Abstract
The mouse Igf2/H19 locus is regulated by genomic imprinting, in which the paternally methylated H19 imprinting control region (ICR) plays a critical role in mono-allelic expression of the genes in the locus. Although the maternal allele-specific insulator activity of the H19 ICR in regulating imprinted Igf2 expression has been well established, the detailed mechanism by which the H19 ICR controls mono-allelic H19 gene expression has not been fully elucidated. In this study, we evaluated the effect of H19 ICR orientation on imprinting regulation in mutant mice in which the H19 ICR sequence was inverted at the endogenous locus. When the inverted-ICR allele was paternally inherited, the methylation level of the H19 promoter was decreased and the H19 gene was derepressed, suggesting that methylation of the H19 promoter is essential for complete repression of H19 gene expression. Unexpectedly, when the inverted allele was maternally inherited, the expression level of the H19 gene was lower than that of the WT allele, even though the H19 promoter remained fully hypomethylated. These observations suggested that the polarity of the H19 ICR is involved in controlling imprinted H19 gene expression on each parental allele, dependent or independent on DNA methylation of the H19 promoter.
Similar content being viewed by others
Introduction
Genomic imprinting in mammals is an epigenetic phenomenon in which a subset of genes is expressed only when inherited from either the father or the mother. Because imprinted genes have diverse functions in development, growth, and behavior, disruption of their mono-allelic expression causes human diseases such as Beckwith–Wiedemann and Silver–Russell syndromes1,2. Imprinted genes form clusters on the genome, and their accompanying imprinted control regions (ICRs) regulate the expression patterns of multiple genes within each locus. Because most ICRs acquire DNA methylation in either the sperm or the egg, they are also called germline differentially methylated regions (gDMRs). After fertilization, differential methylation of ICRs between alleles is maintained throughout development, and, at some loci, induces allele-specific DNA methylation of secondary DMRs or somatic DMRs (sDMRs)3,4. Methylation of gDMRs and sDMRs regulates allele-preferential binding of transcription factors, resulting in unilateral allelic expression of genes within the locus.
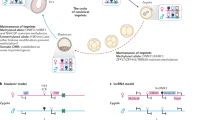
Mono-allelic expression of the Igf2/H19 gene locus is controlled by paternal allele-specific DNA methylation of the H19 ICR, located 2–4 kb upstream of the H19 gene, and its insulator activity is involved in the imprinted expression of the distally-located Igf2 gene (Supplementary Fig. 1a). On the maternal allele, the CTCF insulator protein bound to the unmethylated H19 ICR sequence inhibits the action of the enhancers, located downstream of the H19 gene, on the Igf2 gene. On the other hand, in the paternally inherited allele, the absence of CTCF binding to the methylated H19 ICR allows activation of Igf2 by the enhancers5,6,7,8,9,10.
It is generally believed that regulation of the H19 gene involves an epigenetic change at the H19 promoter that is secondary to the methylation status of the H19 ICR. On the paternal allele, both the H19 ICR and the H19 promoter are DNA methylated, and transcription of H19 is repressed11. In mice lacking the paternal H19 ICR, the methylation level of the H19 promoter decreases, and H19 is expressed12. In addition, when demethylation of the H19 ICR was forcibly induced by epigenome editing in ES cells, the methylation level of the H19 promoter decreased, and H19 expression was activated in mice derived from these ES cells13. Thus, DNA methylation of the H19 promoter seems mandatory for the repression of H19 expression.
On the other hand, it has been reported that the paternal H19 ICR represses H19 expression via a mechanism independent of its promoter methylation status. Deletion of a 1.2 kb region of the paternal H19 ICR resulted in derepression of the H19 gene in cis, although methylation status of the rest of the H19 ICR and of the H19 promoter region remained unchanged14. Deletion of a ~0.9 kb region between CTCF binding sites 2 and 3 or mutation of the eight CpGs outside the CTCF binding sites of the paternal H19 ICR also resulted in expression of H19, without altering the methylation status at the H19 ICR and promoter15. These observations suggest that silencer activity exists in the paternal H19 ICR or that the overall size or (methylated) CpG density is critical for conferring full repression activity on the paternal H19 gene. Furthermore, Gebert et al. reported that knock-in of the H19 ICR sequence at the mouse Afp locus repressed paternal Afp gene expression without altering the methylation level of the Afp gene promoter16.
As described above, H19 gene expression appears to be regulated by multiple mechanisms (i.e., H19 promoter methylation and the H19 ICR itself). To date, however, no experiments have been able to directly determine whether methylation of the H19 promoter is required for repression of the H19 gene. This is largely because no experiments have been able to decrease H19 promoter methylation without altering either the size or hypermethylation status of the H19 ICR12,13.
Methylation of the paternal H19 promoter changes dynamically during development. The H19 locus, including the H19 ICR, its downstream region, and the H19 gene body, is highly and broadly methylated in sperm, with the exception of the H19 promoter5,6. After fertilization and before implantation, the H19 locus becomes extensively demethylated in the embryo, leaving only the H19 ICR sequence methylated on the paternal allele. During postimplantation development, de novo DNA methylation occurs at the region downstream of the H19 ICR through the H19 promoter to the gene body7,11,17. Because this methylation requires the presence of a paternally hypermethylated H19 ICR, the methylation seems to spread from the H19 ICR in the downstream direction.
We previously generated transgenic mice (TgM) carrying a YAC transgene in which the mouse H19 ICR fragment was inserted into the normally non-imprinted human beta-globin locus. Using these mice, we found that the Tg H19 ICR was not methylated in sperm, but was de novo methylated after fertilization, only when paternally inherited18. This result demonstrated that the H19 ICR has an intrinsic ability to acquire paternal allele-specific DNA methylation. In the establishment of post-fertilization imprinted methylation at the Tg H19 ICR, the 5′ portion of the H19 ICR was first methylated, and the 3′ portion became methylated later19. Furthermore, we identified a 118 bp sequence at the 5′ end of the H19 ICR that was responsible for post-fertilization DNA methylation20,21,22.
