Abstract
Colistin is one of the last-resort antibiotics in treating infections caused by multidrug-resistant (MDR) pathogens. Unfortunately, the emergence of colistin-resistant gram-negative strains limit its clinical application. Here, we identify an FDA-approved drug, valnemulin (Val), exhibit a synergistic effect with colistin in eradicating both colistin-resistant and colistin-susceptible gram-negative pathogens both in vitro and in the mouse infection model. Furthermore, Val acts synergistically with colistin in eliminating intracellular bacteria in vitro. Functional studies and transcriptional analysis confirm that the combinational use of Val and colistin could cause membrane permeabilization, proton motive force dissipation, reduction in intracellular ATP level, and suppression in bacterial motility, which result in bacterial membrane disruption and finally cell death. Our findings reveal the potential of Val as a colistin adjuvant to combat MDR bacterial pathogens and treat recalcitrant infections.
Similar content being viewed by others
Introduction
As the underpinning of modern medicine, antibiotics have increased human life expectancy and are crucial for invasive surgery by preventing and treatment of postoperative infections. Unfortunately, cumulative antibiotic consumption has stimulated and accelerated the emergence and spread of antibiotic resistance1. To date, numerous mobile antimicrobial resistance (AMR) genes conferring resistance to last-resort antibiotics such as carbapenems, colistin, and tigecycline have been identified2,3,4. The remarkable propensity of bacteria to horizontally transfer genetic elements both within and across genera has led to the emergence of multidrug-resistant (MDR), extensively drug-resistant (XDR), and even pandrug-resistant (PDR) strains, which further limits the clinical options for treatment5.
Colistin, a cationic lipopeptide antibiotic also known as polymyxin E, is currently considered one of the last-resort antibiotics due to its high efficacy and low resistance rates against severe infections caused by MDR and even PDR gram-negative strains6. The positively charged residues of colistin bind to the negatively charged phosphate groups of lipid A on lipopolysaccharide anchored to the bacterial membrane, leading to membrane disruption and cellular content leakage, and finally cell death7. Regrettably, the clinical use of colistin is further limited due to its adverse effects, including nephrotoxicity and neurotoxicity8. In addition, gram-negative bacteria exhibiting intrinsic colistin resistance is mainly associated with chromosomal mutations in genes encoding two-component systems such as phoPQ, pmrAB and ccrAB, as well as the mgrB gene encoding lipid A phosphoethanolamine (pEtN) transferase, which catalyze the transfer of pEtN to the headgroups of lipid A, reducing the affinity of colistin for lipid A9,10. In recent years, a transferable gene, mcr, encoding pEtN transferase that mediates colistin resistance, has been reported worldwide11. Worse still, the mcr gene has been increasingly reported to be carried by MDR strains such as carbapenem-resistant Enterobacteriaceae bearing the blaKPC12, blaNDM13, and blaVIM genes14, leading to the emergence of XDR bacteria that are resistant to almost all known antibiotics.
Antibiotic adjuvants are compounds that have no or weak antibacterial effect but could restore or enhance the activity of antibiotics against bacteria exhibiting antibiotic resistance phenotypes15. The development of colistin adjuvant offers several therapeutic benefits, most notably the potential to reduce the required dosage of antibiotics in clinical applications. This reduction is pivotal in mitigating the emergence of antimicrobial resistance, a critical concern in contemporary medicine. Additionally, it addresses the issue of off-target toxicity, thereby enhancing the overall safety profile of antibiotic treatment regimens16,17. For instance, several natural phytochemicals have been identified to act synergistically with colistin. Catechol-type flavonoids and kaempferol were reported to potentiate colistin’s activity against gram-negative bacteria by disrupting iron homeostasis18,19. Besides, melatonin has been shown to potentiate the activity of colistin against MCR-producing, colistin-resistant pathogens by promoting oxidative damage and inhibiting efflux pumps20. Otilonium bromide, an Food and Drug Administration (FDA)-approved antispasmodic drug, has been reported to enhance the activity of colistin against gram-negative pathogens and their persisters by dissipating bacterial membrane proton motive force (PMF) and suppressing efflux pumps21. These findings revealed that the strategy of identifying colistin adjuvants from an FDA-approved library is safer and more cost-effective compared to developing new adjuvants. To date, no colistin adjuvant is commercially available, thus there remains an urgent need for the development of potential colistin adjuvants.
Valnemulin (Val), an FDA-approved pleuromutilin antibiotic, exhibited high activity against mycoplasmas and Pasteurella species, but lacked antibacterial activity against Enterobacteriaceae22,23. Val has been extensively used in the prevention and treatment of swine dysentery, colitis, and ileitis24. It has been shown that Val can bind to the 50S ribosomal subunit, resulting in the inhibition of protein synthesis25. Val has also been reported to act synergistically with doxycycline in combating Acinetobacter baumannii26. In addition, Val at concentrations ranging from 0 to 25 μg/ml shows no cytotoxicity to mammalian cells, indicating that its clinical use within this dose range could be safe27. In this study, we identified that Val acted synergistically with colistin in eliminating both colistin-resistant and colistin-susceptible gram-negative pathogens including Escherichia coli, Klebsiella pneumonia, and A. pittii both in vitro and in the mouse infection model. We also investigated the mechanisms underlying this synergistic bactericidal effect. Based on these findings, we hypothesize that Val could serve as a potential colistin enhancer in combating colistin-resistant gram-negative pathogens.
Materials and methods
Bacterial strains and reagents
All strains used in this study were listed in Supplementary Table S1. Strains were incubated in Luria–Bertani (LB) broth or on LB agar plates overnight at 37 °C. Val (Meilunbio, Dalian, China) was initially dissolved in double-distilled water and diluted in the culture medium to different working concentrations.
