Abstract
Toll-like receptor 7 (Tlr7) deficiency-accelerated severe COVID-19 is associated with reduced production of interferons (IFNs). However, the underlying mechanisms remain elusive. To address these questions, we utilize Tlr7 and Irf7 deficiency mice, single-cell RNA analysis together with bone marrow transplantation approaches. We demonstrate that at the early phase of infection, SARS-CoV-2 causes the upregulation of Tlr7, Irf7, and IFN pathways in the lungs of the infected mice. The deficiency of Tlr7 and Irf7 globally and/or in immune cells in mice increases the severity of COVID-19 via impaired IFN activation in both immune and/or non-immune cells, leading to increased lung viral loads. These effects are associated with reduced IFN alpha and gamma levels in the circulation. The deficiency of Tlr7 tends to cause the reduced production and nuclear translocation of interferon regulatory factor 7 (IRF7) in the lungs of the infected mice, indicative of reduced IRF7 activation. Despite higher amounts of lung viral antigen, Tlr7 or Irf7 deficiency resulted in substantially reduced production of antibodies against SARS-CoV-2, thereby delaying the viral clearance. These results highlight the importance of the activation of TLR7 and IRF7 leading to IFN production on the development of innate and adaptive immunity against COVID-19.

Similar content being viewed by others
Introduction
Despite significant progress in controlling the COVID-19 pandemic through global vaccination efforts, SARS-CoV-2 continues to mutate, and variants of the virus are still spreading and causing severe COVID-19 in vulnerable populations1,2,3. Therefore, it is still imperative to dissect the pathogenesis of acute COVID-19 and long COVID4,5. The wide spectrum of severity observed in COVID-19 in different aging populations with various comorbidities remains an elusive phenomenon. This knowledge gap is not only an academic pursuit but a critical necessity for addressing the immediate challenges posed by the pandemic and strengthening defenses against future viral threats. Impaired interferon (IFN) activity has been recognized to contribute to the development of severe COVID-196,7. As an essential part of innate immunity, IFN, a group of signal proteins consisting of 12 different types I, II, and III IFNs work against viral infections6,8. The specialized IFN action in COVID-19 has potential diagnostic and therapeutic implications8. Autoantibodies neutralizing type I IFNs explain 10 to 20% of severe COVID-19 cases9,10. Moreover, IFNs have been studied for their potential therapeutic implications for COVID-19. Previously vaccinated individuals with pegylated IFN Lambda early treatment exhibited a markedly reduced incidence of hospitalization or necessitated an emergency department visit in comparison to those who received a placebo11. The activation of IFNs typically involves a series of steps triggered by the detection of viral or microbial components. Cells contain specialized receptors called Toll-like receptors (TLRs), one group of pattern recognition receptors (PRRs) that are able to identify particular chemical patterns linked to these invaders when a virus or other infection enters the body12.
Emerging clinical evidence indicates that loss of toll-like receptor 7 (TLR7) function is associated with severe COVID-19. TLR7, encoded by an X-linked gene, is a PRR that is primarily expressed by immune cells such as macrophages, dendritic cells, and B cells13,14,15. TLR7 signaling activation is implicated in innate and adaptive immunity16,17. Rare loss-of-function mutations in a single Tlr7 gene can predispose men under 60 without known risk factors to severe COVID-1918,19. In male patients, Tlr7 loss-of-function variants contribute to disease susceptibility in up to 2% of severe COVID-19 cases20,21. The underlying mechanism by which Tlr7 deficiency mediates severe COVID-19 remains elusive. An adjuvant containing TLR7 nanoparticles stimulates a broader immune response—including antigen-specific CD8+ T cells–against SARS-CoV-2 and heterologous influenza viruses22. TLR7 signaling also causes the isotype shift to IgG2b/2c, as well as the acceleration of germinal center formation in response to virus-like particles or SARS-CoV-2 vaccination23,24. Activation of TLR7 has been implicated in COVID-19 prophylaxis and treatment25. Using CRISPR/Cas9-editing of human stem cell-derived plasmacytoid dendritic cells (pDCs), Sluis et al., demonstrated that TLR7-MyD88 signaling is crucial for the production of antiviral IFNs, whereas TLR2 is responsible for the inflammatory IL-6 responses to SARS-CoV-2 infection26. Furthermore, research has shown that rare variants in Tlr7 result in functional impairment and downregulation of cytokine-mediated signaling in COVID-19 patients27. Despite these advances, there remains a lack of in vivo studies to understand the downstream events following TLR7 activation in response to SARS-CoV-2 infection in animal models of COVID-19. How and where TLR7 is specifically activated after SARS-CoV-2 infection is not clear. Further, whether TLR7 and its downstream signaling are involved in the production of antibodies against SARS-CoV-2 infection remains unclear. Our study addresses these gaps by utilizing a mouse-adapted strain (MA30) infection model28,29.
A family of transcription factors known as IFN regulatory factors (IRFs) is essential to modulating the immune system, especially when it involves immune cell development and antiviral responses30,31. Upon activation, typically through TLRs like TLR7, IRF7 stands out from other IRFs like IRF3 or IRF9, and undergoes phosphorylation and nuclear translocation, leading to the induction of type I IFN32,33. However, whether TLR7 activation is related to the activation of IRF7, a master regulator of IFN production34, and how this signaling activation impacts the innate and adaptive immunity against SARS-CoV-2 infection remains unknown and requires experimental investigation. Inborn errors of IRF7-mediated type I IFN immunity in patients with life-threatening COVID-19 were demonstrated in a 659-patient cohort35, indicating that IRF7-mediated type I IFN immunity is essential to control SARS-CoV-2 infection. Conversely, a study using a mouse model of SARS-CoV-2 based on adeno-associated virus (AAV)-mediated expression of hACE2 shows that deficiency of Irf3 and Irf7 and type I IFN signaling activation did not influence SARS-CoV-2 infection36. Further, Sokal, et al investigated the B cell response and measured RBD-specific IgG serological response to mRNA vaccination in SARS-CoV-2 naive patients with inherited Tlr7, Irf7, or Ifnar1 deficiency, as well as young patients with autoantibodies neutralizing type I IFNs and older individuals with age-associated autoantibodies to type I IFNs37. They found that induction of type I IFN is not required for efficient generation of humoral response against SARS-CoV-2 by mRNA vaccines37. These contradictory results indicate that there is a strong need to experimentally dissect the role of Tlr7, Irf7, and IFN production in the development of innate and adaptive immunity against COVID-19. To address these questions, we utilized Tlr7 and Irf7 deficiency mice, single-cell RNA analysis together with bone marrow transplantation approaches. We found that Tlr7 and Irf7 deficiencies increase the severity of COVID-19 through reduced IRF7 activation leading to reduced IFN and antibody production. Our work sheds light on the importance of the TLR7-IRF7-IFN pathway on innate and adaptive immunity against COVID-19.
Results
Upregulation of Tlr7, Irf7, and IFN pathways in the early phase of SARS-CoV-2 infection
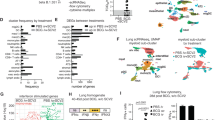
To explore the normal expression pattern of Tlr7 and Irf7 genes and their induction as well as the activation of the IFN pathways, a downstream event of IRF7 induction, to SARS-CoV-2 infection in the pulmonary cell population at the early phase, we performed single-cell RNAseq analysis of the lungs of non-infected and-infected mice, using MA30, a mouse-adapted strain infection model28. In the lungs of non-infected mice, Tlr7, Irf7, but not Tlr3, another pattern recognition receptor critical for sensing double-strand RNA38, are highly expressed in macrophages/dendritic cells (DCs) (Supplementary Fig. 1). Of note, our results document that Tlr7 is expressed on a variety of innate immune cells, including DCs, monocytes, and macrophages in mice. In contrast, in humans, Tlr7 expression is mostly restricted to pDCs and B cells17,39. Next, wild-type (B6) mice were intranasally infected with a sublethal dose of MA30, a mouse-adapted SARS-CoV-2 strain (1 × 104 TCID50)28,29, or PBS. At 2 DPI, scRNAseq was performed on whole lung tissue. A total of 10 clusters were identified in uniform manifold approximation and projection for dimension reduction (UMAP) (Fig. 1a). At 2 DPI, SARS-CoV-2 infection dramatically up-regulated Tlr7 expression but not Tlr3 in myeloid cells as well as in B and erythroid cells, compared with uninfected mice (Fig. 1b, c). Irf7 was globally upregulated in pulmonary immune and non-immune cells as well as parenchymal cells after infection (Fig. 1d). SARS-CoV-2 infection-induced activation of various pathways including IFN alpha and gamma, inflammatory response, and allograft rejection in myeloid cells, endothelial, and epithelial cells as GSEA pathway analysis showed in Fig. 1e, f, and g. Among the activated signaling pathways, IFN alpha and gamma pathways stood out as the highest and most consistent activation in multiple cell populations in the infected lungs as compared to the non-infected lungs (Fig. 1e, f, and g). Together, these results demonstrated that there is an upregulation of Tlr7 and Irf7 in pulmonary myeloid cells and activation of IFN pathways in pulmonary myeloid, endothelial, and epithelial cells, which are the target population of SARS-CoV-2 Orf10 expression in the early phase of infection.
a Major clusters and respective cell types for two 12-week-old female MA30 infected B6 mice (1×104 TCID 50) at 2 DPI and two 12-week-old non-infected B6 mice by scRNA-seq data. Uniform manifold approximation and projection (UMAP) for dimension reduction plot with major cell types of scRNA-seq. Single-cell suspensions from whole infected lungs at 2 DPI and non-infected lungs were processed and sequenced. We identified 10 major clusters including T cells, Myeloid cells, B cells, NK cells, Fibroblast, Endothelial, Erythroid, Non-specific, B & T cells, Epithelial. b–d Expression of Tlr7, Tlr3, and Irf7 in the infected and non-infected lungs. e–g Pathway analysis in myeloid cells, epithelial, and endothelial cells in the infected compared with non-infected lungs.
