Abstract
Eukaryotic translation release factor eRF1 is an important cellular protein that plays a key role in translation termination, nonsense-mediated mRNA decay (NMD), and readthrough of stop codons. The amount of eRF1 in the cell influences all these processes. The mechanism of regulation of eRF1 translation through an autoregulatory NMD-dependent expression circuit has been described for plants and fungi, but the mechanisms of regulation of human eRF1 translation have not yet been studied. Using reporter constructs, we studied the effect of eRF1 mRNA elements on its translation in cell-free translation systems and HEK293 cell culture. Our data indicate the absence of an NMD-dependent autoregulatory circuit for human eRF1 expression. We found that the translation of the eRF1 coding sequence is most strongly influenced by the 5′ untranslated region of eRF1 mRNA and the start codon of the upstream open reading frame. According to the transcription start database, eRF1 mRNA is characterized by high heterogeneity of the transcription start and a variable 5' untranslated region in length. In addition, the start codon of the CDS in eRF1 mRNA is located within the known translational regulator of short 5' untranslated regions (TISU), which also stimulates mRNA transcription of genes with high transcription start heterogeneity. We hypothesize that regulation of human eRF1 synthesis occurs at both the transcriptional and translational levels. At the transcription level, the length of the eRF1 5' untranslated region and the number of the upstream open reading frames in it are regulated. This regulation in turn, regulates the production of eRF1 at the translation level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
The translation release factor eRF1 is essential in mediating eukaryotic translation termination. eRF1 consists of three functional domains: the N domain is involved in stop codon recognition [1–6], the M domain ensures the hydrolysis of peptidyl tRNA at the P site of the ribosome [7], and the C domain binds to the additional translation release factor eRF3 [8–13]. In addition, eRF1 is involved in the regulation of stop codon readthrough [14–16], nonsense-mediated decay (NMD) of mRNA [17–20], and premature termination at some rare sense codons [21, 22].
Termination is an important step in translation, which regulates the rate and efficiency of the release of nascent peptides. It has been shown that termination of translation at the upstream open reading frame (uORF) in the 5' untranslated region (5'-UTR) of mRNAs of a number of genes provide regulation of translation of the main product [23]. The study of translation termination also has practical significance. Up to 10% of human orphan genetic diseases are associated with the occurrence of premature stop codons (PTCs), and methods of treating such diseases involve increasing the readthrough of PTCs [24].
Stop codon readthrough and premature termination are opposite processes that depend on the concentration of eRF1 and competing tRNAs [21, 25]. In the readthrough process, the protein product is extended from the C-terminus due to recognition of the stop codon with a competing closely related tRNA, which, in some cases, may be part of the normal physiological processes of the cell [26–28]. The process of premature termination at sense codons is less studied and described for prokaryotes, fungi, and fruit flies, but its wider occurrence in eukaryotes is debated [21, 22, 29]. It is assumed that the physiological meaning of premature termination at sense codons is the release of ribosomes stuck on the mRNA, and this mechanism is an alternative to No-Go mRNA decay (NGD).
NMD is a process of mRNA degradation caused by the presence of PTC, which can be stimulated by the Exon Junction Complex (EJC) at a certain distance after the stop codon. The main function of NMD is to control the quality and remove aberrant mRNAs with premature stop codons resulting from splicing abnormalities. In addition, there is ample evidence for the involvement of NMD in the normal regulation of gene expression. mRNAs can undergo NMD even in the absence of erroneous exon joining and the appearance of a premature stop codon. Such events may be due to either an EJC-independent NMD or a non-canonical EJC location. The longer the 3'-UTR, the higher the probability of NMD [19].
Thus, studying the production of eRF1 is crucial since the level of this protein in the cell will determine the efficiency of the important cellular processes described above. The expression of the ETF1 gene, encoding human eRF1, has been studied at the transcriptional level [30], however, there are no studies on the regulation of eRF1 translation in mammals. It is worth noting that an autoregulatory circuit of eRF1 expression has been described for plants and fungi. At a high content of eRF1, the mechanism of decay of its own mRNA is triggered due to the presence of EJC after the first stop codon, which is competent to stimulate NMD. Because of this, the amount of eRF1 in the cell decreases, and the stop codon of its own mRNA is readthrough, which inhibits its NMD [31, 32].
In this work, using firefly luciferase (Fluc) and nanoluciferase (Nluc) reporters, we study the effect of eRF1 mRNA elements (5'-UTR, CDS, 3'-UTR) on translation. We have shown that the main contribution to translation regulation is made by the 5'-UTR of eRF1 mRNA and additional start codons located in it. The most likely mechanism for controlling eRF1 translation is regulation of the length of its 5'-UTR due to the frequency of use of transcription initiation sites.
