Abstract
Numerous studies have shown that the Krüppel-like factors (KLFs) family of transcription factors regulate various eukaryotic physiological processes including the proliferation, differentiation, senescence, death, and carcinogenesis of animal cells. In addition, they are involved in the regulation of key biological processes such as cell cycle, DNA repair, and immune response. Current studies focus on investigating the role of KLFs in normal physiological conditions and the incidence and development of diseases. However002C the significance of KLFs family genes in clear cell renal cell carcinoma (ccRCC) remains partly understood; therefore, an in-depth investigation of their role and clinical value in this cancer is desired. The study aimed to investigate the role of KLF family genes in the incidence, development, and prognosis of ccRCC, and to identify the related potential biomarkers and therapeutic targets. The expression of KLFs in the RNA sequencing data of 613 ccRCCs from the TCGA database was analyzed using R software, and UALCAN and GEPIA assessed the expression of KLF genes in ccRCC. Real-time fluorescence quantitative PCR analysis was performed using 10 pairs of paired ccRCC sample tissues and renal cancer cell lines from the First Affiliated Hospital of Nanchang University. Overall survival (OS), progression-free interval (PFI), and disease-specific survival (DSS) of Kidney Clear Cell Carcinoma (KIRC) samples at differential expressions of KLFs in the TCGA database were analyzed using the R software, followed by generating a nomogram prediction model. GSCALite assessed the interactions of KLF genes with miRNAs and generated network maps. Protein interaction network maps of 50 neighboring genes associated with KLF mutations were analyzed using STRING with GO and KEGG functional enrichment analyses. The cBioPortal determined the probability of KLF gene mutations and their impact on OS and disease-free survival (DFS) in patients with ccRCC. Immune cell infiltration of KLFs was analyzed using TIMER. Finally, GSCALite was used to analyze the drug sensitivity and associated pathways of action of KLFs. Correlation validation using cellular experiments. KLF3/5/9/15 were significantly downregulated in ccRCC tissues, whereas KLF16/17 were upregulated compared with the adjacent tissues. Patients with high mRNA levels of KLF16/17 showed significantly lower OS, PFI, and DSS, whereas KLF3/5/9 showed a reverse trend. In patients with ccRCC, a significant correlation was observed between KLF mutations and OS and DSS. Furthermore, the correlation of KLF3/5/9 with immune cell infiltration was stronger than that of KLF15/16, while KLF17 was significantly associated with the Epithelial-Mesenchymal Transition (EMT) pathway. Overexpression of KLF5 inhibits the proliferative and migratory capacity of renal cancer cells (786-O and OS-RC-2), as well as their sensitivity to relevant small molecule drugs. Our research revealed the expression levels and biological significance of KLF genes in ccRCC, particularly highlighting the potential of KLF5 as a promising biomarker and therapeutic target for effective prognosis and diagnosis of ccRCC.
Similar content being viewed by others
Introduction
The underlying mechanisms for the development of clear cell renal cell carcinoma (ccRCC), particularly the incidence and progression, remain poorly understood. ccRCC is one of the prevalent causes of cancer-related deaths globally1. According to global cancer research, a total of 371,700 cases of kidney cancer were reported worldwide in the year 2019, with an age-standardized incidence rate (ASIR) of 4.6 per 100,0002. Moreover, the global incidence of kidney cancer increased by 154.78% compared with that in the year 1990. During these three decades, the age-standardized mortality rate (ASMR) was persistently high3, with ccRCC being the most prevalent form4. Although a majority of the patients with ccRCC undergo surgery, local spread or distant metastasis occurs in approximately 30%, thereby reducing the overall 5-year survival rate to approximately 58%5. Early diagnosis and prompt treatment are effective in treating ccRCC. Therefore, the identification of more precise biomarkers and therapeutic targets for ccRCC is urgently required.
Recently, the role of Krüppel-like factors (KLFs) family genes in ccRCC has been widely investigated. KLFs are a class of zinc finger-containing transcription factors that are crucial in regulating various biological processes, including cell proliferation, differentiation, and survival6. KLFs promote or inhibit the expression of target genes by binding to specific DNA sequences. Particularly in ccRCC, the expression of several KLF family genes is associated with the occurrence, development, invasion, and prognosis of the tumor7. For example, KLF6, an important member of the KLF family, has a potential oncogenic role in tumorigenesis and progression. In patients with ccRCC, a downregulated expression of KLF6 was associated with poor prognosis8. Similarly, KLF4 is crucial in the normal functioning of renal cells and is associated with the progression of ccRCC; therefore, it may serve as a potential therapeutic target. Nevertheless, in vitro and in vivo studies have reported that overexpression of KLF4 inhibits the proliferation of renal cancer cells and induces apoptosis, highlighting its potential application in anti-tumor therapy9.
