Abstract
To retrospectively assess the effectiveness of deep learning (DL) model, based on breast magnetic resonance imaging (MRI), in predicting preoperative lymphovascular invasion (LVI) status in patients diagnosed with invasive breast cancer who have negative axillary lymph nodes (LNs). Data was gathered from 280 patients, including 148 with LVI-positive and 141 with LVI-negative lesions. These patients had undergone preoperative breast MRI and were histopathologically confirmed to have invasive breast cancer without axillary LN metastasis. The cohort was randomly split into training and validation groups in a 7:3 ratio. Radiomics features for each lesion were extracted from the first post-contrast dynamic contrast-enhanced (DCE)-MRI. The Least Absolute Shrinkage and Selection Operator (LASSO) regression method and logistic regression analyses were employed to identify significant radiomic features and clinicoradiological variables. These models were established using four machine learning (ML) algorithms and one DL algorithm. The predictive performance of the models (radiomics, clinicoradiological, and combination) was assessed through discrimination and compared using the DeLong test. Four clinicoradiological parameters and 10 radiomic features were selected by LASSO for model development. The Multilayer Perceptron (MLP) model, constructed using both radiomic and clinicoradiological features, demonstrated excellent performance in predicting LVI, achieving a high area under the curve (AUC) of 0.835 for validation. The DL model (MLP-radiomic) achieved the highest accuracy (AUC = 0.896), followed by DL model (MLP-combination) with an AUC of 0.835. Both DL models were significantly superior to the ML model (RF-clinical) with an AUC of 0.720. The DL model (MLP), which integrates radiomic features with clinicoradiological information, effectively aids in the preoperative determination of LVI status in patients with invasive breast cancer and negative axillary LNs. This is beneficial for making informed clinical decisions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Lymphovascular invasion (LVI), defined as the infiltration of cancer cells into the lymphatic or vascular intraluminal area at the periphery of the primary tumor site1, is widely recognized as an unfavorable prognostic factor for patients with lymph node (LN)-negative breast cancer2. LVI is a crucial step in LN metastasis, and LVI-positive tumors have a higher potential for local recurrence and distant metastasis than LVI-negative ones3,4,5. Specifically, for patients with LN-negative breast cancer, preoperative detection of LVI status can predict metastatic potential, informing decisions regarding adjuvant treatment6,7,8. Comprehensive regional nodal irradiation is recommended for patients with LVI according to the NCCN guidelines9. Furthermore, the presence of LVI is related to non-sentinel lymph node involvement in patients with a positive sentinel node biopsy10. Currently, LVI status is determined using surgical pathology specimens, which can lead to complications such as delayed treatment, additional surgeries, unnecessary anxiety, and diminished quality of life. Therefore, early identification of LVI during treatment is crucial. While preoperative needle biopsy is the preferred method for most patients to determine tumor characteristics, accurate detection of LVI through partial sampling from core biopsy is challenging due to tissue shrinkage and mechanically induced cellular displacement11.
Multiparametric MRI is an exceptionally reliable, comprehensive, and accurate diagnostic technique for preoperative staging, boasting the highest sensitivity and a reasonably high specificity for detecting breast malignancy among current imaging patterns12,13. Several studies on breast MRI have indicated that certain MRI features such as mass margins, adjacent-vessel sign (AVS), peritumoral edema, tumor size, background parenchymal enhancement (BPE), apparent diffusion coefficient (ADC) patterns, and contrast enhancement patterns are associated with LVI2,12,14,15. Nevertheless, their results are sometimes inconsistent due to subjective factors during feature assessment.
Radiomics, a recent branch of science, can aid radiologists in making highly accurate diagnoses through deep mining, prediction, and analysis of large data sets16. Deep learning (DL), in contrast, has made significant strides in the field of breast imaging diagnosis, particularly in the detection and classification of breast cancer17.
While several studies have utilized MRI-based radiomics to predict LVI in patients with breast cancer18,19,20,21, the cohort sizes in these studies were small, potentially affecting the outcomes. To the best of our knowledge, few studies have integrated the clinicoradiological features with DL signatures to establish LVI-predictive models19,21,22, and data on DL techniques for determining LVI status in a node-negative invasive breast cancer cohort are scarce.
Accordingly, in this study, we collected data from a large cohort (280 patients). Our aim was to investigate whether a DL model incorporating MRI-based radiomic features, clinicopathologic characteristics, and conventional MRI manifestations could serve as a preoperative tool for LVI prediction in patients with invasive breast cancer and negative axillary LNs.
