Abstract
Background
In mice, germ cells are specified through signalling between layers of cells comprising the primitive embryo. The function of Dppa3 (also known as Pgc7 or stella), a gene expressed in primordial germ cells at the time of their emergence in gastrulating embryos, is unknown, but a recent study has claimed that it plays a central role in germ cell specification.
Results
To test Dppa3's role in germ cell development, we disrupted the gene in mouse embryonic stem cells and generated mutant animals. We were able to obtain viable and fertile Dppa3-deficient animals of both sexes. Examination of embryonic and adult germ cells and gonads in Dppa3-deficient animals did not reveal any defects. However, most embryos derived from Dppa3-deficient oocytes failed to develop normally beyond the four-cell stage.
Conclusion
We found that Dppa3 is an important maternal factor in the cleavage stages of mouse embryogenesis. However, it is not required for germ cell specification.
Similar content being viewed by others
Background
Among the many specialized cell types present in adult mammals, the first to be programmed or specified during embryogenesis are germ cells, which give rise to eggs and sperm. Which molecules direct this programming of germ cells? In many other animals, including flies and worms, material known as "germ plasm" is laid down in the egg before fertilization, and its subsequent passage to a subset of embryonic cells dictates their fate as germ cells [1, 2]. In mammalian embryos, germ cells are specified in a very different manner, through signalling between layers of cells comprising the primitive embryo [3, 4].
Recently, Saitou, Barton and Surani proposed a molecular pathway by which these intercellular signals are translated into germ cell fate in mice [5, 6]. Central to this proposed program of germ cell specification is stella / PGC7 / Dppa3, a gene expressed in primordial germ cells and their descendants, including oocytes [5, 7, 8]. Here we will use the name Dppa3, as approved by the Mouse Genome Informatics Database, when referring to this gene. Saitou and colleagues' model of Dppa3's role in germ cell specification was based on the timing and site of the gene's expression, not on functional analysis. Nonetheless, the model makes clear predictions as to the phenotype of mice lacking Dppa3 function: such embryos should not form germ cells. We tested this prediction and sought to clarify the gene's importance by generating Dppa3-deficient mice and examining their germline development.
Results and Discussion
We disrupted the Dppa3 gene in cultured embryonic stem (ES) cells and thereby generated Dppa3-deficient mice. Specifically, we replaced the entire open reading frame of Dppa3 in mouse V6.5 ES cells [9] with a hygromycin-thymidine kinase selection cassette flanked by loxP sites (Figure 1A). The selection cassette was subsequently removed via transient expression of Cre recombinase in targeted ES cells. The resulting heterozygous Dppa3 tm1WHT/+ ES cells were used to generate chimeric mice, which transmitted the mutation to offspring. Intercrosses between Dppa3 tm1WHT/+ heterozygous animals yielded Dppa3 tm1WHT/Dppa3 tm1WHThomozygotes as well as Dppa3 tm1WHT/+ heterozygotes and +/+ offspring, demonstrating that zygotic function of Dppa3 is not essential for viability (Figure 1B). This allowed us to characterize germ cell development in animals lacking Dppa3.
Generation of Dppa3 -deficient animals. A, Schematic representation of genomic ablation of Dppa3. The gene's four exons are shown; non-coding regions of the first and last exons are shaded gray. The hygromycin-thymidine kinase (Hygro-TK) cassette replaces the entire open reading frame of the gene. Cre-mediated excision of the selection cassette leaves only the non-coding portions of the gene, together with a single loxP site (white triangle). Also shown are the locations of genotyping primers p1, p2 and p3 in wild-type and mutated Dppa3 alleles. B, PCR genotyping of the offspring of an intercross between Dppa3 tm1WHT/+ animals. Inferred genotypes are shown above the gel image. The wild type allele yields a PCR product of 304 bp with primers p1 and p2. The mutant allele (Dppa3 tm1WHT) yields a PCR product of 492 bp with primers p1 and p3. M, DNA molecular weight marker.