Based on these observations, we assumed that methylation at the H19 locus initiates from the 5′ portion of the H19 ICR during the post-fertilization period in a 118 bp sequence-dependent manner, which then spreads further toward the H19 promoter region during the post-implantation period. If this assumption is correct, we would predict that reversing the direction of the H19 ICR would result in reduced methylation of the H19 promoter region. This would allow us to investigate the relationship between methylation state of the H19 promoter and the expression of the H19 gene without altering the size of the H19 ICR itself or its imprinted methylation status.
In this study, we generated mice in which the orientation of the H19 ICR was inverted at the endogenous locus by genome editing. When the inverted ICR allele was paternally inherited, the methylation level of the H19 promoter was reduced and the H19 gene was derepressed, suggesting that methylation of the H19 promoter itself is required for complete repression of H19 expression. Unexpectedly, when the inverted allele was maternally inherited, the expression level of the H19 gene was lower than that of the WT allele, even though the H19 promoter remained fully hypomethylated. These observations, both expected and unexpected, should provide important insight into the mechanism of transcriptional regulation of imprinted genes in which polarity of the ICR plays a role within the locus.
Results
Generation of mutant mice carrying the H19 ICR-inverted allele
To investigate the effect of H19 ICR orientation on the imprinted expression and DNA methylation status of the H19 locus, inversion of the H19 ICR sequence was induced at the mouse endogenous locus by genome editing. To this end, gRNA and Cas9 protein were introduced into fertilized mouse eggs to cleave both sides of the ICR (Supplementary Fig. 1b). We also included single-stranded oligodeoxynucleotides (ssODNs) with sequences that correspond to the junctional sequences after correct inversion, expecting that this would facilitate the inverted ligation of the ICR fragment (Supplementary Fig. 1b). We introduced artificial restriction enzyme recognition sites into these ssODNs, allowing the use of the ssODNs in the inverted ligation reaction to be evaluated. Screening of F0 mice by PCR and DNA sequencing identified that although many of them carried a deletion between the two cleavage sites, three carried correctly inverted ICR alleles; two of those three carried the artificial enzyme sites (Supplementary Fig. 1c). We established mutant mouse lines (lines 21 and 23) from these two F0 mice.
H19 ICR inversion derepresses H19 gene expression on the paternal allele
First, we examined the effect of altering the orientation of the H19 ICR on expression of the imprinted gene in cis at the paternal allele. To distinguish the parental origin of the genes, we used single-nucleotide polymorphisms (SNPs) between mouse strains (C57B6/6J [B6] and JF1/Msf [JF1] background). To analyze H19 and Igf2 gene expression when the inverted ICR allele was inherited from the father, we crossed B6 males carrying the heterozygous mutant allele with wild-type JF1 females (Fig. 1a) and extracted RNA from embryonic tissues. RT-PCR products generated under quantitative amplification conditions were digested with restriction enzymes whose recognition sites are present only in the B6 allele (Fig. 1b). In the fetal liver (E12.5, E18.5), the paternal H19 gene was repressed on the wild-type allele, while it was reproducibly expressed from the inverted ICR allele in two independent litters, although the expression level was much lower than from the maternal wild-type allele (Fig. 1c and Supplementary Fig. 2, H19). In the placenta (E18.5), expression of H19 remained repressed on the inverted ICR allele, as it was on the wild-type allele. The Igf2 genes were expressed on the inverted ICR, as were the wild-type alleles, only when they were paternally inherited (Fig. 1c and Supplementary Fig. 2, Igf2). These results indicated that ICR orientation did not affect Igf2 gene activation on the paternal allele, but it did have an effect on the expression of H19 in a tissue-specific manner.
a Breeding scheme. In order to distinguish parental origin of the alleles by using SNPs between inbred mouse strains, inverted-ICR heterozygous male mice (WT/INV; C57BL/6J [B6] background) were mated with wild-type female mice (WT/WT; JF1/Msf [JF1]), and embryos were obtained. b, c The allele-specific expression of the Igf2 and H19 genes was examined by RFLP analysis. Total RNA was extracted from livers (at E12.5 and E18.5) and placentas (E18.5), and H19 and Igf2 gene transcripts were amplified by RT-PCR followed by Cac8I or BstUI digestions, respectively. Parental origin of transcripts was discriminated by allele-specific restriction sites. The asterisks in (b) indicate restriction sites introduced into primer sequence to monitor complete digestion of PCR products. The signal intensity of the bands was quantified and the ratio of H19 or Igf2 expression to that of Gapdh (arbitrary unit) was displayed on the graph. The means ± SD are shown for genes expression in E12.5 liver (*p < 0.05).
Effect of H19 ICR inversion on methylation level inside the H19 ICR
To determine whether derepression of paternal H19 caused by the inversion of the H19 ICR in fetal liver was due to a change in the DNA methylation status of the locus, we conducted bisulfite sequencing analysis of tissue samples. First, we analyzed the inside of the inverted ICR and found that it was highly methylated in both liver (E12.5 and E18.5) and placenta (E18.5), as in the wild-type ICR (Supplementary Fig. 3, regions I and II in WT, and regions III and IV in inverted ICR). Internal methylation of the ICR was stably maintained throughout development, and postnatal tail DNA was also highly methylated at the paternal allele (Supplementary Fig. 4). In addition, the methylation state was reprogrammed across generations and was consistent between two independent strains (Supplementary Fig. 4). Consistent with our previous observation that the H19 ICR has the intrinsic ability to establish and maintain imprinted DNA methylation after fertilization18,19,23, the inside of the H19 ICR was properly methylated regardless of its orientation within the locus.
