Abstract
The gut microbiota plays an important role in the immunology, physiology and growth and development of animals. However, currently, there is a lack of available sequencing data on the gut microbiota of Asian badgers. Studying the gut microbiota of Asian badgers could provide fundamental data for enhancing productivity and immunity of badgers’ breeding, as well as for the protection of wild animals. In this study, we first characterized the composition and structure of the gut microbiota in the large intestines of wild and captive Asian badgers during summer and winter by sequencing the V3-V4 region of the 16S ribosomal RNA gene. A total of 9 dominant phyla and 12 genera among the bacterial communities of the large intestines exhibited significant differences. Our results showed that Firmicutes and Proteobacteria were the most predominant in both wild and captive badgers, regardless of the season. Romboutsia, Streptococcus and Enterococcus may represent potential sources of zoonoses, warranting further attention and study. Our findings indicated that the diversity and availability of food resources were the most important influencing factors on the gut microbiota of Asian badgers, providing fundamental data for the protection and conservation of wild animals. Variation in the gut microbiota due to season, age and sex in both wild and captive Asian badgers should be considered in future research directions. Furthermore, combined multi-omics studies could provide more information for wild animal conservation, and enhancing our understanding of the molecular mechanism between the microbiota and host.
Similar content being viewed by others
Introduction
The Asian badger (Meles leucurus) is a species of Mustelidae widely distributed across central Asia, including the southern portion of Russia east of the Urals, Mongolia, Kazakhstan, China and the Korean Peninsula1. The taxonomic status of badgers is comparatively controversial due to the collision between traditional taxonomy2,3,4 and modern molecular biology5,6. Although classified as least concern class by the International Union for the Conservation of Nature (IUCN)7, the Asian badger is on the list of terrestrial wildlife that is beneficial or of important economic or scientific research value under state protection in China. However, significant declines in the badger population have been observed due to increased human activity, illegal hunting, habitat loss and fragmentation8,9. Fortunately, with the establishment of the National Park for the Amur Tiger and Amur Leopard in China, the detection rate of Asian badgers is gradually increasing. The variation in population and distribution range of Asian badgers, considered crucial secondary consumers, has a significant impact on the balance and stability of the entire ecosystem. Moreover, Asian badgers play other important roles in the economy and traditional Chinese medicine10.
Studies on European badgers were conducted relatively earlier and in more depth than those on Asian badgers, covering various aspects of their ecology, physiology and behavior11,12,13,14, as well as population structure and dynamics15,16. Currently, most previous studies on Asian badgers focus on habitat selection, behavioral ecology, activity rhythms and the breeding industry17,18,19. Although studies of Asian badgers using molecular biology methods, including phylogenetic analysis and genetic diversity assessment, are increasing20,21, a deep understanding of this species and appropriate protection strategies remain insufficient, especially in China.
In the past decade, the gut microbiota has become a research focus due to its role in nutrient availability, food digestion and host protection from pathogens, as well as its influence on host behavior, development, reproduction, and health22,23. The composition and dynamics of the host gut microbiota are determined by external factors, such as the environment, food resources and behavior, as well as internal factors, including diet, age, sex and health status24,25. However, gut microbiota data of Asian badgers have not yet been reported.
The primary objective of this study is to characterize the basic bacterial community composition and structure of the large intestine contents from both captive and wild Asian badgers for the first time, and to compare the gut microbiota of badgers in different seasons. While it’s true that having data on basic gut microbiome composition may not directly translate into practical conservation or management strategies, it can still serve as a valuable baseline for understanding the health and ecology of the Asian badger population. By establishing this benchmark, researchers can monitor changes in the gut microbiota over time, which may reflect broader shifts in the environment or health status of the badgers. Additionally, understanding the gut microbiota of Asian badgers can provide insights into their dietary habits, disease susceptibility, and overall well-being, all of which are critical factors for effective conservation and management efforts.
Materials and methods
Sample collection
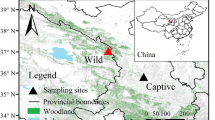
A total of 25 large intestine samples from Asian badgers were collected between September 2018 and December 2019. Sample information and sampling locations are provided in Table S1 and Figure S1 (Oviframe Interactive Map, Version V9.9.8.31753, https://www.ovital.com/product/), respectively. Samples from captive Asian badgers during both summer (CS1-CS7) and winter (CW1-CW8) were collected from Haodong Farm in Cangzhou, Hebei Province, China. The farmed badgers were fed mainly corn, chicken and modified feed for foxes, which ensured a sufficient food supply in both summer and winter. The captive badgers were housed individually in outdoor enclosures and were euthanized by electrocution. The large intestines were collected immediately from deceased Asian badger individuals on the farm. The entire sampling process was conducted by a professional veterinarian. All captive individuals have not received veterinary treatments or antibiotics.
