Abstract
Tissue-resident memory T cells (TRMs) play a critical role in cancer immunity by offering quick and effective immune responses. However, the cellular heterogeneity of TRMs and their significance in cervical cancer (CC) remain unknown. In this study, we generated and analyzed single-cell RNA sequencing data from 12,945 TRMs (ITGAE+ CD3D+) and 25,627 non-TRMs (ITGAE− CD3D+), derived from 11 CC tissues and 5 normal cervical tissues. We found that TRMs were more immunoreactive than non-TRMs, and TRMs in CC tissues were more activated than those in normal cervical tissues. Six CD8+ TRM subclusters and one CD4+ TRM subcluster were identified. Among them, CXCL13+ CD8+ TRMs were more abundant in CC tissues than in normal cervical tissues, had both cytotoxic and inhibitory features, and were enriched in pathways related to defense responses to the virus. Meanwhile, PLAC8+ CD8+ TRMs were less abundant in CC tissues than in normal cervical tissues but had highly cytotoxic features. The signature gene set scores of both cell subclusters were positively correlated with the overall survival and progression-free survival of patients with CC following radiotherapy. Of note, the association between HLA-E and NKG2A, either alone or in a complex with CD94, was enriched in CXCL13+ CD8+ TRMs interacting with epithelial cells at CC tissues. The in-depth characterization of TRMs heterogeneity in the microenvironment of CC could have important implications for advancing treatment and improving the prognosis of patients with CC.
Similar content being viewed by others
Introduction
Cervical cancer (CC) is the fourth most common malignant tumor in women worldwide and is a leading cause of malignant tumor-related deaths1. Persistent infection with high-risk human papillomavirus (HPV) subtypes, particularly HPV 16 and 18, can alter cervical cells, leading to the development of pre-cancerous lesions and eventually CC2. Conventional cancer therapies, including surgical resection, radiotherapy, and chemotherapy, for CC are currently insufficient. Resistance to therapy causes CC recurrence and metastasis in approximately one-third of patients after treatment completion3,4. In recent years, immunotherapy has brought new hope to cancer patients and is especially changing the treatment landscape of advanced CC5. These immunotherapies include immune checkpoint inhibitors (ICIs), therapeutic vaccines, and adoptive cell transfer therapy, all of which aim to boost anti-tumor immune responses mediated by immune cells, especially T cells6.
Tissue-resident memory T cells (TRMs) are a special subset of T lymphocytes that reside indefinitely in tissues7. The presence of an infected or malignant cell initiates a T-cell-mediated immune response, following which, some T cells, which retain specific information relating to this encounter, survive. When the same infection or tumor cell re-emerges, these so called “memory T cells” can respond more quickly and effectively than their naive counterparts8. Among the various subsets of memory T cells, those residing in tissues and organs are called TRMs and are distinguished by the expression of the integrin CD103 (encoded by ITGAE)9,10. CD103+ T cells have been demonstrated to play a vital role in the immune response against solid tumors, including breast cancer11, lung cancer12, endometrial adenocarcinoma13, and bladder urothelial carcinoma14. CD49a and CD69 are important markers of TRMs but their expressions are not as widespread and consistent across different tissues as CD103. CD69 is a T cell activation marker that can be expressed in other T cell subsets beyond TRMs, including activated effector T cells[15, 16]. CD49a expression might be used to distinguish TRM subsets with different effector functions and putative roles in immunopathology. In skin from patients with vitiligo, CD8+CD49a+ TRMs that constitutively express perforin and GZMB accumulates in the epidermis and dermis. Conversely, CD8+CD49a- TRMs from psoriasis lesions predominantly generate IL-17 responses that promote local inflammation in this skin disease[17]. Therefore, we chose CD103 as the most widely recognized and strongly associated single marker to ensure the reliability of our research and the clarity of the results in this study. Currently, researchers are seeking ways to activate and enhance the efficacy of TRM responses as a potential strategy for cancer immunotherapy. Moreover, CD8+ TRMs express high levels of immune checkpoint molecules and are thought to be early responders to ICIs treatment15. In addition, cancer vaccines, which prime TRMs to specifically target cancer cells, are being developed and have shown objective efficacy in inhibiting the growth of melanoma tumors in mucosal tissues16,17. Furthermore, a novel adoptive T cell therapeutic strategy, which aims to collect TRMs from resected tumor tissues, also brings new challenges and opportunities to the field of cancer therapy18. Thus, the use of TRMs in cancer immunotherapy is a field of active research, and continued studies are necessary to determine their feasibility as an effective cancer treatment.
In recent times, the field of cancer research has been advanced by the emergence of single-cell RNA sequencing (scRNA-seq) technology19. Compared with bulk RNA sequencing, scRNA-seq enables the study of individual cells, providing researchers with a measure of cellular heterogeneity within complex tissues or cell populations. scRNA-seq can identify rare cell types or states, which may otherwise be missed in a bulk RNA analysis20. However, there are currently no scRNA-seq studies deciphering the TRMs landscape of CC. A thorough dissection of the cellular heterogeneity and molecular characteristics of TRMs in the tumor microenvironment (TME) of CC at the single-cell level is crucial for developing novel immunotherapies and improving patient prognosis.
In our study, we analyzed scRNA-seq data from 38,572 T cells, derived from 11 CC and 5 normal cervical tissues. Our data revealed that TRMs demonstrated significantly higher levels of immunoreactivity than non-TRMs; and TRMs within CC tissues were more activated than those in normal cervical tissues. Furthermore, we identified two subclusters of CD8+ TRMs, expressing either CXCL13 or PLAC8, which had important roles in the TME of CC but differed in their abundance, molecular characteristics, cell interactions, and prognostic significance. We believe that these findings will help guide the development of diagnostic and/or therapeutic approaches to improve the prognosis of patients with CC.
