Abstract
Inflammatory monocytes (iMO) are recruited from the bone marrow to the brain during viral encephalitis. C-C motif chemokine receptor (CCR) 2 deficiency substantially reduces iMO recruitment for most, but not all encephalitic viruses. Here we show CCR7 acts synergistically with CCR2 to control this process. Following Herpes simplex virus type-1 (HSV-1), or La Crosse virus (LACV) infection, we find iMO proportions are reduced by approximately half in either Ccr2 or Ccr7 knockout mice compared to control mice. However, Ccr2/Ccr7 double knockouts eliminate iMO recruitment following infection with either virus, indicating these receptors together control iMO recruitment. We also find that LACV induces a more robust iMO recruitment than HSV-1. However, unlike iMOs in HSV-1 infection, LACV-recruited iMOs do not influence neurological disease development. LACV-induced iMOs have higher expression of proinflammatory and proapoptotic but reduced mitotic, phagocytic and phagolysosomal transcripts compared to HSV-1-induced iMOs. Thus, virus-specific activation of iMOs affects their recruitment, activation, and function.
Similar content being viewed by others
Introduction
Leukocyte infiltration into the brain is associated with La Crosse virus (LACV)-induced encephalitis in humans1. LACV is the leading cause of pediatric arboviral encephalitis in North America and causes dozens of clinical cases annually2, some of which can have severe outcomes3. Mice infected with LACV exhibit a similar age-dependent predisposition to encephalitic disease as humans, which lessens as the peripheral immune response develops while aging4,5. In weanling mice infected with a lethal dose of LACV, leukocytes were also found associated with disease6,7. Flow cytometry analysis identified inflammatory monocytes (iMOs) as the largest subset of brain-infiltrating leukocytes suggesting these cells may play a role in disease7 and may represent a therapeutic target.
Hematopoietic progenitors in the bone marrow give rise to iMOs released into the blood in response to virus infection8. iMOs travel to sites of infection, infiltrate infected tissues and influence pathogenesis either directly9 or by maturing into effector cells that contribute to the immune response10. Appropriate recruitment of iMOs is critical during viral encephalitis to effectively clear virus from the brain, while not damaging neurons. Viral pathogenesis studies have shown that iMOs can have ameliorating11,12,13,14 or exacerbating effects on disease15,16, depending on the pathogen. Despite their potential as a therapeutic target to treat disease, our understanding of the mechanisms involved in the recruitment of iMOs from the bone marrow and their activation state once they reach the brain, remains incomplete.
One of the main initiators of monocyte recruitment is the signaling of the chemokine C-C motif ligand (CCL) 2 through the G protein-coupled chemokine receptor (GPCR) C-C motif receptor (CCR) 217,18. For encephalitic viruses, CCL2 has been demonstrated to be produced in the periphery early in infection19 and by neurons20 or glial cells7,21 later in infection. This signal must travel to the bone marrow where it is thought to activate CCR2-dependent desensitization of a CXCL12-CXCR4 stay signal in maturing monocytes22 to facilitate their release into the blood. This role for CCR2 has been validated in multiple studies where Ccr2−/− mice have been infected with encephalitic viruses such as herpes simplex virus-1 (HSV-1)11,12, West Nile virus14 and Tahyna virus (TAHV)7, and iMO recruitment to the blood and/or brain is impaired. Yet, Ccr2−/− mice infected with the encephalitic LACV or Jamestown Canyon virus, iMO recruitment was not impaired7, suggesting an alternative pathway for iMO recruitment out of the bone marrow.
It is possible that multiple signaling pathways control virus-induced iMO recruitment, which could explain the variable dependence for CCR2 in this process. Cellular immune responses commonly integrate multiple signaling pathways to control both the intensity and timing of the response23. Additionally, different viruses may more strongly activate specific signaling molecules thus shifting the overall contribution of that molecule in the response24,25. Here we examined iMOs recruited to the brain during a CCR2-independent (LACV) versus a CCR2-dependent (HSV-1) infection for GPCR transcript expression. We found that Ccr7 transcripts and protein were more highly expressed on iMOs from LACV-infected mice. We then examined the role of both CCR2 and CCR7 in iMO recruitment in either single (Ccr2−/− or Ccr7−/−), or double knockout (Ccr2−/− × Ccr7−/−) mice during either LACV or HSV-1 infection and identified that both receptors contribute to iMO egress from the bone marrow and entry to the brain. Furthermore, we determined that LACV-recruited iMOs do not influence disease in contrast to HSV-1-recruited iMOs11,12. To address this difference, we transcriptomically analyzed LACV- and HSV-1-recruited iMOs and found differences in proinflammatory, phagocytic and phagolysomal pathways that could influence iMO function.
Results
CCR7 expression is increased on iMOs during LACV infection, but is not necessary for iMO recruitment to the brain
In previous studies, we found that LACV induced iMO recruitment from the bone marrow to the brain in the absence of CCR27, suggesting other receptors or signaling pathways may compensate for CCR2 in this process. To identify other receptors that might compensate for CCR2 in iMO recruitment, we compared transcriptional expression of an array of GPCRs, some of which are known to be involved in immune cell trafficking26,27,28,29, in iMOs recruited to the brain during HSV-1 (CCR2-dependent iMO recruitment) and LACV (CCR2-independent iMO recruitment) infections. iMOs were isolated from the brains of three clinical HSV-1 and LACV infected mice by FACS (Fig. 1a–d), processed for RNA and then analyzed for GPCR transcriptional expression via RT2 Profiler PCR Array (Fig. 1e). Multiple transcripts were significantly differentially expressed between iMOs isolated from LACV- versus HSV-1- infected mouse brains (Supplementary Data 1). The transcript with the largest fold change was Ccr7 (Fig. 1e, Supplementary Data 1), which has previously been linked to iMO migration to sites of parasitic infection30. CCR7 protein expression was also increased on iMOs from the brain of LACV infected mice relative to HSV-1 infected mice (Fig. 1f, g). However, in a previous analysis looking at different chemokine receptor knockouts, we had found that CCR7 deficiency alone did not inhibit iMO recruitment to the blood following a lethal dose of LACV (103 PFU)7. Similarly, when iMO recruitment to the brain was analyzed at the same clinical time point, no difference was observed in the percentage of iMOs in the brain between WT or Ccr7−/− mice during LACV infection (Fig. 1h). Thus, despite the increased expression of CCR7 by LACV-induced monocytes, CCR7 alone does not control iMO recruitment from the bone marrow to the brain during LACV infection.
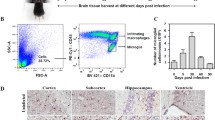
Representative FACs plots of iMO isolated (populations associated with the black arrows) from brains of (a, b) HSV-1 and (c, d) LACV IP infected mice. Gray = microglia, blue = granulocytes/neutrophils and red = iMOs. e Volcano plot of a RT2 Profiler PCR Array of 370 GPCR transcripts from FACs isolated iMOs. Data are plotted as the expression fold change of LACV vs HSV-1 iMO transcripts (x-axis) relative to p-value as determined by a Student’s t test between groups assuming equal variances (y-axis). Transcripts of highest significance and/or fold change are labeled and the complete data set is shown in Supplementary Table 1. n = 3 mice per group, all males. f Histogram of fluorescent intensity measurements of iMOs taken from the brain of LACV (black) and HSV-1 (gray)-infected, clinical mice. Data are normalized to peak values for ease of comparison due to the higher numbers of iMOs present in the brain of LACV relative to HSV-1 infected mice. g The mean fluorescent intensity (MFI) of iMOs from the brains of clinical LACV (black circles) and HSV-1 (gray squares) mice are plotted. A two-tailed, unpaired t-test was used to examine differences in MFI between iMOs from LACV and HSV-1 clinical brains with the following statistics: t = 4.035, df = 5, p = 0.0100. ** indicates a p value ≤ 0.01. h The percent of Ly6chi iMOs relative to microglia in the brain of IP 103 LACV-infected WT and Ccr7−/− mice at the 7dpi/clinical timepoint. This timepoint was selected to best correlate with the RT2 Profiler PCR Array (e) and with peak of iMO infiltration into the brain7. Data point dispersion is likely accounted for by only ~50% of animals in each group reaching clinical disease. A two-tailed, unpaired t-test was performed to examine differences in iMO recruitment between WT and Ccr7−/− mice with the following statistics: t = 0.2226, df = 7, p = 0.8302. Results from individual animals are plotted with the black horizontal bar representing the mean.