Global Tlr7 deficiency increases the severity of COVID-19 in mice
To investigate the role of TLR7 in the pathogenesis of COVID-19, we infected sex-matched 12-week-old Tlr7 deficient (Tlr7−/−) or sufficient (Tlr7+/+) mice with a sublethal dose of MA3028 as determined in our recent publication29. After inoculation, we monitored body weight daily (Fig. 2a and b) as well as survival rates (Fig. 2c, d). Additionally, qRT-PCR was used to assay subgenomic N viral transcripts (Fig. 2e, f). Infected Tlr7−/− males and females both lost significantly more body weight (Fig. 2a, b) and had higher pulmonary viral subgenomic RNA loads (Fig. 2e, f) than Tlr7+/+ males and females, respectively. We did not detect any viral load differences in other tissues including the heart, spleen, brain, kidney, or gut (Supplementary Fig. 2). SARS-CoV-2 spike protein staining of the lungs at 6-10 DPI showed higher virus presence in Tlr7−/− males compared with Tlr7+/+ males (with no or very few positive cells) (Fig. 2g, h, representative images of all samples in Supplementary Fig. 3). Further, Tlr7−/− mice had more severe bronchiolar necrosis (Fig. 2I, j, Supplementary Fig. 4), and a tendency to more severe alveolar edema than Tlr7+/+ mice (Fig. 2I, k, Supplementary Fig. 5). Taken together, these results indicate that the deficiency of Tlr7 increased the severity of COVID-19 phenotypes with a higher mortality rate only seen in males (Fig. 2c, d). This finding is consistent with clinical observation that in male but not female patients, Tlr7 loss-of-function variants contribute to disease susceptibility in up to 2% of severe COVID-19 cases18,19,20,21.
a Male Tlr7–/– mice lost more body weight compared to age-matched male Tlr7+/+ mice post-MA30 infection. 12-week-old male Tlr7–/– mice (n = 4) and age-matched Tlr7+/+ (n = 4) male mice were intranasally inoculated with MA30 infection (1 × 104 TCID 50). Body weights were monitored daily. Results are shown as mean ± SEM. The body weight difference is analyzed by a mixed-effects model. P = 0.0030. b Female Tlr7–/– mice lost more body weight compared to age-matched female Tlr7+/+ mice post-MA30 infection. 12-week-old female Tlr7-/- mice (n = 10) and age-matched Tlr7+/+ (n = 9) female mice were administrated with MA30 infection (1 × 104 TCID 50). Body weights were monitored daily. Results are shown as mean ± SEM. The body weight difference is analyzed by a mixed model. P < 0.0001. c Survival analysis of male Tlr7–/– mice and age-matched male Tlr7+/+ mice post MA30 infection. 12-week-old male Tlr7–/– mice (n = 4) and age-matched Tlr7+/+ (n = 5) male mice were administrated with MA30 infection (1 × 104 TCID 50). The comparison of survival curves is analyzed by the Log-rank (Mantel-Cox) test. P = 0.0504. d Survival analysis of female Tlr7–/– mice and age-matched male Tlr7+/+ mice post MA30 infection. 12-week-old female Tlr7–/– mice (n = 10) and age-matched Tlr7+/+ (n = 9) female mice were administrated with MA30 infection (1 × 104 TCID 50). The comparison of survival curves is analyzed by the Log-rank (Mantel-Cox) model. P = 0.1682. e Male Tlr7–/– mice were detected with higher viral load in lungs compared to age-matched male Tlr7+/+ mice post MA30 infection at 6-10 DPI. 12-week-old male Tlr7–/– mice (n = 4) and age-matched Tlr7+/+ (n = 4) male mice were administrated with MA30 infection (1 × 104 TCID 50) and lung tissue was collected at 6–10 DPI. Subgenomic viral load was detected by qRT-PCR. Results are shown as mean ± SEM. Comparison between Tlr7–/– mice and Tlr7+/+ mice is analyzed by unpaired t-test. P = 0.0229. f Female Tlr7–/– mice were detected with higher viral load in lungs compared to age-matched female Tlr7+/+ mice post MA30 infection at 5–7 DPI and 14 DPI. 12-week-old female Tlr7–/– mice (n = 5 for 5-7 DPI and 14 DPI) and age-matched Tlr7+/+ (n = 4 for 7 DPI, n = 5 for 14 DPI) male mice were administrated with MA30 infection (1 × 104 TCID 50) and lung tissue was collected at endpoints. Subgenomic viral load was detected by qRT-PCR. Results are shown as mean ± SEM. Comparison between Tlr7–/– mice and Tlr7+/+ mice is analyzed by unpaired t-test. P = 0.0317 for 5-7 DPI; P = 0.0009 for 14 DPI. g, h Representative images and quantitative analysis of SARS-CoV-2 spike protein (green) immunofluorescence staining. Nuclei were stained with DAPI (white). Auto-fluorescence is presented with a yellow signal. 12-week-old male Tlr7–/– mice (n = 4) at 6-10 DPI and age-matched Tlr7+/+ (n = 4) at 6 DPI were included for analysis. Comparison is analyzed by unpaired t-test. P = 0.0429. i–k Representative images and quantitative histology analysis of H and E staining of infected Tlr7+/+ and Tlr7–/– mice. 12-week-old male Tlr7–/– mice (n = 4) at 6–10 DPI and age-matched Tlr7+/+ (n = 4) at 6 DPI were included for analysis. Tlr7+/+ mice have minimal to mild pulmonary pathology characterized by scattered interstitial inflammation. Tlr7–/– mice exhibit mild to moderate pathology with widespread interstitial infiltrate and edema. Interstitial inflammation in Tlr7+/+ mice is limited to isolated foci of histiocytic and neutrophilic inflammation (red arrow). In Tlr7–/– mice, all pulmonary compartments are affected with alveoli containing edema, inflammatory cells infiltrate alveolar septae and surround vessels (black arrow), and bronchioles are lined by epithelial cells exhibiting degeneration and necrosis (black asterisks). Inset: higher magnification of bronchiolar epithelial degeneration and necrosis. Comparison is analyzed by unpaired t-test. P = 0.0321 of bronchiolar necrosis. P = 0.207 for alveolar edema.
Tlr7 deficiency in hematopoietic cells increases the severity of COVID-19
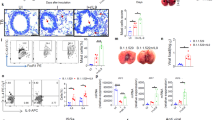
As shown in Fig. 1 and Supplementary Fig. 1, Tlr7 is mainly expressed in immune cells such as DCs and macrophages in mice. To dissect the impact of Tlr7 deficiency in the immune cells on COVID-19, we generated chimeric mice carrying Tlr7−/− (B6-Tlr7−/-BM) or Tlr7+/+ hematopoietic cells (B6-Tlr7+/+BM) by transplanting bone marrow (BM) cells from Tlr7−/− or Tlr7+/+ CD45.2 + B6 mice into Tlr7+/+ CD45.1 + B6 mice, respectively. At 5- and 8-weeks post-BM transplantation (BMT), the reconstituted rates in B6-Tlr7−/-BM and B6-Tlr7+/+BW mice were determined by CD45.1/CD45.2 ratios with flow cytometry analysis were around 98% and 80% in PBMC and BAL respectively (Supplementary Fig. 6). This result documented the successful establishment of the chimeric B6-Tlr7−/-BM or B6-Tlr7+/+BM mice in the circulation and tissues. The mice in both groups at 10 weeks post-BMT were intranasally inoculated with 1×104 TCID50 MA30 and body weights were monitored daily. Consistent with the body weight changes of global Tlr7−/− mice shown in Fig. 2a, the infected B6-Tlr7−/-BM lost significantly more body weight (Fig. 3a) and had a higher pulmonary viral subgenomic RNA load (Fig. 3b) than the infected B6-Tlr7+/+BM. Moreover, there was a trend of a higher number of viral S protein-positive pulmonary cells detected in the lungs of B6-Tlr7−/-BM mice, as compared with that of B6-Tlr7+/+BM mice (Fig. 3c, d). B6-Tlr7−/-BM mice also showed more severe alveolar edema than B6-Tlr7+/+BM mice at 7DPI (Fig. 3e, f, Supplementary Fig. 5).