Large intestine samples from wild Asian badgers in summer (WS1-WS6) and winter (WW1-WW4) were obtained from deceased badgers stored at the Forest Public Security Bureau of Hunchun, Jilin Province. Police officers collected the corpses of wild Asian badgers that died of natural causes or road accidents during their daily patrols and temporarily stored them at −20 ℃ within 1–2 h after discovering these bodies. The large intestine sampling process and scientific research on deceased badgers were permitted by the Forest Public Security Bureau of Hunchun. All necropsies were conducted by professionals in a sterile environment within the laboratory.
All the large intestine samples were stored in a customized freezer at − 20 ℃ and then transferred to − 80 ℃ in the laboratory for further experiments.
DNA extraction
All samples were collected from feces in the large intestine of badgers under aseptic conditions. The QIAamp® Stool Mini Kit (QIAGEN, Germany) was used to extract total genomic DNA following the manufacturer’s protocols. Subsequently, the extracted DNA was analyzed using 1% agarose gel electrophoresis.
16S rRNA gene PCR and sequencing
The V3-V4 region of the bacterial 16S ribosomal RNA gene was amplified by PCR using the following protocol: initial denaturation at 95 ℃ for 3 min; followed by 25 cycles at 95 ℃ for 30 s, 55 ℃ for 30 s, and 72 ℃ for 30 s; and a final extension at 72 ℃ for 5 min The primers used were338F (5′-barcode-ACTCCTACGGGAGGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′). The PCR reaction was performed in a total volume of 50 μL containing the following: 6 μL of the template DNA, 25 μL of 2 × Taq PCR Master Mix, 2 μL of each primer (10 mM) and 15 μL of ddH2O to complement the reaction system. After detection by 2% agarose gel electrophoresis and recovery using the AxyPrep DNA Gel Extraction Kit (AXYGEN, USA), the PCR products were quantified using the Quantified Fluor™ ST Blue Fluorescence Quantification system (Promega, USA).
Sequencing libraries were generated using the TruSeq™ DNA Sample Prep Kit (Illumina, USA) and sequencing was conducted on an Illumina MiSeq platform following the manufacturer’s recommendations.
Sequence processing and data analysis
Raw fastq files were demultiplexed, quality-filtered and assembled using QIIME (version 1.9.1)26 and FLASH (version 1.2.11). Chimeric sequences were identified and removed using UCHIME27. Subsequently, sequences were clustered into the same operational taxonomic units (OTUs) with a 97% sequence identity cutoff value using UPARSE (version 7.0.1 http://drive5.com/uparse/). Taxonomy and annotation of the obtained sequences were performed by RDP Classifier (version 2.11) against the SILVA (SSU132) 16S rRNA database28.
The sequencing depth was 28,763 for rarefaction. Alpha diversity index values, including the Sobs (observed species richness), Shannon, Simpson, Chao 1, ACE and Good’s coverage indices, were generated and analyzed by Mothur (version 1.30.2), along with rarefaction curves. Rank-abundance curves were calculated and displayed using R software (version 3.3.1). Stacked histogram of relative abundance and the heatmap showing clustering of species abundance were created using R software (version 3.3.1). Hierarchical clustering trees were generated and displayed using QIIME (version 1.9.1) and R software (version 3.3.1) based on the unweighted pair-group method with arithmetic mean (UPGMA). Wilcoxon rank-sum test for alpha diversity indices, principal component analysis (PCA), principal coordinate analysis (PCoA) and nonmetric multidimensional scaling (NMDS) were also performed and demonstrated using R software (version 3.3.1). We used the Euclidean distance matrix for PCA and weighted unifrac distance matrix for both PCoA and NMDS. Permutational multivariate analysis of variance (PERMANOVA) was conducted in QIIME (version 1.9.1) to test the grouping of the samples. To determine the species with significant differences, we first performed a log10 transformation on the abundance data before conducting the relevant tests, then the Kruskal–Wallis H test was performed using R software (version 3.3.1). Finally, linear discriminant analysis effect size (LEfSe) was performed using LEfSe software, with a filter value of the LDA score was set as 4 by default29.