EXPERIMENTAL
Cloning of the 3'-UTR of eRF1 mRNA. Total HEK293 RNA was isolated using ExtractRNA (Evrogen, BC032) according to the manufacturer’s protocol. This RNA was used to generate cDNA using the AMV Reverse Transcription System (Promega, A3500) with oligo(dT)15 or random primers (Table S1). Next, using primers CDS F and oligo(dT)15 (Table S1), a PCR product was obtained and cloned into the TA vector pGEM®-T Easy Vector (Promega, A1360). The cloned product contained part of the eRF1 CDS and the 3'-UTR sequence of eRF1 mRNA, including 861 nt after the UAG stop codon of the eRF1 CDS and then A(15).
Creation of experimental designs and synthesis of mRNA. Nl constructs with PTCs and controls with coding codons. Plasmid pNl-gl was previously described [33]. A partial sequence of β-globin CDS with PTC UGA and context CUAGUA (weak), enhancing stop codon readthrough, was inserted into the pNl-gl construct at the NcoI restriction site [34, 35]. The start codon in the β-globin sequence was replaced with ATA using the QuikChange Site-Directed Mutagenesis Kit (Agilent) to obtain the UGA_Nl plasmid. Using the same set, all other plasmids UGU_Nl, UAA_Nl/AAA_Nl, UAG_Nl/UAU_Nl were obtained.
The studied Nl constructs for readthrough of eRF1 mRNA stop codons. The PCR product Nl was obtained from the plasmid pNl-gl with primers pNl F/R. The corresponding PCR products were obtained using primers 5'1 F/R, 5'2 F/R, 5'2 F, and 5'2sense R (Table S1). Using the Gibson Assembly system (NEB, E2611L), plasmid 5'_Nl (1) was obtained from the PCR products Nl, 5'1 and 5'2. Plasmid 5'sense_Nl (1), in which the TAG stop codon of the eRF1 mRNA uORF was replaced by a TCG encoding serine, was obtained from the PCR products Nl, 5'1 and 5'2sense (Table S2). The Nl-gl PCR product was obtained from the pNl-gl plasmid with primers pNl F and pNl-gl R (Table S1). Using the Gibson Assembly system (NEB, E2611L), plasmid 5'_Nl (3) was obtained from the PCR products Nl-gl and 5'2, and plasmid 5'sense_Nl (3) was obtained from the PCR products Nl-gl and 5'2sense (Table S2). PCR products CDS and CDSsense were synthesized from the petSUMO-eRF1 plasmid, using primer pairs pNl_eRF1 F/R, pNl_eRF1 F and pNl_eRF1-Ser R (Table S1). Using the Gibson Assembly system (NEB, E2611L), the plasmid CDS_Nl was obtained from the PCR products Nl-gl and CDS, and the plasmid CDSsense_Nl, in which the TAG stop codon of the eRF1 CDS was replaced with TCG, encoding serine, was obtained from the PCR products Nl-gl and CDSsense (Table S2). The construct with the eRF1 CDS additionally contained a natural 3' context of 9 nucleotides after the UAG stop codon.
Fluc constructs to study the effect of the 5'- and 3'‑UTR of eRF1 mRNA on translation. The Fl PCR product was obtained from the pGEM-Fluc plasmid (Promega, cat. no. E1541) using primers Fl F/R. 3'GAPDH PCR product was obtained using the 3'GAPDH F/R primers. Plasmid Fl_3'GAPDH was obtained from the PCR products of Fl and 3'GAPDH using the Gibson Assembly system (NEB, E2611L) (Table S2). 3'eRF1 PCR product was obtained from the pGEM-3'eRF1 plasmid using primers 3' F and 3' R. Plasmid Fl_3'eRF1 was obtained from the PCR products of Fl and 3'eRF1 using the Gibson Assembly system (NEB, E2611L) (Table S2). PCR products 5'_Fl and 5'sense_Fl were obtained using primers 5'_Fl F/R and plasmids 5'_Nl (1) and 5'sense_Nl (1). The PCR product 5'_Fl was inserted into the plasmids Fl_3'GAPDH and Fl_3'eRF1 at the NheI and XbaI restriction sites to obtain the constructs 5'eRF1_Fl_3'GAPDH and 5'eRF1_Fl_3'eRF1, and the PCR product 5'sense_Fl was inserted into the plasmid at Fl_3'eRF1 to obtain 5'eRF1sense_Fl_3'eRF1 (Table S2).
The mRNAs used in the work were obtained from the corresponding PCR products containing the T7 promoter at the 5' end and the poly-A(50) tail at the 3' end, which were introduced using primers and the T7 RiboMAX™ Large Scale RNA Production System kit (Promega) with addition of an mRNA cap structure analogue (ARCA, NEB) according to the manufacturer’s protocol. A list of mRNAs and the primers used for their synthesis are given in the Supplementary Information (Table S2).
