Abstract
Eyes absent (Eya) is a highly conserved transcriptional coactivator and protein phosphatase that plays vital roles in multiple developmental processes from Drosophila to humans. Eya proteins contain a PST (Proline-Serine-Threonine)-rich transactivation domain, a threonine phosphatase motif (TPM) and a tyrosine protein phosphatase domain. Using a genomic rescue system, we find that the PST domain is essential for Eya activity and Dac expression and the TPM is required for full Eya function. We also find that the threonine phosphatase activity plays only a minor role during Drosophila eye development and the primary function of the PST and TPM domains is transactivation that can be largely substituted by the heterologous activation domain VP16. Along with our previous results that the tyrosine phosphatase activity of Eya is dispensable for normal Eya function in eye formation, we demonstrate that a primary function of Eya during Drosophila eye development is as a transcriptional coactivator. Moreover, the PST/TPM and the threonine phosphatase activity are not required for in vitro interaction between retinal determination factors. Finally, this work is the first report of an Eya-Ey physical interaction. These findings are particularly important because they highlight the need for an in vivo approach that accurately dissects protein function.
Similar content being viewed by others
Introduction
Drosophila eye development depends on a network of retinal determination (RD) genes, which encode conserved nuclear proteins that play critical roles in Drosophila eye development1. The core RD genes include twin of eyeless (toy)2, eyeless (ey)3, eyes absent (eya)4, sine oculis (so)5 and dachshund (dac)6. These genes are involved in interconnected feedback loops and their protein products are necessary and sufficient for inducing retinal fate. ey and toy, which encode paired-type homeobox genes, lie atop the genetic hierarchy controlling eye development. Ey activates the expression of eya and so, which in turn induce dac7. Eya and So expression in the eye begins during the second instar larval stage and is highest near the posterior margin. After morphogenetic furrow (MF) initiation, Eya and So are co-expressed within and posterior to the MF, as well as in a zone immediately anterior to it4,5. Once established, So maintains its own expression, as well as that of eya, dac and ey. So associates with Eya and it is thought that Eya act as a transcriptional coactivator upon recruitment by So, since Eya has no recognized DNA binding activity, but possesses a transactivation domain8,9,10,11. Eya can also physically interact with Dac to regulate target genes during eye development7,11,12,13,14,15. A similar interaction has been reported between their mouse counterparts EYA2 and DACH216.
As a key member of the RD gene network, Eya acts as a transcriptional coactivator and also contains both tyrosine and threonine phosphatase activities17,18,19. Eya regulates multiple developmental processes throughout the metazoans15. In the Drosophila eye, loss of eya function blocks MF initiation, causes massive apoptosis in eye discs and the complete failure of eye development. This cell death phenotype resembles those seen in the ear and kidney primordia of Eya1 mutant mouse embryos20. In contrast to these loss-of-function phenotypes, ectopic overexpression of eya in other imaginal discs is sufficient to cause the formation of ectopic eyes21.
Drosophila Eya contains a highly conserved C-terminal domain called the Eya Domain (ED) and a moderately conserved threonine phosphatase motif (TPM) embedded in a proline-serine-threonine-rich (PST) domain. Throughout the remainder of this paper, “PST/TPM”, “PST” and “TPM” represent the PST domain with the TPM, the PST domain alone and the TPM alone, respectively. Eya and So bind to each other through the ED of Eya and the Six domain of So8,11 to form a transcriptional activator complex. In addition, a series of Drosophila S2 cell-based transcriptional activation assays defined the PST/TPM domain as essential for Eya/So-mediated transactivation of a reporter. UAS-eya transgenes that lack both the PST-rich region and the TPM have drastically reduced ectopic eye-inducing capacity, with induction efficiency dropping from 98% to 1.5%10.
In addition to regulating transcription, Eya has predicted tyrosine and threonine phosphatase activities in the ED and TPM, respectively17,18,19,22,23,24. In Drosophila, tyrosine phosphatase-dead mutations lead to strongly reduced activities in ectopic eye induction and in vivo genetic rescue using the GAL4-UAS system18,19,24. In contrast to these studies, our previous findings revealed that eya genomic rescue (GR) constructs carrying mutations in two key tyrosine phosphatase active-site residues fully restore viability as well as eye formation and function in an eya null mutant background25. In mouse and Drosophila, the threonine phosphatase activity has been suggested to play an important role in the innate immune system17 and a recent study using the GAL4-UAS system reported that Eya threonine phosphatase activity is not required for normal Drosophila eye development24.
Although previous cell culture and in vivo GAL4-UAS based expression studies have suggested specific functions for conserved Eya domains, we have shown that such assays may not always be reliable. In particular, we have developed a genomic rescue (GR) system that provides an accurate method for assessing the functional significance of individual protein domains in vivo25,26. In this study, we have used the GR strategy to conduct functional studies of Eya domains during Drosophila eye development. Interestingly, we found that a major function of Eya is transcriptional coactivation, while the threonine phosphates activity plays only a minor role during Drosophila development.