Recent research has also revealed the role of KLFs in regulating the immune response and inflammatory reactions, particularly those crucial in the microenvironmental regulation of ccRCC. Reportedly, KLF2 and KLF4 influence the tumor microenvironment and associated immune cells10. In addition, the role of KLFs in metabolic alterations of renal cancer cells has presented potential signs for developing novel metabolic intervention strategies11. Owing to their multifaceted roles in the pathophysiology of renal cell carcinoma, KLFs could serve as valuable targets in future therapeutic strategies.
Materials and methods
Patients and tumor samples
We analyzed the mRNA expression of KLFs using R software (“stats” and “ggplot2” packages) on 613 RNAseq results obtained from the public TCGA database (https://portal.gdc.cancer.gov), including 541 ccRCC samplesand 72 adjacent tissue samples followed by data visualization. Simultaneously, 10 pairs of ccRCC and adjacent tissue samples were collected at the Department of Urology, the First Affiliated Hospital of Nanchang University between February 2021 and December 2022. Ethics approval was obtained from the Ethics Committee of the First Affiliated Hospital of Nanchang University, reference number: (2022) CDYFYYLK (10–011). All methods were performed in accordance with the relevant guidelines and regulations, and written informed consent was obtained from all subjects. These tissue specimens were confirmed as ccRCC by 2–3 pathologists and stored in liquid nitrogen for further use. Total RNA extraction from the tissues utilized the “Trizol” method, followed by reverse transcription using the EasyScript® All-in-One First-Strand cDNA Synthesis SuperMix for qPCR kit. Real-time fluorescence quantitative PCR (qRT-PCR) utilized the PerfectStart® Green qPCR SuperMix (+ Dye I) kit. The mRNA expression of KLF genes in ccRCC and adjacent tissues was performed thrice for validation.
Cell lines and cell culture
The six cell lines used in the present study (HK-2, A-498, ACHN, OS-RC-2, 786-O, and Caki-1) were obtained from ATCC. Following the RNA extraction and reverse transcription, qRT-PCR was performed, as mentioned above. Subsequently, the mRNA expression levels of KLFs were validated in both HK-2 (human renal cortical proximal tubular epithelial cells) and human renal cancer cell lines.
RNA and real-time fluorescent quantitative PCR
The total RNA from tissues and cell lines was dissolved in RNase-free water, and the concentration was adjusted to approximately 500 ng/μl. The primer sequences utilized for KLFs (Table 1) were provided by ShangHai Sangon Biotech, with β-actin as an internal reference. The average amplification CT values and corresponding dissolution curves of three replicates were obtained to calculate the expression of respective members of KLFs.
Introduction and application of relevant online databases
GEPIA
The GEPIA database (http://gepia.cancer-pku.cn/) was employed to analyze the RNA sequencing data from TCGA and GTEx databases. The database compiles data from 9,736 tumor tissue and 8,587 normal tissue samples, and the project is technically supported by the Advanced Innovation Center for Genomics at Peking University. We utilized this database for the comparative analyses of the correlation of KLFs at various stages of ccRCC.
UALCAN
UALCAN (http://ualcan.path.uab.edu/index.html) includes post-methylation and differential expression data of KLFs in ccRCC and adjacent tissues. This database analyzes data from TCGA, MET500, CPTAC, and CBTTC databases, and provides the expression profiles of protein-coding, lincRNA-coding, and miRNA-coding genes with patient survival information. In addition, it presents data analyses from the Clinical Proteomic Tumor Analysis Consortium, including total and phosphorylated protein data. We utilized this database for the analysis of KLFs' methylation levels, status of lymph node metastasis, and expression in ccRCC subtypes and grades.
cBioPortal
cBioPortal (https://www.cbioportal.org/) offers online visualization tools for the analysis of cancer-related genetic data and to identify molecular data based on cancer histological and cytological studies. It compiles data from the TCGA, GDAC, UCSC, and others, to provide information on network connections and interactions between cancer mutations. We employed this database to analyze the probability, patterns, and potential sites of mutations in KLFs. In addition, we examined the effect of such mutations on the overall and disease-specific survival (DSS) before and after the mutation. Subsequently, 50 genes were identified as most relevant to the KLF mutations.
GSCALite
GSCALite is an online analytical tool developed by Professor Guo Anyuan's team at Huazhong University of Science and Technology, which provides data on differential expression, methylation, and survival analysis. We used this tool to identify miRNA molecules associated with KLFs and to create a network diagram. Additionally, we integrated drug sensitivity and gene expression profile data from cancer cell lines in the CTRP database to study the drug sensitivity of KLFs. Spearman’s coefficient was used to correlate the expression of each gene in the gene set with small molecule/drug sensitivity (IC50). KLF17 could not be analyzed due to its unavailability in the CTRP database. Finally, pathway activity modules were used to analyze KLFs-related pathways and generate a visual representation of the global percentage.