Effect of H19 ICR inversion on methylation levels of the paternally inherited H19 locus
Next, we investigated whether derepression of H19 gene was due to a change in the DNA methylation status of the H19 promoter. In the liver (E12.5, E18.5), where H19 expression was derepressed, the methylation level of the H19 promoter (regions V, VI, and COBRA in Fig. 2a) was lower in the inverted ICR allele than in the wild-type allele in two independent litters (Fig. 2b, c and Supplementary Fig. 5). This observation indicated that inverting the orientation of the ICR caused a modest loss of DNA methylation at the H19 promoter, even though the H19 ICR itself was correctly methylated. On the other hand, in placenta, the H19 promoter remained properly methylated in the inverted ICR allele (Fig. 2b, c and Supplementary Fig. 5). Thus, reduced methylation of the H19 promoter coincided with derepression of the paternally derived H19 gene, suggesting that methylation of the H19 promoter is required for complete repression of the H19 gene.
a Map of wild-type and inverted-ICR alleles. Regions analyzed by bisulfite sequencing in (b) and COBRA in (c) were indicated by gray bars below the map. b, c DNA methylation status of the H19 promoter on the paternally inherited wild-type and inverted H19 ICR alleles in livers (E12.5 and E18.5) and placentas (E18.5). b Bisulfite sequencing. For analyses of paternally inherited wild-type (WT) allele, genomic DNA of tissues from INV(B6)mat/WT(JF1)pat embryos, which were identical to ones analyzed in Fig. 3c (E12.5, No. 5−7; E18.5, No. 13−16) were pooled. For analyses of paternally inherited inverted-ICR allele, genomic DNA of tissues from WT(JF1)mat/INV(B6)pat embryos, which were identical to ones in Fig. 1c (E12.5, No. 4−6; E18.5, No.12−13) were pooled. The parental origin of the alleles was determined by SNPs between B6 and JF1. Each horizontal row represents a single DNA template molecule. Methylated and unmethylated CpG motifs are shown as filled and open circles, respectively. The methylation levels (%) of CpGs excluding the allele-specific sites (*) are shown for each cluster and were statistically compared (WT vs inverted at region V, p = 0.1860 [E12.5 liver], region VI, p = 0.0174 [E12.5 liver], p = 0.0187 [E18.5 liver], p = 0.1566 [E18.5 placenta]). c COBRA. Genomic DNA of tissues from embryos which were analyzed in Fig. 1c or Supplementary Fig. 2 was pooled according to their genotypes (E12.5 WT/WT, No. 1−3, WT/INV, No. 4−6; E18.5 liver and placenta (left) WT/WT, No. 10−11, WT/INV, No. 12−13 in Fig. 1c; E18.5 placenta (right) WT/WT, No. 21-22, WT/INV, No. 23−24 in Supplementary Fig. 2). The PCR products were digested with HpyCH4IV (Hp) (+) to assess the methylation status of the original DNA. The asterisks indicate restriction sites introduced into primer sequence to monitor the complete digestion of PCR products. The signal intensity of the bands was quantified and the ratio of the intensity of the digested fragments (i.e., methylated, M) to that of the overall fragments (U + M) was calculated, as the value of WT/WT set as 1. The mean ± SD for three technical replicates was displayed on the graph (**p < 0.01).
We then looked for regions other than the H19 promoter where DNA methylation was altered by reversal of the H19 ICR. In both liver and placenta, although the methylation level of the upstream region tended to be higher in the inverted ICR than in the wild-type alleles (regions VII and XI in WT and inverted ICR alleles, respectively, in Supplementary Figs. 6 and 7), differences in methylation levels between wild-type and inverted alleles were not statistically significant. Immediate downstream region of the ICR was highly and equally methylated in the inverted ICR, as well as in the wild-type alleles, in both liver and placenta (regions VIII and XII in WT and inverted ICR alleles, respectively, in Supplementary Fig. 6). Even if the activity of the H19 ICR involved in establishing DNA methylation has polarity, it seems capable of introducing DNA methylation at the immediate vicinity in both directions. Based on reports that methylation of the H19 gene body may regulate expression of the H19 gene24, we also compared its methylation status between the wild-type and inverted ICR alleles in both liver and placenta, but found no significant differences (regions IX and X in Supplementary Fig. 6). Thus, the methylation status of these regions (immediate downstream of the H19 ICR and H19 gene body) does not appear to regulate paternal H19 gene expression because it was consistent between (wild-type and inverted ICR) alleles and tissues.
Based on these observations, we infer that DNA methylation of the paternally inherited H19 gene promoter is regulated by orientation-dependent H19 ICR activity, and that promoter hypermethylation is required for complete repression of the paternal H19 gene.
H19 ICR inversion represses the H19 gene expression on the maternal allele
We next examined the effect of inverting the H19 ICR on the maternal allele (Fig. 3a). Allele-discriminating gene expression analysis using SNPs (Fig. 3b) revealed that the expression level of the H19 gene from the inverted ICR allele was lower than that from the wild-type allele in both liver (E.12.5, 18.5) and placenta (E18.5) in two independent litters (Fig. 3c and Supplementary Fig. 8a); this result was confirmed by RT-qPCR (Fig. 3d and Supplementary Fig. 8b). We then analyzed the DNA methylation status of the H19 promoter and found that the promoter was hypomethylated on both the inverted and wild-type alleles (regions V and VI in Fig. 4 and Supplementary Fig. 9). The methylation level of the H19 gene body also did not differ between wild-type and inverted ICR alleles (regions IX and X in Fig. 4). Therefore, the orientation of the H19 ICR does not affect the methylation status of the H19 promoter or gene body on the maternal allele, and the methylation statuses of these regions are not likely to regulate the H19 expression on the maternal allele.
a Breeding scheme. In order to distinguish parental origin of the alleles by using SNPs between inbred mouse strains, inverted-ICR heterozygous female mice (WT/INV; B6 background) were mated with wild-type male mice (WT/WT; JF1) to obtain embryos. b, c The allele-specific expression of the Igf2 and H19 genes was examined by RFLP analysis. Total RNA was extracted from livers (at E12.5 and E18.5) and placentas (E18.5), and H19 and Igf2 gene transcripts were amplified by RT-PCR followed by Cac8I or BstUI digestions, respectively. Parental origin of transcripts was discriminated by allele-specific restriction sites. The asterisks in (b) indicate restriction sites introduced into primer sequence to monitor complete digestion of PCR products. The signal intensity of the bands was quantified and the ratio of H19 or Igf2 expression to that of Gapdh was calculated (arbitrary unit) and the means ± SD was displayed on the graph (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001). d The expression levels of the H19 gene in the samples analyzed in (c) were also examined by RT-qPCR. The ratio of H19 expression to that of Gapdh was calculated (arbitrary unit) and the means ± SD are shown (**p < 0.01, ***p < 0.001).