Dppa3is not required for germ cell specification
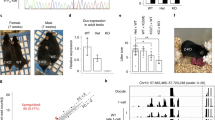
Our findings do not support the proposed centrality of Dppa3 in germ cell programming. First, the gonads of Dppa3 tm1WHT/Dppa3 tm1WHTembryos contained germ cells, identified by expression of alkaline phosphatase, in numbers comparable to those of Dppa3 tm1WHT/ + and +/+ embryos (Figure 2A). Second, the ovaries of Dppa3 tm1WHT/Dppa3 tm1WHTadult females expressed Oct4, a marker of oocytes [10, 11], despite the absence of Dppa3 expression (Figure 2B). Third, histological examination of the gonads of Dppa3 tm1WHT / Dppa3 tm1WHTadults revealed no morphological defects; spermatogenesis in males and ovarian follicle development in females appeared to be normal (Figure 2C,2D). Finally, we obtained fertile Dppa3 tm1WHT/Dppa3 tm1WHTmice of both sexes (though litters from Dppa3 tm1WHT/Dppa3 tm1WHTfemales were small, as described below). Each of these findings demonstrates that the Dppa3 gene is not required for germ cell specification.
Normal germ cell development in the absence of Dppa3 . A, Gonads from E12.5 embryos (above: wild type; below: Dppa3 tm1WHT/Dppa3 tm1WHT) stained for alkaline phosphatase to reveal primordial germ cells. B, RT-PCR analysis of gene expression in wild-type and Dppa3 tm1WHT/Dppa3 tm1WHTadult ovaries. C,D, Dppa3 tm1WHT/Dppa3 tm1WHTtestis (C) and ovary (D) are histologically normal.
Moreover, this function is not readily ascribed to a gene closely related to Dppa3. We electronically searched the sequenced mouse genome for Dppa3 homologues. We identified several processed (intron-less) pseudogenes of Dppa3, but no functional, full-length homologue. As judged by RT-PCR analysis, the Dppa3 pseudogenes are not expressed in embryonic or adult tissues (data not shown).
Dppa3is a potent maternal factor
We found that Dppa3 plays an important role in early embryonic development as a maternal factor. While Dppa3 tm1WHT/Dppa3 tm1WHTmales were fully fertile, Dppa3 tm1WHT/Dppa3 tm1WHTfemales had small litters. This was true regardless of whether such females were crossed with Dppa3 tm1WHT/Dppa3 tm1WHT, Dppa3 tm1WHT/+ or wild type males (3.5 ± 1.5, 3.1 ± 2.1, or 3.0 ± 0.9 viable pups/litter, respectively). By contrast, Dppa3 tm1WHT/+ females of the same (mixed) genetic background had large litters when mated to Dppa3 tm1WHT/Dppa3 tm1WHTor Dppa3 tm1WHT/+ males (9.4 ± 3.5 or 10.1 ± 3.2 viable pups/litter, respectively).
We attribute the small litters from Dppa3 tm1WHT/Dppa3 tm1WHTmothers to abnormalities that manifest early in embryogenesis, during the cleavage stages of pre-implantation development. While nearly all embryos derived from Dppa3-deficient oocytes developed to the 2-cell or 4-cell stage (Figure 3A,3B,3C,3D), subsequent development was severely compromised in most such embryos (Figure 3E,3F). Some embryos derived from Dppa3-deficient oocytes failed to reach the 8-cell stage and instead showed evidence of compaction at the 4-cell stage. Other embryos derived from Dppa3-deficient oocytes cleaved to form 8 to 16 blastomeres, but failed to compact (Figure 3E,3F). These observations suggest that maternally supplied Dppa3 function is important in the cleavage stages of pre-implantation development.
Abnormal pre-implantation development of embryos derived from Dppa3 –deficient oocytes. A,C, Cultured 2-cell (A) and 4-cell (C) control embryos derived from wild-type matings. B,D, Cultured 2-cell (B) and 4-cell (D) embryos produced by crossing Dppa3 tm1WHT/Dppa3 tm1WHTfemales with wild-type males. E,F, E3.5 control embryos derived from wild-type matings have progressed to the blastocyst stage (E). By contrast, most E3.5 embryos produced by crossing Dppa3 tm1WHT/Dppa3 tm1WHTfemales with wild-type males have not progressed to the blastocyst stage and instead cleave abnormally and degenerate (F). G,H, Many embryos produced by crossing Dppa3 tm1WHT/Dppa3 tm1WHTfemales with Dppa3 +/+, Tg (Pou5f1 ΔPE-GFP)10WHT/Tg (Pou5f1 ΔPE-GFP)10WHTmales fail to develop normally beyond the 4-cell stage (G) but nonetheless express the Oct4-GFP marker (H).