Results
The threonine phosphatase activity of Eya plays only a minor role in normal Drosophila development
To study eya function in vivo, we introduced a series of eya genomic rescue constructs (eyaGR) via site-specific transgenesis27,28 to investigate the transcriptional activation and threonine phosphatase activity of Eya. A wild-type eya genomic rescue construct (eya+GR) is known to fully rescue viability and eye formation in an eya null mutant background, therefore serving as a positive control throughout our studies25,26,29. The eyaY4GR construct has tyrosine-to-alanine substitutions for four key tyrosine residues known to be required for threonine phosphatase activity17,24 (Fig. 1a). The eyaΔTPMGR construct has the entire TPM deleted but leaves the PST domain intact. Surprisingly, a single copy of each construct is able to substantially rescue eya2 or eyacliIID mutant phenotypes, restoring viability and rescuing eye size to ~90% (Y4) or ~60% (ΔTPM) of wild-type, albeit with some mild disorganization (Fig. 1d,e and Fig. S1). While there appears to be a largely normal complement and arrangement of rhabdomeres in ommatidia of eya2; eyaY4GR/+ flies (Fig. 2e), eye discs from late third instar larvae (Fig. 2h) and 24 hrs after puparium formation (Fig. 2j) show defects in the number of cone cells and/or ommatidial fusion. Larval eye discs from eyaY4GR and eyaΔTPMGR rescued animals are smaller and show a reduction in Eya and So staining anterior to and within the MF while expression levels are normal posteriorly (Fig. 2b’”), suggesting that the threonine phosphatase activity does play a role during Drosophila eye development but this role is relatively minor as the eyaY4GR construct can restore up to 90% of the eye size. The expression of the core RD genes Dachshund (Dac) and Eyeless (Ey) appear similar in eye discs of positive control and eyaY4GR-rescued larvae (Figs 2a–c and 3a,b). In addition, we found no difference in photoreceptor axon projections between wild-type and eya2; eyaY4GR/+ flies (Fig. S2c), which show a regular pattern of projections in the lamina of the optic lobe.
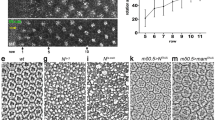
eya genomic rescue constructs rigorously define Eya functional domains.
(a) Schematic view of the eya genomic rescue (GR) constructs assayed in this study. The percent rescue indicated is for animals with one copy of each GR construct in an eya2 homozygous mutant background. At least 100 adult eyes were scored for each GR and the penetrance of the transgene-induced phenotypes for all GRs are 100%, with minor variation in expressivity. Each construct is inserted at the same genomic docking site such that all constructs are directly comparable. (b–k) Representative images of adult eyes for each genotype tested are shown. PST: grey box, TPM: purple box, Y4: red vertical lines, VP16: green box, ED1: orange box.
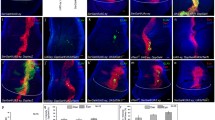
Threonine phosphatase activity is required for normal anterior expression of eya and cone cell development.
Expression of Ey (a–c), Eya (a’–c’) and So (a”–c”) proteins are shown in eya+GR, eyaY4GR and eyaΔTPMGR rescued animals. (d–f) Adult plastic sections in flies rescued with one copy of eya+GR, eyaY4GR and eyaΔTPMGR, respectively. (g,h) Third instar eye discs by Cut staining. (I,j) Eye discs prepared from 48 hrs after puparium formation and stained with Dlg.
Loss of threonine phosphatase activity does not strongly affect Dac, Ato, or Cyclin B expression.
Dac (a,b), Ato (c,d) and the cell cycle marker Cyclin B (e,f) expression in animals rescued with a single copy of eya+GR or eyaY4GR are shown. Expression patterns and levels of Eya, Ato and Cyclin B are similar in both genotypes although minor disruption of Ato is observed.
eya plays an important role in the developmental events associated with morphogenetic furrow movement. Specifically, clonal analysis has shown that eya is required for the initiation and propagation of the MF and for regulation of the cell cycle4,8,30,31. Since loss of threonine phosphatase activity leads to a reduction of Eya expression anterior to and within the MF, we analyzed the effects of eyaY4 on both G1-arrest and induction of the proneural gene atonal (ato). We used the cell cycle marker Cyclin B to monitor G1 arrest. Normally, Cyclin B is exclusively expressed in cells in the G2 and M phases32. Immunohistochemistry shows eyaY4GR rescued animals have largely normal Cyclin B and Ato expression patterns (Fig. 3c–f), implying that the threonine phosphatase-inactive mutations do not adversely affect G1 arrest and initiation of retinal differentiation. This is not surprising since the loss of retinal cells in flies rescued with a single copy of eyaY4GR is relatively mild; therefore, strong alterations in the expression of markers of cell cycle progression or photoreceptor differentiation are not expected.
The Eya threonine phosphatase-inactive mutation does not abolish interaction of Eya with Ey, So, or Dac
Phosphorylation is well known in other systems to regulate protein complex formation and protein stability via ubiquitin-mediated degradation33,34. Accordingly, we hypothesized that one or more of the RD proteins are direct substrates for Eya threonine phosphatase and that loss of this activity either disrupts the formation of RD protein complexes and/or destabilizes the RD proteins themselves. Furthermore, this effect may be specific to complexes involving Eyeless (Ey), thereby limiting effects anterior to the MF where Ey is expressed. We tested this hypothesis by doing co-immunoprecipitation (co-IP) in S2 cultured cells transiently transfected with epitope-tagged Eya, Ey, So and Dac expression constructs. Similar amounts of RD proteins are expressed in transfected cells with or without Eya threonine phosphatase activity and the Y4 mutation or the TPM deletion do not affect Eya protein expression levels in S2 cells (data not shown). As shown in Fig. 4, EyaY4 and EyaΔTPM co-IP with Ey, So and Dac without obviously altered efficiency as wild-type Eya. Notably, this is the first report that Eya can bind to Ey. Previous studies also found that both Eya and Ey proteins interact with So8,35, suggesting Ey, Eya and So may form a complex to mediate Drosophila eye development. Taken together, these observations suggest that the threonine phosphatase activity of Eya is not essential for interactions with other RD proteins.
Threonine phosphatase activity is not required for Eya interaction with Ey, So, or Dac.
