www.ncbi.nlm.nih.gov/nuccore/OQ747867, https://www.ncbi.nlm.nih.gov/nuccore/OQ747868, https://www.ncbi.nlm.nih.gov/nuccore/OR591761, https://www.ncbi.nlm.nih.gov/nuccore/OR591762, https://www.ncbi.nlm.nih.gov/nuccore/OR591763. PB1: https://www.ncbi.nlm.nih.gov/nuccore/OQ747878, https://www.ncbi.nlm.nih.gov/nuccore/OQ747879, https://www.ncbi.nlm.nih.gov/nuccore/OQ747880, https://www.ncbi.nlm.nih.gov/nuccore/OQ747881, https://www.ncbi.nlm.nih.gov/nuccore/OQ747882, https://www.ncbi.nlm.nih.gov/nuccore/OQ747883, https://www.ncbi.nlm.nih.gov/nuccore/OR591525, https://www.ncbi.nlm.nih.gov/nuccore/OR591526, https://www.ncbi.nlm.nih.gov/nuccore/OR591527. NP: https://www.ncbi.nlm.nih.gov/nuccore/OQ747889, https://www.ncbi.nlm.nih.gov/nuccore/OQ747890, https://www.ncbi.nlm.nih.gov/nuccore/OQ747891, https://www.ncbi.nlm.nih.gov/nuccore/OQ747892, https://www.ncbi.nlm.nih.gov/nuccore/OQ747893, https://www.ncbi.nlm.nih.gov/nuccore/OQ747894, https://www.ncbi.nlm.nih.gov/nuccore/OR591730, https://www.ncbi.nlm.nih.gov/nuccore/OR591731, https://www.ncbi.nlm.nih.gov/nuccore/OR591732. PA: https://www.ncbi.nlm.nih.gov/nuccore/OQ747873, https://www.ncbi.nlm.nih.gov/nuccore/OQ747874, https://www.ncbi.nlm.nih.gov/nuccore/OQ747875, https://www.ncbi.nlm.nih.gov/nuccore/OQ747876, https://www.ncbi.nlm.nih.gov/nuccore/OQ747877, https://www.ncbi.nlm.nih.gov/nuccore/OR591529, https://www.ncbi.nlm.nih.gov/nuccore/OR591530, https://www.ncbi.nlm.nih.gov/nuccore/OR591531. MP: https://www.ncbi.nlm.nih.gov/nuccore/OQ747895, https://www.ncbi.nlm.nih.gov/nuccore/OQ747896, https://www.ncbi.nlm.nih.gov/nuccore/OQ747897, https://www.ncbi.nlm.nih.gov/nuccore/OQ747898, https://www.ncbi.nlm.nih.gov/nuccore/OQ747899, https://www.ncbi.nlm.nih.gov/nuccore/OR591733, https://www.ncbi.nlm.nih.gov/nuccore/OR591734, https://www.ncbi.nlm.nih.gov/nuccore/OR591735. NS: https://www.ncbi.nlm.nih.gov/nuccore/OQ747884, https://www.ncbi.nlm.nih.gov/nuccore/OQ747885, https://www.ncbi.nlm.nih.gov/nuccore/OQ747886, https://www.ncbi.nlm.nih.gov/nuccore/OQ747887, https://www.ncbi.nlm.nih.gov/nuccore/OQ747888, https://www.ncbi.nlm.nih.gov/nuccore/OR591754, https://www.ncbi.nlm.nih.gov/nuccore/OR591755, https://www.ncbi.nlm.nih.gov/nuccore/OR591756. Additional data is available upon request.
References
Webster, R. G., Bean, W. J., Gorman, O. T., Chambers, T. M. & Kawaoka, Y. Evolution and ecology of influenza A viruses. Microbiol. Rev. 56, 152 (1992).
Hutchinson, E. C. & Yamauchi, Y. Understanding influenza. Methods Mol. Biol. 1836, 1–21 (2018).
