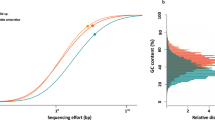
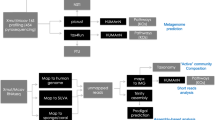
Abstract
Marine Porifera host diverse microbial communities, which influence host metabolism and fitness. However, functional relationships between sponge microbiomes and metabolic signatures are poorly understood. We integrate microbiome characterization, metabolomics and microbial predicted functions of four coexisting Mediterranean sponges –Petrosia ficiformis, Chondrosia reniformis, Crambe crambe and Chondrilla nucula. Microscopy observations reveal anatomical differences in microbial densities. Microbiomes exhibit strong species-specific trends. C. crambe shares many rare amplicon sequence variants (ASV) with the surrounding seawater. This suggests important inputs of microbial diversity acquired by selective horizontal acquisition. Phylum Cyanobacteria is mainly represented in C. nucula and C. crambe. According to putative functions, the microbiome of P. ficiformis and C. reniformis are functionally heterotrophic, while C. crambe and C. nucula are autotrophic. The four species display distinct metabolic profiles at single compound level. However, at molecular class level they share a “core metabolome”. Concurrently, we find global microbiome-metabolome association when considering all four sponge species. Within each species still, sets of microbe/metabolites are identified driving multi-omics congruence. Our findings suggest that diverse microbial players and metabolic profiles may promote niche diversification, but also, analogous phenotypic patterns of “symbiont evolutionary convergence” in sponge assemblages where holobionts co-exist in the same area.
Similar content being viewed by others
Introduction
Sponges are ubiquitous and functionally key components of marine communities, providing habitats for a variety of species, and coupling the benthic and pelagic ecosystems through intense seawater filtration and nutrient re-cycling1,2. These organisms have often been investigated as model holobiont systems to understand host-microbe and metabolic interactions, in the context of functional biological entities3,4,5. From this perspective, the present work focused on four widespread Mediterranean sponge holobionts (Petrosia ficiformis, Chondrosia reniformis, Crambe crambe and Chondrilla nucula) coexisting in high abundances across a study area of the Tyrrhenian Sea. These species were selected on the basis of distinctive bio-ecological potential range traits, involving reproductive strategies, microbial abundance, preferred habitat, general morphology, coloration, growth-type and allelochemistry (summarized in Table 1 and Fig. 1).
Petrosia ficiformis: A Classical massive form. B Reticular and bleached form (darkest habitats). C Reticular and colored form. D Chondrilla nucula in its preferred habitat (photic zone). Chondrosia reniformis: E Dark type, F Brownish type, G Bleached type (darkest habitats). H Crambe crambe patch in its preferred habitat (photic zone). The phenotypes collected to perform our study were represented in the pictures A, E, D, H for P. ficiformis, C. reniformis, C. nucula and C. crambe respectively.
Sponges frequently host dense microbial assemblages as intracellular symbionts in specialized cells (bacteriocytes) and/or as free unicellular partners in the mesohyl, which in some cases account for up to 40-50% of the biomass6,7. Regarding the quantitative importance of the microbial compartment, sponges have been classified as “low microbial abundance”—LMA (105 to 106 microbial cells/g tissue) and “high microbial abundance”—HMA (ca. 108 to 1010 microbial cells/g tissue) species6,8,9. In general, HMA sponges –like P. ficiformis, C. reniformis, and C. nucula– compared with those LMA –such as C. crambe– display a denser mesohyl, lower pumping rates, and exhibit different mechanisms to process carbon and nitrogen compounds10,11,12. Notwithstanding this dichotomy, both groups of sponge holobionts display equivalent capabilities to colonize and flourish in the same range of habitat types13,14.
The diversity, structure and composition of sponge microbiota communities have been widely investigated in the past decades6,7. Similar microbiomes have been frequently reported in the same host species from geographically distant locations15, but still, there are cases (e.g. P. ficiformis) reporting microbial differences across different biogeographic regions16,17,18,19. Depending on the species and the environmental context, hosts can acquire microbial associates by two major mechanisms: via horizontal acquisition from the surrounding seawater (as majorly described for P. ficiformis), or by vertical transmission from parents to offspring in diversified reproductive strategies and modes (e.g., through brooding larvae as in C. crambe, or within oocytes in C. nucula, C. reniformis)20,21,22,23,24,25. Assemblages acquired horizontally are in general more spatially and temporally variable (depending on time, location and environmental conditions), and can occur simultaneously involving similar taxa in diverse host species. Instead, inherited symbionts are more stable and are often species-specific15,20,24. All in all, it is broadly accepted that sponges combine different ratios of both microbial input types, relying on the ecological needs20,24; and that symbiont prokaryote communities reflect high degree of host species-specificity based on collections worldwide6,7,15.
Microbial symbionts can supply nutrients and energy to their hosts, while processing waste products, and can as well produce secondary metabolites, useful to avoid predators and fouling, or to successfully compete for space26. Moreover, sponge microbiomes are involved in the C, N and S geochemical cycles, and in the nutrient re-cycling of the sponge loop, where DOM is filtered by sponges and subsequently released as POM in the form cellular detritus, thus becoming available for the neighboring trophic web levels2,27. Microhabitat acclimating aspects could be related to host-microbiome-metabolome interactions, and their joint potential. Petrosia ficiformis and C. reniformis for instance, flourish in large populations, either in illuminated areas as pigmented forms, or in shaded or even dark caves as bleached morphotypes often in reticulated growth (e.g., P. ficiformis)28,29. While C. crambe seem more abundant in the presence of some light or at most shade but not dark, and are colorful and charged of secondary metabolites to deter putative predators and competitors30,31. Chondrilla nucula still, are clearly photophilic and intolerant to shade-to-dark conditions32,33. There are still many gaps of knowledge on the metabolic functional role of diversified microbiomes in how these and other holobionts are adapted to similar or distinct environments34,35,36,37,38. Sponge microbiomes are known to benefit the holobiont via metabolic pathways of specific taxa9,26,39. In particular, cyanobacteria have been investigated for their involvement in the carbon cycle, and for supplying their hosts with >50% of energy requirements in certain cases –as suggested for C. nucula32. Nitrogen metabolism in sponge holobionts has been mainly attributed to ammonia-oxidizing archaea9,40, while associated nitrogen fixation is considered to be a negligible source of nitrogen in Mediterranean sponges41. The exclusively sponge-associated genus Poribacteria, instead, has been found to be involved in nitrification and denitrification pathways42. In this regard, C. reniformis has been studied for its capability to oxidize exogenous ammonium, mediated by its associated microbial community43. Other sponge holobionts (e.g. P. ficiformis) undergo dark fixation processes that may involve anaplerotic carbon assimilation, which seem to be performed by Acidobacteria, Alphaproteobacteria and Chloroflexi44. Metabolic integration studies within several sponge holobionts reported carnitine derived from sponge cell debris, as a common microbial source of C, N and energy45; in concomitance with a clear involvement of microbial genes in the production of (primary and secondary) metabolites required for the host46,47,48. Predicted genomic functions of microbiomes linking metabolic signatures have been previously investigated44,49,50,51, but it remains a poorly studied topic with still the requirement of rigorous experimental validations.
Generally, there is an equivocal belief that variance in host metabolome can be explained by microbiome modulation52, but to what extent and which are the major drivers, and how this happens in marine holobionts is still poorly understood52. We found a wide gap in addressing microbiome-metabolome relatedness in Porifera, with regards the linkage between the microbial structure and taxonomy and the metabolic profiling phenotype. Therefore, networks of microbial taxa supplying access to metabolic pathways involved in phenotypes that afford host ecological success, are at the moment largely undiscovered, even if recent approaches are providing eager advances in the topic48,49.
In this research we combined microscopy, metabarcoding and metabolomics approaches, along with predicted functions algorithms, to investigate the relationships between sponge microbiomes and metabolic signatures, in the context of host specificity, and considering several distinctive bio-ecological traits, characterizing the four holobiont systems selected. Multivariate correlation and distance-based methods were applied to characterize the strength of the dynamic relationship between the microbiome and the metabolome, and to identify key contributors to the overall concordance. To the best of our knowledge ours is one of the few studies analyzing microbiome-metabolomics relatedness in sponge holobionts, representing a novelty in the field of sponge ecology.
Results
Transmission electron microscopy observations
Petrosia ficiformis and C. nucula revealed numerous microbial cells with different morphologies, enclosed in host cells—“bacteriocytes” (Fig. S2A, B; G, H; Fig. S3B). Abundant cyanobacterial-like cells were remarkably present in C. nucula, whereas a lower number of cyanobacteria was observed in P. ficiformis (Fig. S2A, B; Fig. S3B–D). C. crambe and C. reniformis were characterized by the absence of bacteriocytes and by the presence of prokaryotic cells occupying the mesohyl, which were morphologically similar, in the sponge C. crambe (Fig. S2C–F).
Microbiome diversity and community composition
Sequence data were rarefied to 7038 reads (Fig. S4) per sample resulting in 408204 reads distributed over 58 samples (sample details in Table S1). Sequence reads ranged between a minimum of 7 and a maximum of 64215, distributed over 1181 ASVs. Petrosia ficiformis reported the highest number of ASVs (total = 425; mean = 174), followed by C. crambe (total = 343; mean = 95), C. reniformis (total = 225; mean = 106) and C. nucula (total = 121; mean = 79) (Fig. 2A, Fig. S6), and this pattern was also partially observed for the Shannon diversity index. Indeed C. nucula displayed in this case a higher alpha diversity (Fig. 2B). Microbial diversity in the seawater was higher with respect to any sponge investigated (Kruskal–Wallis, p < 0.01; Tables S2–S3). PCoA analysis based on Bray-Curtis dissimilarity (relative abundance) highlighted a clear separation in microbial composition among sponge species and with respect to the seawater (Fig. 2C, PERMANOVA, p < 0.01; Table S4). Similar results were highlighted also by nMDS analysis based on the Jaccard index (presence/absence), although with a certain overlap between C. crambe and the seawater (PERMANOVA, p < 0.01, Fig. 2D; Table S5).
A Average number of ASVs per sample versus total ASVs across all samples. B Bacterial diversity (as Shannon index) across the different sponge species. PCoA and nMDS (2D stress = 0.085) of beta diversity ordinations based on Bray–Curtis (C) and Jaccard resemblances (D). Ellipses represent 95% confidence interval. Observed ASVs boxplots are reported in Fig. S6.
Compositional RPCA (Robust Principal Component Analysis) based on Aitchison distance revealed four clusters of microbiomes significantly different among the four sponge species (Fig. 3A; PERMANOVA, p < 0.01, Table S6), and the twenty most significant ASVs driving sample grouping in the ordination space were identified (Fig. 3A). Log ratios from DEICODE feature-loadings grouped by lowest best taxonomy annotation level, and which best explained the separation of sample groupings along axis 2 in the RPCA space, were composed by 81 ASVs in the numerator and 50 in the denominator (Fig. 3C). In the numerator these differential taxa belonged to Subgroup 6 (Acidobacteria), SBR1031 (Chloroflexi), Bdellovibrio (Proteobacteria) and in the denominator to Rhodothermaceae, Cyanobiaceae, PAUC34f and EC94 (Betaproteobacteriales). C. reniformis displayed the highest log ratios, followed by P. ficiformis, C. nucula and C. crambe (Fig. 3B), and the differences in log ratios of the selected bacterial consortium were statistically significant for every comparison (Welch’s Test, p < 0.01, Table S7).