Bone marrow transplantation was performed on recipient B6 mice at 8 weeks old. After 10 weeks of reconstitution, B6-Tlr7-/-BM (n = 4) and B6-Tlr7+/+BM (n = 4) male mice shown in Figs. 3a to 3f were administrated with MA30 infection (1 × 104 TCID 50) and euthanized at 7 DPI. a B6-Tlr7-/-BM mice lost more body weight compared to B6-Tlr7+/+BM mice post MA30 infection. Body weights were monitored daily. Results are shown by mean ± SEM. Body weight difference is analyzed by two-way ANOVA. P < 0.0001. b B6-Tlr7-/-BM mice were detected with higher viral load in lungs compared to B6-Tlr7+/+BM mice post MA30 infection at 7 DPI. Subgenomic viral load was detected by qRT-PCR. Results are shown by mean ± SEM. Comparison between B6-Tlr7-/-BM mice and B6-Tlr7+/+BM mice is analyzed by unpaired t-test. P = 0.0459. c, d Representative images and quantitative analysis of SARS-CoV-2 spike protein (green) immunofluorescence staining. Nuclei were stained with DAPI (white). Auto-fluorescence is presented with yellow signal. Results are shown as mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.1333. e–f Representative images and quantitative histology analysis of H and E staining of infected B6-Tlr7-/-BM mice and B6-Tlr7+/+BM mice. B6-Tlr7+/+BM mice have minimal pulmonary pathology compared to B6-Tlr7-/-BM with mild to moderate pulmonary pathology. Pathology in B6-Tlr7+/+BM mice is composed of small areas of perivascular to interstitial inflammation (red arrow). Pathology in B6-Tlr7-/-BM mice consists predominately of pulmonary edema (black arrowhead) and perivascular inflammation (black arrow). Results are shown as mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.0321 of bronchiolar necrosis. P = 0.0020. Bone marrow transplantation was performed on recipient K18 mice at 8 weeks old. After 10 weeks of reconstitution, K18-Tlr7-/-BM (n = 4) and K18-Tlr7+/+BM (n = 4) male mice shown in Figs. 3g to 3l were administrated with SARS-CoV-2 WA (2.5 × 102 TCID 50) and euthanized at 7 DPI. g K18-Tlr7-/-BM mice lost more body weight compared to K18-Tlr7+/+BM mice post MA30 infection. Body weights were monitored daily. Results are shown as mean ± SEM. Body weight difference were analyzed using two-way ANOVA. P = 0.0126. h K18-Tlr7-/-BM mice were detected with higher viral load in lungs compared to K18-Tlr7+/+BM mice post MA30 infection at 7 DPI. Subgenomic viral load were detected by qRT-PCR. Results are shown as mean ± SEM. Comparison between K18-Tlr7-/-BM mice and K18-Tlr7+/+BM mice were analyzed using an unpaired t-test. P = 0.0286. i, j Representative images and quantitative analysis of SARS-CoV-2 spike protein (green) immunofluorescence staining. Nuclei were stained with DAPI (white). Auto-fluorescence is presented with a yellow signal. Results are shown as mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.0010. k, l Representative images and quantitative histology analysis of H and E staining of infected K18-Tlr7-/-BM mice and K18-Tlr7+/+BM mice. Comparison is analyzed by unpaired t-test. P = 0.0300.
We further evaluated the role of TLR7 in immune cells in the humanized human ACE2 transgenic mice (K18 mice) infected with the original SARS-CoV-2 strain. K18 mice express human ACE2, a viral receptor for SARS-CoV-2 in mouse epithelial cells under the control of the Keratin 18 promotor40,41. K18 mice infected with SARS-CoV-2 WA1/2020 (SARS-CoV-2 WA), an ancestral SARS-CoV-2 strain, developed severe COVID-19 phenotypes41. This murine model has been widely used for mechanistic, therapeutic, and vaccine COVID-19 research. Previously, we successfully established severe acute and chronic K18 mice by titrating the WA1/2020 strain41. To evaluate the impact of Tlr7 deficiency in immune cells on the development of COVID-19 in the infected K18 mice, we simultaneously generated chimeric K18 mice carrying Tlr7−/− (K18-Tlr7−/-BM)−or Tlr7+/+ hematopoietic cells (K18-Tlr7+/+BM) by transplanting BM from Tlr7−/− or Tlr7+/+ mice into the irradiated K18 mice. At 10 weeks post-BMT, the chimeric K18 mice were infected with a sublethal dose of SARS-CoV-2 WA (2.5 X 102 TCID 50). K18-Tlr7−/-BM mice lost more body weight (Fig. 3g) and had higher pulmonary viral subgenomic RNA loads (Fig. 3h) compared to K18-Tlr7+/+BM mice. Immunofluorescence staining demonstrated a greater number of S protein-positive cells in the lungs of K18-Tlr7−/-BM compared to K18-Tlr7+/+BM (Fig. 3I, j). Histological analysis revealed that K18-Tlr7−/-BM mice had more severe peribronchial inflammation than K18-Tlr7+/+BM mice (Fig. 3k, l, Supplementary Fig. 7).
scRNA seq mapping shows down-regulated Irf7 and IFN pathways in multiple cell populations of infected-Tlr7 −/− mice compared with the infected Tlr7 +/+ mice
To explore the transcriptomic changes in Tlr7−/− mice at the single cell level, we performed scRNA seq analysis of the lungs of the infected Tlr7−/− and Tlr7+/+ female mice at 2 DPI. Only female mice were used to minimize sex confounding the changes observed. Eight cell clusters were identified (Fig. 4a). Higher viral genomic transcripts (orf10) were detected in multiple cell populations of SARS-CoV-2 infected Tlr7−/− mice as compared with Tlr7+/+ mice (Fig. 4b). This further demonstrated that the loss of Tlr7 increased viral infection during the early phase of the infection. As expected, we found that the infected-Tlr7−/− mice had no significant difference in the expression level of Tlr3 (Fig. 4c) and did not express Tlr7 transcripts in almost all cell populations as compared with the infected-Tlr7+/+ mice (Fig. 4d). Specifically, in myeloid cells, B cells, and erythroid cells that specifically increased Tlr7 expression at 2 DPI as shown in Fig. 1b, we only found an upregulation of Tlr7 expression in these three cell populations in Tlr7+/+ mice compared with Tlr7−/− mice, further demonstrating the induction of Tlr7 after infection is specific to these, and not other, cells (Fig. 4d). As compared with the infected Tlr7+/+ mice, the infected-Tlr7−/− mice had significantly lower expression of Irf7 but not Irf3, Irf8, and Irf9 in many cell populations including myeloid cells, and B cells, but not in epithelial cells (Fig. 4e–i). Interestingly, the infected-Tlr7−/− mice had significantly lower levels of the IFN alpha and/or gamma pathways in myeloid, B, T, epithelial, and endothelial cells compared with infected- Tlr7+/+ mice (Fig. 5a–e). This result led us to hypothesize that in the response to SARS-CoV-2 infection, TLR7 may mainly modulate IRF7 activation, thereby controlling the activation of IFN alpha and gamma pathways in the immune cells.
a Major clusters and respective cell types for 12-week-old female MA30 infected Tlr7+/+ mice (n = 2) at 2DPI (1 × 104 TCID 50) and 12- week-old female MA30 infected Tlr7-/- mice (n = 2) at 2DPI (1 × 104 TCID 50) by scRNA-seq data. UMAP for dimension reduction plot with major cell types of scRNA-seq. Single-cell suspensions from whole infected lungs at 2 DPI were processed and sequenced. We identified 10 major clusters including Myeloid cells, T cells, B cells, Endothelial, Erythroid, NK cells, Fibroblast, and Epithelial. b–h Expression of Orf10, Tlr7, Tlr3, Irf7, Irf3, Irf8 and Irf9 in the Tlr7+/+ and Tlr7–/– lungs.
a–e Pathway analysis in myeloid cells, B cells, T cells, epithelial, and endothelial cells in the Tlr7+/+ compared with Tlr7–/– lungs for 12-week-old female MA30 infected Tlr7+/+ mice (n = 2) at 2DPI (1 × 104 TCID 50) and 12- week-old female MA30 infected Tlr7–/– mice (n = 2) at 2DPI (1 × 104 TCID 50) by scRNA-seq data.
Irf7 deficiency also increases the severity of COVID-19
To directly test this hypothesis, we further explored the role of IRF7 in COVID-19 using Irf7 knockout mice (Irf7−/−). We infected Irf7−/− mice with 1×104 TCID50 MA30 (IN). The infected Irf7−/− males lost more body weight and had a higher mortality rate than the infected-Irf7+/+ males (Fig. 6a, b). Viral load was higher in the lungs of Irf7−/− mice than in the Irf7+/+ mice at 4-7 DPI (Fig. 6c). Viral staining showed higher viral spike protein level in Irf7−/− mice as compared to Irf7+/+ mice which showed no or very few viral positive cells (Fig. 6d, e, representative images of each sample in Supplementary Fig. 8). Histologically, Irf7−/− mice had higher bronchiolar necrosis scores than Irf7+/+ mice (Fig. 6f, g, Supplementary Fig. 4). These results demonstrated that the IRF7 signaling activation indeed protects against SARS-CoV-2 infection.
