Abstract
Various animal and plant species exhibit allometric relationships among their respective traits, wherein one trait undergoes expansion as a power-law function of another due to constraints acting on growth processes. For instance, the acknowledged consensus posits that tree height scales with the two-thirds power of stem diameter. In the context of human development, it is posited that body weight scales with the second power of height. This prevalent allometric relationship derives its nomenclature from fitting two variables linearly within a logarithmic framework, thus giving rise to the term “power-law relationship.” Here, we challenge the conventional assumption that a singular power-law equation adequately encapsulates the allometric relationship between any two traits. We strategically leverage quantile regression analysis to demonstrate that the scaling exponent characterizing this power-law relationship is contingent upon the centile within these traits’ distributions. This observation fundamentally underscores the proposition that individuals occupying disparate segments of the distribution may employ distinct growth strategies, as indicated by distinct power-law exponents. We introduce the innovative concept of “multi-scale allometry” to encapsulate this newfound insight. Through a comprehensive reevaluation of (i) the height-weight relationship within a cohort comprising 7, 863, 520 Japanese children aged 5–17 years for which the age, sex, height, and weight were recorded as part of a national study, (ii) the stem-diameter-height and crown-radius-height relationships within an expansive sample of 498, 838 georeferenced and taxonomically standardized records of individual trees spanning diverse geographical locations, and (iii) the brain-size-body-size relationship within an extensive dataset encompassing 1, 552 mammalian species, we resolutely substantiate the viability of multi-scale allometric analysis. This empirical substantiation advocates a paradigm shift from uni-scaling to multi-scaling allometric modeling, thereby affording greater prominence to the inherent growth processes that underlie the morphological diversity evident throughout the living world.
Similar content being viewed by others
Introduction
Allometry explores the proportional relationship among different biological traits within organisms, often revealing patterns of growth and development. It provides insights into the fundamental principles governing biological form and function by examining how one trait changes in relation to another. Allometric scaling is typically depicted through a power-law function \(B=C\cdot A^{\alpha }\). A and B denote two biological traits, while \(\alpha\) and C represent scaling exponents and constants derived through statistical analysis, respectively. Allometric scaling has been found to exist across the living world and has led to the well-established metabolic scaling laws1,2,3, the nexus between brain size and cognitive acumen4,5,6, and the principles dictating the influence of respiratory surface on lifespan7,8. Rigorous examinations within biomechanics9,10,11,12,13,14, cellular dynamics15,16, the scaling dynamics of plant growth17,18,19, immune system dynamics20,21, reproductive strategies22,23,24,25,26,27, and intricate ecosystem interactions28,29 underscore the far-reaching implications of allometric relationships. These relationships highlight the foundational principles governing life on Earth, ranging from the microcosmic realm of molecules and mitochondria to the mesoscopic world of cells and mammals and extending to the macroscopic domains of communities and ecosystems30,31,32.
A meticulous examination exposes a notable heterogeneity in allometric relationships across organisms and ecological contexts. For instance, considerable variation in the scaling of metabolic rate to body mass has been documented among animals, unicellular organisms, and plants33; ontogenetic shifts in the metabolic intensity of vital processes, including growth, reproduction, locomotion, and heat production, have emerged as pivotal factors contributing to these divergent patterns of metabolic scaling, with organs exhibiting differential scaling in response to body mass25. Furthermore, the allometric link between body mass and metabolic rates showcases variability between ectotherms and endotherms and among metabolic states such as rest, field activity, and exercise34, and the stage of development35. Even the seemingly robust relationship between stem diameter and tree height is not impervious to alteration. Recent studies have reported that these allometric patterns might depend on factors like resource availability, notably drought in semi-arid forests36, and the growth stage of trees37. A mosaic of conflicting evidence has accumulated, fuelling intense debates regarding the adequacy of a universal scaling exponent to encapsulate the allometric relationship between a given pair of biological traits. These nuanced deviations from universal scaling underscore the pressing need for a more nuanced and context-specific comprehension of allometric relationships, thus challenging the notion of a one-size-fits-all approach in biology.
In the present study, we systematically evaluate the legitimacy of allometric relationships by examining the consistency of scaling exponents across the entire spectrum of centile curves within the trait-B-for-trait-A distribution. This uniform scaling exponent, called “uni-scaling,” forms the cornerstone of traditional allometric analysis. Conversely, varying exponents across centile curves lead to what we term ”multi-scaling.” Traditional allometric analysis has implicitly assumed uni-scaling, thus largely neglecting the potential existence of multiple scaling regimes. Based on the quantile regression technique38, we pioneer an innovative methodology for assessing uni-scaling and multi-scaling attributes within trait-B-for-trait-A distributions. To bolster our investigation, we submitted expansive datasets to multi-scaling allometric analysis; these datasets include (i) 7, 863, 520 records of Japanese children aged 5–17 years for both sexes, with measurements including height and weight; (ii) 1, 552 mammalian species with measurements including body size and brain size; (iii) 498, 838 georeferenced and taxonomic records of individual trees, sampled across diverse geographical locations, with measurements including stem diameter, crown radius, and height. Compared to the traditional uni-scaling allometric analysis, we show that the multi-scaling allometric analysis offers a fresh and more accurate perspective on the diversity of growth processes that underlie the morphological diversity evident throughout the living world.
Multi-scaling allometric analysis
The allometric scaling relationship \(B=C\cdot A^{\alpha }\) can be described as a solution to the differential equation
where dB/B represents the rate of change in trait B and dA/A represents the rate of change in trait A. Therefore, the scaling exponent quantifies the ratio between the rate of changes in trait B and trait A:
The traditional “uni-scaling allometric analysis” assumes that the relationship between trait A and trait B can be encapsulated by a singular scaling exponent denoted as \(\alpha\). Hence, any specific centile curve within the trait-B-for-trait-A distribution can be approximated through a power-law function \(B=CA^{\alpha }\), maintaining a consistent scaling exponent \(\alpha\) throughout. To empirically scrutinize this assumption, we conducted a rigorous assessment. Specifically, we fitted the power-law function to the qth centile curve, as visually represented by the solid line in Fig. 1, utilizing the quantile regression method38. In this analysis, the scaling exponent \(\alpha (q)\) can vary, introducing a nuanced perspective that adapts to different centiles (q). This extended analytical approach is denoted as the “multi-scaling allometric analysis.”
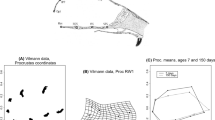
Schematic illustration of qth centile regression. In the multi-scaling allometric analysis, the qth centile (quantile) curve of a Trait-B-for-Trait-A distribution (a, b) is fitted by an extended power-law function \(B_{q}(A)=C_{q}\cdot A^{\alpha (q)}\) (c). This demonstration is based on synthetic data.
When the estimated exponents, \({\alpha (q)}\), exhibit independence from the centile index, denoted as q, this property is termed “uni-scaling” (parallel horizontal lines in Fig. 2a). In such instances, standardized allometric centile curves remain constant and unaffected by variations in height (parallel horizontal lines in Fig. 2c). Conversely, when the estimated exponents \({\alpha (q)}\) depend on the centile index, this property is referred to as “multi-scaling” (oblique lines in Fig. 2b). In multi-scaling scenarios, standardized allometric centile curves exhibit a discernible dependence on height (oblique lines in Fig. 2d). In uni-scaling scenarios, two points, denoted as \(c_{1}\) and \(c_{2}\), sharing the same standardized index, fall on the same 90th centile curve (Fig. 2c). In contrast, under multi-scaling conditions, \(c_{1}\) and \(c_{2}\) lie on distinct centile curves (Fig. 2d).
Schematic illustration of uni- and multi-scaling. (a) Uni-scaling in trait-B-for-trait-A distribution. In a double-logarithmic plot, all centile curves are parallel straight lines. (b) Multi-scaling in trait-B-for-trait-A distribution. In a double-logarithmic plot, slopes of the centile curves vary with position. (c) Uni-scaling centile curves in Index-for-trait-A distribution. Two points with the same Index, \(c_{1}\) and \(c_{2}\), lie on the same 90th centile curve. (d) Multi-scaling centile curves in Index-for-trait-A distribution. Two points with the same Index, \(c_{1}\) and \(c_{2}\), lie on distinct centile curves.