A Compositional RPCA biplot of beta diversity of bacterial communities associated with the four sponge species based on Aitchison distance. Sponge species are depicted by colors and points represent individuals. Twenty most relevant ASVs driving differences in the ordination space are illustrated by colored vectors labeled with the respective taxonomy, reported to the lowest best taxonomic annotation. B Box plots of log-ratios of the microbial balance generated by the selected bacterial consortium plotted across the four sponge species along axis 2 of the compositional RPCA. C Feature rankings, with the ASVs selected as numerator and denominator taxa to generate the microbial balance that maximized group separations by sponge species, along axis 2 of the compositional RPCA. *For other dimensional views, including the third ordination component PC3 of the RPCA plot (A) see Supplementary Fig. S7.
A total of 27 microbial phyla were annotated across the four sponge species, distributed in 49 classes, 109 orders, 151 families and 225 genera (Fig. 4, Fig. S5, Table S8). Betaproteobacteriales, Flavobacteriales, SAR11 (Alphaproteobacteria), SAR202 clade (Chloroflexi) and Synechococcales were the most abundant bacterial orders across all samples. In P. ficiformis the microbiome was composed by 22 phyla, 40 classes, 68 orders, 81 families, and 99 genera. The five most abundant bacterial orders were Caldilineales (20%), SAR202 clade (Chloroflexi—15%), Gemmatimonadetes (8%), TK17 (Chloroflexi—5%) and Microtrichales (4%). Cyanobacteria and Archaea were only present in low percentages and accounted for 1.6 and 0.65% respectively.
Taxa were represented at the best annotated taxonomic rank (i.e. order). Each label in the x axis represents a sample (details reported in the Table S1). *For whole bacterial community including rare background taxa see Fig. S5.
C. reniformis harbored 18 phyla, 29 classes, 49 families, 54 orders and 61 genera. The most abundant bacterial orders were SAR202 clade (Chloroflexi—15%), Nitrosococcales (10%), Subgroup 6 (Acidobacteria—9%), SBR1031 (Chloroflexi—7%) and Myxococcales (7%). Archaea and Cyanobacteria accounted together <1%. The microbiome of C. crambe was composed by 18 phyla, 28 classes, 69 orders, 96 families and 133 genera. This species was dominated by an ASV belonging to the order Betaproteobacteriales, which accounted for ~73%. Cyanobacteria accounted for 10%, mostly belonging to the order Synechococcales. Finally, C. nucula were characterized by the presence of 16 prokaryotic phyla, 27 classes, 46 orders, 50 families and 55 genera. Synechococcales (Cyanobacteria) was the most abundant taxa associated with this species (18%). Other relevant taxa of C. nucula were Caldilineales (10%), Gammaproteobacteria (7%), TK17 (Chloroflexi, 7%) and Oceanospirillales (6%). Finally, Archaea accounted for <0.5%.
Seawater samples were characterized by the presence of 11 prokaryotic phyla, 17 classes, 57 orders, 79 families and 119 genera. The five most relevant ASVs belonged to Flavobacteriales (31%), SAR11 clade (Alphaproteobacteria—14%), Rhodobacterales (11%), Synechococcales (11%) and SAR86 clade (Gammaproteobacteria—8%). No archaea were detected in the seawater.
C. nucula displayed the highest relative abundance of cyanobacteria (7–26%), represented by genus Synechococcus (spongiarum group). Synechococcus (strain CC9902) was also dominant in C. crambe (4–19%), along with genus Cyanobium (~1%). In P. ficiformis the most abundant cyanobacterial taxa belonged to family Cyanobiaceae (0.1–0.5%), followed by genera Synechococcus (strain CC9902), Cyanobium and Prochlorococcus (~0.1%). In C. reniformis, cyanobacteria accounted for a very low fraction of the total prokaryotic assemblages and were represented by the genus Synechococcus (strain CC9902) (~0.1%).
The four sponge species did not share any ASV (100% of the samples). C. nucula displayed the largest core microbiome in terms of ASV richness (55 ASVs), followed by C. reniformis (47), P. ficiformis (38), and C. Crambe (13; Fig. 5). Crambe crambe was the only host sharing core taxa with the seawater, sharing eight core ASVs. The size of the core microbiomes (percentage of the core microbiome with respect to the total) accounted for ~95% in C. nucula, ~81% in C. crambe, ~74% in C. reniformis and ~46%, in P. ficiformis (Fig. 5). The output of the network analysis indicated that each sponge species was characterized by the presence of several exclusive ASVs (Fig. 6A). In particular, P. ficiformis displayed the highest number of exclusive ASVs (337) followed by C. reniformis (179), C. crambe (146) and C. nucula with (99) (Fig. 6B). C. crambe shared the largest number of ASVs with seawater (175), followed by P. ficiformis, C. reniformis and C. nucula (64, 36 and 20 ASVs, respectively; Fig. 6B).
A Network showing bacterial ASVs shared between the four sponge species and the seawater. B Venn diagram showing the numbers of shared and exclusive ASVs among the four sponge species and the surrounding seawater. C PCA ordination of putative functions of microbiomes of each sponge species. Axes represent the first two components with the percentage of variation explained. D Putative functions of the microbiomes of the four sponge species.
SIMPER analysis performed to estimate the contribution of each microbial taxon (%) to the dissimilarity between groups, confirmed high dissimilarity percentages (~98–99%) among microbiome composition of the four sponge species studied. Remarkable dissimilarities were also reported between all sponge microbiomes and a prokaryotic assemblages of the seawater. Instead, similarity of the microbiomes among individuals of the same sponge species was 76.4% for C. nucula, 75.8% for C. crambe, 68.5% for C. reniformis, 54.3% for P. ficiformis, and 61.8% for the seawater.
Putative functions of prokaryotic assemblages
The ASV table of the full dataset was converted into putative functions table via cross-annotation against the Faprotax database53 (https://pages.uoregon.edu/slouca/LoucaLab/archive/FAPROTAX/lib/php/index.php). The PCA ordination revealed three main clusters of putative functions patterns: one composed by C. crambe specimens, one by C. nucula and another one by C. reniformis and P. ficiformis (Fig. 6C; PERMANOVA, p < 0.05; Table S9).
The analysis of the putative functions revealed that the cluster composed by P. ficiformis and C. reniformis (Fig. 6D) was mostly related to functions of chemo-heterotrophy and aerobic chemo-heterotrophy (accounting together for 56 and 54%, respectively). Other relevant functions were nitrification-related processes, which accounted for ~32%, in P. ficiformis and ~20% in C. reniformis. C. reniformis was further characterized by predatory or exoparasitic related functions (16%) and fermentation (9%). The other two clusters formed by C. crambe and C. nucula were dominated by phototrophy, photo-autotrophy and cyanobacteria-related functions (>75%), with heterotrophy representing ~15% in both species. Nitrification-related functions accounted for ~4% in C. nucula and were negligible in C. crambe (ca. 1%). Other minor functions were exclusive of C. crambe.
Metabolomics profiling
Metabolomics analysis carried out on 37 samples discriminated ~5000 chemical features grouped in 33 metabolite classes (further details in Table S12). Phosphatidylcholines (PC) were the most abundant organic compound class (Fig. 7A; Table S10), ranging from 33 to 49% in P. ficiformis, 63–66% in C. reniformis, 19–59% in C. crambe and 57–66% in C. nucula, followed by tryglicerides (TG) ranging from 18 to 38% in P. ficiformis, 6–9% in C. reniformis, 0.6–7% in C. nucula and 0.05–55% in C. crambe. Phosphatidylethanolamines (PE) ranged from 7 to 16% in P. ficiformis, 8–11% in C. reniformis, 4–14% in C. crambe and 3–7% in C. nucula. Another relevant class of compounds were ceramides (Cer) which ranged from 2 to 8% in P. ficiformis, 2–4% in C. reniformis, 6–70% in C. crambe and 4–8% in C. nucula. Wax esters (WE) and methyl phosphatidylcholines (MePC) were detected only in the sponges P. ficiformis and C. nucula, ranging from 0.7 to 3% and 0.5 to 17%, respectively (Fig. 7A).
A Metabolic profiles plotted by molecular classes in the four sponge species. Data were plotted by individual to show the intraspecific variability (for sample details see Table S1). B Network of chemical features associated with the four sponge species. C Network of presence/absence data of chemical classes associated with each sponge. D PCA ordination plot showing metabolomics feature data from the four different sponge species. Each dot represents a sample. Each arrow represents a class of metabolites.
Network analysis highlighted major differences in the metabolomic profiles of the four sponges investigated (dissimilarity ~90%), with many chemical features being exclusive of each species, and a few being shared (Fig. 7B). Metabolic/chemical profiles based on molecular classes (functional biochemical affinity) were compared by presence/absence across species and visualized through network analysis (Fig. 7C). The four sponge species shared 14 molecular classes, including: triglycerides (TG), monoglycerides (MG), diglycerides (DG), phosphatidylcholines (PC), phosphatidylethanolamines (PE), phosphatidylserines (PS), phosphatidylglycerols (PG), coenzymes (Co), cholesterol esters (ChE), ceramides (Cer), ceramides phosphate (CerP), sitosterol esters (SiE), lysophosphatidylcholines (LPC), zymosterol ester (ZyE). Seven molecular classes (sulfatides ST, sphingosines SPH, wax esters WE, phosphatidylmethanols PMe, methyl phosphatidylcholines MePC, acyl carnitines AcCa, hexosylceramides Simple Glc Series) were shared only among P. ficiformis, C. nucula and C. reniformis, while three (hex1cer, hex2cer, phosphatidylinositols PI) were common to P. ficiformis, C. reniformis and C. nucula. Lysophosphatidylglycerols (LPG) were only shared between P. ficiformis and C. reniformis. Stigmasterol esters (StE) were found in P. ficiformis, C. reniformis and C. crambe, while sphingomyelins (SM) were detected in C. reniformis, C. crambe and C. nucula. A few molecular classes were exclusive in each sponge species. C. nucula and C. crambe showed two exclusive molecular classes: monogalactosyl-diacylglycerol (MGDG) and acylGlcStigmasterol ester (AcHexStE) and phosphatidylinositols (PI) and lysophosphatidylinositol (LPI), respectively. P. ficiformis and C. reniformis were characterized by one exclusive molecular class each, campesterol ester (CmE) and lysophosphatidylethanolamines (LPE), respectively (Fig. 7C). Finally, differences in the overall metabolomics profiles among sponge species were highlighted by PCA (Fig. 7D) and PERMANOVA analyses (p < 0.05, Table S11).
Procrustes distance based multi-omics concordance
Strong global concordance between the microbiome and the metabolome was found when considering the overall sample set encompassing the four sponge hosts (Fig. 8), according to the symmetric Procrustes test (p = 1.5 e-5). Contrastingly, overall data structure association was not significant within any of the sponge species based on the respective paired PCAs (C. nucula – p = 0.795; C. reniformis – p = 0.84; C. crambe – p = 0.78; P. ficiformis – p = 0.59; Fig. 8).
A Full data set including all four sponge species (n = 37) and B Sponge species panels: Chondrilla nucula (n = 10), Chondrosia reniformis (n = 10), Crambe crambe (n = 9), Petrosia ficiformis (n = 8). There was significant overall correlation between the microbiome and the metabolome when considering the full dataset comprising all four sponge species, however this association faded at the single species.
