Abstract
Several pathogenic Escherichia coli strains cause diarrhea. Enteroaggregative E. coli (EAEC) strains are one of the diarrheagenic pathotypes. EAEC cells form a “stacked-brick” arrangement over the intestinal epithelial cells. EAEC isolates express, among other virulence determinants, the AggR transcriptional activator and the aggregative adherence fimbriae (AAF). Overexpression of the aggR gene results in increased expression of virulence factors such as the aff genes, as well as several genes involved in specific metabolic pathways such as fatty acid degradation (fad) and arginine degradation (ast). To support the hypothesis that induction of the expression of some of these pathways may play a role in EAEC virulence, in this study we used a murine infection model to evaluate the impact of the expression of these pathways on infection parameters. Mice infected with a mutant derivative of the EAEC strain 042, characterized by overexpression of the aggR gene, showed increased disease symptoms compared to those exhibited by mice infected with the wild type (wt) strain 042. Several of these symptoms were not increased when the infecting mutant, which overexpressed aggR, lacked the fad and ast pathways. Therefore, our results support the hypothesis that different metabolic pathways contribute to EAEC virulence.
Similar content being viewed by others
Introduction
Escherichia coli is one of the most important causes of diarrhea worldwide1,2. Diarrheagenic E. coli (DEC) strains are classified into distinct pathotypes, each distinguished by several features including virulence traits, host preferences, disease burden and mode of transmission3. Enteroaggregative Escherichia coli (EAEC) is one such pathotype4. Within the EAEC pathotype, strains exhibit genetic heterogeneity5,6,7, posing a challenge to understanding the mechanisms by which they cause disease6. Specific virulence determinants shared by all EAEC strains characterized to date include the plasmid-encoded aggR gene, and the aggR-regulated gene aafA. These genes encode the transcriptional activator AggR (a member of the AraC/XylS family) and the aggregative adherence fimbriae (AAF), respectively. AAF fimbriae play a crucial role in EAEC cell adhesion, facilitating attachment to each other and to host intestinal epithelial cells during infection. This adherence results in the formation of a characteristic stackedbrick pattern when cultured on Hep-2 cells8.
EAEC strains are etiological agents of enteric infections, leading to both acute and persistent diarrhea in both children and adults9,10,11,12. They are also implicated in traveller’s diarrhea and extraintestinal infections such as urinary tract infections13,14. An outbreak of foodborne hemorrhagic colitis in Germany in 2011 was traced back to an EAEC strain displaying specific genomic characteristics15. The strain identified, belonging to the O104:H4 serotype, expressed several typical EAEC virulence factors. However, it also expressed the Stx2a toxin, due to the presence of a phage in its genome encoding the stx2 gene. This genotypic characteristic led to the classification of the strain as a hybrid EAEC/STEC strain. This led to a wide range of works and the reclassification of several strains into hybrid strains.
The prototypical EAEC strain 042, which induced diarrhea in a volunteer trial, has been extensively studied16. Strain 042 harbors the IncFIIA plasmid pAA217,18 which encodes various virulence determinants, including the AggR virulence regulator and the AAF/II variant of the fimbrial adhesion determinant, which is essential for EAEC cell adherence to human intestinal mucosa17,19.
The microarray analysis technology was first used to identify the genes subject to AggR regulation in the 042 strain20. Both chromosomal and plasmidic genes were found to be regulated by AggR. AggR-dependent plasmid-encoded virulence factors include aggregative adherence fimbriae (AAF), the Aap dispersin along with its type I secretion system, the polysaccharide deacetylase encoded by the shf gene, and the Shigella flexneri virulence protein VirK20,21,22,23. Additionally, chromosomal determinants under AggR regulation, such as a type VI secretion system, are mainly located in chromosomal islands20. In subsequent studies, RNA-seq technology was used to study the AggR regulon in strain 04221. Comparative analysis of the transcriptome between the wt strain and an aggR mutant, revealed the regulation of 112 genes by AggR, including all but one of the genes identified in the previous microarray study20.
While investigating specific regulatory features of the aggR gene encoded by the plasmid pAA2, we demonstrated the role of sequences within the 3’UTR of the aggR gene in AggR expression. Insertion of foreign DNA sequences into this region resulted in constitutively high expression of AggR, and consequently increased expression of the AggR-regulated genes24. We compared the transcriptomes of the wt strain and that of an aggR::3’UTR mutant, specifically a clone with the FRT sequence following the aggR stop codon (strain aggR + FRT3’UTR), which exhibited abnormally high levels of AggR when cultured in nutrient rich medium. As expected, several genes previously identified as downregulated in the aggR mutant21, were upregulated in the 042 aggR + FRT3’UTR strain24. Unexpectedly, among the genes displaying the highest foldchange values in the comparative transcriptomic studies were those encoding enzymes associated with specific metabolic pathways, such as the ast (arginine degradation) and fad (fatty acid degradation) pathways. Inactivation of these pathways in the 042 aggR + FRT3’UTR strain resulted in reduced virulence in a mouse infection model24.
Considering that these are relevant aspects of the mechanisms by which EAEC cause disease, we decided in this study, to further investigate the correlation between the overexpression of the ast and fad pathways in the 042 aggR + FRT3’UTR strain and its pathogenic potential. To achieve this, we evaluated the outcomes of infection of mice with the wt 042 strain and its mutant derivatives characterized by constitutively expressing increased levels of AggR, both with and without functional ast and fad enzymes. We investigated the immune response, the gut microbiota composition, and the concentration of fecal short-chain fatty acids (SCFAs). Our results show that several of the disease symptoms observed in mice infected with the 042 aggR + FRT3’UTR strain are not observed in mice infected with its ast fad derivative. In addition, we observed a different sex-dependent response to infection by the 042 wt strain and its mutant derivatives.