a Map of wild-type and inverted-ICR alleles. Regions indicated by gray bars below the map were analyzed by bisulfite sequencing in (b). b For analyses of maternally inherited wild-type (WT) allele, genomic DNA of tissues from WT(JF1)mat/INV(B6)pat embryos which were identical to ones analyzed in Fig. 1c (E12.5, No. 4−6; E18.5, No. 12−13) were pooled. For analyses of maternally inherited inverted allele, genomic DNA of tissues from INV(B6)mat/WT(JF1)pat embryos which were identical to ones in Fig. 3c (E12.5, No. 5−7; E18.5, No.13−16) were pooled. The parental origin of the alleles was determined by SNPs between B6 and JF1. Each horizontal row represents a single DNA template molecule. Methylated and unmethylated CpG motifs are shown as filled and open circles, respectively. The methylation levels (%) of CpGs excluding the allele-specific sites (*) are shown for each cluster. There was no significant difference in methylation levels between wild-type and inverted alleles, except for region VI in E18.5 placenta (p = 0.0300).
Effect of H19 ICR inversion on methylation levels of the maternally inherited H19 locus
We then conducted methylation analysis of inside of the H19 ICR and found that the sequence was hypomethylated in the inverted ICR, as well as in the wild-type alleles, in both liver and placenta. However, the extent of the hypomethylated region in the inverted ICR was smaller than in the wild-type ICR (regions I/II and III/IV in the WT and inverted ICR, respectively, in Supplementary Fig. 10). In addition, the methylation level in regions immediately downstream of the ICR was significantly higher in the inverted ICR allele than in the wild-type allele (regions VIII (WT) and XII (inverted) in Supplementary Figs. 10 and 11). Because this ectopic DNA methylation occurred in both liver and placenta, we hypothesized that it is responsible for the reduction in H19 expression from the maternally inherited inverted ICR allele. In other words, the downstream region of the H19 ICR may be involved in transcriptional regulation of the H19 gene in a DNA methylation-dependent manner. Such an activity has not been previously reported in this region of the locus.
Search for a transcriptional regulatory activity around the H19 downstream region
We then hypothesized that the activity of the previously unidentified H19 enhancer is located downstream of the ICR and becomes attenuated when it is methylated by ICR inversion, resulting in reduced H19 transcription. To test this hypothesis in cell transfection assays, we cloned the H19 ICR downstream sequence (ICR-DS in Supplementary Fig. 12) and linked it to H19 or SV40 promoters on luciferase reporter plasmids. However, the ICR-DS fragment did not show any transcriptional activity in transiently transfected MEF cells (Supplementary Fig. 12). Therefore, it seems unlikely that a transcriptional regulatory element in this region acquires DNA methylation in the maternally inherited inverted ICR allele and causes downregulation of H19 gene expression. This suggested that in the maternal allele, the direction of the H19 ICR affects the transcription of the H19 gene through a mechanism that is independent of DNA methylation.
Discussion
H19 gene encodes a long noncoding (lnc) RNA that is highly expressed in developing embryo, and then repressed immediately after birth except in muscle25. Because H19 lncRNA and its processed product, miR-675, are associated with suppression of fetal/placental growth and cell cycle regulation, abnormal expression of these RNAs may cause tumorigenesis and other diseases26,27,28,29. Thus, elucidation of the mechanism underlying transcriptional regulation of H19 is fundamental to clarifying the causes of these diseases and to develop new therapies for treatment.
In this study, we found that inversion of the paternally inherited H19 ICR decreased the methylation level of the H19 promoter and that the paternal H19 gene was concurrently expressed in fetal liver, while the H19 ICR itself remained properly methylated. Although the transcriptional expression level of a gene and its DNA methylation status affect each other, we believe that the reduced methylation of the H19 gene promoter in our mutant mice was not the consequence of ectopic H19 gene activation in the paternal inverted ICR allele. In previously reported mice with partial deletion of the H19 ICR sequence, the paternal H19 gene was derepressed (presumably due to a loss of transcriptional repressive activity of the H19 ICR), despite the fact that the H19 promoter in cis remained correctly methylated14,15, i.e., H19 transcription does not necessarily cause demethylation of the promoter. We, therefore, assume that the paternal H19 gene was derepressed as a result of reduced methylation of the H19 promoter caused by inversion of the orientation of the H19 ICR. In other words, hypermethylation of the H19 promoter is necessary for complete repression of the H19 gene. Taking these observations together, the repression of the paternal H19 gene seems to involve multiple mechanisms: the transcriptional repressive activity of the H19 ICR itself and the methylation of the H19 promoter. Formally, we cannot rule out the possibility that inversion of the H19 ICR placed the hypothetical silencer element too far away from the H19 gene in our mutant mice, resulting in derepression of the H19 gene independent of its promoter methylation status. In accordance with this notion, Gebert et al. reported that knock-in of the H19 ICR sequence at the paternal Afp locus repressed the gene even without changing promoter methylation16; however, this result may not be applicable here, as the Afp promoter contains far fewer CpGs and the distance to the H19 ICR is much shorter.
The effect of the H19 ICR inversion on H19 promoter methylation and gene expression in the paternal allele was tissue-specific, i.e., it was observed in liver but not in placenta. Epigenetic regulation at multiple imprinted loci differs between extraembryonic and embryonic tissues30,31,32,33, suggesting the existence of tissue-specific regulatory factors. The H19 locus may also be regulated by these factors.