STRING
Designated as a core data resource by ELIXIR and the Global Alliance for Genomics and Health, STRING (https://string-db.org/) comprises a comprehensive collection of known and predicted protein/gene interactions, and is capable of generating interaction network diagrams for known genes or proteins. We utilized this database to analyze the 50 genes associated with KLF mutations and to generate an interaction network diagram.
TISCH
Tumor Immune Single-cell Hub (TISCH, http://tisch1.comp-genomics.org) is a scRNA-seq database designed to characterize the tumor microenvironment (TME) at single-cell resolution. TISCH provides detailed cell type annotations at the single-cell level, thus enabling the exploration of TME in different cancer types. TISCH provides detailed cell type annotations at the single-cell level, enabling the exploration of the TME in different cancer types, and its data are mainly derived from the GEO database.
TIMER
TIMER (https://cistrome.shinyapps.io/timer/) is designed to identify the immune cell infiltration in tumor tissues. Based on the RNA-Seq expression profile, it provides comprehensive information on the infiltration of B cells, CD8 + T cells, CD4 + T cells, macrophages, neutrophils, and dendritic cells. We utilized this tool to analyze the relationship between KLs expression levels, overall immune infiltration, and levels of the six immune cell types. Additionally, we examined the levels of infiltration in the somatic cells expressing KLFs.
Statistical analysis
Data analysis utilized SPSS 26.1 and R 4.1.3 software. Continuous variables were represented as mean and standard deviation, whereas percentage (%) was used for categorical variables. Further, Wilcoxon signed-rank and rank-sum tests were utilized for comparing the paired and unpaired samples, respectively.
Ethical approval
Our research protocol was approved by the Ethics Committee of the First Affiliated Hospital of Nanchang University. Ethics number: (2022) CDYFYYLK (10–011). Data were retrieved from online databases, and tissue samples were collected from patients who had given permission for their samples to be used in research and admitted to the Department of Urology in hospital.
Results
mRNA expression levels of KLFs in tumor and adjacent tissues
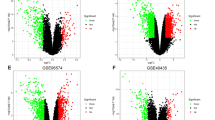
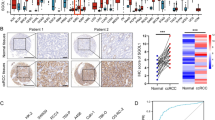
We analyzed the expression of KLFs in TCGA ccRCC and adjacenttissue samples followed by data visualization using R (by “stats” and “ggplot2” packages, respectively; Fig. 1A and B for non-paired and paired samples, respectively). Notably, the expressions of KLF1/6/7/8/14/16 were significantly higher in all the specimens of tumor compared with the adjacent tissue samples (p < 0.001). Conversely, the expressions of KLF5/9/15 were significantly lower in tumor samples relative to the adjacent tissues (p < 0.001). Similarly, for non-paired samples, the expressions of KLF3 and KLF17 were significantly downregulated (p < 0.001) and upregulated (p < 0.05), respectively, in tumors compared with the adjacent tissue samples.
Subsequently, the qPCR results of 10 pairs of renal cancer and adjacent tissues revealed significant differences in the expression of KLFs. While KLF3/5/9/15 were downregulated in the renal cancer tissues (Figs. 1C,E and 2A,C), the expression of KLF16 was upregulated (Fig. 2E) and that of KLF17 remained unaltered (Fig. 2G). Moreover, the expressions of KLF3/5/9/15 were downregulated in the majority of the renal cancer cell lines (Figs. 1D,F and 2B,D), while the expression of KLF16/17 in various cell lines varied non-significantly (Fig. 2F,H).
