Abstract
At least 40% of human cancers are associated with aberrant ERK pathway activity (ERKp). Inhibitors targeting various effectors within the ERKp have been developed and explored for over two decades. Conversely, a substantial body of evidence suggests that both normal human cells and, notably to a greater extent, cancer cells exhibit susceptibility to hyperactivation of ERKp. However, this vulnerability of cancer cells remains relatively unexplored. In this review, we reexamine the evidence on the selective lethality of highly elevated ERKp activity in human cancer cells of varying backgrounds. We synthesize the insights proposed for harnessing this vulnerability of ERK-associated cancers for therapeutical approaches and contextualize these insights within established pharmacological cancer-targeting models. Moreover, we compile the intriguing preclinical findings of ERK pathway agonism in diverse cancer models. Lastly, we present a conceptual framework for target discovery regarding ERKp agonism, emphasizing the utilization of mutual exclusivity among oncogenes to develop novel targeted therapies for precision oncology.
Similar content being viewed by others
Background
Cancer targeting: from chemotherapy to precision oncology
In 1904, after a series of studies on different model organisms, Paul Erlich, a highly meritorious German medical scientist, coined chemotherapy to address his approach when chemicals were used to target infectious diseases1,2 (see Fig. 1). Ehrlich humbly stated: Indeed, from the very origin of the art of healing, chemotherapy has existed since almost all the medicaments we employ are chemicals1. However, Ehrlich’s concept of chemotherapy unprecedently linked the chemical structure and chemoreceptors on the target cells to make sense of the pharmacological activity3,4. He addressed chemicals that selectively exert their detrimental effect on the target cells (e.g., parasites) and not on the treated organism cells as Zauberkugeln or magic bullets1,2. Erlich had tested compounds, such as alkylating agents, to target cancer, which, in his hands, showed limited effects, far from being considered Zauberkugeln. Still, Ehrlich remained optimistic that effective chemotherapeutics could be explored in the future1. He explained this lack of effect by the high similarity between cancer and non-cancer cells1.
During World War II, in a mouse xenograft lymphoma model, two American pharmacologists, Alfred Gilman, and Louis Goodman, showed that nitrogen mustard, a warfare gas, had (chemo)therapeutic effects5,6. Their work immediately justified the treatment of a non–Hodgkin’s lymphoma patient and other cancers with nitrogen mustard5,7. Despite further knowledge about its limited efficacy and proneness to resistance, nitrogen mustard’s ephemerous success laid the foundation for establishing cancer chemotherapy as a valid field. Since then, several cancer chemotherapeutics such as Alkylating, Antimicrotubular agents, and Antimetabolites have been developed8. Cancer chemotherapy is a term whose definition was progressively elucidated long after it was coined, considering the expanding knowledge about chemotherapeutics’ mechanism of action and limitations. Compared to more selective therapeutics, Chemotherapeutics are considered one-size-fits-all treatments targeting different cancer types carrying distinct driver oncogenes8. Despite their narrow therapeutic window and the possibility of targeting non-cancerous dividing cells, chemotherapeutics have saved lives in some cancer populations, such as pediatric hematologic cancers, or among adult malignancies, such as testicular cancer9. Until today, chemotherapy, surgery, and radiotherapy are on the list of available neoadjuvant, adjuvant, or combined cancer treatments8,10.
Followed by the burst of knowledge about cancer-related gene mutations in the 1990s, targeting oncogenes laid the foundation for targeted therapy in cancer11. Indeed, since the New Era of Personalized Medicine in 199911, cancer treatment has been revolutionized. We have seen the mainstream shift from “one‐size‐fits‐all” approaches, such as classical chemotherapy, to individualized treatments according to patients’ tumor genetic profiles12,13. A milestone in personalized cancer therapy was the success of Imatinib, a small molecule inhibitor of the BCR‐ABL protein tyrosine kinase. This inhibitor was initially conceptualized and profiled by Nicholas Lydon and colleagues14. Imatinib demonstrated remarkable clinical advantages during phase II trials, particularly among patients with chronic myeloid leukemia (CML)15,16.
By 2002, to explain mechanisms behind the sensitivity of some cancers to mutation-specific targeting, Bernard Weinstein coined the term oncogene addiction to describe a phenomenon that, despite extensive genetic alterations, cancer cells can depend on a single oncogene activity17,18,19,20,21,22,23. One can analogize eliminating that very oncogene activity in the addicted cell to surpassing the cell’s system biology threshold, ultimately leading to a lethal trade-off favoring proapoptotic vs. the prosurvival signals. Various therapeutics have been developed against other oncogenes to which different cancers are addicted. EGFR- or ALK-altered lung cancers and BRAF mutant melanoma are some success stories of personalized cancer treatments24,25,26. All these therapeutics work by inhibiting a driver oncogene or its downstream effector, based on the rationale that withdrawal from oncogene addiction leads to deleterious oncogenic shock27. Indeed, target inhibition is a shared feature of all these treatments. Although the targets are often kinase, non-kinase protooncogenes, such as KRASG12C, are also actionable28. Sooner or later, however, all these treatments are doomed to the rise of resistance mechanisms, leading to the relapse of the treated cancer25,29,30,31,32,33.
A theoretical alternative of oncogene inhibition as a therapeutic concept is the re-introduction or restoration of a lost tumor suppressor (TS) activity, as cancer cells might be sensitive to that lost function34,35. However, this strategy has been proven challenging as often, if not always (e.g., some p53 variants), TS genes are lost due to various genetic and epigenetic alterations and, therefore, non-targetable in cancer cells. The p53 targeted therapies, conceptualized according to the p53 status of the cancer cell, are being explored36. These approaches can involve different strategies, from stabilizing the wild-type p53 and unleashing its wide range of TS functions in cells with existing wild-type copies to restoring the wild-type conformation among responsive mutant p53 variants36. In the bargain, numerous efforts have been made to re-express tumor suppressors by viral and non-viral gene therapy in tumor cells37. Viral tumor suppressor gene transfer has even been combined with the TS-loss-targeted-oncolytic capacity of therapeutic viruses or synergistic effects of TS re-expression with immunotherapeutics to double-punch the cancer cells37,38,39,40. Despite all these cumbersome efforts, none has yet borne fruit in the clinic among large patient populations. Moreover, in the late stages of cancer, due to the highly divergent genetic build-up of late-stage tumor cells, it is to be further elucidated whether cells have already circumvented their native sensitivity and response to the lost tumor suppressors.
By 1945, Theodosius Dobzhansky, a Russian-American geneticist, had coined the term synthetic lethality, describing a scenario when two distinctive chromosomes in Drosophila pseudoobscura were tolerated if existing alone but became lethal when they co-existed synthetically, as being imposed in the lab, in cross-over flies41. In 2009, Nobel Prize laureate William Kaelin Jr. put synthetic lethality forward as a conceptual framework for exploring novel anticancer treatments42. The cancer research field further translates this as the genetic build-up in certain cancers conferring sensitivity to specific chemical or genetic perturbations, opening novel avenues for indirectly targeting previously non-druggable targets43,44. Today, synthetic lethality is described as the cellular lethality of at least two co-occurring contextual (genetic) or chemical perturbations targeting at least two separate genes. In contrast, these perturbations are tolerated at variance45. As a result, within the scope of personalized medicine, treatments have been developed by targeting specific non-oncogenes based on the concept of synthetic lethality43,44. An example of such an approach in the clinic is the adjuvant as well as palliative PARP inhibitor treatment of BRCA1/2-mutant HER2-negative breast cancer46. An exciting feature of synthetic lethality is that it can be applied to both an oncogene and a tumor suppressor as long as they confer sensitivity to a synthetically lethal target44. An interesting reconceptualization of synthetic lethality as a cancer-targeting approach, namely the one-two punch47 model, is being pioneered and explored by René Bernards and colleagues. In this model, the first-line treatment is designed to induce senescence-associated vulnerabilities in cancer cells that are to be targeted with the second-line senolytic compounds47.
Notably, in recent years, targeting cancer-related immune cells or their interaction with cancer cells has opened a new front in the battle against cancer48,49,50. For instance, in BRAF mutant melanoma, in which targeted inhibitors are known to be at their highest performance, immune checkpoint inhibitors showed a 20% benefit over targeted compounds in terms of overall survival among therapy-naïve and metastatic patients51. As such, oncogene-targeted therapy might seem on the verge of losing momentum in personalized medicine. However, despite the striking responses to different cancer immunotherapeutics in a group of patients, many patients remain non-responsive or fast-resistant to such therapies48,49,50,52.
While it is essential to work towards enhancing existing therapeutic strategies, it is also highly justified to put forward and explore innovative and unconsumed concepts.
By 2015, two significant studies showed that mutual exclusivity among BRAF/KRAS and EGFR/KRAS oncogenes can result from synthetic lethality or senescence43,53.
The British fairy tale “Three Bears” (Eleanor Mure, 1831) revolves around a greedy individual who, in the absence of the three bachelor bears, each of varying sizes, enters their cottage54. She samples their trio of distinct milk portions, chairs, and beds. Repeatedly, amidst these triads, she discovers contentment solely when the element is precisely balanced – not too much, not too little, but just right54. In later biological and medical contexts, the widely spread but modified version of the original story “Goldilocks and the Three Bears,” in which a young girl replaces the elderly woman, has inspired the Goldilocks principle. This principle signifies that, for a biological system to function optimally, its components should possess the correct levels of abundance and activity and be temporally right – avoiding both excess and deficiency55.
Amit Dipak Amin and Jonathan Schatz, inspired by the Goldilocks principle in Biology, introduced the term and the concept of “Oncogene Overdose.” This concept compares cancer cells to drug addicts, as both can die due to an overwhelming excess of what they are addicted to55.
In 2016, Nobel laureate Harold Varmus introduced the concept of harnessing mechanisms elicited by the co-induction of mutually exclusive genes as a therapeutic model56. Furthermore, Xu et al. from Deng’s laboratory have explored a striking KRAS agonist for targeting KRAS mutant cancers57. More recently, Sugiura et al. presented the therapeutic concept of ERKp activators58,59,60. Concurrently, René Bernards and colleagues described Paradoxical Intervention, signifying the simultaneous activation of mitogenic signals and the employment of stress-inducing compounds in cells with pre-existing ERK pathway-activating alterations61,62. Remarkably, they have demonstrated that such treatment enforces unprecedented tumor-suppressive resistance mechanisms62.
Dostoevsky once remarked, ‘We all come out from Gogol’s ‘Overcoat,” paying homage to Nikolai Gogol as the iconic predecessor of Russian literature preceding his own generation. In doing so, he employed wordplay by referring to one of Gogol’s renowned short stories, namely, ‘The Overcoat.’“ Based on chronological order and logical reasoning, all the above concepts revolve around over-activating the already activated pathway as a therapeutic approach, akin to stepping out from oncogene overdose overcoat.
In this writing, we review the evidence and insights that reinforced ERK pathway effectors activity in cells with aberrantly increased ERK activity is damaging to these cells. The pathway is also known as mitogen-activated protein kinase (MAPK) or Ras-Raf-MEK-ERK pathway. In this writing, for simplicity, we will mention the pathway by its major effector ERK without distinguishing between the two human ERK genes, ERK1 and ERK2. Furthermore, we will present a conceptual framework for therapeutic targeting of such vulnerabilities.
The double-life of ERKp in human cancer and the curious case of the ERK proteins
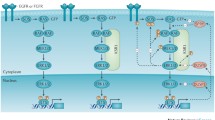
The ERK pathway is one of the extensively studied signal transduction cascades. Essential effectors of this signaling cascade in humans fall into three types of eukaryotic protein kinases: (1) HER family Receptor Tyrosine Kinases (RTKs, EGFR, HER2-4), RAF family serine/threonine kinases (A/B/CRAF), and (3) dual specificity protein kinase families MEK (MEK1/2) and ERK (ERK1/2)63,64,65. In humans, the RAS family, consisting of three essential members, H/N/KRAS, serve as the principal non-kinase effectors within the ERK pathway66.