The data were analyzed using the free online platform of Majorbio Cloud Platform (www.majorbio.com). The dataset from our study is available in the Sequence Read Archive (SRA) of NCBI under the accession number PRJNA751208.
All methods were carried out in accordance with relevant guidelines and regulations. All methods are reported in accordance with ARRIVE guidelines. This study was performed in accordance with the permission from the Ethics and Animal Welfare Committee of Beijing Normal University (approval reference number: CLS-EAW-2019-029).
Results
Overview of the sequencing data
A total of 1,072,316 reads were obtained after quality control and filtration from 25 of large intestine contents, comprising 10 from wild and 15 from captive badgers. Subsequently, the high-quality reads were classified into 1,759 OTUs with a 0.97 identity cutoff.
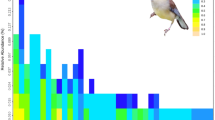
The alpha diversity indices (Sobs, Shannon, Simpson, ACE, Chao 1 and Good’s coverage) are presented in Table 1. The Shannon and Sobs indices of each sample are displayed in Figure S2. Rarefaction curves (Figure S3a) are depicted to assess whether the quantity of OTUs used in the study was sufficient. To analyze the richness and evenness of species in the contents of the large intestines of badgers, rank-abundance curves are shown in Figure S3b.
Bacterial composition and relative abundance
Overall, 33 phyla, 85 classes, 192 orders, 321 families and 572 genera were detected in the microbiota of the large intestine contents from 25 badgers.
At the phylum level (Fig. 1a), for the captive badgers in the summer (CS group), Firmicutes (58.663% ± 27.453%), Proteobacteria (40.367% ± 27.569%) and Actinobacteria (0.671% ± 0.491%) were the predominant phyla, followed by Bacteroidetes (0.160% ± 0.287%) and Fusobacteria (0.045% ± 0.102%). For the captive badgers in the winter (CW group), the top three predominant phyla were also Firmicutes (82.881% ± 9.204%), Proteobacteria (16.435% ± 8.656%) and Actinobacteria (0.279% ± 0.309%). Fusobacteria (0.247% ± 0.567%) and Camplyobacteria (0.079% ± 0.213%) were ranked fourth and fifth, respectively.
For the wild badgers, in the summer group (WS group), the top five phyla were Firmicutes (60.013% ± 41.412%), Proteobacteria (18.128% ± 14.961%), Actinobacteria (13.072% ± 16.595%), Verrucomicrobia (3.064% ± 4.141%) and Chloroflexi (1.638% ± 2.239%). For the wild badgers in the winter (WW group), the predominant phyla were Firmicutes (71.122% ± 26.140%), Proteobacteria (16.003% ± 12.584%), Bacteroidetes (7.372% ± 14.734%), Actinobacteria (3.664% ± 4.281%) and Deltaproteobacteria (0.929% ± 1.853%).
At the genus level (Fig. 1b), Escherichia-Shigella (39.007% ± 26.443%), Streptococcus (23.429% ± 27.924%) and Enterococcus (12.703% ± 11.590%) were the predominant genera in the CS group. For the CW group, Streptococcus (26.567% ± 14.787%) was the predominant genus, followed by Clostridium_sensu_stricito_1 (17.256% ± 7.117%) and Escherichia-Shigella (15.985% ± 8.812%).Interestingly, the most predominant genus in the wild badgers was Romboutsia (44.196% ± 35.401% WS and 46.256% ± 19.527%WW). The genera Terrisporobacter (5.761% ± 13.714%) and Clostridium_sensu_stricto_1 (4.682% ± 7.382%) were ranked second and third in the WS group, while Escherichia-Shigella (10.674% ± 14.815%) and Terrisporobacter (9.707% ± 10.975%) were ranked second and third in the WW group.
To analyze the similarities and differences in the microbiota community composition of the badger large intestine content, a cluster heatmap for species abundance at the phylum level is shown in Fig. 2. There were obvious differences in the community structure between the captive and wild groups, especially at the genus level. Hierarchical clustering trees using UPGMA at the phylum and genus levels are shown in Fig. 3a,b respectively, which indicated similar results to Fig. 2. Samples CS2 and CS6 were clustered more closely with the WS group than other samples in the CS group.
The heatmap of clustering for species abundance. The color gradient of the color block represents the variation of the abundance of different species in the sample. The value represented by the color gradient is on the right of the figure. The software and algorithm used to generate this chart: R language (version 3.3.1) vegan package (https://cran-archive.r-project.org/bin/windows/base/old/3.3.1/).