Spearman and Sparse canonical correlation (sCCA) analyses
A total of 120 (95.2%) microbial genera and 33 (100%) metabolite classes revealed significant correlations in the full dataset (34.94% of total pairwise comparisons), including all four sponge species (Spearman, FDR controlled q-value < 0.05). This correlations pattern was strongly reduced when considering the subsets by the single sponge species. The number of significant correlations reported for represented microbial genera and metabolite classes were 12 (17%) and seven (21%) in C. crambe, seven (10%) and six (18%) in P. ficiformis, six (11%) and 12 (36%) in C. nucula, and four (8%) and six (18%) in C. reniformis (Spearman, FDR controlled q-value < 0.05). Overall multivariate correlation revealed significant results in the full dataset considering all four sponge species (permutation p-value = 0), and non-significance in the species subsets (C. reniformis permutation p-value = 0.995; C. nucula permutation p-value = 0.749; P. ficiformis permutation p-value = 0.131; C. crambe permutation p-value = 0.055). The canonical correlation was marginally higher in P. ficiformis (correlation: 0.989 [0.97–1]) than in C. crambe (correlation: 0.986 [0.955–1]), followed by C. nucula (correlation: 0.935 [0.925–0.998]), with non-significant appertaining differences due to overlapping confidence intervals. Chondrosia reniformis displayed the lowest correlation (0.816 [0.941–0.998]), and the degree of concordance was significantly lower than in P. ficiformis and C. crambe.
A suit of core microbial genera and metabolite classes leading the multivariate correlation were selected by sCCA (Fig. 9). Petrosia ficiformis revealed the largest set of informative microbial genera (12), of which only six were significantly correlated with three metabolite classes, being Sva0996 and TK17 correlated with ChE, NS3a and AEGEAN-169 with Co and SAR202 and Cryomorphaceae correlated with PMe. In C. crambe, out of the 11 genera selected by sCCA, only five revealed significant correlations with two metabolite classes. DG were correlated with Romboutsia, Rubritalea, Blastopirellula and OCS116; and Cer were correlated with Romboutsia, Cryomorphaceae and OCS116. All seven microbial genera selected by sCCA in C. nucula were significantly correlated with at least one metabolite class: Oligoflexaceae, TK17, Synechococcus spongiarum, UBA10353 and HOC36 were linked to DG; Oligoflexaceae, TK17, Synechococcus spongiarum and HOC36 were correlated with SiE; Nitrospira, Cytophagales and HOC36 were linked with ChE; and TK17 and Synechococcus spongiarum were correlated with MePC. Finally, in C. reniformis only two of the seven selected microbial genera by sCCA revealed significant correlations with two metabolite classes, NS4 and Gammaproteobacteria were linked to PI and Clade la with ZyE.
A Heatmap depicting Total pairwise Spearman correlation of class metabolites and microbiome genera across the full dataset including all four sponge species (126 microbial genera, 33 metabolite classes, n = 37 sponge samples). B-E Panels display heatmaps restricted to microbial genera and metabolite classes selected by the sparse CCA procedure in the single sponge species, with significant Spearman correlations assigned by *: B: P. ficiformis (n = 8), C: C. reniformis (n = 10), D: C. crambe (n = 9), E: C. nucula (n = 10).
Discussion
Transmission electron microscopy observations reinforced previous findings that P. ficiformis and C. nucula host abundant and morphologically diverse microbes contained in bacteriocytes54,55,56,57. Crambe crambe was characterized by the presence of low abundant and dispersed microbes in the mesohyl, as previously reported11,58,59,60, dominated by a single morphological type. In C. reniformis, our observations did not reveal the presence bacteriocytes. Previous studies instead, reported nurse cells charged with symbiotic bacteria during the development20. Molecular investigations highlighted that C. crambe hosts one or very few bacterial taxa, mostly (80–90%) belonging to phylum Proteobacteria23,24,59,60. Our results based on SILVA v.128 release 69, expand such taxonomic resolution, by identifying a dominant Betaproteobacteriales ASV in Family EC94 (~70–80%). Recently, predominant Gammaproteobacteria associating with several sponges including C. crambe, were proposed to take part of a new order named Ca. Tethybacterales, and in particular to the species Ca. Beroebacter blanensis, in C. crambe hosts from Spain61. Along this dominant symbiont, we found a moderate representation of cyanobacteria, and a high number of minor bacterial taxa. C. crambe specimens, despite hosting low microbial abundances, revealed values of ASVs richness comparable to those of HMA sponges, but with lower diversity. This outcome suggests that microbial densities and taxa richness are not always correlated62, and highlights the relevance of the rare taxa to the overall diversity, especially in microbiota dominated by one or few symbionts (but see below)63. In LMA microbiomes dominated by one or few ASVs (as in C. crambe) dominant microbes are transmitted from adults to embryos with high fidelity thanks to brooding reproduction, as these symbionts are essential for fitness25,64,65,66. Instead, background taxa are mainly acquired horizontally from the water column63,67. These may include symbiont microbes retained with high fidelity, as the eight core taxa shared with the seawater recorded in our C. crambe, as well as other rare ASVs, some potentially representing transient microbes or food, accounting for up to ~175, in these hosts. The functional role of rare taxa is poorly understood, but they might be involved in host acclimatization processes. In this sense, background and/or newly introduced taxa may become operationally relevant, providing new metabolic pathways to face changing conditions, or replace lost functionalities during symbiont shuffling68,69,70.
In general, LMA species have higher pumping rates than HMA sponges71. This entails filtering larger volumes of seawater and environmental microbes therein, enhancing the chances for transient microorganisms to enter and/or to establish72,73,74. However, there are cases of HMA sponges, like P. ficiformis, in which symbionts are largely acquired from the environment25,54,75, along a likely minor fluctuant rate being transmitted within bacteriocytes to the oocytes20,54. This pattern is hypothesized to confer plasticity to colonize different biogeographical habitats15,18,75. On this line, Björk and collaborators76 reported highly dynamic microbiomes in P. ficiformis, mostly dominated by temporally transient taxa. In our P. ficiformis specimens collected in the same day from several sampling sites, the prokaryotic assemblages displayed the highest intraspecific variability and the smallest core (microbiome) size. At the same time, this species harbored the richest and most diverse microbiome, dominated by phyla Chloroflexi and Gemmatimonadetes. Chloroflexi has been consistently found associated with P. ficiformis from diverse geographical locations and morphotypes, suggesting a co-evolutionary relationship between the sponge host and this microbial taxon18. Instead, Proteobacteria77, and Gammaproteobacteria were predominant in other studies18, again indicating the fluctuant microbial communities even in the major phyla, in correlation with a diversity of growth forms (massive, reticular) described in these sponges. Within the genus Chloroflexi, Caldilineales, SAR202 clade and TK17 were the dominant orders in our sponges, being SAR202 clade commonly reported by other authors15,18,75. Older studies based on morphological approaches, described conspicuous presence of host specialized cells filled with vacuoles rich in cyanobacteria32,78. In our study, cyanobacteria (i.e. Synechococcales) displayed low relative abundances in P. ficiformis and were not a core taxon, which could indicate transitory partnerships. Low and variable presence of cyanobacteria have been similarly documented in recent surveys18,24,77, suggesting minor tropic functional roles of this bacterial group, and potentially related to divergent growth morphologies, colouration or habitat preference44,75,79 (see below).
Despite being widely investigated as a model organism, C. reniformis has received scarce attention regarding microbiome dynamics and composition76,80,81. Microbial assemblages in C. reniformis with pigmented morphotype investigated in the present study were mostly affiliated to the phyla Chloroflexi, Acidobacteria and Proteobacteria and exhibited rather low intraspecific variability. Such microbial steadiness could be linked to the uniformity in growth typologies under diversified habitat conditions (of light, depth, or other natural gradients). All the above may indicate that in C. reniformis there is a large proportion of bacteria transmitted in maternal bacteriocytes to the oocytes and related nursing cells20,78.
The most prevailing phylum in C. nucula was Cyanobacteria (over 25% relative abundance), followed by Chloroflexi and Proteobacteria. The association of C. nucula with cyanobacteria was noticed in previous morphological studies, proposing them as key beneficial symbionts82,83,84. And on the basis of histological analyses, it has been demonstrated that cyanobacteria, different heterotrophic bacteria and yeasts can be acquired by vertical transmission21,22. In our study C. nucula displayed the highest intraspecific similarity in microbiome composition, and the largest core microbiome in terms of both core ASV number and core size. A similar microbiome composition was reported using clone library sequencing on C. nucula collected in other locations of the Mediterranean85 and on the congeneric C. caribensis from the Caribbean (previously identified as C. nucula). On this regard, both species are restricted to illuminated habitats and have identical lobulated “chicken liver” brownish to purple morphotypes86, which may indicate a linkage between symbiosis and stable host morphology. All this information suggests that C. nucula harbors stable core bacterial assemblages across different biogeographic provinces, probably maintained via transgenerational transmission within oocytes.
The analysis of the microbiomes in the sponges investigated in the present study revealed a notable species-specific assemblage composition, which was also different from that of the surrounding seawater. Most of the prokaryotic taxa were exclusive of each sponge species and no shared ASVs were identified among the four species. Microbial assemblages chiefly modulated by host taxonomy have been previously reported in Porifera15,34,39,87, even when sharing similar environmental conditions17,18,46. There are though cases of microbial shifts associated with sponge hosts living in different habitats69, or in response to environmental factors19. Further studies, sampling diverse intraspecific phenotypes across diverse geographies and environmental scenarios should bring light on the phylosymbiosis, and level of within-species steadiness of microbial assemblages associated with our four target holobionts.
We identified a consortium of bacteria which were major drivers of the variability among sponge host microbiomes. In particular, taxa belonging to Subgroup 6 (Acidobacteria), SBR1031 (Chloroflexi), Bdellovibrio (Proteobacteria) were mainly associated to the HMA species (C. reniformis, P. ficiformis and C. nucula), whereas taxa belonging to Rhodothermaceae, Cyanobiaceae, PAUC34f and EC94 (Betaproteobacteriales) were mostly associated to C. crambe.
The bacterial taxon SBR1031 (Chloroflexi), mainly encountered in association with C. reniformis and P. ficiformis, belongs to the bacterial class Anaerolinae. It is involved in carbohydrate degradation and other degradation pathways of organic compounds (e.g. inositol88;), thus allowing the utilization of different carbon sources, and likely linked to the capability of living an heterotrophic lifestyle in the absence of light. Subgroup 6 belongs to the phylum Acidobacteria, which is widespread in marine environments89,90, and involved in the degradation of organic compounds and use of inorganic nutrients (i.e. sulfur and nitrogen), thus relevant for nutrient cycling89,90. The almost absence of this taxon in the C. crambe corroborates previous findings about the different sources of nitrogen compounds used by LMA sponges (mostly planktonic N) vs. HMA sponges (mostly nitrogenous waste)13. Within Proteobacteria, a dominant phylum in marine sponges7,26, genus Bdellovibrio was correlated to HMA species. Bdellovibrio, due to their capability to prey on other bacterial taxa, can exert an important role on microbe-microbe interactions and community structure modulation in complex microbiota assemblages91, thus potentially contributing to stabilize and explain differences among HMA microbiomes.
Among bacteria mostly associated with C. crambe, we found the EC94 group (Betaproteobacteriales), which has been previously reported as an important taxon in other LMA sponges from a variety of habitats of Antarctic ecosystems92, and in corals93. The maintenance of EC94 (Ca. Tethybacterales61) group over evolutionary time, across generations and geographic locations, may suggest that these symbionts could play a major nutritional role in the heterotrophic metabolism of LMA sponges, despite the ecological significance of this taxon is still unknown61,94.
Cumulative information on sponge microbiomes may allow soon to discriminate specific taxa driving co-divergence processes in Porifera, according to host taxonomies and/or habitat context.
