Abstract
Pancreatic neuroendocrine tumors (PanNETs) are a heterogeneous group of tumors that exhibit an unpredictable and broad spectrum of clinical presentations and biological aggressiveness. Surgical resection is still the only curative therapeutic option for localized PanNET, but the majority of patients are diagnosed at an advanced and metastatic stage with limited therapeutic options. Key factors limiting the development of new therapeutics are the extensive heterogeneity of PanNETs and the lack of appropriate clinically relevant models. In that context, genomic sequencing of human PanNETs revealed recurrent mutations and structural alterations in several tumor suppressors. Here, we demonstrated that combined loss of MEN1, ATRX, and PTEN, tumor suppressors commonly mutated in human PanNETs, triggers the development of high-grade pancreatic neuroendocrine tumors in mice. Histopathological evaluation and gene expression analyses of the developed tumors confirm the presence of PanNET hallmarks and significant overlap in gene expression patterns found in human disease. Thus, we postulate that the presented novel genetically defined mouse model is the first clinically relevant immunocompetent high-grade PanNET mouse model.
Similar content being viewed by others
Introduction
Pancreatic neuroendocrine tumors (PanNETs) are a relatively rare neuroendocrine malignancy with an incidence of < 1 per 100,000 per year1 but account for approximately 2–5% of pancreatic malignancies and 6–7% of all neuroendocrine tumors and exhibit the most aggressive behavior among the latter. Although falsely considered benign neoplasia due to long periods of indolent growth, PanNETs are a heterogeneous group of tumors with often unpredictable and varying degrees of malignancy2,3. As many as 50–80% of PanNETs are associated with local and distal metastatic disease4. As a consequence of initially asymptomatic progression, the majority of patients are diagnosed at an advanced stage with limited therapeutic options. Thus, there is a critical unmet need for novel therapeutics and diagnostic modalities. Key factors limiting the development of new therapeutics are the extensive heterogeneity of PanNETs and the lack of appropriate clinically relevant in vivo models.
PanNETs are tumors that originate from endocrine cells within the islet of Langerhans5. These cells produce distinct hormones and are involved in many regulatory functions6. PanNETs that preserve hormone-producing attributes are called functional tumors (i.e. insulinomas, glucagonomas, somatostatinomas)7. Functional tumors often cause severe clinical syndromes, for example, insulinomas may lead to hypoglycemia1,5. In contrast, non-functional PanNETs do not secrete hormones and do not present with any specific clinical characteristics3,7. Despite these clinical differences, both functional and non-functional PanNETs can be aggressive, develop metastases, acquire treatment resistance, and have low overall survival rates7. The overall 5-year survival rate for PanNET patients is 53% with surgical resection. However, in patients with metastatic disease or for those unable to undergo surgery, the 5-year survival rate drastically drops to 23%1. For decades, surgery has remained the first line of defense8 and standard chemotherapeutics have been used in advanced patients with little to no avail9,10. Current targeted therapeutics include Sunitinib and Everolimus11,12, which show limited efficacy due to the development of acquired resistance13. Thus, there is a critical unmet need for translational tools to develop novel and effective precision therapeutics.
Recent sequencing efforts focused on elucidating the genomic landscape of PanNETs identified recurrent mutations and structural alterations in several tumor suppressors and chromatin modulators14,15,16. The most frequently mutated genes are MEN1 (multiple endocrine neoplasia type 1), ATRX (alpha-thalassemia/mental retardation X-linked), DAXX (death-domain associated protein), and PTEN (phosphatase and tensin homolog). Mutations in MEN1 are commonly associated with neuroendocrine malignancies and have been studied before17,18,19,20,21,22,23. MENIN is a scaffold protein encoded by the MEN1 gene that functions in multiple biological processes controlling gene expression, DNA damage repair, cell proliferation, and motility. Those factions are mediated by MENIN interactions with a plethora of proteins24,25; most notably, MENIN is a component of a histone methyltransferase KMT2A complex that methylates lysine 4 of Histone H3 and functions as a transcriptional regulator21,26. Although MEN1 loss is commonly associated with pancreatic neuroendocrine tumorigenesis, the deletion of MEN1 in mouse pancreatic beta-cells results in a relatively modest phenotype, with indolent tumors developing with long latency22. The second and third most common mutations found in human PanNETs are found in ATRX and DAXX, which together form a histone chaperone complex that deposits histone variant H3.3 into repetitive heterochromatin and telomeric regions of the genome27,28. Mutations in the ATRX-DAXX complex are found in a variety of tumors, including adult lower-grade gliomas, pediatric glioblastoma multiforme, pediatric adrenocortical carcinoma, osteosarcoma, and neuroblastoma28. Deletion of DAXX or ATRX in pancreatic neuroendocrine cells fails to trigger neoplastic transformation29,30,31,32. Additional mutations found commonly in human PanNETs target mTOR pathway genes, including PTEN, TSC2, and PIK3CA14,33. The PI3K-mTOR pathway is critical in the regulation of cell growth, apoptosis, differentiation, and migration and has been extensively studied in many solid tumors, including PanNETs. In addition to the loss of function mutations, half of the PanNET patients exhibit decreased PTEN expression and 35% with downregulation of TSC2, which significantly correlates with poor survival34.
Here, we developed and characterized a novel PanNET mouse model based on the combined deletion of Men1, Atrx, and Pten tumor suppressors.