Feedback loops regulate ERK signaling during development and in normal physiologic conditions for cell proliferation, survival, and homeostasis63,64,65,66.
Due to genetic alterations of the ERK pathway effectors and/or its regulatory components, ERK signaling may become pathologically deregulated. Aberrant down-regulation of the ERK pathway can be associated with some neurodegenerative or autoimmune disorders, while its abnormal upregulation is associated with human cancers, rasopathies, and Erdheim-Chester disease66,67,68,69.
Under normal cellular conditions, upon ligands binding, HER receptors can undergo conformational changes, transitioning to an active state and forming homo- or hetero-dimers70. The induced conformational alterations in the tyrosine kinase domain of dimerization partners facilitate ATP binding in their ATP binding cleft and, as suggested, in the case of EGFR, might lead to the relief of cis-autoinhibition and trans-autophosphorylation of the tyrosine kinase domains70,71.
Under physiological circumstances, the mature RAS protein, when residing in the inner surface of the plasma membrane, undergoes conformational changes mediated by the upstream HER signaling or regulatory feedback signals66,72. RAS alternates between a guanosine triphosphate (GTP)-bound state and a guanosine diphosphate (GDP)-bound state28,66,73. The oscillation of RAS into its active conformation involves GTP loading28,66,73. This process can be initiated by recruiting GRB2 proteins to the plasma membrane, where the GRB2 is activated via its SH2 domain by an active RTK66. Subsequently, SOS is recruited, forming a complex involving at least GRB2, SOS, and RAS66. This complex facilitates the displacement of GDP and promotes the loading of GTP onto the RAS molecule66. RAS can hydrolyze its GTP as a GTPase protein, converting it to GDP and becoming inactive28,66,73. This process is facilitated by GTPase-activating proteins (GAPs)28,66,73. Feedback loops play a regulatory role in the GTPase activity of RAS66. The GTP-bound RAS triggers the activation of downstream RAF by engaging in a physical (allosteric) interaction with RAF or mediating the release of its autoinhibition74. RAFs, in turn, phosphorylate the MEK1/2 through phosphotransferase activity, and MEKs, in turn, phosphorylate the ERKs, which, due to the Erks’ vast array of network, major cellular events such as proliferation and survival can take place63,64,65,66. At least 659 of ERKs’ direct substrates are discovered75.
Approximately 40% of human cancers are linked to the abberant upregulation of ERKp76. Notably, cancers that harbor genetic alterations in an ERKp effector can be associated with worse clinical outcomes than cancers where such alterations are absent, emphasizing the significance of these alterations on cancer prognosis77. For over two decades, the effective inhibition of the ERKp has remained a key focus in cancer research and targeted therapy efforts12,24,26,28,77.
However, in an ironic twist, a wealth of evidence underscores the reality that human cells, including cancer cells, cannot tolerate excessive levels of ERKp activity. While upstream effectors of the ERKp, such as RTKs, RAS, and RAFs, are subject to activating and oncogenic mutations in humans, the mutations in downstream effectors, such as MEK and even to a greater extent ERK, are infrequent26,73,77,78. Before the ERK small molecule inhibitors’ development and clinical employment, ERK mutations were hardly reported in human cancer79. These mutations can be associated with secondary resistance mechanisms that lead to loss-of the protein affinity to ERK inhibitors in cells treated with these compounds79,80,81. As illustrated in Fig. 2, among the conventional ERK signaling pathway effectors, the ERK mutations still rank at the bottom of the list regarding the frequency of mutations in human cancer.
In July 2023, the query of 69223 samples belonging to 65853 patients in 213 curated and non-redundant studies in cBiportal for only mutations among conventional effectors of the ERKp yielded the approximate frequencies as displayed. Note that this oversimplified schematic is not meant to communicate complex signaling and structures of the effectors. Unlike typical oncogenes, ERK1/2 mutations are distributed along the ERK protein’s conserved protein kinase and non-conserved regions, with no hotspot mutational site. Figure generated in Biorender.
The ERKs need two critical phosphorylation events before full conformational activation, none triggered by either cis- or trans-autophosphorylation mechanisms82. As opposed to RTKs, RAFs, or even MEKs, ERKs protein conformations have evolved in such a way to be less poised for autoactivation and instead rely on direct phosphotransferase activity of MEKs for full activation82,83.
Several activating variants that cause increased kinase-dependent or -independent activity of the ERK orthologues, such as Sevenmaker variants, are studied in Yeast and Drosophila models83,84,85. However, how such mutations can induce or contribute to human carcinogenesis remains enigmatic. Several investigations have concentrated on characterizing a few purportedly activating ERK variants by expressing them in mammalian cells82,86,87. These studies have looked into activating variants of ERK that render the ERK protein an autoactivation capacity82,86,87. In particular, the activating variant ERK1R84H, reported twice in human cancer, was found to transform NIH3T3 cells87. While introducing these presumably activating ERK variants into non-cancerous mammalian cells was occasionally feasible, accomplishing the same in human cancer cell lines appeared to pose challenges. Markedly, Goetz et al. had shown in the past that overexpression of wild-type ERK1/2 and not the kinase-dead or low-kinase ERK1/2 variants in BRAFV600E mutant melanoma cell line A-375 leads to growth inhibitory effects88. Interestingly, the ERK1/2 variants with increased kinase activity, which were found to confer resistance against RAF inhibitors (RAFi) and MEK inhibitors (MEKi) during a random mutation screen, even exerted more potent growth inhibition when overexpressed in A-375 cells. The most active ERK1/2 variants exhibited growth-suppressive effects in two other BRAF mutant melanoma cell lines beyond A-375 cells(SKMEL-19, and WM266.4)88. It was also shown that inducible overexpression of ERK2 in A-375 cells was selectively detrimental to these cells vs. cells with wild-type BRAF and caused anti-tumor effects in vitro and in vivo89. Such detrimental effects could only be rescued upon ERK2 or BRAF knock-down89. The ERK2 overexpression in these cells was associated with the induction of ER stress and DNA damage in addition to proapoptotic signals89. The fact that some ERK-associated cancers, such as BRAF mutant melanoma, are sensitive to ERK activation is well-established.
Interestingly, the negative effect of putative Gain-of-Function (GOF) mutations on the proliferation of these cells has been utilized as a model in an ERK saturation mutagenesis study90. Such a comprehensive approach by Brenan et al. has shed light on the relevance of ERK mutations in human cancer. It is worth noting that saturation mutagenesis of ERK was unsuccessful in identifying the direct GOF impact of activating variants, unless upon MEKi and BRAFi treatment. Indeed, expression of GOF sevenmaker ERK mutations ERK2D321N and ERK2E322K or other supposedly GOF variants in the A-375 cell line had a robust anti-proliferative effect in these cells90. Interestingly, the sevenmaker variant and those that phenocopy its effect could only be tolerated by the BRAFV600E cells under MEK or RAF inhibition90. These variants are proposed to enhance ERK activity by disrupting ERK’s interaction with inhibitory DUSP phosphatases90.
The rarity of ERK mutations in human cancer raises an intriguing question, prompting consideration of at least two alternative scenarios to explain this phenomenon.
One possibility is that the ERK protein, by default, exhibits low basal activity and has a weak impact on its network. Consequently, even an activating mutation might not be sufficient to manifest an effective GOF phenotype leading to cell transformation. An illustrative example of such a scenario is the case of CRAF mutations in human cancer, which are rarer than mutations of another RAF isoform, BRAF91,92. Past explanations attribute this rarity to the low basal activity of CRAF compared to BRAF93. Conversely, the effectors, such as EGFR, which are more upstream and have a wider array of targets belonging to distinctive pathways, exhibit a higher frequency of mutations in human cancer. From a vertical signaling cascade standpoint, the three commonly mutated effectors of the ERKp in human cancer are arranged as EGFR, KRAS, and BRAF. EGFR and BRAF mutations exhibit a comparable frequency in human cancer (see Fig. 2). Notably, KRAS mutations occur at a frequency equivalent to the combined occurrences of EGFR and BRAF oncogenes (Fig. 2). The higher mutational frequency of KRAS over BRAF can be explained by its vertical rank along the signaling cascade, a rationale not applicable to the other two effectors82.
Another explanation could be that while ERK mutations capable of generating an effective GOF phenotype may exist synthetically, they are negatively selected in the real world due to being poorly tolerated by human cells. In contrast to, for instance, KRAS, which oscillates between active and inactive states under normal physiological conditions and may become constitutively active in human cancer73, human cells cannot tolerate ERKs in a constitutively active state. It is crucial to distinguish between two types of ERK mutations. The first type comprises mutations that occur in the real world. As the above evidence suggests, these mutations exhibit low transforming capacity. Then comes the synthetic ERK mutations, which do not occur in the real world. Indeed, as mentioned in the above paragraphs, such synthetic variants have been studied in the past, and it has been shown that they are not easily tolerated by human cells, particularly cancer cells79,81,82,84,85,86,88,89,90. More interestingly, in addition to some ERK mutants, even wild-type ERK overexpression is not tolerated in such human cellular models88,90. Indeed, one of the vital pieces of evidence is ERK saturation mutagenesis by Brenan et al., as they observed in human melanoma cell line model with BRAFV600E, expression of wild-type ERK and activating ERK variants could be tolerated only in the presence of ERK pathway inhibitors90. This evidence, at least, can rule out the possibility that ERK, by nature, cannot render gain-of-function. The scenario that ERK-activating mutations are not easily tolerated by human cells aligns with one aspect of ERK protein activity autoregulation, namely its reduced potential for autoactivation, in contrast to other ERK pathway effectors like EGFR82,83. The ERK protein is shown to tolerate synthetic mutations that can render it increased autophosphorylation and kinase activity to levels comparable to ERK582.
The RTK/RAS/RAF pathway and its inhibitors exhibit a Janus-faced immunomodulatory effect in cancer (reviewed here94). In brief, inhibition of the ERKp can enhance the activity of associated T-helper cells and cytotoxic T-cells, thereby potentially enhancing the immune response against cancer cells94. Additionally, it can enhance dendritic cell activity, which plays a crucial role in antigen presentation and immune activation. Conversely, this inhibition may hamper tumor infiltration of immunosuppressive regulatory T-cells, monocytes, and macrophages, potentially limiting their immunosuppressive functions94. On the other hand, long-term ERKp inhibition might eventually have an immunosuppressive effect95. Of note, oncogenic RAS can contribute to immune evasion by stabilizing the PD-L1 mRNA and subsequently favoring the PD-L1 expression on tumor cells96.
ERK pathway hyperactivation can be lethal to cells in various ways
The ERKp activity can lead to cell toxicity and death (see Fig. 3). Numerous studies have unraveled such a role in different model organisms60,97,98. The ERK-induced cell apoptosis can involve extrinsic or intrinsic apoptotic pathways60,97,99. ERK activity affects cell death by influencing multiple cellular processes, including mitochondrial dysfunction100,101,102, DNA Damage Response103, Endoplasmic Reticulum stress104, Autophagy modulation60,105, Metabolic imbalance106, and accumulation of Reactive Oxygen Species(ROS)97,103. ERKp activity can trigger senescence in vitro and in vivo97. An interesting example of such a phenomenon is the existence of benign human naevi with BRAFV600E mutation107,108. In cases where this mutation occurs in isolation and without the concurrent loss-of the p53 tumor suppressor, it has been proposed to result in irreversible cellular senescence rather than malignancy108. More recently, an alternative mechanism involving non-coding RNAs has come to light, linking BRAFV600E-carrying benign naevi to occurrences of mitotic failure and reversible proliferation arrest107.