Isolation and purification of recombinant eRF1. Recombinant eRF1 was obtained from our previously created petSUMO-eRF1 construct as described for NSP1 [33]. Then, the eRF1 was further purified using ion exchange chromatography on a HiTrap Q HP column (Cytiva).
In vivo and in vitro translation in cell-free translation systems and HEK293 cells. Translation of reporter mRNAs was carried out in 10 μL of a mixture containing 50% HEK293 lysate, 20 mM HEPES-KOH pH 7.5, 2 mM DTT, 0.25 mM spermidine, 0.6 mM MgAc2, 16 mM creatine phosphate, 0.06 U/μL creatine kinase, 1 mM ATP, 0.6 mM GTP, 60 mM KAc, 0.05 mM each amino acid, 0.2 U/μL RiboLock, 5 mM D-luciferin (or 1% Nano-Glo® (Promega)). eRF1 was added to a final concentration of 0.6 μM. Luminescence was measured at 30°C in a Tecan Infinite 200 Pro plate reader for 100 min. Translation efficiency was calculated as the maximum derivative of the growing linear portion of the luminescence curve (v0, RLU/min). Stop codon readthrough was calculated as the ratio of the translation efficiency of the template with PTC to the translation efficiency of the control template with a sense codon at a given position.
In vivo translation of reporters was carried out in HEK293 cells in accordance with the previously described FLERT (Fleeting mRNA Transfection technique) [37]. HEK293 cells were cultured in DMEM with 10% fetal bovine serum at 37°C and 5% CO2. The day before transfection, exponentially growing HEK293 cells were seeded into 24-well plates at 50% coverage. After 12–16 h of growth, when the cell density reached 70–90%, transfection was performed using GenJector™-U (Molecta, Russia). To do this, 0.5 pmol of reporter mRNA and 0.05 pmol of control mRNA were incubated with 0.2 μl of transfection reagent in 60 μL of Opti-MEM (Gibco, 11058-021) for 10 min, and then added to the culture medium of Hek293T cells in the appropriate well. After two hours, cells were harvested and luciferase activity was analyzed using the Dual Luciferase® Assay System (Promega). All transfections were repeated several times in different cell passages.
Statistical analysis. Experiments were carried out in three or more repetitions. Data are presented as mean ± standard error of the mean (SE). An unpaired two-tailed t test was used to compare means between two groups. Multiple comparisons were performed using the Holm correction for p value [34]. Asterisks indicate statistically significant differences (*p < 0.05; **p < 0.01; ****p < 0.0001).
RESULTS AND DISCUSSION
eRF1 mRNA Organization
The human eRF1 mRNA reference sequence in the NCBI RefSeq database has the identification number NM_004730. A diagram of this mRNA with elements annotated in the GenBank database is presented in Fig. 1a. eRF1 mRNA has a long 3' UTR located within one exon, that is, there are no EJCs after the stop codon of CDS. Thus, the mechanism of expression autoregulation due to EJC-dependent NMD is not applicable for human eRF1.
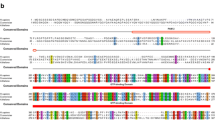
Structure of eRF1 mRNA. (a) Scheme of organization of the reference eRF1 mRNA NM_004730.The arrows indicate the primers used to obtain the 3'-UTR of eRF1 mRNA. (b) Scheme of organization of the eRF1 5'-UTR and usage frequency of transcription start sites according to the DBTSS database. Green rectangles indicate start codons, and red rectangles indicate stop codons. The number inside the rectangles indicates the reading frame relative to the CDS. The number next to the green squares means the AUG serial number relative to the start codon of the CDS, taken as 0. The sequence of the TISU regulatory element is marked in light green. The shortened sequence of the eRF1 5'-UTR studied in the work, along with the full-length one, is highlighted in gray.
The 3'-UTR of eRF1 mRNA was cloned from the human embryonic kidney cell line HEK293. Total mRNA was isolated from HEK293 cells (in three biological replicates) and two types of cDNA were synthesized on each sample using oligo-dT or random primers. PCR products approximately 1600 bp long were synthesized from cDNA samples obtained using oligo-dT and a specific forward primer, and then cloned into a vector. The resulting products were sequenced and found to correspond to the region between the forward primer and polyadenylation site 1, which is a shortened fragment of the 3' UTR (Fig. 1a). However, larger PCR products were not detected. Next, we tried to synthesize PCR products from each cDNA variant obtained with a forward specific primer and one of the reverse primers for the 3'‑UTR of eRF1 mRNA (3'R 1−5). Moreover, in all cases, PCR products were obtained only with reverse primers 3'R 4−5 (Fig. 1a). Sequencing of these PCR products showed their correspondence to the previously cloned truncated 3'-UTR. Thus, HEK293 predominantly contains a truncated version of the 3'‑UTR of eRF1 mRNA, which was subsequently used in experiments.