Results and discussion
Beyond body mass index (BMI): multi-scaling allometry in childhood growth
Body mass index (BMI) originates from a scaling law denoted as \(w=C\cdot h^{\alpha }\), wherein w and h represent the weight and height, respectively. It is empirically observed that the resting energy expenditure exhibits proportionality to \(w^{3/4}\) across a broad spectrum of animals, encompassing unicellular entities, metazoans, and homeothermic species—known as Kleiber’s law or the 3/4-power-law scaling39. Building upon this premise and under the assumption that energy expenditure is contingent upon body surface area to maintain a consistent body temperature40, it can be mathematically deduced that an individual’s weight is proportional to \(h^{2}\). Early investigations, incorporating relatively large sample sizes comprising thousands of cases, have substantiated that the scaling exponent relating weight and height across diverse racial and ethnic adult human cohorts approximates the value of 241,42. In this context, we raise a fundamental query concerning the uni-scaling scaling exponent relating weight to height.
We applied the multi-scaling allometric analysis to a public health dataset encompassing 7, 863, 520 records of Japanese children aged 5–17 years, documenting crucial parameters such as sex, age, height, and weight in a national study by the Japanese Ministry of Education, Culture, Sports, Science, and Technology. (This section methodologically represents several key findings relevant to the current study, which have been extensively detailed and discussed in other parts of a previously published report43.) Fig. 3 depicts the outcomes for males and females across various age groups, spanning from the 5th to the 95th centiles of weight-to-height distribution. Remarkably, our analysis revealed a diverse range of scaling exponents for 5-year-old children, deviating from the conventional assumption of 2 in BMI calculation (i.e., BMI=\(w/h^{2}\)). This variability becomes more pronounced as young children enter a phase of accelerated growth, with the most significant divergence occurring in children aged 8–10. Beyond the age of 10, the scaling exponents gradually converge, ultimately narrowing down to a tight range for males around 15 years and females as early as 12. Subsequently, the scaling exponent diminishes to smaller than 2 for 17-year-old males and 14-year-old females. This observation suggests that the traditional definition of BMI, assuming a uni-scaling exponent of 2, is applicable during the young adult phase but not during the rapid growth phases in both males and females.
Age dependence of allometric scaling exponents relating weight to height in growing children. (a) Males. (b) Females. The convergence of the scaling exponents for different qth centile weight-for-height distributions suggests a uni-scaling allometric relationship, while divergence signifies a multi-scaling allometric relationship.
We focus on the relationship and the BMI-for-height relationship within the cohort of 17-year-old females, a pivotal transitional phase on the verge of adulthood. Notably, all regressions for the 10th, 50th, and 90th centiles in the weight-for-height distribution boast parallel slopes when observed in the log-log domain. This pattern unequivocally signifies a well-defined uni-scaling allometric relationship between weight and height (Fig. 4a). By establishing standardized BMI as \(w/h^{\alpha (50)}[\textrm{kg}/\textrm{m}^{\alpha (50)}]\), with \(\alpha (50)=1.86\), we effectively stabilize the 10th, 50th, and 90th centile positions in the BMI-for-height distribution, making them impervious to variations in height (Fig. 4b). This underlines the substantial independence of BMI centiles from height fluctuations. Furthermore, the estimated BMI distributions across distinct height strata exhibit an impressive uniformity, a stark departure from the weight distributions shown in Fig. 4c, which manifest distinct shapes across various height strata. These findings testify to the credentials of BMI as an objective metric for appraising thinness and obesity, irrespective of height strata, as portrayed in Fig. 4d. In short, the uni-scaling allometric relationship between weight and height makes BMI a useful index for body-build assessment in 17-year-old females.
Older children, in this case, 17-year-old females, showed a uni-scaling allometric relationship between weight and height. (a) Weight-for-height distribution in the log-log space. (b) Standardized BMI, \(w/h^{1.85}\)-for-height distribution in the linear space. In (a) and (b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10nd, 50th, and 90th centile points in each height stratum defined within \(median\pm 3SD\). (c) The weight distribution for each height stratum. (d) The standardized BMI, \(w/h^{1.86}\), distribution for each height stratum. In (c) and (d), circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each height stratum.
However, as noted above, conventional BMI is not always an accurate, objective index of body-build assessment because the scaling exponents \({\alpha (q)}\) in males and females under 17 years of age do not converge around any specific scaling exponent. The non-parallel regression lines in Fig. 5a demonstrate the multi-scaling property of the weight-for-height distribution in 7-year-old females. In this example, the estimated slope of the median regression line was \(\alpha (50)=2.60\); therefore, we considered standardized BMI as \(w/h^{2.60}\). Unlike Fig. 4b, this result shows that the 10th and 90th regression lines are no longer horizontal (Fig. 5b), implying that the weight and standardized BMI distributions vary depending on the height strata (Fig. 5c, d). This result indicates that no unified formula for \(w/h^{\alpha }\), such as the BMI, can assess thinness and obesity in developing children.
Younger children, in this case, 8-year-old females, showed a multi-scaling allometric relationship between weight and height, questioning the validity of BMI. (a) Weight-for-height distribution in the log-log space. (b) Standardized BMI, \(w/h^{2.60}\)-for-height distribution in the linear space. In (a,b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each height stratum defined within \(median\pm 3SD\). (c) The weight distribution for each height stratum. (d) The standardized BMI, \(w/h^{2.60}\), distribution for each height stratum. In (c,d), circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each height stratum.
In summary, as the age of males and females approaches 17, an intriguing convergence emerges; all scaling exponents, \(\alpha (q)\), closely approximate a value of 2, as depicted in Fig. 4. However, during the earlier stages of development, a more complex picture unfolds, characterized by multiscale dynamics featuring distinct scaling exponents, \(\alpha (q)\). The age-dependent behavior of \(\alpha (q)\), which corresponds to the conventional allometric uni-scaling exponent (indicated by the red lines), exhibits a noteworthy trend. It commences at \(\alpha (50)=2\) around 5 years, progressively ascends to 3 around 10 years, and reverts to 2 in adulthood44,45. Intriguingly, the scaling exponents for higher (\(\alpha (10)\), light blue line) and lower (\(\alpha (90)\), dark blue lines) centiles present differing values relative to \(\alpha (50)\) during the age range of 5 to 13 years for males and 5 to 11 years for females. This complex age-dependent variation in scaling exponents adds a nuanced dimension to our understanding of growth dynamics. Together, these results unveil the intricacies of the weight-for-height relationship in children, challenging prior assumptions and the practice of uni-scaling allocentric relationship between weight and height. This prompts a critical reevaluation of the suitability of BMI as a truly objective measure for assessing body composition in children (cf. Refs.46,47,48,49).
Allometric multi-scaling of brain size in mammals: moving beyond the three-fourths power-law relationship
Brain and body size scaling offers a tantalizing avenue for unifying two foundational facets of biology: energy and information dynamics50. Conventionally, the scaling of brain size with body size adheres to a power-law expression: \(Brain\;size=C\cdot Body\;size^{\alpha}\), where C and \(\alpha\) are constants denoting the intercept and slope, respectively. Within mammals, one study encompassing a cohort of 1552 species spanning 28 distinct extant taxonomic orders reported a uni-scaling scaling exponent of 0.7551. This stands in contrast to other previous work, which proposed an exponent of 0.6652,53, but is congruent with others advocating for a three-fourths scaling relationship54,55,56. This allometric scaling, where brain size relates to body size with a three-fourths power, mirrors the fundamental underpinnings of metabolic theory in ecology, which governs the scaling of metabolic rate with body size50,57,58. Yet, the connection between brain size and metabolic scaling remains an intriguing enigma. Several investigations have examined the correlation between deviations in brain size and metabolic rate while meticulously controlling for body size. The prevailing consensus underscores a positive relationship between relative brain size and metabolic rate59,60. However, it is essential to underscore that this correlation, though discernible, tends to be relatively feeble61. Furthermore, its manifestation exhibits variations across taxonomic scales and exhibits sensitivity to phylogenetic considerations62. For instance, an extensive comparative analysis employing phylogenetic methods reported a scaling exponent of 0.563, based on the idea that slopes within taxonomic orders typically exhibit shallower trends than the median values encompassing various orders64. There are, however, exceptions to this general trend. Bats and primates display steeper scaling slopes, while lagomorphs and soricomorphs manifest a three-fourths scaling pattern with intercepts falling beneath the allometric trajectory observed for mammals. Prior scholars have engaged in extensive discourse concerning the diverse selection pressures that simultaneously mold brain size and body size, offering a myriad of scenarios for the evolution of correlated traits63,64,65,66. Yet, the recurring theme in all these studies is a uni-scaling allometric relationship between brain and body size.