Results
To identify key genetic alternations in human PanNETs, we analyzed the genomic sequencing database (GENIE v14.1)35. We found that mutations in tumor suppressors MEN1, ATRX, DAXX, and PTEN are the most frequent (Fig. 1A). Consistent with ATRX and DAXX forming one functional protein complex36, mutations in encoding genes are found mutually exclusive in PanNET samples (Fig. 1B). Next, we found that mutations in MEN1, PTEN and ATRX or DAXX co-occur with a high statistical likelihood, suggesting that co-mutation of those genes is one of the core determinants of pancreatic neuroendocrine tumors pathogenesis (Fig. 1B). To explore the tumorigenic potential of the combined loss of MEN1, ATRX, and PTEN in vivo, we interbred conditional Men1, Atrx and Pten knockout strains with pancreas-specific Pdx1-CreER strain (Fig. 2A). The resulting Men1LoxP/LoxP; AtrxLoxP/LoxP; PtenLoxP/LoxP; Pdx1-CreER mutant mice (hereafter termed MAP) were confirmed by PCR to carry the mutated alleles (Supplementary Fig. 1). PDX1 is expressed in pancreatic multipotent progenitor cells during embryonic development37,38, but is restricted to neuroendocrine cells in adult tissue39. To remain consistent with the onset of human PanNET in adults and avoid potential embryonic phenotype, we performed tamoxifen injections to induce Cre-mediated recombination in MAP mice at 2 months of age (Fig. 2B). Successful depletion of MENIN, ATRX and PTEN was confirmed in tissue lysates (Fig. 2C). Next, MAP mutant animals were aged and analyzed at 8 months post tumor induction and at humane endpoint when the animals become moribund. Based on gross pathology examination, all analyzed MAP animals developed macroscopic multifocal tumors of variable size in the pancreas (Fig. 2D). Further histological evaluation revealed a significant tenfold increase in pancreatic neuroendocrine cell area in the MAP model when compared to wildtype controls at 8 months and progressively larger tumors were observed at endpoint (Fig. 2D, E). To systematically assess PanNETs pathogenesis, we applied the WHO grade system based on the proliferative index (percent of Ki67 positive cells), which clinically serves as a robust prognostic factor in the multivariate analyses and independent predictor for patients survival outcomes40,41. The median Ki67 index of MAP tumors ranged from 7.2 to 14.8% (mean 10.9%) at the 8-month time point (Fig. 2F), which is consistent with intermediate grade 2 (G2) neuroendocrine tumors (Ki67 index of 3–20%). Of note, we performed similar tumor burden and proliferation analyses of various combinations of Men1, Atrx, and Pten double-knockouts (Supplementary Fig. 2). Our results indicate that combinations of Men1;Pten or Men1;Atrx deletion lead to the development of relatively benign well-differentiated grade 1 (G1) tumors (Ki67 index below < 3%), whereas Atrx;Pten does not exhibit signs of neoplastic transformation at 8 months post-induction (Supplementary Fig. 2A–C). Overall, these data support the notion that a combined loss of MEN1, ATRX, and PTEN represents an important step in PanNET pathogenesis.
Mutations of MEN1, ATRX, and PTEN co-occur in human PanNETs. (A) MEN1, ATRX, and PTEN tumor suppressor genes are frequently mutated and co-occur in human PanNETs. Oncoprint plot based on AACR GENIE Cohort v14.1 dataset (n = 413 patients). (B) Mutual exclusivity and co-occurrence of mutations in human PanNETs. In statistical analyses of the co-occurrence and exclusivity of the most common mutations found in PanNETs, the most notable findings were: (1) the mutual exclusivity between ATRX and DAXX, which encode proteins forming one functional complex, and (2) the statistically significant co-occurrence of MEN1, ATRX/DAXX and PTEN mutations in PanNET patients. For odds ratio calculations (see Methods)35. P values calculated by Fisher's exact test.
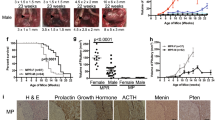
Combined deletion of Men1, Atrx, and Pten in pancreatic islet cells triggers the development of pancreatic neuroendocrine tumors in mutant mice. (A) Schematic illustrating conditional mutant alleles utilized in the generation of PanNET mouse model. To assess the effects of combined Men1LoxP/LoxP, Atrx LoxP/LoxP, and Pten LoxP/LoxP (MAP) ablation on neoplastic transformation of the pancreatic neuroendocrine cells. Tamoxifen-induced recombination of mutant alleles is mediated using pancreatic neuroendocrine cells-specific Pdx1-CreER strain. (B) Experimental design to investigate tumorigenesis in the MAP model following tamoxifen-induced mutant gene recombination (three intraperitoneal injections of 1 mg tamoxifen per mouse every other day) at 8 months and endpoint. (C) Representative macroscopic pathology, HE-stained sections and immunohistochemical (IHC) staining with indicated antibodies of pancreas tissue from normal (wildtype) and MAP mouse model at 8 months and endpoint. IHC stainings for insulin and Ki67 (a marker of proliferating cells) are shown. Scale bars, 1 cm (macroscopic), 200 µm (histology), 50 µm (Ki67 IHC), representative of n = 10 mice for each experimental group. (D) Quantification of neuroendocrine area per total pancreas area in wildtype and MAP mutant mice at indicated time points reveals significant expansion of neuroendocrine compartment. P-values were calculated by one-way ANOVA with Tukey’s multiple comparisons test. (E) Analysis of Ki67 index in normal pancreatic islets and MAP mouse model tumors at 8 months and endpoint, revealing a significant increase in tumor cell proliferation. P values were calculated by one-way ANOVA with Tukey’s multiple comparisons test. (F) Survival analysis of MAP mutant mice (n = 10, median survival of 363 days; see Supplementary Fig. 3A for detailed analysis of the clinical spectrum of disease observed in MAP mutant model). (G) Immunoblotting analysis with indicated antibodies of representative protein lysates of tissue biopsies from wildtype pancreas and MAP model tumors. Vinculin is shown as a loading control. (H) Fasting blood glucose levels in MAP mutant and wildtype (WT) control mice (males and females) at 8 months of age. P values were calculated by Student’s t test.