Results
Single-cell transcriptomes revealed that TRMs are more immunoreactive than non-TRMs
We obtained scRNA-seq data from 38,572 T cells from 11 CC and 5 normal cervical samples (Fig. 1a). T cells were classified as TRMs (12,945 cells; ITGAE+ CD3D+) or non-TRMs (25,627 cells; ITGAE− CD3D+) on the basis of ITGAE expression (Fig. 1b), consistent with TRMs described in previous researches21,22. In addition, we compared the expression of ITGAE in TRMs, non-TRMs, and T cells in peripheral blood mononuclear cells (PBMC), further validating the specificity of ITGAE for defining TRMs (Supplementary Fig. 1d). Differentially expressed genes (DEGs) analysis revealed that TRMs expressed higher levels of genes associated with tissue residence (TGFB1 and ITGB7), cytotoxicity (GZMB and PRF1), and chemokine production (CXCL13 and CCL5) than non-TRMs (Fig. 1c, d). The gene ontology (GO) term analysis showed that DEGs related to the cell adhesion pathway were enriched in TRMs than non-TRMs, which is associated with the retention of T cells in tissues. In addition, pathways associated with T cell activation, lymphocyte-mediated immunity, and antigen processing and presentation were also enriched in TRMs than non-TRMs (Fig. 1e, f and Supplementary Fig. 1c). Moreover, TRMs were more abundant in CC tissues than in normal cervical tissues (p = 0.0053; Fig. 1g, h). Subsequently, we confirmed that the infiltration degree of TRMs in CC was higher than that in normal cervical tissues by performing immunofluorescent labeling on formalin-fixed and paraffin-embedded (FFPE) samples of CC and normal cervical tissues (p = 0.0113; Fig. 1i). These findings indicate that TRMs are more abundant and immunoreactive than non-TRMs in the TME of CC.
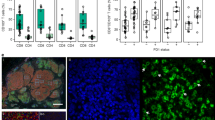
a Overview of the experimental workflow. b t-Distributed Stochastic Neighbor Embedding (tSNE) plots showing subcluster, sample origin, and marker gene expression. The color key shows the gradient of normalized expression. c Volcano plot displaying differentially expressed genes (DEGs) between TRMs (red) and non-TRMs (blue). d Violin plots illustrating the expression of the indicated genes in TRMs (red) and non-TRMs (blue). *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 (two-sided Wilcoxon test). e Gene ontology (GO) term analysis in TRMs. The intensity of color indicates p-value. f Gene Set Enrichment Analysis (GSEA) showing that TRMs were enriched in gene sets associated with the immune response. NES, normalized enrichment score. g Bar plots depicting the proportions of TRMs (red) and non-TRMs (blue) in each sample. h Box plots illustrating the proportions of cell subclusters in normal cervix and cervical cancer (CC) tissues. i Representative immunofluorescent labeling of CD103 (green), CD3 (red), and 4,6-diamidino-2-phenylindole (DAPI, blue) in sections from CC (up) and normal cervical tissues (down). Box plots showing the percentage of CD103+ CD3+ TRMs and CD103- CD3+ non-TRMs in CC and normal cervical tissues based on immunofluorescent labeling results. Scale bar in figure, 80 μm. The p-values were generated using the Wilcoxon test.
Single-cell transcriptome analysis revealed that TRMs are more activated in CC tissues than in normal cervical tissues
Based on the higher infiltration of TRMs in CC tissues than in normal ones, we next explored the transcriptomes of TRMs in CC tissues and in normal cervical tissues (Fig. 2a). We first identified DEGs between TRMs from CC tissues and those from normal cervical tissues (Fig. 2b). Functional gene set analysis revealed that TRMs from CC tissues not only expressed more genes related to cytotoxicity and proliferation, but also expressed higher levels of genes encoding immune checkpoint molecules than TRMs from normal cervical tissues. By contrast, the TRMs from normal cervical tissues had a more naïve-like T cell signature (Fig. 2c). The GO term analysis of DEGs showed that TRMs in CC tissues were enriched in pathways related to T cell activation, antigen processing and presentation of peptide antigen via MHC class II, and lymphocyte-mediated immunity. They were also enriched in pathways associated with defense response to virus (Fig. 2d and Supplementary Fig. 2). The gene set enrichment analysis (GSEA) results confirmed the enrichment of gene sets related to T cell activation, antigen processing and presentation of peptide antigen via MHC class II, as well as the response to type I interferon and defense response to virus (Fig. 2e). These results suggest that TRMs residing in the TME of CC are more activated than those localized to normal cervical tissues; moreover, TRMs may play a crucial role in the anti-viral immune response in the cervix.
a tSNE plots showing distribution of TRMs from CC tissues (blue) and those from normal cervical tissues (pink). b Volcano plot showing DEGs between TRMs from CC tissues and those from normal cervical tissues. c Heatmap showing the expression of functional signature genes, whereby color intensity indicates average gene expression. d GO term analysis in TRMs from CC tissues. The color intensity indicates p-value. e GSEA showing differences in the activation of various immune cell pathways in TRMs from CC tissues and in those from normal cervical tissues.
Single-cell transcriptome analysis of TRMs heterogeneity
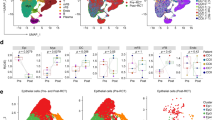
To further dissect the molecular heterogeneity of TRMs, we extracted data from all the TRMs for use in a sub-clustering analysis. We obtained seven TRM subclusters: six CD8+ TRM subclusters, differentiated by CXCL13, PLAC8, ITM2C, STMN1, FCER1G, or XIST expression; and one CD4+ TRM cluster (Fig. 3a). These TRM subclusters exhibited widely divergent gene expression profiles (Fig. 3b and Supplementary Fig. 3a). Among them, CXCL13+ CD8+ TRMs demonstrated elevated expression of genes encoding inhibitory molecules (LAG3, TIGIT, PDCD1, HAVCR2, and CTLA4) and had a higher inhibitory score than other TRM subclusters. Additionally, CXCL13+ CD8+ TRMs also expressed genes encoding cytotoxic molecules (GZMB and PRF1; Fig. 3c and Supplementary Fig. 3b). In addition, PLAC8+ CD8+ TRMs demonstrated elevated expression of genes encoding cytotoxic molecules (GZMA, GNLY, NKG7, and PRF1; Fig. 3c) and had a higher naïve score than other CD8+ TRM subclusters (Supplementary Fig. 3b). The relative abundances of the seven major TRM subclusters in the CC tissues and normal cervical tissues were shown in Fig. 3d. Further comparisons revealed that the proportion of infiltrating CXCL13+ CD8+ TRMs was significantly higher in CC tissues than in normal cervical tissues (p = 0.0022), whereas the proportion of infiltrating PLAC8+ CD8+ TRMs was significantly lower in CC tissues than in normal cervical tissues (p = 0.00092; Fig. 3e).
a tSNE plots showing the TRM subclusters and the expression of marker genes; the color key shows the gradient of normalized expression. Heatmaps displaying the relative expression of the top 3 DEGs (b) and functional signature genes (c) in the seven TRM subclusters; the color intensity reflects the average level of gene expression. d The proportions of the seven TRM subclusters in each tissue and type (CC tissues vs. normal cervical tissues), colored by cell types. e Box plots illustrating the proportions of TRM subclusters in normal cervical (pink) and CC (red) tissues. The p-values were calculated using the paired Wilcoxon test.