12-week-old male and female Irf7-/- mice (n = 8) and age-matched male and female Irf7+/+ (n = 8) mice were administrated with MA30 infection (1×104 TCID 50). a Irf7-/- mice lost more body weight compared to Irf7+/+ mice post MA30 infection. Body weights were monitored daily. Results are shown by mean ± SEM. Because of the data missing at 6 and 7 DPI, body weights from 0 to 5 DPI are analyzed by mixed-effects analysis. P < 0.0001. b Survival analysis of Irf7-/- mice and age-matched Irf7+/+ mice post MA30 infection. Comparison of survival curves is analyzed by Log-rank (Mantel-Cox) test. P = 0.0008. c Irf7-/- mice were detected with higher viral load in lung compared to Irf7+/+ mice post MA30 infection at 4-7 DPI. Subgenomic viral load of 12-week-old male Irf7-/- mice (n = 8) and age-matched male Irf7+/+ (n = 8) mice was detected by qRT-PCR. Results are shown by mean ± SEM. Comparison is analyzed by unpaired t-test. P < 0.0001. d, e Representative images and quantitative analysis of SARS-CoV-2 spike protein (green) immunofluorescence staining. Nuclei were stained with DAPI (white). Auto-fluorescence is presented with yellow signal. Comparison of 12-week-old male Irf7-/- mice (n = 4) and age-matched male Irf7+/+ (n = 4) mice is analyzed by unpaired t-test. P = 0.0003. f, g Representative images and quantitative histology analysis of H and E staining of 12-week-old male Irf7-/- mice (n = 4) and age-matched male Irf7+/+ (n = 4) mice. Irf7+/+ mice and Irf7-/- mice exhibit mild pulmonary pathology. Pathology in Irf7+/+ mice consists of perivascular to interstitial infiltration by mononuclear cells. Inset: Normal bronchiolar epithelium. Pathology in Irf7-/- mice is characterized by moderate to severe bronchiolar necrosis with interstitial to perivascular inflammation. Inset: necrosis of bronchiolar epithelial cells in Irf7-/- mouse. Comparison is analyzed by unpaired t-test. P = 0.0020.
We further conducted the Bulk RNA analysis of the lungs of the infected Irf7+/+ and Irf7−/− mice at 2DPI (Fig. 7a, b) to monitor the transcriptomic changes of IFN-related genes and pathway activation in these mice. We demonstrated 965 genes up-regulated and 467 genes down-regulated in the infected Irf7+/+ mice compared with the infected Irf7−/− mice (Supplementary data 2). As expected, the dramatic upregulation of Irf7 expression in the infected Irf7+/+ mice compared with the infected Irf7−/− mice further confirms the deficiency of Irf7 gene expression in Irf7−/− mice (Fig. 7a). Additionally, IFN-related genes such as IFN beta 1(Ifnb1), IFN alpha-inducible protein 27 like 2 A (Ifi27l2a), and IFN-induced transmembrane protein 6 (Ifitm6) are on the top list of fifteen up-regulated genes in the Irf7+/+ mice as compared with the infected Irf7−/− mice (Fig. 7a and Supplementary data 2). Consistently, pathway analysis showed the activation of IFN alpha response and IFN gamma response in Irf7+/+ mice compared with Irf7−/− mice (Fig. 7b).
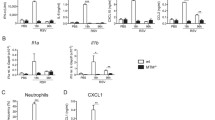
a, b 12-week-old Irf7-/- (n = 3) and age-matched Irf7+/+ (n = 3) males were inoculated with MA30 infection (1 × 104 TCID 50). Lungs were collected at 2 DPI for Bulk RNA analysis. 965 genes were up-regulated and 467 genes down-regulated in Irf7+/+ mice compared with Irf7–/– mice (a) (also shown in Supplementary data 2). Pathway analysis showed activation of 9 pathways in Irf7+/+ mice compared with Irf7–/– mice (b). c, d. 12-week-old B6, Tlr7-/- and lrf7-/- mice were inoculated with MA30 infection (1 × 104 TCID 50). Serums were collected at 2 DPI (n = 5 for each group) or 4 DPI (n = 5 for each B6 mice, n = 4 for Tlr7-/- mice, n = 3 for Irf7-/- mice). IFN-alpha, IFN-beta, IFN-gamma levels in serums were detected by ProcartaPlex Immunoassays at 2DPI and 4DPI. One-tailed unpaired t-test was performed to evaluate the statistical significance between two groups. Values are presented as mean ± SEM. *P < 0.05. **P < 0.01. P = 0.0015 between B6 IFN alpha level and Tlr7-/- IFN alpha level at 2 DPI. P = 0.0015 between B6 IFN alpha level and Irf7-/- IFN alpha level at 2 DPI. P = 0.1021 between B6 IFN beta level and Tlr7-/- IFN beta level at 2 DPI. P = 0.1021 between B6 IFN beta level and Irf7-/- IFN beta level at 2 DPI. P = 0.0137 between B6 IFN gamma level and Tlr7-/- IFN gamma level at 2 DPI. P = 0.0477 between B6 IFN gamma level and Irf7-/- IFN gamma level at 2 DPI.
Reduced IFN Alpha and Gamma levels in the infected Tlr7 −/− and Irf7 −/− mice at 2 DPI
As described above, we demonstrated that both the infected Tlr7 and Irf7 deficient mice had significantly reduced expression of IFN response genes and the activation of IFN alpha and gamma pathways in immune cells and epithelial cells or globally in the lungs. To further evaluate the IFN levels in the circulation, we measured IFN-alpha, IFN-beta, and IFN-gamma levels in the sera collected from the infected age-matched B6, Tlr7−/− and Irf7−/− mice at 2 DPI and 4 DPI by ProcartaPlex Immunoassays. As revealed in Fig. 7c, d, the infected Irf7−/− and Irf7−/− mice at 2 but not 4 DPI had significantly lower levels of serum IFN alpha and gamma but not beta than the infected B6 mice. Moreover, by immunostaining of IRF7 in these infected mice, we documented that the infected Tlr7−/− mice had significantly less Irf7 cytoplasm staining area and tended to have reduced Irf7 nuclear staining area in the lungs than the infected B6 mice at 2 DPI (Supplementary Fig. 9), which is indicative of decreased Irf7 production and activation in the infected Tlr7−/− mice.
Deficiency of Tlr7 and Irf7 results in impaired adapted immunity
We have demonstrated that the infected Tlr7−/−, B6-Tlr7−/-BM, K18-Tlr7−/-BM or Irf7−/− mice had higher viral load determined by qRT-PCR and increased CoV2 protein staining in the lung at 5- to 7-, or 14-DPI compared to control mice (Figs. 2g, h, 3c, e, i, j, 6d and e). These results prompted us to investigate whether deficiency of Tlr7 and Irf7 results in impaired adapted immunity to clear SARS-CoV-2 viruses in the infected mice. We used the direct ELISA method to measure the level of IgG against SARS-CoV-2 spike (S) protein in those infected mice42. We have determined a half maximal IgG absorbance serum dose (AD50) for the infected mice (Fig. 8). We found the infected Tlr7+/+ mice had 1.68-fold higher anti-S IgG levels than the infected Tlr7−/− mice at 14 DPI (Fig. 8a). Additionally, the infected Tlr7+/+ mice had significantly higher or tendency to higher absorbance values than the infected Tlr7−/− mice at 1/31250 and 1/6250 serum dilutions, doses higher and lower to AD50, respectively (Fig. 8b). These findings were also observed in the infected B6-Tlr7−/-BM and K18- Tlr7−/-BM mice as compared to B6-Tlr7+/+BM and K18-Tlr7+/+BM at 7 DPI respectively (Fig. 8c–f). Interestingly, infected Irf7−/− mice did not produce anti-S IgG antibodies compared to Irf7+/+ mice (Fig. 8g, h). Together, these results indicate that deficiency of Tlr7 and Irf7 results in impaired adapted immunity, which explains the delayed viral clearance of SARS-CoV-2 infection in the infected Tlr7−/− and Irf7−/− mice.