Results
Signs of infection in male and female mice after administration of the wt E. coli 042 strain and its mutant derivatives
In the present work, we initially investigated the immune response in mice following infection with the wt E. coli 042 and its two mutant derivatives, based on previous findings indicating that infection of mice with these strains alters cytokine expression in immune tissues24. Both female and male mice were used to assess potential sex differences in response to infection with the E. coli 042 wt, 042 aggR + FRT3′UTR and 042 aggR + FRT3′UTR ast fad strains.
Male mice have lower leukocyte counts in the MLN than female mice (Fig. 1a). Nevertheless, strain 042 aggR + FRT3′UTR ast fad induced lower leukocyte recruitment in the MLN than strain 042 aggR + FRT3′UTR in both sexes.
Effect of the infection by the different E. coli strains on the presence of leukocytes in MLN from mice. Panel (a), leukocytes count into MLN, panel (b), percentage of Th lymphocytes, panel (c), percentage of Ts/c lymphocytes, panel (d), activated Th lymphocytes, panel (e), percentage of Regulatory Th lymphocytes and panel (f), ratio of activated/regulatory Th lymphocytes. Results are expressed as mean ± SEM (n = 5–6 animals/condition).
For Th lymphocytes, no significant differences were observed between the different strains in either in female or in male mice (Fig. 1b). With respect to Ts/c lymphocytes, the only effect observed was a reduction in the percentage of this subset in female mice infected with the 042 aggR + FRT3′UTR ast fad strain (Fig. 1c). Although Th lymphocyte activation also showed sex-dependent variation (Fig. 1d), both female and male mice exhibited an increased percentage of activated Th lymphocytes when infected with both the 042 aggR + FRT3′UTR and 042 aggR + FRT3′UTR ast fad strains compared to both the control group and those infected with the wt strain.
Regarding regulatory Th (Treg) lymphocytes, female mice had a lower percentage of Treg lymphocytes after infection with each strain compared to controls. In male mice, infection with either the 042 wt or with the 042 aggR + FRT3′UTR ast fad strains did not alter the levels of Treg lymphocytes. In contrast, infection with the 042 aggR + FRT3′UTR strain resulted in reduced levels (Fig. 1e). Furthermore, the ratio of activated/Treg lymphocytes was higher in female mice infected with both mutant strains compared to uninfected mice and to mice infected with the 042 wt strain. In male mice, this ratio was highest in mice infected with the 042 aggR + FRT3′UTR strain, with moderate increases observed in mice infected with the 042 wt and 042 aggR + FRT3′UTR ast fad strains (Fig. 1f).
Regarding the expression of proinflammatory cytokines, no significant changes in the expression of Il-6, Tnf-α, Il-1β, Ifn-γ or Il-4 were observed in female mice infected with any of the three strains. However, some differences in the expression of Il-17 were observed (Fig. 2). Only female mice infected with E. coli strain 042 aggR + FRT3′UTR showed a significant increase in Il-17 expression, compared to control animals. Conversely, mice infected with the 042 aggR + FRT3′UTR ast fad strain showed lower Il-17expression compared to those infected with 042 wt or 042 aggR + FRT3′UTR.
In male mice, Il-1β expression increased when challenged with the 042 wt or 042 aggR + FRT3′UTR ast fad strains compared to control mice. Mice infected with the 042 aggR + FRT3′UTR strain showed no significant difference in Il-1β expression compared to control mice. In addition, male mice infected with the 042 aggR + FRT3′UTR and 042 aggR + FRT3′UTR ast fad strains showed significantly increased expression of Il-6 and Tnf-α compared to control mice and those infected with the wt strain. There were no changes in the expression of the cytokines Il-4 and Ifn-γ in male mice. In contrast, the expression of Il-17 was profoundly modified in animals infected with the 042 aggR + FRT3′UTR strain, resulting in significantly different levels compared to all other groups, including control mice, and those infected with the 042 wt or 042 aggR + FRT3′UTR ast fad strains.
With regard to Il-10 expression in the MLN, in both males and females, it follows a profile very similar to that observed for the percentage of regulatory T cells, reinforcing the data already observed. In female mice, its expression is reduced in animals infected with any of the E. coli, strains, although this reduction it is only significant in animals infected with strains 042 aggR + FRT3′UTR and 042 aggR + FRT3′UTR ast fad (Fig. 3). In male mice, animals infected with the strain 042 aggR + FRT3′UTR showed reduced expression compared to the other groups.
Concentration of SCFAs in the fecal samples from mice infected with the different 042 strains
Considering that strain 042 aggR + FRT3′UTR has been shown to overexpress the fad enzymes in vitro24, we hypothesized that infection with this strain would lead to a reduced concentration of SCFAs in the gut, due to increased fad activity. To support this hypothesis, we determined the presence of acetic, propionic, and butyric acids in fecal samples from both uninfected and infected mice. Samples from female and male mice were analyzed separately, and the results are shown in Fig. 4.
Significant changes in the SCFAs concentrations were only observed in the fecal samples of male mice, affecting all three SCFAs analyzed. Unexpectedly, samples from mice infected with the 042 aggR + FRT3′UTR strain showed the highest levels of acetic, propionic, and butyric acids compared to those observed in samples collected from mice infected with either off the other two strains. Acetic and propionic acid levels in control mice were similar to those observed in mice infected with the 042 wt strain. In contrast, a modest increase in the concentration of butyric acid was observed in mice infected with the 042 wt strain compared to control mice.