CXCL13 + CD8+ TRMs are enriched in CC tissues and express transcripts related to antigen processing and presentation and defense responses to the virus
To determine the unique molecular signature of CXCL13+ CD8+ TRMs, we identified DEGs between CXCL13+ CD8+ TRMs and the other TRM subclusters. The result exhibited that CXCL13+ CD8+ TRMs showed high expression of cytotoxic genes (GZMB, GNLY, and PRF1) and inhibitory genes (HAVCR2, TIGIT, LAG3, CTLA4, and PDCD1) (Fig. 4a, b). The GO term analysis revealed that CXCL13+ CD8+ TRMs were enriched in pathways associated with T cell activation, cytokine production, as well as response to virus and response to type I interferon (Fig. 4c). Moreover, CXCL13+ CD8+ TRMs expressed higher level of genes related to defense response to virus, including IFI6, IFIT3, BST2, MX1, PLSCR1, ISG15, OAS1, STAT1, HERC5, and IRF7, than the other TRM subclusters (Fig. 4d). The GSEA results also confirmed that CXCL13+ CD8+ TRMs were enriched in gene sets associated with defense response to virus and response to type I interferon (Fig. 4e and Supplementary Fig. 4). These findings suggest that CXCL13+ CD8+ TRMs may exert anti-viral effects upon stimulation by type I interferon.
a tSNE plot showing the distribution of CXCL13+ CD8+ TRMs (red) relative to that of the other TRM subclusters (blue). b Volcano plot depicting DEGs between CXCL13+ CD8+ TRMs (red) and the other TRM subclusters (blue). c GO term analysis of CXCL13+ CD8+ TRMs. The color intensity indicates p-value magnitude. d Violin plots illustrating the expression of the indicated genes in CXCL13+ CD8+ TRMs and the other TRM subclusters. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 (two-sided Wilcoxon test). e GSEA showing differences in pathway activity between CXCL13+ CD8+ TRMs and the other TRM subclusters. f Volcano plot depicting the DEGs between CXCL13+ CD8+ TRMs from CC tissues (red) and those from normal cervical tissues (blue). g Violin plots displaying the expression of the indicated genes in CXCL13+ CD8+ TRMs from CC tissues and in those from normal cervical tissues. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 (two-sided Wilcoxon test). h GO term analysis of CXCL13+ CD8+ TRMs from CC tissues; color intensity indicates p-value magnitude. i GSEA results showing differences in the pathway activities of CXCL13+ CD8+ TRMs from CC tissues and those from normal cervical tissues. j Kaplan–Meier survival analysis of patients with CC following radiotherapy from TCGA database, stratified based on high or low signature gene set score of CXCL13+ CD8+ TRMs. The p-values were calculated using the two-sided log-rank test.
Because CXCL13+ CD8+ TRMs were predominantly found in CC tissues, we explored the differences in the signatures of CXCL13+ CD8+ TRMs from CC tissues and normal cervical tissues. DEGs analysis revealed that MHC class II molecule genes (HLA-DRB5 and HLA-DRA) and chemokine-related genes (CCL4L2 and CCL5) were highly expressed in CXCL13+ CD8+ TRMs from CC tissues (Fig. 4f, g). The GO term analysis revealed that CXCL13+ CD8+ TRMs from CC tissues were enriched in pathways related to antigen processing and presentation, cell adhesion, and chemotaxis (Fig. 4h); this was confirmed by the results of the GSEA (Fig. 4i). We therefore next evaluated the relationship between CXCL13+ CD8+ TRM signature and the survival outcomes of patients with CC following radiotherapy using bulk RNA-seq data obtained from The Cancer Genome Atlas (TCGA). We found that the signature gene set score of CXCL13+ CD8+ TRMs was positively correlated with the overall survival (OS) and progression-free survival (PFS) of patients with CC following radiotherapy (Fig. 4j).
In summary, these findings indicate that CXCL13+ CD8+ TRMs exhibit cytotoxic and inhibitory characteristics, enriched in pathways associated with defense response to virus and the type I interferon response than the other TRM subclusters, and are associated with the improved outcome of patients with CC following radiotherapy.
PLAC8 + CD8+ TRMs are less enriched in CC tissues than in normal cervical tissues but exhibit highly cytotoxic characteristics
We next dissected the molecular signature of PLAC8+ CD8+ TRMs, the subcluster which was less abundant in CC tissues than in normal cervical tissues (Fig. 5a). DEGs analysis exhibited that PLAC8+ CD8+ TRMs showed high expression of cytotoxic genes (GZMH, GNLY, GZMM, KLRD1, KLRF1, KLRG1, NKG7, and PRF1) and low expression of genes encoding inhibitory molecules (PDCD1, CTLA4, TIGIT, and HAVCR2) (Fig. 5b, c), exhibiting similar characteristics to cytotoxic T cells. Further GO term analysis revealed that PLAC8+ CD8+ TRMs were enriched in pathways associated with T-cell-mediated immune effector processes, and also enriched cytoplasmic translation and ribosome biogenesis pathways (Fig. 5d, e).
a tSNE plot showing the distribution of PLAC8+ CD8+ TRMs (red) relative to that of the other TRM subclusters (blue). b Volcano plot showing DEGs between PLAC8+ CD8+ TRMs (red) and the other TRM subclusters (blue). c Violin plots displaying the expression of the indicated genes between PLAC8+ CD8+ TRMs and the other TRM subclusters. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 (two-sided Wilcoxon test). d GO term analysis of PLAC8+ CD8+ TRMs; the color intensity indicates p-value magnitude. e Violin plots showing differences in pathway activity between PLAC8+ CD8+ TRMs (red) and the other TRM subclusters (blue). f Volcano plot showing DEGs in PLAC8+ CD8+ TRMs between CC tissues (red) and normal cervical tissues (blue). g Violin plots illustrating the expression of indicated genes in PLAC8+ CD8+ TRMs between CC tissues and normal cervical tissues. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 (two-sided Wilcoxon test). h GO term analysis in PLAC8+ CD8+ TRMs of CC tissues. The intensity of color indicates p-values. i GSEA showing differences in the pathway activities of PLAC8+ CD8+ TRMs from CC tissues and those from normal cervical tissues. j Kaplan–Meier survival analysis of patients with CC following radiotherapy from TCGA database, stratified based on high or low signature gene set score of PLAC8+ CD8+ TRMs. The p-values were calculated using a two-sided log-rank test.