Analysis of differences in community composition
The Wilcoxon rank-sum test for alpha diversity indices (ACE, Chao 1, Shannon and Simpson indices at the OTU level) among the four groups is shown in Fig. 4. The p values of the test for ACE and Chao 1 indices between the CS and WS groups were both 0.005. Significant differences in the Shannon and Simpson indices were also detected between the CS and CW groups, respectively (Wilcoxon rank-sum test: Shannon diversity, CS vs. CW, W = 8, p = 0.024; Simpson diversity, CS vs. CW, W = 47, p = 0.032). However, there were no significant differences among the other combinations (the CW and WW group, the WS and WW group).
To demonstrate the discrepancies among the four groups more intuitively, the principal component analysis (PCA) plot, the principal coordinates analysis (PCoA) plot and the nonmetric multidimensional scaling (NMDS) plot are depicted in Fig. 5. All samples within each group were clustered in the three plots, indicating that the bacterial composition and structure of the gut microbiota were similar. Additionally, the captive (CS and CW) and wild (WS and WW) groups showed distinct clustering patterns, further supporting the differences between captive and wild badgers. We also performed PERMANOVA to conduct inter-group similarity analysis on the grouped samples and test the significance of inter-group differences. The PERMANOVA results of wild/captive (R = 0.306, p = 0.001) and summer/winter (R = 0.093, p = 0.035) indicated that the groupings were statistically significant and in line with expectations, consistent with the findings presented in Fig. 4.
PCA (a), PCoA (b) and NMDS (c) of intestianl bacterial community structures of the wild and captive Asian badgers. The different shape with colors represented all samples of badgers respectively. (b) and (c) were generated with weighted unifrac distance. The test results in figure are all from the ANOSIM test.
Then the Kruskal–Wallis H test (Fig. 6) and linear discriminant analysis effect size (LEfSe) (Fig. 7) were used to calculate and detect species with significant differences at different taxonomic levels. At the phylum level (Fig. 6a), significant differences were observed in the relative abundances of Verrucomicrobia (3.064% ± 4.141%, H = 16.745, p < 0.001), Planctomycetes (0.880% ± 1.173%, H = 13.975, p = 0.003), Patescibacteria (0.685% ± 0.855%, H = 10.465, p = 0.015), Cyanobacteria (0.158% ± 0.161%, H = 12.097, p = 0.007) and Gemmatimonadetes (0.162% ± 0.230%, H = 10.471, p = 0.015) in the WS group. Only the phylum Fusobacteria (0.247% ± 0.567%, H = 14.524, p = 0.002) showed a significant difference in the CW group. Other species that exhibited significant differences at the genus level are shown in Fig. 6b. Different color nodes in the cladogram in Fig. 7a represent microbial groups that were significantly enriched in the corresponding groups and had a notable influence on the differences between groups. The LEfSe bar plot, depicted in Fig. 7b, shows the taxa (CS: 10, CW: 10, WS: 17 and WW: 4) with significant differences among the four groups.
The results of LEfSe analysis. The different color nodes in cladogram (a) represent the microbial groups that are significantly enriched in the corresponding groups and have significant influence on the differences between groups. The LEfSe bar chart (b) counts the microbial groups with significant effects in multiple groups. The greater the LDA score, the greater the impact of species abundance on the difference effect.
Discussion
Over the past decade, the role of the gut microbiota in animal health has been extensively studied, encompassing developmental, immunological and physiological functions30,31,32. Diet plays a crucial role in modulating the composition of the gut microbiota33,34. Additionally, for many wild animals, seasonal dietary shifts influence changes in gut microbiota composition due to fluctuations in food availability and energy intake28,34,35. However, studies on the gut microbiota of Asian badgers are currently lacking.
In this study, we first characterized the gut microbiota of both captive and wild Asian badgers and compared it across different seasons. Our findings align with previous studies on Mustelidae species like the sable (Martes zibellina), mink (Mustela vison), and North American river otter (Lontra canadensis)36,37,38,39, showing Firmicutes, Proteobacteria, Actinobacteria, and Bacteroidetes as the predominant phyla, which are commonly found in mammals.
At the phylum level, regardless of the season, Firmicutes and Proteobacteria were consistently the most abundant phyla in both captive and wild Asian badgers. Notably, these two phyla accounted for over 98% of the microbiota in captive Asian badgers, whereas their proportions were lower than 90% in wild individuals. The varying relative abundance of the third and subsequent dominant phyla indicated differences in bacterial community richness between captive and wild Asian badgers. The composition of the bacterial microbiota is known to be closely related to long-term dietary habits40. The feeding strategy of the Asian badger largely depends on food availability and environmental factors.