Concentrations of acetic, butyric, and propionic acids were lower in samples from male mice infected with the 042 aggR + FRT3′UTR ast fad strain than in samples from male mice infected with either the 042 wt or the 042 aggR + FRT3′UTR strains.
Gut microbiota profiles of mice infected with the different E. coli strains
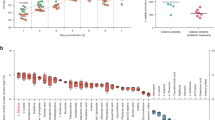
At the phylum level, Bacteroidota and Firmicutes largely predominated, with Proteobacteria, Cyanobacteria and Actinobacteria also present (Fig. 5). Interestingly, while the Bacteroidota/Firmicutes ratio is similar between male and female mice in uninfected animals, a sex-specific response is observed when mice are infected with the 042 strain and its derivatives: the ratio increases in female mice (1.54–2.03) and decreases in male mice (1.43–1.22) (Fig. 5). Additional analysis at the family and genus level showed little or no differences for infected female mice compared to control female mice (Supplementary Figs. S1 and S2). However detectable differences were observed between infected and control male mice, which are discussed below.
At the family level, the bacterial composition of male mice infected with the 042 wt strain was enriched in members of the Lachnospiraceae family (23.3%) compared to uninfected animals (16.6%) (Fig. 6a). Regarding animals infected with the two different 042 mutant derivatives, male mice infected with the 042 aggR + FRT3′UTR mutant showed increased levels of Muribaculaceae (41.55%) and Lactobacillaceae (6.7%), and decreased levels of Lachnospiraceae (19.5%) compared to male mice infected with the wt 042 strain (38.56%, 4.8% and 23.3%, respectively). However, animals infected with the 042 aggR + FRT3′UTR ast fad strain showed decreased levels of Lactobacillaceae (1.95%) and Muribaculaceae (35.38%), and increased levels of Lachnospiraceae (20.26%) compared to mice infected with 042 aggR + FRT3’UTR mutant strain. In fact, the changes in the bacterial composition of the gut microbiota of male mice infected with the 042 aggR + FRT3′UTR mutant strain compared to mice infected with the wt strain were not detected in mice infected with the 042 aggR + FRT3′UTR ast fad strain.
At the genus level, Bacteroides increased in all groups of infected male mice when compared to uninfected male mice, while Lactobacillus and Muribaculaceae genera decreased. When comparing the three groups of infected male mice, mice infected with the 042 aggR + FRT3′UTR strain showed higher levels of Lactobacillus (6.7%) and genera of Muribaculaceae (40.3%) and lower levels of the Lachnospiraceae NK4A136_group (7.75%) than the other two groups (Fig. 6b). Differentially abundant fecal bacterial taxa were further identified by LEfSe analysis. Mice infected with the 042 aggR + FRT3′UTR strain exhibit significant differences in the relative abundance of Clostridia UCG-014 and Enterococcaceae bacterium RF39 compared to the other two infected groups (Supplementary Fig. S3).
Despite the differences observed in relative abundances, no significant differences were found between infected and uninfected male mice when alpha diversity was tested using the Shannon and Chao indexes (Fig. 7). In addition, beta-diversity using Principal Coordinate Analysis (PCoA) based on weighted and unweighted UniFrac distances yielded similar data points on the plots of healthy and infected mice (Fig. 8).
Discussion
The results obtained in this study may contribute to a better understanding of the mechanisms by which EAEC strains cause disease. Unexpectedly, the murine infection model used revealed significant sex differences. Previous studies have also reported sex differences in murine infection models, with evidence suggesting a correlation between the differential response of male and female mice to an infection and the microbiota25,26. In our murine infection model using the EAEC strain 042 and its derivatives, significant changes in proinflammatory cytokines were observed exclusively in male mice. This pattern persisted when analyzing both the concentration of SCFAs in fecal samples and the gut microbiota response to infection. The differential response of male and female mice to 042 infection can be partially explained by examining the impact of infection on the gut microbiota at the phylum level. While the Bacteroidota/Firmicutes ratio remains constant between uninfected male and female mice, it increases in infected female mice and decreases in infected male mice. Increasing the Bacteroidota/Firmicutes ratio may enhance the resistance of female mice to EAEC strain 042 infection.
We took advantage of the higher susceptibility of male mice to E. coli 042 infection to assess whether the wt 042 strain and its two mutant derivatives have similar effects on the immune response to infection, fecal SCFAs concentration and gut microbiota composition. The relationship between E. coli infection, SCFAs and gut microbiota has been extensively studied in recent years27,28,29,30,31,32,33,34,35. In some cases, SCFAs such as butyrate can trigger virulence, as has been observed with enterohaemorrhagic E. coli29. Conversely, colonic SCFAs can inhibit the proliferation of infectious microorganisms by different mechanisms, including butyrate cellular signaling34 or intracellular acidification35. Comparative studies between healthy and EAEC-diarrheic neonatal dairy calves, have shown that diarrheic calves have reduced levels of butyrate and acetate compared to healthy animals27. In addition, in an experimental infection model with calves and E. coli O1, diarrheic calves were found to have increased Proteobacteria and decreased Firmicutes relative abundance. At the genus level, the relative abundance of Escherichia-Shigella was increased, while that of Lactobacillus was decreased, along with the SCFA content27. Recent work has shown that infection of mice with the EAEC strain 042 causes dysbiosis36.