Generation of Ccr2−/− RFP x Ccr7−/− double knockout mice (DKO)
CCR2 and CCR7 may compensate for each other during LACV-induced iMO recruitment. To test this, we generated Ccr2−/− RFP × Ccr7−/− double knockout (hereafter DKO) mice through selective breeding. Genotyping with specific primers to amplify wildtype and knockout gene segments identified mice that were either fully wildtype (WT), heterozygous for both Ccr2 and Ccr7 (hereafter HET) or DKO (Supplementary Fig. 1a, b). Ccr2−/− RFP mice have been reported to be monocytopenic under normal conditions31. To examine whether DKO mice were also monocytopenic, we compared iMOs in the blood of mock infected HET (Supplementary Fig. 1c–f) and Ccr2−/− RFP (Supplementary Fig. 1g–j) mice to DKO (Supplementary Fig. 1k–n) mice. DKO mice had a clear reduction in the iMO population compared to either group indicating that DKO mice were more severely monocytopenic even than Ccr2−/− RFP mice. DKO mice did retain other important blood cell types such as CD11c+ dendritic cells (Supplementary Fig. 1c, g vs k), and Ly6G+ granulocyte/neutrophils (Supplementary Fig. 1e, i vs m). Importantly, DKO mice had similar numbers of iMO progenitors in their bone marrow compared to HET mice indicating that the decrease of iMOs in the blood of DKO mice was not the result of impaired progenitor survival (Supplementary Fig. 2). The relatively few iMOs in the blood of DKO mice retained their RFP expression (Supplementary Fig. 1f vs n), confirming they were CCR2 deficient and could be tracked.
Either CCR2 or CCR7 can mediate iMO recruitment to the blood during LACV infection but each receptor contributes additively to recruitment
To determine the individual contribution of CCR2 and CCR7 to iMO recruitment during LACV infection, Ccr2−/− RFP, Ccr7−/−, HET and DKO mice were infected with a lethal 103 PFU dose of LACV. iMOs were measured in the blood at different days post infection (dpi) (Fig. 2a–f). In HET mice, the proportion of iMOs in blood increased from a mock baseline during infection to a peak at 5dpi (Fig. 2a, e), followed by a return to near mock baseline at 7dpi (Fig. 2f). As previously reported, mock Ccr2−/− RFP mice were monocytopienic17,32 relative to mock HET and Ccr7−/− mice. However, during infection, the proportion of iMOs increased in the blood of both Ccr2−/− RFP and Ccr7−/− mice at a similar rate to HET mice, but with only approximately half the magnitude (Fig. 2e, f). In contrast, there was no iMO recruitment in DKO mice, with no proportional increase in iMOs and levels lower than all other strains throughout infection (Fig. 2e, f). Thus, iMO recruitment was largely abolished suggesting that iMO recruitment from the bone marrow to the blood is strongly dependent on signaling through either CCR2 or CCR7 during lethal LACV infection, and that these receptors play a synergistic role to generate robust iMO recruitment.
Representative flow cytometric plots of Ly6Chi, Ly6G- iMOs from the blood of mice with the highest (a and b HET) and lowest (c and d, DKO) iMO recruitment during 103 PFU LACV IP infection at 5dpi. The light gray labeled cells represent iMOs. e Time course analysis from mock to 5dpi of iMO recruitment to the blood of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice infected IP with 103 PFU LACV. f iMO recruitment to the blood of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice infected IP with 103 PFU LACV at the 7dpi/clinical time point. Data points with black boarders indicate nonclinical animals. All HET animals had clinical disease. g Time course analysis from mock to 5dpi of iMO recruitment to the blood of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice infected IP with 107 FFU HSV-1. For e–g, data are presented in box-whisker plots with individual data points represented by each symbol within the plot for each mouse genotype. The plots are color-coded such that magenta represents data from Ccr2−/− RFP, cyan from Ccr7−/−, black/gray from HET and orange from DKO mice. The following numbers of mice were analyzed for iMO recruitment to blood (e and f) during lethal LACV infection: Ccr2−/− RFP, n = 6 mock, n = 8 at 3dpi, n = 7 at 5dpi and n = 11 for 7dpi/clinical; Ccr7−/−, n = 5 mock, n = 6 at 3dpi, n = 5 at 5dpi and n = 8 at 7dpi/clinical; HET, n = 8 mock, and n = 6 at 3, 5 and 7dpi/clinical; DKO, n = 16 mock and at 3dpi, n = 14 at 5dpi and n = 15 at 7dpi/clinical. For g HSV-1 infection, the following numbers of mice were analyzed for each genotype: Ccr2−/− RFP mice, n = 5 mock, n = 8 at 3dpi and n = 6 at 5dpi/clinical. Ccr7−/− and HET mice, n = 5 at each time point. DKO mice, n = 5 mock, n = 9 at 3dpi and n = 6 at 5dpi/clinical. For e LACV-infected and g HSV-1 infected mice, a two-way ANOVA mixed-effects analysis with a Geisser-Greenhouse correction was used to compare the proportion of iMOs in the blood of (F(2.293,34.78) = 14.35, p < 0.0001) LACV and (F(1.237,21.03) = 10.61, p = 0.0023) HSV-1 mice from mock to 5dpi with a Tukey’s multiple-comparison test to compare four groups over multiple time points with an alpha of 0.05. Specific statistics can be found in Table 1. *, #, % or & symbols associated with a data point indicate the p-value for the statistically comparison between that data point and the time point-associated Ccr2−/− RFP, Ccr7−/−, HET or DKO data point respectively is below 0.05. Because not all mice developing clinical disease, a non-parametric Kruskal–Wallis test was used to compare the percentage of iMOs in the blood (f, Kruskal–Wallis statistic = 23.87, p < 0.0001) of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice at the 7dpi/clinical time point with a Dunn’s multiple comparisons test with an alpha of 0.05 to compare specific groups. P-values of comparisons to DKO mice are shown. All other statistics can be found in Table 1. Error bars represent SE for all groups. Dpi = days post infection.
CCR2 and CCR7 contribute additively to iMO recruitment during HSV-1 infection
To determine whether CCR2 and CCR7 play a synergistic role in iMO recruitment for other encephalitic viruses, the proportion of iMOs in the blood (Fig. 2g) of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice was measured throughout HSV-1 infection after inoculation with 107 FFU. There was an increase in iMOs in the blood of HET mice following HSV-1 infection, although considerably lower than that observed with LACV infection (Fig. 2e). The relative iMO percentages in each of the single knockouts were reduced compared to HET mice at all time points, with only a modest increase in iMO proportions in the blood at 3dpi that returned to near mock baseline by the 5dpi/clinical time point (Fig. 2g). Similar to LACV-infected mice, the proportions of iMOs in DKO infected mice was substantially lower than either single KO strain. Taken together, these data provide strong evidence that CCR2 and CCR7 play a synergistic role to mediate iMO recruitment during encephalitic virus infection, the latter of which has not been previously reported to be involved in iMO recruitment during viral infection.
DKO mice have impaired iMO recruitment to the brain during LACV and HSV-1 infection
iMOs have been shown to influence disease during encephalitis or brain inflammation by migrating to the brain and exerting an effector function9,11,12,14,32. Thus, iMO recruitment to the brain of HET, single KO and DKO mice was examined at multiple time points during LACV infection (Fig. 3a-f). In mock-inoculated control mice, the proportion of iMOs were similar in the brains of HET, Ccr2−/− RFP and DKO mice but were significantly elevated in Ccr7−/− mice by comparison (Fig. 3e). However, by 3dpi in LACV-infected mice, iMO infiltration had increased in Ccr2−/− RFP mice and were similar to Ccr7−/− mice. By 5 dpi, iMO proportions were increased in the brains of LACV-infected HET, Ccr2−/− RFP and Ccr7−/− mice, albeit with high variability. In contrast, iMO infiltration in brains of LACV-infected DKOs remained low. By the 7dpi timepoint (Fig. 3f), the majority of mice had developed clinical disease (solid symbols) with only a few non-clinical animals (open symbols) that had much lower iMO infiltration than clinical animals. At this time point, HET, Ccr2−/− RFP and Ccr7−/− mice had significantly higher proportions of iMOs in the brain relative to DKO mice (Fig. 3f). Thus, there was a lack of iMO recruitment to the blood and the brain in DKO mice, relative to single KO or HET mice during LACV infection.