The relative abundances of Bacteroidetes, which play an important role in the degradation of high molecular weight substances and carbohydrates secreted by the intestines41,42, present a remarkable proportion in the WW group, while in the other groups they were almost undetectable. Moreover, it was also shown that an increase of Bacteroidetes was detected when the weight of obese mice decreased43. The Firmicutes/Bacteroidetes ratio is closely related to the dietary habits and physiological function of the host44,45. First, in the wild, the food resources and availability in winter were limited compared to those in summer. However, the farmed badgers were fed mainly corn, chicken and modified feed for foxes, which ensured a sufficient food supply in winter. Second, studies have shown that hibernation is an adaptive mechanism for badgers to maintain their energy balance when winter climatic conditions lead to food shortages and increased heat loss14. Although the lower body temperature could save considerable energy and reduce fat storage requirements during hibernation46, weight loss of badgers would occur after long and cold winters at high latitudes. Nevertheless, due to the difficulties of sampling, the body weight data of the Asian badgers in this study were insufficient.
Furthermore, the relative abundance of Verrucomicrobia in the WS group was also significantly higher than that in other groups. Due to the difficulties in isolation and the deficiency of genomic data, the ecological and metabolic roles of Verrucomicrobia are poorly understood. Several studies have suggested that Verrucomicrobia is a potential target for inducing regulatory immunity47, and some relatively rare taxa of this phylum may be highly effective in polysaccharide degradation48. The higher abundance of Verrucomicrobia in the WS group may indicate that more polysaccharides in food need to be degraded. However, at present, due to the lack of fresh samples and the strict protection of wild animals, we know very little about the actual health of wild badgers. Therefore, the potential role of Verrucomicrobia should be studied further with more wild samples.
At the genus level, Romboutsia was the only genus that was significantly higher in the WW group. This genus is associated with the host’s healthy status and is also regarded as a biomarker of intestinal dysbiosis49. In the present study, the relative abundance of Romboutsia was low in captive Asian badgers. It is well known that the primary objective of breeding farms is to profit maximization. The Asian badger is a valuable economic animal, prized for its skin, fur and meat, especially in traditional Chinese medicine. Nevertheless, scientific breeding strategies and effective monitoring methods for badger health are still lacking. This result indicates that existing breeding strategies may not be entirely suitable for captive Asian badgers.
Another genus deserving attention and showing a significantly higher abundance in the CW group was Streptococcus. Group B Streptococcus (GBS) has been identified as the primary cause of mastitis in dairy herbs, directly impacting milk productivity50,51. Other species within this genus, such as Streptococcus iniae, are commonly found infecting finfish, leading to Streptococcosis52. The transmission routes of Streptococcus agalactiae from bovines vary and include milk, water contaminated by milk, dairy workers and consumers of dairy products. Therefore, given the hard lessons learned from the spread of COVID-19 worldwide, the prevention and control of zoonotic diseases should be further reinforced, regardless of whether captive or wild animals are involved.
In addition, Enterococcus was identified at higher level in captive Asian badgers than in the wild groups. Enterococcus typically colonizes various environments and animal intestines and is implicated in many diseases, such as urinary tract infections, hepatobiliary sepsis and surgical wound infection53. Some studies have indicated that Enterococcus is common nosocomial pathogens and have been detected in the fecal microbiota of birds that underwent surgery54,55. Although there were no abnormal conditions in the badger farm, the health conditions of badgers should be evaluated carefully based on the most recent developments in epidemiology and zoology.
In summary, the predominant phyla of the gut microbiota related to the digestion of fat and carbohydrates, as well as the degradation of monosaccharides and polysaccharides, indicated that the diversity and availability of food resources of wild Asian badgers in summer were higher than those for captive badgers. Our results also revealed that wild badgers in summer had a higher richness of the bacterial community, consistent with reports that wild animals harbor a more abundant gut microbiota. Thus, we inferred that dietary diversity is the most important factor influencing the composition and structure of the gut microbiota in Asian badgers. Next, the health of captive badgers must be thoroughly examined using more advanced methods, such as metagenomics and proteomics analyses. Moreover, further analysis of the gut microbiota from more wild samples could help improve the optimal dietary conditions for captive Asian badgers in farming systems.