Next, we performed a survival study and found that MAP mice develop signs of morbidity and become moribund at approximately 12 months post-tumor induction (median survival of 351 days) (Fig. 2G). Analysis of tumors collected at the endpoint revealed further advancement of PanNET malignancy with a robust over twofold increase in tumor burden and approximately twofold increase in Ki67 index (mean 19.5%) compared to tumors collected at 8 months post-induction (Fig. 2D, E). Notably, 50% of mice at the endpoint developed high-grade tumors (G3) with a Ki67 index > 20% (Fig. 2F), indicative of progressive disease malignancy. However, we have not observed the formation of macroscopic metastasis at the endpoint analysis. Consistent with Pdx1-CreER expression in pancreatic β-cells, the MAP model developed PanNET insulinomas (see insulin expression (Fig. 2D). Expectedly, the advancing tumor burden was accompanied by the development of hypoglycemia as monitored by fasting blood glucose measurements (Fig. 2H). Congruent with prolonged hypoglycemia, we observed the deterioration of overall animal health, leading to moribund condition, including limb weakness, partial paralysis or tremors of the hindquarters, and weight loss (Supplementary Fig. 3A-B). In addition, in 1 out of 12 animals, we observed jaundice, which resulted in moribund condition. Together, these in vivo data support a key enabling function for a combined loss of MEN1, ATRX, and PTEN in the initiation and progression of PanNET.
To further characterize the tumors developed in MAP mutant mice, we performed RNA sequencing on micro-dissected tumor biopsies from three independent animals. We observed robustly increased expression of neuroendocrine and PanNET marker genes, including Insulin, Chromogranin A, Synaptophysin, PAX6, ISL1, SSTR2, ASCL1, and NCAM1, in MAP tumors compared to normal wildtype pancreas or pancreatic ductal adenocarcinoma biopsies from KrasG12D; p53Lox; Ptf1-Cre (KPC) mouse model (Fig. 3A). We further validated the expression of Chromogranin A, Synaptophysin, PAX6 and NCAM1 using immunohistochemistry staining on MAP model tumor sections (Fig. 3B). Elevated expression of these markers is consistent and diagnostic for human PanNETs42,43, thus indicating the MAP model recapitulates biology of pancreatic neuroendocrine malignancy.
Evaluation of neuroendocrine differentiation and PanNET markers expression in MAP mouse model tumors. (A) Heatmap of neuroendocrine differentiation and PanNET marker genes expression in RNA-seq datasets obtained from wildtype pancreas (n = 3), MAP model of PanNETs (n = 3), and KPC model of pancreatic ductal adenocarcinoma (PDAC, n = 3). (B) Representative immunohistochemistry images for Chromogranin A (CHGA), Synaptophysin (SYP), PAX6 and NCAM1 staining in wildtype pancreas and MAP tumors collected at 8 months. Scale bars, 200 µm.
Next, we utilized RNA-seq data in single-sample gene set enrichment analysis (ssGSEA) to gain insights into molecular mechanisms involved in MAP model tumorigenesis. The score derived from ssGSEA reflects the degree to which specific hallmark gene signatures and pathways are coordinately up or downregulated in analyzed samples. To provide relevant comparison groups, we utilized the same method to analyze RNA-seq datasets of tumor samples from: (1) human PanNETs44, (2) RIP-Tag2 PanNET mouse model45, and (3) KPC PDAC mouse model. Our computational analyses revealed a high level of agreement between transcriptional signatures found in human PanNETs and MAP model tumors, in particular, enrichment in oxidative phosphorylation, reactive oxygen species, fatty acid and xenobiotic metabolism, angiogenesis, MYC and MTOR pathways (Fig. 4A). Notably, MAP enriched molecular programs are consistent with human PanNET biology14,15,16. In contrast, we noted that tumors from RIP-Tag2 mutant mice, a commonly studied PanNET model, are characterized by transcriptional signatures distinct from human and mouse MAP tumors, for instance, different enrichment in pathways involved in cell proliferation (G2M checkpoint, mitotic spindle, E2F targets), xenobiotic metabolism and IL6-JAK-IL6-JAK-STAT3 signaling. As expected, human PanNETs and MAP model tumors exhibit distinct transcriptional programs from those found in the KRAS-driven pancreatic adenocarcinoma KPC model, including those associated with inflammatory response, allograft rejection, NFkB, STAT3, and KRAS pathways. The most characteristic of human and MAP PanNETs is the enrichment of neuroendocrine cell differentiation gene signature, which was absent in RIP-Tag2 and KPC models (Fig. 4A).
Transcriptional programs observed in MAP mouse model tumors are congruent with human PanNETs. (A) Single-sample gene set enrichment analysis (ssGSEA) of RNA-seq datasets of PanNETs obtained from patient samples (n = 22), MAP (n = 3), RIP-Tag2 (n = 31) and pancreatic ductal adenocarcinoma (PDAC) model KPC (n = 3). ssGSEA was performed using cancer hallmarks and neuroendocrine cell differentiation signature (GO0061101) from the MSigDB Molecular Signatures Database. RNA-Seq datasets of human PanNETs44 and RIP-Tag2 mouse model45 were previously published. (B) Spearman correlation analysis of transcription profiles between the human PanNET patient samples and MAP mouse PanNET tissues and. The values indicate Spearman correlation coefficient between variables.