Materials and methods
Patients
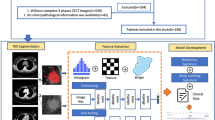
This retrospective study was approved by the Tianjin cancer hospital medical ethics committee, which waived the requirement for informed consent and all research was performed in accordance with relevant guidelines. We recruited a total of 1275 consecutive female patients diagnosed with invasive breast carcinoma between January 2020 and June 2022. The exclusion criteria are detailed in Appendix E1. Ultimately, 280 patients with 289 lesions were included. These patients were randomly grouped into training (n = 202) and validation (n = 87) datasets at a ratio of 7:3. The overall study design is illustrated in Fig. 1.
Image acquisition protocol
MRI examinations were conducted with patients in the prone position using a 1.5 T scanner (Signa HDxt, GE Healthcare, USA) and a 3.0 T scanner (Signa Pioneer and Discovery 750, GE Healthcare, USA), each equipped with a dedicated four- or eight-channel phased-array breast coil. Our center’s routine protocols included the following sequences: axial fast spin-echo (FSE) T1-weighted imaging (T1WI), axial fat-suppressed spin-echo T2-weighted imaging (T2WI), axial echo-planar diffusion-weighted imaging (DWI), and a sagittal dynamic contrast-enhanced (DCE) sequence. Six dynamic sagittal sequences were acquired once before and five times after the injection of contrast medium (Gd-DTPA, 0.2 mL/kg body weight, flow rate 2.0 mL/s) at 60-s intervals, immediately followed by an axial delayed-enhancement sequence. The relevant parameters for each sequence are detailed in Appendix E2.
Image interpretation
Two radiologists, each with 6 and 8 years of experience in breast imaging diagnosis, independently analyzed all MRI manifestations according to the 2013 Breast Imaging Reporting and Data System (BI-RADS) lexicon of the American College of Radiology. Both observers were blinded to the histopathological information. They evaluated conventional imaging findings of the lesions, including lesion type, tumor size, lesion location, tumor margin, time-intensity curve (TIC) patterns, BPE, AVS, and tumor ADC value. All images were sent to the Advantage Workstation (AW 4.6 and AW4.7, GE Healthcare) for further post-processing, including TIC and ADC value measurements, using Functool and READY View software. Circular ROIs, accounting for approximately 75% of each lesion after avoiding areas with hemorrhage or necrosis, were drawn and used. TIC patterns were assessed and categorized into three types (I, II, and III): progressive enhancement, plateau, and washout patterns. BPE was divided into two categories: extremely minimal (or mild enhancement) and moderate (or marked) enhancement. Peritumoral edema was defined as high signal intensity surrounding the tumor on T2WI23. AVS positivity was defined as either one or more vessels entering the lesion on the T1WI post-enhancement sequence24. The largest tumor diameter in the image showing the largest lesion was measured and recorded. For the semi-quantitative kinetic curve parameters automatically computed by the Functool software, we measured SI0 (signal intensity values in the pre-contrast images), SI1 (signal intensity values in the first post-contrast images), and SImax (the maximum signal intensity values in the post-contrast images). The early enhancement ratio (EER) was calculated as follows: EER = (SI1 − SI0)/ SI0 × 100. The peak enhancement ratio (PER) was calculated as follows: PER = (SImax − SI0)/SI0 × 100. Time-to-peak enhancement (TTP) was also obtained. The reliability of the observations was assessed using the intraclass correlation coefficient (ICC). Features with ICCs larger than 0.75 were considered to have satisfactory reproducibility and were reserved for further analysis.
Histopathological assessment
The expression levels of estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), and Ki-67, along with the presence of IMPCs, were pathologically evaluated using hematoxylin and eosin (H&E) and immunohistochemical analysis. ER and PR were considered positive if their expression level was greater than 1%. HER2 was considered negative if its value was 0 or 1, and positive if its value was 3 or greater. If the HER2 value was 2, fluorescence in situ hybridization (FISH) was required for further confirmation of HER2 status. High expression of Ki-67 was defined as ≥ 14%, and low expression as < 14%. All cases were classified into four immunohistochemical subtypes based on the 2013 St. Gallen Consensus Conference25: luminal A, luminal B, HER2-positive, and triple-negative.