Might maternal Dppa3 induce zygotic expression of Oct4/Pou5f1, which encodes a transcription factor that is crucial to pre-implantation development [10, 12]? To test this possibility, we crossed Dppa3 tm1WHT/Dppa3 tm1WHTfemales with Dppa3 +/+, Tg (Pou5f1 ΔPE-GFP)10WHT/Tg (Pou5f1 ΔPE-GFP)10WHTmales, the latter transmitting an Oct4-GFP transgene, with the Oct4 promoter driving expression of GFP. We retrieved the resulting embryos at the 2-cell stage and cultured them in vitro for 72 hours to monitor expression of the Oct4-GFP transgene. All such embryos were observed to express the fluorescent marker, regardless of the degree to which the embryos developed or failed to develop during the culture period (Figure 3G,3H). Thus, the poor development of many embryos derived from Dppa3-deficient oocytes cannot be attributed to the absence of zygotic expression of Oct4. Further analysis of the maternal-effect phenotype of Dppa3 should illuminate the molecular and biological context and consequences of the gene's activity.
Conclusions
We conclude that Dppa3 is not required for germ cell specification in mice. The identity of the mammalian gene or genes that program germ cells remains an open question. Dppa3 appears to function as a maternal factor, with an important role early in embryogenesis, during cleavage.
Methods
Generation of Dppa3-deficient animals
The Dppa3 targeting construct contained 1.3-kb and 3-kb segments of mouse genomic DNA, the former located 5' of Dppa3's translation initiation site and the latter located 3' of the termination codon (Figure 1). At the center of the construct was a 3-kb hygromycin-thymidine kinase selection cassette (Hygro-TK) flanked by two loxP direct repeats. V6.5 (C57BL/6 × 129/Sv)F1 ES cells [9] were transfected by electroporation, and recombined clones were selected in the presence of hygromycin (Invitrogen). Correctly targeted clones were identified by long-distance genomic PCR. The Hygro-TK cassette was removed via transient transfection of ES cells with a Cre-expressing plasmid in the presence of ganciclovir (Sigma). The final genomic structure of the resulting clones was verified by Southern analysis. Two independently targeted ES cell clones were microinjected into Balb/c blastocysts to generate chimeras. Animals used in this study were of a mixed C57BL/6 × 129/Sv genetic background.
Primers for PCR genotyping were as follows: p1 (5' TAG CCT GGG GGT AGA CTC GGC TGT AT 3'); p2 (5' AAC GAG AAG AGA AGG GAG GGC TTC 3'); and p3 (5' TCA CAT AAA TCT GGA TCG TTG TGC ATC 3'). The wild type allele gives rise to a PCR product of 304 bp with primers p1 and p2. The mutant allele (Dppa3 tm1WHT) gives rise to a PCR product of 492 bp with primers p1 and p3.
RNA isolation and RT-PCR
Total RNAs were isolated from mouse tissues, and expression of Dppa3, Oct4, and Gapd was assayed by RT-PCR, all as described previously[8].
Alkaline phosphatase staining of primordial germ cells
Gonads were dissected from wild type and Dppa3 tm1WHT/Dppa3 tm1WHTembryos on day 12.5 of gestation and stained for alkaline phosphatase as described previously [13].
Histology
Dissected adult testes and ovaries were fixed overnight in Bouin's solution, imbedded in paraffin, sectioned, and stained with hematoxylin and eosin.
Generation of Oct4-GFP transgenic animals
Mice bearing an Oct4-GFP transgene were generated by microinjection of a 14-kb Oct4Δ PE-GFP linear DNA fragment into C57Bl/6 × SJL F2 hybrid mouse eggs. This construct essentially reproduces the previously described GOF18Δ PE-lacZ construct [14] but contains a gene for enhanced green fluorescent protein (EGFP, Clontech) in place of lacZ at the ATG of Oct4. Mice bearing transgene Tg (Pou5f1 ΔPE-GFP)10WHTaccurately reproduced the previously reported Oct4 expression pattern [14] and were bred to generate Tg (Pou5f1 ΔPE-GFP)10WHT/Tg (Pou5f1 ΔPE-GFP)10WHThomozygous animals.