a Nonlinear fit curves of IgG antibody reactivity of MA30 (1×104 TCID 50) infected female Tlr7+/+ mice (n = 4) and Tlr7-/- mice (n = 5) at 14 DPI. The half-maximal IgG absorbance serum dose (AD50) titer of Tlr7+/+group is 15699. The AD50 titer of Tlr7-/- group is 9349. b Comparison of absorbance at 450 nm between Tlr7+/+ mice and Tlr7-/- mice at 6250 and 31250 times dilution. Results are shown by mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.1027 for 6250 times dilution; p = 0.0292 for 31250 times dilution. c Nonlinear fit curves of IgG antibody reactivity of MA30 (1 × 104 TCID 50) infected male B6-Tlr+/+BM (n = 4) and B6-Tlr7-/-BM(n = 4) mice at 7 DPI. The AD50 titer of B6-Tlr+/+BM group is 716.9. The AD50 titer of B6-Tlr7-/-BM group is 372.1. d Comparison of absorbance at 450 nm between B6-Tlr7-/-BM (n = 4) and B6-Tlr7+/+BM(n = 4) mice at 250 and 1250 times dilution. Results are shown by mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.0322 for 250 times dilution; p = 0.0486 for 1250 times dilution. e Nonlinear fit curves of IgG antibody reactivity of MA30 (1 × 104 TCID 50) infected male K18-Tlr7+/+BM (n = 4) and K18-Tlr7-/-BM (n = 4) mice at 7 DPI. The AD50 titer of K18-Tlr7+/+BM group is 329.1. The AD50 titer of K18-Tlr7-/-BM group is 273.7. f Comparison of absorbance at 450 nm between K18-Tlr7+/+BM(n = 4) and K18-Tlr7-/-BM (n = 4) mice at 250 and 1250 times dilution. Results are shown by mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.0278 for 250 times dilution; p = 0.0557 for 1250 times dilution. g Nonlinear fit curves of IgG antibody reactivity of MA30 (1 × 104 TCID 50) infected male and female Irf7+/+ mice (n = 8) and Irf7-/- mice (n = 4) at 5–7 DPI. The AD50 titer of Irf7+/+group is 490.0. The IgG levels of Irf7-/- group are too low to fit a nonlinear curve. h Comparison of absorbance at 450 nm between Irf7+/+ mice and Irf7-/- mice at 250 and 1250 times dilution. Results are shown by mean ± SEM. Comparison is analyzed by unpaired t-test. P = 0.0088 for 250 times dilution; p = 0.0397 for 1250 times dilution.
Given the known role of T follicular helper cells as well as co-stimulation in adaptive immune responses43,44, we analyzed expression levels of the T follicular helper cell markers Cxcr5, Il21r, Icos, and Bcl6 (in CD4+ T cells) in infected Tlr7+/+ and Tlr7−/− mice at 2DPI at the single cell level. The expression of Icos in the T cell cluster in Tlr7−/− mice was significantly lower compared with Tlr7+/+ mice (Supplementary Fig. 10a). In myeloid cells, we observed higher levels of Cd80, Cd86, and Cd274, in infected Tlr7+/+ than Tlr7−/− mice at 2DPI. These results provide a potential molecular mechanism by which Tlr7 deficiency mediates impaired adaptive immunity.
Discussion
We document that the deficiency of Tlr7, an X-linked gene globally or in immune cells increases the severity of COVID-19 via impaired Type I and Type II IFN pathway activation in immune and non-immune cells in the infected lung. These results are consistent with the clinical findings obtained from male COVID patients with loss of function Tlr7 variant who tend to develop severe COVID-1918,19,20,21,27,45,46. By using the patient’s PBMC samples, these studies documented that the impaired innate immunity such as the reduced production of IFNs due to Tlr7 deficiency in the circulating immune cells may contribute to the development of severe diseases in these patients18,19,20,21,27,45,46. Of note, Tlr7 gene dose difference between males (one copy) and females (two copies) has been attributed to the observed male-skewed severity seen in COVID-19 patients. Tlr7 is subject to a gene-dose effect stemming from X chromosome inactivation escape47. In our study, since the experiments with Tlr7−/− and Tlr7+/+ male and female mice were infected at different times, we did not compare sex impact on COVID-19. The Tlr7 impact on the male-skewed severity seen in COVID-19 patients is beyond the scope of the current study and warrants further investigation.
Additionally, we demonstrate that Tlr7 expression in immune cells is one of the most important sensors for SARS-CoV-2 infection in the initiation of innate immunity against infection. By single-cell RNA analysis of the lungs of the MA30-infected WT mice and naïve mice at 2 DPI, we directly monitored Tlr7 and its family genes such as Tlr3 expression pattern, and its downstream IFN pathway activation. We found that Tlr7 but not Tlr3 are highly expressed in macrophages/DC and B cells. The expression patterns of Tlr7 on DC and B cells in mice are like those in humans39,48. At an early phase of SARS-CoV-2 infection, Tlr7, but not Tlr3, is upregulated in pulmonary myeloid cells, and activation of type I and II IFN pathways occurs in pulmonary myeloid, epithelial, and endothelial cells. By sc-RNA seq mapping, we observe down-regulated Irf7 and type I and type II IFN pathways and increased viral penetration or spread evidenced by higher viral transcript expression in multiple cell populations of the infected-Tlr7−/− mice compared with the infected Tlr7+/+ mice at 2 DPI. These results underscore that Tlr7 expression in myeloid cells not only senses viral infection but also initiates IFN pathway activation for controlling the viral infection in the early phase of the infection. Of note, the differences observed from the scRNA analysis in our studies cannot illustrate the higher mortality in the male mice. This is because we conducted the scRNA analysis of the lungs of infected Tlr7+/+ and Tlr7−/− females but not both genders to minimize the sex confounding effect on the transcriptomic changes.
It is widely recognized that IRF7 is a master regulator of IFN production34. However, how IRF7 is activated in SARS-CoV-2 infection is not clear. Here, we document that at the early phase of COVID, TLR7 is critical for controlling IRF7 activation for regulating IFN production. First, in non-infected B6 mice, Tlr7 and Irf7 are expressed in myeloid cells. After infection, Tlr7 and Irf7 are highly expressed together in immune and non-immune cells. Second, in the infected B6 mice, myeloid, B and erythroid cells but not T cells, endothelial cells, NK cells, fibroblasts, and epithelial cells specifically express Tlr7 (Fig. 4d). The observed IRF7 activation and IFN responses on the other five clusters that do not express Tlr7 are likely due to autocrine/paracrine effects (Figs. 4 and 5) of IFN production and signaling. Third, after infection, we find that Tlr7 deficiency results in the specific downregulation of Irf7 but no other family genes such as Irf8 and Irf9. The deficiency of Tlr7 and Irf7 reduced the pathway activation and the production of IFN alpha and gamma (Fig. 4 and Fig. 7). Fourth, the infected Tlr7−/− mice had significantly less Irf7 cytoplasm staining positive area and tended to have reduced IRF7 nuclear staining positive area in the lung than the infected B6 mice (Supplementary Fig. 9). Considering that IRF7 nuclear translocation is the key step to regulate IFN production34,49,50, these results indicate that TLR7 may be a main regulator of IRF7 activation for IFN alpha and gamma production in the response to SARS-CoV-2 infection. This indication is consistent with the clinical observation that patients with severe COVID-19 carrying a Tlr7 loss of function mutation had downregulated type I IFN signaling as measured by significantly decreased mRNA expression of Irf719. Finally, the deficiency of Irf7 markedly increases the severity of COVID-19, confirming the critical role of Irf7 in regulating IFN production to enhance the innate immunity against COVID-19 infection. Together, these results indicate that TLR7 regulates the activation of IRF7 for controlling Type I and II IFN production in myeloid cells against COVID-19 infection.
In our studies, we have used multiple approaches including sc RNA seq, qRT-PCR, and immunostainings of S protein for detecting viral load in the lungs collected from both MA30-infected and SARS-CoV-2 WA-infected K18 model at multiple time intervals. The severity of the disease was measured by the loss of body weight, mortality rates, and lung pathology after the infection of Tlr7 and Irf7 deficient mice. Our overall results also demonstrated that the increase in the severity of COVID-19 in Tlr7 and Irf7 deficient mice is related to increased viral load and severe lung pathology in the infected mice. Consistently, in our study, we evaluated the viral loads in Tlr7−/− mice by scRNA seq and qRT-PCR analyses of the infected lungs respectively collected from the infected Tlr7−/− and Tlr7+/+ mice at the same time intervals (2 DPI and 14 DPI). We found that the Tlr7−/− mice had a significantly higher viral load in the lung than Tlr7+/+ mice at 2 DPI and 14 DPI (Fig. 2). However, it is important to note that some of our other studies included variability in our time point comparisons which is an important caveat when interpreting viral load and lung pathology as shown in Figs. 2e, f, h, j, k, 6c, e, and g. The samples for B6 mice were collected at a fixed day (at 7 DPI or 10 DPI), but samples for Tlr7 and Irf7 deficient mice were collected at different days (4-7 DPI or 6-10 DPI). This variability resulted from the more severe diseases phenotype in infected deficient mice compared with the infected B6 mice that resulted in infected deficient mice reaching endpoint criteria earlier than scheduled. For viral loads, previous study28 has shown that the viral replication drops dramatically in the lungs of infected mice from day 2 onward. Thus, the lung viral loads and histological changes of the infected Tlr7 and Irf7 deficient mice for these time intervals remain to be investigated as compared with B6 mice in a different experimental setting.