We applied the multi-scaling allometric analysis to a dataset encompassing brain and body sizes derived from an extensive review of the scientific literature, encompassing 1, 552 species spanning across 28 distinct extant taxonomic orders51. The conventional brain-body size allometric relationship does not consistently serve as an objective measure for evaluating the energy and information dynamics across mammalian species, as the scaling exponents \({\alpha (q)}\) systematically deviate from the canonical value of 0.75, indicating multi-scaling allometric relationship. The non-parallel regression lines in Fig. 6a vividly illustrate this multi-scaling within the brain-size-for-body-size distribution across mammalian species. Because the estimated slope of the median regression line measures \(\alpha (50)=0.76\), we defined standardized brain-size-body-size as \(Brain\;size/Body\;size^{0.76}\). Notably, the 10th and 90th centile regression lines no longer remain horizontal, as depicted in Fig. 6b. This implies that the body size and standardized index distributions vary depending on the body size strata, as evidenced by the wide dispersion in the centile points in Fig. 6c, d. Hence, no single, unified allocentric relationship like \(Brain\;size\propto Body\;size^{0.75}\) in the log-log space can effectively characterize the energy and information dynamics across mammalian species.
Mammalian species showed a multi-scaling relationship between brain size and body size. (a) Brain-size-for-body-size distribution in the log-log space. (b) Standardized index, \(Brain\;size/Body\;size^{0.76}\)-for-brain-size distribution in the linear space. In (a,b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each brain size stratum defined within \(median\pm 2.5S.D.\) (from blue to yellow). (c) The brain size distribution for each body size stratum. (d) The standardized index, \(Brain\;size/Body\;size^{0.76}\), distributions for each body size stratum. In (c,d), circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each body size stratum.
We found a multi-scaling allometric relationship between brain and body size across mammalian species. The implications of this finding are profound and invite a reevaluation of the complex relationship between brain size and body size in the context of energy and information dynamics across mammalian species. Conventionally, a power-law expression has been used to describe the scaling of brain size with body size, with an exponent of 0.75. This relationship is rooted in the allometric scaling exponent relating metabolic rate to body mass (\(Metabolic\;rate\propto mass^{\alpha }\)) around 3/4. Nonetheless, the present findings challenge the presumed universality of the allometric relationship, revealing a systematic deviation from the conventional exponent of 0.75, indicative of a phenomenon known as multi-scaling. This departure from a singular scaling exponent underscores the intricate and nuanced nature of the brain-body size relationship, necessitating a more profound exploration of the underlying biological dynamics. A comprehensive meta-analysis encompassing 127 interspecific allometric relationships across diverse taxa, including birds, mammals, fish, reptiles, amphibians, arthropods, and unicellular organisms, spanning metabolic states from basal to maximum aerobic, exposes substantial heterogeneity in the impact of mass on metabolic rate34. However, it is important to acknowledge that allometric inquiries have played a pivotal role in shaping two fundamental debates that significantly influence our comprehension of brain evolution: the functional significance of brain size and the adaptive potential of brain structure52,67,68. These debates pivot on whether the observed patterns of scaling relationships, be it between brain and body size or different brain components, stem from selection pressures striving to maintain functional harmony or are constrained by shared developmental programs69. As we strive to enhance our understanding of the underlying factors driving the co-variation and co-evolution among diverse brain components, we gain valuable insights into how brains adapt to shifting selection pressures. It is crucial to appreciate that the relative importance of coordinated (concerted) and independent (mosaic) evolution of brain components may fluctuate over time and among distinct taxonomic groups, contingent upon the specific selection pressures exerted on brain size and structure. Discerning the circumstances favoring alternative pathways of phenotypic evolution presents a formidable challenge but is essential for unraveling the intricate mechanisms governing brain evolution70. Our findings highlight the imperative need to consider the statistical variability in allometric relationships resulting from the multifaceted selection pressures influencing body size and design, as illustrated by Ref.71.
The complexity of multi-scaling allometric relationships in tree growth: a comparative analysis of oak and pine strategies
Allometric relationships play a pivotal role in diverse ecological disciplines, offering a practical framework for scaling tree height, h, as a function of stem diameter, d, within and across plant species. These relationships typically rest on the foundational assumption that logarithmically transformed stem diameters and tree heights exhibit linear interdependencies, with a consistent slope maintained across varying environmental gradients30,72,73. Furthermore, there is a prevailing notion that the elastic similarity model, encapsulated by the equation \(h\propto d^{2/3}\), universally applies to all tree species74,75,76 and describes mechanical stability against gravitational forces arising from the self-weight of the trees (see77 for competing explanations; also see78). We questioned the established allometric uni-scaling relationship within this context, conventionally relating tree height and stem diameter. Our exploration aims to shed light on the potential variations in the geometrical and mechanical constraints underpinning the tree-height-to-stem-diameter relationship, particularly across shorter and taller trees.
We applied the multi-scaling allometric analysis to a publicly available dataset encompassing 498, 838 georeferenced records of individual trees across 61, 856 geographically diverse locations79. Remarkably, a significant proportion of trees documented within this database (\(88\%\)) have been accurately identified down to the species level, aggregating information about 5, 163 distinct species distributed across a taxonomic hierarchy comprising 1, 453 genera and 187 different plant families. We considered the seven families with data for at least 10, 000 trees. Figure 7 presents the outcomes for seven groups: Betulaceae, Cupressaceae, Fabaceae, Fagaceae, Lauraceae, Myrtaceae, and Pinaceae. As can be observed from the solid lines in Fig. 7, the regression lines for the 2th, 10th, 50th, 90th, and 98th centiles in Betulaceae, Cupressaceae, Fabaceae, Lauraceae, Myrtaceae, and Pinaceae exhibit a remarkable degree of uniform scaling, characterized by a scaling exponent close to 2/3. In stark contrast, the same regression lines in Fig. 7 reveal a conspicuous multi-scaling characteristic for Fagaceae.
Centile dependence of allometric scaling exponents relating tree height to stem diameter across different families. Stable scaling exponent, \(\alpha (q)\), across the entire range of centiles suggests a uni-scaling allometric relationship. In contrast, centile dependence of \(\alpha (q)\)—as in the case of Fagaceae, which includes oaks, beeches, and chestnuts—signifies a multi-scaling allometric relationship.
We first delved into the tree-height-for-stem-diameter relationship within the Pinaceae family, which includes pine, spruce, and fir, among other genera. When we examine within the log-log domain, all regressions for the 10th, 50th, and 90th centiles in the tree-height-for-stem-diameter distribution reveal parallel slopes, as depicted in Fig. 8a. The standardized tree-height-stem-diameter relationships in the form of \(h/d^{\alpha (50)}\), with \(\alpha (50)=0.73\), stabilizes the 10th, 50th, and 90th centiles in the tree-height-for-stem-diameter distribution against the influence of variations in stem diameter, as depicted in Fig. 8b. Moreover, when examining the standardized tree-height-stem-diameter distributions across various strata, we meet with a remarkable uniformity with the distinct shapes observed in the stem diameter distributions shown by the closely clustered centile points in Fig. 8d instead of spread-apart centile points in Fig. 8c. These results support the traditional tree-height-stem-diameter allometric relationship as a robust index for assessing tree growth in Pinaceae.
The Pinaceae family showed a uni-scaling allometric relationship between tree height, h, and stem diameter,d. (a) Tree-height-for-stem-diameter distribution in the log-log space (\(N=22,802\)). (b) Standardized index, \(h/d^{0.73}\)-for-stem-diameter distribution in the linear space. In (a,b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each tree height stratum defined within \(median\pm 2.5S.D.\) (c) The tree height distribution for each stem diameter stratum. (d) The standardized index, \(h/d^{0.73}\), distribution for each stem diameter stratum. In (c,d), circles, triangles, and squares represent the 10th, 50th, and 90th centile in each stem diameter stratum.