Isolation, culture and analysis of cleavage stage embryos
2-cell embryos were flushed from oviducts at E1.5 and cultured for up to 72 hours in microdrops of KSOM (Specialty Media) under light mineral oil (Squibb) with 5% CO2 in air. E3.5 embryos were flushed from uteri.
References
Extavour CG, Akam M: Mechanisms of germ cell specification across the metazoans: epigenesis and preformation. Development. 2003, 130: 5869-5884. 10.1242/dev.00804.
Wylie C: Germ cells. Curr Opin Genet Dev. 2000, 10: 410-413. 10.1016/S0959-437X(00)00105-2.
McLaren A: Primordial germ cells in the mouse. Dev Biol. 2003, 262: 1-15. 10.1016/S0012-1606(03)00214-8.
Lawson KA, Dunn NR, Roelen BA, Zeinstra LM, Davis AM, Wright CV, Korving JP, Hogan BL: Bmp4 is required for the generation of primordial germ cells in the mouse embryo. Genes Dev. 1999, 13: 424-436.
Saitou M, Barton SC, Surani MA: A molecular programme for the specification of germ cell fate in mice. Nature. 2002, 418: 293-300. 10.1038/nature00927.
Saitou M, Payer B, Lange UC, Erhardt S, Barton SC, Surani MA: Specification of germ cell fate in mice. Philos Trans R Soc Lond B Biol Sci. 2003, 358: 1363-1370. 10.1098/rstb.2003.1329.
Sato M, Kimura T, Kurokawa K, Fujita Y, Abe K, Masuhara M, Yasunaga T, Ryo A, Yamamoto M, Nakano T: Identification of PGC7, a new gene expressed specifically in preimplantation embryos and germ cells. Mech Dev. 2002, 113: 91-94. 10.1016/S0925-4773(02)00002-3.
Bortvin A, Eggan K, Skaletsky H, Akutsu H, Berry DL, Yanagimachi R, Page DC, Jaenisch R: Incomplete reactivation of Oct4-related genes in mouse embryos cloned from somatic nuclei. Development. 2003, 130: 1673-1680. 10.1242/dev.00366.
Eggan K, Akutsu H, Loring J, Jackson-Grusby L, Klemm M, Rideout W. M., 3rd, Yanagimachi R, Jaenisch R: Hybrid vigor, fetal overgrowth, and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc Natl Acad Sci U S A. 2001, 98: 6209-6214. 10.1073/pnas.101118898.
Pesce M, Scholer HR: Oct-4: gatekeeper in the beginnings of mammalian development. Stem Cells. 2001, 19: 271-278. 10.1634/stemcells.19-4-271.
Pesce M, Wang X, Wolgemuth DJ, Scholer H: Differential expression of the Oct-4 transcription factor during mouse germ cell differentiation. Mech Dev. 1998, 71: 89-98. 10.1016/S0925-4773(98)00002-1.
Nichols J, Zevnik B, Anastassiadis K, Niwa H, Klewe-Nebenius D, Chambers I, Scholer H, Smith A: Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell. 1998, 95: 379-391. 10.1016/S0092-8674(00)81769-9.
Ginsburg M, Snow MH, McLaren A: Primordial germ cells in the mouse embryo during gastrulation. Development. 1990, 110: 521-528.
Yeom YI, Fuhrmann G, Ovitt CE, Brehm A, Ohbo K, Gross M, Hubner K, Scholer HR: Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development. 1996, 122: 881-894.
Acknowledgements
A.B. was a Leukemia & Lymphoma Society Special Fellow. Supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
AB conducted molecular biological, ES cell culture and embryological studies, and co-wrote the manuscript. MG carried out blastocyst injections. ML assisted in mouse and embryological studies. DP coordinated the study and co-wrote the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Bortvin, A., Goodheart, M., Liao, M. et al. Dppa3 / Pgc7 / stella is a maternal factor and is not required for germ cell specification in mice. BMC Dev Biol 4, 2 (2004). https://doi.org/10.1186/1471-213X-4-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-213X-4-2