The immunological response of male mice to infection by the 042 strain and its derivatives shows differences between the different E. coli strains. The number of leukocytes in the MLN increased in male mice infected with the 042 aggR + FRT3′UTR mutant strain. This increased leukocyte recruitment suggests a more intense immune response in these males. Additionally, mice infected with the 042 aggR + FRT3′UTR strain showed an increased proportion of activated Th lymphocytes. The combination of higher leukocyte recruitment and a greater proportion of activated Th lymphocytes points to a robust inflammatory response37. Notably, this strain caused a diminished anti-inflammatory response, as evidenced by the reduced presence of Tregs cells. In male mice, this duality suggests that the 042 aggR + FRT3′UTR strain induces a more pronounced inflammatory milieu.
In contrast, male mice infected with the 042 aggR + FRT3′UTR ast fad strain showed reduced leukocyte recruitment into the MLN and displayed a balance between pro-inflammatory Th cells and anti-inflammatory Tregs. This balance may suggest a modulatory effect of the inactivated ast and fad pathways, potentially dampening the inflammatory response and promoting a more regulated immune environment38.
The increased expression in infected mice of proinflammatory cytokines, including Il-1β, Il-6, and Tnf-α, is consistent with the increased leukocyte recruitment and Th cell activation. In particular, male mice infected with strain 042 aggR + FRT3′UTR ast fad exhibited a significant increase in Il-1β and Il-6 expression, although they showed a moderate Tnf-α response. This is consistent with the observed balanced Th/Treg ratio. In addition, the expression patterns of Ifn-γ, Il-4, and Il-17 provide further insight into the immune modulation of these bacterial strains. None of the strains exhibited modulation of the expression of cytokines relevant to either the Th1 (Ifn-γ) or Th2 (Il-4) response, suggesting that the immune response is not biased towards either the Th1 or Th2 pathway.
Il-17, which is associated with the Th17 response, was found to be significantly elevated in mice infected with strain 042 aggR + FRT3′UTR. Th17 lymphocytes are more directly involved in fighting intracellular pathogens and promoting inflammation39. This increase in Il-17 levels provides a robust inflammatory response, as Th17 cells are essential for recruiting neutrophils and mediating the clearance of extracellular pathogens40. Conversely, animals infected with strain 042 aggR + FRT3′UTR ast fad exhibited diminished Il-17 production.
Given that the 042 aggR + FRT3′UTR and 042 aggR + FRT3′UTR ast fad strains differ only in the inactivation of the ast and fad pathways, it is reasonable to expect that several infection parameters would be very similar, as observed. Nevertheless, the specific changes in some infection parameters exclusive to the 042 aggR + FRT3′UTR strain support the hypothesis that specific metabolic pathways play a role in the infection properties of this strain.
Based on our previous work with the 042 strain and the 042 aggR + FRT3′UTR mutant derivative24, we expected that proliferation of both the 042 wt and especially the 042 aggR + FRT3′UTR mutant (with upregulated expression of the fad pathway) would result in decreased levels of SCFAs in our male mouse infection model. Nevertheless, this was not the case. Compared to both to-uninfected animals and mice infected with the 042 wt strain, those infected with the 042 aggR + FRT3′UTR strain showed a significantly increased concentration of both acetic and propionic acids. In addition, animals infected with either strain show a significantly increased concentration of butyric acid when compared to non-infected animals. Conversely, animals infected with the 042 aggR + FRT3′UTR ast fad strain showed reduced levels of SCFAs compared to animals infected either with the 042 wt strain or with the 042 aggR + FRT3′UTR strain. These results can be interpreted as a depletion of SCFAs in the gut caused by the metabolic activity of the 042 aggR + FRT3′UTR strain, leading to a response from the gut microbiota that not only restores but also increases the concentration of these SCFAs. This in turn, may result in a less severe infection.
In contrast to other models using longer treatment times36, the effect of infection with the different 042 E. coli strains on the gut microbiota was not very significant, probably due to the short treatment time. This is supported by the analysis of alpha and beta diversity. However, some differences were observed. Compared to uninfected male mice, infected males showed increased levels of Lachnospiraceae, and lacked Muribaculaceae and Lactobacillaceae. Taking into account that animals infected with the 042 aggR + FRT3′UTR strain showed more significantly altered infection parameters and higher SCFAs production than those infected with the 042 wt strain, we also compared the microbiota composition of both groups. Increased levels of Muribaculaceae and Lactobacillaceae, and decreased levels of Lachnospiraceae were found in animals infected with the 042 aggR + FRT3′UTR strain. These changes were not observed in mice infected with the 042 aggR + FRT3′UTR ast fad derivative. These data indicate that the increased production of SCFAs in animals infected with the 042 aggR + FRT3′UTR strain can be correlated with higher levels of Muribaculaceae and Lactobacillaceae. It cannot be excluded that the observed predominance of Clostridia UCG-014 and Enterococcaceae bacterium RF39 in the fecal samples of these infected mice may also contribute to the increased production of SCFAs. The changes in the microbiota composition of male mice following infection with the 042 wt strain and its mutant derivatives fit well with previously reported data in calves and mice27,36.
We interpret the above data as follows: under the experimental conditions used, administration of the 042 wt strain induces mild signs of infection in male mice. When the infecting strain is the 042 aggR + FRT3′UTR strain, the symptoms of infection are more pronounced, but the infection is far from severe. Changes in the microbiota (either in its composition or in its metabolic activity) may contribute to the avoidance of severe infection symptoms in mice infected with the 042 aggR + FRT3′UTR strain, partly because of the increased production of SCFAs.
The association between the increased expression of the ast and fad pathways in the 042 aggR + FRT3′UTR strain24 and (i) an enhanced immune response, (ii) increased levels of SCFAs, and (iii) changes in the microbiota, compared to mice infected with the 042 wt strain, is supported by the fact that several of these distinguishing features are absent in male mice infected with the 042 aggR + FRT3′UTR ast fad strain.