The 5'-UTR of eRF1 mRNA contains additional start and stop codons in different reading frames (Fig. 1b). Interestingly, the second start (−2) and stop codons organize the minimal uORF (AUGUGA), located in a different reading frame relative to the CDS. In this case, the third start and stop codons form another uORF (hereinafter also 5'ORF) in the same reading frame with the CDS. Detailed data on transcriptional starts of NM_004730 (and the size of the 5'-UTR) are available in the DBTSS database (Data Base Transcriptional Start Sites, https://dbtss.hgc.jp) [35]. According to these data, the full-length 5'-UTR of eRF1 in HEK293 (and other samples available in the database) is synthesized extremely rarely, and transcription starts are located in a wide range, mainly before and after additional start codons (Fig. 1b). We obtained the full-length 5'-UTR of eRF1 (hereinafter referred to as the “entire 5'-UTR” of eRF1), as well as a shortened version of the 5'-UTR of eRF1 (hereinafter referred to as the “5'-UTR” of eRF1), found in the bulk NM_004730 mRNA transcripts and does not include the first two additional start and stop codons (Fig. 1b). Thus, eRF1 mRNA in cells is most often shortened at the 5' and 3' ends relative to the reference sequence, and the size of its 5'-UTR varies widely.
The Length of the eRF1 5' -UTR Influences Translation and Readthrough Efficiency. eRF1 Does Not Affect Its Own Translation
To study the effect of the eRF1 5'-UTR and eRF1 protein on translation and stop codons readthrough, we generated a series of constructs with the Nluc reporter (Fig. 2b). In these constructs, Nluc was devoid of its own start codon. In the 5'_Nl (1-3) constructs, the Nluc reporter detected translation of the eRF1 5'-UTR uORF (5'-ORF). To study readthrough, corresponding control constructs were created for these ones, in which the UAG stop codon was replaced by the serine-encoding codon UCG. 5'_Nl (1) contained the entire eRF1 5'-UTR from the reference sequence NM_004730, and 5'_Nl (2) contained a truncated version (marked in gray in Fig. 1b). 5'_Nl (3) contained a hybrid 5'-UTR in which the 5'-UTR of the β-globin (HBB) sequence was located immediately before the 5'-ORF and did not contain additional start and stop codons. CDS upstream of the reporter contains the HBB 5′-UTR followed by the complete eRF1 CDS and the 9 nucleotide natural 3' context of its stop codon.
Effect on readthrough and translation of the 5'-UTR and eRF1 CDS, as well as excess eRF1. (a) General diagram of constructs with premature stop codons before the nanoluciferase CDS. The construct includes the 5'-UTR and the first 13 codons of the β-globin mRNA CDS followed by a stop/sense codon (UAA/AAA, UAG/UAU, UGA/UGU) and the corresponding 6‑nt 3' context (GGGCUG, GAUAAU, CUAGUA ) to enhance readthrough, followed by an Nluc CDS without its own start codon in the same reading frame. The 3'-UTR is artificial and is produced from ~170 nucleotides after the stop codon of the Nluc CDS sequence of the original plasmid. (b) Scheme of model constructs for studying readthrough of eRF1 mRNA stop codons. (c) Level of readthrough of model mRNAs with a reporter containing nanoluciferase measured in cell-free translation systems. (d) Effect of excess eRF1 on readthrough of stop codons in models with 5'-UTR and CDS of self-mRNA. e—Effect of the length of the 5'-UTR of eRF1 on translation. The compared templates are identical to the sequences after the start of translation and differ only in the length of the 5'-UTR. R.L.U.—relative luminescence units. n.s., differences are not significant.
Independent constructs were created to evaluate readthrough at different stop codons, the general scheme of which is presented in Fig. 1a. These constructs in front of the Nluc CDS contain a 5'-UTR and the first 13 codons of the HBB CDS, followed by a stop/sense codon (UAA/AAA, UAG/UAU, UGA/UGU) and the corresponding 6-nt 3' context (GGGCUG, GAUAAU, CUAGUA) to enhance readthrough [36].
The data obtained on the readthrough of the UAG 5'-ORF stop codon show that it is influenced by the preceding sequence: the maximum efficiency of readthrough is observed on the shortened 5'-UTR of eRF1 (more common in the cell), the stop codon with the full-length 5'-UTR of eRF1 is read slightly worse, and the worst is the readthrough of the stop codon with the 5'-UTR of HBB (Fig. 2c). At the same time, the readthrough of the UAG 5'-ORF in the presence of the eRF1 5'-UTR is higher than that of the model construct with a premature termination codon (PTC) UAG_Nl, but the HBB 5'-UTR reduces the efficiency of the readthrough. The readthrough of the UAG CDS eRF1 does not differ from the readthrough of the UAG_Nl. The lack of increased readthrough of the eRF1 CDS provides further evidence against the possibility of autoregulation of eRF1 translation by NMD described for plants and fungi. Moreover, the addition of excess eRF1 does not affect the readthrough of stop codons in the 5'-ORF and the main ORF of eRF1 and does not affect the translation of the corresponding constructs (Figs. 2d, 2e). These data contradict the possibility of autoregulation of eRF1 translation. Translation efficiency in the presence of the full-length eRF1 5'-UTR is lower than with the truncated version, which is most likely due to the presence of a larger number of additional start codons (Fig. 2e).