Clinical relevance of members of KLFs
Furthermore, we analyzed the expression of KLFs in patients with ccRCC based on TCGA data to assess their prognostic value and to generate the Kaplan–Meier survival curves (Fig. 3A–L). Lower mRNA levels of KLF3 (HR = 0.66, 95% CI 0.48–0.89 and p = 0.006), KLF5 (HR = 0.63, 95% CI 0.46–0.85 and p = 0.003), KLF9 (HR = 0.47, 95% CI 0.34–0.64 and p < 0.001), and KLF15 (HR = 0.67, 95% CI 0.49–0.90 and p = 0.009) were associated with poor overall survival (OS) in patients with ccRCC (Fig. 3A,D,G,J). Similarly, lower mRNA levels of KLF3 (HR = 0.50, 95% CI 0.33–0.74 and p < 0.001), KLF5 (HR = 0.65, 95% CI 0.44–0.95 and p = 0.025), KLF9 (HR = 0.38, 95% CI 0.25–0.57 and p < 0.001), and KLF15 (HR = 0.57, 95% CI 0.39–0.84 and p = 0.005) were associated with a shorter DSS (Fig. 3B,E,H,K), whereas lower mRNA levels of KLF3 (HR = 0.59, 95% CI 0.43–0.81 and p = 0.001), KLF5 (HR = 0.57, 95% CI 0.41–0.78 and p < 0.001), and KLF9 (HR = 0.45, 95% CI 0.33–0.63 and p < 0.001) were significantly correlated with a shorter progression-free interval (PFI; Fig. 3C,F,d,I). In contrast, upregulated expressions of KLF16 (HR = 1.52, 95% CI 1.12–2.06 and p = 0.007) and KLF17 (HR = 1.72, 95% CI 1.27–2.33 and p < 0.001) were associated with a poor OS (Fig. 4A,D). Similarly, higher mRNA levels of KLF16 (HR = 1.75, 95% CI 1.19–2.59 and p = 0.005) and KLF17 (HR = 1.82, 95% CI 1.23–2.68 and p = 0.003) were associated with a poor DSS (Fig. 4B,E). Moreover, higher mRNA levels of KLF16 (HR = 1.57, 95% CI 1.14–2.15 and p = 0.006) and KLF17 (HR = 1.39, 95% CI 1.02–1.90 and p = 0.039) were associated with a shorter PFI (Fig. 4C,F). Collectively, these results suggested a significant correlation between the expression of KLF3/5/9/15/16/17 genes and the prognosis of ccRCC, making them potential biomarkers for predicting the survival probability of patients with ccRCC.
Next, we analyzed the correlation between the mRNA levels of KLFs and clinical-pathological features (individual pathological staging). As shown in Fig. 4G–L, expression of KLF16 was not significantly correlated with the pathological stages of ccRCC (p > 0.05). However, the expression of KLF3/5/9/15 decreased with the advancing pathological stages of ccRCC (p < 0.001). Conversely, the expression of KLF17 increased with the higher pathological stage of ccRCC (p < 0.001). These results suggested the potential of KLF3/5/9/15 as tumor-suppressing and KLF17 as tumor-promoting genes in ccRCC.
Subsequently, the methylation levels of KLFs mRNA, lymph node metastasis, and expression in different tumor subtypes and grades were evaluated by using the UALCAN database. As shown in Fig. 5A–F, the methylation levels of KLF3/5/15 were significantly higher in ccRCC than in adjacent tissues (p < 0.001), whereas those of KLF16/17 were significantly lower (p < 0.001). In Fig. 5J–L, compared with the adjacent tissues, expressions of KLF3/5/9/15 were decreased in ccRCC tissues with increased lymph node metastasis. Moreover, in different ccRCC subtypes, KLF3/5/9/15 were significantly downregulated (Fig. 6A–F), while KLF16/17 were significantly upregulated in ccRCC. In addition, the levels of KLF3/9/15/17 varied significantly in different subtypes (p < 0.001). Increased tumor grades were accompanied by decreased levels of KLF3/5/9/15 (Fig. 6G–L) and increased levels of KLF16. These results further suggested the potential roles of KLF3/5/9/15 and KLF16/17 as tumor-suppressing and tumor-promoting genes, respectively, in ccRCC.
Combined analysis of differential expression of the KLF genes and tumor grading predicted the survival time of patients with ccRCC. Results revealed that lower expression of KLF3/5/9/15 and higher expression of KLF16/17 were significantly correlated with a decreased survival period (Fig. 7A–D, P < 0.001).
Development and validation of a prognostic model for column-line diagrams
After selecting independent prognostic factors including age, pathological M stage, and KLF5 through Cox regression analysis, we constructed a forest plot prognostic model (Fig. 8A). The AUC values for the ROC curves at 1, 3, and 5 years were 0.89 (0.95–0.83), 0.82 (0.88–0.75), and 0.82 (0.88–0.75), respectively (Fig. 8B). The calibration curves of the prediction model confirmed a better fit of the curves for 1, 3, and 5 years with the diagonal line (Fig. 8C), indicating a high accuracy of the prediction model. Further validation using an external database (E-MTAB-1980)12 was satisfactory (Fig. 8D–E). The proportional hazards assumption test for the constructed model was performed followed by visualization using R (“ggplot2” package). Results suggested that the model met the proportional hazards assumption (Supplementary Fig. 1A,B).