Antimicrobial susceptibility tests
The minimum inhibitory concentrations (MIC) of the Val and colistin were determined using the broth dilution method and the results were analyzed with reference to the Clinical & Laboratory Standards Institute guideline28. The synergistic antimicrobial effect of Val and colistin was further analyzed using a checkerboard assay29. Briefly, Val and colistin were serially diluted with 100 μL of cation-adjusted MH broth (CAMHB) to create an 8 × 6 matrix. Bacterial suspension at 106 CFU/ml was then inoculated into the broth containing different concentrations of Val and colistin and incubated at 37 °C for 16 h. The EnSight™ microplate reader (PerkinElmer, Waltham, MA, USA) was used to determine the absorbance of bacterial suspension at 600 nm (OD600). The fractional inhibitory concentration index (FICI) was calculated as follows to plot isobolograms: FICI = FICVal + FICcolistin = (MIC of colistin in combination with Val)/(MIC of colistin alone) + (MIC of Val in combination with colistin)/(MIC of Val alone). FICI ≤ 0.5 indicated synergy. All experiments were performed in biological triplicate. Isobolograms were generated following the descriptions previously with minor modifications30. FICVal and FICColistin was plotted as x and y coordinates, respectively. A straight line connecting the FIC values represents synergy.
Time-kill kinetic studies
The synergistic bactericidal effect of Val and colistin was evaluated by contracting the time-dependent killing curves following previous descriptions with some modifications31. Briefly, the overnight culture of colistin-resistant strains was 100-fold diluted in 3 ml LB broth and then incubated at 37 °C for 2 h with continuous shaking at 200 rpm. Bacterial suspension was treated with different concentrations of Val, colistin, and their combination for 24 h. Viable cells at each time point were counted in triplicate and the time-kill curves were constructed by plotting log10 CFU/ml against time using GraphPad Prism 9.0 (San Diego, CA, USA).
Resistance development assay
The development of colistin resistance was evaluated according to the methods reported previously32. Briefly, sequential culture of colistin-susceptible E. coli Bw25113, K. pneumoniae 80, and A. pittii P87 in the treatment of sub-inhibitory levels of colistin in the absence and the presence of 2 μg/ml of Val was performed to obtain resistant mutants. The serial passage was sustained over a period of 10 days.
Scanning electron microscopy (SEM) assay
SEM imaging was performed to visualize the cellular morphology of colistin-resistant E. coli in the treatment of Val, colistin, and the combination of both as described previously33. Briefly, mcr-1-bearing E. coli P47 at the late exponential phase was treated with Val, colistin, and the combination of both for 2 h. The bacterial cells were washed twice with sterile PBS, and then fixed overnight with 2.5% glutaraldehyde. The fixed cells were centrifuged (10,000 rpm, 1 min) and dehydrated with 100% ethanol. Cellular morphology was then observed using a Hitachi SU8010 SEM (Tokyo, Japan).
Cytotoxicity evaluation
Cytotoxicity study of Val and colistin on mammalian cells was performed using HEK293T cells34. Briefly, HEK293T was cultured at 37 °C in a 5% CO2 supplementing incubator until the density reached 6000–8000 cells/well in a 96-well microplate. After incubation, different concentrations of Val, colistin, and the combination of both were added to the medium for 24 h. The Culture medium without the addition of cells or drugs was used as the blank control. Cell viability was determined using a MTT Cell Proliferation and Cytotoxicity Assay Kit (Beyotime, Shanghai, China) following the manufacturer’s instructions, and the absorbance at 570 nm was measured using an EnSight™ microplate reader.
Intracellular bacterial determination
The synergistic bactericidal effect of Val and colistin on intracellular bacteria was determined using the cell infection model following previous descriptions with minor modifications31,35. Briefly, Vero cells were infected with E. coli P47 at an MOI of 100. Plates were centrifuged for 5 min at 1000 rpm and incubated at 37 °C with 5% CO2 for 1 h. Vero cells were co-cultured with Val, colistin, and the combination of both for 6 h at 37 °C in 5% CO2. Extracellular bacteria were eliminated with 2 μg/ml of meropenem, followed by washing twice with PBS. Subsequently, cells were lysed by DMEM supplemented with 0.1% BSA and 0.1% Triton X-100, and serial dilutions of the lysates were plated on MH agar plates for CFU counting.
Mouse infection model
The synergistic antibacterial effect of Val and colistin in vivo was evaluated in a mouse infection model29,36,37. Briefly, the BALB/c mice aged 5–8 weeks were divided into four groups (five mice per group). All mice were subjected to immunosuppression induced by cyclophosphamide treatment. Subsequently, all mice were infected with the colistin-resistant and carbapenem-resistant E. coli P80 at the dosage of 4.0 × 108 CFU/ml to establish a sepsis infection model for further investigation. At 1 h post-infection, the mice were treated with 2 mg/kg of Val, 4 mg/kg of colistin, and a combination of both every 12 h for 48 h, respectively. All animal experiments were approved by the Institutional Animal Care and Use Committee of Jiangsu University. We have complied with all relevant ethical regulations for animal use.
Membrane permeability test
The synergistic effect of Val and colistin on bacterial membrane permeability was determined using SYTOX Green (Thermo Fisher Scientific, Waltham, USA)38. Briefly, E. coli P47 at the exponential phase was treated with different concentrations of Val, colistin, and the combination of both for 2 h. After treatment, bacterial cells were collected by centrifugation and washed twice with sterile PBS, followed by re-suspending in PBS to adjust OD600 to 0.2. The bacterial suspension was stained with a final concentration of 1 μM SYTOX Green and incubated in the dark for 10 min. An EnSight™ microplate reader was used to monitor the fluorescence intensity of the test groups with an excitation and emission wavelength of 488 and 523 nm, respectively.
Membrane depolarization
Bacterial membrane potential was monitored using a fluorescent probe DiSC3(5) (Thermo Fisher Scientific, Waltham, USA)39. Overnight culture of E. coli P47 was 100-fold diluted in 3 ml LB broth and incubated until the exponential phase was reached. Bacterial cells were collected by centrifugation and resuspended in the same volume of sterile PBS. The bacterial suspension was labeled with a final concentration of 1 μM of DiSC3(5) and incubated at 37 °C in the dark with continuously shaking at 100 rpm for 5 min. Different concentrations of Val, colistin, and the combination of both were added to the stained bacterial cells, followed by the measurement of the fluorescence level of test groups using an EnSight™ microplate reader.