Volume of interest (VOI) segmentation and radiomic feature extraction
Segmentations for all MRI images were performed by a radiologist with 6 years of experience in MRI interpretation. Subsequently, 100 randomly selected cases were assigned to another radiologist with 8 years of experience in MRI interpretation. Both radiologists were blinded to the patients’ clinical data. An example of the segmentation process is shown in Fig. S1. Feature extraction was performed using the radiomics module in the 3D Slicer software. A total of 851 radiomic features were extracted from each ROI. These features included 107 original features and 744 wavelet-based features (comprising first-order statistics, gray-level co-occurrence matrix [GLCM], gray-level size-zone matrix [GLSZM], gray-level run-length matrix [GLRLM], gray-level dependence matrix [GLDM], neighboring gray tone difference matrix [NGTDM], and shape-based features). A description of the initial 851 features is provided in Appendix E3.
Radiomic and clinicoradiological feature selection
Feature selection and ML/DL model development were performed on the training cohort, while the validation cohort was kept completely independent and invisible until the final model performance was evaluated. Feature standardization (standard deviation) was performed before feature selection. The selection of robust radiomic features included three steps. First, to reduce the risk of overfitting, features with highly pairwise correlations at the level of |r| ≥ 0.9 were removed by Pearson’s correlation analysis. Second, we performed univariate selection using the SelectKBest method for every feature, and features with p values < 0.05 were retained for further analysis. Third, the Least Absolute Shrinkage and Selection Operator (LASSO) method was adopted to identify the optimal log (λ) value by fivefold cross-validation and obtain the robust radiomic features with a non-zero coefficient for the differentiation of the LVI and non-LVI groups. Independent clinicoradiological features were selected from the variables found to be significant in the univariate analysis using multivariate logistic regression analysis.
Model construction and validation
Four different ML classification algorithms (random forest (RF), logistic regression (LR), support vector machine (SVM) and stochastic gradient descent (SGD)), and a DL classification algorithm (Multilayer Perceptron (MLP)) were used to build models for predicting LVI in the training cohort. We tuned the parameters of the different algorithms using fivefold cross-validation. The optimal hyperparameters for each algorithm were obtained based on the evaluation results. The relevant hyperparameters are presented in Appendix E4. Predictive models for the radiomic signature (Model 1), selected clinicoradiological variables (Model 2), and integrated features (combining the above two parameters, Model 3) were constructed for the training cohort and were validated in the validation cohorts.
The models’ performance was evaluated based on discrimination metrics, which included the receiver operating characteristic (ROC) curve, area under the ROC curve (AUC), accuracy, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV). The AUCs of the models were compared using DeLong’s test.
Statistical analysis
Statistical analyses were conducted using MedCalc® version 20.0.3 (MedCalc Software Ltd, Ostend, Belgium) and R software version 4.2.2 (http://www.Rproject.org). Continuous variables are presented as the mean ± standard deviation (SD), and categorical variables as frequencies or percentages, unless otherwise specified. The Kolmogorov–Smirnov test was utilized to test the normality of the data distribution. Continuous variables were compared using the Mann–Whitney U test, while dichotomous qualitative variables were assessed using the Chi-squared test or Fisher’s exact test. Univariate and multivariate logistic regression methods were employed to determine the association between clinicopathological and radiological features and LVI. Radiomic feature selection and model development were performed using the Shukun Medical Research Platform (https://medresearch.shukun.net/project; Appendix E5). All tests were performed two-sided, and a p-value of < 0.05 was considered statistically significant.
Results
Patient and MRI characteristics
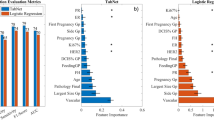
The ICCs for all MRI characteristics exceeded 0.75 between the two radiologists. A comparison of the clinicoradiological features between the LVI-positive and LVI-negative patient groups is presented in Table 1. In total, 280 consecutive patients were included (median age, 47 years; range, 22–74 years). LVI was found in 140 patients with 148 lesions and was absent in 140 patients with 141 lesions. In the univariate analysis, patient age, peritumoral edema, TTP, and IMPCs were significantly correlated with LVI (p < 0.05). In the multivariate logistic regression analysis, patient age (odds ratio [OR] = 1.03; 95% confidence interval [CI] 1.00–1.05; p = 0.02), peritumoral edema (OR = 2.44; 95% CI 1.18–5.08; p = 0.02), TTP (OR = 1.00; 95% CI 0.99–1.00; p = 0.04) and IMPCs (OR = 0.35; 95% CI 0.14–0.87; p = 0.02) remained independent predictors of LVI (Table 2 and Fig. 2).