Neighboring genes associated with KLF mutations
GSCALite generated a network diagram showing the interactions between KLF3/5/9/15 and numerous miRNAs (Fig. 9A). Additionally, the cBioPortal identified 50 adjacent genes associated with KLF mutations and created a corresponding protein interaction network using the STRING tool (Fig. 9B). Subsequently, R software (“ggplot2” package) was employed to perform KEGG and GO functional enrichment analyses for KLFs and the 50 adjacent genes (Fig. 9C). Biological processes (BP) included genes for acyl-CoA biosynthesis (GO:0,071,616), thioester biosynthesis (GO:0,035,384), acetyl-CoA metabolism (GO:0,006,084), and acetyl-CoA biosynthesis (GO:0,006,085). In addition, cellular components (CC) included GO:0,007,044 (cell-substrate junction component), GO:0,048,041 (focal adhesion component), GO:0,035,579 (specific granule membrane), and GO:1,905,286 (serine-type peptidase complex). Moreover, KEGG pathways included hsa05202 (transcriptional misregulation in cancer) and hsa00410 (butanoate metabolism). Notably, these processes are associated with KLF mutations in ccRCC.
Single-cell analysis of the KLFs family in ccRCC
The selected single-cell analysis dataset (KIRC_GSE121636) contained a significant number of CD4Tconv cells (Fig. 10A). While a majority of the immune cells expressed KLF3/9 genes, a relatively higher distribution was observed for CD4 Tconv cells, CD8 T cells, and NK cells (Fig. 10B,C,E). Conversely, cells expressing KLF5/9 genes were less prevalent (Fig. 10D,F).
Association of KLF gene mutations with OS and PFS
We employed cBioPortal to assess the correlation of mutations in KLFs with OS and PFS in patients with ccRCC. Figure 11A shows the genetic alterations in KLFs associated with ccRCC. The KM curves showed that the KLF mutations were significantly related to shorter OS (Fig. 11C, P = 4.415e−3) and DFS (Fig. 11C, P = 3.571e−3) in patients with ccRCC. Consequently, mutations in KLFs may significantly affect the patient survival prognosis of ccRCC.
Association of immune cell infiltration with the KLFs in ccRCC
The TIMER database evaluated the correlation between immune cell infiltration and KLF gene expression. As shown in Fig. 12A–F, infiltration of B cells, CD8 + T cells, CD4 + T cells, macrophages, neutrophils, and dendritic cells positively correlated with the KLF3/5 expression. Similarly, KLF9 expression positively correlated with the infiltration of CD8 + T cells, CD4 + T cells, macrophages, neutrophils, and dendritic cells, and KLF15 expression positively correlated with the infiltration of CD4 + T cells, neutrophils, and dendritic cells. Conversely, expression of KLF16 negatively correlated with the infiltration of B cells, CD4 + T cells, and dendritic cells. Similarly, KLF17 expression negatively correlated with the infiltration of CD4 + T cells, macrophages, neutrophils, and dendritic cells.
Drug sensitivity and related pathways of action in members of KLFs
We employed GSCALite (A total of 481 molecules selected from the CTRP database) to analyze the drug sensitivity of KLFs. Spearman’s coefficient analyzed the correlation between the individual expression of KLF genes and their sensitivity to small molecules. Since the drug sensitivity data for KLF17 was not available in the CTRP database, the respective results were not displayed. The results (Fig. 13A) revealed that the expression of KLF3/5 was positively correlated with the small molecules/drugs, suggesting that a higher expression of KLF3/5 was associated with a greater resistance to these drugs. Conversely, the expression of KLF15/16 was negatively correlated with small molecules/drugs, suggesting that a higher expression of KLF15/16 was associated with increased sensitivity. However, the association of KLF9 with drug resistance was non-significant.
We utilized the pathway activity module in GSCALite to analyze the relevant pathways of KLFs (Fig. 13B). While KLF3 was associated with the Receptor Tyrosine Kinase (RTK) pathway, KLF9 was involved in the DNA Damage Response pathway, KLF15 in the Hormone ER pathway, KLF16 in the RTK pathway, and KLF5 and KLF17 influenced the Epithelial-Mesenchymal Transition (EMT) pathway.
Cellular experiments verified the phenotypic changes of renal cancer cells after overexpression of KLF5
After overexpression of the KLF5 gene in renal cancer cells 786-O and OS-RC-2, both cells had reduced value-adding (Fig. 14A) and migration abilities (Fig. 14B) and were more sensitive to small molecule drug inhibitors, such as our predicted inhibitor of CCT036477 (Fig. 14C).