(a,b) Co-immunoprecipitation (co-IP) studies between wild-type and threonine phosphatase-dead Eya (EyaY4 and EyaΔTPM) and Ey or So are shown. Flag-tagged Eya was co-expressed with HA-Ey or Myc-So in S2 cells and co-IP with anti-Flag beads followed by western blotting (WB) was performed. Ey, So and Eya (wild-type and mutants) were detected by anti-HA, anti-Myc and anti-Flag antibodies, respectively. Lanes 1, 2 and 3 show that EyaWT, EyaY4 and EyaΔTPM can pull down So and Ey, respectively. Empty vector is the negative control (Lane 4). (c) co-IP analysis of Eya and Dac. Lane 1, Flag-EyaWT/HA-Dac; lane 2, Flag-EyaY4/HA-Dac; lane 3, Flag-EyaΔTPM/HA-Dac; lane 4 Flag-EyaΔPST/TPM/HA-Dac; lane 5, empty vector. Anti-FLAG is used for IP. The red arrow indicates Dac protein. All proteins are expressed at similar levels in crude cell lysates (the bottom panel of each set and data not shown). Western blots presented in a-c were cropped to improve clarity and full-length blots are presented in Supplementary Figs S7–9. All gels were run under the same experimental conditions.
The threonine phosphatase motif of Eya has transcriptional activation function
In addition to threonine phosphatase activity, previous cell culture transactivation reporter assays showed that the TPM has transcriptional activation function10. To test the hypothesis that this function is biologically relevant in vivo, we replaced the TPM only with VP16, a well-known heterologous transcriptional activation domain (Chasman et al., 1989). The resulting construct, eyaΔTPM+VP16GR, was tested for rescue activity. Remarkably, while eyaΔTPM can restore about 60% of eye size, VP16 is able to largely complement loss of the TPM and restore eye development to approximately 90% of wild-type, both in eya2 (Fig. 1a,e,f) and eyacliIID mutant backgrounds (Fig. S3a). The external eye morphology of eyaΔTPM+VP16GR rescued eyes shows only minor disorganization compared to eyaΔTPMGR.
As shown in Fig. 2f, loss of the TPM causes abnormal ommatidial morphology in adult compound eyes. Flies rescued by one copy of eyaΔTPMGR have a reduced number and unusual arrangement of rhabdomeres compared with the normal trapezoidal array of photoreceptors in wild-type animals. Tangential sections of eya2; eyaΔTPM+VP16GR/+ adult eyes reveal ommatidia with the correct number and largely normal arrangement of rhabdomeres (Fig. S3b). Moreover, in contrast to wild-type (Fig. S2a) and eya2; eyaΔTPM+VP16GR/+ (Fig. S2e) flies, axon terminations in the lamina plexus have irregular gaps and breaks (yellow arrows) in eyaΔTPMGR rescued flies, reminiscent of the photoreceptor axon defects in eya loss-of-function mutants36. These observations suggest that a major role of the TPM during Drosophila eye development is to provide transactivation function, that this activity is required for normal ommatidial development and photoreceptor axon projections and that this function can be largely substituted by the VP16 domain.
The entire PST/TPM domain of Eya is critical for transcriptional activation during eye development
The PST/TPM domain of Eya is critical for transactivation in cell culture reporter assays10. In order to characterize the Drosophila Eya transcriptional activity in its native context in vivo, we generated four genomic rescue constructs: eyaΔPST/TPMGR (deletion of the PST/TPM domain), eyaΔPSTGR (deletion of the PST domain alone), eyaΔPST/TPM +VP16GR (substitution of both the PST and TPM domains with the VP16 activation domain) and eyaΔPST+VP16GR (substitution of the PST domain alone with the VP16 activation domain) (Fig. 1a). We found that eyaΔPST/TPMGR completely fails to rescue eya2 or eyacliIID mutant phenotypes, even when the transgene is present in two copies (Fig. 1h and data not shown). We can readily detect the predicted, truncated eyaΔPST/TPM transcript and protein (Fig. 5a–f) in late second instar eye discs prior to MF initiation, suggesting that although the transgene is expressed, at least initially, the EyaΔPST/TPM protein is non-functional. While the eyaΔPST/TPMGR construct completely fails to rescue eya2 mutant animals, the eyaΔPSTGR retains slightly more function and can rescue about 5% of normal eye size (Fig. 1j). Previous S2 cell culture studies have suggested that both the PST and TPM domains contribute transcription activation function10 and our GR data are consistent with these results. In addition to eyaΔTPM+VP16GR, our other VP16 substitution genomic rescue results also confirm these findings. Specifically, the eyaΔPST/TPM +VP16GR is sufficient to rescue about 5% of eye size in an eya2 background (Fig. 1i), similar to that of the eyaΔPSTGR construct alone. eyaΔPST+VP16GR is able to restore eye development to ~30% of wild-type (Fig. 1k). Two copies of eyaΔPST/TPM +VP16GR or eyaΔPST+VP16GR consistently rescue eya2 eye size better than one copy (Fig. S4). Moreover, eyaΔPST/TPM +VP16GR, eyaΔPSTGR and eyaΔPST+VP16GR fail to rescue eyacliIID mutants. These functional dissection studies reveal that the transactivation domain PST/TPM is essential for eye formation and viability in Drosophila. In addition, the PST domain is likely playing a more significant role than the TPM during Drosophila development since eyaΔTPMGR rescues 60% of the eye size compared to 5% of the eye size rescued by eyaΔPSTGR and eyaΔTPMGR is able to restores viability to eyacliIID null mutants.
eyaΔPST/TPMGR fails to rescue eya mutant defects even though mutant transcript and protein are detected.