5'-UTR and 3'-UTR of eRF1 Ensure Efficient Translation of Its CDS
To study the effect of eRF1 5'- and 3'-UTRs on eRF1 translation, model mRNAs were generated based on the Fluc reporter (Fig. 3a). In contrast to the small Nluc CDS, the Fluc CDS has a similar length and GC composition to the eRF1 CDS. The 5'-UTR of HBB and the 3'-UTR of glyceraldehyde phosphate dehydrogenase (GAPDH) were used as controls. These proteins are present in cells in high concentrations, and their UTRs should facilitate efficient translation. In vitro in HEK293 cell lysate, the eRF1 5'-UTR increased translation efficiency, even relative to the HBB 5'-UTR, while the eRF1 3'-UTR did not improve translation of the Fluc reporter (Fig. 3b). The 5'- and 3'-UTRs of eRF1 jointly mediate the maximum level of reporter translation. We repeated the same experiment in vivo in HEK293 cells, transfecting them with the same mRNAs in accordance with the previously described FLERT (Fleeting mRNA Transfection technique) [37]. In vivo, the eRF1 5'- and 3'-UTRs together significantly improved translation of the Fluc reporter, while the eRF1 3'-UTR alone did not significantly affect translation (Fig. 3c). Thus, the 5'-UTR of eRF1 most strongly affects the translation of the release factor, and the effect of the 3'-UTR of eRF1 on its translation depends on the presence of the eRF1 5'-UTR.
Effect on translation of the 5'-UTR and 3'-UTR of eRF1 mRNA. (a) Scheme of constructs encoding firefly luciferase used in the experiments. (b) Effect of 5'- and 3'-UTR of eRF1 mRNA on Fluc translation in HEK293 lysate. (c) Effect of 5'- and 3'-UTR of eRF1 mRNA on Fluc translation in HEK293 cell culture.
Start Codon of eRF1 5'-UTR Inhibits Translation
We studied the influence of the additional start and stop codons that form the uORF in the 5'-UTR of eRF1 on the translation of the CDS using variants of the Fluc CDS mRNA with the 5'- and 3-UTR of eRF1, in which the start and/or a stop codon was replaced by a sense codon (AAG and UCG, respectively). These constructs were used for in vivo experiments in HEK293 cell culture using FLERT [37]. Removal of the alternative start codon from the eRF1 5′-UTR resulted in a more than twofold increase in the translation efficiency of the reporter, while deletion of the stop codon in the eRF1 5′-UTR had virtually no effect on translation (Fig. 4). Thus, the most prominent translation regulator in eRF1 mRNA is the start codon of the uORF in the eRF1 5′-UTR.
Effect of start and stop codons in the uORF of the eRF1 mRNA 5'-UTR. (a) General diagram of the constructs, representing 4 variants of mRNA: “WT”—wild type 5'-UTR of eRF1 is not changed and contains the start and stop codon of the uORF, “-start in the 5'-UTR”—the start codon of the uORF is replaced by codon AAG, “-stop in the 5'-UTR”—the stop codon of the uORF is replaced by UCG, “-start and stop in the 5'-UTR”—the start codon of the uORF is replaced by the AAG codon, and the stop codon by UCG. (b) Efficiency of in vivo translation in HEK293 cells of reporter mRNA with mutations in the start and stop codon of the uORF 5'-UTR from eRF1 mRNA.
Mechanism of eRF1 mRNA Translation Regulation by Non-Coding Regions
Our data demonstrate that the most pronounced effect on translation is exerted by the eRF1 5'-UTR, as well as its combination with the eRF1 3'-UTR, stimulating the translation of its CDS. At the same time, the start codon of the uORF in the 5'-UTR of eRF1 inhibits translation, since its removal additionally increases the translation efficiency by more than 2 times, which is generally consistent with the generally accepted ideas about the regulation of uORF translation [38–41]. According to our findings, we propose that eRF1 translation is regulated at two levels: the transcriptional level and the translational level. As can be seen from Figure 1b, eRF1 mRNA is heterogeneous, has leaders of different lengths, and multiple transcription start sites are concentrated in the region of additional start codons in the 5'-UTR. Thus, due to transcriptional regulation, eRF1 mRNA with varying numbers of additional start codons and uORFs can be produced. In turn, additional start codons and uORFs regulate translation according to known mechanisms, inhibiting the translation of the CDS due to the “capture” of translation initiation factors. Thus, the highest translation efficiency of eRF1 should be ensured by transcriptional variants of mRNA with a short 5'-UTR that do not include additional start codons. It should be noted that the regulation of translation of the CDS by uORFs is being actively studied in other objects, and our data are consistent with the generally accepted view that uORFs suppress the translation of the CDS [38–41]. In addition, a number of studies raises the question of translation regulation due to the heterogeneity of the transcription start of some mRNAs, leading to the appearance of a set of 5'-UTRs of different lengths in them [42–44], which corresponds to our findings about eRF1 transcription.