ERKp activity is crucial in determining divergent cellular outcomes, including cell proliferation, growth arrest, or cell death. Notably, Hong et al. from Park’s lab have elucidated an exclusive threshold for ERK activity109. Previously, it was known that excessive activity of ERKs, like other effectors such as MEKs, CRAF, and BRAF, can cause growth arrest110,111,112,113. However, Hong et al. show that very high activity levels of ERKp, exceeding those that cause growth arrest, can lead to apoptosis109. These ultra-high ERKp levels in HEK293 and U251 cells could only be achieved by combining ERKs overexpression and the presence of tamoxifen-inducible active CR3 catalytic domain of CRAF109. As proposed by Hong et al., this cell death-inducing characteristic is unique to the ERKs compared to the upstream kinase effectors of the ERKp109.
Overall, the evidence is ample that excessive ERK activity is toxic to cancer cells with different tissue backgrounds. First, we highlight a few critical early studies among such evidence. Early evidence of ERK’s proapoptotic activity was observed in the human breast cancer cell line (MCF-7), as RAF depletion could desensitize these cells to Paclitaxel114. During the characterization of Phenethyl Isothiocyanate anti-proliferative effects against p53-deficient prostate cancer cell line PC-3, ERKp prolonged activation was responsible for the compound’s growth inhibitory effects115. Moreover, as shown in mouse embryonic fibroblasts and MCF-7, the DNA Damage (DD) caused by different stimuli, including Etoposide, could trigger ERKp activation independent of p53 but rest on Ataxia-Telangiectasia Mutant (ATM)116. The ERK activity was essential for DD-triggered growth arrest and apoptosis116. Later, it was shown that growth inhibitory and proapoptotic effects of Asiatic acid against breast cancer cell lines (MCF-7 and MDA-MB-231) were associated with the induction of ERKp117. In another study investigating Lauryl-gallate’s growth-suppressive and proapoptotic effect on three breast cancer cell lines MCF-7, MCF-7 ADR, and MDA-MB-231, the observed phenotype was accompanied by ERKp activity and p21-induced Cell Cycle (CC) arrest118. As one of the investigated cell lines was p53 mutant (MDA-MB-231) and the other possessed a multidrug-resistant phenotype (MCF-7 ADR), authors conclude that none of these conditions affects the sensitivity of cells to the tested compound118. Interestingly, MEK inhibition and the resulting ERKp suppression were associated with rescuing the drug effect118. In another study, sustained ERKp activity due to exogenous expression of active MEK1/2 led to G1 CC arrest, marked by p21WAF expression119. Prolonged ERK activity was associated with the activation of cellular protein biosynthesis regulator p70S6K, accompanied by an increased translation of p21 protein120. Nonetheless, none of these studies has fully elucidated the mechanisms elicited by ERK activation to lead to the observed phenotypes. Several other studies demonstrate that reinforced ERK activity in cancer cells potentiates the cytotoxic effects of various chemotherapeutical agents (reviewed here60,97). In many of these studies, the MEK inhibitor compounds, if among other means, were employed to investigate the rescue effects60,97. Moreover, as recently highlighted by Sugiura et al.60 and taking into account subsequent revelations regarding MEK inhibitors off-targets and mechanisms of action, these factors could potentially undermine the precision of the abovementioned findings.
A revisit of the precedent evidence reveals an intriguing aspect of ERKp-induced cell death and senescence, as it can happen in both p53 wild-type and p53-altered backgrounds114,115,116,117,118,119,120. Numerous studies have indicated crosstalk and interactions between the p53 and ERKp, suggesting that they can influence each other’s functions and responses under specific cellular conditions. As a major tumor suppressor known as “the guardian of the genome121,” p53 governs safeguard mechanisms that prevent the accumulation of DNA damage and the proliferation of damaged cells122. The p53 protein primarily functions as a transcription factor, regulating a plethora of genes that control cell proliferation, apoptosis, DNA repair, and other cellular processes123. Additionally, p53 possesses multiple transcription-independent activities in cell death124,125,126, metabolism127,128, autophagy129, DNA replication130, and repair131, all of which contribute to tumor suppression and the maintenance of genomic integrity. The aberrant activation of the ERKp leads to the induction of the p53 response, which can trigger senescence108,132,133,134,135,136 or cell death109,137. The classical mechanism of p53 activation by oncogenes is mediated by the p14ARF protein, which sequesters the p53 inhibitory ubiquitin ligase Mdm2 (HDM2 in humans), allowing for the accumulation of the p53 protein138. Additionally, studies have shown that ERK can phosphorylate p53 on Ser15 under stress conditions, inhibiting its interaction with Mdm2 and promoting the stabilization of the p53 protein139,140,141. The phosphorylation of p53 by ERK2 on Thr55 was reported to induce p53 stabilization and activation in doxorubicin-treated MCF-7 cells142. However, it remains unclear to what extent the direct phosphorylation of p53 by ERK contributes to the induction of senescence and apoptosis in tumor cells without additional stress stimuli.
In the process of malignant transformation, cells with hyperactive ERK signaling experience strong negative selective pressure from p53132,143,144,145,146,147,148,149 which they can counteract by blunting p53 activity, often through the acquisition of TP53 mutations that entirely or partially inactivate the tumor suppressor150. Large-scale genomic studies have demonstrated that TP53 mutations are found in all types of cancers151. The frequency of TP53 genetic alterations, estimated at an average of 50%, varies significantly depending on the tumor entity, ranging from 5% in neuroblastoma to over 90% in ovarian and small-cell lung cancer151,152,153,154,155,156.
Somatic mutations that inactivate TP53 are predominantly observed in the later stages of tumorigenesis, indicating its role in advanced cancer progression157. It is estimated that 80% of genetic alterations in the TP53 gene are missense mutations158,159 that lead to the expression of mutant p53 proteins. These mutants can possess oncogenic properties, driving cancer progression and conferring therapy resistance160,161. However, the functional impact of the majority of cancer-associated TP53 missense mutations (approximately 70%) remains poorly characterized, and their role in tumorigenesis and therapy response remains unclear162.
TP53 mutations often co-occur with oncogenic EGFR, BRAF, and RAS mutations, but the extent of cooperation between TP53 and oncogenes may vary even within the same tumor type. For instance, in lung adenocarcinoma (LUAD), TP53 mutations are prevalent in EGFR mutant tumors but are strongly underrepresented in KRAS-driven LUAD151,163, indicating different paths of tumor evolution driven by different oncogenes. The intricate interplay between various oncogenes within the ERKp and its impact on the residual activity of wild-type or partial Loss-of-Function (LOF) p53, retained in tumors, which can potentially lead to tumor-suppressive effects, remains an area of ongoing investigation162,164. Moreover, different TP53 variants are suggested to cause distinctive LOF, dominant negative, or even GOF phenotypes, which needs to be acknowledged in further exploration of ERKp-induced lethality in different cellular contexts with distinctive TP53 status165,166,167,168.
ERK pathway in light of emerging knowledge about vulnerability of cancers to replication stress targeting
In eukaryotes, transmitting genetic material to daughter cells is a crucial event and is thus tightly regulated. Cyclin-dependent kinases (CDKs) are pivotal in advancing the CC during interphase and M phases169,170,171. The CC represents a series of sequential decisions and commitments169,170,171. Each CC stage is intricately governed by a complex interplay involving CDKs coupled with E3 ubiquitin ligases like APC/C and their activating proteins169,170,171. Varied levels of CC-related proteins can establish decision windows that impact entry, prevention of re-entry, or even the deceleration of CCs169,170,171. Although tightly controlled, the CC can face undesired events due to DD, Replication Stress (RS), or spindle assembly malfunctions169,170,171. Eukaryotic cells rely on distinct checkpoints to tackle each scenario169,170,171. RS emerges when diverse factors slowdown or stall the replication fork. DD and RS differ, but the latter can trigger DD as the collapse of the stalled fork can lead to double-strand breaks169,170,171,172. RS checkpoints function to avoid RS-induced DNA damage response, while DD checkpoints are meant to prevent the accumulation of DNA damage and thereby protect cells from subsequent complications170. A cancer’s hallmark is uncontrolled proliferation, hampering apoptosis and avoiding long-term CC exits34. Oncogenes like RAS and MYC can negatively affect DNA replication licensing and firing, inciting RS169,170,172. Deregulation of DD and growth pathways due to cancer-related mutations causes excessive S phase entry and subsequent RS. While a temporary check-out from the Mitosis/entry is possible, leaving the CC is not favored169,170,171,172. Cancer cells hold a higher basal level of RS vs. normal cells170. The severity of the damage and the context determines the cell fate, which can be apoptosis, quiescence, or senescence. It deserves to be mentioned that DD checkpoint responses can lead to a temporary exit or, as opposed to RS, an eternal exit of the CC169,170,171,172. However, DD responses and RS are so intertwined that, ultimately, RS can lead to such events169,170,171,172. Curiously, cancer cells highlight a distinct approach to DD and RS checkpoints. DD checkpoints are often surrogated in cancer; cancer cells tolerate defects in DD response and perhaps even benefit from such a compromise in favor of increased selection pressure, all while sidestepping unfavorable exits from the cell cycle169,170,171,172. Conversely, RS checkpoints are somehow intact169,170,171,172. Indeed, cancer cells are more sensitive to DD responses than RS responses170. Cancer cells tolerate and even favor some Chromosomal instability (CIN) levels170. In contrast, they cannot tolerate excessive CIN, which can be caused by excessive RS and DD, leading to catastrophic mitotic defects, loss-of-essential genes, and cell death170. Therefore, cancer relies on RS checkpoints to avoid too much CIN. Consequently, targeting RS tolerance in cancer holds promise as a viable cancer therapy approach169,170,171,172,173. Of note, studies have demonstrated that elevated oncogenic RAS activity can trigger RS by ubiquitously enhancing cellular transcription events174,175,176. This augmented transcription activity increases the likelihood of collisions between replication and transcription processes174,176. In addition to RS induction, oncogenic RAS (HRASG12V) can circumvent the p53 activity, thereby sensitizing cancer cells to RS-inducing compounds176.
Withdrawing ERK pathway inhibitors from addicted cells: lethal consequences of excessive pathway activity
There are several pieces of evidence that ERK-related cancer cells can tolerate upregulation of the ERKp only in the presence of ERKp inhibitors. Resistance mechanisms against ERKp inhibitors predominantly involve ERKp effectors and regulators30,31,81,177,178,179,180,181. Different RAF and MEK inhibitors can trigger clonal expansion of drug-tolerant cells, which maintain a proliferative advantage, perhaps preferably in the presence of inhibitors by virtue of enhanced ERKp activity182,183. This elevated activity could be lethal upon inhibitor cessation, potentially resulting in ERK-related cellular toxicities183. It is suggested ERKp inhibitors can create a window upon drug removal, in which cells lose their fitness advantage gained during drug treatment and may even experience growth disadvantage due to excessive ERKp activity and adaptive switching182,184. In an elegant study by Kong et al. conducted in Peeper lab185, a CRISPR knock-out screen of melanoma cells resistant and addicted to BRAFi revealed a phenotypic switch dependent on ERK2 kinase and JUNB and FRA1 transcription factors accompanied by suppression of microphthalmia-associated transcription factor (MITF)185. The ERK2 dependency of the observed phenotype was supported by in vivo and clinical findings185. This dependency was also observed in lung cancer cells resistant and addicted to EGFRi185. The addicted Melanoma cells experienced grave cell death upon withdrawal from BRAFi, which could be rescued by ERK2 targeting or restoring MITF activity185. Another intriguing aspect of this study was that the authors opted to investigate the intermittent drug treatment with the chemotherapeutic agent dacarbazine, and they showed BRAFi-addicted melanoma cells were sensitized to this compound accompanied with MITF inhibition185.