E. coli infection has been previously associated with changes in the metabolome, both in humans and in calves41,42. In our model, we assume that the metabolic activity of the infecting E. coli strain is responsible for the changes in the metabolome that favors infection. To support this, we used the double ast fad mutant instead of the single ast and fad mutants in order to amplify and detect in vivo differences between male mice infected with the 042 aggR + FRT3′UTR strain and its derivative with altered function of specific metabolic pathways. The specific contribution of ast, fad or other metabolic pathways whose expression is altered in the 042 aggR + FRT3′UTR strain, awaits further research.
We noticed limitations in the study regarding microbiota analysis. The number of mice is limited to be able to make more robust statistical analyzes and, due to the sex differences found, this is more evident. Furthermore, it is possible that the treatment time with the E. coli 042 strain and its derivatives was not sufficient to displace the commensal microbiota of the mice and be able to see obvious changes in its composition.
Despite these limitations, certain trends can be seen in the composition of the microbiota depending on the E. coli derivative with which the infection was carried out, affecting the production of SCFAs. These changes can also be reflected in the activation of Th lymphocytes and the expression of cytokines, demonstrating the role that certain metabolic pathways can play in the infectious process.
Materials and methods
Bacterial strains, plasmids, and growth conditions
Bacterial strains and plasmids used in this study are outlined in Supplementary Table S1. The 042 aggR + FRT3′UTR mutant harbors an insertion of the FRT sequence after the stop codon of the aggR gene on the pAA2 plasmid. The ast mutation was generated by deleting the two first genes of the ast operon (astE and astB). The fad mutation was generated by deleting the fadA and fadB genes. Bacterial cultures were grown in either Luria broth (LB) medium (tryptone 10 g/L, yeast extract 5 g/L and sodium chloride 10 g/L), DMEM (supplemented with 0.45% glucose), or M9 minimal medium (supplemented with 0.4% glucose). Cultures were maintained at temperatures of either 25 °C or 37 °C with vigorous shaking at 200 rpm (Innova 3100 water bath shaker, New Brunswick Scientific). Liquid cultures were inoculated with a 1:100 dilution of cells previously cultured overnight (16 h at 37 °C) in LB with vigorous shaking at 200 rpm. When required, the media were supplemented with specific antibiotics at the following concentrations: carbenicillin (Cb) at 50 μg/mL, kanamycin (Km) at 50 μg/mL, chloramphenicol (Cm) at 25 μg/mL, or tetracycline (Tc) at 12.5 μg/mL.
In vivo infection of mice
C57BL/6 mice were purchased from Envigo (Bresso, Italy) and maintained under stable temperature and humidity conditions with a 12-h light 12-h dark cycle and free access to food and water. Mice were fed a standard chow diet. All animal experiments were carried out in strict adherence to the Guide for the Care and Use of Laboratory Animals, and all protocols were approved by the Ethics Committee for Animal Experimentation of the University of Barcelona and the Catalan government (Refs. 290/19 and 10,969, respectively). The study is in accordance with the ARRIVE guidelines.
At 8 weeks of age, a total of 48 mice were randomly divided into four groups (6 males and 6 females per group). The experimental groups were E. coli 042 wt infection, 042 aggR + FRT3′UTR infection, 042 aggR + FRT3′UTR ast fad infection, and a control group. Mice were inoculated by oral gavage either with 108 bacterial CFU of the respective bacterial strain or sterile saline with phosphate buffer (PBS), respectively. Additionally, three hours prior to bacterial inoculation, all mice received intraperitoneal (i.p.) administration of cimetidine (50 mg/kg; Sigma-Aldrich) to reduce acid secretion and enhance bacterial survival43.
Fecal samples collection
Fecal samples were collected at 4 days postinfection (dpi). Samples were immediately frozen in liquid nitrogen and stored at − 80 °C until further analysis. Aliquots were used both for DNA isolation and for the quantification of SCFAs.
Leukocyte isolation from mesenteric lymph nodes (MLN)
On the fifth dpi, mice were anesthetized with an intraperitoneal injection of ketamine/xylazine (100/10 mg/kg). Following anesthesia, euthanasia was performed by cervical dislocation, and mesenteric lymph nodes (MLN) were harvested and processed as previously described44. Briefly, MLN were finely minced and then incubated in a digestion solution composed of RPMI-1640 (Invitrogen, Carlsbad, CA, USA) supplemented with 5% inactivated fetal bovine serum (FBS), 100,000 U/L penicillin, 100 mg/L streptomycin, 10 mM HEPES, 2 nM L-glutamine, and 150 U/mL collagenase (Invitrogen, Carlsbad, CA, USA). This incubation step was carried out at 37 °C with agitation (Thermomixer Comfort Eppendorf, Heuppauge, NY, USA). MLN were mechanically disaggregated and passed through a stainless-steel mesh. The cell suspension was then centrifuged at 500 g for 10 min at 4 °C, and the pelleted cells were resuspended in PBS-FBS for further analysis.
Cell staining of leukocytes
The cell staining protocol for leukocytes was performed as described before45. Briefly, the staining was performed on 1.5 × 104 cells. For extracellular markers, cells were incubated with the primary antibodies for 30 min at 4 °C see Supplementary Table S2 for details). Intracellular markers were stained following fixation with 4% paraformaldehyde and permeabilization using 0.1% Triton-X-100® (Sigma-Aldrich). Subsequently, cells were washed and incubated with the primary antibodies specific to intracellular markers (see Supplementary Table S2). Finally, cells were washed and maintained in paraformaldehyde until further analysis using the Gallios Flow cytometer (Beckman Coulter, Miami, FL, USA), located at the Cytometry Unit of the Scientific-Technical Services of the Barcelona Science Park. Data analysis was performed using Flowjo Software (version 7.6.5, Treestar Inc., Ashland, OR, USA), with lymphocytes and non-lymphocytic leucocytes differentiated based on forward/side scatter characteristics.