In addition, we explored signature differences between PLAC8+ CD8+ TRMs from CC tissues and those from normal cervical tissues. DEGs analysis revealed that HLA-DRB5, CD74, HLA-DRA, and HLA-DQA2 were more highly expressed in PLAC8+ CD8+ TRMs from CC tissues than in those from normal cervical tissues (Fig. 5f, g). The GO term analysis demonstrated that PLAC8+ CD8+ TRMs from CC tissues were enriched with immune response-related pathways, including T cell activation, cytokine-mediated signaling pathway, antigen processing and presentation, and cell adhesion (Fig. 5h and Supplementary Fig. 5). The GSEA confirmed that PLAC8+ CD8+ TRMs were enriched in gene sets associated with antigen processing and presentation, as well as cell adhesion (Fig. 5i). Moreover, survival analysis revealed a positive correlation between the signature gene set score of PLAC8+ CD8+ TRMs and the OS and PFS of patients with CC following radiotherapy (Fig. 5j). These findings suggest that although PLAC8+ CD8+ TRMs were not abundant in the TME of CC, they were a highly cytotoxic, readily activated, clinically important TRM subset.
CXCL13 + CD8+ TRMs and PLAC8 + CD8+ TRMs interact with epithelial cells in the TME of CC
To further investigate the interaction between epithelial cells and TRMs, we used CellChat to interrogate the scRNA-seq data and gather evidence of cell-cell communication through the analysis of ligand-receptor complexes. We found that the interactions between epithelial cells and TRMs were more robust in CC tissues than in normal cervical tissues, both in terms of their number and strength (Fig. 6a). Specifically, epithelial cells, as senders, showed more active interactions with CXCL13+ CD8+ TRMs and PLAC8+ CD8+ TRMs than the other TRM subclusters in CC tissues, both in terms of number and strength (Fig. 6b and Supplementary Fig. 6a).
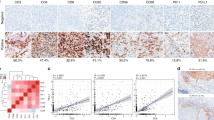
a Bar graphs displaying the number and strength of interactions between epithelial cells and TRMs in normal cervical tissues and CC tissues, respectively. b CellChat analysis of scRNA-seq data to identify cell-cell communications between epithelial cells and TRM subclusters in CC tissues. The color intensity indicates the number (top panel) and strength (bottom panel) of the cell-cell interactions. c Bubble plots showing differences in ligand-receptor pairs involved in interactions between epithelial cell-CXCL13+ CD8+ TRM and epithelial cell-PLAC8+ CD8+ TRM in normal cervical tissues and CC tissues. Bubble size and color intensity indicate p-value magnitude and communication probability, respectively. d Circle plots showing the indicated ligand-receptor pairs involved in interactions between epithelial cells and either CXCL13+ CD8+ TRMs, or PLAC8+ CD8+ TRMs in CC tissues (left) and normal cervical tissues (right); color-coded by cell type. e Representative immunofluorescent labeling of Pan-CK (yellow), CXCL13 (red), CD103 (green), CD8 (white), and DAPI (blue) to identify epithelial cells, CXCL13+ CD8+ TRMs, and their spatial relationship in CC sections. Scale bar in figure, 20 μm.
Next, we explored the specific ligand-receptor pairs which participated in the interactions between epithelial cells and either CXCL13+ CD8+ TRMs or PLAC8+ CD8+ TRMs both in CC tissues and in normal cervical tissues (Fig. 6c). We found that MHC class I molecules-related pairs (HLA-A/B/C/E/F - CD8A/ CD8B), T cell proliferation and activation-related pairs (MIF-(CD74+CD44)(CD74+CD44) and MDK-NCL) were more active both in epithelial cell-CXCL13+ CD8+ TRM and epithelial cell-PLAC8+ CD8+ TRM at CC tissues than normal cervical tissues (Fig. 6c and Supplementary Fig. 6b). This result suggested that tumor antigens may be presented to TRMs via MHC class I molecules in the TME of CC, leading to the rapid activation of this T cell subset and the induction of anti-tumor immune responses. In addition, HLA-E - KLRC1 and HLA-E - CD94:NKG2A pairs were enriched in epithelial cell-CXCL13+ CD8+ TRM at CC tissues, rather than in epithelial cell-PLAC8+ CD8+ TRM (Fig. 6c, d). Of note, both of these interactions are associated with T cell inhibition and increased cancer growth23,24. We confirmed the presence and spatial relationship of epithelial cells and CXCL13+ CD8+ TRMs within the TME of CC by multiplex immunofluorescent staining by labeling for pan-cytokeratin (Pan-CK), CXCL13, CD103, and CD8. The proximity observed between epithelial cells and CXCL13+ CD8+ TRMs on tissue sections supports their potential cellular interactions (Fig. 6e).
In summary, we detected active interactions between epithelial cells and either CXCL13+ CD8+ TRMs, or PLAC8+ CD8+ TRMs in the TME of CC. Moreover, the immunosuppressive HLA-E - KLRC1 and HLA-E - CD94:NKG2A pairs were enriched in CXCL13+ CD8+ TRMs interacting with epithelial cells at CC tissues.
Discussion
TRMs play a vital role in the immune response against tumors9. Previous studies have dissected the cellular heterogeneity and molecular profiles of conventional T cells within the TME of CC25,26,27; however, the same in-depth characterization of TRMs has not been performed in the CC setting. Thus, in this study, we employed scRNA-seq to investigate the population characteristics of TRMs in CC. Our findings reveal that TRMs readily infiltrate CC tissues and exhibit high levels of immune activity. Furthermore, our data indicate that TRMs generate effective anti-tumor immune responses, recruit other immune cells, and respond to viral stimuli.
We initially observed that TRMs were more immunoreactive than non-TRMs, as evidenced by the high expression of immune-related genes, including those associated with cytotoxicity and chemokine production. This suggested that TRMs may kill tumor cells and recruit other immune cells to elicit effective anti-tumor immune responses; these properties of TRMs have also been reported in the context of breast cancer28 and lung cancer22. Additionally, we observe a higher level of TRM infiltration into CC tissue than normal cervical tissues, which aligns with the findings of a prior study29. The factors contributing to the development of tumor-infiltrating TRMs are not yet fully understood9. However, it is generally believed that the upregulation of migratory molecules, such as the chemokine receptors CXCR3 and CXCR6, or tissue-specific homing molecules, may play a crucial role in facilitating the recruitment of memory precursor-like cells into tumors30. We found that TRMs were enriched in genes associated with antigen processing and presentation, indicating that the activation of TRMs in the TME of CC may rely on MHC class II-mediated antigen presentation. Notably, the enrichment of gene sets associated with the response to type I interferon and defense response to virus in TRMs from CC tissues suggests that TRMs might exert anti-viral effects upon stimulation with type I interferon. This finding is especially relevant in the context of CC, where HPV infection plays a crucial role in tumor development31,32.