Discussion
The KLF family comprises 18 members (KLF1–KLF18) and represents a subfamily of mammalian Sp/KLF zinc-finger proteins. They play crucial roles in transcription through the interactions of their highly conserved DNA-binding domains (DBD) or C-terminal regions with the G/C and CACCC boxes of the target genes13. KLFs are involved in tumor cell proliferation, invasion, and metastasis by serving as transcriptional activators or repressors, depending on the type of regulatory proteins they bind14. The KLF family proteins are associated with the formation of fat tissue and muscles, nervous system development, tumor formation, cellular and tissue metabolism, and supporting damage repair at cellular, tissue, and systemic levels15,16. Each KLF gene has a unique structure and is specifically expressed and regulated in different tissues, times, and environments17. While the role of some of the KLF genes has been reported in ccRCC, the impact of expression on ccRCC remains unclear18. The present study investigated the effects of mRNA expression, methylation, protein expression, gene mutation, and immune infiltration of KFLs on ccRCC.
Under normal physiological conditions, KLF3 acts as a transcriptional repressor and is involved in various cellular processes, including adipocyte differentiation19, epidermal differentiation20, erythropoiesis, and B-cell development21. Previous studies suggest that KLF3 is primarily expressed in CD4Tconv, CD8T, monocytes/macrophages, endothelial, and malignant cells in most tumor microenvironments22. KLF3 is aberrantly expressed in various diseases and is involved in related pathways. Therefore, its study would contribute to an in-depth understanding of the pathogenesis of these diseases and will provide new insights and avenues for the treatment of related diseases. In the present study, patients with differential expressions of KLF3 exhibited significantly different survival periods. Specifically, those with a decreased expression showed significantly decreased OS, PFS, and DSS, suggesting its tumor-suppressing role. Additionally, a similar effect of KLF3 was evident in the subgroup analysis of ccRCC. Moreover, Our study suggests KLF3 was associated with a higher immune cell infiltration. Studies have shown that KLF3 expression is closely related to the infiltration of CD4 + T cells, CD8 + T cells, neutrophils, myeloid dendritic cells, mononuclear/macrophages, and endothelial cells in the tumor microenvironment. It is suggested that KLF3 plays an important role in the tumor microenvironment22. Therefore, it can be a potential diagnostic target for ccRCC in future studies.
Studies also suggest the ability of KLF5 to modulate the tumor microenvironment23. KLF5 regulates the expression of various target genes and participates in cellular functions, including stemness, proliferation, apoptosis, autophagy, and migration24. It is an essential transcription factor in cardiovascular remodeling, thereby serving as a potential therapeutic target25. In our study, KLF5 was expressed at lower levels in paired and unpaired ccRCC tissue samples collected from the TCGA database. Similarly, it was downregulated in renal cancer cell lines. Moreover, our results confirmed a downregulated KFL5 expression in ccRCC tissue compared with the adjacent tissues. Elevated levels of KFL5 were associated with longer OS, DSS, and PFI in ccRCC, suggesting its role as a tumor suppressor. Additionally, our results revealed a positive correlation between KLF5 expression and immune cell infiltration in ccRCC, suggesting its role in regulating tumor immunity. The potential of immune cells in regulating tumor growth is well-established, and immune cell infiltration around tumors is highly significant. Reports suggest that KLF5 inactivation delays the growth of basal-like breast tumors in a CD8 T cell-dependent manner26. Knocking down KLF5 can make tumors sensitive to PD-1 blocking by increasing CD4+ and CD8+T cells while reducing bone marrow-derived cells27. Collectively, these findings suggest KLF5 as a promising prognostic and therapeutic target for patients with ccRCC.
KLF9 is involved in transcriptional regulation and plays a crucial role in various cellular processes such as proliferation, differentiation, and the development of tissues and organs28. KLF9 is downregulated in various tumor tissues and cancer cells, thereby regulating cancer cell proliferation and apoptosis29. According to a study, KLF9 also participates in tumor cell invasion and metastasis30. Our results also reported downregulated levels of KLF9 in non-paired and paired tumor tissues. Moreover, patients with high KLF9 mRNA expression showed a significantly increased OS, PFS, and DSS. Additionally, its expression decreased with an increase in the tumor grade.
KLF15 plays a crucial role in various biological processes, including the regulation of lipid metabolism, plasma corticosteroid transport, inflammatory responses31, and inhibition of cardiac hypertrophy32. Our results reported downregulated expression of KLF15 in both non-paired and paired tumor tissues and increased OS and DSS with increased KLF15 mRNA levels. This demonstrates the anti-cancer effects of KLF15 and its potential as a discriminatory factor for malignant tumors in ccRCC based on its relative expression levels.
KLF16 regulates the dopaminergic transmission, metabolism, and related endocrinology. Recent studies have shown the involvement of KLF16 in various disease mechanisms, including insulin resistance and hepatic steatosis33, oxidative stress, and inflammation34. In our study, the mRNA levels of KLF16 were upregulated in non-paired and paired tumor tissues. Furthermore, an upregulated expression of KLF16 was associated with increased OS, DSS, and PFI. Consequently, KLF16 has a significant oncogenic role, and its relative expression levels can be used to identify ccRCC.