Furthermore, our results highlight the importance of the activation of TLR7 and IRF7 leading to the production of type I and II IFN on the development of humoral immune response to COVID-19. Tlr7 or Irf7 deficiency respectively results in the reduced production of antibodies or no antibody production against SARS-CoV-2 infection, thereby delaying the viral clearance in the late phase of the infection. T follicular helper cells are a distinct subset of CD4+ T cells, and their primary function involves facilitating B cell maturation and fostering the production of antibodies targeting foreign pathogens51. Inducible costimulator (Icos), a member of the CD28 family of costimulatory molecules, exhibits heightened expression upon the activation of CD4+ T cells. Icos was found elevated expressed in T cells in infected Tlr7 sufficient mice than in Tlr7 deficient mice, implying that TLR7 might play a role in ICOS-involved T follicular helper cell maturation and B cell maturation. Moreover, several B7 family proteins such as Cd80, Cd86, and Cd274 had increased expression in myeloid cells and B cells in infected Tlr7 sufficient mice than in Tlr7 deficient mice, indicating TLR7 plays a role in the co-stimulatory process to activate T cells and B cells52. No antibody detection in the infected Irf7−/− mice and reduced antibody production in the infected Tlr7−/− mice as compared with control mice shown in Fig. 7 clearly indicates that there is also unknown TLR7 independent signaling involved in the antibody production, which requires further investigation. Together, these results underscore the interaction between TLR7 and IRF7 not only in the realm of innate immunity but also in adaptive immune responses. Consistently, Toll-like receptors have been recognized to play a critical role in linking the innate immune to the adaptive immune responses17,53. A recent study shows that TLR7 signaling in B cells is required for antibody responses to SARS-CoV-2 vaccines24. However, there is no report so far to explore the role of TLR7, a downstream event of IRF7 and IFN production in humoral response to COVID-19 infection. Instead, a previous study conducted by Sokal, et al documents that TLR7 activation and induction of type I IFN is not required for efficient generation of a humoral response against SARS-CoV-2 by mRNA vaccines37. In contrast, we report the first experimental results showing that activation of TLR7 and IRF7 and production of IFNs participate in the adaptive immunity directed against COVID-19 infection in the lung. Our finding also underscores the critical role of local innate and adaptive immunity against COVID, highlighting the emerging need for developing an effective intranasal vaccine to control SARS-CoV-2 variant infection with a much longer and more efficient protection than current vaccines such as mRNA vaccines. It is conceivable that this finding may be attributed to the predominant expression of TLR7 and IRF7 on pulmonary myeloid cells, T cells, and B cells after the COVID-19 infection, which play a pivotal role in antigen presentation, thus initiating adaptive immune response and producing antibodies, respectively. However, the exact roles of TLR7, IRF7, and IFN activation in the development of humoral immune response to COVID-19 remains unknown and warrants further investigation. Dissection of these mechanisms requires the utilization of a combination of approaches, such as conditional Tlr7 and Irf7 knock out, specific cell ablation, signal cell and spatial transcriptomics54,55,56,57.
Methods
Mice
Irf7−/− (RBRC01420) mice were purchased from RIKEN, Japan. Tlr7−/−(008380), K18+/−(034860) and C57BL/6 J (B6) were purchased from the Jackson Laboratory and housed in an animal facility at Tulane University.
Study approval
All animal experiments were approved by the Institutional Animal Care and Use Committee at Tulane University. We have complied with all relevant ethical regulations for animal use.
SARS-CoV-2 infection
For SARS-CoV-2, the USA-WA1/2020 isolate was obtained as a seed stock (NR-52281 strain deposited by the Centers for Disease Control and Prevention and obtained through BEI Resources, NIAID, NIH). We passaged the virus in VeroE6-TMPRSS2 cells in DMEM media with 2% FBS and sequence verified to ensure homology with the original patient sequence. Mice were infected intranasally with SARS-CoV-2 (2 X 105 TCID50 for acute lethal dose infection or 1 X 104 TCID50 for sub-lethal dose infection) in an ABSL3 facility. The MA30 strain was obtained as a seed stock from Dr. Stanley Perlman (U. Iowa) and expanded and characterized as described for the USA-WA1/2020 isolate.
Tissue collection and process
The health status of the mice was monitored daily. Euthanasia was performed if the mice exhibit a body weight loss of 25% or upon reaching the predetermined necropsy date. Blood was drawn via cardiac puncture into a BD microtainer and allowed to clot at room temperature for a minimum of 30 min. Subsequently, the blood samples were centrifuged at 3000 rpm for 10 min, and the serum was collected from the upper layer. One-half of the lung was fixed in Z-FIX buffer at room temperature. The remaining lung tissue was divided as follows: half of the residual lung (left side) was immersed in 1 mL of Trizol reagent and stored at −80°C for RNA extraction; the other half of the residual lung (left side) was frozen on dry ice without any medium and stored at −80°C.
Histological analysis and quantification of histopathologic lesions
Lung sections were processed according to standard protocols, stained with hematoxylin and eosin (H&E), and subsequently scanned using the Axio Scan.Z1 (Zeiss, Thornwood, NY). Image acquisition was performed using the Figure Maker tool in HALO (Indica Labs, Albuquerque, NM).
RNA isolation
Tissues were collected in 1 mL of Trizol reagent (Invitrogen, Cat. No. 15596026) and RNA was extracted using the RNeasy Mini Kit (QIAGEN, Hilden, Germany, Cat. No. 74104) according to the manufacturer’s protocol. The RNA concentration was measured using the NanoDrop 2000 spectrophotometer.
Subgenomic N viral copy number detection
A total of 100 ng of RNA was mixed with TaqPath 1-Step Multiplex Master Mix (Thermo Fisher, Cat. No. A15299, Waltham, MA) and FAM-labeled primers (sgm-N-FOR: 5’-CGATCTCTTGTAGATCTGTTCTC-3’, sgm-N-Probe: 5’-FAM-TAACCAGAATGGAGAACGCAGTGGG-TAMRA-3’, sgm-N-REV: 5’-GGTGAACCAAGACGCAGTAT-3’), following the manufacturer’s instructions. The subgenomic N viral copy number was calculated based on standard Cq values. The assay was performed using the ABI QuantStudio 6 system (Thermo Fisher, Waltham, MA).
Bone marrow transplantation
Recipient mice were irradiated with a dose of 950 Rad. We harvested femurs & tibias from donor mice by cleaning muscles near the bones. In the biosafety cabinet, we cut the ends of the femurs and flushed the cells with PBS (with 1%P/S) using a 27 G needle. The solution was passed through a sterile 40-um-nylon Cell Strainer and collected in a 50-ml tube. Then we centrifuged and washed the cells twice with cold PBS and resuspended them in the required volume of PBS. Finally, we injected the cell solution into the recipient mice via the tail vein.
Preparation of single cells from mouse and flow cytometry
Blood collected from the tail using heparinized capillary tubes was transferred into EDTA anti-clotting tubes, and then lyzed with 1x RBC lysis buffer. The mixture was centrifuged and the supernatant was decanted. The cell pellet was resuspended in staining buffer (PBS with 2% BSA). Mice were sacrificed, and the bronchoalveolar lavage (BAL) was gently washed out through the trachea with PBS. The lavage was centrifuged, and cell pellets were resuspended with staining buffer. The samples were blocked with 1:200 CD16/CD32 (eBioscience, 14-0161) for 30 min. They were then stained with an antibody cocktail: Live/Dead staining (Thermo Fisher, L34957) at 1:200; CD45.1, APC, Clone-104 (Invitrogen, 17-0453-82) at 1:100; and CD45.2, eFluor 450, Clone-A20 (eBioscience, 48-0454-82). In general, a 1:100 dilution of the above antibodies was used for the staining. The antibody cocktails were added and incubated at 4 ◦C in the dark for 30 min. After washing twice with FACS buffer, the samples were acquired on BDLSR Fortessa and analyzed using FACS Diva v.6.1.3 software.
Single-cell RNA sequencing and 10x assay
Lung tissues were minced using forceps and small scissors and then digested in 2 mL of serum-free medium containing 2 mg/mL collagenase (MilliporeSigma) and 80 U/mL DNase I (MilliporeSigma) for 60 min at 37 °C41. A target of 5000 live cells per sample was set using the 10× Single Cell RNAseq technology provided by 10× Genomics (10X Genomics)41. Full-length barcoded cDNAs were then generated and amplified by PCR to obtain sufficient mass for library construction41. Pooled libraries, at a final concentration of 1.8 pM, were sequenced using the paired-end single index configuration on the Illumina NextSeq 2000 platform. The raw sequencing data were processed using Cell Ranger version 7.1.0 (10X Genomics), and differentially expressed genes between specified cell clusters were identified using Loupe Cell Browser (10X Genomics)41. Additionally, Seurat suite version v5 was used for further quality control and downstream analysis41. Filtering was conducted to eliminate multiplets and broken cells, and sources of uninteresting variation were regressed out. Variable genes were identified through iterative selection based on the dispersion versus average expression of each gene. For clustering, principal component (PC) analysis was performed for dimensionality reduction. The top 10 PCs were selected using a permutation-based test implemented in Seurat and subsequently passed to UMAP for clustering visualization41. Single-cell RNA sequencing data have been deposited to the GEO database (GSE274686).