Representative flow cytometric plots of Ly6Chi, Ly6G− iMOs from the brains of mice with the highest (a, b HET) and lowest (c, d, DKO) iMO recruitment during 103 PFU LACV IP infection at 7dpi. The dark gray labeled cells represent microglia and the light gray represent iMOs. e Time course analysis from mock to 5dpi of iMO recruitment to the brain of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice infected IP with 103 PFU LACV. f iMO recruitment to the blood of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice infected IP with 103 PFU LACV at the 7dpi/clinical time point. Data points with black boarders indicate nonclinical animals. All HET animals had clinical disease. g Time course analysis from mock to 5dpi of iMO recruitment to the blood of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice infected IP with 107 FFU HSV-1. For e LACV infection, the same Ccr2−/− RFP, Ccr7−/− and HET mice described in Fig. 2e, f were analyzed for iMO recruitment to the brain. Only a subset of the DKO mice used for blood analysis in Fig. 2f were used to analyze iMO recruitment to the (f) brain, totaling: n = 13 mock, n = 6 at 3dpi, n = 13 at 5dpi and n = 12 at 7dpi/clinical. For g HSV-1 infection, the same mice described in Fig. 2g were analyzed for iMO recruitment to the brain. For e–g, data are presented in box-whisker plots with individual data points represented by each symbol within the plot for each mouse genotype. The plots are color-coded such that magenta represents data from Ccr2−/− RFP, cyan from Ccr7−/−, black/gray from HET and orange from DKO mice. For e LACV-infected and g HSV-1 infected mice, a two-way ANOVA mixed-effects analysis with a Geisser-Greenhouse correction was used to compare the percentage of iMOs in the brain of (F(1.005,39.20) = 12.64, p < 0.0010) LACV and (F(1.496,36.64) = 5.215, p = 0.0167) HSV-1 mice from mock to 5dpi with a Tukey’s multiple-comparison test to compare four groups over multiple time points with an alpha of 0.05. Specific statistics can be found in Table 2. *, #, % or & symbols associated with a data point indicate the p-value for the statistically comparison between that data point and the time point-associated Ccr2−/− RFP, Ccr7−/−, HET or DKO data point respectively is below 0.05. Because not all mice developing clinical disease, a non-parametric Kruskal–Wallis test was used to compare the percentage of iMOs in the brain (f, Kruskal–Wallis statistic = 23.63, p < 0.0001) of Ccr2−/− RFP, Ccr7−/−, HET and DKO mice at the 7dpi/clinical time point with a Dunn’s multiple comparisons test with an alpha of 0.05 to compare specific groups. P-values of comparisons to DKO mice are shown. All other statistics can be found in Table 2. Error bars represent SE for all groups. Dpi = days post infection.
The lack of iMO recruitment in DKO mice occurred despite similar levels of CCR2 and CCR7 ligand transcript expression in HET and DKO lymph node (Supplementary Fig. 3a–d) at 3dpi when iMO recruitment to the blood is underway7 and in the brain at the 3 and 5dpi time (Supplementary Fig. 3e–h) when iMO brain recruitment is increasing in all groups (Fig. 3e). Furthermore, at least one Ccr2 or Ccr7 ligand transcript was elevated in most infected mice relative to mock controls in both tissues demonstrating these recruitment signals were elevated despite receptor knockout. Thus, the lack of iMO recruitment to the brain in DKO animals does not appear to be due to a lack of recruiting signal.
The recruitment of iMOs to the brains of HET, Ccr2−/− RFP, Ccr7−/− and DKO mice was also examined during HSV-1 infection (Fig. 3g). Again, iMOs were elevated in the brains of mock Ccr7−/− mice relative to all other groups (Fig. 3e), albeit without reaching statistical significance. During HSV-1 infection, the proportion of iMOs in the brain increased the most in HET mice at the 3 and 5dpi timepoints. This increase was not significantly different from single knockout mice but was significantly higher than DKO mice. The proportion of iMOs in the brain of infected HET mice was similar to Ccr7−/− mice at 3 and 5 dpi although this may be influenced by the higher basal proportion of iMOs observed in Ccr7−/− mice. iMO proportions in Ccr2−/− RFP mice were intermediate of HET and DKO mice similar to observations in the blood (Fig. 2e, f, g). Collectively this data indicates that dual deletion of CCR2 and CCR7 strongly inhibits iMO recruitment to the brain during infection with either LACV or HSV-1.
iMOs do not affect LACV pathogenesis
HSV-1 pathogenesis can be modulated by inhibiting iMO recruitment to the brain11,12. To determine if iMOs play a role in LACV-induced neuropathogenesis, HET and DKO weanling mice were infected with a lethal 103 PFU dose of LACV and followed for the development of neurological disease. Interestingly, HET and DKO mice developed neurological disease with similar timing and frequency, suggesting that iMO recruitment to the brain during LACV infection did not significantly influence viral pathogenesis (Fig. 4a).
a n = 3 HET and n = 3 DKO mice injected with mock inoculum and n = 13 HET and n = 15 DKO mice infected IP with high dose (103 PFU) LACV were followed for the development of clinical signs of neurological disease. A Log-rank Mantel–Cox curve comparison test was used to examine differences in survival between HET and DKO animals but did not reach significance (chi Square value = 0.1148, df = 1, p = 0.7347). b n = 30 Ccr2−/− RFP, n = 19 Ccr7−/−, n = 25 HET and n = 38 DKO mice infected IP with a low dose (102 PFU) of LACV were followed for the development of clinical signs of neurological disease. A Log-rank Mantel-Cox curve comparison test was used to examine differences in survival between Ccr2−/− RFP, Ccr7−/−, HET and DKO mice but did not reach significance (chi Square value = 4.221, df = 3, p = 0.2386).
Possibly, iMOs recruited with a lethal dose of LACV virus may not influence disease because the high virus dose is overwhelming, and a subtle effect of iMOs on the development of neurological disease would be missed. To address this, single knockout Ccr2−/− RFP and Ccr7−/− mice, along with HET and DKO mice were infected with a ~ LD50, 102 PFU dose of LACV. Again, all strains developed disease with similar timing and frequency with Ccr7−/− mice developing disease slightly less, and DKO slightly more often (Fig. 4b). No statistical difference was observed between these groups suggesting iMOs play a negligible role in LACV-induced neuropathogenesis.
LACV-recruited iMOs have higher proinflammatory, proapoptotic and less promitotic transcript expression than HSV-1-recruited iMOs
LACV induces a large influx of iMOs to the brain, relative to that seen with HSV-1 (Fig. 3f vs g). To examine if iMOs from LACV-infected mice were functionally different than the ones recruited during HSV-1 infection, iMOs from the brains of lethally infected LACV and HSV-1 mice were FACs isolated at the clinical time point and their transcriptomes compared via RNAseq analysis (Figs. 5 and 6). To validate the iMO isolation, expression of known iMO (Fig. 5a) and non-iMO (Fig. 5b) transcripts were compared across all samples. The known iMO transcripts were consistently highly expressed in both iMO populations (Fig. 5a), while non-iMO related gene transcripts were relatively low, indicating the iMOs from each virus infection were highly enriched and highly similar for iMO markers.
iMOs from (square) 3 HSV-1- and (circle) 3 LACV-infected brains were assayed for (a) iMO associated or (b) non-iMO associated transcripts. Data are plotted as fragments per kilobase of transcript per million reads (FPMK). c Volcano plot of the top 436 up-regulated and 334 down-regulated transcripts expressed by FACs-isolated iMOs in the brains of LACV and HSV-1 infected mice. Data are plotted as the expression fold change of LACV vs HSV-1 iMO transcripts (x-axis) relative to p-value as determined by a Student’s t test between groups assuming equal variances (y-axis). Heatmaps showing FPMK normalized expression of known (d) proinflammatory and antiviral, (e) proapoptotic and (f) mitotic prometaphase transcripts in iMOs from the brains of 3 LACV and 3 HSV-1 infected mice. Replicate mice from each infection group indicated (HSV-1 = green, LACV = magenta) had increased or decreased expression as shown by red or blue colored boxes respectively. Transcripts from the (d, e and f) heatmaps that were also in the top differentially expressed genes shown in c are highlighted in the volcano plot as either (magenta) up- or (green) down-regulated in LACV relative to HSV-1 iMOs.