Intracellular adenosine triphosphate (ATP) measurement
The effects of Val and colistin on intracellular ATP were measured using the Enhanced ATP Assay Kit (Beyotime, Shanghai, China) according to the manufacturer’s instructions with slight modifications40. Briefly, after treatment with Val, colistin, and the combination of both for 2 h, bacterial cells were lysed, and the supernatants were prepared for measurement using an EnSight microplate reader.
Swimming assay
The effects of Val and colistin on the swimming motility of E. coli P47 were investigated according to the previously reported method with minor modifications41. Briefly, cultures of E. coli P47 were inoculated into LB medium supplemented with 0.3% agar and different concentrations of Val, colistin, and the combination of both. The swimming halos were filmed after incubation at 37 °C for 48 h.
Transcriptomic analysis
Transcriptomic analysis of E. coli P47 treated with 1 μg/ml colistin in the absence of Val or the presence of 2 μg/ml Val was performed42. Briefly, Overnight cultures of E. coli P47 were 100-fold diluted in 3 ml CAMHB and then treated with Val, colistin, and the combination of both at 37 °C until an OD600 of 0.6 was reached. After treatment, total RNA from samples was extracted using the Bacteria RNA Extraction Kit (Vazyme, Nanjing, China) and sent to Shanghai Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China) for sequencing using the Illumina Hiseq system. Bioinformatics analysis was performed using the Majorbio Cloud Platform (www.majorbio.com). Differential expression analysis was performed with the DESeq2 package43. Functional annotation was performed with the GO and KEGG databases44,45.
Statistical analysis
All data collected from at least triplicates were presented as means ± SD. Statistical analysis was performed using GraphPad Prism 9.0 to calculate the p values (*p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001) with unpaired t-test between two groups or one-way ANOVA.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Val acted synergistically with colistin in eliminating both colistin-resistant and colistin-susceptible isolates and slow down the development of colistin resistance in vitro
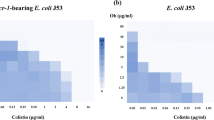
In this study, we constructed a checkerboard assay to determine the synergistic bactericidal effect of Val and colistin using colistin-resistant isolates and colistin-susceptible isolates in vitro (Fig. 1A and Supplementary Fig. S1). Four clinical colistin-resistant strains were used for screening, including E. coli P47 carrying mcr-1, K. pneumoniae strains P30 and 80 both with mgrB inactivation, and A. pittii CS17 with pmrC mutation. As shown in the isobolograms, the synergistic bactericidal effects of Val and colistin on the four colistin-resistant strains were observed with FICIs being 0.312, ≤0.0625, ≤0.0938, and ≤0.0156, respectively. The effects of Val and colistin on colistin-susceptible strains were also tested using three strains including E. coli J53, K. pneumoniae MGH78578, and A. pittii P87. As illustrated in the isobolograms, Val also acted synergistically with colistin in eliminating colistin-susceptible strains, with FICIs of 0.3125, ≤0.28125, and ≤0.375, respectively, whereas the bactericidal efficacy was lower than that on colistin-resistant strains.
A Isobolograms showing the synergistic effect of Val and colistin in colistin-resistant isolates and colistin-susceptible isolates (Supplementary Table S1). Experiments were performed in biological triplicate. The x axis and y axis are FICVal and FICColistin, respectively. B Time-kill curve of colistin-resistant strains in the treatment of Val, colistin, and the combination of both. Experiments were performed in biological triplicate, and the data were presented as means ± SD. C The addition of Val prevented the emergence and development of colistin resistance in vitro. E. coli J53 was incubated with colistin (0.5, 1, 2, 4, 8 μg/ml) and the combination of Val (2 μg/ml) and colistin (0.125, 0.25, 0.5, 1, 2 μg/ml). K. pneumoniae 78578 and A. pittii CS17 was incubated with colistin (0.5, 1, 2, 4, 8 μg/ml) and the combination of Val (2 μg/ml) and colistin (0.25, 0.5, 1, 2, 4 μg/ml). A. pittii CS17 was incubated with colistin (1, 2, 4, 8, 16 μg/ml) and the combination of Val (2 μg/ml) and colistin (0.25, 0.5, 1, 2, 4 μg/ml). Val valnemulin HCl, CT colistin.
To further evaluate the efficacy of the combination of Val and colistin on colistin-resistant strains, a time-kill kinetic assay was conducted using three of the aforementioned colistin-resistant strains including E. coli P47, K. pneumoniae P30 and clinical A. pittii CS17 with colistin MIC at 8, 64, and 16 μg/ml, respectively (Fig. 1B). The growth of the test strains could not be suppressed with monotherapy of Val or colistin, suggesting that neither Val at 2 μg/ml nor colistin at 2 μg/ml (32 μg/ml for K. pneumoniae P30 and 16 μg/ml for A. pittii CS17) could effectively kill the strains within 24 h. In contrast, the combinational use of Val and colistin could effectively eradicate almost all strains by reducing the loads of bacteria by ≈109-fold within the 24-h treatment.
To explore the possibility of Val in preventing the emergence and development of colistin resistance, resistance development assays were performed using colistin-susceptible strains (Fig. 1C). Although E. coli J53 treated with colistin alone or combined with Val did not develop colistin resistance, serial passage of K. pneumoniae MGH78578 and A. pittii P87 with sub-minimum inhibitory (sub-MIC) of colistin caused a rapid increase in colistin MIC by 64-fold and 16-fold, respectively. By contrast, the addition of 2 μg/ml Val mitigates the increase in colistin MIC by 16-fold and 1-fold, respectively, indicating that the combinational use of Val and colistin could be a promising strategy for preventing the development of colistin resistance.