We found that the conventional tree-height-stem-diameter allometric relationship might not consistently serve as an objective measure for evaluating tree growth in Fagaceae, as the allometric scaling exponents \({\alpha (q)}\) systematically deviate from the expected value of 0.66, indicating a multi-scaling allometric relationship. The non-parallel regression lines in Fig. 9a vividly illustrate this multi-scaling within the tree-height-for-stem-diameter distribution in Fagaceae. In this particular case, the estimated slope of the median regression line measures \(\alpha (50)=0.37\). Consequently, we define standardized height-diameter as \(h/d^{0.37}\). Notably, in contrast to the pattern seen in Fig. 9b, the 10th and 90th regression lines no longer remain horizontal, as depicted in Fig. 9b. This implies that the tree height and standardized tree-height-stem-diameter distributions vary depending on the height strata, as evidenced by the closely clustered centile points in Fig. 9c, d. This outcome emphatically indicates that no single, unified allocentric relationship exists like \(h/d^{\alpha }\) that can characterize tree growth within the Fagaceae family.
The Fagaceae family showed a multi-scaling allometric relationship between tree height, h, and stem diameter, d. (a) Tree-height-for-diameter distribution in the log-log space (\(N=76,773\)). (b) Standardized index, \(h/d^{0.37}\)-for-stem-diameter distribution in the linear space. In (a,b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each tree height stratum defined within \(median\pm 2.5S.D.\) (c) The tree height distribution for each stem diameter stratum. (d) The standardized index, \(h/d^{0.37}\), distribution for each stem diameter stratum. In (c,d), circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each stem diameter stratum.
Exploring the same set of families for the allometric relationship between tree height, h, and crown radius, r—typically found to be around 2/3 (e.g., Ref80)—revealed some extent of multi-scaling allometric relationships in all these families (Fig. 10). Specifically, the scaling exponent for Betulaceae, Cupressaceae, Fabaceae, Lauraceae, Myrtaceae, and Pinaceae did not depend on the tree height stratum (e.g., Fig. 11). In contrast, the scaling exponent increased with q for Fagaceae (Fig. 12). This suggests that the relationship between crown radius and tree height is even more complex than stem diameter and tree height. Together, these results suggest that the uni-scaling allometric relationship between stem diameter and tree height, or between the crown radius and tree height, often regarded as a benchmark in forestry and ecology30,72,73, is not universally applicable.
Centile dependence of scaling exponents relating tree height to crown radius across different families. Stable scaling exponent, \(\alpha (q)\), across the entire range of centiles suggests uni-scaling. In contrast, all families show some extent of centile dependence of \(\alpha (q)\), signifying multi-scaling.
The family Pinaceae showed a multi-scaling allometric relationship between tree height, h, and crown radius, r. (a) Tree-height-for-crown-radius distribution in the log-log space (\(N=22,802\)). (b) Standardized index, \(h/r^{0.43}\)-for-radius distribution in the linear space. In (a,b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each tree height stratum defined within \(median\pm 2.5S.D.\) (c) The tree height distribution for each crown radius stratum. (d) The standardized index, \(h/r^{0.43}\), distributions for each crown radius stratum. In (c,d), circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each crown radius stratum.
The family Fagaceae showed a multi-scaling allometric relationship between tree height, h, and crown radius, r. (a) Tree-height-for-crown-radius distribution in the log-log space (\(N=76,773\)). (b) Standardized index, \(h/r^{0.40}\)-for-crown-radius distribution in the linear space. In (a,b), solid grey lines represent the contours; red lines represent the 10th, 50th, and 90th centile curves; circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each tree height stratum defined within \(median\pm 2.5S.D.\) (c) The tree height distribution for each crown radius stratum. (d) The standardized index, \(h/r^{0.40}\), distributions for each crown radius stratum. In (c,d), circles, triangles, and squares represent the 10th, 50th, and 90th centile points in each crown radius stratum.
The structural differences between the Fagaceae and Pinaceae families might explain the multi-scaling allometric relationships or a lack thereof. Fagaceae trees, characterized by spreading crowns with broad leaves, contrast with Pinaceae trees, which generally exhibit conical shapes with needle-like leaves81,82. This variation in crown architecture impacts light interception and distribution within the canopy, consequently affecting tree growth rates and development83. Trees with broader crowns may exhibit different biomass allocation patterns than narrower ones, potentially altering the tree-height-tree-diameter relationship. Fagaceae trees typically possess denser wood composed of vessels, fibers, and rays, whereas Pinaceae trees feature lighter, softer wood with tracheids84,85. These differences in wood density and structure influence the mechanical properties of the trees, such as their ability to support vertical growth and resist bending forces86. Hence, the growth rate for the diameter relative to height might vary, impacting the allometric relationship between these two parameters. Furthermore, the root systems of Fagaceae and Pinaceae trees differ in depth, extent, and anchorage mechanisms. Fagaceae trees often have extensive lateral root systems, providing enhanced anchorage in shallow soils87. In contrast, Pinaceae trees may develop deeper taproots or mycorrhizal associations, improving nutrient uptake and anchorage88. These root architecture differences might influence how trees allocate resources between height and diameter growth89,90, potentially modifying the allometric relationship. As a result, Fagaceae trees may depend more on stem diameter and crown radius than Pinaceae trees. Therefore, the allometric scaling exponent could diverge for trees with varying stem diameters and crown radii within Fagaceae more so than within Pinaceae.
In summary, the present results challenge the notion of a universal allometric relationship in tree growth and underscore the imperative of adopting multi-scaling assessment approaches in forestry and ecology. The contrasting growth strategies of oaks and pines are a testament to nature’s adaptability, each species optimizing its development based on its ecological niche and long-term survival objectives91,92. These complexities underscore the need to consider the nuanced biology and ecological context of specific tree families when evaluating the applicability of the canonical allometric relationship in understanding and managing forest ecosystems. They highlight the ecological context and growth strategies as pivotal factors in comprehending allometric relationships within tree growth. Indeed, the global biomass estimates based on tree crown radius (cf.93) might need reevaluation as they might have been underestimated in some cases and overestimated in others, necessitating further exploration using the multi-scaling allometry method proposed in this study.
General discussion
The results of the present investigation challenge the conventional assumption that a singular power-law equation sufficiently encapsulates the allometric relationship between any two biological traits related to growth processes. We demonstrated that the exponent characterizing this power-law relationship is contingent upon the centile position of a trait within its respective distribution. This revelation fundamentally underscores the proposition that individuals in distinct distribution segments may employ disparate growth strategies, as evidenced by discernible power-law exponents for each segment. We introduce the novel concept of “multi-scale allometry” to encompass this newfound insight. Through a comprehensive reevaluation of the allometric relationship across three distinct domains:
-
1.
A cohort of 7, 863, 520 Japanese children aged 5–17 years, for whom age, sex, height, and weight data were recorded as part of a national study.
-
2.
An extensive dataset encompassing brain and body sizes for 1552 mammalian species.
-
3.
A substantial sample of 498, 838 georeferenced and taxonomically standardized records of individual trees spanning diverse geographical locations, with stem diameter, crown radius, and tree height measurements.
Our analysis firmly establishes the validity of multi-scale allometry. The discovery of multi-scaling allometric relationships, as opposed to uniform allometric relationships, across diverse organismal contexts, shown in the present study, holds significant implications for comprehending growth processes. The observed variations in scaling relationships between correlated organismal traits across different centiles suggest that the allocation of resources may be contingent upon an individual’s position within the entire trait distribution. This observation underscores the existence of discrete growth strategies that individuals, species, or taxa may adopt. In essence, these findings underscore the intricate adaptability of biological systems and provide insights into the complexities of resource allocation for growth and survival. This nuanced perspective challenges the oversimplified concept of a singular, universal growth model that underlies so-called “universal” scaling exponents (cf.94) and emphasizes the necessity of accounting for context-specific variables when examining growth dynamics in the natural world.
It is crucial to clarify that the aforementioned claim centers around identifying a novel phenomenon in multi-scaling allometry rather than introducing a new analytical method. Therefore, while our current analysis methodology could certainly be refined in the future, the core focus remains on this groundbreaking discovery. Nonetheless, it is essential to recognize that even with methodologies like least squares95,96, which boast a rich history spanning over two centuries, novel phenomena remain unearthed and published today. Quantile regression38, distinct from the least squares method, is a generalization of the least absolute value method, yielding a solution by minimizing the sum of absolute residuals. Both least squares and least absolute value approaches can be applied to nonlinear fitting across various functions, including linear and quadratic polynomials. Notably, quantile regression enables the estimation of centile curves within the distribution, a capability absent in the least squares-based framework. While the origins of the least squares method trace back over two centuries to Gauss, quantile regression, with a history of approximately 50 years, necessitates substantial computational power for discovering optimal solutions, rendering it a realistic option only in recent times. Although quantile regression has not attained the same ubiquity as the least squares method, its potential utility across diverse studies is considerable, as evidenced by our successful application in allometry.