Flow cytometrically identified iMOs from LACV and HSV-1 infected mouse (a, b) blood or (c, d) brain were assayed for their ability to phagocytose labeled bioparticles. Representative histogram plots of negative control (gray plots) and bioparticle-fed (black plots) samples from (a) blood and (c) brain demonstrate the positive bioparticle signal intensity in iMOs taken from LACV and HSV-1 infected brain. The percent of positive cells in (b) blood and (d) brain are plotted for each mouse with the black horizontal bar representing the mean. A two-tailed, unpaired t-tests was performed to examine differences in iMO phagocytosis collected from blood (c, t = 0.6497 df = 17, p = 0.5246) or brain (e, t = 0.3862, df = 9, p = 0.7083) of LACV and HSV-1 infected mice, that did not reach significance. Heatmaps showing FPMK normalized expression of known (e) phagosome-, (f and g) phagolysosomal-forming transcripts in iMOs from the brain of 3 LACV and 3 HSV-1 infected mice. Replicate mice from each infection group indicated (HSV-1 = green, LACV = magenta) and increased or decreased expression are shown by red or blue colored boxes, respectively.
Direct comparison of gene transcripts showed substantial differences between iMOs from LACV and HSV-1-infected mice (Fig. 5c–f). Transcript expression of cytokines, chemokines and interferon stimulated genes known to be upregulated during viral encephalitis were consistently higher in iMOs from LACV-infected mice compared to HSV-1-infected mice, while type I IFN was decreased (Fig. 5d). Concordantly, proapoptotic factor transcripts including Cycs, Apaf1, Fas, FasL and Casp7 were increased in LACV iMOs while the antiapoptotic Bcl2 was decreased (Fig. 5c, e) suggesting enhanced cell death. Surprisingly, other proapoptotic factors including Casp3, Casp6 and Bax were largely unchanged between groups indicating only select cell death pathways may be activated during viral encephalitis.
Correlative to the proapoptotic phenotype, transcripts for promitotic factors were largely decreased in LACV-recruited iMOs (Fig. 5c, f) which may indicate impaired immune cell development. Monocyte-derived macrophages have been shown to self-renew at sites of infection33 which may be critical for controlling infection in the brain. Interestingly, expression of Ifna2 and Ifnb1 transcripts was also decreased in LACV iMOs (Fig. 5c, d) which could be an indication of negative feedback inhibition34 as the type I interferon response has clearly been initiated as evidenced by high interferon-stimulated gene expression. Likewise, expression of the antimicrobial Mpo transcript was decreased in LACV monocytes which could indicate impaired phagolysosomal function35.
LACV-recruited iMOs are phagocytic, but have decreased transcription of phagolysosomal machinery
The decreased expression of Mpo transcript in LACV iMOs could suggest that phagocytosis or other downstream antiviral effector functions may be impaired in LACV-recruited iMOs in the brain. To address this, iMOs collected from the blood and brain of LACV infected mice were evaluated for their phagocytic ability compared to iMOs from HSV-1 infected mice (Fig. 6a–d). FACS isolated iMOs from blood (Fig. 3b, c) or brain (Fig. 3d, e) of clinical mice infected with either HSV-1 or LACV were incubated with fluorescent bioparticles to measure phagocytic uptake and fluorescent intensity was compared to cells incubated without bioparticles (Fig. 3b, d, gray line). The majority of iMOs isolated from both HSV-1 or LACV infected mice were capable of phagocytosing bioparticles indicating iMOs from LACV could be functional effectors. In support of this finding, Fcgr1 and Lamp2 transcripts, which are important for phagosome formation36, were increased in LACV iMOs (Fig. 6e). However, the ability to phagocytose does not confirm function. In fact, expression of multiple other transcripts involved in phagosome formation, function and fusion to the lysosome were decreased in LACV iMOs relative to HSV-1 (Fig. 6e–g). These included the cell surface receptors Fcgr2b, Clec7a, P2rx7, Manba, Lrp1 and CD36 which can initiate phagocytosis37,38, as well as multiple intracellular effectors of phagosome internalization and maturation including well known effectors such as Rab37, Atg5, Pikfyve and Eea139,40,41. Transcripts of critical effectors that mediate the fusion of the phagosome with the lysosome into the degradative phagolysosome were also largely decreased in LACV iMOs relative to HSV-1, including components of the HOPS (Fig. 6f) and SNARE (Fig. 6g) complex which facilitate fusion of the two organelles42. Thus, these data suggest that despite being functionally phagocytic, LACV-recruited iMOs have impaired transcription of critical factors involved in the formation of the phagosome and phagolysosome which may contribute for their inability to influence disease as has been shown for HSV-1-recruited iMOs.
Discussion
iMO monocytes have been shown to influence encephalitic disease11,12,14 and represent a potential therapeutic target16, but how they are recruited from the bone marrow to the brain is not fully understood. Here, we identified CCR7 as an important receptor for iMO recruitment from the bone marrow to the brain following virus infection. iMO recruitment was reduced, but was still robust, in both CCR2 and CCR7 single KO mice following LACV infection (Fig. 2). However, the absence of both receptors nearly eliminated iMO recruitment to the blood or brain (Figs. 2 and 3). Thus, iMO recruitment from the bone marrow to the brain during LACV infection can be mediated by either CCR2 or CCR7, with an additive effect of these receptors to induce substantial recruitment of iMOs to the brain.
CCR2 has been shown to be important for iMO recruitment during homeostatic and pathologic states for nearly two decades43. Many studies, including our own work with the encephalitic viruses HSV-1 and TAHV, have validated this finding7,11,12,14,22,32,43,44,45. However, the current study uncovered a previously unappreciated role for CCR7 in iMO recruitment to the blood and brain during encephalitic viral infection and provides a more complete view of how CCR2 and CCR7 have complementary roles in this process under homeostatic conditions. In uninfected Ccr2−/− RFP mice, iMOs account for ~1.0–2.5% of CD45+ cells in the blood while they account for ~2.8–4.8% of cells in Ccr7−/− mice (Fig. 2e). These amounts summate closely to the 3.9–6.9% of iMOs in blood found in HET controls (Fig. 2e)7 demonstrating the receptors are complementary even in the absence of an inflammatory signal. Furthermore, iMOs are found at lower levels in DKO mice compared to Ccr2−/− RFP or Ccr7−/− in mock or infected mice (Fig. 2e). Thus, our data demonstrate that CCR2 and CCR7 cooperate to contribute to the release of iMOs to the blood both during health and disease.
CCR7 expression by monocytes and monocyte-derived dendritic cells has been associated with their migration to the draining lymph node or target tissue in response to inflammation46,47,48 or infection30. These studies have focused on CCR7 function in monocyte-lineage cells following maturation or after arriving in target tissues. Our current findings suggest that CCR7 also functions upstream of trafficking to work in tandem with CCR2 to promote the initial recruitment of iMOs out of the bone marrow. Little is known about the expression of CCR7 by monocytes in the bone marrow, but transcriptional evidence suggests the receptor is lowly expressed under normal conditions49. However, expression on monocytes substantially increases as they reach the blood26 suggesting it could be involved in the iMO recruitment process. Furthermore, CCR7 expression cannot be induced following maturation of monocytes into macrophages50 indicating CCR7 expression is associated with the recruitment and migration but not subsequent maturation of iMOs into effector cells.
The involvement of both CCR2 and CCR7 in the recruitment of iMOs could be an advantage to the immune response in multiple ways. First, redundancy in the signaling response controlling iMO recruitment is beneficial as it limits the ability of pathogens to evolve evasion strategies to escape the immune response51. Additionally, the involvement of multiple receptors in this process allows for the response to be tunable, such that a stronger response can be elicited by a weak signal because an overall larger number of receptors could respond to the infection52. Conversely, the fact that only CCR7 is required to drive a robust iMO response during a lethal LACV infection7 indicates that the system can be saturated which would protect the host from an overwhelmingly harmful immune response. This is advantageous as the brain is a very delicate tissue and iMOs can be damaging in certain contexts15,16. Finally, it may be beneficial from an evolutionary perspective to have redundancy in immune response signaling that is not centered around a single genomic locus. A recent study demonstrated that deletion of the genomic locus containing the genes for Ccr1, Ccr2, Ccr3 and Ccr5 on chromosome 9 had a minimal impact on monocyte recruitment beyond the phenotype observed by Ccr2 knockout alone44 leading the authors to conclude CCR2 is the primary controller of recruitment. However, our data indicate that CCR7, which is located on chromosome 11, plays a complementary role in recruitment that could account for the CCR2-independent monocytes observed by Dyer et al.44 in Ccr2 and genome locus knockout tissues during resting and inflamed states.