Finally, we sought to investigate similarities in overall gene expression profiles of MAP tumors and PanNET samples representing localized and metastatic disease44. Spearman's correlation analysis revealed the statistically significant (p < 0.01) strong positive correlation (Spearman’s correlation coefficient > 0.7) between MAP and human PanNET transcriptomes (Fig. 4B). Altogether, molecular characterization indicates that MAP model tumors resembled molecular features of human PanNETs.
Discussion
Pancreatic neuroendocrine tumors are a rare and clinically challenging group characterized by high phenotypic and molecular heterogeneity. Although standard-of-care surgical resection8 and chemotherapy9,10 improved the prognosis of low-grade PanNETs, high-grade tumors are highly lethal. Therefore, efforts to decipher molecular mechanisms of PanNET pathogenesis are critical in developing novel therapeutic modalities. However, the lack of cellular and animal models of PanNETs hinders research progress. Currently utilized animal models have not faithfully recapitulated human PanNET disease as current models yield mixed tumor types (e.g. pancreatic and pituitary neuroendocrine tumors)15,46,47,48 which have severe phenotypes that confound preclinical findings49, or utilize mutations that are not common in PanNETs47,50,51. The latter includes RIP-Tag transgenic mice the most commonly used PanNET models, based on the expression of SV40 T antigen, which blocks p53 and Rb family tumor suppressors, mutations rarely found in PanNETs. In this context, sequencing efforts within the last decade have revealed the genetic mutation profile for this disease. These results have shown that loss of MEN1, DAXX, ATRX, and genes in PI3K/mTOR pathway (in particular PTEN and TSC2 tumor suppressors) are most frequently associated with PanNET tumorigenesis and predict poor prognosis14,15,16,52,53. Further, our analysis shows that MEN1, DAXX or ATRX and PTEN mutations commonly co-occur, suggesting that combined loss of those tumor suppressors is likely a key event in pancreatic neuroendocrine cell tumorigenesis. However, we do not have a genetically defined animal model that recapitulates the full spectrum of mutations co-occurring in human PanNETs. Since genetically engineered mouse models of cancer are instrumental in understanding the mechanisms of tumor progression, we sought to establish a model based on the most common and co-occurring PanNET mutations. Our results demonstrated that the combined loss of Men1, Atrx, and Pten, tumor suppressors, triggers the initiation and progression of pancreatic neuroendocrine cell tumorigenesis. The histopathological evaluation of MAP tumors at various time points indicates the development of progressive disease, which resembles intermediate to high-grade human PanNETs54. In contrast, previous models based on ablation of MEN1 or a combination of MEN1 and PTEN or DAXX deletion yield only low-grade tumors that rarely show signs of disease progression30,32,48. Further, our comparative transcriptomic analyses revealed that MAP model tumors recapitulate the expression of biomarkers diagnostic for human PanNET including Chromogranin A, Synaptophysin, PAX6 and NCAM1 (Fig. 3)42,43. In addition, MAP model tumors closely mimic gene expression programs found in human disease (Fig. 4), unlike commonly utilized RIP-Tag model exhibiting distinct transcriptomic programs. Altogether, we present here a novel genetically defined animal model of PanNETs that recapitulates human disease at phenotypic, genetic, and transcriptomic levels allowing further application in investigating PanNET biology and therapeutic response.
Materials and methods
Animal models
Conditional knockout strains Men1LoxP/LoxP, PtenLoxP/LoxP and AtrxLoxP/LoxP have been previously described55,56,57. Tissue-specific recombination of conditional alleles was achieved by interbreeding the conditional knockout alleles with Cre strain expressed from pancreas neuroendocrine cell specific promoter Pdx1-CreER obtained from Jackson Laboratory (#024968) and previously described58. The genotype of mice was determined by PCR on DNA obtained from tail snips of new weanlings using specific primers: (a) Pdx1-CreER-F: 5′ACCAGCCAGCTATCAACTCG3′, Pdx1-CreER-R: 5′TTACATTGGTCCAGCCACC, Control-F: -5′CTAGGCCACAGAATTGAAAGATCT3′, Control-R: 5′GTAGGTGGAAATTCTAGCATCATCC; Men1-WT-F: 5′ATTGAATAGCCAGAGGGATCTG, Men1-WT-R: 5′AGATGCTTGCTCAGTACATTGC, Men1-Lox-F: 5′GCCATTTCATTACCTCTTTCTCC; Atrx-WT-F: 5′TCAACTGCCCTACATACTGGTG, Atrx-Neo-R: 5′CGTGATATTGCTGAAGAGCTTG, Atrx-WT-R: 5′GCACGCAAGATAAGAGTGTCTG; Pten-F: 5′CAAGCACTCTGCGAACTGAG, Pten-R: 5′AAGTTTTTGAAGGCAAGATGC. To induce Cre-recombination and initiate PanNET tumorigenesis Pdx1-CreER; Men1 LoxP/LoxP; Atrx LoxP/LoxP; PtenLoxP/LoxP (MAP) mutant mice were injected intraperitoneally with 1 mg tamoxifen (Sigma-Aldrich) diluted in 100 μL sunflower oil at 8 weeks of age. For tumor development studies, animals were sacrificed at 8 months after tamoxifen injection and tissues were processed for histological and immunohistochemical analysis. For survival studies, mutant mice were continuously monitored for signs of disease progression and moribund conditions development. At the endpoint, tumors were processed for histological and immunohistochemical analysis. For blood glucose measurement MAP mutant mice at 8 months post-tumor induction and age- and sex-matched wildtype mice were fasted for 4 h before measurement. Blood glucose was measured with ONE TOUCH Ultra2 blood glucose meter (Lifescan, Inc., USA) according to the manufacturer’s instructions. In all experiments, all animals were numbered, and experiments were conducted in a blinded fashion. After data collection, genotypes were revealed, and animals assigned to groups for analysis. None of the mice with the appropriate genotype were excluded from this study or used in any other experiments. All mice were co-housed with littermates (2–5 per cage) in pathogen-free facility with standard controlled temperature of 72 °F, with a humidity of 30–70%, and a light cycle of 12 h on/12 h off set from 7 a.m. to 7 p.m. and with unrestricted access to standard food and water under the supervision of veterinarians, in an AALAC-accredited animal facility at the University of Texas M.D. Anderson Cancer Center (MDACC). Mouse handling and care followed the NIH Guide for Care and Use of Laboratory Animals. All animal procedures followed the guidelines of and were approved by the MDACC Institutional Animal Care and Use Committee (IACUC protocol 00001636, PI: Mazur).