We wish to address a minor limitation of the multi-scaling allometric analysis in its current form. While highly informative, this approach demands substantial data resources, potentially confining its applicability primarily to intraspecific datasets and necessitating comprehensive and precise data across various centiles. This challenge is exacerbated when examining scaling relationships at extreme centiles, such as the 2nd and 98th centiles. Even our smallest dataset, comprising brain and body sizes for 1552 mammalian species, was notably extensive, enabling successful analysis across multiple centiles. Nonetheless, scenarios with more limited data may arise where the relationship between traits diverges significantly, warranting future advancements in multi-scaling allometric analysis methodologies to address such situations effectively.
We would also like to contrast the multi-scaling allometric analysis with a commonly used technique in allometry called the standardized major axis (SMA). The SMA fitting is particularly useful in allometric studies40,97,98 because it considers errors in both variables and does not assume a dependent-independent relationship, treating both variables symmetrically. This is crucial in biological studies where distinguishing between dependent and independent variables can be challenging, and both variables are often subject to measurement error99,100. However, SMA fitting relies on several assumptions to ensure accurate and meaningful results. (i) The relationship between the two variables is assumed to be linear. This means the data points should roughly align in a straight line when plotted on a graph. (ii) The data points should follow a bivariate normal distribution. This implies that the joint distribution of the two analyzed variables is approximately normal. (iii) The variances of the variables should be homogeneous across the range of data. This assumption ensures that the spread of data points is consistent along the entire line. (iv) Unlike simple linear regression, which assumes that only the dependent variable has measurement error, SMA fitting acknowledges that both variables may have measurement error. This is particularly important in allometry, where both the dependent and independent variables are biological measurements that can have associated errors. (v) The ratio of the standard deviations of the measurement errors (if present) should be constant across the range of data. This means that the relative accuracy of measurements for both variables remains consistent. (vi) The presence of significant outliers can distort the SMA fitting, although SMA assumes that no significant outliers disproportionately influence the fit. However, real-world data often violate one or more of these assumptions, making SMA an inappropriate technique in such cases. In contrast, multi-scale allometric analysis, which does not rely on these assumptions, yields valid results, as observed in the present study.
The concept of allometric relationships has played a pivotal role in enriching our understanding of biological phenomena and the underlying growth processes. It has specifically shed light on how various organismic traits scale with one another, often defined by power functions within specific physical constraints. Whether we consider complex growth patterns in human development or the towering majesty of trees, the prevalence of these allometric relationships has intrigued scientists for decades. Traditionally, the consensus held that a single power-law equation could effectively capture these intricate relationships. However, we have boldly challenged this conventional wisdom. We have harnessed the power of quantile regression analysis to illuminate a fascinating aspect of allometric relationships: the exponent characterizing these power laws is not a fixed constant; instead, it varies depending on the centile within the distribution of a specific trait. This revelation underscores a fundamental truth—individual organisms at different points along the distribution spectrum employ distinct growth strategies, leading to variations in the scaling exponent. We have introduced the concept of “multi-scale allometry” to encapsulate this newfound understanding, acknowledging that there is no one-size-fits-all approach to modeling these complex allometric relationships. Our journey through comprehensive reevaluations, from the heights and weights of millions of Japanese children to the intricate measurements of trees across diverse landscapes and even the morphological diversity of animal species, has firmly substantiated the concept of multi-scale allometry. This empirical validation advocates for a paradigm shift in approaching allometric modeling—moving from uni-allometric to multi-allometric perspectives. In embracing the nuances of growth processes, our results invite us to appreciate the rich tapestry of life’s diversity, where each organism may follow its unique development path.
Methods
Simulated dataset
We simulated two linearly correlated variables representing the association between two hypothetical biological traits when viewed in logarithmic space. We systematically adjusted the correlation coefficient to explore a range of values from 0.05 to 1, that is, \(r=0.05,0.1,\dots ,1\). This analysis allowed examining the impact of varying levels of correlation on these traits.
Empirical datasets
Dataset 1: comprehensive demographic analysis of school-age children: a decade-long study from 2008 to 2019
We sourced data from the School Health Statistics Research Initiative of the Japanese Ministry of Education, Culture, Sports, Science, and Technology. This national survey employs a rigorous stratified random sampling approach. It methodically stratifies schools within each prefecture based on student enrollment numbers, ensuring representation from small and large institutions. Within these size strata, it randomly selects children in the surveyed schools. It also surveys children using a systematic sampling method that considers age and gender. The survey is conducted annually, spanning from April to June. The obtained dataset consisted of demographic factors such as age, gender, height, and weight of 8, 083, 466 children aged 5–17 years collected from 2008–2019. The exclusion of \(2.72\%\) samples for missing values resulted in 7, 863, 520 samples.
Dataset 2: mammalian brain and body sizes: a taxonomic database of 1552 species
The second dataset comprised brain and body sizes derived from an extensive review of the scientific literature, encompassing a total of 1552 species spanning across 28 distinct extant taxonomic orders51. The compilation of this dataset adhered to rigorous criteria. The brain and body size measurements were drawn from the same primary sources whenever feasible and independently sourced from distinct references when this was not attainable. The sex of the specimens and pertinent sample size details were documented and reported, ranging from discrete values to open-ended ranges (e.g., \(1<n<10\)), wherever such information was available in the literature. Although typically, the mean values for adult individuals of both sexes were employed, for lineages exhibiting sexual size dimorphism, the authors relied on adult females’ brain and body size data per the established methodology101. The compiled data was subjected to a taxonomically driven standardization process, aligning with the taxonomy system102. This standardization was essential to facilitate compatibility with established comparative databases dealing with mammalian life history and ecological parameters103,104. When subspecies information was reported, weighted mean values for the species were computed, factoring in sample sizes. When diverse units of measurement were encountered, all measurements were converted to a consistent unit, transforming grams (1 g) to cubic centimeters (1 cm\(^{3}\)), as per prior studies conforming to this convention55.
Dataset 3: Tallo database: a comprehensive repository of georeferenced tree records for biodiversity and ecosystem research
The third dataset utilized in this study comprises entries from the “Tallo” database79. This repository consists of 498, 838 georeferenced records standardized in taxonomy, specifically focusing on individual trees. Critical parameters such as stem diameter, crown radius, and height have been systematically measured within these records. This dataset was compiled from a vast array of 61, 856 geographically diverse locations, encompassing a broad spectrum of forested and nonforested biomes: 5163 distinct species distributed across a taxonomic hierarchy comprising 1453 genera and 187 different plant families.
Multi-scaling allometric relationship for any given centile of weight-for-height distribution
We extended the allometric scaling to any centile of the weight-for-height distribution for each sex and age (Dataset 1), the brain-size-for-body-size distribution (Dataset 2), and the height-for-stem-diameter and height-for-crown-radius for each plant family (Dataset 3). The qth centile of a continuous random variable X is the value x such that \(\textrm{Prob}(X\le x)=q/100\). We considered that the conditional centile of the weight, given height (Dataset 1), brain size, given body size (Dataset 2), height, given stem diameter or crown radius (Dataset 3) follows an extended power-law model described as
where \(y_{q}(x)\) represents the qth centile body weight in kilograms (kg) given height x in meters (m; Dataset 1), the qth centile brain size in grams (g) given body size x in grams (g; Dataset 2), the qth centile tree height in meters (m) given stem diameter in centimeters (cm) or crown radius x in meters (m; Dataset 3), and the model parameters \(\alpha (q)\) and \(C_{q}\) represent the scaling exponent and proportionality constant, respectively. We employed the quantile regression approach to estimate the model parameters using custom Matlab 2023a (Mathworks Inc., Natick, MA)38.