(a) Lanes 1–3 show RT-PCR on RNA prepared from eyes discs 68 hrs after egg laying (AEL). Lane 1: wild-type; Lane 2: eya2; eyaΔPST/TPMGR; Lane 3: eya2; Lane 4: water. A truncated ΔPST/TPM transcript is readily detected (Lane 2). (b) An anti-Eya Western blot on extracts prepared from 68 hrs after egg laying (AEL) eye discs (n = 40/lane) from either eyacliIID/CyO; eya+GR/+ or eyacliIID/CyO; eyaΔPST/TPMGR/+ animals shows a readily detectable, truncated ΔPST/TPM protein (*). Heterozygous eyacliIID animals were used to obtain enough tissue for the experiment. Western blot presented in b is cropped to improve clarity and full-length blot is presented in Supplementary Fig. S10. (c–f) Eye discs prepared from larvae 68 hrs AEL are stained for Eya expression. (g–j) Eya staining of 56 hrs AEL eye discs.
The PST/TPM domain regulates retinal determination gene expression
Eya can act as a transcriptional coactivator and physically interact with other RD proteins to regulate multiple developmental processes7,8,9,10,37. Therefore, we were interested in understanding the role of PST/TPM in RD gene regulation since it is critical for Eya function. Since eyaΔPST/TPMGR fails to rescue the eye phenotype of eya2 animals and little Eya expression is detected at late third instar (data not shown), we used second instar larvae to assess the function of the PST/TPM when EyaΔPST/TPM protein is still expressed (Fig. 5f). eya2 flies rescued with two copies of eyaΔPST/TPMGR show slightly lower Eya expression compared to wild-type or eya GR-rescued animals at 68 hrs after egg laying (AEL) (Fig. 5c–f). We also found that Eya expression in eya2; eyaΔPST/TPMGR eye discs is lower than that of wild-type discs at 56 hrs AEL (Fig. 5g–j). Similar reductions are observed for the expression of the retinal determination protein Dac, a known downstream target of Eya (Fig. 6a–h). In addition, in eyaΔPST/TPM clones (eyacliIID null clones rescued by a single copy of eyaΔPST/TPMGR) at 72 hrs AEL, Dac expression is reduced while the expression of EyaΔPST/TPM is normal (Fig. 6i–l, yellow arrows). Taken together, these data imply that the PST/TPM domain of Eya is required for normal Dac expression.
PST/TPM is required for normal Dac expression.
(a–d) Dac expression in Canton-S (a), eya2 (b), eya2; eya+GR (c) and eya2; eyaΔPST/TPMGR (d) eye imaginal discs from 68 hrs AEL. (e–h) Immunostaining of Dac on 56 hrs AEL eye discs. (i–k) Dac, Eya and GFP expression in eyaΔPST/TPM rescued eyacliIID null clones. Yellow arrow indicates one of the larger, more posterior clones in which Dac expression is reduced. (l) Merge of channels.
Moreover, ey-Gal4 induced So expression in eya2 animals rescued by one copy of eyaΔPST/TPMGR partially restores Dac expression (Fig. 7a–d), but has no effect on expression of Eya (Fig. 7e–h). These observations suggest that the PST/TPM positively regulates expression of Dac through the Eya binding partner So.
Overexpression of So in eya2; eyaΔPST/TPMGR animals causes increased Dac expression.
(a–d) Dac staining in third instar larvae after inducing so expression with ey-Gal4/UAS-so. Yellow arrows (d) indicate region of the disc in which Dac expression is increased. (e–h) Eya expression after ey-Gal4/UAS-so induction. eya2; eya+GR (a,e) and eya2 (b,f) are used as positive and negative controls.
To test if the PST/TPM deletion affects Ey regulation and photoreceptor differentiation, we assayed Ey and Elav expression in eyaΔPST/TPM rescued eya null mutant clones. We found that eyaΔPST/TPM clones show a complete loss of Elav expression, a marker of photoreceptor differentiation38, posterior to the MF (Fig. 8a–d). In eyaΔPST/TPM clones posterior to the furrow, we found strong Ey expression (Fig. 8a’–d’), suggesting the PST/TPM domain of Eya is required for Ey suppression. Additionally, eyaΔPST/TPM clones result in the loss of photoreceptor development and black overgrowths in adults (Fig. S5d).
The PST/TPM of Eya is necessary for ey repression posterior to the morphogenetic furrow.
(a–a’”) Wild-type clones. (b–b’”) eyacliIID null clones. (c–c’”) eya+GR rescued eyacliIID null clones. (d–d’”) eyaΔPST/TPMGR rescued eyacliIID null clones. Grayscale images of Elav, Ey and GFP expression are shown in grayscale (a–d”) and as red, blue and green, respectively, in a’”–d’”; Elav marks differentiating photoreceptors and complete loss of GFP expression marks homozygous mutant clones.
Deletion of the PST/TPM does not abolish interactions between Eya and Ey, So, or Dac
So and Dac are known binding partners of Eya7,8,11. Since eyaΔPST/TPMGR rescued flies have no eyes, similar to the loss-of-function phenotypes of the core RD genes (ey, so and dac), we hypothesized that the PST/TPM domain may mediate specific, essential interactions between Eya and Ey, So, or Dac. To test this hypothesis, we carried out co-immunoprecipitation (co-IP) experiments. As shown in Figs 4 and 9, both wild-type and EyaΔPST/TPM can co-IP with Ey, So and Dac, suggesting that deletion of the PST/TPM does not abolish the interactions between Eya and these three RD proteins. These observations are consistent with previous findings that Eya-So and Eya-Dac interaction is mediated via the ED of Eya7,8,11. The Eya domain that mediates Eya-Ey physical interaction remains to be determined.