We observed that iMOs play little-to-no role in LACV-induced pathogenesis regardless of infectious dose administered (Fig. 4). This was a surprising result considering these cells influence the pathogenesis of multiple other encephalitic viruses, including HSV-111,12,14,16. Furthermore, iMOs enter the brain of LACV-infected animals in exceptionally high numbers compared to other encephalitic viruses (Fig. 3f vs g)7 and are functionally phagocytic (Fig. 6a–d). The limited effect of LACV-recruited iMOs may be explained by their substantially different transcriptomics profile (Figs. 5 and 6), which had higher expression of proinflammatory and proapoptotic transcripts. This heavily proinflammatory/proapoptotic phenotype could be the result of an overwhelming lethal viral infection in neurons53,54,55 that is beyond the ability of infiltrating immune cells to control. LACV-recruited iMOs also had lower expression of transcripts associated with type I IFN production, mitosis, phagocytosis and phagolysosomal fusion, which are likely indicative of an immature, ineffective immune phenotype. Thus, infiltration of large numbers of iMOs into the LACV infected brain, particularly ones that may not be fully mature or possibly overstimulated, may simply be insufficient to combat an exceptionally aggressive encephalitic virus. It is possible, that the strong signals driving robust iMO recruitment during LACV infection could lead to this impaired state.
Methods
Ethics statement
Mouse experiments were approved by the Rocky Mountain Laboratories (RML) Animal Care and Use Committee and adhered to the National Institutes of Health guidelines and ethical policies. The RML facility is fully accredited by AAALAC International, and we have complied with all relevant ethical regulations for animal use.
Virus stocks and plaque assay
LACV (human, 1978)6 and McKrae HSV-156 stocks were made using Vero (CCL-81, ATCC) cells by infecting a confluent T75 flask (Corning) with a multiplicity of infection of 0.01 of either virus and harvesting the supernatant of each flask 5 days later. Supernatants were titered using plaque assay55 for LACV or focus forming assay for HSV-1 to determine viral concentrations. For both plaque and focus forming assays, Vero cells were plated 1 day in advance into 24-well plates (Corning) to confluency. Viral stock supernatants were diluted serially at 10-fold decreasing concentrations in DMEM (Gibco) supplemented with 2% FBS and penicillin/streptomycin and 200 μl of each dilution was applied the Vero cultures in duplicate. Plates were incubated 1 h to allow viral attachment and then each well was overlayed with 0.5 mL of 1.5% carboxymethyl cellulose in MEM (Gibco). Plates were incubated for 5 days and then fixed with 10% formaldehyde to a final concentration ≥4% formaldehyde per well for 1 h. HSV-1 wells were rinsed and refilled with 1xPBS to count endogenous viral-GFP foci for each sample dilution. LACV plates were rinsed with water stained with 0.35% crystal violet and rinsed with water again. LACV plates were air dried, and plaques were counted for each sample dilution. Sample titers were calculated as focus (HSV-1) or plaque (LACV)-forming units (FFU or PFU) per mL of supernatant.
Mice, husbandry and inoculations
Ccr2−/− RFP (B6.129(Cg)-Ccr2tm2.1lfc/J)32,57, Ccr7−/− (B6.129P2(C)-Ccr7tm1Rfor/J)58 and C57BL/6J (B6) mice were purchased from The Jackson Laboratory and maintained in RML breeding colonies. Ccr2−/− RFP Ccr7−/− double knockout (DKO) mice were generated in-house by crossing single knockouts and selective breeding of genotyped progeny. Confirmation of DKO mice is shown in the relevant figure and the primers used are reported in Supplementary Table 1. Heterozygous Ccr2+/− RFP Ccr7+/− (HET) controls were generated by crossing DKO mice with B6 mice. Mice were co-housed with same-sex littermates in ventilated plastic cages (Innovive IVC Caging Systems) with autoclaved bedding (Sani-Chips®) and enrichment (Shepherd Specialty Papers). Food and water were provided ad libitum. Animal holding rooms were kept at 22 ± 2 °C with 50 ± 10% relative humidity, and a 12-h light-dark cycle. All experimental groups were mixed-sex, except the RT2 Profiler PCR Array where only male mice were used to minimize sex-specific transcriptional hits per the manufacturer’s recommendations. Experimental mouse numbers are indicated in the associated figure legend.
Animals were reared in a specific pathogen free facility maintained by biannual sentinel sampling. Experimental animals were housed in an approved animal biosafety level 2 (ABSL-2) vivarium. Virus inoculations were performed intraperitoneally (IP) in weanling mice between the age of 20–22 days old. Virus stocks were diluted in phosphate-buffered saline (PBS) to the required concentration in a 200 μl volume for IP administration. Supernatant from uninfected Vero cultures equivalent diluted in 200 μl of PBS and administered via the same route served as a mock control. A lethal dose of LACV was 103 PFU/mouse and an ~50% lethal dose (LD50) was 102 PFU/mouse. HSV-1 was given at 107 FFU/mouse for all experiments.
For the RT2 Profiler PCR array, RNA-seq, flow cytometry, phagocytosis and qRT experiments, the investigator was responsible for mouse infection and harvesting of tissues and was thus unblinded for this process. However, after tissue collection, all experimental animal tissues were assigned a de-identified sequential tracking number which followed the specific tissue through the processing, experimentation, and analysis pipeline. Only after the final analysis was completed were the tracking numbers cross-reference with the experimental groups to unblind the investigator. For survival studies, the investigator was not blinded because they were responsible for randomizing groups by sex and treatment group and performed the injections. However, monitoring animals for development of disease is not subjective and any clinical signs of neurological disease including impaired ambulation, paralysis, ataxia, seizures, or repetitive tics resulted in a clinical score. Clinical disease progresses rapidly in mice infected with LACV and a mouse showing mild neurological signs will progress to severe disease within a matter of hours, thus a clinical score is consistently accurate to that day. Mice were twice monitored daily for presentation of clinical neurological signs.
Flow cytometry
At indicated timepoints post-infection, brain or whole bone marrow was collected into ice cold PBS, and blood was collected in a 1 mL syringe attached to a 27-gauge 1/2” needle with the needle barrel filled with 1000 U/mL heparin. Whole bone marrow was passed through a 70 μm filter to generate a single cell suspension and was then incubated with LIVE/DEAD fixable blue (ThermoFisher) at 1 μl of working solution per 106 cells for 30 min (min) to identify live cells prior to further processing. Bone marrow cells and immune cells that were isolated from brain and blood were then incubated with CD16/CD32 FcγIII/II (BD Biosciences, clone 2.4G2) to block Fc-receptors and immunolabeled with the primary conjugate antibodies indicated in Table 3, each at a 1:200 dilution for 30 min at 4 °C in the dark5,7. RFP fluorescence expression in the place of CCR2 on iMOs was observed in Ccr2−/− RFP, DKO and HET mice. Cells were then fixed with 2% paraformaldehyde for 30 min, washed 3× with PBS and analyzed using a BD LSRII or FACsSymphony A5 (BD Biosciences). The gating strategy for identifying iMO progenitors in bone marrow is shown in Supplementary Fig. 2 and the gating strategy for identifying iMOs in blood is shown in Supplementary Fig. 1. iMO identification in the brain has been published7 but utilizes the same gating strategy as in blood with the exception that microglia cells must be excluded as is shown in Fig. 1a, c. Antibodies listed in Table 3, but not shown in the gating strategy were used solely for confirmation of negative expression within iMOs and were not part of the core gating strategy. Single-stain and fluorescence minus one controls were performed for all experiments to establish gating boundaries. Data analysis was performed using FCS Expression Research Edition version 5 (Denovo software).