Sponge microbiomes can be involved in a variety of processes including photosynthesis, methane oxidation, nitrification, nitrogen fixation, sulfate reduction, and dehalogenation26. Microbiomes of the sponges investigated showed putative functions mostly related to heterotrophic processes (e.g. chemoheterotrophy, aerobic heterotrophy) in P. ficiformis and C. reniformis, in accordance with the presence of different heterotrophic bacterial taxa (e.g. Chloroflexi), and the capability to be cave dweller sponges and live in dark conditions44,79,95. Conversely, the main putative functions of the microbiomes of C. crambe and C. nucula were related to autotrophic processes. These were mediated by the cyanobacterial component already documented32,85,86,96, and identified in the present study, in correlation with the preference for illuminated habitats.
HMA and LMA sponges can display different functional traits, probably due to intrinsic factors, such as divergent pumping rates50. In this sense, the LMA species C. crambe, revealed a wide array of exclusive metabolic functions, which were probably related to the rare taxa acquired from the surrounding environment. Processes as nitrification and denitrification can be instead found in both sponge typologies97. In our study, nitrification related functions (nitrification, aerobic nitrite oxidation, ammonia oxidation) were more relevant in microbiomes of P. ficiformis and C. reniformis than in those of C. nucula and C. crambe (in this latter negligible), independently from the HMA and LMA dichotomy. Nonetheless, relevant nitrification functional traits are normally due to a high presence of ammonia-oxidizing microbes in HMA hosts either heterotrophic or autotrophic9,97,98,99. Predatory or exoparasitic functions are related to predatory bacteria like Bdellovibrionales and Myxococcales, which were abundant in C. reniformis. These taxa feed on gram negative bacteria and can modulate microbial communities in sponges100, likely contributing to the microbiome stability of this species80. Fermentation functions were preeminent in C. reniformis, likely due to the presence of members of the phylum Poribacteria101. Overall, these findings suggest the presence of a complex network of metabolic functions of microbiomes, whose relative importance changes in relation with the sponge species considered, and that may have different implications for the holobionts’ fitness.
In the present study, we expand the current knowledge on compounds involved in the primary metabolism (i.e. lipids) of the sponge holobionts, being the majority of investigations focused on secondary metabolites of biotechnological interests102. Previous findings carried out on P. ficiformis revealed the predominance of phospholipids (phosphatidyl ethanolamines PE, phosphatidylcholin PC, phosphatidylglycerols PG, phosphatidylserine PS and phosphatidylinositols PI), along with other constituents related to bacterial component (9-hexadecenoic, 15-methyl-9-hexadecenoic and l-octadecenoic acid)103. A single survey on C. reniformis highlighted the presence of 57 fatty acids, some of which with an odd number of carbon atoms putatively of bacterial origin104, while for C. nucula, the only one published work revealed that 5,9,23-triacontatrienoic acid was the more relevant lipid compound105. In the present study, the four sponge species were characterized by the presence of a “core metabolome” (lipids shared among all samples) represented in order of relative importance by phosphatidylcholines (PC), triglycerides (TG), phosphatidylethanolamines (PE), ceramides (Cer) and diglycerides (DG). Phospholipids, such as phosphatidylcholine and phosphatidylethanolamine and others (e.g. phosphatidylserine, phosphatidylglycerin, phosphatidylinosite and diphosphatidylglycerin)106, are key components of cellular membranes and their diversity in sponges has been proposed to contribute to the remarkable plasticity and acclimatization capability of this phylum107. Besides the shared lipids, we observed marked differences in metabolomics profiles of the different sponge holobionts, likely due to a combination of specific compounds characterizing the host and its microbiome. Indeed, in our samples, certain lipids were at times shared among some of the investigated holobionts or were exclusive of a single species. Phosphatidylinositols (PI), for instance, were found in all sponges except in C. crambe. High concentrations of phosphatidylinositols and phosphatidylglycerols are indicators of the presence of symbiotic bacteria107. Thus, the lack of phopsphatidylinositols signal in C. crambe could be related to the low microbial abundances characterizing this species (LMA). Lysophosphatidylglycerols are also integrant lipids of bacterial membranes108, and were detected only in P. ficiformis and C. reniformis, suggesting the presence of particular large groups of bacteria shared between these two species (e.g. SAR202 clade). One characteristic compound type exclusive of C. nucula were the monogalactosyldiacylglycerols (MGDG), which are ubiquitous lipids in the thylakoids of cyanobacteria109. These findings agree with the intimate partnership established between C. nucula and these photosymbionts32,85,86,96, and are further supported by our microbiome results, revealing cyanobacteria as the dominant phylum in this species. Another exclusive class of sterols were the campesterols (CmE), only found in P. ficiformis specimens, and considered specific markers of genus Petrosia110,111. Finally, sphingomyelins, which are sphingolipids of animal cell membranes related to the myelin sheath of nerve cells112 and never found before in sponges107, were detected in all the sponges’ samples investigated with the exception of P. ficiformis.
Overall, findings reported in the present study indicate that some lipids, such as those constituents of cellular membranes (e.g. phosphatidylcholine and phosphatidylethanolamine) or involved in energy storage (e.g. triglycerides), are ubiquitously present in the investigated sponges independent of their taxonomy, and derived from the host cells and/or microbes providing consonant pathways. While a wide variety of other compounds appear to be species-dependent and related to specific microbial components.
Our correlations analysis based on multi-omic profiling via paired targeted sequencing of the 16 S rRNA gene and LC-MS metabolomics revealed high level of global congruence across all four species. Most microbes in the overall dataset were involved in demarcating metabolic signatures representing each of the host species. However, at the single species subsets inter-omic concordance was weak, with few microbial genera displaying relevant correlations with a reduced number of class metabolites. The vanishment in microbial–metabolic correlation signal within host species, may reflect a substantial functional redundancy, already reported in sponge microbiomes (113,114,115). On top of this, technical aspects in our approach could be masking microbiome-metabolome coupling cues. On the one hand, probable infra-generic intra strain diverseness might not be captured with taxonomical classification based on amplicon sequencing in place of whole genomes116; whereas on the other, the metabolite classes used might be too generic and hence poorly resolutive117. We are enthusiastic that both perspectives will improve as microbial, and particularly metabolomic annotations, will become more accessible in the reference databases.
Results recognized some degree of difference in microbiome-metabolome coupling across species. Chondrosia reniformis had significantly lower canonical correlation than P. ficiformis and C. crambe, and also than C. nucula, but against this last one the difference was not significant. This could indicate higher functional redundancy and metabolic stability in C. reniformis’ promoted by a diverse and constant microbiome. This is in agreement with previous work reporting microbial stability from specimens collected in several natural habitats and from laboratory experimentations80,81. In contrast, P. ficiformis seems to rely on more fluctuant microbial communities modulating the metabolome, where large inputs of environmental symbionts of potentially different functionalities may narrow functional redundancy48. There is scarce previous information about the other two species on this topic. But, in the case of C. crambe the functional rare biosphere, mostly acquired by horizontal transmission, may as in P. ficiformis uphold more shifting metabolic signatures, yielding increased correlation signal.
Small clusters of core microbes and metabolites with non-zero loading coefficients in sCCA and significant pairwise Spearman ranks, were able to explain intraspecific interactions within each host. All the selected microbial taxa exhibiting significant metabolic coupling in C. crambe were negatively correlated with either Cer or DG. Cer and DG are structural lipids of the bilayered membrane of eukaryotic cells controlling its permeability, and acting as signaling molecules in immune responses118,119. Correlation with microbial groups suggest mechanisms of symbiont recognition120. Previous studies in anemones detected positive intensities of Cer and DG with the presence of intracellular symbionts120.Attending to our data and previous work59, negative correlations found in C. crambe could be related to sparse microorganisms living in the mesohyl. In C. nucula negative associations with DG seem to co-occur also with extracellular symbionts. These include Oligoflexaceae, an extracellular partner taxon in corals121, and Synechoccocus spongiarium122, which had also negative association with MePC, phosphatidylcholine derivatives typical of eukaryote membranes, and reported in intracellular bacteria to avoid host immune responses123. Instead, HCO36, UBA10353 and TK17 microbes were positively related with DG, being TK17 also correlated with MePC. We cannot, though, ascribe these taxa as intracellular symbionts with the available information (see88). Interestingly, Synechococcus spongiarium (Cyanobacteria) and TK17 (Chloroflexi) revealed opposite correlations with the same class metabolites, including phytosterol-like SiE, which could indicate a functional interplay. Actually, fermentation coupling between Cyanobacteria and Chloroflexi as oxygenic and anoxygenic phototrophs respectively were demonstrated across hypersaline microbial mats124. Several cholesterol derivatives revealed microbial correlations, as such ChE with described antibacterial properties125 interacting with several taxa in C. nucula and P. ficiformis, and ZyE, which was associated to Clade Ia in C. reniformis. Finally, cytosolic membrane related PI compounds exerted positive interactions with NS4 and selected Gammaproteobacteria microbes in C. reniformis. We can conclude that most metabolite classes captured in the correlation approaches were cell membrane affiliated components, presumably involved in host-symbiont recognition mechanisms. It is highly probable that future advances in microbial and metabolic analytics will retrieve other functional interactions. This will allow us to understand how sponge microbiomes participate in the multifactorial metabolic pathways, leading to intraspecific phenotypic diversification and species acclimatization to challenging conditions.
Our study performed on samples of a single phenotype appertaining to four different target species, allowed us to encounter trends in the microbiomes, metabolomes and/or their inter-omics interactions that seem to correlate with the potentiality of range traits for each sponge. Crambe crambe, is a LMA species displaying a large core microbiome dominated by a single symbiont, of vertical brooding acquisition. The presence of a notable arsenal of secondary metabolites, and a moderate representation of cyanobacteria and other rare taxa of likely environmental origin may allow it to colonize habitats with wide-range of light exposure. Petrosia ficiformis has the most variable microbiome and displayed the smallest core microbiome. This variability in the microbiome (mainly acquired horizontally), together with the plasticity in the growth typologies is likely to contribute to the colonization of different habitats (e.g. marine caves, shaded habitats at different depths) and to the production of a diversified group of secondary metabolites.
Chondrosia reniformis is a HMA species with a stable microbiome. Its growth typology is massive, with a certain variability in the pigmentation, being independent from its associated microbial communities. Such pigmentation variability coupled with a large and diverse core microbiome, and the characteristic to reproduce asexually by budding, could be the key for this species to spread in different habitats (caves, coralligenous, shaded or dark habitats).
Chondrilla nucula belongs to the HMA sponge group and has a large and diversified core microbiome. The characteristic to vertically transmit most of its microbiome to the offspring might be crucial for this species to be ubiquitous in shallow environments. But may also delay colonization in new habitats with different conditions, to which parental symbionts might not be acclimatized. Additionally, its strict trophic relationship with cyanobacteria constraints this sponge to colonize mainly photophilous habitats, with rare cases of rachitic individuals, found in shaded conditions.
Correlations outcomes indicate an overall relatedness between microbial genera and metabolite classes in the full dataset comprising all four sponge species. But at the single species, this degree of coherence vanishes. This in part could be explained by the strong functional redundancies of sponge microbiomes sustaining homeostasis, in particular in such hosts, constantly exposed to multitude and dynamic microbial partnerships. But furthermore, the metabolite classes analyzed might not be illustrative of the intricate coupling between microbiomes and metabolomes, for being somewhat structural or inclusive, probably holding relatedness with many taxa concurrently. It is probable that more specific compound arrays may yield fitter correlations. Further, the use whole genomes instead of partial 16 S rRNA amplicons for accurate microbial taxonomic characterization may have resolved potential strain level variabilities.