Radiomic feature selection
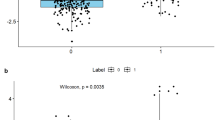
In this study, a total of 851 radiomics features for each lesion were extracted, including 107 original features and 744 wavelet-based features. 282 features remained after feature preselection using the Pearson correlation analysis. Then, using the single-variable feature selection method (SelectKBest), 117 features remained. After further feature dimensionality reduction by the LASSO regression algorithm, 10 significant radiomic features were finally selected for the subsequent establishment of the ML and DL models (Figs. S2 and Fig. 3).
Predictive models assessment
Radiomics predictive models were built using different ML algorithms and a DL algorithm, which demonstrated good diagnostic performance in the training sets. The predictive performance was validated using the validation dataset. The optimal model was the MLP model (AUC = 0.896; 95% CI 0.806–0.964). The corresponding ROC curves, AUC, accuracy, sensitivity, specificity, PPV, and NPV values are shown in Table 3 and Fig. 4.
The models based on the selected clinicoradiological variables applied to the four ML algorithms and MLP algorithm showed relatively good performance in predicting LVI after fivefold cross-validation. The best performance was observed in the RF model (AUC = 0.720; 95% CI 0.611–0.830). The corresponding ROC curves, AUC, accuracy, sensitivity, specificity, PPV, and NPV values are presented in Table S1 and Fig. S3.
In the models based on combined features, including 10 selected radiomic signatures and f4 clinicoradiological variables, the MLP model (AUC = 0.835; 95% CI 0.719–0.929) was the optimal model after fivefold cross-validation. The corresponding ROC curves, AUC, accuracy, sensitivity, specificity, PPV, and NPV values are shown in Table 4 and Fig. 5.
The performances of model 1, 2 and 3 in the optimal classifier (RF and MLP algorithm) are displayed in Fig. 6. In the validation cohort, the Delong test showed that the model 1 achieved the highest accuracy (AUC = 0.896), being significantly superior to the model 2 (AUC = 0.720) or model 3 (AUC = 0.835) (p < 0.001). Model 3 exhibited better performance than models 2 (p < 0.001) (Table 5).
Discussion
In this study, we investigated the relationship between LVI and invasive breast cancer in node-negative patients. We constructed MRI-based radiomic models that demonstrated a highly robust ability to predict LVI. Four different ML algorithms and one DL algorithm were used for model construction based on the 10 selected radiomic signatures, and the DL model outperformed the other models in terms of identifying the LVI status. Furthermore, the combined model, which integrated clinicoradiological characteristics with radiomic signatures, exhibited significantly better predictive performance than the clinical model. This is expected to offer a relatively accurate and objective approach for preoperative LVI prediction.
Some studies have shown that LVI is associated with age, tumor size, mass margins, positive lymph nodes, BPE, TIC patterns, ADC values, peritumoral edema, AVS, DWI rim sign, and histologic grade1,2,12,15,26. Our study found that the risk of LVI increased with a lower mean age. The mean age of the patients with LVI was lower than that of the patients without LVI. A previous study revealed that age is a high-risk factor for LVI in invasive breast cancer27. Yang et al.28 reported that premenopausal status was associated with an increased risk (4.59-fold) of early-onset breast cancer (< 40 years). Early-onset breast cancer is more aggressive in premenopausal women than in older women, due to the gradual decrease in blood estrogen and progesterone levels with age29.
We also identified peritumoral edema as a predictor of LVI; patients with peritumoral edema had a higher risk of LVI, which aligns with previous study results21,26. A significantly higher incidence of peritumoral edema was observed in patients with more biologically aggressive tumors23. One possible reason for peritumoral edema may be an increase in vascular permeability and obstruction of lymphatic drainage by tumor emboli30. TTP, a semiquantitative DCE-MRI parameter, was found to be an independent predictive factor for LVI. We found that patients with a longer TTP had a higher probability of positive LVI, which might be related to the reduced focal blood flow caused by tumor emboli. Numerous studies have shown that TTP is a diagnostic indicator of malignant breast lesions31 and an independent predictive factor of pathological complete response (pCR) after neoadjuvant chemotherapy (NAC)32,33,34. One recent study analyzed the relationship between TTP and LVI, but found no significant differences between patients with and without LVI35. Our study identified IMPC as a predictive indicator of LVI. To the best of our knowledge, this is the first study to include IMPCs to construct a predictive model for detecting the LVI status in patients with invasive breast cancer. IMPC is a rare subtype of an epithelial tumor of the breast listed in the 2003 World Health Organization (WHO) histological classification of breast tumors36. Previous studies demonstrated that breast cancers with IMPCs have a relatively high incidence of LVI37. LVI has also been found to be more common among patients with IMPC (pure or mixed with IDC), as reported by Tang et al.38, with 14.7% versus only 0.1% in the IDC group and 94.7% versus 71.9% in the control group reported by Gokce et al.39. In our study, we found a higher frequency of IMPC in patients with LVI than in those without LVI, which is consistent with a previous study40.