Histology
Fixed tissues were processed, cut, and stained by Anatomic Pathology Core at Tulane National Primate Research Center. The histological injury level was scored by a pathologist.
Immunostaining
Lung tissues fixed in zinc formalin (Z-fixed) and embedded in paraffin were sectioned, deparaffinized, and subjected to heat-induced epitope retrieval using both high-pH (Vector Labs H-3301, Olean, NY) and low-pH solutions (Vector Labs H-3300, Olean, NY) 3. Sections were blocked with either 10% goat serum for 40 min at room temperature, followed by incubation with primary antibodies overnight at 4 °C and secondary antibodies for 60 min at room temperature3. Slides were digitally scanned using the Zeiss Axio Scan.Z13. Detailed antibody information is provided in the Reporting Summary Table.
Digital image analysis for quantification of fluorescent marker expression
Slides were stained with DAPI and either SARS-CoV-2 or IRF7 antibodies, and imaged in three channels, including one empty channel for autofluorescence detection3. Regions of interest were delineated around the entire lung section3. Quantification of SARS-CoV-2 fluorescence was conducted using the HighPlex FL v4.1.3 module in HALO. Quantification of IRF7 nuclear and cytoplasmic localization was performed using the Area Quantification FL v2.3.4 module in HALO3. Thresholds for marker detection were established and subsequently verified for accuracy post-analysis by the same pathologist3.
Bulk RNA sequencing
Isolated lung tissue RNA was quantified using the Qubit 3.0 Fluorometer (ThermoFisher Scientific, Waltham, MA). The Agilent 4150 TapeStation (Santa Clara, CA) was utilized to determine the RNA integrity number and fragment sizes (DV200 metrics)3. The Illumina NEBNext Ultra II directional RNA library prep kit (San Diego, CA) was employed for library preparation. For cluster generation, the cDNA libraries were pooled to a final concentration of 1.8 pM3. Sequencing was conducted on the Illumina NextSeq 2000 P2 100 (San Diego, CA) with a minimum input of 750 pM RNA. Gene expression and nucleotide variation were assessed following the processing and mapping of raw reads as previously described [54]. Raw read counts were normalized across all samples3. Differential expression analysis was performed using Cuffdiff, EdgeR, and DESeq (Slug Genomics, University of California, Santa Cruz) 3. Volcano plots were generated in R using the Cuffdiff output. All curated datasets were deposited in the Sequence Read Archive under BioProject number: GSE274586.
ProcartaPlex Immunoassays
A Procarta 3Plex plate (target: IFN alpha, IFN beta, and IFN gamma) for mouse was purchased from Thermo Fisher Scientific. The experiment was performed by the Pathogen Detection & Quantification Core Lab (PDQC) at Tulane National Primate Research Center strictly following the instructions. Briefly, the procedure is analogous to ELISA, with the primary difference being that antibody-coated beads are employed to capture the analyte in mouse serum. Each run included standards of known concentration to establish a standard curve. After fully incubated on a shaker, the beads were completely washed by placing the 96-well plate on a flat magnet for 30 s. The magnet was then removed, and the beads were resuspended in the detection antibody. After an additional incubation and wash, streptavidin-R-phycoerythrin (SAPE) was added. The beads underwent another wash and were then ready for analysis. Data was analyzed with ProcartaPlex Analysis App (Thermo Fisher Scientific).
ELISA of serum antibody activity test
The plates were coated with antigen (SARS-CoV-2 spike protein, BEI Catalog No. NR-53769) 100 ng/50 uL in carbonate coating buffer (Thermo Scientific, CB01100), then were incubated the plate at 4 °C overnight. After that, the plates were washed 2 times with wash buffer (1x PBS + 0.05% Tween20) and blocked for 2 h at RT using the blocking buffer [1% bovine serum albumin (BSA) and 0.1% Tween 20 in PBS], They were then washed twice with wash buffer. We applied the 100uL samples appropriately diluted with assay buffer (0.5% BSA and 0.05% Tween 20 in PBS) and incubated for 1 h at 37 °C. The plates were washed four times, and then 100uL of diluted detection antibody (1 in 4000 diluted goat anti-mouse IgG, HRP conjugated (SouthernBiotech, 1030-05)) in assay buffer was applied. After the plates were incubated for 30 min at 37 °C, they were washed five times. Finally, we applied the 100uL of TMB for color development in the dark and added 100uL of stop solution (2 M HCL). The plate was then read at 450 nm.
Statistics and reproducibility
The results presented depict the output of independent mice (12–18 weeks old males and females, n = 3–10). Infected mice were analyzed at multiple time intervals post-infection to minimize bias and increase reproducibility. Data are shown as mean ± SEM. One-way analysis of variance (ANOVA) was used to compare values obtained from multiple groups. A two-way analysis of variance (ANOVA) was used to compare values obtained from multiple groups over time. To compare values obtained from the two groups, the unpaired Student’s t-test was performed. The comparison of survival curves are analyzed by Log-rank (Mantel-Cox) test. Non-linear regression is utilized for AD50 titer calculation. Statistical significance was taken at the P < 0.05 level. * means P < 0.05. ** means P < 0.01. *** means P < 0.001. **** means P < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Bulk RNA and Single-cell RNA sequencing data reported in this paper have been deposited to the GEO database (accession numbers: GSE274586 and GSE274686). The source data are available as [Supplementary Data 1] or available upon request to the first author or corresponding author.
References
Polatoğlu, I., Oncu-Oner, T., Dalman, I. & Ozdogan, S. COVID-19 in early 2023: Structure, replication mechanism, variants of SARS-CoV-2, diagnostic tests, and vaccine & drug development studies. MedComm (2020) 4, e228 (2023).
Carabelli, A. M. et al. SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat. Rev. Microbiol. 21, 162–177 (2023).
Wang, C. et al. COVID-19 and influenza infections mediate distinct pulmonary cellular and transcriptomic changes. Commun. Biol. 6, 1265 (2023).
Beck, D. B. & Aksentijevich, I. Susceptibility to severe COVID-19. Science 370, 404–405 (2020).
Davis, H. E., McCorkell, L., Vogel, J. M. & Topol, E. J. Long COVID: major findings, mechanisms and recommendations. Nat. Rev. Microbiol. 21, 133–146 (2023).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Akamatsu, M. A., de Castro, J. T., Takano, C. Y. & Ho, P. L. Off balance: Interferons in COVID-19 lung infections. EBioMedicine 73, 103642 (2021).
Galbraith, M. D. et al. Specialized interferon action in COVID-19. Proc. Natl Acad. Sci. USA 119, e2116730119 (2022).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 370, eabd4585 (2020).
Chauvineau-Grenier, A. et al. Autoantibodies neutralizing Type I interferons in 20% of COVID-19 deaths in a French Hospital. J. Clin. Immunol. 42, 459–470 (2022).
Reis, G. et al. Early treatment with pegylated Interferon Lambda for Covid-19. N. Engl. J. Med. 388, 518–528 (2023).
Li, D. & Wu, M. Pattern recognition receptors in health and diseases. Signal Transduct. Target. Ther. 6, 291 (2021).
Kawai, T. & Akira, S. Toll-like receptors and their crosstalk with other innate receptors in infection and immunity. Immunity 34, 637–650 (2011).
Blasius, A. L. & Beutler, B. Intracellular toll-like receptors. Immunity 32, 305–315 (2010).
Lim, K. H. & Staudt, L. M. Toll-like receptor signaling. Cold Spring Harb. Perspect. Biol. 5, a011247 (2013).
Takagi, H. et al. Plasmacytoid dendritic cells orchestrate TLR7-mediated innate and adaptive immunity for the initiation of autoimmune inflammation. Sci. Rep. 6, 24477 (2016).
Duan, T., Du, Y., Xing, C., Wang, H. Y. & Wang, R. F. Toll-like receptor signaling and its role in cell-mediated immunity. Front. Immunol. 13, 812774 (2022).
van de Veerdonk, F. L. & Netea, M. G. Rare variants increase the risk of severe COVID-19. Elife 10, e67860 (2021).
van der Made, C. I. et al. Presence of genetic variants among young men with severe COVID-19. JAMA 324, 663–673 (2020).
Fallerini, C. et al. Association of Toll-like receptor 7 variants with life-threatening COVID-19 disease in males: findings from a nested case-control study. Elife 10, e67569 (2021).
Asano, T. et al. X-linked recessive TLR7 deficiency in ~1% of men under 60 years old with life-threatening COVID-19. Sci. Immunol. 6, eabl4348 (2021).
Yin, Q. et al. A TLR7-nanoparticle adjuvant promotes a broad immune response against heterologous strains of influenza and SARS-CoV-2. Nat. Mater. 22, 380–390 (2023).