Aissa et al. elegantly showed at the single-cell level that drug-resistant EGFR mutant lung cancer cell clusters exhibited markers indicative of activated ERKp180. Recent work by Nuria Gutierrez-Prat et al.186 reported similar findings concerning ERK and MITF dependency on drug withdrawal toxicity. Moreover, the authors find that the knock-down of DUSP4, an ERK phosphatase and negative regulator, was lethal by causing excessive ERK activity186. This effect was observed not only in melanoma cells addicted to inhibitors of the ERKp but also, intriguingly, in drug-naïve cells186. Xue et al. in Piro Lito’s lab found the link between oncogenic BRAF protein dosage in cells, the depth of ERKp inhibition and the related resistance mechanisms. In their patient-derived xenograft lung cancer and melanoma models, they discovered the more robust the ERK inhibition is, the higher the oncogene dosage required for cells to retain proliferation advantage in the presence of inhibitors183. They proposed a fitness threshold model, suggesting that cells treated with regimens with a higher threshold, such as upon combination of RAFi, MEKi, and ERKi vs. ERKi monotherapy, might face a disadvantageous outcome due to sustained and excessive ERKp activation when the drug treatment is stopped183.
Some preclinical and early clinical evidence suggested the benefits of intermittent treatment with RAFi and MEKi vs. sequential treatments, in particular in melanoma182,183,184,187,188,189,190,191,192,193,194,195,196,197. This evidence laid the foundation for clinical trials examining intermittent treatment regimens in individuals with BRAF mutant melanoma198,199. Contrary to the expectations, these trials did not show any overall survival benefits from the intermittent therapies, and even worse progression-free survival outcomes were reported upon intermittent treatments198,199. Nevertheless, it is still uncertain whether those intermittent therapies meant to produce that high fitness threshold as recommended by Xue et al. to cause selection disadvantages in tumor cells during drug removal effectively.
Mutual exclusivity of highly activating variants of BRAF, KRAS and EGFR oncogenes in cancer: induction of synthetic lethality or senescence
In 2006, Carlotta Petti et al.200 showed that synthetic expression of NRASQ61R oncogene in a metastatic melanoma clone, which natively harbored the mutually exclusive BRAFV600E oncogene, resulted in senescence. Later, Cisowski et al. discovered that co-expression of BRAFV600E and KRASG12D under their endogenous promotors provides a selective disadvantage compared to single oncogene expression in mouse lung cells53. The decrease in tumor burden in double oncogene-expressing tumors was associated with hyperactivated ERK and AKT signaling and a decrease in proliferating cells53. Further analysis demonstrated enhanced β-galactosidase expression and increased p15, p16, and p19 levels upon oncogenes co-expression, suggesting that double oncogene-expressing cells become senescent53. Meanwhile, Unni et al. exogenously induced the expression of KRASG12V or EGFRL858R in EGFREX19Del and KRASG12C LUAD cell lines, respectively43. They observed decreased cell viability, indicating that mutant KRAS and EGFR co-expression are not tolerated in cells43. Additionally, they generated genetically engineered mice with co-induction of KRAS and EGFR mutants in lung epithelium. The established lung tumors did not grow faster than those harboring only one of the oncogenic mutations did43. Further analysis indicated that only one of these oncogenes could be activated in the tumor cells43. Furthermore, Unni et al. revealed that DUSP6 prevents ERK activity from exceeding critical thresholds in EGFR and KRAS mutant cell lines201. They found that targeting DUSP6 reduced cell viability due to unleashing the excessive and toxic levels of RAS-mediated ERK activity in cancer cells harboring mutations in EGFR and KRAS201. Markedly, Ambrogio et al. showed that conditional induction of an EGFRL858R allele in KRASG12V knock-in mouse LUAD models led to decreased tumor burden, increased mice survival, and reversible cell toxicity in remaining tumor cells202. The latter could further be recovered through ERKp activity reduction202. Of note, all these consequences were associated with hyperactivation of ERKp signaling43,201,202.
Harold Varmus, a Nobel Prize laureate, and his colleagues, who played key roles in some of the aforementioned studies, articulated and championed the intriguing concept that, unlike how previously considered, not the redundancy of functions but synthetic lethality or senescence could lie behind mutual exclusivity of certain oncogenes in cancer56.
We argue that the pathway redundancy and synthetic lethality, senescence, or any other constraining phenomenon should not necessarily be seen as conflicting scenarios. One can consider that pathway redundancy and synthetic lethality could both play roles in explaining mutual exclusivity. When two proteins with overlapping functions in a pathway are excessively active, the likelihood of both events being mutually exclusive within a cancer cell is increased.
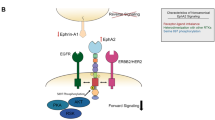
On co-induction of activating EGFR and BRAF events in the same cell, it has been shown that exogenous expression of wild-type EGFR in a Melanoma cell line with native BRAFV600E leads to decreased proliferation of these cells in a dose-dependent manner, in-vitro and in-vivo203. The slowdown in proliferation was associated with cellular senescence as suggested by hypophosphorylation of RB1 and induction of CDKN1A, CDKN1B, and Beta-galactosidase203. Concerning mutant EGFR and mutant BRAFs, two studies have been inspired so far by the emergence of BRAFV600E in treatment-refractory EGFRL858R lung cancers as they become resistant to EGFR-targeted therapies177,179. In one study, exogenous expression of BRAFV600E in a polyclonal pool of EGFR mutant lung cancer cells led to no differences in cell proliferation and cell death rate compared to empty vector control177. Of note, the BRAFV600E protein levels and mRNA levels showed an indispensable increase only in the presence of an ERKp inhibitor, suggesting that in the polyclonal pool, cells with BRAFV600E induction are not clonally expanded unless the ERKp activity is suppressed177. As such, further investigation is required to determine the existence and the mechanism behind such clonal disfavor. Overall, the findings of these two studies align with the results of two independent GOF CRISPRa screens in Vemurafenib-treated A-375 cells (BRAFV600E), showing that overexpression of EGFR, among others, is a resistance mechanism to BRAFi204,205.
Three BRAF mutational classes26 include Class I variants, like BRAFV600E, which often function independently of upstream effectors as constitutively active monomers. Class II variants, exemplified by BRAFG469A, activate the ERK pathway independently of RAS and CRAF as active homodimers. Class III involves kinase-dead BRAF mutants activating the ERK pathway through RAS-dependent allosteric transactivation of CRAF.
Cancer-relevant KRAS mutations are classified into three categories73 based on their impact on KRAS protein functions: Class I (Hydrolysis) includes mutations leading to the loss-of the GTP-hydrolyzing feature of KRAS, Class II (Exchange) involves mutations causing a gain in KRAS Exchange function facilitated by Guanine Nucleotide Exchange Factors, and Class III (Hybrid) encompasses mutations affecting both functions.
A recent classification of EGFR mutations24 considers both the structural effects of mutations on the EGFR protein, specifically its drug-binding pocket (DBP), and the implications of mutations on drug response. Accordingly, one of the EGFR mutational classes is Classical-like (relevant to this writing), where mutations like L858R have minimal impact on DBP and the affinity for corresponding Tyrosine Kinase Inhibitors.
Recently, the variant-specific landscape of mutual exclusivity among BRAF, KRAS, and EGFR mutations in cancer has been unraveled206. We learn which oncogenic variants can co-occur in the same cancer sample while certain driver events are mutually exclusive. The authors conclude that class I BRAF(in line with another recent report206,207), Hydrolysis KRAS206, and classical-like EGFR206 class mutations are less likely to co-occur. When they dissected the analyses into variants, they discovered novel instances of mutual exclusivity involving unconventional yet common oncogenic variants. They showed that specific classical-like EGFR and BRAF mutations, often the most frequent ones, are mutually exclusive in human cancer206.
Leveraging oncogenes mutual exclusivity for precision oncology: a target discovery framework for RTK/RAS/RAF pathway agonism
Reinforced activation of the ERKp could serve as a therapeutic strategy for ERK-associated cancers. As previously explained, these cancers exhibit susceptibility to elevated ERKp activity beyond conventional oncogenic levels. Importantly, this vulnerability can be selectively targeted because normal cells and ERK-associated cancer cells differ in their baseline ERKp activities. Thus, reinforcing ERK activation to levels intolerable for cancer cells may not necessarily push normal cells beyond the tolerable threshold (see Fig. 4).
This schematic is oversimplified and does not acknowledge the spatiotemporal context of ERK activity and regulation. The figure was generated using a template provided in https://www.slideegg.com.
At first glance, the primary concern with this approach is the potential undesired activation of the ERKp in non-cancerous tissues. Interestingly, this potential adverse event is not unfamiliar in precision oncology. Patients with BRAFV600E mutant cancers have been treated with various RAF inhibitors for years, initially as single agents and later combined with MEK inhibitors. Type I RAF inhibitors like Vemurafenib and Dabrafenib can cause paradoxical ERKp activation in non-cancerous cells with wild-type B/CRAF. Some patients develop benign teratomas like keratoacanthoma, but most do not26,208,209. Adding MEK inhibitors to therapy can significantly reduce the occurrence of such adverse events, although MEK inhibition is also associated with toxic effects210. Furthermore, predictive markers and signatures can help identify patients at risk of these adverse events. On the other hand, developing selective ERKp activators with exclusive effects on cancer cells could mitigate this challenge from the outset.
Our proposed model recognizes three non-detrimental and two detrimental stages and four thresholds of ERKp activity (Fig. 4). The “normal low” occurs when cells are at rest. The “normal high” occurs when cells are in proliferating status or are stressed due to intrinsic or extrinsic signals. ERKp activity spatiotemporally surpassing “normal high” physiological levels enters the “oncogenic window.” Levels below the “normal low” and above the “oncogenic window” can be detrimental. For any potential therapy, it will be necessary to set pathway activity exceeding the “oncogenic window” and entering the detrimental stage in the target cancer cells. Ideally, the therapy should not push pathway activity beyond “normal high” levels in normal cells, adhering to the Goldilocks principle.
This discussion will not delve into the chemistry and structural aspects of potential therapeutics, including small molecule activators, monoclonal antibodies, monobodies, etc. We propose three strategies to reinforce the ERKp or ERK-independent signals for detrimental effects (Fig. 5a, b).
Perturbation 1 (P1)-global activators
Potential therapeutics activate the ERKp through various means, such as ligand-mimetic molecules targeting EGFR, irrespective of the specific oncogenic mechanism of the treated cell. While not selective towards their direct targets, these therapeutics can exert selective detrimental effects on cancer cells. Recently, Xu et al. from Deng’s lab have introduced and characterized the first-in-class small molecule KRAS agonist that activates both the wild-type and mutant KRAS molecules and has shown selective effects towards KRAS mutant lung cancer in-vitro and in-vivo57. Another tactic involves directly targeting the ERK1/2 proteins. Exciting research in Sugiura’s lab and colleagues has explored ERKp agonists and lead compounds that show selective activity against ERK-associated cancer cell lines compared to normal cells58,59,60. Unsupervised screens involving perturbations to activate different activating effectors of the ERKp can be applied to find the most effective targets.
P2-Mutation-specific targeting
Targeting protooncogenes like BRAFV600, EGFRL858R, or KRASG12V with mutation-specific agonists can elevate basal ERK activity beyond the oncogenic window in cancer cells, leading to events like apoptosis or senescence. These agonists should ideally be selective (if they ever could be) against oncogenic variants of the targeted- versus the wild-type proteins, aligning with Paul Ehrlich’s magic bullet concept. It deserves to be mentioned that as oncogenes, like any other protein, have a threshold of conformational stability; it cannot be ruled out that oncogene agonists might end up destabilizing and further orchestrating protooncogene degradation processes before even exerting the envisioned effects.