Several studies have confirmed that the DCE-MRI-based radiomics ML model has robust power for predicting the LVI status of patients with invasive breast cancer18,19,20,21. Zhang et al.21 identified that the proposed nomogram, incorporating MRI-based radiomic signatures and MRI-reported peritumoral edema, achieved satisfactory preoperative prediction of LVI and clinical outcomes in patients with IDC. A recent study used an SVM classifier to establish a prediction model for LVI based on the ADC radiomic signature, and its AUC was 0.77 in the test set20. Liu and colleagues19 confirmed that a radiomics model based on DCE-MRI utilizing a multivariate logistic regression method significantly improved the performance for discriminating LVI-positive from LVI-negative lesions. Another study found that an ML-based radiomics model based on 3D segmentation of ADC maps could be used to predict the LVI status in breast cancer patients18. These studies, however, only employed classical non-network methods for classification. In contrast, our research introduced and a deep learning model (MLP) for comparison. Yang et al.22 developed a diagnostic model that combined MRI morphological features, Radiomics, and DL features to determine the LVI status in 206 breast cancer patients. They found that the combined model was more efficient in distinguishing between LVI positive and negative cases compared to individual models. A recent study41 developed a novel DL framework, “Prior Clinico-Radiological Features Informed Multi-Modal MR Images Convolutional Neural Network (PCMM-Net)”, for predicting LVI in breast cancer, achieving a higher AUC of 0.843 than clinico-radiological features alone. This aligns with our study’s findings. However, their studies included both lymph node-positive and -negative patients, while ours only included lymph node-negative patients. We selected this sample group due to its clinical significance. The 16th St. Gallen consensus established that the inclusion of chemotherapy for N0 patients should also be based on the presence of LVI42. LVI is also currently of interest for predicting non-sentinel lymph node metastases43. In clinical practice, axillary node dissection is typically recommended for effective local treatment of patients with positive sentinel nodes44. Nevertheless, since over 40% of patients with positive sentinel nodes have no other nodal metastases, predicting non-sentinel node involvement has been conducive to avoiding unnecessary axillary node dissection45. Consequently, the prediction of non-sentinel node involvement associated with the presence of LVI might allow for appropriate treatment (sentinel node biopsy or axillary node dissection) of patients.
This study has several limitations. First, this was a retrospective single-center study, highlighting the necessity for a large-scale prospective multicenter study to further validate the effectiveness of the proposed radiomics model. Second, only the first phase after enhancement was used to extract radiomic features in this study, suggesting the need for other sequences, such as T2W, DWI images, and ADC maps, for feature extraction in further studies. Third, our institution originally designed a sagittal DCE sequence for breast MRI examination, meaning the functional tumor volume (FTV) of the lesions could not be obtained. Similarly, our study only analyzed semi-quantitative kinetic curve parameters for LVI prediction. Quantitative pharmacokinetic parameters (such as ktrance, kep, and Ve) were not measured using our equipment. Further exploration of these parameters is required, if conditions permit.
In conclusion, the radiomic features derived from DCE-MRI are robust biomarkers for predicting LVI. The combined DL model, incorporating the radiomic signature and clinicoradiological-based variables, exhibited a highly acceptable predictive efficacy for LVI status in patients with node-negative invasive breast cancer.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
Code availability
The essential analysis code used to produce the results is available from the GitHub repository at https://github.com/Sarah147369/BreastCancerLVI-Prediction.