Quantile regression
Quantile regression, a statistical method introduced by Koenker et al. in 197838, estimates a response variable’s conditional quantiles (centiles). In the context of an allometric model, this means we minimize the sum of weighted absolute residuals:
where \(y_i\) serves as our response variable, with i representing the individual index. The allometric model, characterized by \(f(x_i;C,\alpha )=Cx_i^{\alpha }\), encapsulates the relationship between x and y. We estimated the model parameters as \(\hat{C}\) and \(\hat{\alpha }\). To tailor the analysis to the desired centile (q), we set \(\tau\) to q/100 when applying the q-th quantile regression. For the task of parameter identification, we employed a straightforward search algorithm105.
Data availability
The height-weight data on Japanese school children are available from Japan’s Ministry of Education, Culture, Sports, Science, and Technology. Still, restrictions apply to the availability of these data, which were used under license for the current study and are not publicly available. Communication regarding this data must be directed to Ken Kiyono. The dataset for the mammalian brain and body sizes is publicly available51 (https://doi.org/10.1093/jmammal/gyz043) and the “Tallo” database can be obtained from a publicly available repository of georeferenced tree records for biodiversity and ecosystem research79 (https://doi.org/10.5281/zenodo.6637599).
References
Darveau, C.-A., Suarez, R. K., Andrews, R. D. & Hochachka, P. W. Allometric cascade as a unifying principle of body mass effects on metabolism. Nature 417, 166–170. https://doi.org/10.1038/417166a (2002).
Minias, P. & Podlaszczuk, P. Longevity is associated with relative brain size in birds. Ecol. Evolut. 7, 3558–3566. https://doi.org/10.1002/ece3.2961 (2017).
West, G. B., Woodruff, W. H. & Brown, J. H. Allometric scaling of metabolic rate from molecules and mitochondria to cells and mammals. Proc. Natl. Acad. Sci. 99, 2473–2478. https://doi.org/10.1073/pnas.012579799 (2002).
Beaulieu-Laroche, L. et al. Allometric rules for mammalian cortical layer 5 neuron biophysics. Nature 600, 274–278. https://doi.org/10.1038/s41586-021-04072-3 (2021).
Deaner, R. O., Isler, K., Burkart, J. & Van Schaik, C. Overall brain size, and not encephalization quotient, best predicts cognitive ability across non-human primates. Brain Behav. Evolut. 70, 115–124. https://doi.org/10.1159/000102973 (2007).
Kapellou, O. et al. Abnormal cortical development after premature birth shown by altered allometric scaling of brain growth. PLoS Med. 3, e265. https://doi.org/10.1371/journal.pmed.0030265 (2006).
Brown, J. H., Burger, J. R., Hou, C. & Hall, C. A. The pace of life: Metabolic energy, biological time, and life history. Integr. Comp. Biol. 62, 1479–1491. https://doi.org/10.1093/icb/icac058 (2022).
Lindstedt, S. & Calder, W. III. Body size, physiological time, and longevity of homeothermic animals. Q. Rev. Biol. 56, 1–16 (1981).
Biewener, A. A. Biomechanical consequences of scaling. J. Exp. Biol. 208, 1665–1676. https://doi.org/10.1242/jeb.01520 (2005).
Demes, B. & Günther, M. M. Biomechanics and allometric scaling in primate locomotion and morphology. Folia Primatol. 53, 125–141 (1989).
Rubenson, J. et al. Reappraisal of the comparative cost of human locomotion using gait-specific allometric analyses. J. Exp. Biol. 210, 3513–3524. https://doi.org/10.1242/jeb.000992 (2007).
West, B. J. & Griffin, L. Allometric control, inverse power laws and human gait. Chaos Solit. Fractals 10, 1519–1527. https://doi.org/10.1016/S0960-0779(98)00149-0 (1999).
Wilson, R. S., Franklin, C. E. & James, R. S. Allometric scaling relationships of jumping performance in the striped marsh frog Limnodynastes peronii. J. Exp. Biol. 203, 1937–1946. https://doi.org/10.1242/jeb.203.12.1937 (2000).
Zoeller, R. F. et al. Allometric scaling of isometric biceps strength in adult females and the effect of body mass index. Eur. J. Appl. Physiol. 104, 701–710. https://doi.org/10.1007/s00421-008-0819-2 (2008).
Miettinen, T. P. & Björklund, M. Mitochondrial function and cell size: An allometric relationship. Trends Cell Biol. 27, 393–402. https://doi.org/10.1016/j.tcb.2017.02.006 (2017).
Stone, K. C. et al. Allometric relationships of cell numbers and size in the mammalian lung. Am. J. Respir. Cell Mol. Biol. 6, 235–43. https://doi.org/10.1165/ajrcmb/6.2.235 (1992).
Enquist, B. J., Brown, J. H. & West, G. B. Allometric scaling of plant energetics and population density. Nature 395, 163–165. https://doi.org/10.1038/25977 (1998).
Enquist, B. J., West, G. B., Charnov, E. L. & Brown, J. H. Allometric scaling of production and life-history variation in vascular plants. Nature 401, 907–911. https://doi.org/10.1038/44819 (1999).
Muller-Landau, H. C. et al. Testing metabolic ecology theory for allometric scaling of tree size, growth and mortality in tropical forests. Ecol. Lett. 9, 575–588. https://doi.org/10.1111/j.1461-0248.2006.00904.x (2006).
Althaus, C. L. Of mice, macaques and men: Scaling of virus dynamics and immune responses. Front. Microbiol. 6, 355. https://doi.org/10.3389/fmicb.2015.00355 (2015).
Cable, J. M., Enquist, B. J. & Moses, M. E. The allometry of host–pathogen interactions. PloS One 2, e1130. https://doi.org/10.1371/journal.pone.0001130 (2007).
Gosden, R. & Telfer, E. Numbers of follicles and oocytes in mammalian ovaries and their allometric relationships. J. Zool. 211, 169–175. https://doi.org/10.1111/j.1469-7998.1987.tb07460.x (1987).
Kaplan, R. H. & Salthe, S. N. The allometry of reproduction: An empirical view in salamanders. Am. Nat. 113, 671–689 (1979).
McBride, R. S. et al. Energy acquisition and allocation to egg production in relation to fish reproductive strategies. Fish Fish. 16, 23–57. https://doi.org/10.1111/faf.12043 (2015).
Müller, M. J. et al. Effect of constitution on mass of individual organs and their association with metabolic rate in humans—A detailed view on allometric scaling. PloS One 6, e22732. https://doi.org/10.1371/journal.pone.0022732 (2011).
Savage, V. M., Deeds, E. J. & Fontana, W. Sizing up allometric scaling theory. PLoS Comput. Biol. 4, e1000171. https://doi.org/10.1371/journal.pcbi.1000171 (2008).
Weiner, J., Campbell, L. G., Pino, J. & Echarte, L. The allometry of reproduction within plant populations. J. Ecol. 97, 1220–1233. https://doi.org/10.1111/j.1365-2745.2009.01559.x (2009).
Brose, U., Williams, R. J. & Martinez, N. D. Allometric scaling enhances stability in complex food webs. Ecol. Lett. 9, 1228–1236. https://doi.org/10.1111/j.1461-0248.2006.00978.x (2006).
Mulder, C., Vonk, J. A., Den Hollander, H. A., Hendriks, A. J. & Breure, A. M. How allometric scaling relates to soil abiotics. Oikos 120, 529–536. https://doi.org/10.1111/j.1600-0706.2011.18869.x (2011).
West, G. B., Brown, J. H. & Enquist, B. J. A general model for the origin of allometric scaling laws in biology. Science 276, 122–126. https://doi.org/10.1126/science.276.5309.122 (1997).
West, G. B., Brown, J. H. & Enquist, B. J. The fourth dimension of life: Fractal geometry and allometric scaling of organisms. Science 284, 1677–1679. https://doi.org/10.1126/science.284.5420.1677 (1999).
West, G. B. & Brown, J. H. The origin of allometric scaling laws in biology from genomes to ecosystems: Towards a quantitative unifying theory of biological structure and organization. J. Exp. Biol. 208, 1575–1592. https://doi.org/10.1242/jeb.01589 (2005).
Glazier, D. S. Beyond the ‘3/4-power law’: Variation in the intra-and interspecific scaling of metabolic rate in animals. Biol. Rev. 80, 611–662. https://doi.org/10.1017/S1464793105006834 (2005).