We are aware of and acknowledge that other factors may have influenced the results of this study. Firstly, we cannot ascertain the health status of wild badgers prior to their demise. Therefore, we cannot rule out the possibility that wild badgers may have died due to illness. Additionally, recent research has shown variation in the gut microbiota of other mustelids according to breeding and non-breeding seasons56. Secondly, we have not yet considered the potential effects of age or sex on the gut microbiota, both of which could contribute to variation in microbial composition. Additionally, due to the differences in sampling areas, there is significant biogeographical variation between the captive and wild populations. Therefore, in further research and analysis, we will continue to take these factors into account to ensure the accuracy and reliability of our study results.
While the results of our study offer primary data on the gut microbiota of Asian badgers, the scarcity and value of wild samples limited the information we could gather. To enhance our understanding, it is essential to collect a wider variety of samples, including fresh feces and urine, along with detailed information on individual animals, in a more scientific and systematic manner in the future.
Conclusions
In general, we have reported for the first time the bacterial composition and structure of the gut microbiota from large intestine samples of wild Asian badgers using high-throughput sequencing of the V3-V4 region of the 16S rRNA gene. We observed that Firmicutes and Proteobacteria were the most predominant phyla in the gut microbiota of both wild and captive Asian badgers, regardless of the season. We speculated that the diversity and availability of food resources are the most important factors influencing the gut microbiota of Asian badgers. Several genera may serve as potential sources of zoonoses, warranting further attention and study. Certainly. This study could significantly contribute to conservation and management efforts in several ways. Firstly, by understanding the gut microbiota of Asian badgers, we can gain insights into their overall health status and ecosystem interactions, which are crucial for effective conservation measures. Secondly, identifying any patterns or changes in the microbiota could help in early detection and prevention of disease outbreaks among badger populations, thus aiding in their conservation. Additionally, this research can inform wildlife management strategies by providing valuable information on the factors influencing badger populations and their habitats. Overall, a better understanding of the gut microbiota of Asian badgers can lead to more informed conservation and management decisions aimed at protecting this species and their ecosystems.
Data availability
The dataset from our study is available in the Sequence Read Archive (SRA) of NCBI under the accession number PRJNA751208.
References
Abramov, A. V. & Puzachenko, A. Y. Sexual dimorphism of craniological characters in Eurasian badgers, Meles spp. (Carnivora, Mustelidae). Zool. Anz. A J. Comp. Zool. 244, 11–29 (2005).
Baryshnikov, G. & Potapova, O. Variability of the dental system in badgers (Meles, Carnivora) of the USSR fauna. Zool. Ž. 69, 84–97 (1990).
Lynch, J. M. Morphometric variation in the badger (Meles meles): Clinal variation in cranial size and shape across Eurasia. Small Carniv. Conserv. 10, 6–7 (1994).
Nowak, R. M. & Walker, E. P. Walker’s Mammals of the World (JHU Press, 1999).
Abramov, A. Notes on the taxonomy of the Siberian badgers (Mustelidae: Meles). Trudy Zool. Inst. Ross Akad Nauk 288, 221–233 (2001).
Del Cerro, I. et al. Nuclear and mitochondrial phylogenies provide evidence for four species of Eurasian badgers (Carnivora). Zool. Scr. 39, 415–425 (2010).
Abramov, A.V. Meles leucurus. The IUCN Red List of Threatened Species. 2016, e.T136385A45221149 (2016). https://doi.org/10.2305/IUCN.UK.2016-1.RLTS.T136385A45221149.en. Accessed on 05 August 2024.
Clark, E.; Munkhbat, J.; Dulamtseren, S.; Baillie, J.; Batsaikhan, N.; King, S.; Samiya, R.; Stubbe, M. Summary Conservation Action Plans for Mongolian Mammals. Regional Red List Series Vol. 2. Zoological Society of London, London. English and Mongolian 2006, 165.
Lau, M. W. N., Fellowes, J. R. & Chan, B. P. L. Carnivores (Mammalia: Carnivora) in South China: A status review with notes on the commercial trade. Mamm. Rev. 40, 247–292 (2010).
Cui, H. T. et al. Traditional Chinese medicine for treatment of coronavirus disease 2019: A review. Tradit. Med. Res. 5, 65 (2020).
Kruuk, H. & Parish, T. Feeding specialization of the European badger Meles meles in Scotland. J. Animal Ecol. 50, 773–788 (1981).
Zagainova, O. & Markov, N. The diet of Asian badger, Meles leucurus Hodgson, 1847, in Samarovskii Chugas Nature Park Western Siberia. Russ. J. Ecol. 42, 414–420 (2011).