P3-Unsupervised target discovery and targeting aided by the variant-specific landscape of mutual exclusivity among BRAF, EGFR, and KRAS oncogenes in human cancer
We discussed how induction of mutually exclusive BRAF, EGFR, and KRAS oncogenes could be detrimental in affected cells. Nature itself is presenting these scenarios to us. While all the hypothesis-driven approaches mentioned above show promise for further investigation, we would like to underscore an unbiased target discovery approach guided by the uncovered variant-specific landscape of mutual exclusivity among BRAF, EGFR, and KRAS oncogenes. Oncogene overdose55, mimicking the co-expression of EGFR and KRAS oncogenes56, paradoxical intervention61, and ERK-Dependent Apoptosis60 as therapeutic models proposed by different research groups share fundamental similarities. Nevertheless, they vary in certain specific details. The recent unveiling of the variant-specific landscape of mutual exclusivity among ERKp oncogenes introduces a new layer of complexity to existing concepts. This newfound complexity offers insights into the hyperactivation of specific signaling pathways that certain cancer cells circumvent to avoid most hostile conditions.
Consequently, replicating these mutually exclusive scenarios could lead to highly effective yet personalized therapies tailored for each oncogenic variant (BRAF, KRAS, and EGFR). Therefore, in the proposed model, the toxic effects of two oncogenes’ joint expression would be limited to specific variants, or at least more likely in those mutually exclusive scenarios. Consequently, in our proposed model, both target (Fig. 5b) and co-target discovery (Fig. 6) can be a matter of precision. Considering this, our model suggests that the initial point of target discovery and potentially resulting therapy, for instance, for the BRAFG469V variant, may differ from that of BRAFV600E. Unbiased approaches, such as bulk RNA sequencing and single-cell RNA sequencing, can help explore differentially expressed genes and altered pathways resulting from the induction of mutually exclusive BRAF, EGFR, and KRAS oncogenes. Genetic screens such as CRISPR screens, especially dual LOF and GOF screens (i.e., CRISPRi and CRISPRa), may identify the signaling nodes that govern the response and dictate cell fate when mutually exclusive oncogenes are co-induced. These screens are intended to elucidate targeting approaches that phenotypically replicate the co-induction of two synthetically lethal and mutually exclusive gene mutations. Merely Droup-out screens like KO screens may fall short in identifying negative regulators of the ERKp as potential signaling nodes.
Of note, this approach can ideally be tailored to each tumor. For example, for target discovery of cancer cells with BRAFV600E, one can refer to the variant-specific landscape of mutual exclusivity database207 and find the most mutually exclusive scenario with BRAFV600E, and as such, design a model with inducible expression of KRASG12D or EGFRL858R in BRAFV600E background. In a suppressor CRISPR screen, those genes whose activation or inhibition will lead to the rescue of senescence or synthetic lethality would be the governing nodes and potential targets whose antagonism (when activation rescues) or agonism (when inhibition rescues) replicate the synthetically lethal co-induction of these mutually exclusive oncogenes (Fig. 5b). This approach widens the targeting possibilities and may fit both the Goldilocks principle and the magic bullet concept. Besides, this approach may fit the concept of precision oncology.
It is crucial to emphasize that the RTK/RAS/RAF effectors, the inhibitors designed to target them, and their regulators can demonstrate functions and interactions independently of one another, their kinase activity, or even the ERKp itself211,212,213. Further investigation exploring the RTK/RAS/RAF agonism as a therapeutic approach must address whether these functions can be leveraged to enhance oncogenic activity beyond tolerable levels for potential therapeutic benefits or, conversely, whether they might counteract such a strategy. Therefore, any proposed approach to exploit the susceptibility of ERK-associated cancers to hyperactivation of these effectors should consider this notion. If the desired detrimental effect would be ERK-independent, the P1 might fail to address it.
Widespread and highly active oncogenic variants (gene 1), in mutually exclusive relationships with other variants (gene 2), align with the third approach for target discovery (Fig. 5a, b: P3). Conversely, cells harboring oncogenes with co-occurring tendencies206 (variants of gene 1 and gene 2) may align with P1 and P2 (Fig. 5a).
Combined targeting
Cancer targeting resembles a time loop: as monotherapies are developed, resistance emerges, driving the search for more effective combinatorial treatments that not only offer a stronger initial impact but may also delay the development of further resistance. Therefore, it would be wishful thinking to predict that P1-3 would be exempt from the rise of resistance. As such, concurrent, sequential, or intermittent combinatorial treatments can be explored depending on the ultimate phenotypes exerted by putative therapeutics. Based on the evidence revisited in this writing, opportunities for combinatorial treatments with chemotherapeutics can be studied, whether as double-punch or one-two-punch with senolytics. Suppose ERK agonism would lead to immune evading phenotypes. In that case, a one-two-punch model can be explored to address whether, after ERK agonism, the remaining tumor could be sensitized to immune checkpoint inhibitors. The impact of ERK agonism on cancer stem cells and in-parallel or subsequent sensitivity of this subpopulation to chemotherapy or stem cell-targeted therapies is also an avenue worthy of exploration. Notably, P1-P3 hold promise to be explored in combination with therapeutics targeting the RS tolerance.
For co-target discovery, we propose that the initial conditions and ongoing perturbations will be pre-determined using one of the P1-P3, as we discussed for single-target discovery (Figs. 5a, b and 6). In this context, dual loss- and GOF screens (e.g., CRISPR screens) will prove valuable in identifying co-targets when ERK-associated cancer cells undergo the pre-determined ERK-activating perturbation (P1-P3). This approach aims to uncover synthetically lethal co-perturbations (SP) with P1-3, enhancing the detrimental effects to the levels warranting higher fitness threshold191 and the emergence of tumor-suppressive drug resistance mechanisms62. In this respect, our proposed model differs from the paradoxical intervention model, where co-perturbations are identified within the context of hypothesis-driven constant perturbation, such as stress-inducing agents61,62. Consequently, we envision unbiased target discovery for both the discovery of the ERK-activating signaling nodes that govern the detrimental effects of excessive ERKp activation and the subsequent co-target discovery phase (P3 combined with SP, see Fig. 6). Therefore, our model may precisely recapitulate the most hostile conditions cancer cells are avoiding, which are the co-induction of specific oncogenic scenarios.
Conclusions
Despite ample evidence that cancer cells are sensitive to excessive RTK/RAS/RAF pathway activity, this approach has not been widely explored. Perhaps years of relentless efforts to develop efficient ERK inhibitory treatments have established a psychological barrier within us, the scientific community, which, if not overcome, could become a dogma.
Learning from the past, we humbly recommend that all the proposed strategies in this writing undergo unbiased preclinical exploration without any premature preference for a specific strategy over the others.
Despite the primary focus of this writing being exclusively on the ERKp, the mutual exclusivity of cancer-related genetic events offers a wealth of information regarding unexplored synthetic lethality scenarios. Harnessing these scenarios for therapeutic purposes could open new horizons in targeting cancer-related vulnerabilities. Apart from targeting oncogenes, this approach can also encompass events related to tumor suppressors, such as p53, especially when cancer cells remain sensitive to restoring the related functions. Exploring these approaches and identifying targets demands a collaborative effort involving collective benchwork and shared intellectual contributions.
References
Ehrlich, P. The collected papers of Paul Ehrlich, including a complete bibliography/comp. and ed. by F. Himmelweit, with the assistance of the late M. Marquardt, under the editorial direction of Sir H. Dale. (Pergamon Press, 1956).
Valent, P. et al. Paul Ehrlich (1854-1915) and his contributions to the foundation and birth of translational medicine. J. Innate Immun. 8, 111–120 (2016).
Schweitzer, H. Ehrlich’s chemotherapy–a new science. Science 32, 809–823 (1910).
Bosch, F. & Rosich, L. The contributions of Paul Ehrlich to pharmacology: a tribute on the occasion of the centenary of his nobel prize. Pharmacology 82, 171–179 (2008).
Gilman, A. & Philips, F. S. The biological actions and therapeutic applications of the B-chloroethyl amines and sulfides. Science 103, 409–415 (1946).
Gilman, A. Therapeutic applications of chemical warfare agents. Fed. Proc. 5, 285–292 (1946).
Goodman, L. S. & Wintrobe, M. M. Nitrogen mustard therapy; use of methyl-bis (beta-chloroethyl) amine hydrochloride and tris (beta-chloroethyl) amine hydrochloride for Hodgkin’s disease, lymphosarcoma, leukemia and certain allied and miscellaneous disorders. J. Am. Med. Assoc. 132, 126–132 (1946).
Amjad, M. T., Chidharla, A. & Kasi, A. Cancer chemotherapy. In: StatPearls (2022).
Miller, K. D. et al. Cancer treatment and survivorship statistics, 2022. CA. Cancer J. Clin. 72, 409–436 (2022).
Anand, U. et al. Cancer chemotherapy and beyond: current status, drug candidates, associated risks and progress in targeted therapeutics. Genes Dis. 10, 1367–1401 (2023).
Langreth, R. & Waldholz, M. New era of personalized medicine: targeting drugs for each unique genetic profile. Oncologist 4, 426–427 (1999).
Waarts, M. R., Stonestrom, A. J., Park, Y. C. & Levine, R. L. Targeting mutations in cancer. J. Clin. Invest. 132, e154943 (2022).
Blass, E. & Ott, P. A. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat. Rev. Clin. Oncol. 18, 215–229 (2021).
Druker, B. J. et al. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr–Abl positive cells. Nat. Med. 2, 561–566 (1996).
Ottmann, O. G. et al. A phase 2 study of imatinib in patients with relapsed or refractory Philadelphia chromosome-positive acute lymphoid leukemias. Blood 100, 1965–1971 (2002).
Sawyers, C. L. et al. Imatinib induces hematologic and cytogenetic responses in patients with chronic myelogenous leukemia in myeloid blast crisis: results of a phase II study. Blood 99, 3530–3539 (2002).
Weinstein, I. B. & Joe, A. K. Mechanisms of disease: oncogene addiction–a rationale for molecular targeting in cancer therapy. Nat. Clin. Pract. Oncol. 3, 448–457 (2006).
Weinstein, I. B. Cancer. Addiction to oncogenes–the Achilles heal of cancer. Science 297, 63–64 (2002).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Sjöblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Weinstein, I. B. Disorders in cell circuitry during multistage carcinogenesis: the role of homeostasis. Carcinogenesis 21, 857–864 (2000).
Weinstein, I. B. et al. Disorders in cell circuitry associated with multistage carcinogenesis: exploitable targets for cancer prevention and therapy. Clin. Cancer Res. 3, 2696–2702 (1997).
Weinstein, I. B. & Joe, A. Oncogene addiction. Cancer Res. 68, 3077–80 (2008). discussion 3080.
Robichaux, J. P. et al. Structure-based classification predicts drug response in EGFR-mutant NSCLC. Nature 597, 732–737 (2021).
Cooper, A. J., Sequist, L. V. & Lin, J. J. Third-generation EGFR and ALK inhibitors: mechanisms of resistance and management. Nat. Rev. Clin. Oncol. 19, 499–514 (2022).
Dankner, M., Rose, A. A. N. N., Rajkumar, S., Siegel, P. M. & Watson, I. R. Classifying BRAF alterations in cancer: new rational therapeutic strategies for actionable mutations. Oncogene 37, 3183–3199 (2018).
Sharma, S. V., Fischbach, M. A., Haber, D. A. & Settleman, J. ‘Oncogenic shock’: explaining oncogene addiction through differential signal attenuation. Clin. Cancer Res. 12, 4392s–4395s (2006).
Moore, A. R., Rosenberg, S. C., McCormick, F. & Malek, S. RAS-targeted therapies: is the undruggable drugged? Nat. Rev. Drug Discov. 19, 533–552 (2020).