The PST/TPM domain is not required for interaction between Eya and Ey, So, or Dac.
(a,b) S2 cells were transfected as described in Materials and Methods, lysates were immunoprecipitated (IP) with anti-Flag beads and then immunoblotted (WB) with anti-Myc, anti-HA and anti-Flag antibodies to detect Myc-So, HA-Ey and Flag-Eya, respectively. Co-IP for EyaΔPST/TPM and Dac is shown in Fig. 4c. Western blots presented in a-b were cropped to improve clarity and full-length blots are presented in Supplementary Fig. S11–12. All gels were run under the same experimental conditions.
Discussion
In this paper we report that loss of threonine phosphatase activity has little effect on Drosophila eye development, since eye development in eyaY4GR rescued flies proceeds relatively normally. On the other hand, the essential function of the PST and the threonine phosphatase motif (TPM) is transcriptional activation that can be largely complemented by the heterologous activation domain VP16. Together with our findings that the PST and TPM are required for normal Drosophila eye development, we conclude that a major function of Eya during Drosophila eye development is as a transcriptional coactivator. Although the tyrosine phosphatase activity of the Eya Domain (ED) is dispensable for Eya function25, the specific role the ED plays in vivo has not been reported.
The retinal determination (RD) network is a small group of highly conserved transcriptional regulators that are both necessary for eye development and sufficient to trigger ectopic eye formation when overexpressed in other imaginal discs1,2,3,4,5,6,7,8,14,21,39. As a vital member of the RD network, a unique feature of the Eya proteins is that they have several distinct biochemical activities. In Drosophila, previous cell culture reporter assays and cDNA-based Gal4-UAS genetic rescue studies suggested that the PST-rich region is a transactivation domain and plays a role in ectopic eye induction, while the TPM and ED possess threonine and tyrosine phosphatase activity, respectively10,18,19,24. Intriguingly, our results using genomic rescue constructs are consistent with previous studies of the PST/TPM transactivation domain, but are contrary to previous reports that the tyrosine phosphatase domain, but not the threonine phosphatase domain, governs Drosophila eye development.
In our work, we have found that both the TPM and PST contribute transcriptional activation for normal eye development. Substituting the heterologous activation domain VP16 for the TPM and PST domain substantially restores Eya function. Two reasons could account for the failure of complete rescue by VP16. First, the TPM or PST have other, distinct functions. Although we have excluded the possibility that the TPM and PST are required for Eya binding with Ey, So, or Dac in this report, we cannot rule out other possibilities. For example, previous findings identified the PST/TPM domain of Eya as the primary target of Nmo and Abl-mediated phosphorylation in kinase assays36,40. Second, there may be insufficient activation function provided by VP16 - perhaps due to an inability to make specific contacts with other proteins, or that the fusion proteins do not have the proper conformation to interact properly via other domains.
The transcriptional role of Eya has been studied in Drosophila through genetic and/or biochemical interaction with the transcription factors So and Dac7,8. In this paper, we further indicate that the PST/TPM domain positively regulates Dac expression and this regulation may be mediated via So. Moreover, the PST/TPM is required to suppress Ey expression posterior to the furrow. These observations are consistent with previous reports that dac expression requires both so and eya7,14,39,41 and both Eya and So are necessary to mediate Ey repression posterior to the MF42. Our studies localize these functions of Eya to the PST/TPM domain.
Although genetic interactions between Eya and Ey have been widely reported, physical interactions between these two RD proteins have not. In this paper, we report that Eya physically interacts with Ey for the first time. Previous studies also found physical interactions between Eya-So8 and Ey-So35, suggesting that Ey-Eya-So may form a ternary complex. In addition, previous findings show that ectopic eye induction by Ey requires the presence of Eya and So43 and the expression patterns of all three genes overlap extensively and are nearly identical anterior to the MF43. Moreover, misexpression of Eya and So induces the formation of ectopic eyes; however, this effect is lost in an ey mutant background8,21. Finally, ey is a direct target of Eya and So11,44 and vice versa - eya and so are direct targets of Ey45,46. Since Groucho is a repressor of the Eya-So complex10, Ey may act as an activator of Eya-So to increase transcriptional output of Dac. Consistent with this hypothesis, loss of ey, eya, or so function causes loss of Dac expression, suggesting that Ey, So and Eya are primary regulators of Dac7,8,47. Similar relationships have been observed with Pax6, Eya1/2 and Six3, mouse orthologs of ey, eya and so, respectively. Specifically, mouse Pax6 mutants have reduced levels of Eya1 and eya2 in the optic vesicle and overlying ectoderm48,49 and Pax6 induces expression of Six3 when ectopically expressed in mice50. In addition, we used STRING51, a database of known and predicted protein interactions, to predict protein-protein interactions for Ey, Eya and So. As expected, we found equally high associations for all three pairs of complexes (Fig. S6), providing further evidence of strong interactions among these RD proteins, which may act together in a ternary complex.
In addition, our genomic rescue assays show that the threonine phosphatase activity is largely but not entirely dispensable for Drosophila eye development. Our threonine-phosphatase inactive GRs can robustly rescue eye formation in eya null mutants, but the rescued eyes show disorganized external and internal morphology as compared to wild-type rescue controls. This result is in contrast to another report based on the GAL4-UAS system that finds the threonine phosphatase activity of Eya to be dispensable during eye development24. The reason for this difference is that our GR system offers higher resolution thereby allowing detection of more subtle defects in morphology, while the GAL4-UAS system is a less accurate approach. In particular, Liu et al. did in fact observe a disorganized eye phenotype in eya2 flies rescued by UAS-eyaY4. However, this phenotype appeared similar to the imperfect rescue achieved with the wild-type UAS-eya transgene. For this reason, they could not uncover the requirement for the threonine phosphatase activity during differentiation. This report highlights the need for careful interpretation of results based on the GAL4-UAS system and the superior sensitivity of the GR method. Although the threonine phosphatase activity of Eya plays only a minor role during eye development, it has been reported to be involved in the innate immune response in both Drosophila and mouse17,24.