Determination of cytokine expression by Real-Time PCR
RNA was extracted from MLN leukocytes as described previously44. Total RNA extracted was then reverse-transcribed using the High-capacity cDNA Reverse Transcription Kit (ThermoFisher Scientific, Waltham, MA, USA). Real-time PCR was performed on a MiniOpticon Real-Time PCR System (Bio-Rad, Hercules, CA, USA), employing primers listed in Supplementary Table S3. Product fidelity was confirmed through melt curve analysis. Each PCR run included duplicates of reverse transcription for each sample and appropriate negative controls (reverse transcription-free samples, RNA-free sample). Quantification of target gene transcripts was performed using hypoxanthine phosphoribosyl transferase 1 (Hprt1) gene expression as reference and was carried out using the 2-ΔΔCT method46.
DNA extraction and microbial profiling by 16S rRNA amplicon sequencing
DNA was isolated from fecal samples (80–100 mg per mouse) using the previously described protocol47.
For 16S rRNA sequencing, DNA samples were amplified by PCR targeting the V3 and V4 hypervariable regions of the bacterial 16S rRNA gene using specific primers: forward 5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3′ and reverse 5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3′. High-through sequencing was done with the Illumina MiSeq platform (Illumina, San Diego, CA, USA) at the Genomics and Bioinformatics Service, Universitat Autònoma de Barcelona (Bellaterra, Spain).
Bioinformatic analysis
Bioinformatic analysis of bacterial diversity was performed following stringent procedures. Raw reads underwent quality and length filtering, which involved removing of low-quality nucleotides at the 3′ end in windows of 20 nucleotides with low average quality, as well as eliminating of sequences with less than 200 nucleotides, using prinseq48and joined using fastq-join with a minimum overlap of 30 base pairs (bp)49. Primers and distal bases were trimmed, and singletons were removed using USEARCH v1150. zOTUs mapping to the human genome (GRCh38) were filtered out using the Burrow–Wheeler Aligner in Deconseq v0.4.3 (B6). The resulting reads were denoised and chimeras were filtered with UNOISE350.
Taxonomic assignment of zero-radius operational taxonomic units (zOTUs) was carried out using the Ribosomal Database Project51. The zOTUs were aligned with MAFFT52, and a phylogenetic tree was constructed with FASTTREE53, which was then midpoint-rooted. Reads classified as mitochondria and chloroplast were removed prior to statistical analysis. The taxonomic assignment of these zOTUs was obtained with a Naïve-Bayes classifier based on the scikit-learn system and the Silva v13854.
The samples were filtered out, with less than 30,000 reads for the final data analysis.
Quantification of SCFAs profiles in fecal samples
The concentration of short-chain fatty acids (SCFAs) was determined using an Agilent GC 7890B-5977 GC–MS system, equipped with a multipurpose sampler (Gerstel MPS). The GC column employed was an Agilent DB-FATWAX with dimensions of 30 m × 0.25 mm × 0.25 μm, operated in split mode Helium was employed as the carrier gas with a flow rate of 1 mL per minute. Samples from female and male mice were analyzed separately, with standard calibration curves previously established for all SCFAs analyzed.
The GC–MS analysis followed the protocol outlined55. As an internal standard, 3-methylvaleric acid was used at a concentration of 5 mM. Standard curves for acetic, propionic, and butyric acid were utilized to quantify SCFAs in the samples. For sample preparation, 200 mg of feces from each mouse were weighed, resuspended in 1 mL of PBS and centrifuged at 13,000 rpm for 5 min. Subsequently, 200 μL of supernatant samples were combined with 800 μL of the internal standard solution, 1 mL of diethyl ether, and a spoonful of Na2SO4. The mixture was vortexed for 10 s and then centrifuged at 1500×g for 2 min at 4 °C. Finally, 200 μL of the upper phase was transferred to a chromatography vial for analysis.
Statistical analysis
Data are expressed as mean ± standard error of the mean (SEM). Statistical analysis was performed using GraphPad Prism version 9 (GraphPad Software, Inc., La Jolla, CA, USA). First, Grubb’s test was applied for the detection and exclusion of outliers. Moreover, Levene’s test was used to assess the homogeneity of variance and the Shapiro–Wilk test to verify data normality across groups. For data adhering to a normal distribution, group comparisons were made using a two-way ANOVA, considering both the group condition and sex as factors. Subsequent post hoc analyses were conducted using Fisher’s Least Significant Difference (LSD) test. In contrast, the Kruskal–Wallis test was used for data that did not present a normal distribution. Statistical significance was considered when P values were < 0.05.
Alpha and Beta diversities (Unifrac matrix)56 were determined using the phyloseq R package57. Statistical analysis was performed using ANOVA (Alpha diversity) and PERMANOVA test (Beta-diversity) from the vegan R package58. Furthermore, a linear discriminant analysis effect size (LEfSe) was performed to identify specific bacterial biomarkers associated with health and disease states59 with the zOTUs assigned to genus taking the relative abundances of the samples infected with E. coli 042 and its derivatives. Bacteria occurring less than 20 times in at least 20% of the samples were filtered out.
Data availability
The datasets generated and analyzed during the current study are available in Zenodo repository at https://zenodo.org/records/12095587.
References
Troeger, C. et al. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: A systematic analysis for the Global Burden of Disease Study 2015. Lancet Infect. Dis. 17, 909–948 (2017).