In addition, we noticed that the start codon of the CDS eRF1 is part of the translational-transcriptional regulatory element TISU (Translation Initiator of Short 5′-UTR). Previously, it was believed that translation of the first start codon of an mRNA with a too short 5'-UTR occurs inefficiently, with “leakage” of the translation start to the next AUG [45]. But, later, a regulatory element was described that promotes efficient translation from the first start codon in mRNA templates with a short 5'-UTR, called TISU [46]. The localization of TISU partially overlaps with the Kozak sequence. In addition, TISU has been described as a cis-transcription regulator capable of binding to the transcription factor YY1 (Yin Yang 1) and effectively stimulating the synthesis of mRNA from gene promoters with high heterogeneity of transcription start sites [46]. The presence and position of TISU in eRF1 mRNA is consistent with our hypothesis of a two-level transcriptional-translational regulation of eRF1 translation.
In general, we can conclude that the translational regulation of the human eRF1 CDS is fundamentally different from the translational regulation of plant and fungal eRF1, where the existence of an NMD-dependent autoregulatory expression circuit is described [31, 32]. The production of human release factor eRF1 does not depend on its concentration in the cell and is regulated by its own 5'- and 3'-UTRs.
ABBREVIATIONS AND NOTATION
NMD | nonsense-mediated mRNA decay |
5'-UTR and 3'-UTR | untranslated region |
CDS | coding sequence |
uORF | upstream open reading frame |
EJC | exon junction complex |
Fluc (Fl) | firefly luciferase |
Nluc (Nl) | nanoluciferase |
HBB | β-globin |
ETF1 | eukaryotic translation termination factor 1—a gene coding eRF1 |
GAPDH | Glyceraldehyde 3-phosphate dehydrogenase |
PTC | premature termination codon |
TSS | transcription start site |
RLU | Relative Light Units |
REFERENCES
Bertram G., Bell H.A., Ritchie D.W., Fullerton G., Stansfield I. 2000. Terminating eukaryote translation: domain 1 of release factor eRF1 functions in stop codon recognition. RNA. 6, 1236–1247.
Frolova L., Seit-Nebi A., Kisselev L. 2002. Highly conserved NIKS tetrapeptide is functionally essential in eukaryotic translation termination factor eRF1. RNA. 8, 129–136.
Bulygin K.N., Khairulina Y.S., Kolosov P.M., Ven’yaminova A.G., Graifer D.M., Vorobjev Y.N., Frolova L.Y., Kisselev L.L., Karpova G.G. 2010. Three distinct peptides from the N domain of translation termination factor eRF1 surround stop codon in the ribosome. RNA. 16, 1902–1914.
Bulygin K.N., Khairulina Y.S., Kolosov P.M., Venꞌyaminova A.G., Graifer D.M., Vorobjev Y.N., Frolova L.Y., Karpova G.G. 2011. Adenine and guanine recognition of stop codon is mediated by different N domain conformations of translation termination factor eRF1. Nucleic Acids Res. 39, 7134–7146.
Kryuchkova P., Grishin A., Eliseev B., Karyagina A., Frolova L., Alkalaeva E. 2013. Two-step model of stop codon recognition by eukaryotic release factor eRF1. Nucleic Acids Res. 41, 4573–4586.
Brown A., Shao S., Murray J., Hegde R.S., Ramakrishnan V. 2015. Structural basis for stop codon recognition in eukaryotes. Nature. 524, 493–496.
Frolova L.Y., Tsivkovskii R.Y., Sivolobova G.F., Oparina N.Y., Serpinsky O.I., Blinov V.M., Tatkov S.I., Kisselev L.L. 1999. Mutations in the highly conserved GGQ motif of class 1 polypeptide release factors abolish ability of human eRF1 to trigger peptidyl-tRNA hydrolysis. RNA. 5, 1014–1020.
Zhouravleva G., Frolova L., Le Goff X., Le Guellec R., Inge-Vechtomov S., Kisselev L., Philippe M. 1995. Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3. EMBO J. 14, 4065–4072.
Frolova L., Le Goff X., Zhouravleva G., Davydova E., Philippe M., Kisselev L. 1996. Eukaryotic polypeptide chain release factor eRF3 is an eRF1- and ribosome-dependent guanosine triphosphatase. RNA. 2, 334–341.