Histology and immunohistochemistry
Mice were euthanized by CO2 asphyxiation followed by cervical dislocation at the indicated time points or when moribund conditions developed (endpoint). Tissue specimens were then fixed in 4% zinc-buffered formalin for 48 h and stored in 70% ethanol until processing and paraffin embedding. 1.5 μm sections were prepared and stained with Hematoxylin and Eosin (HE) or used for immunohistochemical studies (IHC). IHC was performed on formalin-fixed, paraffin-embedded mouse tissue section using a biotin-avidin method as previously described59. The following antibodies were used (at the indicated dilutions): insulin (CST #3014, 1:2000), synaptophysin (Abcam #ab32127, 1:1000), Ki67 (CST #12202, 1:800), CD56/NCAM (Proteintech #14255-1-AP, 1:2000) and ChgA (Abcam #ab15160, 1:1000). Sections were developed with DAB and counterstained with hematoxylin. Pictures were taken using a PreciPoint M8 microscope equipped with PointView software. The percentage of islet area or tumor area per pancreas was quantified based on HE staining utilizing Image J software. Cell proliferation was quantified using Ki67 marker. Ki67-positive and negative cells were quantified for each tumor field at 400 × magnification (5 representative tumor fields per tumor sample were analyzed). The Ki67 or mitotic index was calculated as the percentage of positively stained cells per the total number of cancer cells assessed.
Immunoblotting analysis
For immunoblotting analysis, wildtype pancreas and MAP model tumor biopsies were collected and homogenized in RIPA buffer with 1 mM PMSF and a complete protease inhibitor cocktail (Roche) to prepare whole-cell protein lysates. Protein concentration was determined using the BCA protein assay kit (Pierce #23227). Protein samples were resolved by SDS-PAGE and transferred to a PVDF membrane (0.45 µm). The following antibodies were used (at the indicated dilutions): MEN1 (1:1000, CST #19893), PTEN (1:1,000, CST #9188), ATRX (1:1000, CST #10321), Vinculin (1:10,000, CST #13901) and HRP-conjugated secondary antibody (1:10,000, CST #7074). Protein bands were visualized using Amersham ECL or Amersham ECL Prime western blotting detection reagent.
RNA isolation and sequencing
RNA samples were extracted from fresh tumor biopsies from the MAP model and pancreatic ductal adenocarcinoma model: KrasG12D; p53Lox/lox; Ptf1a-Cre (KPC) previously described60. Briefly, biopsied tumors were homogenized with TRIzol reagent (Life Technologies) and centrifuged at 16,000 RPM for 15 min at 4 °C. The samples received additional precipitation with a 1:2 chloroform:isoamyl alcohol pH 8.0 solution and centrifuged at 12,000 RPM for 15 min to promote phase separation of RNA. Three layers formed from this precipitation: an upper aqueous phase (containing RNA), a white interphase (containing DNA), and a lower organic phenol:chloroform phase (consisting of proteins). The upper aqueous phase was collected and mixed 1:1 with 100% ethanol. The Direct-zol RNA miniprep kit (cat. no. R2050) was utilized. RNA samples were sequenced at 20 M reads, PE150 (Novogene).
Human and mouse PanNET RNA-seq analyses
Previously reported RNA-seq datasets of human PanNETs44 were obtained from Gene Expression Omnibus (accession no. GSE178398) and RIP-Tag2 mouse model45 from the ZENODO database (access: doi.org/https://doi.org/10.5281/zenodo.4160441). RNA-seq data of MAP and KPC models were generated in-house and were mapped, counted, and normalized to TPM. Shared gene homologs between the human and mouse samples were kept for downstream processing. Correlation analysis was performed in R, and the results were plotted using the R package Corrplot.
Single-sample gene set enrichment analysis (ssGSEA)
Single-sample gene set enrichment analysis (ssGSEA) for each pancreatic tumor sample was performed using the R package GSVA on cancer hallmarks61 and GO neuroendocrine gene set (GO0061101) from the MSigDB Molecular Signatures Database.
Human PanNET mutation analysis
To identify genetic alternations critical for PanNET development, we performed mutation analysis of recurring mutations in human PanNETs using AACR GENIE Cohort v14.1 dataset (n = 413 patients) from cBioPortal v5.3.1335,62. To identify patterns of mutual exclusivity or co-occurrence, we performed statistical analysis for each pair of queried genes (MEN1, ATRX, DAXX and PTEN) that computes the likelihood that the mutations in each pair of genes are mutually exclusive or co-occurrent across the selected PanNET patient samples. The odds ratio (OR) is calculated with formula: OR = (A * D)/(B * C), where A = number of cases altered in both genes; B = number of cases altered in gene 1 but not gene 2; C = number of cases altered in gene 2 but not gene 1; and D = number of cases altered in neither genes.