On the other hand, H19 expression level was reduced on the maternally inherited inverted ICR allele, even though the H19 promoter remained hypomethylated, as on the WT allele. We, therefore, assumed that the DNA methylation status of transcriptional regulatory sequences other than the H19 promoter may have been altered on the maternal allele, resulting in H19 expression. Although we found a candidate sequence that was significantly more methylated on the maternal inverted ICR allele than on the WT allele immediately downstream of the H19 ICR, reporter assays revealed no transcriptional regulatory activity in this sequence. The maternally inherited H19 ICR sequence itself is required for full activation of the H19 gene34, and CTCF binding sites in the H19 ICR are essential for the formation of maternal allele-specific chromatin modification status and appropriate interaction between the enhancer and H19 promoter35,36,37. These processes may be somehow impaired by the inversion of the H19 ICR.
Although inversion of the H19 ICR altered the expression levels of H19 in both parental alleles, it had no obvious effect on the weights of either embryos or placentas (Supplementary Fig. 13). We, therefore, concluded that the change in H19 expression level induced by the H19 ICR inversion was not sufficient to affect embryonic growth.
In contrast to H19, the expression of Igf2 was not altered by inversion of the H19 ICR. The mouse H19 ICR contains four CTCF/cohesin-binding motifs8,9,38, and the binding of the complexes to sites on hypomethylated maternal H19 ICR represses Igf2 expression via its insulator activity. Insulator activity arises from the functional isolation of specific genomic regions by chromatin loops formed by association of two CTCF/cohesin sites39; the orientation of the CTCF binding sequence is normally in opposite directions and regulates the mode of loop formation. In the Pcdh and β-globin loci, reversing the orientation of one of the CTCF sites in either the promoter or enhancer region changed the topology of the loops and hence the pattern of gene expression40. Recent work showed that the maternal H19 ICR formed loop structures with several downstream CTCF clusters, each of which contains CTCF binding sites in the opposite direction to the ones within the H19 ICR41. The expression of maternally inherited Igf2 was not affected on the inverted ICR allele in our study, implying that the spatial structure of the locus may have a trivial effect on Igf2 gene regulation on the maternal allele or that the gross structure of the gene locus is not altered by inversion of the entire H19 ICR. It would be interesting to investigate whether the H19 ICR inversion altered the spatial structure of the locus. Curiously, slight derepression of Igf2 was observed on the inverted maternal allele in placenta (Fig. 3c and Supplementary Fig. 8a). This may be due to the unusual increase in methylation inside the ICR (E18.5 placenta, Supplementary Fig. 10b) and reduced binding to CTCF insulator protein, which then allows interaction between the enhancer and Igf2 promoter.
In this study, we hypothesized that H19 promoter methylation on the paternal allele is a result of methylation spreading downwards from the 118-bp sequence of the H19 ICR. Based on this, we analyzed the effect of inverting the H19 ICR on genomic imprinting in mice. Although the methylation level of the paternal H19 promoter was decreased as expected, the spreading of methylation, if any, may not have been rigorously unidirectional, as the region between the ICR and the H19 promoter remained highly methylated. Generally, it is accepted that the presence of transcription and the H3K36me histone mark in oocytes and sperm are prerequisite for de novo DNA methylation during gametogenesis42,43,44. It remains to be fully elucidated where de novo DNA methylation initiates in somatic cells (i.e., is there a DNA methylation origin that recruits methyltransferases?) and how DNA methylation states propagate to the surrounding regions. Our mutant mice may help to obtain insights into such molecular mechanisms and to understand the etiology of diseases involving DNA methylation abnormalities, such as genomic imprinting disorders and oncogenesis.
Methods
Mice
Mice were housed in a pathogen-free barrier facility in a 12 h light/12 h dark cycle, and fed standard rodent chow. Animal experiments were performed in a humane manner and approved by the Institutional Animal Experiment Committee of the University of Tsukuba. Experiments were conducted in accordance with the Regulation of Animal Experiments of the University of Tsukuba and the Fundamental Guidelines for Proper Conduct of Animal Experiments and Related Activities in Academic Research Institutions under the jurisdiction of the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Generation of “inverted-ICR” mice by CRISPR/Cas9 genome editing
The oligonucleotides were annealed, phosphorylated, and ligated to the BbsI site of pX330 (plasmid 42230; Addgene)45 for generating Cas9/sgRNA expression vectors. In the following sequences, overhanging nucleotides are shown in lowercase letters. The sequences for the 5′ border are 5′-caccGGCAGTAAGCTTTGGCGGGG-3′ and 5′-aaacCCCCGCCAAAGCTTACTGCC-3′; for the 3′ border they are 5′-caccGTTTCGGTGGACGCACGCACG-3′ and 5′-aaacCGTGCGTGCGTCCACCGAAAC-3′. We used ssODNs (IDT, Coralville, IA, USA) with sequences that correspond to the junctional sequences after correct inversion. The sequence of ssODN for 5′ junction is 5′-ctgtttgcccaccagctgctagccatcacctagtcctcaatgtcacgtactattacaatggccaaaacagactagacttgaccccaagagcccccctcgagcgcggcagtttctatgtctcccgcctataaccgattctgtattgagtttggattgaacagatctggctagcttgaggagtcccaaggcagaaggggacc-3′, and for 3′ junction that is 5′-atagagctagatctcttcttccagaaacaagttaggcatgcctttgtcaatctggggactgccagggcagaaagtacaatgagggcagtaagctttggatccgtgcgtccaccgaaaccccatagccataaaagcagaggctggggttcaaccattgcaatgtcccaggtaacctaggaactgtagcaagaagttgcaaa-3′ (artificially introduced XhoI and BamHI sites are underlined, respectively). The plasmids and ssODNs were microinjected into the pronuclei of fertilized eggs of C57BL/6J mice (Charles River Laboratories Japan, Kanagawa, Japan). Tail DNA from 27 founder offspring was screened by PCR and sequencing.