Stansfield I., Jones K.M., Kushnirov V.V., Dagkesamanskayal A.R., Poznyakovski A., Paushkin S.V., Nierras C.R., Cox B.S., Ter-Avanesyan M.D., Tuite M.F. 1995. The products of the SUP45 (eRF1) and SUP35 genes interact to mediate translation termination in Saccharomyces cerevisiae. EMBO J. 14, 4365–4373.
Paushkin S.V., Kushnirov V.V., Smirnov V.N., Ter-Avanesyan M.D. 1997. Interaction between yeast Sup45p (eRF1) and Sup35p (eRF3) polypeptide chain release factors: implications for prion-dependent regulation. Mol. Cell. Biol. 17, 2798–2805.
Ito K., Ebihara K., Nakamura Y. 1998. The stretch of C-terminal acidic amino acids of translational release factor eRF1 is a primary binding site for eRF3 of fission yeast. RNA. 4, S1355838298971874.
Merkulova T.I., Frolova L.Y., Lazar M., Camonis J., Kisselev L.L. 1999. C-terminal domains of human translation termination factors eRF1 and eRF3 mediate their in vivo interaction. FEBS Lett. 443, 41–47.
Namy O., Hatin I., Rousset J. 2001. Impact of the six nucleotides downstream of the stop codon on translation termination. EMBO Rep. 2, 787–793.
Atkins J.F., Loughran G., Bhatt P.R., Firth A.E., Baranov P.V. 2016. Ribosomal frameshifting and transcriptional slippage: from genetic steganography and cryptography to adventitious use. Nucleic Acids Res. 44, 7007–7078
Rodnina M.V., Korniy N., Klimova M., Karki P., Peng B.-Z., Senyushkina T., Belardinelli R., Maracci C., Wohlgemuth I., Samatova E., Peske F. 2020. Translational recoding: Canonical translation mechanisms reinterpreted. Nucleic Acids Res. 48, 1056–1067.
Brogna S., Wen J. 2009. Nonsense-mediated mRNA decay (NMD) mechanisms. Nat. Struct. Mol. Biol. 16, 107–113.
Chabelskaya S.V., Zhouravleva G.A. 2010. Mutations in the SUP35 gene impair nonsense-mediated mRNA decay. Mol. Biol. (Moscow). 44, 45–53. https://doi.org/10.1134/S0026893310010073
Schweingruber C., Rufener S.C., Zünd D., Yamashita A., Mühlemann O. 2013. Nonsense-mediated mRNA decay—mechanisms of substrate mRNA recognition and degradation in mammalian cells. Biochim. Biophys. Acta, Gene Regul. Mech. 1829, 612–623.
Karousis E.D., Gurzeler L.-A., Annibaldis G., Dreos R., Mühlemann O. 2020. Human NMD ensues independently of stable ribosome stalling. Nat. Commun. 11, 4134.
Yang Q., Yu C.-H., Zhao F., Dang Y., Wu C., Xie P., Sachs M.S., Liu Y. 2019. eRF1 mediates codon usage effects on mRNA translation efficiency through premature termination at rare codons. Nucleic Acids Res. 47, 9243–9258.
Wada M., Ito K. 2019. Misdecoding of rare CGA codon by translation termination factors, eRF1/eRF3, suggests novel class of ribosome rescue pathway in S. cerevisiae. FEBS J. 286, 788–802.
Dever T.E., Ivanov I.P., Sachs M.S. 2020. Conserved upstream open reading frame nascent peptides that control translation. Annu. Rev. Genet. 54, 237–264.
Bidou L., Allamand V., Rousset J.-P., Namy O. 2012. Sense from nonsense: therapies for premature stop codon diseases. Trends Mol. Med. 18, 679–688.
Janzen D.M. 2004. The effect of eukaryotic release factor depletion on translation termination in human cell lines. Nucleic Acids Res. 32, 4491–4502.
Freitag J., Ast J., Bölker M. 2012. Cryptic peroxisomal targeting via alternative splicing and stop codon read-through in fungi. Nature. 485, 522–525.
Schueren F., Lingner T., George R., Hofhuis J., Dickel C., Gärtner J., Thoms S. 2014. Peroxisomal lactate dehydrogenase is generated by translational readthrough in mammals. ELife. 3, e03640
Hofhuis J., Schueren F., Nötzel C., Lingner T., Gärtner J., Jahn O., Thoms S. 2016. The functional readthrough extension of malate dehydrogenase reveals a modification of the genetic code. Open Biol. 6, 160246.
Svidritskiy E., Demo G., Korostelev A.A. 2018. Mechanism of premature translation termination on a sense codon. J. Biol. Chem. 293, 12472–12479.
Dubourg C., Toutain B., Le Gall J.-Y., Le Treut A., Guenet L. 2003. Promoter analysis of the human translation termination factor 1 gene. Gene. 316, 91–101.