In summary, we have shown that both the transcriptional activation and threonine phosphatase activity of Eya are required for normal Drosophila eye development. However, a primary function of Eya during this process is transcriptional coactivation, while the phosphatase activity plays only a minor role. Our study provides an accurate approach to assess the functional significance of individual protein domains in vivo, highlighting the importance of the transactivation function of Eya during Drosophila development. As Eya is conserved and plays important roles in retinal development throughout the metazoa, the underlying mechanisms of Eya function are likely to be conserved in vertebrates as well.
Methods
Fly strains and maintenance
All flies were maintained with standard corn meal and yeast extract medium at 25 °C. Canton-S was used as a wild-type control. Heat shocks were performed at 37 °C as described previously52. To test the function of the mutant eyaGR during eye development, we crossed transgenes into the following mutant backgrounds: eya2, which completely lack eyes due to a deletion of an enhancer required for eya expression during eye development4 and eyacliIID, which is a null allele caused by a premature stop codon that causes recessive embryonic lethality53. Wild-type clones and eyaΔPST/TPM clones were generated by crossing w/Y; FRT40A and w/Y; eyaclillD FRT40A/CyO; eyaΔPST/TPMGR with ywhs-flp; w+ubiGFP, FRT40A animals, respectively.
Recombineering-induced mutagenesis of eya+GR and Drosophila transgenesis
A two-step recombineering method was used to create the Y4, ΔTPM, ΔTPM+VP16, ΔPST/TPM, ΔPST/TPM+VP16, ΔPST and ΔPST+VP16 mutations in the eya+GR construct as described previously54. Recombineering products were verified by DNA sequencing and restriction enzyme fingerprint digestion prior to transgenesis. Constructs were inserted into the attP2 docking site on the third chromosome using PhiC31-mediated transgenesis and site-specific integration was confirmed by genomic PCR with attP/attB primers28. Transgenic flies were confirmed by genomic DNA PCR sequencing. Primer sequences are available on request.
Construction of cell culture expression plasmids
We used the Q5 Site-Directed Mutagenesis Kit (NEB) to introduce a series of mutations in cell culture expression plasmids which were confirmed by DNA sequencing. These mutations include: pMT-Flag-EyaY4, pMT-Flag-EyaΔTPM, pMT-Flag-EyaΔPST/TPM and pMT-HA-Dac. pAHW-Ey was generated from destination vector pAHW and pUAST-Ey (a gift from Dr. Rui Chen, Houston, TX) according to the Gateway protocol provided by the Drosophila Genomics Resource Center. pMT-Flag-Eya, pMT-Myc-So, pMT-dac and pAHW were kindly provided by Dr. Ilaria Rebay (Chicago, IL). Primers used in this report are listed in Table S1.
S2 cell culture and transfection
Drosophila S2 cells were cultured in Schneider’s medium containing 10% heat-inactivated fetal bovine serum and antibiotics at 25 °C. Cells were transiently transfected in 6-well plates using the FuGENE HD Transfection Reagent (Promega) according to the manufacturer’s protocol. 24 hrs after transfection, cells were induced by addition of 0.1 M CuSO4.
Co-IP and western blots
Transfected cells were lysed by rocking at 4 °C for 30 min in Pierce IP lysis buffer (Thermo Fisher Scientific) with a Roche Complete, Mini, EDTA-free protease inhibitor cocktail tablet. The lysates were subjected to immunoprecipitation with anti-Flag-conjugated agarose beads (Sigma) for 2 h at 4 °C. After washing three times with lysis buffer, immunoprecipitates were boiled in 4× NuPAGE LDS sample buffer (Novex) and western blotting was carried out according to the NuPAGE electrophoresis (Novex) protocol with rabbit anti-Flag (1:1000, Sigma), rabbit anti-MYC (1:100, Santa Cruz Biotechnology) and rabbit anti-HA (1:200, Santa Cruz Biotechnology) antibodies.
For tissue preparation, 68 hrs AEL eye discs (n = 40) were collected in cold RIPA lysis buffer (Thermo Fisher Scientific). After centrifuge at 20000 g for 10 min at 4 °C, the supernatant was transferred to a new tube and ready for western blot analysis.
Histology and immunohistochemistry
Staining of eye discs and imaging of the adult eye were conducted as described previously42. Immunohistochemistry on 48 hr pupal eye discs and tangential sections of adult eyes were generated as previously described55. For antibodies used, please reference Table S2.
RT-PCR
RNA was extracted from 56 hrs AEL eye discs using PureLink RNA Mini Kit (Ambion). Reverse transcription was performed according to the instructions of SuperScript One-Step RT-PCR kit (Invitrogen).
Additional Information
How to cite this article: Jin, M. and Mardon, G. Distinct Biochemical Activities of Eyes absent During Drosophila Eye Development. Sci. Rep. 6, 23228; doi: 10.1038/srep23228 (2016).
References
Pappu, K. S. & Mardon, G. Genetic control of retinal specification and determination in Drosophila. Int J Dev Biol 48, 913–924 (2004).