De Greve, J. & Giron, P. Targeting the tyrosine kinase inhibitor-resistant mutant EGFR pathway in lung cancer without targeting EGFR? Transl. Lung Cancer Res. 9, 1–3 (2020).
Proietti, I. et al. Mechanisms of acquired BRAF inhibitor resistance in melanoma: a systematic review. Cancers (Basel). 12, 2801 (2020).
Spagnolo, F., Ghiorzo, P. & Queirolo, P. Overcoming resistance to BRAF inhibition in BRAF-mutated metastatic melanoma. Oncotarget 5, 10206–10221 (2014).
Tanaka, N. et al. Clinical acquired resistance to KRAS(G12C) inhibition through a novel KRAS switch-II pocket mutation and polyclonal alterations converging on RAS-MAPK reactivation. Cancer Discov. 11, 1913–1922 (2021).
Awad, M. M. et al. Acquired resistance to KRAS(G12C) inhibition in cancer. N. Engl. J. Med. 384, 2382–2393 (2021).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Morris, L. G. T. & Chan, T. A. Therapeutic targeting of tumor suppressor genes. Cancer 121, 1357–1368 (2015).
Hassin, O. & Oren, M. Drugging p53 in cancer: one protein, many targets. Nat. Rev. Drug Discov. https://doi.org/10.1038/s41573-022-00571-8 (2022).
Moaven, O., Mangieri, C. W., Stauffer, J. A., Anastasiadis, P. Z. & Borad, M. J. Evolving role of oncolytic virotherapy: challenges and prospects in clinical practice. JCO Precis. Oncol. 432–441 (2021) https://doi.org/10.1200/PO.20.00395.
Bressy, C., Hastie, E. & Grdzelishvili, V. Z. Combining oncolytic virotherapy with p53 tumor suppressor gene therapy. Mol. Ther. Oncolytics 5, 20–40 (2017).
Araki, H. et al. Oncolytic virus-mediated p53 overexpression promotes immunogenic cell death and efficacy of PD-1 blockade in pancreatic cancer. Mol. Ther. Oncolytics 27, 3–13 (2022).
Tian, Y., Xie, D. & Yang, L. Engineering strategies to enhance oncolytic viruses in cancer immunotherapy. Signal Transduct. Target. Ther. 7, 117 (2022).
Dobzhansky, T. Genetics of natural populations; recombination and variability in populations of Drosophila pseudoobscura. Genetics 31, 269–290 (1946).
Kaelin, W. G. Synthetic lethality: a framework for the development of wiser cancer therapeutics. Genome Med. 1, 99 (2009).
Unni, A. M., Lockwood, W. W., Zejnullahu, K., Lee-Lin, S.-Q. & Varmus, H. Evidence that synthetic lethality underlies the mutual exclusivity of oncogenic KRAS and EGFR mutations in lung adenocarcinoma. Elife 4, e06907 (2015).
Beijersbergen, R. L., Wessels, L. F. A. & Bernards, R. Synthetic lethality in cancer therapeutics. Annu. Rev. Cancer Biol. 1, 141–161 (2017).
Nijman, S. M. B. Synthetic lethality: general principles, utility and detection using genetic screens in human cells. FEBS Lett. 585, 1–6 (2011).
Tung, N. & Garber, J. E. PARP inhibition in breast cancer: progress made and future hopes. NPJ Breast Cancer 8, 47 (2022).
Wang, L., Lankhorst, L. & Bernards, R. Exploiting senescence for the treatment of cancer. Nat. Rev. Cancer 22, 340–355 (2022).
Tan, S., Li, D. & Zhu, X. Cancer immunotherapy: pros, cons and beyond. Biomed. Pharmacother. 124, 109821 (2020).
Hamdan, F. & Cerullo, V. Cancer immunotherapies: a hope for the uncurable? Front. Mol. Med. 14, 73 (2023).
Waldman, A. D., Fritz, J. M. & Lenardo, M. J. A guide to cancer immunotherapy: from T cell basic science to clinical practice. Nat. Rev. Immunol. 20, 651–668 (2020).
Atkins, M. B. et al. Combination dabrafenib and trametinib versus combination nivolumab and ipilimumab for patients with advanced BRAF-mutant melanoma: the DREAMseq trial—ECOG-ACRIN EA6134. J. Clin. Oncol. https://doi.org/10.1200/JCO.22.01763 (2022).
Wang, S., Xie, K. & Liu, T. Cancer immunotherapies: from efficacy to resistance mechanisms - not only checkpoint matters. Front. Immunol. 12, 690112 (2021).
Cisowski, J., Sayin, V. I., Liu, M., Karlsson, C. & Bergo, M. O. Oncogene-induced senescence underlies the mutual exclusive nature of oncogenic KRAS and BRAF. Oncogene 35, 1328–1333 (2016).
Mure, E., Library, T. P. & Staff, T. P. L. The story of the three bears: metrically related, with illustrations locating it at cecil lodge, in September 1831. (Toronto Public Library, 2010).
Dipak Amin, A., Rajan, S., Groysman, M. J., Pongtornpipat, P. & Schatz, J. H. Oncogene overdose: too much of a bad thing for oncogene-addicted cancer cells. Biomark. Cancer 7, 7–2 (2015).
Varmus, H., Unni, A. M. & Lockwood, W. W. How cancer genomics drives cancer biology: does synthetic lethality explain mutually exclusive oncogenic mutations? Cold Spring Harb. Symp. Quant. Biol. 81, 247–255 (2016).
Xu, K. et al. Small molecule KRAS agonist for mutant KRAS cancer therapy. Mol. Cancer 18, 85 (2019).
Satoh, R. et al. Discovery of new benzhydrol biscarbonate esters as potent and selective apoptosis inducers of human melanomas bearing the activated ERK pathway: SAR studies on an ERK MAPK signaling modulator, ACA-28. Bioorg. Chem. 103, 104137 (2020).
Satoh, R. et al. Identification of ACA-28, a 1’-acetoxychavicol acetate analogue compound, as a novel modulator of ERK MAPK signaling, which preferentially kills human melanoma cells. Genes Cells 22, 608–618 (2017).
Sugiura, R., Satoh, R. & Takasaki, T. ERK: a double-edged sword in cancer. ERK-dependent apoptosis as a potential therapeutic strategy for cancer. Cells 10, 2509 (2021).
Dias, M. H. & Bernards, R. Playing cancer at its own game: activating mitogenic signaling as a paradoxical intervention. Mol. Oncol. 15, 1975–1985 (2021).
Dias, M. H. et al. Paradoxical activation of oncogenic signaling as a cancer treatment strategy. bioRxiv, https://doi.org/10.1101/2023.02.06.527335 (2023).
Roskoski, R. J. The ErbB/HER family of protein-tyrosine kinases and cancer. Pharmacol. Res. 79, 34–74 (2014).
Roskoski, R. ERK1/2 MAP kinases: Structure, function, and regulation. Pharmacol. Res. 66, 105–143 (2012).
Ullah, R., Yin, Q., Snell, A. H. & Wan, L. RAF-MEK-ERK pathway in cancer evolution and treatment. Semin. Cancer Biol. 85, 123–154 (2022).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170, 17–33 (2017).
Zarrin, A. A., Bao, K., Lupardus, P. & Vucic, D. Kinase inhibition in autoimmunity and inflammation. Nat. Rev. Drug Discov. 20, 39–63 (2021).
Rai, S. N. et al. The Role of PI3K/Akt and ERK in neurodegenerative disorders. Neurotox. Res. 35, 775–795 (2019).
Verschelden, G. et al. Significant response to dabrafenib in a patient with Erdheim–Chester disease with BRAFV600E mutation. Polish. Arch. Intern. Med. 128, 386–388 (2018).
Wee, P. & Wang, Z. Epidermal growth factor receptor cell proliferation signaling pathways. Cancers (Basel). 9, 52 (2017).
Du, Z. & Lovly, C. M. Mechanisms of receptor tyrosine kinase activation in cancer. Mol. Cancer 17, 58 (2018).
Santos, E. & Nebreda, A. R. Structural and functional properties of ras proteins. FASEB J. 3, 2151–2163 (1989).
Johnson, C., Burkhart, D. L. & Haigis, K. M. Classification of KRAS-activating mutations and the implications for therapeutic intervention. Cancer Discov. 12, 913–923 (2022).
Nussinov, R., Tsai, C.-J. & Jang, H. Does ras activate Raf and PI3K allosterically? Front. Oncol. 9, 1231 (2019).
Ünal, E. B., Uhlitz, F. & Blüthgen, N. A compendium of ERK targets. FEBS Lett. 591, 2607–2615 (2017).
Lee, S., Rauch, J. & Kolch, W. Targeting MAPK signaling in cancer: mechanisms of drug resistance and sensitivity. Int. J. Mol. Sci. 21, 1102 (2020).
Sinkala, M., Nkhoma, P., Mulder, N. & Martin, D. P. Integrated molecular characterisation of the MAPK pathways in human cancers reveals pharmacologically vulnerable mutations and gene dependencies. Commun. Biol. 4, 9 (2021).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome Atlas. Cell 173, 321–337.e10 (2018).
Smorodinsky-Atias, K., Soudah, N. & Engelberg, D. Mutations that confer drug-resistance, oncogenicity and intrinsic activity on the ERK MAP kinases-current state of the art. Cells 9, 129 (2020).
Jha, S. et al. Dissecting therapeutic resistance to ERK inhibition. Mol. Cancer Ther. 15, 548–559 (2016).
Jaiswal, B. S. et al. ERK mutations and amplification confer resistance to ERK-inhibitor therapy. Clin. Cancer Res. 24, 4044–4055 (2018).
Sang, D. et al. Ancestral reconstruction reveals mechanisms of ERK regulatory evolution. Elife 8, e38805 (2019).
Taylor, C. A. 4th et al. Functional divergence caused by mutations in an energetic hotspot in ERK2. Proc. Natl Acad. Sci. USA 116, 15514–15523 (2019).
Levin-Salomon, V., Kogan, K., Ahn, N. G., Livnah, O. & Engelberg, D. Isolation of intrinsically active (MEK-independent) variants of the ERK family of mitogen-activated protein (MAP) kinases. J. Biol. Chem. 283, 34500–34510 (2008).
Kushnir, T. et al. An activating mutation in ERK causes hyperplastic tumors in a scribble mutant tissue in drosophila. Genetics 214, 109–120 (2020).
Smorodinsky-Atias, K. et al. Intrinsically active variants of Erk oncogenically transform cells and disclose unexpected autophosphorylation capability that is independent of TEY phosphorylation. Mol. Biol. Cell 27, 1026–1039 (2016).
Soudah, N. et al. A conserved arginine within the αC-helix of Erk1/2 is a latch of autoactivation and of oncogenic capabilities. J. Biol. Chem. 299, 105072 (2023).
Goetz, E. M., Ghandi, M., Treacy, D. J., Wagle, N. & Garraway, L. A. ERK mutations confer resistance to mitogen-activated protein kinase pathway inhibitors. Cancer Res. 74, 7079–7089 (2014).
Leung, G. P. et al. Hyperactivation of MAPK signaling is deleterious to RAS/RAF-mutant melanoma. Mol. Cancer Res. 17, 199–211 (2019).
Brenan, L. et al. Phenotypic characterization of a comprehensive set of MAPK1/ERK2 missense mutants. Cell Rep. 17, 1171–1183 (2016).
Noeparast, A. et al. CRAF mutations in lung cancer can be oncogenic and predict sensitivity to combined type II RAF and MEK inhibition. Oncogene 38, 5933–5941 (2019).
Riaud, M. et al. The role of CRAF in cancer progression: from molecular mechanisms to precision therapies. Nat. Rev. Cancer 24, 105–122 (2024).