Bactericidal effect of Val and colistin combination in mammalian cells and mice infection models
Given the well-established utility of E. coli as a model organism in microbiological research, we have chosen the strain E. coli P47 for our subsequent investigations. After the observation of the synergistic effect of Val and colistin on killing colistin-resistant strains, we then evaluated the bactericidal efficacy of the combinational treatment on intracellular bacteria in the cell infection model. We first evaluated the cytotoxicity of Val and colistin on mammalian cells using MTT assays. The results showed that over 94% of HEK293T cells were viable after the addition of Val, colistin, and the combination of both, suggesting that Val and colistin had no cytotoxic effect on mammalian cells (Fig. 2A). In the cell infection model, intracellular bacteria load in Vero cells in treating Val, colistin, and their combination were compared by counting viable bacterial cells (Fig. 2B). Compared with the untreated group, 2 μg/ml Val could not kill intracellular bacteria and 2 μg/ml colistin could slightly reduce viable intracellular bacteria. However, combinational use of 2 μg/ml Val and 2 μg/ml colistin caused a dramatic reduction in the number of viable bacteria by 1000-fold, indicating that the combination had profound antibacterial effects on intracellular pathogens.
A Cytotoxicity determination of Val on HEK293T cells using the MTT assay. HEK293T cells were incubated in the presence of Val, colistin, and the combination of both. B Determination of the intracellular bacteria load of E. coli P47 in Vero cells treated with Val, colistin, and their combination. C Survival rates of mice infected by colistin-resistant clinical E. coli P80 with the treatment of Val, colistin, and the combination of both (n = 5/group). Val valnemulin HCl, CT colistin. Experiments were performed in triplicate and the data were presented as means ± SD. Statistical analysis was carried out using GraphPad Prism 9.0 to calculate the p values (ns not significant, *p < 0.05, ***p < 0.001, and ****p < 0.0001) with unpaired t-test between two groups.
We further evaluated the bactericidal efficacy of the Val and colistin combinational therapy in vivo using a mouse infection model of sepsis (Fig. 2C). After infected with 4 × 108 CFU/ml colistin-resistant E. coli P80, all mice died within 36 h upon treatment with saline, colistin (4 mg/kg), and Val (2 mg/kg). Combinational therapy of Val and colistin could sharply increase the survival rate by rescuing 60% of the animals at 48 h post-infection, suggesting that Val could effectively restore the bactericidal effect of colistin in vivo. Despite the effect of Val in recovering the antibacterial effect of colistin not reaching statistical significance, a discernible trend was evident. These data highlighted the combinational use of Val and colistin could be a promising potential strategy to combat colistin-resistant pathogens.
Val potentiates the membrane permeabilizing activity of colistin to cause membrane disruption
Having shown that Val could restore the bactericidal effect of colistin in vitro and in vivo, we then performed SEM imaging to visualize the morphology of E. coli P47 during the treatment with Val and colistin (Fig. 3A). No morphological changes were observed when E. coli P47 was treated with sub-MIC of Val (2 μg/ml) and colistin (2 μg/ml). However, the shrinkage of the cell membrane was visualized with the addition of the combination of Val and colistin, indicating that their combination caused the leakage of cellular contents and finally cell death.
A SEM imaging of E. coli P47 treated with Val (2 μg/ml), colistin (2 and 64 μg/ml), and the combination of both (Val 2 μg/ml + colistin 2 μg/ml). B Determination of the bacterial membrane permeability of E. coli P47, K. pneumoniae P30, and A. pittii CS17 treated with Val (8 μg/ml), colistin (0, 1, 2 μg/ml), and their combination using SYTOX Green staining. Val valnemulin HCl, CT colistin. Experiments were performed in biological triplicate and the data were presented as means ± SD. The statistical significance of colistin plus Val versus colistin was analyzed using an unpaired t-test (****p < 0.0001). Statistical significance of colistin plus Val versus Val was determined by one-way ANOVA, and shown as ####p < 0.0001.
Previous studies reported that the bactericidal effect of colistin is dependent on specifically binding to lipid A in lipopolysaccharide in the bacterial cell membrane, resulting in the membrane permeabilization and leakage of cellular contents, and the final cell death46. Meanwhile, resistance to colistin mainly occurs based on lipid A modification, leading to the prevention of colistin binding47. Therefore, we hypothesized that Val might facilitate the colistin damage on the bacterial membrane by potentiating the membrane permeabilizing activity of colistin. Colistin-resistant strains E. coli P47, K. pneumoniae P30, and A. pittii CS17 were stained with SYTOX Green to evaluate the synergistic effects of Val and colistin on the bacterial membrane permeability. Although Val (8 μg/ml) could not permeabilize bacterial membranes, the combinational use of Val (8 μg/ml) and colistin significantly increased the membrane permeability compared to the effect of the monotreatment of colistin (Fig. 3B). These data demonstrated that Val could restore the membrane permeabilizing effect of colistin on colistin-resistant strains.
Val dissipated bacterial PMF
Previous studies have reported that PMF, a fundamental driving force for various physiological functions, was an unprecedented antibacterial target located in bacterial cell membranes48. Considering that colistin is a membrane-targeting antimicrobial cyclic peptide, it is plausible that Val might enhance the bactericidal efficacy of colistin by dissipating PMF. To validate this hypothesis, we determined bacterial PMF using three different approaches. First, since PMF was maintained by the homeostasis of pH controlled by proton pumps and dependent on the membrane potential49, a membrane-permeable fluorescent probe DiSC3(5) was used to determine bacterial membrane potential. DiSC3(5) could be accumulated in polarized E. coli P47 cells with no treatment50. Upon dissipating bacterial membrane potential, DiSC3(5) would be rapidly released into the medium, and strong signals could be monitored fluorometrically39. As shown in Fig. 4A, compared with the monotreatment of colistin ranging from 1 to 16 μg/ml, the addition of Val (8 μg/ml) and colistin resulted in a significant increase in fluorescence intensity, indicating that Val could enhance the activity of colistin in dissipating bacterial membrane potential. Meanwhile, the combinational use of colistin and Val showed a stronger depolarization effect than the monotreatment of Val, which further confirmed that Val and colistin acted synergistically in dissipating bacterial membrane potential.