FACs
B6 mice were infected with either 103 LACV or 107 HSV-1 IP and brain immune cells were isolated as described above. Isolated cells were processed and immunolabeled as described above for identification of iMOs by FACs with either a FACsAria II (BD Biosciences) for PCR analysis59 or, for RNA-seq analysis, using a Miltenyi MACSQuant Tyto cell sorter (Miltenyi Biotec) per the manufacturer’s recommendations. Briefly, for RNA-seq analysis, a MACSQuant Tyto HS cartridge was primed with 0.4 mL of running buffer and brain immune cell samples were loaded into the cartridge. Prior to sorting, a small volume (~50 ul) of each sample were run by Flow Cytometry to define exclusion gates for debris and undesired cell types and to accurately identify iMOs. The antibodies used are shown in Table 3. The gating strategy used for FACs identification of iMOs was the same as for Flow Cytometry and is shown in Supplementary Fig. 1. Antibodies listed in Table 3, but not shown in the gating strategy were used solely for confirmation of negative expression and assessment of sort purity. FACs isolated iMOs were collected directly into 300 ul of RLT buffer for further processing as discussed below.
RT2 profiler PCR array analysis
iMOs were collected from the brain via FACs sorting as described above at the clinical time point. Whole-cell RNA was extracted from iMOs using Zymo RNA isolation kits per the manufacturer’s protocol. Extracted RNA was analyzed by a Nanodrop spectrophotometer (Thermofisher) to ensure sufficient concentration and purify to proceed. Using 200 ng of RNA from each sample, genomic DNA elimination, reverse transcription first strand cDNA synthesis and then real-time qRT (q-RT) analysis of the cDNA was performed per the manufacturer’s RT2 Profiler PCR Array for GPCRs (Qiagen, PAMM-3009ZE) protocol. qRT was performed on a Viia7 thermocycler (Life Technologies). CT values for 370 genes were analyzed using the RT2 Profiler PCR Array data analysis patch. All samples passed internal quality control standards for RT efficiency, specificity and genomic contamination.
Phagocytosis assay
Prior to tissue collection, Escherichia coli (K-12), Alexa Flour 488 conjugated bioparticles (Thermo Fisher, E13231) were reconstituted at 3.3 mg/mL in PBS and sonicated 2 × 15 s at 50 kHz with 65 W. Bioparticles were pre-opsonized by combining 1:1 with mock infected B6 mouse plasma at 37 °C for 30 min and then placed on ice until use. Immune cells from brain and blood were isolated and split between two wells of a 96 well plate (50 μl and 25 μl/well respectively) for each mouse. Each well’s volume was brought up to 140 μl with Hank’s buffered salt solution with 5% fetal bovine serum (FBS). 10 μl of bioparticles were added to one well of each mouse tissue and 10 μl of mouse serum mixed 1:1 with PBS was added to the other well to serve as a negative control. The plate was incubated for 30 min at 37°C and then 100 μl of chilled 0.1% trypan blue solution in PBS was added to each well for 1 min to quench any extracellular fluorescence. Cells were pelleted at 500 × g for 3 min, washed 2X in PBS with 2% FBS and prepared for flow cytometry as described above with the exception that the FITC channel was left open to measure the bioparticles.
RNA isolations and real-time qPCR analysis
RNA was extracted from ½ brains, and inguinal lymph nodes following a Trizol and chloroform protocol60. Briefly, brains and lymph nodes were placed in 2 mL tubes (Sarstedt) containing 1.5 and 1 ml of Trizol respectively and 3–5, 2.3 mm Zirconia/Silica beads (Fisher). Tubes were placed on a bead mill for 5200 rpm for 20 s × 2 with 5 s of dwell to homogenize tissues. Samples were then transferred to 1.5 ml microcentrifuge tubes and 200 ul of chloroform was added. Samples were shaken vigorously, then centrifuged at 12,000 × g for 15 min. The aqueous phase was transferred to a new tube and RNA was precipitated with 600 ul isopropanol for ~15 min–1 h. RNA was then pelleted at 12,000 × g for 10 min and the supernatant was removed. The pellet was washed in 1 ml 70% EtOH, vortexed, and centrifuged at 7600 × g for 5 min. The supernatant was removed and the pellets were allowed to air-dry for ~5 min. Brain and lymph node RNA was resuspended in 100 μl and 50 μl of nuclease-free water respectively and incubated at 55 °C for 10 min. Samples not used immediately were stored at −80 °C. 5 μl of brain and 50 μl of lymph node RNA was then DNase treated using the DNase I kit instructions (Ambion DNase). DNase-treated samples were then cleaned up with the Zymo RNA cleanup kit following the manufacturer’s instructions, with the exception that RNA wash steps were performed with 600 ul Wash Buffer and RNA was eluted in 50 μl warm NF H2O. cDNA was synthesized from the cleaned-up RNA using the BioRad iScript cDNA synthesis kit, following the kit instructions. cDNA samples were diluted five-fold in nuclease-free water and quantitative real-time PCR reactions were set up in triplicate in 384-well plates using SYBR Green SuperMix with ROX (Bio-Rad Laboratories) per the manufacturer’s recommendation with each primer at a 10 mM concentration. cDNA without reverse transcriptase and nuclease-free water with reverse transcriptase were used as negative controls. All samples were run on a Viia7 (Applied Biosystems) with a 95 °C dissociation for 3 min followed by 40 cycles of 95 °C/15 s to 60 °C/1 min amplification. A final 60–95 °C dissociation melt curve was generated to confirm amplification of a single product for each primer pair in each sample. The percentage difference in CT values with the housekeeping gene Gapdh were calculated (ΔCT = (CT Gapdh) − (CT gene of interest)) for each sample (“%gapdh”). Fold changes of virus and chemokines were calculated comparing the %gapdh values of mock controls with infected samples61. The viral and chemokine-specific primers used are shown in Supplementary Table 2.
RNA-seq processing and analysis
Brain immune cells from 3 HSV-1 and 3 LACV (both 2 females and 1 male) infected mice were prepared and labeled for FACs isolation as described above. Isolated iMOs from were lysed in RLT lysis buffer. The lysate was combined with additional RLT buffer and beta mercaptoethanol (MilliporeSigma, St. Louis, MO) to bring the final volume to 700 µL with 1% BME. Samples were passed through QIAshredder column (Qiagen, Valencia, CA) at 21,000 × g for 2 min to homogenize any genomic DNA. RNA was extracted using Qiagen AllPrep DNA/RNA mini columns (Valencia, CA). RNA integrity was assessed using the Agilent 2100 Bioanalyzer using RNA 6000 Pico kit (Agilent Technologies, Santa Clara, CA). RNA was quantitated using a fluorescence assay (Quant-it RiboGreen RNA, Thermofisher Scientific, Waltham, MA) on a Tecan Spark multiplate reader (Tecan, Switzerland). Starting with 200 pg of high-quality total RNA for each sample, volumes were concentrated via speed-vacuum to 9.5 ul. Sequencing libraries were constructed following the Takara SMART-Seq v4 PLUS protocol 011820 (Takara, Mountain View, CA) beginning with oligo(dT) priming/cDNA synthesis and following the manufacturers recommendations. Modifications during cDNA amplification included the use of SeqAmp CB PCR Buffer with 14 cycles of PCR. After AmPure XP bead purification (Agencourt Biosciences, Beverly, MA), cDNA quality was visualized and quantity assessed on a BioAnalyzer HS chip (Agilent Technologies, Inc., Santa Clara, CA). Purified cDNA for each sample was normalized to 1.5 ng in 8 ul volume going into library preparation with the Stem-Loop Adaptor mix. Following 15 cycles of library amplification using balanced SMARTer RNA unique-dual index primers from 96U SetA kit 042121 for Illumina sequencing (Takara, Mountain View, CA), individual libraries were cleaned using an 80% vol of AmPure XP beads, visualized on a BioAnalyzer HS chip, and quantified using the Kapa SYBR FAST Universal qPCR kit for Illumina sequencing (Kapa Biosystems, Wilmington, MA) on the CFX384 Real-Time PCR Detection System (Bio-Rad Laboratories, Hercules, CA). Each final library was diluted to a final concentration of 1.5 nM and pooled together in equimolar concentrations for sequencing. After an initial MiSeq paired-end 2 ×150 cycle sequencing run was completed to confirm proper index balancing, samples were run on a NextSeq2k instrument using 1000pM of manually-denatured final library pool and sequenced on a P2 flowcell following a paired-end 2 ×100 run with 300 cycle chemistry. Resulting Raw fastq files were trimmed to remove adapters and low-quality bases using Cutadapt v1.1862 before alignment to the GRCh39 reference genome and the Gencode v30 genome annotation using STAR v2.7.9a63. PCR duplicates were marked using the MarkDuplicates tool from the Picard v3.1.0 (https://broadinstitute.github.io/picard/) software suite. Raw gene counts were generated using RSEM v1.3.364 and differential expression was evaluated using DESeq265. Heatmaps for gene sets of interest were generated using Clustvis66, with FPKM normalized expression as input and rows mean centered.