Sponges co-existing within local community assemblages are likely to exhibit diverse microbiome networks and metabolic profiles that promote niche diversification, but also analogous phenotypic patterns of “symbiont evolutionary convergence”, yielding holobionts with different bio-ecological traits. Our investigations supply new documentation in the interconnection of multi-omic approaches applied to marine holobionts, and propose their suitability to study phenotypes diversification in the face of changing environments and acclimatization processes.
Methods
Sponge collection and processing
All the samples in the same species collected for our study were of the same phenothype, this is: pigmented and massive for P. ficiformis and C. reniformis, pigmented and lobular for C. nucula and encrusting for C. crambe, and all from illuminated habitats, with analogous conditions as specified in Fig. 1.
Sponge pieces (~10 cm3) of different target species individuals Petrosia ficiformis (n = 12), Crambe crambe (n = 14), Chondrilla nucula (n = 10), and Chondrosia reniformis (n = 15) were collected along with ambient seawater (n = 9) in spring 2018 by scuba diving at 2–5 m depth around the island of Ischia, (Southern Tyrrhenian, Mediterranean Sea, Italy), where all species are highly abundant126,127. Sponges were recovered from three nearby sites with similar conditions: Castello Aragonese (CCO; 40°43'55.9“N–13°57'52.9“E), Grotta Mago external (GF; 40°42'41.6“N–13°57'51.4“E) and Sant’Anna (SA; 40°43'36.5“N–13°57'43.4“E) (Fig. S1). The sampling sites were accessed with local authorization from the marine protected area: Area Marina Protetta Regno di Nettuno (Ischia).
Samples were subdivided and preserved as explained in the Supplementary Material and Methods (SM, section 1.0) for microscope, microbiome and metabolomic analyses.
Transmission electron microscope (TEM) observations
After sample preparation (see SM, section 1.1), ultra-thin sections obtained using an ultra-microtome (Leica, UCT Ultracut Microtome) were observed by TEM (Zeiss, LEO 912 AB).
Microbial profiling based on 16 S rRNA targeted gene sequencing
DNA from all four species (n = 51 total sponge samples) and from seawater filters (n = 9) obtained after filtration was extracted using the DNeasy PowerSoil Pro Kit (Qiagen), following manufacturer’s instructions. For prokaryotic diversity analysis, the V3-V4 hypervariable region of the 16 S rRNA gene (Escherichia coli position: 341-805) was amplified using the primer pairs Bakt_341F 5′-CCTACGGGNGGCWGCAG-3′ and Bakt_805R 5′-GACTACHVGGGTATCTAATCC-3′128. These primers are specific for bacterial communities and may reveal lower capture of Archaea communities128. Sequencing was performed on an Illumina MiSeq sequencer with a Reagent Kit v3 (600-cycle) targeting ~400–450 bp amplification products.
Demultiplexed paired end sequence reads were imported into quantitative insights into microbial ecology pipeline (QIIME2) v.2019.10129. Sequences were filtered and denoised with DADA2130 to obtain “Amplicon Sequence Variants” (ASVs) feature frequency table. Taxonomy was assigned using a pre-trained Naïve Bayes classifier (sklearn)131 against the 16 S SILVA (99% identity) reference database v.128132 (details in SM, sections 1.2–1.3).
Metabolomics profiling based on untargeted LC-MS
For each sponge specimen (P. ficiformis n = 8, C. reniformis n = 10, C. crambe n = 9, C. nucula n = 10), ca. 500 mg of freeze-dried material was crushed and sonicated with 4.5 mL methanol (MeOH) for 30 s. Then, 15 mL of methyl tert-butyl ether (MTBE) were added and left 1 h at room temperature under shaking. Upon addition of 3.75 mL of chromatography water, the sample was mixed and then centrifuged to induce phase separation133. The upper organic phase was recovered, dried and kept in glass vials at -20 °C until use. Dry extracts were resuspended and injected in a Thermo Scientific Dionex UltiMate 3000 chromatograph coupled to a Thermo Scientific Q Exactive focus quadrupole-Orbitrap mass spectrometer (UHPLC-q-Orbitrap, Thermo Fisher Scientific, MA, USA) operated in full scan-data dependent MS2 (Full MS-ddMS2) discovery acquisition mode. Further details about UHPLC-q-Orbitrap configuration are reported in section 1.4 of the SM.
Statistics and reproducibility
Sponge samples from single individuals were collected, along with ambient seawater, in the following number of replicates: Petrosia ficiformis (n = 12), Crambe crambe (n = 14), Chondrilla nucula (n = 10), and Chondrosia reniformis (n = 15), seawater (n = 9). Statistical tests (Kruskal-Wallis, Wilcoxon rank sum test, PERMANOVA, Welch’s test; see below) were used to detect significant differences on alpha and beta diversity and were run in the R environment. Diversity comparisons based on binary and relative abundance transformed data and on center log transformed (CLR) data for differential abundance for compositional data were calculated applying QIIME2 plugins and R packages (for details see SM section 1.4). LipidSearch v4.0 (Thermo Fisher Scientific) and several R packages were used for metabolomics data analysis. Similarity Percentages (SIMPER) on network patterns were statistically evaluated with PRIMER134 (section 1.5, SM).
Microbial community analysis
Diversity analyses were run with core-metrics-phylogenetics command on rarefied data for alpha-diversity (within sample diversity) estimates using Observed ASVs and Shannon Indexes135 and using Bray Curtis and Jaccard distance matrixes for beta-diversity136,137. Classical methods based on relative abundance (rarefaction, alpha and beta diversity, taxa barplots) can lead to misleading results, in which specific features individually carry no trustful information for the absolute abundance138.
In order to address this potential flaw of compositional data, our analysis was complemented with an Aitchison distance based approach for diversity analysis, that can handle high levels of sparsity and zero values. The resulting distance matrix was plotted in ordination robust compositional principal component analysis (RPCA) biplots139, following q2-DEICODE plugin (https://library.qiime2.org/plugins/deicode/19). Input data in DEICODE pipeline, works on non-rarefied tables in a non-supervised method (no co-variates included in the formula computing the model).
This tool has been previously used in order to analyze compositional datasets both in biomedical and ecological studies140,141. Data for this analysis were log transformed and centered to obtain center log-ratios (clr), a transformation widely applied for compositional data investigations142.
Log ratios of the most important features driving the differences in the ordination space were visualized and plotted with Qurro interface to detect sample groupings143 (https://github.com/biocore/qurro). Qurro calculates log-ratios between any chosen sets of numerator features and denominator features. This is achieved by computing, for each sample, the log-ratio between the total counts of the selected numerator features (numerator ASVs) and the selected denominator features (denominator ASVs)143. Certain consortia of features (microbial taxa) could be identified as major drivers (high ranked) of sample type grouping along a selected ordination axis of the RPCA biplot. The resulting selected numerator and denominator differential taxa were then used to generate/calculate new log-ratios of the most explanatory microbial balances separating sample types.
Kruskal-Wallis tests for overall tests and pairwise Wilcoxon rank sum tests for multilevel comparisons were applied for contrasting alpha-diversity indexes. Permutational multivariate analysis of variance (PERMANOVA) was used to detect significant differences on beta diversity based on resemblance matrices. Pairwise Adonis144,145 (https://github.com/pmartinezarbizu/pairwiseAdonis) was used to perform multilevel comparisons. Welch’s Tests were applied to test the statistical significance of log ratios based on groupings from the differential microbe consortia selected in Qurro. Core microbiome and core size were calculated in R (packages “Microbiome”, “Phylosmith” and “Phyloseq”) considering ASVs 100% present across groups146,147,148.
Similarity percentage analysis (SIMPER)149 was run on PRIMER 6 + 134 to calculate turnover diversity and visualize the similarity within and among sample groups. Exclusive and shared ASVs were counted using the CONTA.SE command in Microsoft Excel Software. Network analyses were computed and visualized on Gephi v0.9.2150. FAPROTAX v1.2.4 was used as a database for the predicted functions annotations53, keeping in mind though, its limitations due to the taxonomy resolution151. R Studio (v1.2.5033) was used to perform statistical analyses and to generate plots with specific packages (Qiime2R, Phyloseq, Vegan, Ggplot2 R packages)148,152,153.
Metabolomics data analysis
LipidSearch v4.0 (Thermo Fisher Scientific) was used to process the UHPLC-q-Orbitrap data in non-targeted mode, to identify metabolomic features grouped as molecule class profiles. Class metabolites were represented in Principal component analysis (PCA) ordination from CLR transformed data in R (microbiome, MicroViz packages146,154). PERMANOVA was used to detect significant differences between the sponge species. Pairwise Adonis was used to perform multilevel comparisons (vegan R package)144,145. Networks of single compounds and molecular classes across sponge species were visualized and computed by Gephi v0.9.2150. Similarity percentage analysis (SIMPER) (Clarke, 1993) was run on PRIMER 6+ software134. Metabolomics data were deposited on the EMBL-EBI MetaboLights database155.
Microbiome and metabolome correlation patterns
For our correlations, we filtered out those ASVs in the microbiome data with lower 10% prevalence across all samples156,157. Metabolomics features were clustered into 33 metabolite classes. To deal with issues of data compositionality, both, the microbiome and metabolomics sets were centered log ratio transformed (CLR)158, applying a previous pseudocount of min(relative abundance)/2 (microbiome R package)146. CLR transformation over performs with respect to other transformations in addressing spurious correlations, subcompositional incoherence and lack of isometry, while maintaining scale invariance and straightforward interpretation159,160. Microbial ASVs were agglomerated to the genus level157, resulting in 126 genera across all four sponge species (full data set), and 54, 52, 71 and 77 genera for correlations subsets corresponding to C. nucula (n = 10), C. reniformis (n = 10), C. crambe (n = 9) and P. ficiformis (n = 8) respectively.
Global concordance based on procrustes testing
Principal component analysis (PCA) is an unconstrained method that does not use distance matrix, with the additional benefit that the feature variables (microbes, metabolites) maintain their “loadings” for plotting in the ordination. PCA were performed directly on our CLR transformed microbial and metabolome variables using the rda function from the vegan package in R152. Paired microbiome and metabolomics PCAs, corresponding to the full data set (four species) and for each of the sponge species subsets, were compared based on the first two axes via symmetric Procrustes (protest function, vegan package152). Procrustes procedure measures the degree of concordance between two ordinations normalized to unit variance, by conducting a superimposition fit, in which coordinates are translated and rotated, maintaining the overall data structure. Significance is calculated according to the sum-of-squared differences (m2) against the permuted null (999 permutations). For ecology applications, procrustes has demonstrated to be more powerful than the Mantel test, while reducing type I error161.
Sparse canonical correlation (sCCA) analyses and pairwise spearman correlations
Pairwise Spearman correlations were conducted to obtain overall correlations patterns between metabolite classes and microbial genera in the overall data set, and further combined sparse canonical correlation analysis (sCCA) to identify closer clustered associations within each of the sponge species.