TRMs exhibited an activated and functional phenotype in CC tissues, expressing genes encoding cytotoxic and immune checkpoint molecules, as well as those implicated in cell proliferation. Previous studies in melanoma and lung cancer33,34 have shown that TRMs enriched in immune-checkpoint-related genes are especially sensitive to checkpoint inhibition therapy. TRMs play an important role in cancer immunotherapy in some solid tumors. For instance, in head and neck squamous cell carcinoma patients, neoadjuvant immunotherapy resulted in the preferential expansion of TRMs15. Encouragingly, blockade of PD-1 and/or CTLA-4 effectively restored TRMs proliferation and enhanced their cytokine production35. Moreover, anti-PD-1 therapy promoted TRMs proliferation in melanoma patients, which was associated with improved survival36.
We identified two significant subgroups of TRMs in CC. CXCL13+ CD8+ TRMs were more abundant in CC tissues compared to normal cervical tissues and had both cytotoxic (GZMB, GNLY, and PRF1) and inhibitory (HAVCR2, TIGIT, LAG3, CTLA4, and PDCD1) characteristics. In vitro experiments have demonstrated that blocking immune checkpoints, such as PD-1 and TIM-3, enhances the proliferation of CD8+ TRMs and improves their cytokine-mediated killing capacity37,38. These findings suggest that CXCL13+ CD8+ TRMs, which express immune-checkpoint molecules, could be effectively targeted by ICIs therapy. Interestingly, CXCL13+ CD8+ TRMs were more enriched in pathways associated with defense response to virus and the type I interferon response than the other TRM subclusters. Previous studies have shown the significance of TRMs in the immune against HPV infection, as they can recognize HPV-infected cells and contribute to viral clearance39. The unique profile of CXCL13+ CD8+ TRMs suggests that they may exert anti-HPV effects.
By contrast, PLAC8+ CD8+ TRMs was less abundant in CC tissues compared to normal cervical tissues and exhibited a higher naïve score than other CD8+ TRM subclusters. We hypothesize that upon residency within CC tissues, PLAC8+ CD8+ TRMs undergo gradual differentiation into other TRM subgroups in response to persistent stimulation by tumor-related antigens. PLAC8+ CD8+ TRMs exhibited elevated expression of cytotoxic genes and reduced expression of genes encoding inhibitory molecules. Moreover, they were enriched in pathways associated with T-cell-mediated immune effector processes. These findings indicate that PLAC8+ CD8+ TRMs are important in immune function. And the upregulation of genes associated with cytoplasmic translation and ribosome biogenesis suggests that PLAC8+ CD8+ TRMs have a high protein synthesis capacity. To recognize and kill target cells, including tumor cells, cytotoxic T cells need to rapidly produce and release cytotoxic proteins, such as granzymes and perforin40. This increased demand for protein following T cell activation requires the up-regulation of ribosome biogenesis41. Importantly, the frequencies of CXCL13+ CD8+ TRMs and PLAC8+ CD8+ TRMs exhibited a positive correlation with the prolonged survival of patients with CC, demonstrating their potential as novel prognostic markers.
The results of cell-cell interaction analysis revealed active interactions between epithelial cell-CXCL13+ CD8+ TRM and epithelial cell-PLAC8+ CD8+ TRM in the TME of CC. The ligand-receptor pairs involved in these interactions included MHC class I molecules-related pairs (HLA-A/B/C/E/F - CD8A/CD8B) and T cell proliferation and activation-related pairs (MIF-(CD74+CD44) and MDK-NCL). Tumor cells can express various MHC class I molecules (e.g., HLA-A, HLA-B, and HLA-C) on their surface, which present tumor-specific antigens to CD8+ T cells. In some cases, cancer cells can down-regulate MHC class I expression, thus evading recognition and killing by T cells42,43. Notably, we observed that the interaction between HLA-E and NKG2A (either alone or in a complex with CD94), which is associated with T cell inhibition and increased cancer growth, was enriched in CXCL13+ CD8+ TRMs interacting with epithelial cells at CC tissues. NKG2A and its heterodimeric CD94:NKG2A form are inhibitory receptors that interact with HLA-E on target cells. This interaction transmits inhibitory signals to NK cells and CD8+ T cells, dampening their cytotoxic activity23,24. It has been shown that CC-infiltrating CD8+ T cells exhibit higher expression of CD94:NKG2A than those present in the peripheral blood or normal cervical tissues. Moreover, kinetic co-culture experiments have shown that CC cells promote CD94:NKG2A expression on CD8+ T cells. The increase in CD94:NKG2A expression on CD8+ T cells markedly reduce their intracellular perforin levels44. Therefore, the use of NKG2A inhibitors, typically antibodies which block the HLA-E and NKG2A interaction, has emerged as a promising immunotherapeutic approach in CC.
Although our study has provided valuable insights into the role of TRMs in CC, it is important to acknowledge several limitations. First, the sample size in our study is relatively small, and larger-scale studies are required to validate and reinforce our findings. Second, the HPV infection status of a portion of the normal cervical tissues was unknown in our study. Therefore, it was impossible to directly compare the immunological differences between normal tissues in which the virus had been cleared or was not detectable, and those in which the tissue appeared morphologically normal but the virus had been detected. Third, further functional experiments are needed to validate the characteristics of key TRMs subsets identified in this study. Lastly, additional spatial information is needed to understand the true localization of TRMs in CC.
In summary, this study comprehensively evaluated the heterogeneity and gene expression profiles of TRMs in CC, as well as their potential functions and interactions in the TME of CC at a single-cell level. Our findings revealed that TRMs are highly immunoreactive, capable of infiltrating tumor tissues, and displaying distinct gene expression profiles. Moreover, they play a significant role in CC eradication, potentially by recruiting other immune cells and responding to viral stimuli. These findings significantly contributed to our knowledge of TRM-mediated immunotherapy in CC, underscoring the potential of TRMs as prognostic markers and therapeutic targets. Further studies exploring the functional mechanisms and dynamics of TRMs in the TME of CC are warranted to fully comprehend their roles in the anti-tumor/viral immune response and to develop targeted therapeutic strategies for CC.