Ethics approval
All animal experiments in this study were approved by the MDACC Institutional Animal Care and Use Committee (IACUC protocol 00001636, PI: Mazur) and conducted in compliance with the ARRIVE guidelines.
Data availability
The RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) database under accession number GSE248606. Review access tokens: gnavgwiwvtahfyx.
References
Dasari, A. et al. Trends in the incidence, prevalence, and survival outcomes in patients with neuroendocrine tumors in the United States. JAMA Oncol. 3, 1335–1342. https://doi.org/10.1001/jamaoncol.2017.0589 (2017).
Tang, L. H. & Klimstra, D. S. Conundrums and caveats in neuroendocrine tumors of the pancreas. Surg. Pathol. Clin. 4, 589–624. https://doi.org/10.1016/j.path.2011.03.003 (2011).
de Wilde, R. F., Edil, B. H., Hruban, R. H. & Maitra, A. Well-differentiated pancreatic neuroendocrine tumors: From genetics to therapy. Nat. Rev. Gastroenterol. Hepatol. 9, 199–208. https://doi.org/10.1038/nrgastro.2012.9 (2012).
Ferrone, C. R. et al. Determining prognosis in patients with pancreatic endocrine neoplasms: Can the WHO classification system be simplified?. J. Clin. Oncol. 25, 5609–5615. https://doi.org/10.1200/JCO.2007.12.9809 (2007).
Oronsky, B., Ma, P. C., Morgensztern, D. & Carter, C. A. Nothing but NET: A review of neuroendocrine tumors and carcinomas. Neoplasia 19, 991–1002. https://doi.org/10.1016/j.neo.2017.09.002 (2017).
Da Silva Xavier, G. The cells of the islets of Langerhans. J. Clin. Med. https://doi.org/10.3390/jcm7030054 (2018).
Halfdanarson, T. R., Rabe, K. G., Rubin, J. & Petersen, G. M. Pancreatic neuroendocrine tumors (PNETs): Incidence, prognosis and recent trend toward improved survival. Ann. Oncol. 19, 1727–1733. https://doi.org/10.1093/annonc/mdn351 (2008).
Singh, S. et al. Consensus recommendations for the diagnosis and management of pancreatic neuroendocrine tumors: Guidelines from a Canadian national expert group. Ann. Surg. Oncol. 22, 2685–2699. https://doi.org/10.1245/s10434-014-4145-0 (2015).
Kouvaraki, M. A. et al. Fluorouracil, doxorubicin, and streptozocin in the treatment of patients with locally advanced and metastatic pancreatic endocrine carcinomas. J. Clin. Oncol. 22, 4762–4771. https://doi.org/10.1200/JCO.2004.04.024 (2004).
Moertel, C. G., Lefkopoulo, M., Lipsitz, S., Hahn, R. G. & Klaassen, D. Streptozocin-doxorubicin, streptozocin-fluorouracil or chlorozotocin in the treatment of advanced islet-cell carcinoma. N. Engl. J. Med. 326, 519–523. https://doi.org/10.1056/NEJM199202203260804 (1992).
Raymond, E. et al. Sunitinib malate for the treatment of pancreatic neuroendocrine tumors. N. Engl. J. Med. 364, 501–513. https://doi.org/10.1056/NEJMoa1003825 (2011).
Yao, J. C. et al. Daily oral everolimus activity in patients with metastatic pancreatic neuroendocrine tumors after failure of cytotoxic chemotherapy: A phase II trial. J. Clin. Oncol. 28, 69–76. https://doi.org/10.1200/JCO.2009.24.2669 (2010).
Vandamme, T. et al. Long-term acquired everolimus resistance in pancreatic neuroendocrine tumours can be overcome with novel PI3K-AKT-mTOR inhibitors. Br. J. Cancer 114, 650–658. https://doi.org/10.1038/bjc.2016.25 (2016).
Scarpa, A. et al. Whole-genome landscape of pancreatic neuroendocrine tumours. Nature 543, 65–71. https://doi.org/10.1038/nature21063 (2017).
Sadanandam, A. et al. A cross-species analysis in pancreatic neuroendocrine tumors reveals molecular subtypes with distinctive clinical, metastatic, developmental, and metabolic characteristics. Cancer Discov. 5, 1296–1313. https://doi.org/10.1158/2159-8290.CD-15-0068 (2015).
Jiao, Y. et al. DAXX/ATRX, MEN1, and mTOR pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science 331, 1199–1203. https://doi.org/10.1126/science.1200609 (2011).
Crabtree, J. S. et al. A mouse model of multiple endocrine neoplasia, type 1, develops multiple endocrine tumors. Proc. Natl. Acad. Sci. USA 98, 1118–1123. https://doi.org/10.1073/pnas.98.3.1118 (2001).
Effraimidis, G. et al. Multiple endocrine neoplasia type 1 (MEN-1) and neuroendocrine neoplasms (NENs). Semin. Cancer Biol. 79, 141–162. https://doi.org/10.1016/j.semcancer.2021.04.011 (2022).
Harding, B. et al. Multiple endocrine neoplasia type 1 knockout mice develop parathyroid, pancreatic, pituitary and adrenal tumours with hypercalcaemia, hypophosphataemia and hypercorticosteronaemia. Endocr. Relat. Cancer 16, 1313–1327. https://doi.org/10.1677/ERC-09-0082 (2009).
Jiang, X. et al. Targeting beta-catenin signaling for therapeutic intervention in MEN1-deficient pancreatic neuroendocrine tumours. Nat. Commun. 5, 5809. https://doi.org/10.1038/ncomms6809 (2014).