Kotloff, K. L. et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global enteric multicenter study, GEMS): A prospective, case-control study. The Lancet 382, 209–222 (2013).
Nataro, J. P. & Kaper, J. B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. 11, 142–201 (1998).
Kaper, J. B., Nataro, J. P. & Mobley, H. L. T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2, 123–140 (2004).
França, F. L. S. et al. Genotypic and Phenotypic Characterisation of Enteroaggregative Escherichia coli from Children in Rio de Janeiro Brazil. PLoS One 8, e69971 (2013).
Estrada-Garcia, T. & Navarro-Garcia, F. Enteroaggregative Escherichia coli pathotype: A genetically heterogeneous emerging foodborne enteropathogen. FEMS Immunol. Med. Microbiol. 66, 281–298 (2012).
Boisen, N. et al. Redefining enteroaggregative Escherichia coli (EAEC): Genomic characterization of epidemiological EAEC strains. PLoS Negl. Trop. Dis. 14, e0008613 (2020).
Nataro, J. P. et al. Patterns of adherence of diarrheagenic Escherichia coli to HEp-2 cells. Pediatr. Infect. Dis. J. 6, 829–831 (1987).
Bhan, M. K. et al. Enteroaggregative Escherichia coli associated with persistent diarrhea in a cohort of rural children in India. J. Infect. Dis. 159, 1061–1064 (1989).
Fang, G. D., Lima, A. A. M., Martins, C. V., Nataro, J. P. & Guerrant, R. L. Etiology and epidemiology of persistent diarrhea in northeastern Brazil. J. Pediatr. Gastroenterol. Nutr. 21, 137–144 (1995).
Okeke, I. N., Lamikanra, A., Steinrück, H. & Kaper, J. B. Characterization of Escherichia coli strains from cases of childhood diarrhea in provincial southwestern Nigeria. J. Clin. Microbiol. 38, 7–12 (2000).
Wilson, A. et al. Characterisation of strains of enteroaggregative Escherichia coli isolated during the infectious intestinal disease study in England. Eur. J. Epidemiol. 17, 1125–1130 (2001).
Adachi, J. A. et al. Enteroaggregative Escherichia coli as a major etiologic agent in Traveler’s Diarrhea in 3 Regions of the world. Clin. Infect. Dis. 32, 1706–1709 (2001).
Boll, E. J. et al. Role of enteroaggregative Escherichia coli virulence factors in uropathogenesis. Infect. Immun. 81, 1164–1171 (2013).
Scheutz, F. et al. Characteristics of the enteroaggregative Shiga toxin/verotoxin-producing Escherichia coli O104:H4 strain causing the outbreak of haemolytic uraemic syndrome in Germany, May to June 2011. Eurosurveillance 16, (2011).
Nataro, J. P. et al. Heterogeneity of enteroaggregative Escherichia coli virulence demonstrated. J. Infect. Dis. 171, 465–468 (1995).
Chaudhuri, R. R. et al. Complete genome sequence and comparative metabolic profiling of the prototypical Enteroaggregative Escherichia coli strain 042. PLoS One 5, e8801 (2010).
Nataro, J. P., Scaletsky, I. C., Kaper, J. B., Levine, M. M. & Trabulsi, L. R. Plasmid-mediated factors conferring diffuse and localized adherence of enteropathogenic Escherichia coli. Infect. Immun. 48, 378–383 (1985).
Czeczulin, J. R. et al. Aggregative adherence fimbria II, a second fimbrial antigen mediating aggregative adherence in enteroaggregative Escherichia coli. Infect. Immun. 65, 4135–4145 (1997).
Morin, N., Santiago, A. E., Ernst, R. K., Guillot, S. J. & Nataro, J. P. Characterization of the AggR Regulon in Enteroaggregative Escherichia coli. Infect. Immun. 81, 122–132 (2013).
Yasir, M. et al. Organization and architecture of AggR-dependent promoters from enteroaggregative Escherichia coli. Mol. Microbiol. 111, 534–551 (2019).
Sheikh, J. et al. A novel dispersin protein in enteroaggregative Escherichia coli. J. Clin. Investig. 110, 1329–1337 (2002).
Nishi, J. et al. The export of coat protein from enteroaggregative Escherichia coli by a specific ATP-binding cassette transporter system. J. Biol. Chem. 278, 45680–45689 (2003).
Prieto, A. et al. Modulation of AggR levels reveals features of virulence regulation in enteroaggregative E. coli. Commun. Biol. 4, 1295 (2021).
Dibbern, J., Eggers, L. & Schneider, B. E. Sex differences in the C57BL/6 model of Mycobacterium tuberculosis infection. Sci. Rep. 7, 10957 (2017).
Efron, P. A. et al. Sex differences associate with late microbiome alterations after murine surgical sepsis. J. Trauma Acute Care Surg. 93, 137–146 (2022).
He, L. et al. Effects of pathogenic Escherichia coli infection on the flora composition, function, and content of short-chain fatty acids in Calf Feces. Animals 12, 959 (2022).
Pace, F. et al. The Short-Chain Fatty Acids Propionate and Butyrate Augment Adherent-Invasive Escherichia coli virulence but repress inflammation in a human intestinal enteroid model of infection. Microbiol Spectr 9, (2021).
Nakanishi, N. et al. Regulation of virulence by butyrate sensing in enterohaemorrhagic Escherichia coli. Microbiology (N Y) 155, 521–530 (2009).
He, Z. et al. Gut microbiota-derived ursodeoxycholic acid from neonatal dairy calves improves intestinal homeostasis and colitis to attenuate extended-spectrum β-lactamase-producing enteroaggregative Escherichia coli infection. Microbiome 10, 79 (2022).