A Determination of membrane potential in E. coli P47 upon treatment with Val, colistin, and their combination using DiSC3(5) staining. B Measurement of intracellular ATP level of E. coli P47 with the addition of Val, colistin, and their combination. C Bacterial swimming motility test of E. coli P47 inoculated on semisolid culture medium containing Val, colistin, and their combination. Val valnemulin HCl, CT colistin. Experiments were performed in triplicate and the data were presented as means ± SD. The statistical significance of colistin plus Val versus colistin was analyzed using an unpaired t-test (*p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001). The statistical significance of colistin plus Val versus Val was determined by one-way ANOVA and shown as ####p < 0.0001.
Given that PMF was the driving force of the synthesis of ATP51, we next evaluated the ATP level in E. coli P47 treated with Val, colistin, and their combination (Fig. 4B). Although Val (8 μg/ml) could not cause a significant decrease in the ATP content of bacterial cells, the combinational use of colistin and Val significantly reduced the intracellular ATP level compared to that of monotreatment with Val or colistin, suggesting that Val had a synergistic effect with colistin in inhibiting ATP synthesis.
Subsequently, a bacterial swimming motility test was performed since PMF also drives bacterial motility52. Bacterial cells at ~4 × 108 CFU/ml were inoculated in motility assay, while bacteria at ~105 CFU/ml was inoculated in checkerboard analysis (Supplementary Fig. S1). The killing effects of Val (2 μg/ml), colistin (1 μg/ml), and the combination of both in motility assay were evaluated by plotting the time-kill curves (Supplementary Fig. S2), suggesting that the addition of Val, colistin, and their combination could not decrease bacterial viability. Given the irregular shape of the bacterial colonies, which precludes precise measurement of the migration distance, we speculated that statistical analysis is not appropriate for the current data.
As shown in Fig. 4C, there was no change in the migration distance of E. coli P47 inoculated on a semisolid culture medium containing Val (2 μg/ml) or colistin (1 μg/ml). However, the combinational use of Val (2 μg/ml) and colistin (1 μg/ml) could effectively decrease the migration distance, suggesting that Val and colistin acted synergistically in inhibiting bacterial motility.
To confirm the synergistic bactericidal effect of colistin and PMF dissipater, the MICs of colistin against E. coli P47 and E. coli J53 was also determined in the presence of carbonyl cyanide m-chlorophenylhydrazone (CCCP), a classical PMF disrupter (Supplementary Table S2). CCCP show no antibacterial effect against E. coli P47 and E. coli J53 within the range of tested concentrations (MIC > 16 μg/ml). The colistin MIC was found to decrease with the addition of CCCP in a dose-dependent manner. CCCP showed synergistic antibacterial effects against E. coli P47 and E. coli J53 with colistin with FICI of ≤0.25 and ≤0.375, respectively.
Collectively, these findings demonstrated that Val and colistin showed synergistic effects in dissipating PMF, thereby inhibiting various physiological functions including ATP synthesis and bacterial motility, and finally boosting bactericidal antimicrobial activities.
Val induced alteration in gene expression
To demonstrate the molecular mechanisms underlying the bactericidal effect of the combination of Val and colistin, we performed transcriptomic analysis on E. coli P47 treated with colistin alone and the combination of colistin and Val to evaluate the changes in gene expression levels. A total of 1545 upregulated and 318 downregulated genes were identified in E. coli P47 treated with the combination of Val and colistin compared with that treated with colistin alone (Fig. 5A). GO annotation analysis suggested that these upregulated differentially expressed genes (DEGs) were associated with biological processes (e.g., cell motility and metal ion homeostasis), cellular components (e.g., cell projection), and molecular functions (e.g., carbohydrate transmembrane transporter activity) (Fig. 5B). GO enrichment analysis revealed that these upregulated DEGs were involved in membrane and transport, and downregulated DEGs were enriched in cell motility especially flagellum-dependent cell motility (Fig. 5C, D). The expression of genes associated with the membrane component constituted the most significantly upregulated module in response to the combinatorial treatment. Specifically, the transcriptional upregulation of genes encoding efflux pumps is observed, which encompasses members of the ATP-binding cassette (ABC) superfamily, including yddA, malK, malF, malG, phnC, yciO, and malA. Additionally, the expression of genes from the major facilitator superfamily (MFS), such as ydiM, setC, ydjE, entS, ydiN, as well as those belonging to the resistance-nodulation-division (RND) family (acrE, acrF) are also upregulated53. It is plausible that the induction of efflux mechanisms in bacteria is a response to the synergistic stress exerted by the combination of colistin and Val54. In contrast, the expression levels of genes encoding the flagella biosynthesis proteins (fliT, fliS, flgN, fliQ, fliO, fliR, fliH, fliL, fliJ, flgD) and the structure proteins of flagella body (flgB, flgA, yegR, fliN, fliM, fliD, flgK, flgL, fliE, flgG, flgH, flgC, fliF, flgF, flgE, fliC) were significantly downregulated53, which was consistent with the suppression in bacterial cell motility upon treatment with the combinational use of Val and colistin (Fig. 5E).
A Volcano plot of the distribution of gene expression difference. The x axis represents the difference in expression level (FC value). The y axis represents the corresponding statistically value. An adjusted p value < 0.05 (Student’s t-test) was applied as the cutoff for significantly differently expressed genes (DEGs). B GO (gene ontology) annotation analysis of the DEGs. C GO enrichment analysis of upregulated DEGs. D GO enrichment analysis of downregulated DEGs. The ten most significantly enriched pathways are shown in (C) and (D). E Selected DEGs involved in membrane components and cell motility.