Czerny, T. et al. Twin of eyeless, a second Pax-6 gene of Drosophila, acts upstream of eyeless in the control of eye development. Mol Cell 3, 297–307 (1999).
Quiring, R., Walldorf, U., Kloter, U. & Gehring, W. J. Homology of the eyeless gene of Drosophila to the Small eye gene in mice and Aniridia in humans. Science 265, 785–789 (1994).
Bonini, N. M., Leiserson, W. M. & Benzer, S. The eyes absent gene: genetic control of cell survival and differentiation in the developing Drosophila eye. Cell 72, 379–395 (1993).
Cheyette, B. N. et al. The Drosophila sine oculis locus encodes a homeodomain-containing protein required for the development of the entire visual system. Neuron 12, 977–996 (1994).
Mardon, G., Solomon, N. M. & Rubin, G. M. dachshund encodes a nuclear protein required for normal eye and leg development in Drosophila. Development 120, 3473–3486 (1994).
Chen, R., Amoui, M., Zhang, Z. & Mardon, G. Dachshund and eyes absent proteins form a complex and function synergistically to induce ectopic eye development in Drosophila. Cell 91, 893–903 (1997).
Pignoni, F. et al. The eye-specification proteins So and Eya form a complex and regulate multiple steps in Drosophila eye development. Cell 91, 881–891 (1997).
Ohto, H. et al. Cooperation of six and eya in activation of their target genes through nuclear translocation of Eya. Mol Cell Biol 19, 6815–6824 (1999).
Silver, S. J., Davies, E. L., Doyon, L. & Rebay, I. Functional dissection of eyes absent reveals new modes of regulation within the retinal determination gene network. Mol Cell Biol 23, 5989–5999 (2003).
Bui, Q. T., Zimmerman, J. E., Liu, H. & Bonini, N. M. Molecular analysis of Drosophila eyes absent mutants reveals features of the conserved Eya domain. Genetics 155, 709–720 (2000).
Jemc, J. & Rebay, I. Identification of transcriptional targets of the dual-function transcription factor/phosphatase eyes absent. Dev Biol 310, 416–429 (2007).
Suzuki, T. & Saigo, K. Transcriptional regulation of atonal required for Drosophila larval eye development by concerted action of eyes absent, sine oculis and hedgehog signaling independent of fused kinase and cubitus interruptus. Development 127, 1531–1540 (2000).
Shen, W. & Mardon, G. Ectopic eye development in Drosophila induced by directed dachshund expression. Development 124, 45–52 (1997).
Tadjuidje, E. & Hegde, R. S. The Eyes Absent proteins in development and disease. Cell Mol Life Sci 70, 1897–1931 (2013).
Heanue, T. A. et al. Synergistic regulation of vertebrate muscle development by Dach2, Eya2 and Six1, homologs of genes required for Drosophila eye formation. Genes Dev 13, 3231–3243 (1999).
Okabe, Y., Sano, T. & Nagata, S. Regulation of the innate immune response by threonine-phosphatase of Eyes absent. Nature 460, 520–524 (2009).
Rayapureddi, J. P. et al. Eyes absent represents a class of protein tyrosine phosphatases. Nature 426, 295–298 (2003).
Tootle, T. L. et al. The transcription factor Eyes absent is a protein tyrosine phosphatase. Nature 426, 299–302 (2003).
Xu, P. X. et al. Eya1-deficient mice lack ears and kidneys and show abnormal apoptosis of organ primordia. Nat Genet 23, 113–117 (1999).
Bonini, N. M., Bui, Q. T., Gray-Board, G. L. & Warrick, J. M. The Drosophila eyes absent gene directs ectopic eye formation in a pathway conserved between flies and vertebrates. Development 124, 4819–4826 (1997).
Li, X. et al. Eya protein phosphatase activity regulates Six1-Dach-Eya transcriptional effects in mammalian organogenesis. Nature 426, 247–254 (2003).
Sano, T. & Nagata, S. Characterization of the threonine-phosphatase of mouse eyes absent 3. FEBS Lett 585, 2714–2719 (2011).
Liu, X. et al. Drosophila EYA Regulates the Immune Response against DNA through an Evolutionarily Conserved Threonine Phosphatase Motif. PloS one 7, e42725 (2012).
Jin, M., Jusiak, B., Bai, Z. & Mardon, G. Eyes absent tyrosine phosphatase activity is not required for Drosophila development or survival. PloS one 8, e58818 (2013).
Jusiak, B., Abulimiti, A., Haelterman, N., Chen, R. & Mardon, G. MAPK target sites of eyes absent are not required for eye development or survival in Drosophila. PloS one 7, e50776 (2012).
Groth, A. C., Fish, M., Nusse, R. & Calos, M. P. Construction of transgenic Drosophila by using the site-specific integrase from phage phiC31. Genetics 166, 1775–1782 (2004).
Venken, K. J., He, Y., Hoskins, R. A. & Bellen, H. J. P. [acman]: a BAC transgenic platform for targeted insertion of large DNA fragments in D. melanogaster. Science 314, 1747–1751 (2006).
Karandikar, U. C. et al. Drosophila eyes absent is required for normal cone and pigment cell development. PloS one 9, e102143 (2014).
Dominguez, M. & de Celis, J. F. A dorsal/ventral boundary established by Notch controls growth and polarity in the Drosophila eye. Nature 396, 276–278 (1998).
Dominguez, M. Dual role for Hedgehog in the regulation of the proneural gene atonal during ommatidia development. Development 126, 2345–2353 (1999).
Lopes, C. S. & Casares, F. hth maintains the pool of eye progenitors and its downregulation by Dpp and Hh couples retinal fate acquisition with cell cycle exit. Dev Biol 339, 78–88 (2010).