White, C. R., Cassey, P. & Blackburn, T. M. Allometric exponents do not support a universal metabolic allometry. Ecology 88, 315–323. https://doi.org/10.1890/05-1883 (2007).
Riisgård. No foundation of a “3/4 power scaling law”’ for respiration in biology. Ecol. Lett. 1, 71–73 https://doi.org/10.1046/j.1461-0248.1998.00020.x (1998).
Dai, J. et al. Drought-modulated allometric patterns of trees in semi-arid forests. Commun. Biol. 3, 405. https://doi.org/10.1038/s42003-020-01144-4. (2020).
Osada, N. Height-dependent changes in shoot structure and tree allometry in relation to maximum height in four deciduous tree species. Funct. Ecol. 25, 777–786. https://doi.org/10.1111/j.1365-2435.2011.01833.x (2011).
Koenker, R. & Bassett Jr, G. Regression quantiles. Econometrica 33–50 https://doi.org/10.2307/1913643 (1978).
Kleiber, M. The Fire of Life. An Introduction to Animal Energetics. (Wiley, 1961).
Leibel, R. L., Rosenbaum, M. & Hirsch, J. Changes in energy expenditure resulting from altered body weight. N. Engl. J. Med. 332, 621–628. https://doi.org/10.1056/NEJM199503093321001 (1995).
Heymsfield, S. B. et al. Scaling of adult body weight to height across sex and race/ethnic groups: Relevance to BMI. Am. J. Clin. Nutr. 100, 1455–1461. https://doi.org/10.3945/2Fajcn.114.088831 (2014).
Hood, K. et al. Allometric scaling of weight to height and resulting body mass index thresholds in two Asian populations. Nutr. Diabetes 9, 2. https://doi.org/10.1038/s41387-018-0068-3 (2019).
Ogata, H. et al. Allometric multi-scaling of weight-for-height relation in children and adolescents: Revisiting the theoretical basis of body mass index of thinness and obesity assessment. PloS One 19, e0307238. https://doi.org/10.1371/journal.pone.0307238 (2024).
Cole, T. A critique of the NCHS weight for height standard. Hum. Biol. 57, 183–196 (1985) https://www.jstor.org/stable/41464748.
Cole, T. Weight/height\(^{p}\) compared to weight/height\(^{2}\) for assessing adiposity in childhood: Influence of age and bone age on \(p\) during puberty. Ann. Hum. Biol. 13, 433–451. https://doi.org/10.1080/03014468600008621 (1986).
Jayawardena, R. et al. Novel anthropometric parameters to define obesity and obesity-related disease in adults: A systematic review. Nutr. Rev. 78, 498–513. https://doi.org/10.1093/nutrit/nuz078 (2020).
Keys, A., Fidanza, F., Karvonen, M. J., Kimura, N. & Taylor, H. L. Indices of relative weight and obesity. J. Chronic Dis. 25, 329–343. https://doi.org/10.1016/0021-9681(72)90027-6 (1972).
Nuttall, F. Q. Body mass index: Obesity, BMI, and health: A critical review. Nutr. Today 50, 117. https://doi.org/10.1097/FNT.0000000000000092 (2015).
Smalley, K. J., Knerr, A. N., Kendrick, Z. V., Colliver, J. A. & Owen, O. E. Reassessment of body mass indices. Am. J. Clin. Nutr. 52, 405–408. https://doi.org/10.1093/ajcn/52.3.405 (1990).
Sibly, R. M., Brown, J. H. & Kodric-Brown, A. Metabolic Ecology: A Scaling Approach (Wiley, 2012).
Burger, J. R., George, M. A. Jr., Leadbetter, C. & Shaikh, F. The allometry of brain size in mammals. J. Mammal. 100, 276–283. https://doi.org/10.1093/jmammal/gyz043 (2019).
Jerison, H. Evolution of the Brain and Intelligence (Academic Press, 2012).
Sol, D., Bacher, S., Reader, S. M. & Lefebvre, L. Brain size predicts the success of mammal species introduced into novel environments. Am. Nat. 172, S63–S71. https://doi.org/10.1086/588304 (2008).
Boddy, A. et al. Comparative analysis of encephalization in mammals reveals relaxed constraints on anthropoid primate and cetacean brain scaling. J. Evolut. Biol. 25, 981–994. https://doi.org/10.1111/j.1420-9101.2012.02491.x (2012).
Isler, K. & van Schaik, C. P. The expensive brain: A framework for explaining evolutionary changes in brain size. J. Hum. Evolut. 57, 392–400. https://doi.org/10.1016/j.jhevol.2009.04.009 (2009).
Stankowich, T. & Romero, A. N. The correlated evolution of antipredator defences and brain size in mammals. Proc. R. Soc. B: Biol. Sci. 284, 20161857. https://doi.org/10.1098/rspb.2016.1857 (2017).
Brown, J. H., Gillooly, J. F., Allen, A. P., Savage, V. M. & West, G. B. Toward a metabolic theory of ecology. Ecology 85, 1771–1789. https://doi.org/10.1890/03-9000 (2004).
Brown, J. H., Hall, C. A. & Sibly, R. M. Equal fitness paradigm explained by a trade-off between generation time and energy production rate. Nat. Ecol. Evolut. 2, 262–268. https://doi.org/10.1038/s41559-017-0430-1 (2018).
Armstrong, E. Relative brain size and metabolism in mammals. Science 220, 1302–1304. https://doi.org/10.1126/science.6407108 (1983).
Martin, R. D. Relative brain size and basal metabolic rate in terrestrial vertebrates. Nature 293, 57–60. https://doi.org/10.1038/293057a0 (1981).
Isler, K. & Van Schaik, C. P. Metabolic costs of brain size evolution. Biol. Lett. 2, 557–560. https://doi.org/10.1098/rsbl.2006.0538 (2006).
Sobrero, R., May-Collado, L. J., Agnarsson, I. & Hernández, C. E. Expensive brains: “Brainy’’ rodents have higher metabolic rate. Front. Evolut. Neurosci. 3, 2. https://doi.org/10.3389/fnevo.2011.00002 (2011).
Tsuboi, M. et al. Breakdown of brain-body allometry and the encephalization of birds and mammals. Nat. Ecol. Evolut. 2, 1492–1500. https://doi.org/10.1038/s41559-018-0632-1 (2018).
Pagel, M. D. & Harvey, P. H. The taxon-level problem in the evolution of mammalian brain size: Facts and artifacts. Am. Nat. 132, 344–359. https://doi.org/10.1086/284857 (1988).
Lande, R. Quantitative genetic analysis of multivariate evolution, applied to brain: Body size allometry. Evolution 33, 402–416. https://doi.org/10.2307/2407630 (1979).
Smaers, J. B., Dechmann, D. K., Goswami, A., Soligo, C. & Safi, K. Comparative analyses of evolutionary rates reveal different pathways to encephalization in bats, carnivorans, and primates. Proc. Natl. Acad. Sci. 109, 18006–18011. https://doi.org/10.1073/pnas.1212181109 (2012).
Gould, S. J. Allometry and size in ontogeny and phylogeny. Biol. Rev. 41, 587–638. https://doi.org/10.1111/j.1469-185X.1966.tb01624.x (1966).
White, J. F. & Gould, S. J. Interpretation of the coefficient in the allometric equation. Am. Nat. 99, 5–18. https://doi.org/10.1086/282344 (1965).
Montgomery, S. H., Mundy, N. I. & Barton, R. A. Brain evolution and development: Adaptation, allometry and constraint. Proc. R. Soc. B Biol. Sci. 283, 20160433. https://doi.org/10.1098/rspb.2016.0433 (2016).
Healy, S. D. & Rowe, C. A critique of comparative studies of brain size. Proc. R. Soc. B Biol. Sci. 274, 453–464. https://doi.org/10.1098/rspb.2006.3748 (2007).
Font, E., García-Roa, R., Pincheira-Donoso, D. & Carazo, P. Rethinking the effects of body size on the study of brain size evolution. Brain Behav. Evolut. 93, 182–195. https://doi.org/10.1159/000501161 (2019).
Feldpausch, T. R. et al. Height-diameter allometry of tropical forest trees. Biogeosciences 8, 1081–1106. https://doi.org/10.5194/bg-8-1081-2011 (2011).
Hulshof, C. M., Swenson, N. G. & Weiser, M. D. Tree height-diameter allometry across the United States. Ecol. Evolut. 5, 1193–1204. https://doi.org/10.1002/ece3.1328 (2015).