Emuss, V., Garnett, M., Mason, C. & Marais, R. Mutations of C-RAF are rare in human cancer because C-RAF has a low basal kinase activity compared with B-RAF. Cancer Res. 65, 9719 (2005).
Yang, L., Zheng, L., Chng, W. J. & Ding, J. L. Comprehensive analysis of ERK1/2 substrates for potential combination immunotherapies. Trends Pharmacol. Sci. 40, 897–910 (2019).
Prasad, M. et al. MEK1/2 inhibition transiently alters the tumor immune microenvironment to enhance immunotherapy efficacy against head and neck cancer. J. Immunother. Cancer 10, e003917 (2022).
Coelho, M. A. et al. Oncogenic RAS signaling promotes tumor immunoresistance by stabilizing PD-L1 mRNA. Immunity 47, 1083–1099.e6 (2017).
Cagnol, S. & Chambard, J. C. ERK and cell death: mechanisms of ERK-induced cell death - apoptosis, autophagy and senescence. FEBS J. 277, 2–21 (2010).
Wu, P.-K., Becker, A. & Park, J.-I. Growth inhibitory signaling of the Raf/MEK/ERK pathway. Int. J. Mol. Sci. 21, 5436 (2020).
Yue, J. & López, J. M. Understanding MAPK signaling pathways in apoptosis. Int. J. Mol. Sci. 21, 2346 (2020).
Yuan, Y. et al. Activation of ERK-Drp1 signaling promotes hypoxia-induced Aβ accumulation by upregulating mitochondrial fission and BACE1 activity. FEBS Open Bio 11, 2740–2755 (2021).
He, K. & Aizenman, E. ERK signaling leads to mitochondrial dysfunction in extracellular zinc-induced neurotoxicity. J. Neurochem. 114, 452–461 (2010).
Cook, S. J., Stuart, K., Gilley, R. & Sale, M. J. Control of cell death and mitochondrial fission by ERK1/2 MAP kinase signalling. FEBS J. 284, 4177–4195 (2017).
Rezatabar, S. et al. RAS/MAPK signaling functions in oxidative stress, DNA damage response and cancer progression. J. Cell. Physiol. 234, 14951–14965 (2019).
Luo, Z. et al. Hypoxia signaling in human health and diseases: implications and prospects for therapeutics. Signal Transduct. Target. Ther. 7, 218 (2022).
Zhang, J. et al. ROS and ROS-mediated cellular signaling. Oxid. Med. Cell. Longev. 2016, 4350965 (2016).
Papa, S., Choy, P. M. & Bubici, C. The ERK and JNK pathways in the regulation of metabolic reprogramming. Oncogene 38, 2223–2240 (2019).
McNeal, A. S. et al. BRAF\textsuperscript{V600E} induces reversible mitotic arrest in human melanocytes via microRNA-mediated suppression of AURKB. Elife 10, e70385 (2021).
Michaloglou, C. et al. BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature 436, 720–724 (2005).
Hong, S.-K. K., Wu, P.-K. K. & Park, J.-I. I. A cellular threshold for active ERK1/2 levels determines Raf/MEK/ERK-mediated growth arrest versus death responses. Cell. Signal. 42, 11–20 (2018).
Wu, P.-K. et al. A mortalin/HSPA9-mediated switch in tumor-suppressive signaling of Raf/MEK/extracellular signal-regulated kinase. Mol. Cell. Biol. 33, 4051–4067 (2013).
Arthan, D., Hong, S.-K. & Park, J.-I. Leukemia inhibitory factor can mediate Ras/Raf/MEK/ERK-induced growth inhibitory signaling in medullary thyroid cancer cells. Cancer Lett. 297, 31–41 (2010).
Hong, S.-K., Kim, J.-H., Lin, M.-F. & Park, J.-I. The Raf/MEK/extracellular signal-regulated kinase 1/2 pathway can mediate growth inhibitory and differentiation signaling via androgen receptor downregulation in prostate cancer cells. Exp. Cell Res. 317, 2671–2682 (2011).
Hong, S.-K., Yoon, S., Moelling, C., Arthan, D. & Park, J.-I. Noncatalytic function of ERK1/2 can promote Raf/MEK/ERK-mediated growth arrest signaling. J. Biol. Chem. 284, 33006–33018 (2009).
Blagosklonny, M. V., Schulte, T., Nguyen, P., Trepel, J. & Neckers, L. M. Taxol-induced apoptosis and phosphorylation of Bcl-2 protein involves c-Raf-1 and represents a novel c-Raf-1 signal transduction pathway. Cancer Res. 56, 1851–1854 (1996).
Xiao, D. & Singh, S. V. Phenethyl isothiocyanate-induced apoptosis in p53-deficient PC-3 human prostate cancer cell line is mediated by extracellular signal-regulated kinases. Cancer Res. 62, 3615–3619 (2002).
Tang, D. et al. ERK activation mediates cell cycle arrest and apoptosis after DNA damage independently of p53. J. Biol. Chem. 277, 12710–12717 (2002).
Hsu, Y., Kuo, P., Lin, L. & Lin, C. Asiatic acid, a triterpene, induces apoptosis and cell cycle arrest through activation of extracellular signal-regulated kinase and p38 mitogen-activated protein kinase pathways in human breast cancer cells. J. Pharmacol. Exp. 313, 333–344 (2005).
Calcabrini, A. et al. Inhibition of proliferation and induction of apoptosis in human breast cancer cells by lauryl gallate. Carcinogenesis 27, 1699–1712 (2006).
Ciccarelli, C. et al. p21WAF1 expression induced by MEK/ERK pathway activation or inhibition correlates with growth arrest, myogenic differentiation and onco-phenotype reversal in rhabdomyosarcoma cells. Mol. Cancer 4, 41 (2005).
Guégan, J. P., Ezan, F., Gailhouste, L., Langouët, S. & Baffet, G. MEK1/2 overactivation can promote growth arrest by mediating ERK1/2-dependent phosphorylation of p70S6K. J. Cell. Physiol. 229, 903–915 (2014).
Lane, D. P. p53, guardian of the genome. Nature 358, 15–16 (1992).
Eischen, C. M. Genome stability requires p53. Cold Spring Harb. Perspect. Med. 6, a026096 (2016).
Bieging, K. T., Mello, S. S. & Attardi, L. D. Unravelling mechanisms of p53-mediated tumour suppression. Nat. Rev. Cancer 14, 359–70 (2014).
Vaseva, A. V. et al. P53 opens the mitochondrial permeability transition pore to trigger necrosis. Cell 149, 1536–1548 (2012).
Marchenko, N. D. & Moll, U. M. Mitochondrial death functions of p53. Mol. Cell. Oncol. 1, e955995 (2014).
Timofeev, O. et al. Residual apoptotic activity of a tumorigenic p53 mutant improves cancer therapy responses. EMBO J. https://doi.org/10.15252/embj.2019102096 (2019).
Jiang, P. et al. P53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat. Cell Biol. https://doi.org/10.1038/ncb2172 (2011).
Zhao, Y. et al. p53 translocation to mitochondria precedes its nuclear translocation and targets mitochondrial oxidative defense protein-manganese superoxide dismutase. Cancer Res. https://doi.org/10.1158/0008-5472.CAN-04-3835 (2005).
Tasdemir, E. et al. Regulation of autophagy by cytoplasmic p53. Nat. Cell Biol. 10, 676–687 (2008).
Hampp, S. et al. DNA damage tolerance pathway involving DNA polymerase ι and the tumor suppressor p53 regulates DNA replication fork progression. Proc. Natl. Acad. Sci. USA. https://doi.org/10.1073/pnas.1605828113 (2016).
Linke, S. P. et al. p53 interacts with hRAD51 and hRAD54, and directly modulates homologous recombination. Cancer Res. 63, 2596–2605 (2003).
Serrano, M., Lin, A. W., McCurrach, M. E., Beach, D. & Lowe, S. W. Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 88, 593–602 (1997).
Braig, M. et al. Oncogene-induced senescence as an initial barrier in lymphoma development. Nature 436, 660–665 (2005).
Chen, Z. et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 436, 725–730 (2005).
Collado, M. et al. Tumour biology: senescence in premalignant tumours. Nature 436, 642 (2005).
Courtois-Cox, S. et al. A negative feedback signaling network underlies oncogene-induced senescence. Cancer Cell 10, 459–472 (2006).
Brown, L. & Benchimol, S. The involvement of MAPK signaling pathways in determining the cellular response to p53 activation: cell cycle arrest or apoptosis. J. Biol. Chem. 281, 3832–3840 (2006).
Bates, S. et al. p14ARF links the tumour suppressors RB and p53. Nature 395, 124–125 (1998).
Persons, D. L., Yazlovitskaya, E. M. & Pelling, J. C. Effect of extracellular signal-regulated kinase on p53 accumulation in response to cisplatin. J. Biol. Chem. 275, 35778–35785 (2000).
Okuno, T., Matsuoka, M., Sumizawa, T. & Igisu, H. Involvement of the extracellular signal-regulated protein kinase pathway in phosphorylation of p53 protein and exerting cytotoxicity in human neuroblastoma cells (SH-SY5Y) exposed to acrylamide. Arch. Toxicol. 80, 146–153 (2006).
She, Q. B., Chen, N. & Dong, Z. ERKs and p38 kinase phosphorylate p53 protein at serine 15 in response to UV radiation. J. Biol. Chem. 275, 20444–20449 (2000).
Yeh, P. Y. et al. Phosphorylation of p53 on Thr55 by ERK2 is necessary for doxorubicin-induced p53 activation and cell death. Oncogene 23, 3580–3588 (2004).
Collado, M. & Serrano, M. Senescence in tumours: evidence from mice and humans. Nat. Rev. Cancer. https://doi.org/10.1038/nrc2772 (2010).
Astle, M. V. et al. AKT induces senescence in human cells via mTORC1 and p53 in the absence of DNA damage: implications for targeting mTOR during malignancy. Oncogene 31, 1949–1962 (2012).
Lowe, S. W., Cepero, E. & Evan, G. Intrinsic tumour suppression. Nature 432, 307–15 (2004).
Drosten, M. et al. Loss of p53 induces cell proliferation via Ras-independent activation of the Raf/Mek/Erk signaling pathway. Proc. Natl Acad. Sci. USA 111, 15155–15160 (2014).
Karkhanis, M. & Park, J.-I. Sp1 regulates Raf/MEK/ERK-induced p21(CIP1) transcription in TP53-mutated cancer cells. Cell Signal. 27, 479–486 (2015).
De, S., Campbell, C., Venkitaraman, A. R. & Esposito, A. Pulsatile MAPK Signaling Modulates p53 activity to control cell fate decisions at the G2 checkpoint for DNA damage. Cell Rep. 30, 2083–2093.e5 (2020).
Marchetti, A. et al. p53 can inhibit cell proliferation through caspase-mediated cleavage of ERK2/MAPK. Cell Death Differ. 11, 596–607 (2004).
Sabapathy, K. & Lane, D. P. Therapeutic targeting of p53: all mutants are equal, but some mutants are more equal than others. Nat. Rev. Clin. Oncol. https://doi.org/10.1038/nrclinonc.2017.151 (2017).
Donehower, L. A. et al. Integrated analysis of TP53 gene and pathway alterations in the cancer genome atlas. Cell Rep. 28, 1370–1384.e5 (2019).
Campbell, P. J. et al. Pan-cancer analysis of whole genomes. Nature. https://doi.org/10.1038/s41586-020-1969-6 (2020).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature. https://doi.org/10.1038/s41586-019-1907-7 (2020).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Leroy, B., Anderson, M. & Soussi, T. TP53 mutations in human cancer: database reassessment and prospects for the next decade. Hum. Mutat. https://doi.org/10.1002/humu.22552 (2014).
Steele, C. D. et al. Signatures of copy number alterations in human cancer. Nature 606, 984–991 (2022).