Discussion
With the current situation that the evolution of antimicrobial resistance is overwhelming our antibiotic defenses55, the discovery of new antibiotic adjuvants provides a promising strategy to compensate for the inefficacy of antibiotics and meet clinical needs15. Previous studies have reported that the combinational use of colistin and adjuvants could effectively extend the life of colistin by restoring colistin bactericidal activity in treating infections caused by colistin-resistant pathogens56.
In this study, we found that Val acted synergistically with colistin on dissipating bacterial membrane potential, resulting in the failure in ion and carbohydrate transportation. According to our previous study, several bacterial membrane dissipaters have been identified as colistin adjuvants21,29. Colistin has been reported to bind to LPS and displace membrane-stabilizing magnesium and calcium ions, leading to increased membrane permeability and uptake of colistin into the periplasm57. The strength of electrostatic interactions of divalent cations (Mg2+ and Ca2+) and the head group of LPS was dependent on the electric field58. Val, a PMF dissipater, caused the disruption of bacterial outer membrane potential. The addition of Val weakens the electrostatic interactions of divalent ions and LPS, which could lead to increased disorder in chain packing in lipids and membrane thinning. In addition, a weakening of divalent ions binding could enhance the binding efficacy of colistin to LPS. To this end, Val acts synergistically with colistin in eliminating bacteria. Despite Val having only a weak antibacterial effect on its own, it effectively restored the antibacterial activity of colistin against both colistin-resistant and colistin-susceptible Gram-negative strains including E. coli, K. pneumoniae, and A. pittii, both in vitro and in vivo. It is well known that the significant nephrotoxicity and neurotoxicity associated with colistin in mammalian cells represents a major limitation for its clinical use59. Fortunately, no cytotoxicity of Val was observed on mammalian cells60, thus the application of Val efficaciously reduced the treatment dosage of colistin, which further reduced the risk of side effects of colistin on mammals.
Intracellular pathogens, such as enteroinvasive E. coli, are capable of invading host cells, which allows them to escape and survive the clearance by the host’s immune defenses or antimicrobial agents, leading to the establishment, persistence, and propagation of infections61. Furthermore, intracellular bacteria help the survival of tumor cells since the tumor-resident bacteria plays an important role in regulating and benefiting cancer metastasis. Therefore, intracellular bacteria are regarded as a new potential therapeutic target in cancer treatment62. In this study, the synergistic antibacterial effect of Val and colistin on intracellular bacteria also indicated the potential of combinational use in cancer treatment.
In our mechanistic studies, Val was found to enhance the activity of colistin in permeabilizing bacterial cell membranes, which finally disrupted bacterial membranes and lead to cell death. According to the transcriptomic analysis, it is possible that the upregulation of membrane-associated genes compensated for the attack from Val and colistin combinations. Apart from the membrane disruption, the collapse of PMF caused by the combination of Val and colistin resulted in a reduced intracellular level of ATP. Considering that ATP acts as the primary energy source for almost all biological processes63, ATP could be regarded as the metabolic reporter64. The reduction in intracellular level of ATP caused by Val and colistin combination could cause perturbations to bacterial metabolic homeostasis or even death65.
In addition, bacterial cell motility was also suppressed by the combination of Val and colistin. Both PMF and ATP are essential for flagella formation crucial for bacterial motility. ATP is necessary for the initial step of flagella protein translocation, while PMF accelerates the translocation processes and drives flagella motor rotation52,66. Meanwhile, there was a decrease in the expression levels of motility-related genes, including those associated with flagellum assembly as well as flagellum-dependent swarming and swimming motility. Collectively, considering that flagellum-mediated motility contributes to virulence by enabling strains to colonize and evade the host immune response67, the synergistic suppressive effect of Val and colistin on bacterial motility might enhance the therapeutic efficacy in clinical trials68 (Fig. 6). Other mechanisms, such as a disruption of iron homeostasis or an increased expression of pmrA/pmrB, are not involved.
The combinational use of Val and colistin caused bacterial membrane permeabilization ① and membrane disruption ②, resulting in cytosol leakage and metabolic perturbation. Val also acted synergistically with colistin in dissipating bacterial PMF ③. The collapse of PMF leads to the reduced intracellular ATP level ④ and suppression of flagellum-dependent motility ⑤. The decrease in intracellular ATP level caused the perturbations to bacterial metabolic homeostasis or even death ⑥. The expression of cell motility-related genes was downregulated by Val and colistin combination ⑦. The inhibition in bacterial motility could inhibit the colonization ⑧ and immune evasion of pathogens ⑨, ultimately leading to cell death ⑩. Graphical abstract was created with BioRender.com.
Data availability
Datasets are available on Figshare: https://doi.org/10.6084/m9.figshare.26528215.v1. RNA-sequencing data have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) database (PRJNA1121497). All other data are available from the corresponding authors.
References
Kuehn, B. M. Alarming antimicrobial resistance trends emerge globally. JAMA 324, 223–223 (2020).
Sun, J. et al. Plasmid-encoded tet(X) genes that confer high-level tigecycline resistance in Escherichia coli. Nat. Microbiol. 4, 1457–1464 (2019).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Zhang, R. et al. Nationwide surveillance of clinical carbapenem-resistant Enterobacteriaceae (CRE) strains in China. EBioMedicine 19, 98–106 (2017).
Wright, G. D. & Sutherland, A. D. New strategies for combating multidrug-resistant bacteria. Trends Mol. Med. 13, 260–267 (2007).
Sun, J., Zhang, H., Liu, Y.-H. & Feng, Y. Towards understanding MCR-like colistin resistance. Trends Microbiol. 26, 794–808 (2018).
Boll, J. M. et al. A penicillin-binding protein inhibits selection of colistin-resistant, lipooligosaccharide-deficient Acinetobacter baumannii. Proc. Natl. Acad. Sci. USA 113, E6228–E6237 (2016).