Bertram, J. E. Size-dependent differential scaling in branches: The mechanical design of trees revisited. Trees 3, 241–253. https://doi.org/10.1007/BF00225358 (1989).
McMahon, T. A. & Kronauer, R. E. Tree structures: Deducing the principle of mechanical design. J. Theor. Biol. 59, 443–466. https://doi.org/10.1016/0022-5193(76)90182-X (1976).
Norberg, R. A. Theory of growth geometry of plants and self-thinning of plant populations: Geometric similarity, elastic similarity, and different growth modes of plant parts. Am. Nat. 131, 220–256. https://doi.org/10.1086/284787 (1988).
Niklas, K. J. & Spatz, H.-C. Growth and hydraulic (not mechanical) constraints govern the scaling of tree height and mass. Proc. Natl. Acad. Sci. 101, 15661–15663. https://doi.org/10.1073/pnas.0405857101 (2004).
Sperry, J. S., Meinzer, F. C. & McCulloh, K. A. Safety and efficiency conflicts in hydraulic architecture: Scaling from tissues to trees. Plant Cell Environ. 31, 632–645. https://doi.org/10.1111/j.1365-3040.2007.01765.x (2008).
Jucker, T. et al. Tallo: A global tree allometry and crown architecture database. Glob. Change Biol. 28, 5254–5268. https://doi.org/10.1111/gcb.16302 (2022).
Dai, X. et al. Plant height-crown radius and canopy coverage-density relationships determine above-ground biomass-density relationship in stressful environments. Biol. Lett. 5, 571–573. https://doi.org/10.1098/rsbl.2009.0228 (2009).
Burnham, R. J. & Johnson, K. R. South American Palaeobotany and the origins of neotropical rainforests. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 359, 1595–1610. https://doi.org/10.1098/rstb.2004.1531 (2004).
Niinemets, Ü. & Valladares, F. Tolerance to shade, drought, and waterlogging of temperate northern hemisphere trees and shrubs. Ecol. Monogr. 76, 521–547. https://doi.org/10.1890/0012-9615(2006)076[0521:TTSDAW]2.0.CO;2 (2006).
Hara, T. Dynamics of size structure in plant populations. Trends Ecol. Evolut. 3, 129–133. https://doi.org/10.1016/0169-5347(88)90175-9 (1988).
Carlquist, S. Comparative Wood Anatomy: Systematic, Ecological, and Evolutionary Aspects of Dicotyledon Wood (Springer, 2013).
Shmulsky, R. & Jones, P. D. Forest Products and Wood Science: An Introduction (Wiley, 2019).
Smith, D. D. et al. Deviation from symmetrically self-similar branching in trees predicts altered hydraulics, mechanics, light interception and metabolic scaling. New Phytol. 201, 217–229. https://doi.org/10.1111/nph.12487 (2014).
Mahall, B. E., Tyler, C. M., Cole, E. S. & Mata, C. A comparative study of oak (Quercus, Fagaceae) seedling physiology during summer drought in southern California. Am. J. Bot. 96, 751–761. https://doi.org/10.3732/ajb.0800247 (2009).
Martínez-Vilalta, J., Sala, A. & Piñol, J. The hydraulic architecture of Pinaceae—A review. Plant Ecol. 171, 3–13. https://doi.org/10.1023/B:VEGE.0000029378.87169.b1 (2004).
Eissenstat, D. & Yanai, R. The ecology of root lifespan. In Advances in Ecological Research, vol. 27, 1–60 (Elsevier, Amsterdam, Netherlands, 1997).
Fitter, A. Characteristics and functions of root systems. In Plant Roots. 49–78 (CRC Press, 2002).
Morán-López, T., Poyatos, R., Llorens, P. & Sabaté, S. Effects of past growth trends and current water use strategies on Scots pine and pubescent oak drought sensitivity. Eur. J. For. Res. 133, 369–382. https://doi.org/10.1007/s10342-013-0768-0 (2014).
Martín-Gómez, P., Aguilera, M., Pemán, J., Gil-Pelegrín, E. & Ferrio, J. P. Contrasting ecophysiological strategies related to drought: The case of a mixed stand of scots pine (Pinus sylvestris) and a submediterranean oak (Quercus subpyrenaica). Tree Physiol. 37, 1478–1492. https://doi.org/10.1093/treephys/tpx101 (2017).
Goodman, R. C., Phillips, O. L. & Baker, T. R. The importance of crown dimensions to improve tropical tree biomass estimates. Ecol. Appl. 24, 680–698. https://doi.org/10.1890/13-0070.1 (2014).
Hulbert, A. J. A sceptics view: “Kleiber’s law’’ or the “3/4 rule’’ is neither a law nor a rule but rather an empirical approximation. Systems 2, 186–202. https://doi.org/10.3390/systems2020186 (2014).
Björck, Å. Least squares methods. Handbook of Numerical Analysis 1, 465–652. https://doi.org/10.1016/S1570-8659(05)80036-5 (1990).
Björck, Å. Numerical Methods for Least Squares Problems (SIAM, 1996).
Warton, D. I., Wright, I. J., Falster, D. S. & Westoby, M. Bivariate line-fitting methods for allometry. Biol. Rev. 81, 259–291. https://doi.org/10.1017/S1464793106007007 (2006).
Wright, I. J. et al. Modulation of leaf economic traits and trait relationships by climate. Glob. Ecol. Biogeogr. 14, 411–421. https://doi.org/10.1111/j.1466-822x.2005.00172.x (2005).
Clarke, M. The reduced major axis of a bivariate sample. Biometrika 67, 441–446. https://doi.org/10.1093/biomet/67.2.441 (1980).
Smith, R. J. Use and misuse of the reduced major axis for line-fitting. Am. J. Phys. Anthropol. 140, 476–486. https://doi.org/10.1002/ajpa.21090 (2009).
Isler, K. & van Schaik, C. P. Allomaternal care, life history and brain size evolution in mammals. J. Hum. Evolut. 63, 52–63. https://doi.org/10.1016/j.jhevol.2012.03.009 (2012).
Wilson, D. E. & Reeder, D. M. Mammal Species of the World: A Taxonomic and Geographic Reference Vol. 1 (Johns Hopkins University Press, 2005).
Jones, K. E. et al. PanTHERIA: A species-level database of life history, ecology, and geography of extant and recently extinct mammals: Ecological archives E090-184. Ecology 90, 2648–2648. https://doi.org/10.1890/08-1494.1 (2009).
Myhrvold, N. P. et al. An amniote life-history database to perform comparative analyses with birds, mammals, and reptiles: Ecological archives E096-269. Ecology 96, 3109–3109. https://doi.org/10.1890/15-0846R.1 (2015).
Nelder, J. A. & Mead, R. A simplex method for function minimization. Comput. J. 7, 308–313. https://doi.org/10.1093/comjnl/7.4.308 (1965).
Acknowledgements
Madhur Mangalam was supported by the Center of Research in Human Movement Variability at the University of Nebraska at Omaha, funded by the National Institute of General Medical Sciences (NIGMS; Grant No. P20GM109090). Ken Kiyono was supported by the Japan Ministry of Health, Labour and Welfare (MHLW) FA Program (Grant No. JPMH22FA1023) and the Japan Society for the Promotion of Science (JSPS) KAKENHI (Grant Nos. 22KK0161 and 23H01647).
Author information
Authors and Affiliations
Contributions
Madhur Mangalam: Conceptualization, Methodology, Software, Validation, Formal analysis, Writing—Original draft, Writing—Review & Editing, Visualization; Yosuke Isoyama: Formal analysis; Writing—Review & Editing; Hitomi Ogata: Conceptualization, Writing—Review & Editing; Sayaka Nose-Ogura: Conceptualization, Writing—Review & Editing; Momoko Kayaba: Conceptualization, Writing—Review & Editing; Narumi Nagai: Conceptualization, Writing—Review & Editing; Ken Kiyono: Conceptualization, Methodology, Software, Writing—Review & Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mangalam, M., Isoyama, Y., Ogata, H. et al. Multi-scaling allometry in human development, mammalian morphology, and tree growth. Sci Rep 14, 19957 (2024). https://doi.org/10.1038/s41598-024-69199-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69199-5
- Springer Nature Limited