Bouvier, N. M. & Palese, P. The biology of influenza viruses. Vaccine 12, D49–D53 (2008).
Krammer, F. et al. Influenza. Nat. Rev. Dis. Primers 4, 1–21 (2018).
Bourret, V. Avian influenza viruses in pigs: An overview. Vet. J. 239, 7–14 (2018).
Olsen, B. et al. Global patterns of influenza A virus in wild birds. Science 2006(312), 384–388 (1979).
Medina, R. A. & García-Sastre, A. Influenza A viruses: New research developments. Nat. Rev. Microbiol. 9, 590–603 (2011).
Air, G. M. Influenza neuraminidase. Influenza Other Respir. Viruses 6, 245–256. https://doi.org/10.1111/j.1750-2659.2011.00304.x (2012).
Wilks, S., De Graaf, M., Smith, D. & Burke, D. A review of influenza haemagglutinin receptor binding as it relaets to pandemic properties. Vaccine 30, 4369–4376. https://doi.org/10.1016/j.vaccine.2012.02.076.A (2012).
Gamblin, S. J. & Skehel, J. J. Influenza hemagglutinin and neuraminidase membrane glycoproteins. J. Biol. Chem. 285, 28403–28409. https://doi.org/10.1074/jbc.R110.129809 (2010).
Slemons, R. D., Johnson, D. C., Osborn, J. S., Hayes, F. & Hayesd, F. Type-A influenza viruses isolated from Wild Free-Flying Ducks in California. Source Avian Dis. 18, 119–124 (1973).
Venkatesh, D. et al. Avian influenza viruses in wild birds: Virus evolution in a multihost ecosystem. J. Virol. 92, 1–20. https://doi.org/10.1128/jvi.00433-18 (2018).
Brown, J.D., Stallknecht, D.E. Wild Bird Surveillance for the Avian Influenza Virus. In Avian Influenza Virus. Methods in Molecular Biology (eds Spackman, E.) 85–97, Vol. 436 (2008).
Ma, W., García-Sastre, A. & Schwemmle, M. Expected and unexpected features of the newly discovered bat influenza A-like viruses. PLoS Pathog. 11, e1004819 (2015).
Tong, S. et al. A distinct lineage of influenza A virus from bats. Proc. Natl. Acad. Sci. U S A 109, 4269–4274. https://doi.org/10.1073/pnas.1116200109 (2012).
Alexander, D. J. & Brown, I. H. History of highly pathogenic avian influenza. Rev. Sci. Tech. 28, 19–38 (2009).
Fuller, T. L. et al. Mapping the risk of avian influenza in wild birds in the US. BMC Infect. Dis. 10, 1–13. https://doi.org/10.1186/1471-2334-10-187 (2010).
Lee, D. H., Bertran, K., Kwon, J. H. & Swayne, D. E. Evolution, global spread, and pathogenicity of highly pathogenic avian influenza H5Nx Clade 2.3.4.4. J. Vet. Sci. 18, 269–280 (2017).
Lewis, N. S. et al. Emergence and spread of novel H5N8, H5N5 and H5N1 Clade 2.3.4.4 highly pathogenic avian influenza in 2020. Emerg. Microbes Infect. 10, 148–151 (2021).
Mosaad, Z. et al. Emergence of highly pathogenic avian influenza A virus (H5N1) of clade 2.3.4.4b in Egypt, 2021–2022. Pathogens 12, 90. https://doi.org/10.3390/pathogens12010090 (2023).
Ndumu, D. et al. Highly pathogenic avian influenza H5N8 Clade 2.3.4.4B virus in Uganda, 2017. Infect. Genet. Evol. 66, 269–271. https://doi.org/10.1016/j.meegid.2018.10.010 (2018).