Rivlin, N., Brosh, R., Oren, M. & Rotter, V. Mutations in the p53 tumor suppressor gene: important milestones at the various steps of tumorigenesis. Genes Cancer 2, 466–474 (2011).
Chen, X. et al. Mutant p53 in cancer: from molecular mechanism to therapeutic modulation. Cell Death Dis. 13, 1–14 (2022).
Hainaut, P. & Pfeifer, G. P. Somatic TP53 mutations in the era of genome sequencing. Cold Spring Harb. Perspect. Med. 6, a026179 (2016).
Mantovani, F., Collavin, L. & Del Sal, G. Mutant p53 as a guardian of the cancer cell. Cell Death Differ. https://doi.org/10.1038/s41418-018-0246-9 (2019).
Stiewe, T. & Haran, T. E. How mutations shape p53 interactions with the genome to promote tumorigenesis and drug resistance. Drug Resist. Updat. 38, 27–43 (2018).
Klimovich, B. et al. p53 partial loss-of-function mutations sensitize to chemotherapy. Oncogene 41, 1011–1023 (2022).
Skoulidis, F. & Heymach, J. V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 19, 495–509 (2019).
Klimovich, B. et al. Partial p53 reactivation is sufficient to induce cancer regression. J. Exp. Clin. Cancer Res. 41, 80 (2022).
Kotler, E. et al. A systematic p53 mutation library links differential functional impact to cancer mutation pattern and evolutionary conservation. Mol. Cell 71, 178–190.e8 (2018).
Boettcher, S. et al. A dominant-negative effect drives selection of TP53 missense mutations in myeloid malignancies. Science 365, 599 LP–604 (2019).
Kato, S. et al. Understanding the function–structure and function–mutation relationships of p53 tumor suppressor protein by high-resolution missense mutation analysis. Proc. Natl Acad. Sci. 100, 8424 LP–8429 (2003).
Giacomelli, A. O. et al. Mutational processes shape the landscape of TP53 mutations in human cancer. Nat. Genet. 50, 1381–1387 (2018).
Panagopoulos, A. & Altmeyer, M. The hammer and the dance of cell cycle control. Trends Biochem. Sci. 46, 301–314 (2021).
Matthews, H. K., Bertoli, C. & de Bruin, R. A. M. Cell cycle control in cancer. Nat. Rev. Mol. Cell Biol. 23, 74–88 (2022).
Gaillard, H., García-Muse, T. & Aguilera, A. Replication stress and cancer. Nat. Rev. Cancer 15, 276–289 (2015).
Petropoulos, M., Champeris Tsaniras, S., Taraviras, S. & Lygerou, Z. Replication licensing aberrations, replication stress, and genomic instability. Trends Biochem. Sci. 44, 752–764 (2019).
da Costa, A. A. B. A., Chowdhury, D., Shapiro, G. I., D’Andrea, A. D. & Konstantinopoulos, P. A. Targeting replication stress in cancer therapy. Nat. Rev. Drug Discov. 22, 38–58 (2023).
Bowry, A., Kelly, R. D. W. & Petermann, E. Hypertranscription and replication stress in cancer. Trends Cancer 7, 863–877 (2021).
Kotsantis, P., Petermann, E. & Boulton, S. J. Mechanisms of oncogene-induced replication stress: Jigsaw falling into place. Cancer Discov. 8, 537–555 (2018).
Segeren, H. A., van Liere, E. A., Riemers, F. M., de Bruin, A. & Westendorp, B. Oncogenic RAS sensitizes cells to drug-induced replication stress via transcriptional silencing of P53. Oncogene 41, 2719–2733 (2022).
Schaufler, D. et al. Clonal dynamics of BRAF-driven drug resistance in EGFR-mutant lung cancer. NPJ Precis. Oncol. 5, 102 (2021).
McCubrey, J. A. et al. Roles of the Raf/MEK/ERK pathway in cell growth, malignant transformation and drug resistance. Biochim. Biophys. Acta 1773, 1263–1284 (2007).
Ohashi, K. et al. Lung cancers with acquired resistance to EGFR inhibitors occasionally harbor BRAF gene mutations but lack mutations in KRAS, NRAS, or MEK1. Proc. Natl Acad. Sci. USA 109, E2127–33 (2012).
Aissa, A. F. et al. Single-cell transcriptional changes associated with drug tolerance and response to combination therapies in cancer. Nat. Commun. 12, 1628 (2021).
Awad, M. M. et al. Acquired resistance to KRASG12C inhibition in cancer. N. Engl. J. Med. 384, 2382–2393 (2021).
Xue, Y. et al. An approach to suppress the evolution of resistance in BRAF(V600E)-mutant cancer. Nat. Med. 23, 929–937 (2017).
Tétu, P. et al. Mitogen-activated protein kinase blockade in melanoma: intermittent versus continuous therapy, from preclinical to clinical data. Curr. Opin. Oncol. 33, 127–132 (2021).
Kavran, A. J. et al. Intermittent treatment of BRAF(V600E) melanoma cells delays resistance by adaptive resensitization to drug rechallenge. Proc. Natl Acad. Sci. USA 119, e2113535119 (2022).
Kong, X. et al. Cancer drug addiction is relayed by an ERK2-dependent phenotype switch. Nature 550, 270–274 (2017).
Gutierrez-Prat, N. et al. DUSP4 protects BRAF- and NRAS-mutant melanoma from oncogene overdose through modulation of MITF. Life Sci. Alliance 5, e202101235 (2022).
Stagno, A. et al. Case report: rechallenge with BRAF and MEK inhibitors in metastatic melanoma: a further therapeutic option in salvage setting? Front. Oncol. 11, 645008 (2021).
Matter, A. V., Micaletto, S., Urner-Bloch, U., Dummer, R. & Goldinger, S. M. Long-term response to intermittent binimetinib in patients with NRAS-mutant melanoma. Oncologist 25, e1593–e1597 (2020).
Das Thakur, M. et al. Modelling vemurafenib resistance in melanoma reveals a strategy to forestall drug resistance. Nature 494, 251–255 (2013).
Hong, A. et al. Exploiting drug addiction mechanisms to select against mapki-resistant melanoma. Cancer Discov. 8, 74–93 (2018).
Xue, Y. et al. An approach to suppress the evolution of resistance in BRAFV600E-mutant cancer. Nat. Med. 23, 929–937 (2017).
Holderfield, M., Deuker, M. M., McCormick, F. & McMahon, M. Targeting RAF kinases for cancer therapy: BRAF-mutated melanoma and beyond. Nat. Rev. Cancer 14, 455–467 (2014).
Dooley, A. J., Gupta, A., Bhattacharyya, M. & Middleton, M. R. Intermittent dosing with vemurafenib in BRAF V600E-mutant melanoma: review of a case series. Ther. Adv. Med. Oncol. 6, 262–266 (2014).
Smalley, I. et al. Leveraging transcriptional dynamics to improve BRAF inhibitor responses in melanoma. EBioMedicine 48, 178–190 (2019).
Schreuer, M. et al. Combination of dabrafenib plus trametinib for BRAF and MEK inhibitor pretreated patients with advanced BRAF(V600)-mutant melanoma: an open-label, single arm, dual-centre, phase 2 clinical trial. Lancet Oncol. 18, 464–472 (2017).
Valpione, S. et al. Rechallenge with BRAF-directed treatment in metastatic melanoma: a multi-institutional retrospective study. Eur. J. Cancer 91, 116–124 (2018).
Tietze, J. K. et al. The efficacy of re-challenge with BRAF inhibitors after previous progression to BRAF inhibitors in melanoma: a retrospective multicenter study. Oncotarget 9, 34336–34346 (2018).
Algazi, A. P. et al. Continuous versus intermittent BRAF and MEK inhibition in patients with BRAF-mutated melanoma: a randomized phase 2 trial. Nat. Med. 26, 1564–1568 (2020).
Gonzalez-Cao, M. et al. Intermittent BRAF inhibition in advanced BRAF mutated melanoma results of a phase II randomized trial. Nat. Commun. 12, 7008 (2021).
Petti, C. et al. Coexpression of NRASQ61R and BRAFV600E in human melanoma cells activates senescence and increases susceptibility to cell-mediated cytotoxicity. Cancer Res. 66, 6503–6511 (2006).
Unni, A. M. et al. Hyperactivation of ERK by multiple mechanisms is toxic to RTK-RAS mutation-driven lung adenocarcinoma cells. Elife 7, 1–24 (2018).
Ambrogio, C., Barbacid, M. & Santamaría, D. In vivo oncogenic conflict triggered by co-existing KRAS and EGFR activating mutations in lung adenocarcinoma. Oncogene 36, 2309–2318 (2017).
Sun, C. et al. Reversible and adaptive resistance to BRAF(V600E) inhibition in melanoma. Nature 508, 118–122 (2014).
le Sage, C. et al. Dual direction CRISPR transcriptional regulation screening uncovers gene networks driving drug resistance. Sci. Rep. 7, 17693 (2017).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Vaeyens, F. et al. Variant-specific landscape of mutual exclusivity among BRAF, EGFR, and KRAS oncogenes in human cancer. medRxiv https://doi.org/10.1101/2023.10.21.23297089. (2023).
Zhao, Y. et al. Assessment of RAS sependency for BRAF alterations using cancer genomic databases. JAMA Netw. Open 4, e2035479–e2035479 (2021).
Whittaker, S. R. et al. Combined pan-RAF and MEK inhibition overcomes multiple resistance mechanisms to selective RAF inhibitors. Mol. Cancer Ther. 14, 2700-11 (2015).
Reyes, R. et al. Clinical benefit from BRAF/MEK inhibition in a double non-V600E BRAF mutant lung adenocarcinoma: a case report. Clin. Lung Cancer 20, e219–e223 (2019).
Garutti, M. et al. BRAF and MEK inhibitors and their toxicities: a meta-analysis. Cancers (Basel). 15, 141 (2022).
Maurer, G., Tarkowski, B. & Baccarini, M. Raf kinases in cancer–roles and therapeutic opportunities. Oncogene 30, 3477–3488 (2011).
Dorard, C. et al. RAF1 contributes to cell proliferation and STAT3 activation in colorectal cancer independently of microsatellite and KRAS status. Oncogene 42, 1649–1660 (2023).
Eggermont, C. et al. The EGFR-STYK1-FGF1 axis sustains functional drug tolerance to EGFR inhibitors in EGFR-mutant non-small cell lung cancer. Cell Death Dis. 13, 611 (2022).
Acknowledgements
M.N. is grateful to Dr. Erik Teugels for years of scientific discussion about the idea behind this writing and beyond.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
M.N. conceived and conceptualized the idea for the review and the conceptual framework and authored the manuscript with scientific support, authoring contribution, and intellectual input of O.T., P.G., S.L., and M.P. All authors critically revised the text as well. M.N. and S.L. generated figures.
Corresponding author
Ethics declarations
Competing interests
All authors, including M.N., O.T., P.G., S.L., and M.P., declare no competing (financial or non-financial) interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Timofeev, O., Giron, P., Lawo, S. et al. ERK pathway agonism for cancer therapy: evidence, insights, and a target discovery framework. npj Precis. Onc. 8, 70 (2024). https://doi.org/10.1038/s41698-024-00554-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-024-00554-5
- Springer Nature Limited
This article is cited by
-
Exploring and clinical validation of prognostic significance and therapeutic implications of copper homeostasis-related gene dysregulation in acute myeloid leukemia
Annals of Hematology (2024)
-
HDAC3 genetic and pharmacologic inhibition radiosensitizes fusion positive rhabdomyosarcoma by promoting DNA double-strand breaks
Cell Death Discovery (2024)
-
SHANK3 depletion leads to ERK signalling overdose and cell death in KRAS-mutant cancers
Nature Communications (2024)