Nyikó T., Auber A., Szabadkai L., Benkovics A., Auth M., Mérai Z., Kerényi Z., Dinnyés A., Nagy F., Silhavy D. 2017. Expression of the eRF1 translation termination factor is controlled by an autoregulatory circuit involving readthrough and nonsense-mediated decay in plants. Nucleic Acids Res. 45, 4174–4188.
Kurilla A., Szőke A., Auber A., Káldi K., Silhavy D. 2020. Expression of the translation termination factor eRF1 is autoregulated by translational readthrough and 3'-UTR intron-mediated NMD in Neurospora crassa. FEBS Lett. 594, 3504–3517.
Shuvalov A., Shuvalova E., Biziaev N., Sokolova E., Evmenov K., Pustogarov N., Arnautova A., Matrosova V., Egorova T., Alkalaeva E. 2021. Nsp1 of SARS-CoV-2 stimulates host translation termination. RNA Biol. 18, 804–817.
Sokolova E.E., Vlasov P.K., Egorova T.V., Shuvalov A.V., Alkalaeva E.Z. 2020. The Influence of A/G composition of 3' stop codon contexts on translation termination efficiency in eukaryotes. Mol. Biol. (Moscow). 54, 739–748. https://doi.org/10.1134/S0026893320050088
Biziaev N., Sokolova E., Yanvarev D.V., Toropygin I.Y., Shuvalov A., Egorova T., Alkalaeva E. 2022). Recognition of 3′ nucleotide context and stop codon readthrough are determined during mRNA translation elongation. J. Biol. Chem. 298, 102133.
Holm S. 1979. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70.
Suzuki A., Kawano S., Mitsuyama T., Suyama M., Kanai Y., Shirahige K., Sasaki H., Tokunaga K., Tsuchihara K., Sugano S., Nakai K., Suzuki Y. 2018. DBTSS/DBKERO for integrated analysis of transcriptional regulation. Nucleic Acids Res. 46, D229–D238.
Akulich K.A., Andreev D.E., Terenin I.M., Smirnova V.V., Anisimova A.S., Makeeva D.S., Arkhipova V.I., Stolboushkina E.A., Garber M.B., Prokofjeva M.M., Spirin P.V., Prassolov V.S., Shatsky I.N., Dmitriev S.E. 2016. Four translation initiation pathways employed by the leaderless mRNA in eukaryotes. Sci. Rep. 6, 37905.
Lin Y., May G.E., Kready H., Nazzaro L., Mao M., Spealman P., Creeger Y., McManus C.J. 2019. Impacts of uORF codon identity and position on translation regulation. Nucleic Acids Res. 47, 9358–9367.
Dever T.E., Ivanov I.P., Hinnebusch A.G. 2023. Translational regulation by uORFs and start codon selection stringency. Genes Dev. 37, 474–489.
Starck S.R., Tsai J.C., Chen K., Shodiya M., Wang L., Yahiro K., Martins-Green M., Shastri N., Walter P. 2016. Translation from the 5′ untranslated region shapes the integrated stress response. Science. 351, aad3867
Akulich K.A., Sinitcyn P.G., Makeeva D.S., Andreev D.E., Terenin I.M., Anisimova A.S., Shatsky I.N., Dmitriev S.E. 2019. A novel uORF-based regulatory mechanism controls translation of the human MDM2 and eIF2D mRNAs during stress. Biochimie. 157, 92–101.
Dieudonné F.-X., O’Connor P.B.F., Gubler-Jaquier P., Yasrebi H., Conne B., Nikolaev S., Antonarakis S., Baranov P.V., Curran J. 2015. The effect of heterogeneous transcription start sites (TSS) on the translatome: Implications for the mammalian cellular phenotype. BMC Genomics. 16, 986.
Wang X., Hou J., Quedenau C., Chen W. 2016. Pervasive isoform-specific translational regulation via alternative transcription start sites in mammals. Mol. Syst. Biol. 12, 875
Li H., Bai L., Li H., Li X., Kang Y., Zhang N., Sun J., Shao Z. 2019. Selective translational usage of TSS and core promoters revealed by translatome sequencing. BMC Genomics. 20, 282.
Kozak M. 1991. A short leader sequence impairs the fidelity of initiation by eukaryotic ribosomes. Gene Expression. 1, 111–115.
Elfakess R., Dikstein R. 2008. A translation initiation element specific to mRNAs with very short 5′UTR that also regulates transcription. PLoS One. 3, e3094.
Funding
The research was supported by a grant from the Russian Science Foundation (Project No. 22-24-01019).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
CONFLICT OF INTEREST
The authors of this work declare that they have no conflicts of interest.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This work does not contain any studies involving human and animal subjects.
Additional information
Publisher’s Note.
Pleiades Publishing remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Shuvalov, A.V., Klishin, A.A., Biziaev, N.S. et al. Human eRF1 Translation Regulation. Mol Biol 58, 708–717 (2024). https://doi.org/10.1134/S0026893324700298
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893324700298