Methods
Human samples and data collection
The scRNA-seq data of 38,572 T cells included in this study comprised new data of 20,976 (54.4%) T cells from 3 HPV+ CC samples and previously published data of 17,596 (45.6%) T cells from 8 HPV+ CC and 5 normal cervical samples45,46. This study was approved by the Ethics Committee of Shandong Cancer Hospital (SDTHEC201906009) and conducted in accordance with the guidelines of the Declaration of Helsinki. Written informed consent was obtained from all participants. A total of three patients (age range: 32–53 years) with CC, all of whom were diagnosed with HPV16-positive tumors, were newly enrolled in this study after obtaining their written informed consent. Two patients were diagnosed with stage IIIC and stage IB2 cervical adenocarcinoma, according to the Federation International of Gynecology and Obstetrics classification. The third patient was diagnosed with stage IB2 cervical squamous cell carcinoma (Supplementary Table 1). Fresh tissues were obtained from the primary tumors of these patients through surgery. The tissues were promptly placed in MACS Tissue Storage Solution (Miltenyi Biotec) while kept on ice. Subsequently, the samples were promptly transferred to the laboratory.
Sample preparation
For each tumor specimen, a serious of digestion and sorting steps were performed. First, the samples were enzymatically digested using a medium containing collagenase IV (2 mg ml−1) and Dnase I (1 mg ml−1) (both from Sigma), and then incubated at 37 °C for a duration of 35 minutes. To stop the digestion process, phosphate-buffered saline (PBS), supplemented with 1% bovine serum albumin (BSA) (Absin), was added. The digestion mixture was filtered using a 70-µm cell filter (BD Biosciences) and centrifuged at 400 × g for 5 min. After carefully removing the supernatant, the resulting cell pellet was treated with RBC Lysis Buffer (BioLegend) and incubated at 4 °C for 5 min. Following another centrifugation step at 400 × g for 5 min, the supernatant was discarded, and the resulting cell pellet was resuspended in 100 μl sorting buffer (PBS containing 1% BSA). Mouse anti-human monoclonal antibodies specifically targeting CD45 (Clone 2D1; BD Biosciences) and CD3 (Clone OKT3; BD Biosciences) were used to label the cells. The cells were incubated with the antibodies for 15 min at 4 °C, followed by two washes and resuspension in 100 μl of sorting buffer. Subsequently, the cells were stained with 7-AAD Viability Staining Solution (eBioscience) and sorted on the BD FACSAria III cell sorter, gating on live T cells (7-AAD−CD45+CD3+), for subsequent single-cell library preparation and sequencing.
scRNA‐seq and data preprocessing
The scRNA-seq data were generated using the 10× Genomics platform. The raw sequencing FASTQ files were aligned to the GRCh38 reference genome using Cell Ranger count, utilizing the STAR algorithm were aligned to generate a gene expression library. The gene expression data obtained from the count matrices generated by Cell Ranger were processed using Seurat (version 4.2.0)47. The newly generated data were integrated with high-quality single-cell data from our previous studies45,46. The data was analyzed using Seurat for each batch, and the DoubletFinder R package was utilized to identify doublets. After removing the doublets, the raw gene expression data were filtered by excluding cells with high mitochondrial content or low gene expression counts. For the remaining high-quality cells, the gene expression matrix was normalized using the NormalizeData function. Highly variable genes were then selected using the FindVariableFeatures function with a threshold of 2000 features, and data standardization was performed using the ScaleData function. Next, principal component analysis (PCA) was conducted on the highly variable genes to identify the most significant principal components. To mitigate batch effects between samples, the data were integrated using the “harmony” R package.
Cell type annotation and T cell isolation
The FindClusters function was used to identify cell clusters and subclusters, and the t-Distributed Stochastic Neighbor Embedding (t-SNE) algorithm was employed for dimensionality reduction and cell visualization. The resulting cell clusters were annotated based on the expression of lineage-specific genes in different cell types. After quality control and filtering, scRNA-seq profiles were obtained for 38,572T cells (CD2+ and CD3D+) (Supplementary Fig. 1a, b). Based on the expression of the known TRMs marker gene ITGAE (encoding CD103)10, we classified T cells as TRMs or non-TRMs using normalized expression data. The threshold was ITGAE expression greater than 0.5. Additionally, we obtained the scRNA-seq data of PBMC from three healthy individuals from the Gene Expression Omnibus database (accession number GSE157007). We extracted T cell data for analysis through dimensionality reduction clustering analysis and compared the expression of ITGAE in TRMs, non-TRMs, and T cells in PBMC. The differential proportion analysis for cell subclusters was conducted using the Wilcoxon test.
Differentially expressed genes and pathway enrichment analysis
DEGs were identified using the FindAllMarkers function in Seurat, with the Wilcoxon test and significance threshold at p-value < 0.05 and an absolute log2 fold change >0.25. To investigate the functional relevance and activity of the gene sets, GO term analysis and GSEA were carried out using the ClusterProfiler package in R, focusing on the DEGs48. In addition, the activity levels of signaling pathways were compared between different cell types by scoring gene signatures using the Ucell R package49.
Analysis of data from The Cancer Genome Atlas
Bulk RNA-seq data were used alongside curated clinical data from TCGA database. The top 30 DEGs in TRMs were selected to define the T cell population characteristics. Subsequently, Gene Set Variation Analysis (GSVA) was conducted to evaluate the relative abundance of these cell signatures in TCGA database50. The survcutpoint function from the survminer R package was employed to determine the optimal cutoff value. Using this cutoff, the Patients with CC following radiotherapy were categorized into high- or low-expression groups based on their gene expression profiles. The survival package as used to perform the log-rank test and the Kaplan–Meier survival curves were plotted by the ggsurvplot function.
Predicting putative interactions between cell types
The CellChat R package was utilized to infer putative interactions between different cell types by analyzing receptor-ligand interactions within the scRNA-seq dataset51. We first normalized the Seurat object using the NormalizeData function of the Seurat package, which scales gene expression measurements to obtain comparable data from different cells. After normalization, data were stratified by tumor and normal tissues to explore potential cell interactions within the tumor ecosystem. Preprocessing functions, such as identifying overexpressed genes and interactions, as well as data projection, were executed using default parameters. For the primary analyses, core functions, including computing communication probabilities, computing pathway-specific communication probabilities, and aggregating the network, were executed separately for tumor-free and tumor tissue datasets. The results were merged using the mergeCellChat function. The results obtained were merged using the mergeCellChat function. To visually represent the communication probabilities mediated by ligand-receptor pairs between specific cell groups, the netVisual_bubble function was utilized. Additionally, netVisual_individual was used to depict all the important interactions between a given signal and the associated signaling genes.