Pairwise Spearman correlations, with multiple hypothesis testing adjustment applying the Benjamini-Hochberg correction (false discovery rate, FDR, controlled q-values) method were calculated integrating psych::corr.test and the calculateCorForTwoMatrices function in R (packages psych and multiOmixViz162,163), setting as cutoff for significance pairwise correlations an FDR q-value of 0.05.
sCCA is a very useful tool to biologically integrate multi ‘omics datasets, and select explanatory sets of features associations (e.g. microbe–metabolite). sCCA performs informative variable selection by calculating linear combinations of maximum correlation among the query data matrices, and bounding variable individual weights to generate sparsity. Based on L1-penalized matrix decomposition method of cross-product matrix akin to a LASSO regression problem157,164,165, we calculated the overall multivariate associations given by the first canonical variate correlation coefficient. With this procedure, variables were selected based on their importance to the overall covariance between microbiome and metabolome datasets. CCA.permute (nperms = 50) was used for model fitting, with nboot = 1000 bootstrapped 95% confidence interval and nperm = 1000 permutation test setting, at the 0.05 significance level (PMA R package164). Function randomizeMatrix from R package picante166 preserved data structure across the different permutations. Outcomes of Spearman correlations and selected sCCA associations were combined in heatmaps for each species subset with pheatmap package in R167.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Statistical outputs, Supplementary Figures. and tables can be found within the Supplementary Information file. Numerical source data for Fig. 5 and Fig. 8 can be found respectively within Supplementary Data 1 Table S8 and Supplementary Data 2 Table S10. Numerical source data for boxplots were provided in the Supplementary Data 3 file. Sequencing data for this study can be found under the Bioproject ID PRJNA851757 and can be accessed here: https://www.ncbi.nlm.nih.gov/bioproject/851757. The metagenomic reads for the 16 S V3-V4 region are available at NCBI under the accession numbers SAMN29249576-SAMN29249593, SAMN29249594-SAMN29249614, SAMN29249648-SAMN29249669. Metabolomics data have been deposited to the EMBL-EBI MetaboLights database (https://doi.org/10.1093/nar/gkz1019, PMID:31691833) with the identifier MTBLS5125. The complete dataset can be accessed here: https://www.ebi.ac.uk/metabolights/MTBLS5125.
References
Bell, J. J. The functional roles of marine sponges. Estuar. Coast. Shelf Sci. 79, 341–353 (2008).
De Goeij, J. M. et al. Surviving in a marine desert: the sponge loop retains resources within coral reefs. Science 342, 108–110 (2013).
Dittami, S. et al. A community perspective on the concept of marine holobionts: state-of-the-art, challenges, and future directions. PeerJ 9, e10911 (2019).
Guerrero, R., Margulis, L. & Berlanga, M. Symbiogenesis: the holobiont as a unit of evolution. Int. Microbiol. 3, 133–143 (2013).
Cerrano, C., Giovine, M. & Steindler, L. Petrosia ficiformis (Poiret, 1789): an excellent model for holobiont and biotechnological studies. Curr. Opin. Biotechnol. 74, 61–65 (2022).
Hentschel, U., Usher, K. M. & Taylor, M. W. Marine sponges as microbial fermenters. FEMS Microbiol. Ecol. 55, 167–177 (2006).
Webster, N. S. & Taylor, M. W. Marine sponges and their microbial symbionts: love and other relationships. Environ. Microbiol. 14, 335–346 (2012).
Hentschel, U. et al. Microbial diversity of marine sponges. Prog. Mol. Subcell. Biol. 37, 59–88 (2003).
Hentschel, U., Piel, J., Degnan, S. M. & Taylor, M. W. Genomic insights into the marine sponge microbiome. Nat. Rev. Microbiol. 10, 641–654 (2012).
Erwin, P. M., Coma, R., López-Sendino, P., Serrano, E. & Ribes, M. Stable symbionts across the HMA-LMA dichotomy: low seasonal and interannual variation in sponge-associated bacteria from taxonomically diverse hosts. FEMS Microbiol. Ecol. 91, fiv115 (2015).
Gloeckner, V. et al. The HMA-LMA dichotomy revisited: an electron microscopical survey of 56 sponge species. Biol. Bull. 227, 78–88 (2014).
Ribes, M. et al. Functional convergence of microbes associated with temperate marine sponges. Environ. Microbiol. 5, 1224–1239 (2012).
Morganti, T., Coma, R., Yahel, G. & Ribes, M. Trophic niche separation that facilitates co-existence of high and low microbial abundance sponges is revealed by in situ study of carbon and nitrogen fluxes: trophic niche separation in Sponges. Limnol. Oceanogr. 62, 1963–1983 (2017).
Freeman, C. J., Easson, C. G. & Baker, D. M. Niche structure of marine sponges from temperate hard-bottom habitats within Gray’s Reef National Marine Sanctuary. J. Mar. Biol. Assoc. U. Kingd. 96, 559–565 (2016).
Díez-Vives, C. et al. On the way to specificity–microbiome reflects sponge genetic cluster primarily in highly structured populations. Mol. Ecol. 29, 4412–4427 (2020).
Hentschel, U. et al. Molecular evidence for a uniform microbial community in sponges from different oceans. Appl. Environ. Microbiol. 68, 4431–4440 (2002).
Luter, H. M. et al. Biogeographic variation in the microbiome of the ecologically important sponge, Carteriospongia foliascens. PeerJ 3, e1435 (2015).
Burgsdorf, I. et al. Biogeography rather than association with cyanobacteria structures symbiotic microbial communities in the marine sponge Petrosia ficiformis. Front. Microbiol. 5, 1–11 (2014).
Yang, Q., Zhang, W. & Franco, C. M. M. Response of sponge microbiomes to environmental variations. in Symbiotic Microbiomes of Coral Reefs Sponges and Corals, Vol. 1, 181–247 (Springer, 2019).
Díez-Vives, C., Koutsouveli, V., Conejero, M. & Riesgo, A. Global patterns in symbiont selection and transmission strategies in sponges. Front. Ecol. Evol. 10, 1015592 (2022).
Maldonado, M. et al. Endosymbiotic yeast maternally transmitted in a marine sponge. Biol. Bull. 209, 94–106 (2005).
Maldonado, M. Intergenerational transmission of symbiotic bacteria in oviparous and viviparous demosponges, with emphasis on intracytoplasmically-compartmented bacterial types. J. Mar. Biol. Assoc. U. Kingd. 87, 1701–1713 (2007).
Björk, J. R., Díez-Vives, C., Astudillo-García, C., Archie, E. A. & Montoya, J. M. Vertical transmission of sponge microbiota is inconsistent and unfaithful. Nat. Ecol. Evol. 3, 1172–1183 (2019).
Sipkema, D. et al. Similar sponge-associated bacteria can be acquired via both vertical and horizontal transmission. Environ. Microbiol. 17, 3807–3821 (2015).
Carrier, T. J. et al. Symbiont transmission in marine sponges: reproduction, development, and metamorphosis. BMC Biol. 20, 100 (2022).
Taylor, M. W., Radax, R., Steger, D. & Wagner, M. Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol. Mol. Biol. Rev. 71, 295–347 (2007).
Hudspith, M. et al. Subcellular view of host–microbiome nutrient exchange in sponges: insights into the ecological success of an early metazoan–microbe symbiosis. Microbiome 9, 44 (2021).
Regoli, F., Cerrano, C., Chierici, E., Bompadre, S. & Bavestrello, G. Susceptibility to oxidative stress of the Mediterranean demosponge Petrosia ficiformis: role of endosymbionts and solar irradiance. Mar. Biol. 137, 453–461 (2000).
Bavestrello, G. & Sarà, M. Morphological and genetic differences in ecologically distinct populations of Petrosia (Porifera, Demospongiae). Biol. J. Linn. Soc. 47, 49–60 (1992).
Becerro, M. A., Turon, X. & Uriz, M. J. Natural variation of toxicity in encrusting sponge Crambe crambe (Schmidt) in relation to size and environment. J. Chem. Ecol. 21, 1931–1946 (1995).
Becerro, M. A., Uriz, M. J. & Turon, X. Trends in space occupation by the encrusting sponge Crambe crambe: variation in shape as a function of size and environment. Mar. Biol. 121, 301–307 (1994).
Arillo, A., Bavestrello, G., Burlando, B. & Sarà, M. Metabolic integration between symbiotic cyanobacteria and sponges: a possible mechanism. Mar. Biol. 117, 159–162 (1993).
Wilkinson, C. R. & Vacelet, J. Transplantation of marine sponges to different conditions of light and current. J. Exp. Mar. Biol. Ecol. 37, 91–104 (1979).
Moitinho-Silva, L. et al. The sponge microbiome project. GigaScience 6, 1–7 (2017).
Thomas, T. et al. Diversity, structure and convergent evolution of the global sponge microbiome. Nat. Commun. 7, 11870 (2016).
Fan, L. et al. Functional equivalence and evolutionary convergence in complex communities of microbial sponge symbionts. Proc. Natl Acad. Sci. Usa. 109, E1878–E1887 (2012).
Langille, M. G. I. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Wang, N. et al. Variations in bacterial taxonomic profiles and potential functions in response to the gut transit of earthworms (Eisenia fetida) feeding on cow manure. Sci. Total Environ. 787, 147392 (2021).
Webster, N. S. & Thomas, T. The sponge hologenome. mBio 7, e00135–16 (2016).
Turque, A. S. et al. Environmental shaping of sponge associated archaeal communities. PLoS One 5, e15774 (2010).
Ribes, M., Dziallas, C., Coma, R. & Riemann, L. Microbial diversity and putative diazotrophy in high- and low- microbial-abundance mediterranean sponges. Appl. Environ. Microbiol. 81, 5683–5693 (2015).
Siegl, A. et al. Single-cell genomics reveals the lifestyle of Poribacteria, a candidate phylum symbiotically associated with marine sponges. ISME J. 5, 61–70 (2011).
Nemoy, P., Spanier, E. & Angel, D. L. Nitrification activity of the sponge Chondrosia reniformis under elevated concentrations of ammonium. Front. Mar. Sci. 7, 607979 (2021).
Burgsdorf, I. et al. Lineage-specific energy and carbon metabolism of sponge symbionts and contributions to the host carbon pool. ISME J. 16, 1163–1175 (2022).
Zhang, S. et al. Comparative metabolomic analysis reveals shared and unique chemical interactions in sponge holobionts. Microbiome 10, 22 (2022).
Mohanty, I. et al. Multi-omic profiling of melophlus sponges reveals diverse metabolomic and microbiome architectures that are non-overlapping with ecological neighbors. Mar. Drugs 18, 124 (2020).
Clark, C. M. et al. Relationship between bacterial phylotype and specialized metabolite production in the culturable microbiome of two freshwater sponges. ISME Commun. 2, 22 (2022).
Loureiro, C. et al. Comparative metagenomic analysis of biosynthetic diversity across sponge microbiomes highlights metabolic novelty, conservation, and diversification. mSystems 7, e0035722 (2022).
Botté, E. S. et al. Changes in the metabolic potential of the sponge microbiome under ocean acidification. Nat. Commun. 10, 1–10 (2019).
Lurgi, M., Thomas, T., Wemheuer, B., Webster, N. S. & Montoya, J. M. Modularity and predicted functions of the global sponge-microbiome network. Nat. Commun. 10, 992 (2019).
Lopez, J. V. After the taxonomic identification phase: addressing the functions of symbiotic communities within marine invertebrates. in Symbiotic Microbiomes of Coral Reefs Sponges and Corals, Vol. 1, 105–144 (Springer, 2019).
Turnbaugh, P. J. & Gordon, J. I. An invitation to the marriage of metagenomics and metabolomics. Cell 134, 708–713 (2008).
Louca, S., Parfrey, L. W. & Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science 353, 1272–1277 (2016).
Maldonado, M. & Riesgo, A. Gametogenesis, embryogenesis, and larval features of the oviparous sponge Petrosia ficiformis (Haplosclerida, Demospongiae). Mar. Biol. 156, 2181–2197 (2009).
Usher, K. M., Kuo, J., Fromont, J. & Sutton, D. C. Vertical transmission of cyanobacterial symbionts in the marine sponge Chondrilla australiensis (Demospongiae). Hydrobiologia 461, 15–23 (2001).