Kowalczyk, R., Jȩdrzejewska, B. & Zalewski, A. Annual and circadian activity patterns of badgers (Meles meles) in Białowieża Primeval Forest (eastern Poland) compared with other Palaearctic populations. J. Biogeogr. 30, 463–472 (2003).
Fowler, P. & Racey, P. Overwintering strategies of the badger, Meles meles, at 57 N. J. Zool. 214, 635–651 (1988).
Roper, T., Ostler, J., Schmid, T. & Christian, S. Sett use in European badgers Meles meles. Behaviour 138, 173–187 (2001).
Wilson, G., Harris, S. & McLaren, G. Changes in the British Badger Population, 1988 to 1997 (People’s Trust for Endangered Species London, 1997).
Wang, J. et al. Away from the city: Habitat selection of badgers in mountainous area around Beijing. Biologia 76, 1737–1746 (2021).
Li, F. & Jiang, Z. Is nocturnal rhythm of Asian badger (Meles leucurus) caused by human activity? A case study in the eastern area of Qinghai Lake. Biodivers. Sci. 22, 758 (2014).
Zhou, Y., Meng, Y., Ren, Y., Liu, Z. & Li, Z. A retrospective survey of the abortion outbreak event caused by brucellosis at a blue fox breeding farm in Heilongjiang province. Front. Vet. Sci. https://doi.org/10.3389/fvets.2021.666254 (2021).
Hwang, J. Y., Lee, S.-G. & Kim, E. B. Polymorphic sites in complete genome sequences of Asian badger, Meles leucurus amurensis (Mustelidae, Melinae) mitochondria. Mitochondrial DNA Part B 1, 264–265 (2016).
Kinoshita, E. et al. Hybridization between the European and Asian badgers (Meles, Carnivora) in the Volga-Kama region, revealed by analyses of maternally, paternally and biparentally inherited genes. Mamm. Biol. 94, 140–148 (2019).
Chow, J., Lee, S. M., Shen, Y., Khosravi, A. & Mazmanian, S. K. Host–bacterial symbiosis in health and disease. Adv. Immunol. 107, 243–274 (2010).
Hanning, I. & Diaz-Sanchez, S. The functionality of the gastrointestinal microbiome in non-human animals. Microbiome 3, 1–11 (2015).
Guarner, F. & Malagelada, J.-R. Gut flora in health and disease. Lancet 361, 512–519 (2003).
Nicholson, J. K., Holmes, E. & Wilson, I. D. Gut microorganisms, mammalian metabolism and personalized health care. Nat. Rev. Microbiol. 3, 431–438 (2005).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Amato, K. R. et al. Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J. 7, 1344–1353 (2013).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, 1–18 (2011).
Purchiaroni, F. et al. The role of intestinal microbiota and the immune system. Eur. Rev. Med. Pharmacol. Sci. 17, 323–333 (2013).
Sekirov, I., Russell, S. L., Antunes, L. C. M. & Finlay, B. B. Gut microbiota in health and disease. Physiol. Rev. https://doi.org/10.1152/physrev.00045.2009 (2010).
Rajilić-Stojanović, M. Function of the microbiota. Best Pract. Res. Clin. Gastroenterol. 27, 5–16 (2013).
Brown, K., DeCoffe, D., Molcan, E. & Gibson, D. L. Diet-induced dysbiosis of the intestinal microbiota and the effects on immunity and disease. Nutrients 4, 1095–1119 (2012).
Ren, T., Grieneisen, L. E., Alberts, S. C., Archie, E. A. & Wu, M. Development, diet and dynamism: Longitudinal and cross-sectional predictors of gut microbial communities in wild baboons. Environ. Microbiol. 18, 1312–1325 (2016).
Bergmann, G. T., Craine, J. M., Robeson, M. S. & Fierer, N. Seasonal shifts in diet and gut microbiota of the American bison (Bison bison). PloS ONE 10, e0142409 (2015).
Su, L. et al. Habitat elevation shapes microbial community composition and alter the metabolic functions in wild sable (Martes zibellina) guts. Animals 11, 865 (2021).
Guan, Y. et al. Comparison of the bacterial communities in feces from wild versus housed sables (Martes zibellina) by high-throughput sequence analysis of the bacterial 16S rRNA gene. Amb. Express 6, 1–11 (2016).
Zhao, H. et al. Mink (Mustela vison) gut microbial communities from Northeast China and its internal relationship with gender and food additives. Curr. Microbiol. 74, 1169–1177 (2017).