Multiplex immunofluorescent labeling
To verify the difference in the infiltration degree of TRMs and non-TRMs between CC tissues and normal cervical tissues, we obtained 20 FFPE samples of CC and 20 normal cervical tissue samples. The following markers were used for immunofluorescent labeling: CD103, CD3, and 4,6-diamidino-2-phenylindole (DAPI) for nuclear staining. Immunofluorescence kit designed for multiplex staining was used. FFPE tissue sections of CC and normal cervical tissues were sequentially incubated with the above primary antibodies. The tissues were subsequently incubated with horseradish peroxidase-conjugated secondary antibodies, and signal amplification was achieved using tyramide signal amplification (TSA). Each step was followed by heating in a microwave to facilitate antibody stripping and prepare the sections for subsequent labeling. After all target antigens were labeled, nuclear counterstaining was performed using DAPI. Stained slides were scanned with a Mantra System (PerkinElmer), capturing fluorescence spectra across 420–720 nm at 20-nm intervals, under consistent exposure times. The resulting spectral images were processed using the InForm image analysis software (PerkinElmer). Use Visiopharm Intelligent Full Line AI Digital Pathology Quantitative Analysis Software to perform target cell counting on the entire resulting spectral images.
To identify epithelial cells, CXCL13+ CD8+ TRMs, and their spatial relationship within the TME of CC, a multiplex immunofluorescent labeling technique was utilized. This approach employed markers including Pan-CK to detect epithelial cells, CXCL13, CD103, and CD8 to identify CXCL13+ CD8+ TRMs, and DAPI for nuclear staining. Immunofluorescence kit designed for multiplex staining was used. FFPE tissue sections of CC were sequentially incubated with the above primary antibodies followed by incubation with horseradish peroxidase-conjugated secondary antibodies. Signal amplification was achieved using TSA. Each TSA step was followed by heating in a microwave to facilitate antibody stripping and prepare the sections for subsequent labeling. After all target antigens were labeled, nuclear counterstaining was performed using DAPI. The stained slides were scanned using a Mantra System (PerkinElmer), capturing fluorescence spectra across 420–720 nm at 20-nm intervals, under consistent exposure times. The resulting spectral images were processed using the InForm image analysis software (PerkinElmer).
Data availability
Processed scRNA-seq data were uploaded to the National Genomics Data Center, which are publicly accessible at https://ngdc.cncb.ac.cn/omix (OMIX006446-01).
Code availability
The data analysis R script for this study is publicly accessible at https://github.com/WFhoo/TRM. Conducted using R version 4.1.3, the script includes detailed documentation of the specific parameters and variables utilized.
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. CA Cancer J. Clinicians 72, 7–33 (2022).
Rahangdale, L. et al. Human papillomavirus vaccination and cervical cancer risk. BMJ 379, e070115 (2022).
Yoon, J. W. et al. PET/CT Response Criteria (European Organization for Research and Treatment of Cancer) Predict Survival Better Than Response Evaluation Criteria in Solid Tumors in Locally Advanced Cervical Cancer Treated With Chemoradiation. Clin. Nucl. Med. 41, 677–682 (2016).
Perucho, J. A. U. et al. Association between IVIM parameters and treatment response in locally advanced squamous cell cervical cancer treated by chemoradiotherapy. Eur. Radio. 31, 7845–7854 (2021).
Mauricio, D. et al. Immunotherapy in Cervical Cancer. Curr. Oncol. Rep. 23, 61 (2021).
Ferrall, L. et al. Cervical Cancer Immunotherapy: Facts and Hopes. Clin. Cancer Res. 27, 4953–4973 (2021).
Lin, Y. H. et al. Small intestine and colon tissue-resident memory CD8(+) T cells exhibit molecular heterogeneity and differential dependence on Eomes. Immunity 56, 207–223.e8 (2023).
Kurd, N. S. et al. Early precursors and molecular determinants of tissue-resident memory CD8(+) T lymphocytes revealed by single-cell RNA sequencing. Sci. Immunol. 5, eaaz6894 (2020)
Park, S. L., Gebhardt, T. & Mackay, L. K. Tissue-resident memory T cells in cancer immunosurveillance. Trends Immunol. 40, 735–747 (2019).
Amsen, D., van Gisbergen, K., Hombrink, P. & van Lier, R. A. W. Tissue-resident memory T cells at the center of immunity to solid tumors. Nat. Immunol. 19, 538–546 (2018).
Wang, Z. Q. et al. CD103 and Intratumoral Immune Response in Breast Cancer. Clin. Cancer Res. 22, 6290–6297 (2016).
Ganesan, A. P. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat. Immunol. 18, 940–950 (2017).
Workel, H. H. et al. CD103 defines intraepithelial CD8+ PD1+ tumour-infiltrating lymphocytes of prognostic significance in endometrial adenocarcinoma. Eur. J. Cancer 60, 1–11 (2016).
Wang, B. et al. CD103+ Tumor Infiltrating Lymphocytes Predict a Favorable Prognosis in Urothelial Cell Carcinoma of the Bladder. J. Urol. 194, 556–562 (2015).
Luoma, A. M. et al. Tissue-resident memory and circulating T cells are early responders to pre-surgical cancer immunotherapy. Cell 185, 2918–2935.e29 (2022).
Sun, Y. Y. et al. Local HPV Recombinant Vaccinia Boost Following Priming with an HPV DNA Vaccine Enhances Local HPV-Specific CD8+ T-cell-Mediated Tumor Control in the Genital Tract. Clin. Cancer Res. 22, 657–669 (2016).
Malik, B. T. et al. Resident memory T cells in the skin mediate durable immunity to melanoma. Sci. Immunol. 2, eaam6346 (2017)
Beumer-Chuwonpad, A., Taggenbrock, R., Ngo, T. A. & van Gisbergen, K. The Potential of Tissue-Resident Memory T Cells for Adoptive Immunotherapy against Cancer. Cells. 10, 2234 (2021)
Lei, Y. et al. Applications of single-cell sequencing in cancer research: progress and perspectives. J Hematol Oncol. 14, 91 (2021)
Mereu, E. et al. Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat. Biotechnol. 38, 747–755 (2020).
Savas, P. et al. Single-cell profiling of breast cancer T cells reveals a tissue-resident memory subset associated with improved prognosis. Nat. Med. 24, 986–993 (2018).
Clarke, J. et al. Single-cell transcriptomic analysis of tissue-resident memory T cells in human lung cancer. J. Exp. Med. 216, 2128–2149 (2019).