Hooper, J. N. A., Soest, R. W. M. V. & Willenz, P. Systema Porifera, Vol. 1, 1–1099 (Springer, 2002).
Boury-Esnault, N. Order Chondrosida Boury-Esnault & Lopès, 1985. Family chondrillidae gray, 1872. in Systema Porifera, Vol. 1, 291–297 (Springer, Boston, MA, 2002).
Moitinho-Silva, L. et al. Predicting the HMA-LMA status in marine sponges by machine learning. Front. Microbiol. 8, 752 (2017).
Croué, J. et al. A single betaproteobacterium dominates the microbial community of the crambescidine-containing sponge Crambe crambe. Sci. Rep. 3, 1–8 (2013).
Gantt, S. E., López-Legentil, S. & Erwin, P. M. Stable microbial communities in the sponge Crambe crambe from inside and outside a polluted Mediterranean harbor. FEMS Microbiol. Lett. 364, 1–7 (2017).
Taylor, J. A. et al. Phylogeny resolved, metabolism revealed: functional radiation within a widespread and divergent clade of sponge symbionts. ISME J. 15, 503–519 (2021).
Giles, E. C. et al. Bacterial community profiles in low microbial abundance sponges. FEMS Microbiol. Ecol. 83, 232–241 (2013).
Reveillaud, J. et al. Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J. 8, 1198–1209 (2014).
Carrier, T. J., Schmittmann, L., Jung, S., Pita, L. & Hentschel, U. Maternal provisioning of an obligate symbiont in a sponge. Ecol. Evol. 13, 1–10 (2023).
Song, H., Hewitt, O. H. & Degnan, S. M. Arginine biosynthesis by a bacterial symbiont enables nitric oxide production and facilitates larval settlement in the marine-sponge Host. Curr. Biol. 31, 433–437 (2021).
Knobloch, S., Jóhannsson, R. & Marteinsson, V. Genome analysis of sponge symbiont ‘Candidatus Halichondribacter symbioticus’ shows genomic adaptation to a host-dependent lifestyle. Environ. Microbiol. 22, 483–498 (2020).
Webster, N. S. & Blackall, L. L. What do we really know about sponge-microbial symbioses. ISME J. 3, 1–3 (2009).
Musat, N. et al. A single-cell view on the ecophysiology of anaerobic phototrophic bacteria. Proc. Natl Acad. Sci. USA 105, 17861–17866 (2008).
Weigel, B. L. & Erwin, P. M. Intraspecific variation in microbial symbiont communities of the sun sponge, Hymeniacidon heliophila, from intertidal and subtidal habitats. Appl. Environ. Microbiol. 82, 650–658 (2016).
Pester, M., Bittner, N., Deevong, P., Wagner, M. & Loy, A. A ‘rare biosphere’ microorganism contributes to sulfate reduction in a peatland. ISME J. 4, 1591–1602 (2010).
Morganti, T. M., Ribes, M., Yahel, G. & Coma, R. Size is the major determinant of pumping rates in marine sponges. Front. Physiol. 10, 1474 (2019).
Blanquer, A., Uriz, M. J. & Galand, P. E. Removing environmental sources of variation to gain insight on symbionts vs. transient microbes in high and low microbial abundance sponges: symbionts vs transient bacteria in HMA and LMA sponges. Environ. Microbiol 15, 3008–3019 (2013).
Schmitt, S., Angermeier, H., Schiller, R., Lindquist, N. & Hentschel, U. Molecular microbial diversity survey of sponge reproductive stages and mechanistic insights into vertical transmission of microbial symbionts. Appl. Environ. Microbiol. 74, 7694–7708 (2008).
Gerçe, B., Schwartz, T., Syldatk, C. & Hausmann, R. Differences between bacterial communities associated with the surface or tissue of Mediterranean sponge species. Microb. Ecol. 61, 769–782 (2011).
Britstein, M. et al. Sponge microbiome stability during environmental acquisition of highly specific photosymbionts. Environ. Microbiol. 22, 3593–3607 (2020).
Björk, J. R., O’Hara, R. B., Ribes, M., Coma, R. & Montoya, J. M. The dynamic core microbiome: structure, dynamics and stability. bioRxiv 137885 https://doi.org/10.1101/137885 (2017).
Schmitt, S., Hentschel, U. & Taylor, M. W. Deep sequencing reveals diversity and community structure of complex microbiota in five Mediterranean sponges. Hydrobiologia 687, 341–351 (2012).
Sarà, M., Bavestrello, G., Cattaneo-vietti, R. & Cerrano, C. Endosymbiosis in sponges: Relevance for epigenesis and evolution. Symbiosis 25, 57–70 (1998).
Burgsdorf, I. et al. Rethinking symbiotic metabolism: trophic strategies in the microbiomes of different sponge species. bioRxiv 458021, 1–37 https://doi.org/10.1101/2021.08.28.458021 (2021).
Ribes, M. et al. Restructuring of the sponge microbiome favors tolerance to ocean acidification. Environ. Microbiol. Rep. 8, 536–544 (2016).
Roveta, C. et al. The prokaryotic community of Chondrosia reniformis Nardo, 1847: from diversity to mercury detection. Zoology 158, 126091 (2023).
Liaci, L. & Sara, M. Associazione fra la cianoficea Aphanocapsa feldmanni e alcune demospongie marine. Bolletino di zoologia 31, 55–65 (1964).
Sarà, M. Associazioni di demospongie con zooxantelle e cianelle. Boll. di Zool. 31, 359–365 (1964).
Gaino, E., Pansini, M. & Pronzato, R. Aspetti dell´associazione tra Chondrilla nucula Schmidt (Demospongiae) e microorganismi simbionti (Batteri e Cianoficee) in condizioni naturali e sperimentali. Cah. de. Biologie Mar. 18, 303–310 (1977).
Thiel, V., Leininger, S., Schmaljohann, R., Brümmer, F. & Imhoff, J. F. Sponge-specific bacterial associations of the Mediterranean sponge Chondrilla nucula (Demospongiae, Tetractinomorpha). Microb. Ecol. 54, 101–111 (2007).
Hill, M., Hill, A., Lopez, N. & Harriott, O. Sponge-specific bacterial symbionts in the Caribbean sponge, Chondrilla nucula (Demospongiae, Chondrosida). Mar. Biol. 148, 1221–1230 (2006).
Yang, Q., Franco, C. M. M., Lin, H. W. & Zhang, W. Untapped sponge microbiomes: structure specificity at host order and family levels. FEMS Microbiol. Ecol. 95, fiz136 (2019).
Bayer, K., Jahn, M. T., Slaby, B. M., Moitinho-Silva, L. & Hentschel, U. Marine sponges as Chloroflexi hot spots: genomic insights and high-resolution visualization of an abundant and diverse symbiotic clade. mSystems 3, e00150–18 (2018).
Kielak, A. M., Barreto, C. C., Kowalchuk, G. A., van Veen, J. A. & Kuramae, E. E. The ecology of Acidobacteria: moving beyond genes and genomes. Front. Microbiol. 7, 744 (2016).
Kalam, S. et al. Recent understanding of soil Acidobacteria and their ecological Significance: a critical review. Front. Microbiol. 11, 580024 (2020).
Bonfiglio, G. et al. Insight into the possible use of the predator Bdellovibrio bacteriovorus as a probiotic. Nutrients 12, 2252 (2020).
Cárdenas, C. A., Font, A., Steinert, G., Rondon, R. & González-Aravena, M. Temporal stability of bacterial communities in antarctic sponges. Front. Microbiol. 10, 2699 (2019).
Gonzalez-Zapata, F. L. et al. Holobiont diversity in a reef-building coral over its entire depth range in the mesophotic zone. Front. Mar. Sci. 5, 1–13 (2018).
Gauthier, M. E. A., Watson, J. R. & Degnan, S. M. Draft genomes shed light on the dual bacterial symbiosis that dominates the microbiome of the coral reef sponge Amphimedon queenslandica. Front. Mar. Sci. 3, 1–18 (2016).
Ruiz, C., Villegas-Plazas, M., Thomas, O. P., Junca, H. & Pérez, T. Specialized microbiome of the cave-dwelling sponge Plakina kanaky (Porifera, Homoscleromorpha). FEMS Microbiol. Ecol. 96, fiaa043 (2020).
Usher, K., Toze, S., Fromont, J., Kuo, J. & Sutton, D. A new species of cyanobacterial symbiont from the marine sponge Chondrilla nucula. Symbiosis 36, 183–192 (2004).
Schläppy, M. L. et al. Evidence of nitrification and denitrification in high and low microbial abundance sponges. Mar. Biol. 157, 593–602 (2010).
Hoffmann, F. et al. Complex nitrogen cycling in the sponge Geodia barretti. Environ. Microbiol. 11, 2228–2243 (2009).
Diaz, M. C. & Ward, B. B. Sponge-mediated nitrification in tropical benthic communities. Mar. Ecol. Prog. Ser. 156, 97–107 (1997).
Johnke, J., Fraune, S., Bosch, T. C. G., Hentschel, U. & Schulenburg, H. Bdellovibrio and like organisms are predictors of microbiome diversity in distinct host groups. Microb. Ecol. 79, 252–257 (2020).
Podell, S. et al. Pangenomic comparison of globally distributed Poribacteria associated with sponge hosts and marine particles. ISME J. 13, 468–481 (2019).
Selvin, J. & Lipton, A. P. Biopotentials of secondary metabolites isolated from marine sponges. Hydrobiologia 513, 231–238 (2004).
Ayanoglu, E., Walkup, R. D., Sica, D. & Djerassi, C. Phospholipid studies of marine organisms: III. New phospholipid fatty acids from Petrosia ficiformis. Lipids 17, 617–625 (1982).
Nechev, J. et al. Chemical composition of the sponge Chondrosia reniformis from the Canary Islands. Hydrobiologia 489, 91–98 (2002).
Litchfield, C., Tyszkiewicz, J. & Dato, V. 5,9,23-triacontatrienoic acid, principal fatty acid of the marine sponge Chondrilla nucula. Lipids 15, 200–202 (1980).
Rod’kina, S. A. Fatty acids and other lipids of marine sponges. Russian J. Mar. Biol. 31, S49–S60 (2005).
Genin, E. et al. New trends in phospholipid class composition of marine sponges. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 150, 427–431 (2008).
Shinde, P. B., Park, T., Kim, D. K., Hong, J. & Jung, J. H. New glycerides from a two-sponge association of Jaspis sp. and Poecillastra sp. Pharm. Biol. 46, 814–818 (2008).
Sato, N. Is monoglucosyldiacylglycerol a precursor to monogalactosyldiacylglycerol in all cyanobacteria? Plant Cell Physiol. 56, 1890–1899 (2015).
Wahid Khalil, M., Djerassi, C. & Sica, D. Minor and trace sterols in marine invertebrates XVII. 1 (24R)-24,26-dimethylcholesta-5,26-dien-3β-01, a new sterol from the sponge Petrosia ficiformis. Steroids 35, 707–719 (1980).
Giner, J. L., Gunasekera, S. P. & Pomponi, S. A. Sterols of the marine sponge Petrosia weinbergi: implications for the absolute configurations of the antiviral orthoesterols and weinbersterols. Steroids 64, 820–824 (1999).
Kolesnick, R. N. Sphingomyelin and derivatives as cellular signals. Prog. Lipid Res. 30, 1–38 (1991).
Lesser, M. P., Sabrina Pankey, M., Slattery, M., Macartney, K. J. & Gochfeld, D. J. Microbiome diversity and metabolic capacity determines the trophic ecology of the holobiont in Caribbean sponges. ISME Commun. 2, 1–12 (2022).