Castro-Sanguinetti, G. R. et al. Genetic subtyping and phylogenetic analysis of HA and NA from avian influenza virus in wild birds from Peru Reveals Unique Features among Circulating Strains in America. PLoS ONE https://doi.org/10.1371/journal.pone.0268957 (2022).
Ghersi, B. M. et al. Avian influenza in wild birds, Central Coast of Peru. Emerg. Infect. Dis. 15, 935–938. https://doi.org/10.3201/eid1506.080981 (2009).
Segovia, K., Icochea, E., González, R., Ghersi, B. & González, A. Presence of avian influenza virus in wild birds in the Wetlands of Puerto Viejo, Lima. Rev. Investig. Vet. Peru 24, 98–103 (2013).
Spackman, E. In Animal Influenza Virus: Methods and Protocols. Methods in Molecular Biology (eds Spackman, E.) 3rd Edn., Vol. 2123 (Humana Press, 2020).
Steinhauer, D. A. Role of hemagglutinin cleavage for the pathogenicity of influenza virus. Virology 258, 1–20 (1999).
Icochea, E. et al. Genetic analysis of infectious bronchitis virus S1 gene reveals novel amino acid changes in the GI-16 lineage in Peru. Microorganisms 11, 691. https://doi.org/10.3390/microorganisms11030691 (2023).
Shi, J., Zeng, X., Cui, P., Yan, C. & Chen, H. Alarming situation of emerging H5 and H7 avian influenza and effective control strategies. Emerg. Microbes Infect. 12, 2155072 (2023).
Hill, N. J. et al. Transmission of influenza reflects seasonality of wild birds across the annual cycle. Ecol. Lett. 19, 915–925 (2016).
Li, L. et al. Genetic evidence supports sporadic and independent introductions of subtype H5 low-pathogenic avian influenza A viruses from wild birds to domestic poultry in North America. J. Virol. https://doi.org/10.1128/jvi.00913-18 (2018).
du Sert, N. P. et al. Reporting animal research: Explanation and Elaboration for the Arrive Guidelines 2.0. PLoS Biol. 18, e3000411 (2020).
Simon Andrews FastQC: A Quality Control Tool for High Throughput Sequence Data 2010.
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069. https://doi.org/10.1093/bioinformatics/btu153 (2014).
Olson, R. D. et al. Introducing the Bacterial and Viral Bioinformatics Resource Center (BV-BRC): A resource combining PATRIC, IRD and ViPR. Nucleic Acids Res. 51, D678–D689. https://doi.org/10.1093/nar/gkac1003 (2023).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754 (2001).
Rambaut, A. Figtree v1.4. Molecular Evolution, Phylogenetics and Epidemiology (2012).
Acknowledgements
The authors would like to express their gratitude to the professionals from the National Service for Animal Health (SENASA) and Servicio Nacional Forestal y de Fauna Silvestre (SERFOR) especially to Ms. DVM Wendy Rojas and Mr. Renato Colán. We acknowledge the financial support from the Asociación Peruana de Avicultura (APA) and the Universidad Nacional Mayor de San Marcos.
Author information
Authors and Affiliations
Contributions
G.C-S., E.I., J.M-B., conceptualized the research plan. J.M-B., G.C-S., E.I., R.G-V., designed and developed the methodology. A. A-C., A.C-L., G.C-S., R.G-V. J.J., W.S. performed field investigation (sample collection and wild bird identification). G.C-S., E.I., R.G-V., performed and supervised the laboratory assays. J.M-B., G.C-S., E.I., R.G-V., composed the original draft of the manuscript. All authors contributed to the manuscript revision and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Castro-Sanguinetti, G.R., González-Veliz, R., Callupe-Leyva, A. et al. Highly pathogenic avian influenza virus H5N1 clade 2.3.4.4b from Peru forms a monophyletic group with Chilean isolates in South America. Sci Rep 14, 3635 (2024). https://doi.org/10.1038/s41598-024-54072-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54072-2
- Springer Nature Limited