Kamilaris, C. D. C. & Stratakis, C. A. Multiple endocrine neoplasia type 1 (MEN1): An update and the significance of early genetic and clinical diagnosis. Front. Endocrinol. (Lausanne) 10, 339. https://doi.org/10.3389/fendo.2019.00339 (2019).
Lines, K. E. et al. A MEN1 pancreatic neuroendocrine tumour mouse model under temporal control. Endocr. Connect. 6, 232–242. https://doi.org/10.1530/EC-17-0040 (2017).
Lu, J. et al. Alpha cell-specific Men1 ablation triggers the transdifferentiation of glucagon-expressing cells and insulinoma development. Gastroenterology 138, 1954–1965. https://doi.org/10.1053/j.gastro.2010.01.046 (2010).
Ehrlich, L. et al. A review of the scaffold protein menin and its role in hepatobiliary pathology. Gene Expr. 17, 251–263. https://doi.org/10.3727/105221617X695744 (2017).
Yokoyama, A. et al. The menin tumor suppressor protein is an essential oncogenic cofactor for MLL-associated leukemogenesis. Cell 123, 207–218. https://doi.org/10.1016/j.cell.2005.09.025 (2005).
Killock, D. Menin-KMT2A interaction inhibitor shows promise. Nat. Rev. Clin. Oncol. 20, 284. https://doi.org/10.1038/s41571-023-00759-w (2023).
Elsasser, S. J., Noh, K. M., Diaz, N., Allis, C. D. & Banaszynski, L. A. Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells. Nature 522, 240–244. https://doi.org/10.1038/nature14345 (2015).
Aguilera, P. & Lopez-Contreras, A. J. ATRX, a guardian of chromatin. Trends Genet. 39, 505–519. https://doi.org/10.1016/j.tig.2023.02.009 (2023).
Gaspar, T. B. et al. Characterisation of an Atrx conditional knockout mouse model: Atrx loss causes endocrine dysfunction rather than pancreatic neuroendocrine tumour. Cancers (Basel) https://doi.org/10.3390/cancers14163865 (2022).
Sun, C. et al. Context matters—Daxx and Atrx are not robust tumor suppressors in the murine endocrine pancreas. Dis. Model Mech. https://doi.org/10.1242/dmm.049552 (2022).
Wasylishen, A. R., Estrella, J. S., Pant, V., Chau, G. P. & Lozano, G. Daxx functions are p53-independent in vivo. Mol. Cancer Res. 16, 1523–1529. https://doi.org/10.1158/1541-7786.MCR-18-0281 (2018).
Wasylishen, A. R. et al. Daxx maintains endogenous retroviral silencing and restricts cellular plasticity in vivo. Sci. Adv. 6, eaba8415. https://doi.org/10.1126/sciadv.aba8415 (2020).
Zanini, S., Renzi, S., Giovinazzo, F. & Bermano, G. mTOR pathway in gastroenteropancreatic neuroendocrine tumor (GEP-NETs). Front. Endocrinol. (Lausanne) 11, 562505. https://doi.org/10.3389/fendo.2020.562505 (2020).
Missiaglia, E. et al. Pancreatic endocrine tumors: Expression profiling evidences a role for AKT-mTOR pathway. J. Clin. Oncol. 28, 245–255. https://doi.org/10.1200/JCO.2008.21.5988 (2010).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1. https://doi.org/10.1126/scisignal.2004088 (2013).
Voon, H. P. & Wong, L. H. New players in heterochromatin silencing: Histone variant H3.3 and the ATRX/DAXX chaperone. Nucleic Acids Res. 44, 1496–1501. https://doi.org/10.1093/nar/gkw012 (2016).
Magnuson, M. A. & Burlison, J. S. Caveats and considerations for performing pancreas-specific gene manipulations in the mouse. Diabetes Obes. Metab. 9(Suppl 2), 5–13. https://doi.org/10.1111/j.1463-1326.2007.00771.x (2007).
Magnuson, M. A. & Osipovich, A. B. Pancreas-specific Cre driver lines and considerations for their prudent use. Cell Metab. 18, 9–20. https://doi.org/10.1016/j.cmet.2013.06.011 (2013).
Sachdeva, M. M. et al. Pdx1 (MODY4) regulates pancreatic beta cell susceptibility to ER stress. Proc. Natl. Acad. Sci. USA 106, 19090–19095. https://doi.org/10.1073/pnas.0904849106 (2009).
Shyr, B. S., Shyr, B. U., Chen, S. C., Shyr, Y. M. & Wang, S. E. Impact of tumor grade on pancreatic neuroendocrine tumors. Asian J. Surg. 45, 2659–2663. https://doi.org/10.1016/j.asjsur.2022.01.094 (2022).
Inzani, F., Petrone, G. & Rindi, G. The new world health organization classification for pancreatic neuroendocrine neoplasia. Endocrinol. Metab. Clin. N. Am. 47, 463–470. https://doi.org/10.1016/j.ecl.2018.04.008 (2018).
Bocchini, M. et al. Biomarkers for pancreatic neuroendocrine neoplasms (PanNENs) management-an updated review. Front. Oncol. 10, 831. https://doi.org/10.3389/fonc.2020.00831 (2020).
Hofland, J., Zandee, W. T. & de Herder, W. W. Role of biomarker tests for diagnosis of neuroendocrine tumours. Nat. Rev. Endocrinol. 14, 656–669. https://doi.org/10.1038/s41574-018-0082-5 (2018).
Greenberg, J. et al. Metastatic pancreatic neuroendocrine tumors feature elevated T cell infiltration. JCI Insight https://doi.org/10.1172/jci.insight.160130 (2022).