Weigel, B. L. & Erwin, P. M. Effects of reciprocal transplantation on the microbiome and putative nitrogen cycling functions of the intertidal sponge, Hymeniacidon heliophila. Sci. Rep. 7, 1–12 (2017).
Kelly, J. B., Carlson, D. E., Low, J. S. & Thacker, R. W. Novel trends of genome evolution in highly complex tropical sponge microbiomes. Microbiome 10, 1–16 (2022).
Semenov, M. V. Metabarcoding and metagenomics in soil ecology research: achievements, challenges, and prospects. Biol. Bull. Rev. 11, 40–53 (2021).
Mitchell, J. M., Flight, R. M. & Moseley, H. N. B. Deriving lipid classification based on molecular formulas. Metabolites 10, 122 (2020).
Kumari, A. Ceramide structure and derivatives. in Sweet Biochemistry, Vol. 1, 59–61 (Elsevier, 2018).
Moonen, H. & Bas, H. Mono- and Diglycerides. in Emulsifiers in Food Technology, Vol. 1, 73–92 (2014).
Chan, W. Y., Rudd, D. & Oppen van, M. J. Spatial metabolomics for symbiotic marine invertebrates. Life Sci. Alliance 6, e202301900 (2023).
Aguirre, E. G., Fine, M. J. & Kenkel, C. D. Abundance of Oligoflexales bacteria is associated with algal symbiont density independent of thermal stress in Aiptasia anemones. bioRxiv 536969 https://doi.org/10.1101/2023.04.14.536969 (2023).
Burgsdorf, I., Handley, K. M., Bar-Shalom, R., Erwin, P. M. & Steindler, L. Life at home and on the roam: genomic adaptions reflect the dual lifestyle of an intracellular, facultative symbiont. mSystems 4, e00057–19 (2019).
Aragón-Aranda, B. et al. The phospholipid N-Methyltransferase and phosphatidylcholine synthase pathways and the ChoXWV choline uptake system involved in phosphatidylcholine synthesis are widely conserved in most, but not all brucella species. Front. Microbiol. 12, 614243 (2021).
Lee, J. Z. et al. Fermentation couples Chloroflexi and sulfate-reducing bacteria to Cyanobacteria in hypersaline microbial mats. Front. Microbiol. 5, 64803 (2014).
Jong, T. & Mudgil, P. Exploring antimicrobial properties of cholesterol esters: a systematic literature review. All Life 15, 684–691 (2022).
Teixidó, N. et al. Functional biodiversity loss along natural CO2 gradients. Nat. Commun. 9, 1–9 (2018).
Goodwin, C., Rodolfo-Metalpa, R., Picton, B. & Hall-Spencer, J. M. Effects of ocean acidification on sponge communities. Mar. Ecol. 35, 41–49 (2014).
Herlemann, D. P. et al. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 5, 1571–1579 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Bokulich, N. A. et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6, 90 (2018).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Cutignano, A. et al. Profiling of complex lipids in marine microalgae by UHPLC/tandem mass spectrometry. Algal Res. 17, 348–358 (2016).
Anderson, M. J., Gorley, R. N. & Clarke, K. R. PERMANOVA+ for PRIMER: Guide to Software and Statistical Methods, Vol. 1, 1–214 (Plymouth, 2008).
Lozupone, C. A. & Knight, R. Global patterns in bacterial diversity. Proc. Natl Acad. Sci. USA 104, 11436–11440 (2007).
Bloom, S. Similarity indices in community studies: potential pitfalls. Mar. Ecol. Prog. Ser. 5, 125–128 (1981).
Ross, H. H., Sokal, R. R., Sneath, P. H. A. & Freeman, W. H. Principles of numerical taxonomy. Syst. Zool. 13, 106–108 (1964).
Morton, J. T. et al. Balance trees reveal microbial niche differentiation. mSystems 2, e00162-16 (2017).
Martino, C. et al. A novel sparse compositional technique reveals microbial perturbations. mSystems 4, e00016-19 (2019).
Baker, J. L. et al. Deep metagenomics examines the oral microbiome during dental caries, revealing novel taxa and co-occurrences with host molecules. Genome Res. 31, 64–74 (2021).
Núñez-Pons, L. et al. Hawaiian coral holobionts reveal algal and prokaryotic host specificity, intraspecific variability in bleaching resistance, and common interspecific microbial consortia modulating thermal stress responses. Sci. Total Environ. 889, 164040 (2023).
Pawlowsky‐Glahn, V., Egozcue, J. J. & Tolosana‐Delgado, R. Modelling and Analysis of Compositional Data 1, 1–272 (Wiley, 2015).
Fedarko, M. W. et al. Visualizing ’omic feature rankings and log-ratios using Qurro. NAR Genom. Bioinform. 2, 1–7 (2020).
Martinez Arbizu, P. pairwiseAdonis: Pairwise Multilevel Comparison Using Adonis (R package version 0.0.1, 2017).
Martinez Arbizu, P. pairwiseAdonis: Pairwise Multilevel Comparison Using Adonis (R package version 0.4, 2020).
Leo, L. & Shetty, S. Microbiome R package. https://microbiome.github.io/tutorials/ (2017).
Smith, S. phylosmith: an R-package for reproducible and efficient microbiome analysis with phyloseq-objects. J. Open Source Software 4, 1–3 (2019).
McMurdie, P. J. & Holmes, S. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8, 1–11 (2013).
Clarke, K. R. Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 18, 117–143 (1993).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. In Third International AAAI Conference on Weblogs and Social Media, Vol. 3, 361–362 (2009).
Yang, Y. et al. Unraveling key functional bacteria across land-use types on the Tibetan plateau. Ecosyst. Health and Sust. 9, 1–9 (2023).
Oksanen, J. et al. vegan: Community Ecology Package https://cran.r-project.org/web/packages/vegan/vegan.pdf (2008).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis, Vol. 2, 1–260 (Hadley Wickham, 2016).
Barnett, D. J. M., Arts, I. C. W. & Penders, J. microViz: an R package for microbiome data visualization and statistics. J. Open Source Softw. 6, 3201 (2021).
Haug, K. et al. MetaboLights: a resource evolving in response to the needs of its scientific community. Nucleic Acids Res. 48, D440–D444 (2020).
Mallick, H. et al. Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences. Nat. Commun. 10, 1–11 (2019).
Nguyen, Q. P. et al. Associations between the gut microbiome and metabolome in early life. BMC Microbiol. 21, 1–19 (2021).
Aitchison, J. The statistical analysis of compositional data. J. R. Stat. Soc. Ser. B 44, 139–160 (1982).
Kurtz, Z. D. et al. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput. Biol. 11, e1004226 (2015).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: and this is not optional. Front. Microbiol. 8, 2224 (2017).
Peres-Neto, P. R. & Jackson, D. A. How well do multivariate data sets match? The advantages of a procrustean superimposition approach over the Mantel test. Oecologia 129, 169–178 (2001).
Revelle, W. Procedures for Psychological, Psychometric, and Personality Research [R package psych version 2.3.9]. https://CRAN.R-project.org/package=psych (2023).
multiOmicsViz. Bioconductor http://bioconductor.org/packages/multiOmicsViz/ (2024).
Witten, D. M. et al. A penalized matrix decomposition, with applications to sparse principal components and canonical correlation analysis. Biostatistics 10, 515–534 (2009).
Rodosthenous, T., Shahrezaei, V. & Evangelou, M. Integrating multi-OMICS data through sparse canonical correlation analysis for the prediction of complex traits: a comparison study. Bioinformatics 36, 4616–4625 (2020).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
pheatmap. Pretty Heatmaps. https://CRAN.R-project.org/package=pheatmap (2018).
Zhou, Z. F., Menna, M., Cai, Y. S. & Guo, Y. W. Polyacetylenes of marine origin: Chemistry and bioactivity. Chem. Rev. 115, 1543–1596 (2015).
Garrabou, J. & Zabala, M. Growth dynamics in four Mediterranean demosponges. Estuar., Coast. Shelf Sci. 52, 293–303 (2001).
Bavestrello, G., Burlando, B. & Sarà, M. The architecture of the canal systems of Petrosia ficiformis and Chondrosia reniformis studied by corrosion casts (Porifera, Demospongiae). Zoomorphology 108, 161–166 (1988).
Becerro, M. A. Multiple functions for secondary metabolites in encrusting marine invertebrates. J. Chem. Ecol. 23, 1527–1547 (1997).
Becerro, M. A., Uriz, M. J. & Turon, X. Chemically-mediated interactions in benthic organisms: the chemical ecology of Crambe crambe (Porifera, Poecilosclerida). Hydrobiologia 355, 77–89 (1997).
Sidri, M. Chondrilla nucula (Porifera, Demonspongiae): An example of successful plasticity. Ecological and morphological aspects. 1–225 https://elib.uni-stuttgart.de/handle/11682/1665 (2004).
Acknowledgements
Thanks are due to Mr. Bruno Iacono for his help during the sampling phase of this study. We also thank the technicians from RIMAR microscopy group of the Stazione Zoologica: Franco Iamunno, Rita Graziano and Giampiero Lanzotti. Thanks are due to Dr. Maria Cristina Gambi for sharing important knowledge on the sampling area. We deeply thank Dr. Maria Costantini for her tips and for helping during the sampling and the laboratory activity. Authors deeply thank Mr. Pasquale Vassallo for providing underwater picture 1E. V. Mazzella has been supported by a PhD fellowship co-funded by the Stazione Zoologica Anton Dohrn and Polytechnic University of Marche (PhD course in Life and Environmental Sciences, XXXIII cycle). V. Mazzella was also supported by Assemble Plus transnational acces (Project n° 8437: ‘MetMetMetAOA’: Metabolic Meta-analysis in Metaorganisms adapted to Ocean Acidification) and EMBRIC (European Marine Biological Research Infrastructure Cluster) transnational access – (Project n° 5380, ‘MicroMetOAc’: Microbiome shifts and Metabolic production under Ocean Acidification). Graphical abstract was created with BioRender.com. Map in Fig. S1 was created using the Free and Open Source QGIS. Project was partially funded under the National Recovery and Resilience Plan (NRRP), Mission 4 Component 2 Investment 1.4 - Call for tender No. 3138 of 16 December 2021, rectified by Decree n.3175 of 18 December 2021 of Italian Ministry of University and Research funded by the European Union—NextGenerationEU: Award Number: Project code CN_00000033, Concession Decree No. 1034 of 17 June 2022 adopted by the Italian Ministry of University and Research, CUP C63C22000520001, Project title “National Biodiversity Future Center – NBFC.
Author information
Authors and Affiliations
Contributions
Conceptualization: L.N.P., A.D., V.M.; Methodology: V.M., N.E., B.G.G., G.N., L.N.P.; Investigation: V.M., A.D., N.E., B.G.G., G.N., A.F., L.N.P.; Data curation and visualization: V.M., L.N.P., N.E.; Original draft preparation: V.M., L.N.P.; Review and editing: V.M., A.D., N.E., B.G.G., G.N., A.F., L.N.P.; Supervision: L.N.P., A.D., N.E., A.F.; Project administration: L.N.P., A.D.; Funding acquisition: L.N.P., A.D. All authors have read and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Fan Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Linn Hoffmann and Johannes Stortz. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mazzella, V., Dell’Anno, A., Etxebarría, N. et al. High microbiome and metabolome diversification in coexisting sponges with different bio-ecological traits. Commun Biol 7, 422 (2024). https://doi.org/10.1038/s42003-024-06109-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06109-5
- Springer Nature Limited