Furthermore, research suggests that KLF17 is commonly downregulated in various cancer types, including colorectal, breast, lung, esophageal, hepatocellular, and gastric cancers35,36,37,38,39,40. Our study results revealed an upregulated KLF17 expression in the non-paired tumor tissues. In addition, it was associated with increased OS, DSS, and PFI, contrary to the findings observed in the aforementioned cancers.
Conclusion
We propose that the KLF family genes, especially KLF5, hold significant potential as prospective biomarkers and therapeutic targets for ccRCC, providing crucial implications for the diagnosis and prognosis of this disease.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Dy, G. W., Gore, J. L., Forouzanfar, M. H., Naghavi, M. & Fitzmaurice, C. Global burden of urologic cancers, 1990–2013. Eur. Urol. 71(3), 437–446 (2017).
Tian, Y. Q. et al. Trends and risk factors of global incidence, mortality, and disability of genitourinary cancers from 1990 to 2019: Systematic analysis for the global burden of disease study 2019. Front. Public Health 11, 1119374 (2023).
Zi, H. et al. Global, regional, and national burden of kidney, bladder, and prostate cancers and their attributable risk factors, 1990–2019. Mil. Med. Res. 8(1), 60 (2021).
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Primers 3, 17009 (2017).
Janzen, N. K., Kim, H. L., Figlin, R. A. & Belldegrun, A. S. Surveillance after radical or partial nephrectomy for localized renal cell carcinoma and management of recurrent disease. Urol. Clin. North Am. 30(4), 843–852 (2003).
Black, A. R., Black, J. D. & Azizkhan-Clifford, J. Sp1 and kruppel-like factor family of transcription factors in cell growth regulation and cancer. J. Cell Physiol. 188(2), 143–160 (2001).
Safe, S. & Abdelrahim, M. Sp transcription factor family and its role in cancer. Eur. J. Cancer 41(16), 2438–2448 (2005).
Narla, G. et al. KLF6, a candidate tumor suppressor gene mutated in prostate cancer. Science 294(5551), 2563–2566 (2001).
Wei, D., Kanai, M., Huang, S. & Xie, K. Emerging role of KLF4 in human gastrointestinal cancer. Carcinogenesis 27(1), 23–31 (2006).
Hamik, A. et al. Kruppel-like factor 4 regulates endothelial inflammation. J. Biol. Chem. 282(18), 13769–13779 (2007).
Mallipattu, S. K. et al. Kruppel-like factor 6 regulates mitochondrial function in the kidney. J. Clin. Invest. 125(3), 1347–1361 (2015).
Shi, Z. et al. Identification and validation of a novel ferroptotic prognostic genes-based signature of clear cell renal cell carcinoma. Cancers (Basel) 14(19), 4690 (2022).
Xiong, Q., Ruan, X. Y. & Fang, X. D. Progress on Sp1-like and Kruppel-like factors. Hereditas (Beijing) 32(6), 531–538 (2010).
Kaczynski, J., Cook, T. & Urrutia, R. Sp1- and Kruppel-like transcription factors. Genome Biol. 4(2), 206 (2003).
Moore, D. L. et al. KLF family members regulate intrinsic axon regeneration ability. Science 326(5950), 298–301 (2009).
Wu, Z. & Wang, S. Role of kruppel-like transcription factors in adipogenesis. Dev. Biol. 373(2), 235–243 (2013).
Pearson, R., Fleetwood, J., Eaton, S., Crossley, M. & Bao, S. Kruppel-like transcription factors: A functional family. Int. J. Biochem. Cell Biol. 40(10), 1996–2001 (2008).
Rane, M. J., Zhao, Y. & Cai, L. Krupsilonppel-like factors (KLFs) in renal physiology and disease. EBioMedicine 40, 743–750 (2019).
He, C. et al. Overexpression of Krueppel like factor 3 promotes subcutaneous adipocytes differentiation in goat Capra hircus. Anim. Sci. J. 92(1), e13514 (2021).
Jones, J. et al. KLF3 mediates epidermal differentiation through the epigenomic writer CBP. iScience 23(7), 101320 (2020).
Pearson, R. C., Funnell, A. P. & Crossley, M. The mammalian zinc finger transcription factor Kruppel-like factor 3 (KLF3/BKLF). IUBMB Life 63(2), 86–93 (2011).
Zhu, J. et al. Pan-cancer analysis of Kruppel-like factor 3 and its carcinogenesis in pancreatic cancer. Front. Immunol. 14, 1167018 (2023).