Salomé, B. et al. NKG2A and HLA-E define an alternative immune checkpoint axis in bladder cancer. Cancer Cell 40, 1027–1043.e9 (2022).
Abd Hamid, M. et al. Enriched HLA-E and CD94/NKG2A Interaction Limits Antitumor CD8(+) Tumor-Infiltrating T Lymphocyte Responses. Cancer Immunol. Res. 7, 1293–1306 (2019).
Li, X. et al. Single-cell RNA-sequencing dissects cellular heterogeneity and identifies two tumor-suppressing immune cell subclusters in HPV-related cervical adenosquamous carcinoma. J. Med. Virol. 94, 6047–6059 (2022).
Li, C. et al. Single-cell transcriptomics reveals cellular heterogeneity and molecular stratification of cervical cancer. Commun. Biol. 5, 1208 (2022).
Li, C., Liu, D., Yang, S. & Hua, K. Integrated single-cell transcriptome analysis of the tumor ecosystems underlying cervical cancer metastasis. Front. Immunol. 13, 966291 (2022).
Byrne, A. et al. Tissue-resident memory T cells in breast cancer control and immunotherapy responses. Nat. Rev. Clin. Oncol. 17, 341–348 (2020).
Losurdo, A. et al. Single-cell profiling defines the prognostic benefit of CD39(high) tissue resident memory CD8+ T cells in luminal-like breast cancer. Commun. Biol. 4, 1117 (2021).
Masopust, D. & Schenkel, J. M. The integration of T cell migration, differentiation and function. Nat. Rev. Immunol. 13, 309–320 (2013).
Crosbie, E. J., Einstein, M. H., Franceschi, S. & Kitchener, H. C. Human papillomavirus and cervical cancer. Lancet 382, 889–899 (2013).
Agarwal, C., Hembree, J. R., Rorke, E. A. & Eckert, R. L. Interferon and retinoic acid suppress the growth of human papillomavirus type 16 immortalized cervical epithelial cells, but only interferon suppresses the level of the human papillomavirus transforming oncogenes. Cancer Res. 54, 2108–2112 (1994).
Boddupalli, C. S. et al. Interlesional diversity of T cell receptors in melanoma with immune checkpoints enriched in tissue-resident memory T cells. JCI Insight 1, e88955 (2016).
Djenidi, F. et al. CD8+CD103+ tumor-infiltrating lymphocytes are tumor-specific tissue-resident memory T cells and a prognostic factor for survival in lung cancer patients. J. Immunol. 194, 3475–3486 (2015).
Lee, Y. J. et al. CD39(+) tissue-resident memory CD8(+) T cells with a clonal overlap across compartments mediate antitumor immunity in breast cancer. Sci. Immunol. 7, eabn8390 (2022).
Edwards, J. et al. CD103(+) Tumor-Resident CD8(+) T Cells Are Associated with Improved Survival in Immunotherapy-Naïve Melanoma Patients and Expand Significantly During Anti-PD-1 Treatment. Clin. Cancer Res. 24, 3036–3045 (2018).
Djenidi, F. et al. CD8+CD103+ Tumor–Infiltrating Lymphocytes Are Tumor-Specific Tissue-Resident Memory T Cells and a Prognostic Factor for Survival in Lung Cancer Patients. J. Immunol. 194, 3475–3486 (2015).
Shwetank et al. Maintenance of PD-1 on brain-resident memory CD8 T cells is antigen independent. Immunol. Cell Biol. 95, 953–959 (2017).
Yenyuwadee, S. et al. The evolving role of tissue-resident memory T cells in infections and cancer. Sci. Adv. 8, eabo5871 (2022).
Shi, Y., Lu, Y. & You, J. Unfolded protein response in the activation-induced biological processes of CD8(+) T cells. Pharm. Res. 169, 105654 (2021).
Galloway, A. et al. Upregulation of RNA cap methyltransferase RNMT drives ribosome biogenesis during T cell activation. Nucleic Acids Res. 49, 6722–6738 (2021).
Burr, M. L. et al. An Evolutionarily Conserved Function of Polycomb Silences the MHC Class I Antigen Presentation Pathway and Enables Immune Evasion in Cancer. Cancer Cell 36, 385–401.e8 (2019).
Jongsma, M. L. M., Neefjes, J. & Spaapen, R. M. Playing hide and seek: Tumor cells in control of MHC class I antigen presentation. Mol. Immunol. 136, 36–44 (2021).
Sheu, B. C. et al. Up-regulation of inhibitory natural killer receptors CD94/NKG2A with suppressed intracellular perforin expression of tumor-infiltrating CD8+ T lymphocytes in human cervical carcinoma. Cancer Res. 65, 2921–2929 (2005).
Liu, C. et al. Single-cell dissection of cellular and molecular features underlying human cervical squamous cell carcinoma initiation and progression. Sci. Adv. 9, eadd8977 (2023).
Liu, C. et al. Single-cell RNA-sequencing reveals radiochemotherapy-induced innate immune activation and MHC-II upregulation in cervical cancer. Signal Transduct. Target Ther. 8, 44 (2023).
Butler, A. et al. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Andreatta, M. & Carmona, S. J. UCell: Robust and scalable single-cell gene signature scoring. Comput Struct. Biotechnol. J. 19, 3796–3798 (2021).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 14, 7 (2013).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Acknowledgements
This work was supported by the following grants: Postdoctoral Fellowship Program of CPSF (GZB20230041), Shandong Provincial Natural Science Foundation (ZR2021QH006), National Natural Science Foundation of China (82272753), Shandong Provincial Natural Science Foundation (ZR2021LZL002), Young TaiShan Scholars and Academic Promotion Program of Shandong First Medical University (2019RC003), and Bethune Cancer Radiotherapy Translational Medicine Research Fund (flzh202103).
Author information
Authors and Affiliations
Contributions
C.L., J.B.Y., and C.W. concepted and designed of the study. Q.Y.H. collected sample and clinical data. F.H.W acquired, analyzed and interpreted the data. S.Q.Y and Q.Y.H. drafted the article. T.Y.L and X.H.L revised the manuscript. F.H.W., S.Q.Y and Q.Y.H. contributed equally to the manuscript and should be considered co-first authors. All authors have reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, F., Yue, S., Huang, Q. et al. Cellular heterogeneity and key subsets of tissue-resident memory T cells in cervical cancer. npj Precis. Onc. 8, 145 (2024). https://doi.org/10.1038/s41698-024-00637-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-024-00637-3
- Springer Nature Limited