Higginson, E. E. et al. Microbiome profiling of enterotoxigenic Escherichia coli (ETEC) carriers highlights signature differences between symptomatic and asymptomatic individuals. mBio 13, (2022).
Ducarmon, Q. R. et al. Gut microbiota and colonization resistance against bacterial enteric infection. Microbiol. Mol. Biol. Rev. 83, (2019).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Bäckhed, F. From dietary fiber to host physiology: Short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Byndloss, M. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 1979(357), 570–575 (2017).
Sorbara, M. T. et al. Inhibiting antibiotic-resistant Enterobacteriaceae by microbiota-mediated intracellular acidification. J. Exp. Med. 216, 84–98 (2019).
Moran-Garcia, N. et al. A novel adult murine model of typical enteroaggregative Escherichia coli Infection reveals microbiota dysbiosis, mucus secretion, and AAF/II-mediated expression and localization of β-catenin and expression of MUC1 in ileum. Front. Cell Infect. Microbiol. 12, (2022).
Miró, L. et al. Dietary supplementation with spray-dried porcine plasma attenuates colon inflammation in a genetic mouse model of inflammatory bowel disease. Int. J. Mol. Sci. 21, 6760 (2020).
Das, P., Lahiri, A., Lahiri, A. & Chakravortty, D. Modulation of the arginase pathway in the context of microbial pathogenesis: A metabolic enzyme moonlighting as an immune modulator. PLoS Pathog 6, e1000899 (2010).
Pérez-Bosque, A. et al. Oral serum-derived bovine immunoglobulin/protein isolate has immunomodulatory effects on the colon of mice that spontaneously develop colitis. PLoS One 11, e0154823 (2016).
Fan, X., Shu, P., Wang, Y., Ji, N. & Zhang, D. Interactions between neutrophils and T-helper 17 cells. Front. Immunol. 14, 1 (2023).
Gallardo, P. et al. Gut microbiota-metabolome changes in children with diarrhea by diarrheagenic E. coli. Front. Cell Infect. Microbiol. 10, (2020).
Shi, Z. et al. Multi-omics strategy reveals potential role of antimicrobial resistance and virulence factor genes responsible for Simmental diarrheic calves caused by Escherichia coli. mSystems. https://doi.org/10.1128/msystems.01348-23 (2024).
Ren, W. et al. Mouse intestinal innate immune responses altered by enterotoxigenic Escherichia coli (ETEC) infection. Microbes Infect. 16, 954–961 (2014).
Miró, L. et al. Dietary animal plasma proteins improve the intestinal immune response in senescent mice. Nutrients 9, 1346 (2017).
Pérez-Bosque, A., Miró, L., Amat, C., Polo, J. & Moretó, M. The anti-inflammatory effect of spray-dried plasma is mediated by a reduction in mucosal lymphocyte activation and infiltration in a mouse model of intestinal inflammation. Nutrients 8, 657 (2016).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Moretó, M. et al. Dietary supplementation with spray-dried porcine plasma has prebiotic effects on gut microbiota in mice. Sci. Rep. 10, 2926 (2020).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Aronesty, E. Comparison of sequencing utility programs. Open Bioinf. J. 7, 1–8 (2013).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. https://doi.org/10.1101/081257 (2016).
Cole, J. R. et al. Ribosomal Database Project: Data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642 (2014).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Eberhart, B. L., Wilson, A. S., O’Keefe, S. J. D., Ramaboli, M. C. & Nesengani, L. T. A simplified method for the quantitation of short-chain fatty acids in human stool. Anal. Biochem. 612, 114016 (2021).
Lozupone, C. & Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8, e61217 (2013).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.6–5 (2024).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Acknowledgements
M.B. thanks to the post-PhD grant “Juan de la Cierva” (ref. JDC2022-050269-I) MCIU/AEI and the European Union “Next Generation EU”/PRTR. This work was supported by grant PID2019-107479RB-I00 (AEI/FEDER, UE) and the Ministerio de Economía, Industria y Competitividad, and CERCA Program/Generalitat de Catalunya to A.J. RCR thanks Generalitat-Valenciana for the grant Plan GenT project (CDEIGENT 2020-02). IATA-CSIC authors also acknowledge the Spanish government MCIN/AEI to the Center of Excellence Accreditation Severo Ochoa (CEX2021-001189-S/MCIN/AEI/https://doi.org/10.13039/501100011033). A.P. and L.M. are members of the Institut de Recerca en Nutrició, i Seguretat Alimentària (INSA-UB), which is recognized as a Maria de Maeztu Unit of Excellence and funded by MICIN/AEI/FEDER (CEX2021-001234-M). A.P. and L.M. are members of Consolidated Research Groups (2021SGR300), Generalitat de Catalunya, Spain.
Author information
Authors and Affiliations
Contributions
AP and MH constructed the 042 mutant derivatives. AP and DS prepared the bacterial cultures for the different experiments performed. MB performed the SCFAs analysis. MB and RC performed the bioinformatic analysis. LM and AP performed the mice infection experiments, collected fecal samples, and analyzed the immunological response of mice. AJ and MC wrote the manuscript. AJ leaded the research project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bernabeu, M., Prieto, A., Salguero, D. et al. Infection of mice by the enteroaggregative E. coli strain 042 and two mutant derivatives overexpressing virulence factors: impact on disease markers, gut microbiota and concentration of SCFAs in feces. Sci Rep 14, 16945 (2024). https://doi.org/10.1038/s41598-024-67731-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-67731-1
- Springer Nature Limited