Chang, X. et al. TLR7 signaling shapes and maintains antibody diversity upon virus-like particle immunization. Front. Immunol. 12, 827256 (2021).
Miquel, C. H. et al. B cell-intrinsic TLR7 signaling is required for neutralizing antibody responses to SARS-CoV-2 and pathogen-like COVID-19 vaccines. Eur. J. Immunol. 53, e2350437 (2023).
Szeto, M. D. et al. Interferon and toll-like Receptor 7 Response in COVID-19: Implications of topical imiquimod for prophylaxis and treatment. Dermatology 237, 847–856 (2021).
van der Sluis, R. M. et al. TLR2 and TLR7 mediate distinct immunopathological and antiviral plasmacytoid dendritic cell responses to SARS‐CoV‐2 infection. EMBO J. 41, e109622 (2022).
Mantovani, S. et al. Rare variants in Toll-like receptor 7 results in functional impairment and downregulation of cytokine-mediated signaling in COVID-19 patients. Genes Immun. 23, 51–56 (2022).
Wong, L.-Y. R. et al. Eicosanoid signalling blockade protects middle-aged mice from severe COVID-19. Nature 605, 146–151 (2022).
Ellsworth, C. R. et al. Natural killer cells do not attenuate a mouse-adapted SARS-CoV-2-induced disease in Rag2(-/-) mice. Viruses 16, 611 (2024).
Jefferies, C. A. Regulating IRFs in IFN driven disease. Front. Immunol. 10, 325 (2019).
Taniguchi, T., Ogasawara, K., Takaoka, A. & Tanaka, N. IRF family of transcription factors as regulators of host defense. Annu. Rev. Immunol. 19, 623–655 (2001).
Negishi, H., Taniguchi, T. & Yanai, H. The Interferon (IFN) Class of Cytokines and the IFN Regulatory Factor (IRF) Transcription Factor Family. Cold Spring Harb. Perspect. Biol. 10, a028423 (2018).
Qing, F. & Liu, Z. Interferon regulatory factor 7 in inflammation, cancer and infection. Front. Immunol. 14, 1190841 (2023).
Ma, W., Huang, G., Wang, Z., Wang, L. & Gao, Q. IRF7: role and regulation in immunity and autoimmunity. Front. Immunol. 14, 1236923 (2023).
Zhang, Q. et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 370, eabd4570 (2020).
Israelow, B. et al. Mouse model of SARS-CoV-2 reveals inflammatory role of type I interferon signaling. J. Exp. Med. 217, e20201241 (2020).
Sokal, A. et al. Human type I IFN deficiency does not impair B cell response to SARS-CoV-2 mRNA vaccination. J. Exp. Med. 220, e20220258 (2023).
Lind, N. A., Rael, V. E., Pestal, K., Liu, B. & Barton, G. M. Regulation of the nucleic acid-sensing Toll-like receptors. Nat. Rev. Immunol. 22, 224–235 (2022).
Sun, H. et al. Targeting toll-like receptor 7/8 for immunotherapy: recent advances and prospectives. Biomark. Res. 10, 89 (2022).
McCray, P. B. Jr. et al. Lethal infection of K18-hACE2 mice infected with severe acute respiratory syndrome coronavirus. J. Virol. 81, 813–821 (2007).
Qin, Z. et al. Endothelial cell infection and dysfunction, immune activation in severe COVID-19. Theranostics 11, 8076–8091 (2021).
Bortz, R. H. et al. Single-Dilution COVID-19 Antibody test with qualitative and quantitative readouts. mSphere 6, https://doi.org/10.1128/msphere.00224-00221 (2021).
Vu Van, D. et al. Local T/B cooperation in inflamed tissues is supported by T follicular helper-like cells. Nat. Commun. 7, 10875 (2016).
Cicalese, M. P., Salek-Ardakani, S. & Fousteri, G. Editorial: Follicular helper T cells in immunity and autoimmunity. Front. Immunol. 11, 1042 (2020).
El-Hefnawy, S. M. et al. COVID-19 susceptibility, severity, clinical outcome and Toll-like receptor (7) mRNA expression driven by TLR7 gene polymorphism (rs3853839) in middle-aged individuals without previous comorbidities. Gene Rep. 27, 101612 (2022).
Solanich, X. et al. Genetic screening for TLR7 variants in young and previously healthy men with severe COVID-19. Front. Immunol. 12, 719115 (2021).
Spiering, A. E. & de Vries, T. J. Why females do better: The X Chromosomal TLR7 gene-dose effect in COVID-19. Front. Immunol. 12, 756262 (2021).
Ning, S., Pagano, J. S. & Barber, G. N. IRF7: activation, regulation, modification and function. Genes Immun. 12, 399–414 (2011).
Liang, Q., Deng, H., Sun, C. W., Townes, T. M. & Zhu, F. Negative regulation of IRF7 activation by activating transcription factor 4 suggests a cross-regulation between the IFN responses and the cellular integrated stress responses. J. Immunol. 186, 1001–1010 (2011).
Kim, T. K., Kim, T., Kim, T. Y., Lee, W. G. & Yim, J. Chemotherapeutic DNA-damaging drugs activate Interferon Regulatory Factor-7 by the Mitogen-activated Protein Kinase Kinase-4-c-Jun NH2-Terminal Kinase Pathway1. Cancer Res. 60, 1153–1156 (2000).
Crotty, S. T follicular helper cell differentiation, function, and roles in disease. Immunity 41, 529–542 (2014).
Collins, M., Ling, V. & Carreno, B. M. The B7 family of immune-regulatory ligands. Genome Biol. 6, 223 (2005).
Iwasaki, A. & Medzhitov, R. Toll-like receptor control of the adaptive immune responses. Nat. Immunol. 5, 987–995 (2004).
Feng, D. et al. Cre-inducible human CD59 mediates rapid cell ablation after intermedilysin administration. J. Clin. Invest. 126, 2321–2333 (2016).
Hu, W. et al. Rapid conditional targeted ablation of cells expressing human CD59 in transgenic mice by intermedilysin. Nat. Med. 14, 98–103 (2008).
Liu, F. et al. Versatile cell ablation tools and their applications to study loss of cell functions. Cell Mol. Life Sci. 76, 4725–4743 (2019).
Liu, F. et al. Distinct fate, dynamics and niches of renal macrophages of bone marrow or embryonic origins. Nat. Commun. 11, 2280 (2020).
Acknowledgements
This work was supported by NIH 2 P51OD011104-62 (CW, CRE, ZC, MI, NVAK, MAA, NJM, RVB, XQ), AHA962950 (XQ), R01DK129881 (XQ), R01HL165265 (XQ), R35 HL139930 (JKK), the Louisiana Board of Regents Endowed Chairs for Eminent Scholars Program, Emergent Ventures-Fast Grant, and Tulane start-up funds (XQ). We are grateful to Dr. S. Perlman at the University of Iowa for sharing the SARS-MA30 strain. The following reagents were deposited by the Centers for Disease Control and Prevention and obtained through BEI Resources, NIAID, NIH: SARS-Related Coronavirus 2, Isolate USA-WA1/2020, NR-52281, SARS-CoV-2 spike protein, BEI, NR-53769, and polyclonal anti-SARS coronavirus (antiserum, Guinea Pig), NR-10361. We thank Kejing Song and Haiyan Miller for technical assistance related to Next Generation Sequencing and Angela Birnbaum, Tammy P Bavaret, Nadia A Golden, Carli C Thompson, and Solange Paredes for technical assistance related to BSL3 experiments. We also thank confocal microscopy and molecular pathology core: RRID: SCR_024613; Anatomic Pathology Core: RRID: SCR_024606; Virus Characterization, Isolation, Production and Sequencing Core: RRID: SCR 024679; Confocal Microscopy and Molecular Pathology Core: RRID: SCR 024613; and High Containment Research Performance Core: RRID: SCR_024612; at Tulane National Primate Research Center for technical assistance related to histological analysis of the lung pathology and expanding, characterizing, and providing the SARS-CoV-2 used in these studies and conducting BSL3 experiments.
Author information
Authors and Affiliations
Contributions
J.K.K. and X.Q. developed the concept. C.W., M.S.K., C.R.E., Z.C., M.I., N.V.A.K., M.A.A., S.L., J.E.M., N.J.M., R.V.B., J.K.K., and X.Q. contributed to perform the experiments and analyzed the results. C.W., M.S.K., N.V.A.K., R.V.B., J.K.K., and X.Q. wrote the manuscript and all authors participated in the review and critique of the manuscript. J.K.K. and X.Q. interpreted the results and supervised the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Sachin Bhagchandani, Renee van der Sluis, and the other, anonymous, reviewer for their contribution to the peer review of this work. Primary Handling Editors: Si Ming Man and David Favero. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, C., Khatun, M.S., Ellsworth, C.R. et al. Deficiency of Tlr7 and Irf7 in mice increases the severity of COVID-19 through the reduced interferon production. Commun Biol 7, 1162 (2024). https://doi.org/10.1038/s42003-024-06872-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06872-5
- Springer Nature Limited