Preparation of embryos
Inverted-ICR mice (which was generated by using C57BL/6J strain, and has a genetic background of Mus musculus domesticus) were mated with wild-type JF1/Msf46 mice (which was provided by the RIKEN BRC through the National Bio-Resource Project of the MEXT, Japan, and of which genome is basically from Mus musculus molossinus) to distinguish the parental origin of the alleles in the offspring. Livers are obtained from E12.5 and E18.5 embryos, and placentas are recovered at E18.5. Each tissue was divided into two and used for the preparation of total RNA and genomic DNA, respectively.
Allele-specific expression analysis using restriction fragment length polymorphism (RFLP)
Inverted-ICR mice (C57BL/6J) carrying the heterozygous mutant allele were crossed with wild-type JF1/Msf mice. Total RNA was recovered from livers and placentas of embryos using ISOGEN (Nippon Gene, Tokyo, Japan) and converted to cDNA using ReverTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO, Osaka, Japan). PCR was performed using AmpliTaq Gold 360 Master Mix (Thermo Fisher Scientific, Waltham, MA, USA) and PCR primers listed in Supplementary Table 1 with α-32P-dCTP (NEG513H, PerkinElmer, Waltham, MA, USA) at quantitative amplification condition (94 °C for 3 min, followed by 21 cycles of 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 1 min). The amplified products were digested with Cac8I or BstUI, in order to discriminate the parental origin of the transcripts, and subjected to polyacrylamide gel electrophoresis. The restriction sites were also introduced into primer sequences so that the complete digestion of PCR products can be concomitantly monitored. Gels were dried, subjected to phosphorimaging on a Typhoon 8600 (GE healthcare, Chicago, IL, USA), and then subjected to quantitative analysis using ImageQuant. X-ray autoradiography was also performed. Uncropped and unedited images are shown in Supplementary Figs. 14−16.
RT-qPCR
Total RNA from livers or placentas of E12.5 or E18.5 embryos was converted to cDNA as described above. Quantitative amplification of cDNA was performed with the Thermal Cycler Dice (TaKaRa Bio, Shiga, Japan) using TB Green Premix EX TaqII (TaKaRa Bio). PCR primers are listed in Supplementary Table 1.
DNA methylation analysis by bisulfite sequencing
For DNA methylation analysis of paternally inherited inverted-ICR allele and maternally inherited wild-type allele, inverted-ICR male mice (C57BL/6J background, carrying the heterozygous mutant allele) and wild-type female mice (JF1/Msf) were mated and heterozygous mutant embryos were obtained. Conversely, for analysis of maternally inherited inverted-ICR allele and paternally inherited wild-type allele, heterozygous inverted-ICR females and wild-type males were mated and heterozygous mutant embryos were obtained. Genomic DNA extracted from tissues of embryos was pooled as described in Figure legends and digested with XbaI, and then treated with sodium bisulfite using the EZ DNA Methylation Kit (Zymo Research, Irvine, CA, USA) by following the manufacturer’s instruction. Subregions of the wild-type or inverted H19 ICR alleles were amplified by PCR using EpiTaq HS (TaKaRa Bio). The PCR products were subcloned into the pGEM-T Easy vector (Promega, Madison, WI, USA) for sequencing analyses. PCR primers are listed in Supplementary Tables 2 and 3. Sequencing results were analyzed by using Quantification tool for Methylation Analysis (QUMA, http://quma.cdb.riken.jp). We checked whether each C that is not present in the CpG motif was correctly converted to T, and used only results from clones with a CT conversion efficiency of 95% or higher.
DNA methylation analysis by Southern blotting
Genomic DNA extracted from tail tips of ~1-week-old animals was first digested by NheI and SacI, and then subjected to the methylation-sensitive enzyme BstUI. Following size separation in agarose gels, Southern blots were hybridized with α-32P-labeled probes and subjected to X-ray film autoradiography. Uncropped and unedited images are shown in Supplementary Fig. 15.
Combined bisulfite restriction analysis (COBRA)
Genomic DNA was pooled as described in Figure legends and treated with sodium bisulfite. PCR was performed using EpiTaq HS (TaKaRa Bio) and primers listed in Supplementary Tables 2 and 3 with α-32P-dCTP (Perkinelmer) under quantitative amplification condition (94 °C for 5 min, followed by 28 cycles of 94 °C for 1 min, 60 °C for 2 min, and 72 °C for 2 min). After purification with Autoseq G-50 columns (Cytiva, Tokyo, Japan), the aliquots of PCR products were digested with HpyCH4IV enzyme and subjected to polyacrylamide gel electrophoresis. The restriction sites were also introduced into primer sequence so that complete digestion of PCR products can be concomitantly monitored. Gels were dried, subjected to phosphorimaging on a Typhoon 8600, and then subjected to quantitative analysis using ImageQuant. X-ray autoradiography was also performed. Uncropped and unedited images are shown in Supplementary Fig. 14.
Reporter plasmid construction
Reporter plasmids were generated based on a PGV-B2 or PGV-P2 vectors (Toyo Ink, Tokyo, Japan). The H19 promoter and ICR-DS plus H19 promoter sequences were PCR amplified using the following primer sets and a template plasmid in which the H19 gene locus sequence is subcloned: 5′- TCGGCCTGTCGACTGCTGATGCTG-3′ and 5′- ACCCCCCCTCGAGCTCCCACACC-3′ (SalI and XhoI sites are underlined, respectively) or 5′- AACTGCCTCGAGCGTGCGTC-3′ and 5′- ACCCCCCCTCGAGCTCCCACACC-3′ (XhoI sites are underlined). The products were digested with appropriate enzymes, and ligated to the XhoI site of PGV-B2 to generate PGV-B2/H19pr or PGV-B2/ICRDS-H19pr plasmids, respectively. The ICR-DS sequence was PCR-amplified using the following primer set: 5′- AACTGCCTCGAGCGTGCGTC-3′ and 5′- AGCATCAGTCGACTAAAGGCCGAG-3′ (XhoI and SalI sites are underlined, respectively). The product, digested with XhoI and SalI, was ligated to the XhoI site of PGV-P2 to generate PGV-P2/ICRDS plasmid.