Smelkinson, M. G., Zhou, Q. & Kalderon, D. Regulation of Ci-SCFSlimb binding, Ci proteolysis and hedgehog pathway activity by Ci phosphorylation. Dev Cell 13, 481–495 (2007).
Eblen, S. T. et al. Mitogen-activated protein kinase feedback phosphorylation regulates MEK1 complex formation and activation during cellular adhesion. Mol Cell Biol 24, 2308–2317 (2004).
Zhang, T., Ranade, S., Cai, C. Q., Clouser, C. & Pignoni, F. Direct control of neurogenesis by selector factors in the fly eye: regulation of atonal by Ey and So. Development 133, 4881–4889 (2006).
Xiong, W., Dabbouseh, N. M. & Rebay, I. Interactions with the abelson tyrosine kinase reveal compartmentalization of eyes absent function between nucleus and cytoplasm. Dev Cell 16, 271–279 (2009).
Li, X., Perissi, V., Liu, F., Rose, D. W. & Rosenfeld, M. G. Tissue-specific regulation of retinal and pituitary precursor cell proliferation. Science 297, 1180–1183 (2002).
Koushika, S. P., Lisbin, M. J. & White, K. ELAV, a Drosophila neuron-specific protein, mediates the generation of an alternatively spliced neural protein isoform. Curr Biol 6, 1634–1641 (1996).
Halder, G., Callaerts, P. & Gehring, W. J. Induction of ectopic eyes by targeted expression of the eyeless gene in Drosophila. Science 267, 1788–1792 (1995).
Morillo, S. A., Braid, L. R., Verheyen, E. M. & Rebay, I. Nemo phosphorylates Eyes absent and enhances output from the Eya-Sine oculis transcriptional complex during Drosophila retinal determination. Dev Biol 365, 267–276 (2012).
Chen, R., Halder, G., Zhang, Z. & Mardon, G. Signaling by the TGF-beta homolog decapentaplegic functions reiteratively within the network of genes controlling retinal cell fate determination in Drosophila. Development 126, 935–943 (1999).
Atkins, M. et al. Dynamic rewiring of the Drosophila retinal determination network switches its function from selector to differentiation. PLoS Genet 9, e1003731 (2013).
Halder, G. et al. Eyeless initiates the expression of both sine oculis and eyes absent during Drosophila compound eye development. Development 125, 2181–2191 (1998).
Pauli, T., Seimiya, M., Blanco, J. & Gehring, W. J. Identification of functional sine oculis motifs in the autoregulatory element of its own gene, in the eyeless enhancer and in the signalling gene hedgehog. Development 132, 2771–2782 (2005).
Niimi, T., Seimiya, M., Kloter, U., Flister, S. & Gehring, W. J. Direct regulatory interaction of the eyeless protein with an eye-specific enhancer in the sine oculis gene during eye induction in Drosophila. Development 126, 2253–2260 (1999).
Ostrin, E. J. et al. Genome-wide identification of direct targets of the Drosophila retinal determination protein Eyeless. Genome Res 16, 466–476 (2006).
Anderson, J., Salzer, C. L. & Kumar, J. P. Regulation of the retinal determination gene dachshund in the embryonic head and developing eye of Drosophila. Dev Biol 297, 536–549 (2006).
Ton, C. C. et al. Positional cloning and characterization of a paired box- and homeobox-containing gene from the aniridia region. Cell 67, 1059–1074 (1991).
Xu, P. X., Woo, I., Her, H., Beier, D. R. & Maas, R. L. Mouse Eya homologues of the Drosophila eyes absent gene require Pax6 for expression in lens and nasal placode. Development 124, 219–231 (1997).
Chow, R. L., Altmann, C. R., Lang, R. A. & Hemmati-Brivanlou, A. Pax6 induces ectopic eyes in a vertebrate. Development 126, 4213–4222 (1999).
Szklarczyk, D. et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43, D447–452 (2015).
Anderson, A. M., Weasner, B. M., Weasner, B. P. & Kumar, J. P. Dual transcriptional activities of SIX proteins define their roles in normal and ectopic eye development. Development 139, 991–1000 (2012).
Boyle, M., Bonini, N. & DiNardo, S. Expression and function of clift in the development of somatic gonadal precursors within the Drosophila mesoderm. Development 124, 971–982 (1997).
Thomason, L. et al. Recombineering: genetic engineering in bacteria using homologous recombination. Curr Protoc Mol Biol Chapter 1, Unit 1, 16 (2007).
Pepple, K. L., Anderson, A. E., Frankfort, B. J. & Mardon, G. A genetic screen in Drosophila for genes interacting with senseless during neuronal development identifies the importin moleskin. Genetics 175, 125–141 (2007).
Acknowledgements
We would like to thank past members of Mardon lab (2011–2015) for their kind support and help. We are grateful to Dr. Ming Fa and Dr. Baojun Wu for critical reading of manuscript, Xuan Zhu and Trevor Davis for technical help. We thank the Bloomington Stock Center for providing fly stocks, Dr. Hugo Bellen and Dr. Uwe Walldorf for antibodies and Dr. Justin Kumar, Dr. Rui Chen and Dr. Ilaria Rebay for plasmids.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: M.J. and G.M. Performed the experiments: M.J. Analyzed the data: M.J. and G.M. Contributed reagents/materials/analysis tools: M.J. Wrote the paper: M.J. and G.M.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Jin, M., Mardon, G. Distinct Biochemical Activities of Eyes absent During Drosophila Eye Development. Sci Rep 6, 23228 (2016). https://doi.org/10.1038/srep23228
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23228
- Springer Nature Limited