Abbreviations
- ADC:
-
Apparent diffusion coefficient
- AR:
-
Androgen receptor
- AUC:
-
Area under the curve
- AVS:
-
Adjacent vessel sign
- BI-RADS® :
-
Breast Imaging Reporting and Data System
- BPE:
-
Background parenchymal enhancement
- DL:
-
Deep learning
- EER:
-
Early enhancement ratio
- ER:
-
Estrogen receptor
- HER2:
-
Human epidermal growth factor receptor 2
- ICC:
-
Interclass correlation coefficient
- IDC:
-
Invasive ductal carcinoma
- IMPCs:
-
Invasive micropapillary components
- LASSO:
-
Least Absolute Shrinkage and Selection Operator
- LVI:
-
Lymphovascular invasion
- ML:
-
Machine learning
- NME:
-
Nonmass enhancement
- PER:
-
Peak enhancement ratio
- PR:
-
Progesterone receptor
- ROC:
-
Receiver operating characteristic
- RF:
-
Random Forest
- SLNB:
-
Sentinel lymph node biopsy
- TIC:
-
Time-intensity curve
- TTP:
-
Time to peak
References
Igarashi, T., Furube, H., Ashida, H. & Ojiri, H. Breast MRI for prediction of lymphovascular invasion in breast cancer patients with clinically negative axillary lymph nodes. Eur. J. Radiol. 107, 111–118. https://doi.org/10.1016/j.ejrad.2018.08.024 (2018).
Choi, B. B. Dynamic contrast enhanced-MRI and diffusion-weighted image as predictors of lymphovascular invasion in node-negative invasive breast cancer. World J. Surg. Oncol. 19, 76. https://doi.org/10.1186/s12957-021-02189-3 (2021).
Schoppmann, S. F. et al. Prognostic value of lymphangiogenesis and lymphovascular invasion in invasive breast cancer. Ann. Surg. 240, 306–312. https://doi.org/10.1097/01.sla.0000133355.48672.22 (2004).
Zhang, S. et al. High lymphatic vessel density and presence of lymphovascular invasion both predict poor prognosis in breast cancer. BMC Cancer 17, 335. https://doi.org/10.1186/s12885-017-3338-x (2017).
Lin, Y. et al. Survival and clinicopathological significance of blood vessel invasion in operable breast cancer: A systematic review and meta-analysis. Jpn. J. Clin. Oncol. 53, 35–45. https://doi.org/10.1093/jjco/hyac149 (2023).
Matsuda, N. et al. A nomogram for predicting locoregional recurrence in primary breast cancer patients who received breast-conserving surgery after neoadjuvant chemotherapy. J. Surg. Oncol. 109, 764–769. https://doi.org/10.1002/jso.23586 (2014).
Recht, A. et al. Postmastectomy radiotherapy: An American Society of Clinical Oncology, American Society for Radiation Oncology, and Society of Surgical Oncology Focused Guideline Update. Ann. Surg. Oncol. 24, 38–51. https://doi.org/10.1245/s10434-016-5558-8 (2017).
Lee, S. J. et al. Lymphovascular invasion is an independent prognostic factor in breast cancer irrespective of axillary node metastasis and molecular subtypes. Front. Oncol. 13, 1269971. https://doi.org/10.3389/fonc.2023.1269971 (2023).
Daly, M. B. et al. Genetic/Familial High-Risk Assessment: Breast, ovarian, and pancreatic, version 2.2021, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 19, 77–102. https://doi.org/10.6004/jnccn.2021.0001 (2021).
Pache, B. et al. Predictors of non-sentinel lymph node metastasis in patients with positive sentinel lymph node in early-stage cervical cancer: A SENTICOL GROUP Study. Cancers (Basel) https://doi.org/10.3390/cancers15194737 (2023).
Cheung, S. M. et al. Intra-tumoural lipid composition and lymphovascular invasion in breast cancer via non-invasive magnetic resonance spectroscopy. Eur. Radiol. 31, 3703–3711. https://doi.org/10.1007/s00330-020-07502-4 (2021).
Li, J. et al. Association between MRI background parenchymal enhancement and lymphovascular invasion and estrogen receptor status in invasive breast cancer. Br. J. Radiol. 92, 20190417. https://doi.org/10.1259/bjr.20190417 (2019).
Mann, R. M., Cho, N. & Moy, L. Breast MRI: State of the art. Radiology 292, 520–536. https://doi.org/10.1148/radiol.2019182947 (2019).
Okuma, H. et al. Peritumor to tumor apparent diffusion coefficient ratio is associated with biologically more aggressive breast cancer features and correlates with the prognostication tools. PLoS One 15, e0235278. https://doi.org/10.1371/journal.pone.0235278 (2020).
Ouyang, F. S. et al. A nomogram for individual prediction of vascular invasion in primary breast cancer. Eur. J. Radiol. 110, 30–38. https://doi.org/10.1016/j.ejrad.2018.11.013 (2019).