Wagenlehner, F. et al. Systematic review on estimated rates of nephrotoxicity and neurotoxicity in patients treated with polymyxins. Clin. Microbiol. Infect. 27, 671–686 (2021).
Adams, M. D. et al. Resistance to colistin in Acinetobacter baumannii associated with mutations in the PmrAB two-component system. Antimicrob. Agents Chemother. 53, 3628–3634 (2009).
Giani, T. et al. Large nosocomial outbreak of colistin-resistant, carbapenemase-producing Klebsiella pneumoniae traced to clonal expansion of an mgrB deletion mutant. J. Clin. Microbiol. 53, 3341–3344 (2015).
Arcilla, M. S. et al. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect. Dis. 16, 147–149 (2016).
Falgenhauer, L. et al. Colistin resistance gene mcr-1 in extended-spectrum β-lactamase-producing and carbapenemase-producing gram-negative bacteria in Germany. Lancet Infect. Dis. 16, 282–283 (2016).
Yao, X., Doi, Y., Zeng, L., Lv, L. & Liu, J.-H. Carbapenem-resistant and colistin-resistant Escherichia coli co-producing NDM-9 and MCR-1. Lancet Infect. Dis. 16, 288–289 (2016).
Poirel, L., Kieffer, N., Liassine, N., Thanh, D. & Nordmann, P. Plasmid-mediated carbapenem and colistin resistance in a clinical isolate of Escherichia coli. Lancet Infect. Dis. 16, 281 (2016).
Wright, G. D. Antibiotic adjuvants: rescuing antibiotics from resistance. Trends Microbiol. 24, 862–871 (2016).
Wang, X. et al. NhaA: a promising adjuvant target for colistin against resistant Escherichia coli. Int. J. Biol. Macromol. 268, 131833 (2024).
Li, J. et al. The synergistic antibacterial activity and mechanism of colistin-oxethazaine combination against gram-negative pathogens. Front. Pharmacol. 15, 1363441 (2024).
Gadar, K. et al. Disrupting iron homeostasis can potentiate colistin activity and overcome colistin resistance mechanisms in gram-negative bacteria. Commun. Biol. 6, 937 (2023).
Zhong, Z.-X. et al. Natural flavonoids disrupt bacterial iron homeostasis to potentiate colistin efficacy. Sci. Adv. 9, eadg4205 (2023).
Liu, Y. et al. Melatonin overcomes MCR-mediated colistin resistance in gram-negative pathogens. Theranostics 10, 10697 (2020).
Xu, C. et al. Otilonium bromide boosts antimicrobial activities of colistin against gram-negative pathogens and their persisters. Commun. Biol. 5, 1–11 (2022).
Stipkovits, L., Ripley, P., Varga, J. & Palfi, V. Use of valnemulin in the control of Mycoplasma bovis infection under field conditions. Vet. Rec. 148, 399–402 (2001).
Richter, M. F. & Hergenrother, P. J. The challenge of converting gram‐positive‐only compounds into broad‐spectrum antibiotics. Ann. N. Y. Acad. Sci. 1435, 18–38 (2019).
Dip, R., Nemet, Z., Schiessl, B., Klein, U. & Strehlau, G. Efficacy and tolerability of early administration of valnemulin hydrochloride premix on epizootic rabbit enteropathy. Vet. J. 204, 309–314 (2015).
Poulsen, S. M., Karlsson, M., Johansson, L. B. & Vester, B. The pleuromutilin drugs tiamulin and valnemulin bind to the RNA at the peptidyl transferase centre on the ribosome. Mol. Microbiol. 41, 1091–1099 (2001).
Siricilla, S. et al. A new combination of a pleuromutilin derivative and doxycycline for treatment of multidrug-resistant Acinetobacter baumannii. J. Med. Chem. 60, 2869–2878 (2017).
Zhang, X. et al. Valnemulin downregulates nitric oxide, prostaglandin E2, and cytokine production via inhibition of NF-κB and MAPK activity. Int. Immunopharmacol. 9, 810–816 (2009).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing 33th edn (Clinical and Laboratory Standards Institute, 2023).
Xu, C. et al. Imidazole type antifungal drugs are effective colistin adjuvants that resensitize colistin-resistant Enterobacteriaceae. Adv. Ther. 3, 2000084 (2020).
Adamowicz, E. M. & Harcombe, W. R. Weakest-link dynamics predict apparent antibiotic interactions in a model cross-feeding community. Antimicrob. Agents Chemother. 64, 00465–00420 (2020).
Liu, Y. et al. Metformin restores tetracyclines susceptibility against multidrug resistant bacteria. Adv. Sci. 7, 1902227 (2020).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature 517, 455 (2015).
Xu, C. et al. Bactericidal, anti-biofilm and anti-virulence activity of vitamin C against carbapenem-resistant hypervirulent Klebsiella pneumoniae. iScience 25, 103894 (2022).
Zeng, P. et al. Investigation of antibiofilm activity, antibacterial activity, and mechanistic studies of an amphiphilic peptide against Acinetobacter baumannii. Biochim. Biophys. Acta (BBA) Biomembr. 1863, 183600 (2021).
Ernstsen, C. L. et al. Detection and quantification of intracellular bacterial colonies by automated, high-throughput microscopy. J. Microbiol. Methods 139, 37–44 (2017).
Mazzolini, R. et al. Engineered live bacteria suppress Pseudomonas aeruginosa infection in mouse lung and dissolve endotracheal-tube biofilms. Nat. Biotechnol. 41, 1089–1098 (2023).
Sun, J. et al. Generation of a broadly useful model for COVID-19 pathogenesis, vaccination, and treatment. Cell 182, 734–743. e735 (2020).
Sochacki, K. A., Barns, K. J., Bucki, R. & Weisshaar, J. C. Real-time attack on single Escherichia coli cells by the human antimicrobial peptide LL-37. Proc. Natl. Acad. Sci. USA 108, E77–E81 (2011).