Saghafinia, S. et al. Cancer cells retrace a stepwise differentiation program during malignant progression. Cancer Discov. 11, 2638–2657. https://doi.org/10.1158/2159-8290.CD-20-1637 (2021).
Hunter, K. E., Quick, M. L., Sadanandam, A., Hanahan, D. & Joyce, J. A. Identification and characterization of poorly differentiated invasive carcinomas in a mouse model of pancreatic neuroendocrine tumorigenesis. PLoS ONE 8, e64472. https://doi.org/10.1371/journal.pone.0064472 (2013).
Hanahan, D. Heritable formation of pancreatic beta-cell tumours in transgenic mice expressing recombinant insulin/simian virus 40 oncogenes. Nature 315, 115–122. https://doi.org/10.1038/315115a0 (1985).
Wong, C. et al. Two well-differentiated pancreatic neuroendocrine tumor mouse models. Cell Death Differ. 27, 269–283. https://doi.org/10.1038/s41418-019-0355-0 (2020).
Biondi, C. A. et al. Conditional inactivation of the MEN1 gene leads to pancreatic and pituitary tumorigenesis but does not affect normal development of these tissues. Mol. Cell Biol. 24, 3125–3131. https://doi.org/10.1128/MCB.24.8.3125-3131.2004 (2004).
Glenn, S. T. et al. Conditional deletion of p53 and Rb in the renin-expressing compartment of the pancreas leads to a highly penetrant metastatic pancreatic neuroendocrine carcinoma. Oncogene 33, 5706–5715. https://doi.org/10.1038/onc.2013.514 (2014).
Yamauchi, Y. et al. Rb and p53 execute distinct roles in the development of pancreatic neuroendocrine tumors. Cancer Res. 80, 3620–3630. https://doi.org/10.1158/0008-5472.CAN-19-2232 (2020).
Marinoni, I. et al. Loss of DAXX and ATRX are associated with chromosome instability and reduced survival of patients with pancreatic neuroendocrine tumors. Gastroenterology 146, 453-460.e455. https://doi.org/10.1053/j.gastro.2013.10.020 (2014).
Singhi, A. D. et al. Alternative lengthening of telomeres and loss of DAXX/ATRX expression predicts metastatic disease and poor survival in patients with pancreatic neuroendocrine tumors. Clin. Cancer Res. 23, 600–609. https://doi.org/10.1158/1078-0432.CCR-16-1113 (2017).
Guilmette, J. M. & Nose, V. Neoplasms of the neuroendocrine pancreas: An update in the classification, definition, and molecular genetic advances. Adv. Anat. Pathol. 26, 13–30. https://doi.org/10.1097/PAP.0000000000000201 (2019).
Lesche, R. et al. Cre/loxP-mediated inactivation of the murine Pten tumor suppressor gene. Genesis 32, 148–149. https://doi.org/10.1002/gene.10036 (2002).
Libutti, S. K. et al. Parathyroid gland-specific deletion of the mouse Men1 gene results in parathyroid neoplasia and hypercalcemic hyperparathyroidism. Cancer Res. 63, 8022–8028 (2003).
Berube, N. G. et al. The chromatin-remodeling protein ATRX is critical for neuronal survival during corticogenesis. J. Clin. Investig. 115, 258–267. https://doi.org/10.1172/JCI22329 (2005).
Gu, G., Dubauskaite, J. & Melton, D. A. Direct evidence for the pancreatic lineage: NGN3+ cells are islet progenitors and are distinct from duct progenitors. Development 129, 2447–2457. https://doi.org/10.1242/dev.129.10.2447 (2002).
Mazur, P. K. et al. SMYD3 links lysine methylation of MAP3K2 to Ras-driven cancer. Nature 510, 283–287. https://doi.org/10.1038/nature13320 (2014).
Wang, Z. et al. SETD5-coordinated chromatin reprogramming regulates adaptive resistance to targeted pancreatic cancer therapy. Cancer Cell 37, 834-849.e813. https://doi.org/10.1016/j.ccell.2020.04.014 (2020).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425. https://doi.org/10.1016/j.cels.2015.12.004 (2015).
Cerami, E. et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404. https://doi.org/10.1158/2159-8290.CD-12-0095 (2012).
Acknowledgements
This work was supported in part by grants from the NIH to P.K.M. (R01CA236118, R01CA278940, R01CA272844, R01CA236949, R01CA266280, R01CA272843) and S.H. (K99CA255936), P.K.M. is also supported by the Neuroendocrine Tumor Research Foundation (NETRF), DoD PRCRP Career Development Award (CA181486), CPRIT IIRA (RP220391) and CPRIT Scholar in Cancer Research (RR160078). N.M.F. was supported by the American Cancer Society Fellowship, X. L. and M.E.F. are TRIUMPH Fellows in the CPRIT Training Program (RP210028)
Author information
Authors and Affiliations
Contributions
M.E.F. was responsible for the experimental design, execution, data analysis, and manuscript drafting. X.L. performed RNA-seq bioinformatics and ssGSEA analyses N.M.F. and S.H. assisted with animal studies and data interpretation. S.H. performed RNA-seq experiments. P.K.M. was responsible for supervision of research, data interpretation and manuscript preparation. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
P.K.M. is a co-scientific founder, consultant, and stockholder of Amplified Medicines, Inc.; consultant and stockholder of Ikena Oncology, Inc. and Alternative Bio, Inc. The other authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fuentes, M.E., Lu, X., Flores, N.M. et al. Combined deletion of MEN1, ATRX and PTEN triggers development of high-grade pancreatic neuroendocrine tumors in mice. Sci Rep 14, 8510 (2024). https://doi.org/10.1038/s41598-024-58874-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-58874-2
- Springer Nature Limited