Fusco, R. et al. An informative review of radiomics studies on cancer imaging: The main findings, challenges and limitations of the methodologies. Curr. Oncol. 31, 403–424. https://doi.org/10.3390/curroncol31010027 (2024).
Díaz, O., Rodríguez-Ruíz, A. & Sechopoulos, I. Artificial Intelligence for breast cancer detection: Technology, challenges, and prospects. Eur. J. Radiol. 175, 111457. https://doi.org/10.1016/j.ejrad.2024.111457 (2024).
Kayadibi, Y. et al. MRI radiomics of breast cancer: Machine learning-based prediction of lymphovascular invasion status. Acad. Radiol. 29(Suppl 1), S126–S134. https://doi.org/10.1016/j.acra.2021.10.026 (2022).
Liu, Z. et al. Preoperative prediction of lymphovascular invasion in invasive breast cancer with dynamic contrast-enhanced-MRI-based radiomics. J. Magn. Reson. Imaging 50, 847–857. https://doi.org/10.1002/jmri.26688 (2019).
Nijiati, M. et al. MRI-based radiomics for preoperative prediction of lymphovascular invasion in patients with invasive breast cancer. Front. Oncol. 12, 876624. https://doi.org/10.3389/fonc.2022.876624 (2022).
Zhang, J. et al. Multiparametric MRI-based radiomics nomogram for preoperative prediction of lymphovascular invasion and clinical outcomes in patients with breast invasive ductal carcinoma. Eur. Radiol. 32, 4079–4089. https://doi.org/10.1007/s00330-021-08504-6 (2022).
Yang, X. et al. Assessment of lymphovascular invasion in breast cancer using a combined MRI morphological features, radiomics, and deep learning approach based on dynamic contrast-enhanced MRI. J. Magn. Reson. Imaging https://doi.org/10.1002/jmri.29060 (2023).
Cheon, H. et al. Invasive breast cancer: Prognostic value of peritumoral edema identified at preoperative MR imaging. Radiology 287, 68–75. https://doi.org/10.1148/radiol.2017171157 (2018).
Zhao, S. et al. Association of peritumoral region features assessed on breast MRI and prognosis of breast cancer: A systematic review and meta-analysis. Eur. Radiol. https://doi.org/10.1007/s00330-024-10612-y (2024).
Untch, M. et al. 13th St. Gallen international breast cancer conference 2013: Primary therapy of early breast cancer evidence, controversies, consensus—Opinion of a German team of experts (Zurich 2013). Breast Care (Basel, Switzerland) 8(221–229), 2013. https://doi.org/10.1159/000351692 (2013).
Cheon, H. et al. Preoperative MRI features associated with lymphovascular invasion in node-negative invasive breast cancer: A propensity-matched analysis. J. Magn. Reson. Imaging 46, 1037–1044. https://doi.org/10.1002/jmri.25710 (2017).
Zhang, C. et al. Risk factors for lymphovascular invasion in invasive ductal carcinoma based on clinical and preoperative breast MRI features: A retrospective study. Acad. Radiol. https://doi.org/10.1016/j.acra.2022.10.029 (2022).
Yang, P. J., Hou, M. F., Ou-Yang, F., Tsai, E. M. & Wang, T. N. Association of early-onset breast cancer with body mass index, menarche, and menopause in Taiwan. BMC Cancer 22, 259. https://doi.org/10.1186/s12885-022-09361-2 (2022).
Goodburn, R. et al. Quantitative background parenchymal enhancement and fibro-glandular density at breast MRI: Association with BRCA status. Eur. Radiol. 33, 6204–6212. https://doi.org/10.1007/s00330-023-09592-2 (2023).
Uematsu, T. Focal breast edema associated with malignancy on T2-weighted images of breast MRI: Peritumoral edema, prepectoral edema, and subcutaneous edema. Breast Cancer (Tokyo, Japan) 22, 66–70. https://doi.org/10.1007/s12282-014-0572-9 (2015).
Cao, Y. et al. Multiple parameters from ultrafast dynamic contrast-enhanced magnetic resonance imaging to discriminate between benign and malignant breast lesions: Comparison with apparent diffusion coefficient. Diagn. Interv. Imaging 104, 275–283. https://doi.org/10.1016/j.diii.2023.01.006 (2023).
Panzeri, M. M. et al. Prediction of chemoresistance in women undergoing neo-adjuvant chemotherapy for locally advanced breast cancer: Volumetric analysis of first-order textural features extracted from multiparametric MRI. Contrast Media Mol. Imaging 2018, 8329041. https://doi.org/10.1155/2018/8329041 (2018).