Cell culture and transfection
Mouse embryonic fibroblasts (MEF) cells, generated from JF1/C57BL6J F1 embryo (E13.5), were maintained in Dulbecco’s modified Eagle’s medium (08458-45, nacalai tesque, Kyoto, Japan) containing 10% fetal bovine serum (Biological Industries, Kibbutz Beit-Haemek, Israel) and penicillin-streptomycin (Sigma, St. Louis, MO, USA) on gelatin-coated dishes. For luciferase assays, MEF cells were seeded in gelatin-coated 24-well plates at a density of 2 × 104 cells/well 12 h prior to transfection. Equimolar amounts of test reporter plasmid, 25 ng of pCMV-β-Gal (where CMV is cytomegalovirus and β-Gal is β-galactosidase), and pUC19 (to make a total plasmid weight of 275 ng) were introduced by using 0.75 µl of Gene Juice transfection reagent (Millipore, Burlington, MA, USA). Forty-eight hours after transfection, cells were harvested, and luciferase and β-Gal activities (for the correction of variation in transfection efficiencies) were measured by using a Centro XS3 LB 960 Microplate Luminometer (Berthold Technologies, Bad Wilbad, Germany) and X-Mark spectrophotometer (Bio-Rad, Hercules, CA, USA), respectively.
Statistics and reproducibility
Statistical analysis was performed by two-tailed unpaired Student’s t-test and Welch’s t-test using Prism 7 (GraphPad Software). The means and standard deviation were indicated. In bisulfite sequencing analysis, Mann−Whitney U-test was performed using QUMA. p values < 0.05 were considered significant. The sample numbers were described in each figure legend.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Source data for the graphs are available in Supplementary Data 1. Sanger sequencing data for bisulfite sequencing analysis are available from Dryad (https://doi.org/10.5061/dryad.wdbrv15qb)47. Any remaining information can be obtained from the corresponding author upon reasonable request.
Code availability
The data were plotted graphically by using Prism 7 (GraphPad Software). No custom code was used in this study.
References
Plasschaert, R. N. & Bartolomei, M. S. Genomic imprinting in development, growth, behavior and stem cells. Development 141, 1805–1813 (2014).
Tucci, V., Isles, A. R., Kelsey, G., Ferguson-Smith, A. C. & Erice Imprinting, G. Genomic imprinting and physiological processes in mammals. Cell 176, 952–965 (2019).
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harbor Perspect. Biol. https://doi.org/10.1101/cshperspect.a018382 (2014).
Farhadova, S., Gomez-Velazquez, M. & Feil, R. Stability and lability of parental methylation imprints in development and disease. Genes https://doi.org/10.3390/genes10120999 (2019).
Bartolomei, M. S., Webber, A. L., Brunkow, M. E. & Tilghman, S. M. Epigenetic mechanisms underlying the imprinting of the mouse H19 gene. Genes Dev. 7, 1663–1673 (1993).
Ferguson-Smith, A. C., Sasaki, H., Cattanach, B. M. & Surani, M. A. Parental-origin-specific epigenetic modification of the mouse H19 gene. Nature 362, 751–755 (1993).
Tremblay, K. D., Saam, J. R., Ingram, R. S., Tilghman, S. M. & Bartolomei, M. S. A paternal-specific methylation imprint marks the alleles of the mouse H19 gene. Nat. Genet. 9, 407–413 (1995).
Bell, A. C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405, 482–485 (2000).
Hark, A. T. et al. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 405, 486–489 (2000).
Srivastava, M. et al. H19 and Igf2 monoallelic expression is regulated in two distinct ways by a shared cis acting regulatory region upstream of H19. Genes Dev. 14, 1186–1195 (2000).
Tremblay, K. D., Duran, K. L. & Bartolomei, M. S. A 5’ 2-kilobase-pair region of the imprinted mouse H19 gene exhibits exclusive paternal methylation throughout development. Mol. Cell Biol. 17, 4322–4329 (1997).
Thorvaldsen, J. L., Duran, K. L. & Bartolomei, M. S. Deletion of the H19 differentially methylated domain results in loss of imprinted expression of H19 and Igf2. Genes Dev. 12, 3693–3702 (1998).
Horii, T. et al. Successful generation of epigenetic disease model mice by targeted demethylation of the epigenome. Genome Biol. 21, 77 (2020).
Drewell, R. A. et al. Deletion of a silencer element disrupts H19 imprinting independently of a DNA methylation epigenetic switch. Development 127, 3419–3428 (2000).
Ideraabdullah, F. Y. et al. Novel cis-regulatory function in ICR-mediated imprinted repression of H19. Dev. Biol. 355, 349–357 (2011).
Gebert, C., Rong, Q., Jeong, S., Iben, J. & Pfeifer, K. H19ICR mediated transcriptional silencing does not require target promoter methylation. Biochem. Biophys. Res. Commun. 476, 121–126 (2016).
Tomizawa, S. et al. Dynamic stage-specific changes in imprinted differentially methylated regions during early mammalian development and prevalence of non-CpG methylation in oocytes. Development 138, 811–820 (2011).
Tanimoto, K. et al. Genomic imprinting recapitulated in the human beta-globin locus. Proc. Natl Acad. Sci. USA 102, 10250–10255 (2005).
Matsuzaki, H. et al. De novo DNA methylation through the 5′-segment of the H19 ICR maintains its imprint during early embryogenesis. Development 142, 3833–3844 (2015).
Matsuzaki, H. et al. Synthetic DNA fragments bearing ICR cis elements become differentially methylated and recapitulate genomic imprinting in transgenic mice. Epigenetics Chromatin 11, 36 (2018).
Matsuzaki, H. et al. Recapitulation of gametic DNA methylation and its post-fertilization maintenance with reassembled DNA elements at the mouse Igf2/H19 locus. Epigenetics Chromatin 13, 2 (2020).