Guo, G. et al. The gut microbial community structure of the north american river otter (Lontra canadensis) in the alberta oil sands region in Canada: Relationship with local environmental variables and metal body burden. Environ. Toxicol. Chem. 39, 2516–2526 (2020).
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334, 105–108 (2011).
Salyers, A., Vercellotti, J., West, S. & Wilkins, T. Fermentation of mucin and plant polysaccharides by strains of Bacteroides from the human colon. Appl. Environ. Microbiol. 33, 319–322 (1977).
Thoetkiattikul, H. et al. Comparative analysis of microbial profiles in cow rumen fed with different dietary fiber by tagged 16S rRNA gene pyrosequencing. Curr. Microbiol. 67, 130–137 (2013).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Mariat, D. et al. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol. 9, 1–6 (2009).
Tanaka, H. Winter hibernation and body temperature fluctuation in the Japanese badger Meles meles anakuma. Zool. Sci. 23, 991–997 (2006).
Lindenberg, F. et al. Expression of immune regulatory genes correlate with the abundance of specific Clostridiales and Verrucomicrobia species in the equine ileum and cecum. Sci. Rep. 9, 1–10 (2019).
Martinez-Garcia, M. et al. Capturing single cell genomes of active polysaccharide degraders: An unexpected contribution of Verrucomicrobia. PloS ONE 7, e35314 (2012).
Mangifesta, M. et al. Mucosal microbiota of intestinal polyps reveals putative biomarkers of colorectal cancer. Sci. Rep. 8, 1–9 (2018).
McCracken, G. H. Jr. Group B streptococci: The new challenge in neonatal infections. J. Pediatr. 82, 703–706 (1973).
Botelho, A. C., Ferreira, A. F., Fracalanzza, S. E., Teixeira, L. M. & Pinto, T. C. A perspective on the potential zoonotic role of Streptococcus agalactiae: Searching for a missing link in alternative transmission routes. Front. Microbiol. 9, 608 (2018).
Beck, B. H., Yildirim-Aksoy, M., Shoemaker, C. A., Fuller, S. A. & Peatman, E. Antimicrobial activity of the biopolymer chitosan against Streptococcus iniae. J. Fish Dis. 42, 371–377 (2019).
Fisher, K. & Phillips, C. The ecology, epidemiology and virulence of Enterococcus. Microbiology 155, 1749–1757 (2009).
Lopes, Md. F. S. et al. Antimicrobial resistance profiles of dairy and clinical isolates and type strains of enterococci. Int. J. Food Microbiol. 103, 191–198 (2005).
Guan, Y., Wang, H., Gong, Y., Ge, J. & Bao, L. The gut microbiota in the common kestrel (Falco tinnunculus): A report from the Beijing Raptor Rescue Center. PeerJ 8, e9970 (2020).
DeCandia, A. L. et al. Gut bacterial composition shows sex-specific shifts during breeding season in ex situ managed black-footed ferrets. J. Hered. 115, 065 (2023).
Acknowledgements
We sincerely thank all members of the Forest Public Security Bureau of Hunchun, Yanbian Korean Autonomous Prefecture, Jilin Province, Northeast Tiger Leopard National Park Authority, Hunchun Forestry Co. Ltd of Changbai Mountain Forest Industry Group Co. Ltd and Hunchun forestry bureau.
Funding
National Natural Science Foundation of China,32001093,31700469,31670537,National Scientific and Technical Foundation Project of China,2019FY101700
Author information
Authors and Affiliations
Contributions
Conceptualization, G.Y. and Y.H.; methodology, P.J. and G.Y.; software, P.J. and G.Y.; validation, P.J. and G.Y. and Y.H.; formal analysis, P.J. and G.Y.; investigation, P.J., G.Y. and Y.H.; resources, G.Y., X.W., Y.H., G.J. and F.L.; data curation, P.J. and G.Y.; writing—original draft preparation, P.J. and G.Y.; writing—review and editing, G.Y., X.W., Y.H., F.L.; visualization, P.J. and G.Y.; supervision, G.J., Y.H. and F.L.; project administration, Y.H. and F.L.; funding acquisition, X.W., G.J. and F.L. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pei, J., Guan, Y., Xiao, W. et al. The comparison of gut microbiota between wild and captive Asian badgers (Meles leucurus) under different seasons. Sci Rep 14, 18199 (2024). https://doi.org/10.1038/s41598-024-69277-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69277-8
- Springer Nature Limited