Statistics and reproducibility
Sample size was determined by multiple factors. For the RT2 Profiler PCR array and RNA-seq analysis, the number of mice used in each group (3) was dictated respectively by the array and the sequencing chip to reach an adequate sequencing depth of coverage. For flow cytometry, phagocytosis, qRT and survival analyses, an initial power analysis was performed to estimate the number of animals required to detect a specified mean fold change and standard deviation, that were based on results from previous experiments, between groups assuming an alpha of 0.05 and 80% power. These estimates were 6 mice per group for cytometry, phagocytosis and qRT analyses and 14 mice for survival analyses. The final number of mice used in each experimental group was further dictated by the accumulated mean and variance within a group as additional replicates were added. If means and variance were consistent within a group after analyzing 5 or more mice, no additional mice were used. The breeding success of a specific strain, such as Ccr7−/− mice also limited the number of mice that were available for analysis. Specific animal numbers are reported in figures and figure legends. All measurements were taken from distinct experimental animals at an indicated time point.
Statistical tests were performed using Prism 9.3.1 software (GraphPad) or, Clustvis written in R (The R Project) for the RNA-seq analysis. As required for the experiment, one of the following statistical tests was used: a two-tailed unpaired t-test, a two-way ANOVA with a Sidak’s multiple-comparison or a two-way ANOVA with a Tukey multiple comparison. The specific test(s), degrees of freedom (df) and t-, or F-values and the p-value for each analysis are described in the corresponding figure legend or in the indicated tables. For RNA-seq heatmaps, rows and columns are clustered using Pearson correlation.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for Fig. 1e are provided in Supplementary Data 1. Numerical source data for all Figures can be found in Supplementary Data 2. RNA-seq analysis data from Figs. 5 and 6 have been upload to NCBI Geo and can be accessed with Accession # GSE254432. All other data that support the findings of this study are available from the corresponding author (CWW) upon reasonable request.
References
McJunkin, J. E. et al. La Crosse encephalitis in children. N. Engl. J. Med. 344, 801–807 (2001).
Centers for Disease Control and Prevention. La Crosse Encephalitis Virus (Centers for Disease Control and Prevention; 2022).
Boutzoukas A. E., et al. La Crosse Virus Neuroinvasive Disease in Children: A Contemporary Analysis of Clinical/Neurobehavioral Outcomes and Predictors of Disease Severity. Clin. Infect. Dis. 76, e1114–e1122 (2023).
Taylor, K. G., Woods, T. A., Winkler, C. W., Carmody, A. B. & Peterson, K. E. Age-dependent myeloid dendritic cell responses mediate resistance to la crosse virus-induced neurological disease. J. Virol. 88, 11070–11079 (2014).
Winkler, C. W. et al. Lymphocytes have a role in protection, but not in pathogenesis, during La Crosse Virus infection in mice. J. Neuroinflammation 14, 62 (2017).
Bennett, R. S. et al. La Crosse virus infectivity, pathogenesis, and immunogenicity in mice and monkeys. Virol. J. 5, 25 (2008).
Winkler, C. W. et al. Cutting Edge: CCR2 Is Not Required for Ly6C(hi) Monocyte Egress from the Bone Marrow but Is Necessary for Migration within the Brain in La Crosse Virus Encephalitis. J. Immunol. 200, 471–476 (2018).
Kratofil, R. M., Kubes, P. & Deniset, J. F. Monocyte Conversion During Inflammation and Injury. Arterioscler. Thromb. Vasc. Biol. 37, 35–42 (2017).
Terry, R. L. et al. Inflammatory monocytes and the pathogenesis of viral encephalitis. J. Neuroinflammation 9, 270 (2012).
Yang, J., Zhang, L., Yu, C., Yang, X. F. & Wang, H. Monocyte and macrophage differentiation: circulation inflammatory monocyte as biomarker for inflammatory diseases. Biomark. Res. 2, 1 (2014).
Boivin, N., Menasria, R., Gosselin, D., Rivest, S. & Boivin, G. Impact of deficiency in CCR2 and CX3CR1 receptors on monocytes trafficking in herpes simplex virus encephalitis. J. Gen. Virol. 93, 1294–1304 (2012).
Menasria, R., Canivet, C., Piret, J., Gosselin, J. & Boivin, G. Both Cerebral and Hematopoietic Deficiencies in CCR2 Result in Uncontrolled Herpes Simplex Virus Infection of the Central Nervous System in Mice. PLoS One 11, e0168034 (2016).
Michael, B. D. et al. Astrocyte- and Neuron-Derived CXCL1 Drives Neutrophil Transmigration and Blood-Brain Barrier Permeability in Viral Encephalitis. Cell Rep. 32, 108150 (2020).
Lim, J. K. et al. Chemokine receptor Ccr2 is critical for monocyte accumulation and survival in West Nile virus encephalitis. J. Immunol. 186, 471–478 (2011).
Kim, J. V., Kang, S. S., Dustin, M. L. & McGavern, D. B. Myelomonocytic cell recruitment causes fatal CNS vascular injury during acute viral meningitis. Nature 457, 191–195 (2009).
Getts, D. R. et al. Therapeutic inflammatory monocyte modulation using immune-modifying microparticles. Sci. Transl. Med. 6, 219ra217 (2014).
Tsou, C. L. et al. Critical roles for CCR2 and MCP-3 in monocyte mobilization from bone marrow and recruitment to inflammatory sites. J. Clin. Invest. 117, 902–909 (2007).
Conrady, C. D., Zheng, M., Mandal, N. A., van Rooijen, N. & Carr, D. J. IFN-alpha-driven CCL2 production recruits inflammatory monocytes to infection site in mice. Mucosal. Immunol. 6, 45–55 (2013).
Tumpey, T. M., Fenton, R., Molesworth-Kenyon, S., Oakes, J. E. & Lausch, R. N. Role for macrophage inflammatory protein 2 (MIP-2), MIP-1alpha, and interleukin-1alpha in the delayed-type hypersensitivity response to viral antigen. J. Virol. 76, 8050–8057 (2002).
Howe, C. L., LaFrance-Corey, R. G., Goddery, E. N., Johnson, R. K. & Mirchia, K. Neuronal CCL2 expression drives inflammatory monocyte infiltration into the brain during acute virus infection. J. Neuroinflammation 14, 238 (2017).
Bozzelli, P. L. et al. HIV-1 Tat promotes astrocytic release of CCL2 through MMP/PAR-1 signaling. Glia 67, 1719–1729 (2019).
Jung, H., Mithal, D. S., Park, J. E. & Miller, R. J. Localized CCR2 Activation in the Bone Marrow Niche Mobilizes Monocytes by Desensitizing CXCR4. PLoS One 10, e0128387 (2015).
Zhu, Y., Yao, S. & Chen, L. Cell surface signaling molecules in the control of immune responses: a tide model. Immunity 34, 466–478 (2011).
Zaiss, A. K. et al. Differential activation of innate immune responses by adenovirus and adeno-associated virus vectors. J. Virol. 76, 4580–4590 (2002).
Bouwman, W., Verhaegh, W., Holtzer, L. & van de Stolpe, A. Measurement of Cellular Immune Response to Viral Infection and Vaccination. Front. Immunol. 11, 575074 (2020).
Qu, C. et al. Role of CCR8 and other chemokine pathways in the migration of monocyte-derived dendritic cells to lymph nodes. J. Exp. Med. 200, 1231–1241 (2004).
Bai, Z. et al. CXC chemokine ligand 12 promotes CCR7-dependent naive T cell trafficking to lymph nodes and Peyer’s patches. J. Immunol. 182, 1287–1295 (2009).