Hirakawa, K., Matsuzaki, H. & Tanimoto, K. Transient establishment of imprinted DNA methylation of transgenic human IC1 sequence in mouse during the preimplantation period. Hum. Mol. Genet. 29, 3646–3661 (2021).
Matsuzaki, H., Okamura, E., Shimotsuma, M., Fukamizu, A. & Tanimoto, K. A randomly integrated transgenic H19 imprinting control region acquires methylation imprinting independently of its establishment in germ cells. Mol. Cell Biol. 29, 4595–4603 (2009).
Jinno, Y. et al. Establishment of functional imprinting of the H19 gene in human developing placentae. Nat. Genet. 10, 318–324 (1995).
Gabory, A., Jammes, H. & Dandolo, L. The H19 locus: role of an imprinted non-coding RNA in growth and development. BioEssays: N. Rev. Mol., Cell. Dev. Biol. 32, 473–480 (2010).
Yoshimizu, T. et al. The H19 locus acts in vivo as a tumor suppressor. Proc. Natl Acad. Sci. USA 105, 12417–12422 (2008).
Keniry, A. et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat. Cell Biol. 14, 659–665 (2012).
Lecerf, C., Le Bourhis, X. & Adriaenssens, E. The long non-coding RNA H19: an active player with multiple facets to sustain the hallmarks of cancer. Cell Mol. Life Sci. 76, 4673–4687 (2019).
Chang, S. & Bartolomei, M. S. Modeling human epigenetic disorders in mice: Beckwith−Wiedemann syndrome and Silver−Russell syndrome. Dis. Model Mech. https://doi.org/10.1242/dmm.044123 (2020).
Lewis, A. et al. Imprinting on distal chromosome 7 in the placenta involves repressive histone methylation independent of DNA methylation. Nat. Genet. 36, 1291–1295 (2004).
Sato, S., Yoshida, W., Soejima, H., Nakabayashi, K. & Hata, K. Methylation dynamics of IG-DMR and Gtl2-DMR during murine embryonic and placental development. Genomics 98, 120–127 (2011).
Inoue, A., Jiang, L., Lu, F., Suzuki, T. & Zhang, Y. Maternal H3K27me3 controls DNA methylation-independent imprinting. Nature 547, 419–424 (2017).
Hanna, C. W. et al. Endogenous retroviral insertions drive non-canonical imprinting in extra-embryonic tissues. Genome Biol. 20, 225 (2019).
Thorvaldsen, J. L., Mann, M. R., Nwoko, O., Duran, K. L. & Bartolomei, M. S. Analysis of sequence upstream of the endogenous H19 gene reveals elements both essential and dispensable for imprinting. Mol. Cell Biol. 22, 2450–2462 (2002).
Engel, N., Thorvaldsen, J. L. & Bartolomei, M. S. CTCF binding sites promote transcription initiation and prevent DNA methylation on the maternal allele at the imprinted H19/Igf2 locus. Hum. Mol. Genet. 15, 2945–2954 (2006).
Han, L., Lee, D. H. & Szabo, P. E. CTCF is the master organizer of domain-wide allele-specific chromatin at the H19/Igf2 imprinted region. Mol. Cell Biol. 28, 1124–1135 (2008).
Engel, N., Raval, A. K., Thorvaldsen, J. L. & Bartolomei, S. M. Three-dimensional conformation at the H19/Igf2 locus supports a model of enhancer tracking. Hum. Mol. Genet. 17, 3021–3029 (2008).
Wendt, K. S. et al. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 451, 796–801 (2008).
Ghirlando, R. & Felsenfeld, G. CTCF: making the right connections. Genes Dev. 30, 881–891 (2016).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Lleres, D. et al. CTCF modulates allele-specific sub-TAD organization and imprinted gene activity at the mouse Dlk1-Dio3 and Igf2-H19 domains. Genome Biol. 20, 272 (2019).
Chotalia, M. et al. Transcription is required for establishment of germline methylation marks at imprinted genes. Genes Dev. 23, 105–117 (2009).
Xu, Q. et al. SETD2 regulates the maternal epigenome, genomic imprinting, and embryonic development. Nat. Genet. 51, 844–856 (2019).
Shirane, K., Miura, F., Ito, T. & Lorincz, M. C. NSD1-deposited H3K36me2 directs de novo methylation in the mouse male germline and counteracts Polycomb-associated silencing. Nat. Genet. 52, 1088–1098 (2020).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Koide, T. et al. A new inbred strain JF1 established from Japanese fancy mouse carrying the classic piebald allele. Mamm. Genome 9, 15–19 (1998).
Matsuzaki, H., Miyajima, Y., Fukamizu, A. & Tanimoto, K. Data from: Orientation of mouse H19 ICR affects imprinted H19 gene expression through promoter methylation-dependent and -independent mechanisms. Dryad, Dataset, https://doi.org/10.5061/dryad.wdbrv15qb (2021).
Acknowledgements
We thank Chie Kodama for providing technical support. This work was supported in parts by JSPS KAKENHI grants (grant numbers JP20K06481 to H.M., JP19H03134 to K.T., and JP20K21360 to K.T.), MEXT KAKENHI grants (grant number JP20H05379 to H.M.), and a research grant from the Takeda Science Foundation to K.T.
Author information
Authors and Affiliations
Contributions
H.M., Y.M., and K.T. designed the experiments. H.M. and Y.M. performed the experiments. H.M., Y.M., and K.T. analyzed the data. A.F. provided scientific advice and materials. H.M. and K.T. wrote the manuscript. All authors reviewed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Communications Biology thanks Maxim Greenberg and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editors: Patrick Murphy and George Inglis.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Matsuzaki, H., Miyajima, Y., Fukamizu, A. et al. Orientation of mouse H19 ICR affects imprinted H19 gene expression through promoter methylation-dependent and -independent mechanisms. Commun Biol 4, 1410 (2021). https://doi.org/10.1038/s42003-021-02939-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-02939-9
- Springer Nature Limited