Wei, R. et al. Ketogenesis attenuates KLF5-dependent production of CXCL12 to overcome the immunosuppressive tumor microenvironment in colorectal cancer. Cancer Res. 82(8), 1575–1588 (2022).
Luo, Y. & Chen, C. The roles and regulation of the KLF5 transcription factor in cancers. Cancer Sci. 112(6), 2097–2117 (2021).
Xie, Z. et al. Current knowledge of Krüppel-like factor 5 and vascular remodeling: Providing insights for therapeutic strategies. J. Mol. Cell Biol. 13(2), 79–90 (2021).
Wu, Q. et al. KLF5 inhibition potentiates anti-PD1 efficacy by enhancing CD8(+) T-cell-dependent antitumor immunity. Theranostics 13(4), 1381–1400 (2023).
Li, J. et al. Epigenetic and transcriptional control of the epidermal growth factor receptor regulates the tumor immune microenvironment in pancreatic cancer. Cancer Discov. 11(3), 736–753 (2021).
Avila-Mendoza, J., Subramani, A. & Denver, R. J. Kruppel-like factors 9 and 13 block axon growth by transcriptional repression of key components of the cAMP signaling pathway. Front. Mol. Neurosci. 13, 602638 (2020).
Simmen, F. A., Su, Y., Xiao, R., Zeng, Z. & Simmen, R. C. The Kruppel-like factor 9 (KLF9) network in HEC-1-A endometrial carcinoma cells suggests the carcinogenic potential of dys-regulated KLF9 expression. Reprod. Biol. Endocrinol. 6, 41 (2008).
Li, Y. et al. KLF9 suppresses gastric cancer cell invasion and metastasis through transcriptional inhibition of MMP28. FASEB J. 33(7), 7915–7928 (2019).
Jiang, Z. et al. KLF15 cistromes reveal a hepatocyte pathway governing plasma corticosteroid transport and systemic inflammation. Sci. Adv. 8(10), eabj2917 (2022).
Leenders, J. J. et al. Regulation of cardiac gene expression by KLF15, a repressor of myocardin activity. J. Biol. Chem. 285(35), 27449–27456 (2010).
Sun, N. et al. Hepatic Kruppel-like factor 16 (KLF16) targets PPARalpha to improve steatohepatitis and insulin resistance. Gut 70(11), 2183–2195 (2021).
Xin, Y. et al. Knock out hepatic Kruppel-like factor 16 (KLF16) improve myocardial damage and promoted myocardial protection of myocardial ischemia-reperfusion via anti-oxidative and anti-inflammation effects by TFAM/PPARbeta signal passage. Bioengineered 12(2), 10219–10231 (2021).
Ali, A. et al. Tumor-suppressive p53 signaling empowers metastatic inhibitor KLF17-dependent transcription to overcome tumorigenesis in non-small cell lung cancer. J. Biol. Chem. 290(35), 21336–21351 (2015).
Ismail, I. A., Kang, H. S., Lee, H. J., Kim, J. K. & Hong, S. H. DJ-1 upregulates breast cancer cell invasion by repressing KLF17 expression. Br. J. Cancer 110(5), 1298–1306 (2014).
Ali, A. et al. KLF17 empowers TGF-beta/Smad signaling by targeting Smad3-dependent pathway to suppress tumor growth and metastasis during cancer progression. Cell Death Dis. 6(3), e1681 (2015).
Jiang, X. et al. Clinical significance and biological role of KLF17 as a tumour suppressor in colorectal cancer. Oncol. Rep. 42(5), 2117–2129 (2019).
Gumireddy, K. et al. KLF17 is a negative regulator of epithelial-mesenchymal transition and metastasis in breast cancer. Nat. Cell Biol. 11(11), 1297–1304 (2009).
Peng, J. J. et al. Reduced Kruppel-like factor 17 (KLF17) expression correlates with poor survival in patients with gastric cancer. Arch. Med. Res. 45(5), 394–399 (2014).
Acknowledgements
The authors would like to thank all the investigators who contributed to this study.
Funding
This work was supported by National Natural Science Foundation of China (82060148).
Author information
Authors and Affiliations
Contributions
M.R.F. and Y.Z.D. designed the study. Y.Y. and J.X. supervised the research. M.R.F. performed the analysis, plotted the graphs. M.R.F. and F.L. conducted the experiment. Y.Z.D. collected clinical specimens. M.R.F. and Y.Z.D. wrote the manuscript. L.Z. Y.Z. and Y.J.Y. edited and reviewed the manuscript. All authors read and accepted the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fu, M., Du, Y., Liu, F. et al. Prognostic value of KLFs family genes in renal clear cell carcinoma. Sci Rep 14, 20204 (2024). https://doi.org/10.1038/s41598-024-69892-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69892-5
- Springer Nature Limited