Ricart, B. G., John, B., Lee, D., Hunter, C. A. & Hammer, D. A. Dendritic cells distinguish individual chemokine signals through CCR7 and CXCR4. J. Immunol. 186, 53–61 (2011).
Jakubzick, C., Tacke, F., Llodra, J., van Rooijen, N. & Randolph, G. J. Modulation of dendritic cell trafficking to and from the airways. J. Immunol. 176, 3578–3584 (2006).
Kling, J. C., Mack, M. & Korner, H. The absence of CCR7 results in dysregulated monocyte migration and immunosuppression facilitating chronic cutaneous leishmaniasis. PLoS One 8, e79098 (2013).
Jia, T. et al. Additive roles for MCP-1 and MCP-3 in CCR2-mediated recruitment of inflammatory monocytes during Listeria monocytogenes infection. J. Immunol. 180, 6846–6853 (2008).
Saederup, N. et al. Selective chemokine receptor usage by central nervous system myeloid cells in CCR2-red fluorescent protein knock-in mice. PLoS One 5, e13693 (2010).
Pang, J. & Koh, T. J. Proliferation of monocytes and macrophages in homeostasis, infection, injury, and disease. J. Leukoc. Biol. 114, 532–546 (2023).
Lukhele, S., Boukhaled, G. M. & Brooks, D. G. Type I interferon signaling, regulation and gene stimulation in chronic virus infection. Semin. Immunol. 43, 101277 (2019).
Arnhold, J. The Dual Role of Myeloperoxidase in Immune Response. Int J. Mol. Sci. 21, 8057 (2020).
Pauwels, A. M., Trost, M., Beyaert, R. & Hoffmann, E. Patterns, Receptors, and Signals: Regulation of Phagosome Maturation. Trends Immunol. 38, 407–422 (2017).
Jaumouille, V. & Grinstein, S. Molecular Mechanisms of Phagosome Formation. Microbiol. Spectr. 4, 1–18 (2016).
Gu, B. J. & Wiley, J. S. P2X7 as a scavenger receptor for innate phagocytosis in the brain. Br. J. Pharm. 175, 4195–4208 (2018).
Song, Y., Shang, D., Cheng, H. & Zhou, R. The small GTPase RAB37 functions as an organizer for autophagosome biogenesis. Autophagy 14, 727–729 (2018).
Uribe-Querol, E. & Rosales, C. Phagocytosis: Our Current Understanding of a Universal Biological Process. Front. Immunol. 11, 1066 (2020).
Kim, G. H., Dayam, R. M., Prashar, A., Terebiznik, M. & Botelho, R. J. PIKfyve inhibition interferes with phagosome and endosome maturation in macrophages. Traffic 15, 1143–1163 (2014).
Nguyen, J. A. & Yates, R. M. Better Together: Current Insights Into Phagosome-Lysosome Fusion. Front. Immunol. 12, 636078 (2021).
Serbina, N. V. & Pamer, E. G. Monocyte emigration from bone marrow during bacterial infection requires signals mediated by chemokine receptor CCR2. Nat. Immunol. 7, 311–317 (2006).
Dyer, D. P. et al. Chemokine Receptor Redundancy and Specificity Are Context Dependent. Immunity 50, 378–389 e375 (2019).
Fujimura, N. et al. CCR2 inhibition sequesters multiple subsets of leukocytes in the bone marrow. Sci. Rep. 5, 11664 (2015).
Liu, L., Wei, Y. & Wei, X. The Immune Function of Ly6Chi Inflammatory Monocytes During Infection and Inflammation. Curr. Mol. Med. 17, 4–12 (2017).
Zigmond, E. et al. Ly6C hi monocytes in the inflamed colon give rise to proinflammatory effector cells and migratory antigen-presenting cells. Immunity 37, 1076–1090 (2012).
Le Nouen, C. et al. Low CCR7-mediated migration of human monocyte derived dendritic cells in response to human respiratory syncytial virus and human metapneumovirus. PLoS Pathog. 7, e1002105 (2011).
Monaghan, K. L., Zheng, W., Hu, G. & Wan, E. C. K. Monocytes and Monocyte-Derived Antigen-Presenting Cells Have Distinct Gene Signatures in Experimental Model of Multiple Sclerosis. Front. Immunol. 10, 2779 (2019).
Allaire, M. A., Tanne, B., Cote, S. C. & Dumais, N. Prostaglandin E 2 Does Not Modulate CCR7 Expression and Functionality after Differentiation of Blood Monocytes into Macrophages. Int. J. Inflam. 2013, 918016 (2013).
Nish, S. & Medzhitov, R. Host defense pathways: role of redundancy and compensation in infectious disease phenotypes. Immunity 34, 629–636 (2011).
Lammermann, T. & Kastenmuller, W. Concepts of GPCR-controlled navigation in the immune system. Immunol. Rev. 289, 205–231 (2019).
Mukherjee, P., Woods, T. A., Moore, R. A. & Peterson, K. E. Activation of the innate signaling molecule MAVS by bunyavirus infection upregulates the adaptor protein SARM1, leading to neuronal death. Immunity 38, 705–716 (2013).
Winkler, C. W., Race, B., Phillips, K. & Peterson, K. E. Capillaries in the olfactory bulb but not the cortex are highly susceptible to virus-induced vascular leak and promote viral neuroinvasion. Acta Neuropathol. 130, 233–245 (2015).
Evans, A. B., Winkler, C. W. & Peterson, K. E. Differences in Neuropathogenesis of Encephalitic California Serogroup Viruses. Emerg. Infect. Dis. 25, 728–738 (2019).
Halford, W. P., Balliet, J. W. & Gebhardt, B. M. Re-evaluating natural resistance to herpes simplex virus type 1. J. Virol. 78, 10086–10095 (2004).
Boring, L. et al. Impaired monocyte migration and reduced type 1 (Th1) cytokine responses in C-C chemokine receptor 2 knockout mice. J. Clin. Invest. 100, 2552–2561 (1997).
Forster, R. et al. CCR7 coordinates the primary immune response by establishing functional microenvironments in secondary lymphoid organs. Cell 99, 23–33 (1999).
Winkler, C. W., Evans, A. B., Carmody, A. B. & Peterson, K. E. Placental Myeloid Cells Protect against Zika Virus Vertical Transmission in a Rag1-Deficient Mouse Model. J. Immunol. 205, 143–152 (2020).
Evans, A. B., Winkler, C. W. & Peterson, K. E. Differences in neuroinvasion and protective innate immune pathways between encephalitic California Serogroup orthobunyaviruses. PLoS Pathog. 18, e1010384 (2022).
Butchi, N. B., Woods, T., Du, M., Morgan, T. W. & Peterson, K. E. TLR7 and TLR9 trigger distinct neuroinflammatory responses in the CNS. Am. J. Pathol. 179, 783–794 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMDnetjournal. 17, 1–3 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Metsalu, T. & Vilo, J. ClustVis: a web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 43, W566–W570 (2015).
Acknowledgements
We thank Lara Myers, Carrie Long, Lydia Roberts and Christine Schneider-Lewis for critical reading of the manuscript, the entire Rocky Mountain Veterinary Branch staff and especially Jeff Severson, Shelby Heinz, Kimberly Dillard and Rebecca Charlesworth for outstanding animal husbandry with immune compromised animals. We also thank Kishore Kanakabandi, and Kent Barbian from the Research Technologies Branch at Rocky Mountain Laboratories for their assistance processing and running samples for RNA-seq. This work was funded by the Intramural Research Program at NIAID.
Funding
Open access funding provided by the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
C.W.W. developed study concept, designed, and conducted experiments, wrote the first draft of the manuscript and revised subsequent drafts. A.B.E. designed and conducted experiments. A.B.C. conducted experiments, curated, and analyzed data. J.B.L. analyzed data and generated figures. T.A.W. designed and conducted experiments. K.E.P. developed study concept, reviewed, and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Nadeem Khan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Si Ming Man and Johannes Stortz. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Winkler, C.W., Evans, A.B., Carmody, A.B. et al. C-C motif chemokine receptor 2 and 7 synergistically control inflammatory monocyte recruitment but the infecting virus dictates monocyte function in the brain. Commun Biol 7, 494 (2024). https://doi.org/10.1038/s42003-024-06178-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06178-6
- Springer Nature Limited