Te Winkel, J. D., Gray, D. A., Seistrup, K. H., Hamoen, L. W. & Strahl, H. Analysis of antimicrobial-triggered membrane depolarization using voltage sensitive dyes. Front. Cell Dev. Biol. 4, 29 (2016).
Jia, Y. et al. Melatonin prevents conjugative transfer of plasmid-mediated antibiotic resistance genes by disrupting proton motive force. Pharmacol. Res. 175, 105978 (2022).
Lee, J.-H., Kim, Y.-G., Ryu, S. Y., Cho, M. H. & Lee, J. Ginkgolic acids and Ginkgo biloba extract inhibit Escherichia coli O157: H7 and Staphylococcus aureus biofilm formation. Int. J. Food Microbiol. 174, 47–55 (2014).
Chen, C., Cai, J., Shi, J., Wang, Z. & Liu, Y. Resensitizing multidrug-resistant gram-negative bacteria to carbapenems and colistin using disulfiram. Commun. Biol. 6, 810 (2023).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 1–21 (2014).
Consortium, G. O. The gene ontology (GO) database and informatics resource. Nucleic Acids Res. 32, D258–D261 (2004).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
El-Sayed Ahmed, M. A. E. et al. Colistin and its role in the era of antibiotic resistance: an extended review (2000–2019). Emerg. Microbes Infect. 9, 868–885 (2020).
Poirel, L., Jayol, A. & Nordmann, P. Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin. Microbiol. Rev. 30, 557–596 (2017).
Le, D., Krasnopeeva, E., Sinjab, F., Pilizota, T. & Kim, M. Active efflux leads to heterogeneous dissipation of proton motive force by protonophores in bacteria. mBio 12, e0067621 (2021).
Benarroch, J. M. & Asally, M. The microbiologist’s guide to membrane potential dynamics. Trends Microbiol. 28, 304–314 (2020).
Waggoner, A. Optical probes of membrane potential. J. Membr. Biol. 27, 317–334 (1976).
Meyrat, A. & Von Ballmoos, C. ATP synthesis at physiological nucleotide concentrations. Sci. Rep. 9, 3070 (2019).
Minamino, T. & Namba, K. Distinct roles of the FliI ATPase and proton motive force in bacterial flagellar protein export. Nature 451, 485–488 (2008).
Karp, P. D. et al. The EcoCyc Database (2023). EcoSal Plus 11, eesp-0002–eesp-2023 (2023).
Bhattacharyya, S. et al. Efflux-linked accelerated evolution of antibiotic resistance at a population edge. Mol. Cell 82, 4368–4385. e4366 (2022).
Lewis, K. Recover the lost art of drug discovery. Nature 485, 439–440 (2012).
Tyers, M. & Wright, G. D. Drug combinations: a strategy to extend the life of antibiotics in the 21st century. Nat. Rev. Microbiol. 17, 141–155 (2019).
Mohapatra, S. S., Dwibedy, S. K. & Padhy, I. Polymyxins, the last-resort antibiotics: mode of action, resistance emergence, and potential solutions. J. Biosci. 46, 85 (2021).
Khairalla, B. & Brand, I. Membrane potentials trigger molecular-scale rearrangements in the outer membrane of gram-negative bacteria. Langmuir 38, 446–457 (2021).
Velkov, T. et al. Polymyxins for CNS infections: pharmacology and neurotoxicity. Pharmacol. Ther. 181, 85–90 (2018).
Li, G., Wang, H., Zhang, X., Wu, Z. & Yang, H. A Cas9–transcription factor fusion protein enhances homology-directed repair efficiency. J. Biol. Chem. 296, 100525 (2021).
Lewis, A. J., Richards, A. C. & Mulvey, M. A. Invasion of host cells and tissues by uropathogenic bacteria. Microbiol. Spectr. 4, https://doi.org/10.1128/microbiolspec.UTI-0026-2016 (2016).
Schorr, L., Mathies, M., Elinav, E. & Puschhof, J. Intracellular bacteria in cancer—prospects and debates. npj Biofilms Microbiomes 9, 76 (2023).
Holms, H. Flux analysis and control of the central metabolic pathways in Escherichia coli. FEMS Microbiol. Rev. 19, 85–116 (1996).
Lopatkin, A. J. et al. Bacterial metabolic state more accurately predicts antibiotic lethality than growth rate. Nat. Microbiol. 4, 2109–2117 (2019).
Stokes, J. M., Lopatkin, A. J., Lobritz, M. A. & Collins, J. J. Bacterial metabolism and antibiotic efficacy. Cell Metab. 30, 251–259 (2019).
Berg, H. C. The rotary motor of bacterial flagella. Annu. Rev. Biochem. 72, 19–54 (2003).
Wiles, T. J. et al. Swimming motility of a gut bacterial symbiont promotes resistance to intestinal expulsion and enhances inflammation. PLoS Biol. 18, e3000661 (2020).
Matilla, M. A. & Krell, T. Targeting motility and chemotaxis as a strategy to combat bacterial pathogens. Microb. Biotechnol. 16, 2205 (2023).
Acknowledgements
This work was supported by the China Postdoctoral Science Foundation (2023M731375), the National Natural Science Foundation of China (32300156), the Natural Science Foundation of Jiangsu Province (BK20220493), the Senior Talent Start-up Funds of Jiangsu University (5501280007), and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
N.D., C.X., Yuanyuan Li, and Y.Z. conceived and designed the experiments. C.X., Y.Z., L.M., G.Z., C.L., C.Z., Yunbing Li, and X.Z. performed experiments. N.D. and C.X. analyzed data and wrote the manuscript. N.D. and Yuanyuan Li supervised the research project. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Mei-Ling Han, Ruichao Li, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christina Karlsson Rosenthal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, C., Zhang, Y., Ma, L. et al. Valnemulin restores colistin sensitivity against multidrug-resistant gram-negative pathogens. Commun Biol 7, 1122 (2024). https://doi.org/10.1038/s42003-024-06805-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06805-2
- Springer Nature Limited