Li, Y. et al. Development and validation of a nomogram based on pretreatment dynamic contrast-enhanced MRI for the prediction of pathologic response after neoadjuvant chemotherapy for triple-negative breast cancer. Eur. Radiol. 32, 1676–1687. https://doi.org/10.1007/s00330-021-08291-0 (2022).
Murakami, W. et al. Quantitative predictors of response to neoadjuvant chemotherapy on dynamic contrast-enhanced 3T breast MRI. J. Breast Imaging 4, 168–176. https://doi.org/10.1093/jbi/wbab095 (2022).
Lai, T. et al. Quantitative parameters of dynamic contrast-enhanced magnetic resonance imaging to predict lymphovascular invasion and survival outcome in breast cancer. Cancer Imaging 22, 61. https://doi.org/10.1186/s40644-022-00499-7 (2022).
Walker, R. A. World Health Organization classification of tumours. Pathology and genetics of tumours of the breast and female genital organs. Histopathology 46, 229–229 (2010).
Chen, H. et al. Invasive micropapillary carcinoma of the breast has a better long-term survival than invasive ductal carcinoma of the breast in spite of its aggressive clinical presentations: A comparison based on large population database and case-control analysis. Cancer Med. 6, 2775–2786. https://doi.org/10.1002/cam4.1227 (2017).
Tang, S. L. et al. Clinicopathologic study of invasive micropapillary carcinoma of the breast. Oncotarget 8, 42455–42465. https://doi.org/10.18632/oncotarget.16405 (2017).
Gokce, H. et al. Invasive micropapillary carcinoma of the breast: A clinicopathologic study of 103 cases of an unusual and highly aggressive variant of breast carcinoma. Breast J. 19, 374–381. https://doi.org/10.1111/tbj.12128 (2013).
Kuroda, H., Sakamoto, G., Ohnisi, K. & Itoyama, S. Clinical and pathologic features of invasive micropapillary carcinoma. Breast Cancer (Tokyo, Japan) 11, 169–174. https://doi.org/10.1007/bf02968297 (2004).
Zheng, H., Jian, L., Li, L., Liu, W. & Chen, W. Prior clinico-radiological features informed multi-modal MR images convolution neural network: A novel deep learning framework for prediction of lymphovascular invasion in breast cancer. Cancer Med. 13, e6932. https://doi.org/10.1002/cam4.6932 (2024).
Burstein, H. J. et al. Estimating the benefits of therapy for early-stage breast cancer: The St. Gallen International Consensus Guidelines for the primary therapy of early breast cancer 2019. Ann. Oncol. 30, 1541–1557. https://doi.org/10.1093/annonc/mdz235 (2019).
Van Zee, K. J. et al. A nomogram for predicting the likelihood of additional nodal metastases in breast cancer patients with a positive sentinel node biopsy. Ann. Surg. Oncol. 10, 1140–1151. https://doi.org/10.1245/aso.2003.03.015 (2003).
Gentilini, O. D. et al. Sentinel lymph node biopsy vs no axillary surgery in patients with small breast cancer and negative results on ultrasonography of axillary lymph nodes: The SOUND Randomized Clinical Trial. JAMA Oncol. 9, 1557–1564. https://doi.org/10.1001/jamaoncol.2023.3759 (2023).
Fan, Y. J. et al. Efficacy and safety comparison between axillary lymph node dissection with no axillary surgery in patients with sentinel node-positive breast cancer: A systematic review and meta-analysis. BMC Surg. 23, 209. https://doi.org/10.1186/s12893-023-02101-8 (2023).
Funding
This work was supported by the National Natural Science Foundation of China [grant numbers 82172025]; Chinese National Key Research and Development Project [grant number 2021YFC2500400].
Author information
Authors and Affiliations
Contributions
Rong Liang and Fangfang Li wrote the main manuscript text. Jingyuan Yao and Fang Tong prepared Figs. 1, 2, 3, 4, 5 and 6. Minghui Hua and prepared tables1-2. Junjun Liu prepared table 3. Chenlei Shi and Lewen Sui prepared table 4-5. Hong Lu revised the main manuscript text. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, R., Li, F., Yao, J. et al. Predictive value of MRI-based deep learning model for lymphovascular invasion status in node-negative invasive breast cancer. Sci Rep 14, 16204 (2024). https://doi.org/10.1038/s41598-024-67217-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-67217-0
- Springer Nature Limited