Abstract
Leaf rolling is one of the most significant symptoms of drought stress in plant. Previously, we identified a dominant negative mutant, termed rolled and erect 1 (hereafter referred to rel1-D), regulating leaf rolling and erectness in rice. However, the role of REL1 in drought response is still poorly understood. Here, our results indicated that rel1-D displayed higher tolerance to drought relative to wild type, and the activity of superoxide dismutase (SOD) and drought responsive genes were significantly up-regulated in rel1-D. Moreover, our results revealed that rel1-D was hypersensitive to ABA and the expression of ABA associated genes was significantly increased in rel1-D, suggesting that REL1 likely coordinates ABA to regulate drought response. Using the RNA-seq approach, we identified a large group of differentially expressed genes that regulate stimuli and stresses response. Consistently, we also found that constitutive expression of REL1 alters the expression of biotic and abiotic stress responsive genes by the isobaric tags for relative and absolute quantification (iTRAQ) analysis. Integrative analysis demonstrated that 8 genes/proteins identified by both RNA-seq and iTRAQ would be the potential targets in term of the REL1-mediated leaf morphology. Together, we proposed that leaf rolling and drought tolerance of rel1-D under normal condition might be caused by the endogenously perturbed homeostasis derived from continuous stressful dynamics.
Similar content being viewed by others
Background
Crop yield is adversely challenged by drought stress, one of the most major environmental stresses, of which the occurrence and severity are both increased due to the recent climate change, inadequate water supply and world population growth worldwide. Therefore, improving drought tolerance of crop is an important objective to overcome such issue and provide enough world food (Yamaguchi-Shinozaki and Shinozaki, 2006). One of the most significant symptoms of drought stress in plant is the leaf rolling. Plant leaf generally is polarized along the adaxial-abaxial axis (Itoh et al. 2005), thereby generating two types of leaf forms under unfavorable conditions: abaxially leaf rolling and adaxially leaf rolling. Moderate leaf rolling promotes rice yield by increasing the photosynthetic efficiency and reducing the transpiration (Lang et al. 2004; Zhang et al. 2009; Zou et al. 2011). In addition, moderate leaf rolling also facilitates the survival and development of plant under stress conditions (Kadioglu et al. 2012). Therefore, manipulation of adaxial and abaxial leaf rolling would be one of the most important strategies to increase the rice productivity and tolerance to drought and other stresses in coming years (Zou et al. 2011).
Generally, leaf rolling is largely regulated by several factors: hormones (Fujino et al. 2008; Lee et al. 2006), metabolic changes (Talukdar and Talukdar, 2014) and formation of specific cells, such as bulliform cell (Fang et al. 2012; Xiang et al. 2012). To date, more than 30 genes responsible for the rice leaf rolling have been identified. For example, Abaxially Curled Leaf 1 (ACL1), Abaxially Curled Leaf 2 (ACL2) (Li et al. 2010) and Rice outermost cell-specific gene 5 (Roc5) (Zou et al. 2011) are involved in abaxial leaf rolling, whereas SEMI-ROLLED LEAF 1 (SRL1) (Xiang et al. 2012), ROLLED LEAF 9 (RL9)/SHALLOT-LIKE1 (SLL1) (Zhang et al. 2009; Yan et al. 2008), OsAGO7 (Shi et al. 2007), Rolling-leaf 14 (RL14) (Fang et al. 2012), R2R3-MYB transcription factor (OsMYB103L) (Yang et al. 2014), ADAXIALIZED LEAF 1 (ADL1) (Hibara et al. 2009) and Narrow and Rolled Leaves 1 (NRL1) (Wu et al. 2010) participates in adaxial leaf rolling. In addition, other regulators, such as CONSTITUTIVELY WILTED1 (COW1)/NAL7 (Fujino et al. 2008), has also been well-characterized in term of their involvements in leaf rolling. Even thought, it is still poorly understood whether these leaf rolling genes participate in stress response, in particular drought.
To cope with drought stress, plants have been evolved a sophisticated adaptation mechanisms to increase their chance of survival through coordinating the expression of drought responsive genes, which retarded leaf rolling, wilting, and loss of chlorophyll via ABA-dependent or independent pathways (Yamaguchi-Shinozaki and Shinozaki, 2006; Weiner et al. 2010; Cutler et al. 2010). Under drought stress, ABA accumulates in plant cells and stimulates the core ABA signaling transduction through the ABA receptors PYR/PYL/RCAR, Protein Phosphatases 2C (PP2Cs), and subclass III SNF 1-RELATED PROTEIN KINASE 2 (SnRK2) protein kinases (Weiner et al. 2010). For example, loss of the rice subclass-I and -II SnRK2s (OsSAPK2) confers the more sensitivity to drought stress and insensitive to ABA (Lou et al. 2017). Over-expression of OsDT11 enhanced drought tolerance thought the ABA signaling pathway in rice (Li et al. 2017). In contrast, a group of regulators modulate drought tolerance by ABA-independent pathway. Fox example, OsMADS50 was markedly induced by low water-deficit treatment but not ABA, and the osmads50 knockout mutant exhibited significant delays in flowering compared with the WT, indicating OsMADS50 had an ABA-independent and positive role in drought response (Du et al. 2018).
Here, we were particularly interested in elucidating the relationship between the rel1-D mediated leaf rolling and drought stress in rice. Our results suggested that up-regulation of REL1 confers more drought tolerance by eliminating the ROS level and triggers the ABA response. Similarly, transcriptomic and proteomic profiling of rel1-D and wild type also revealed that most of the differentially expressed genes (DEGs) and differentially expressed proteins (DEPs) are mainly involved in metabolic changes and stress responses. Collectively, we proposed that leaf rolling of rel1-D is caused by the altered dynamics of stress response endogenously.
Methods
Plant materials and growth conditions
All plants (wild type and rel1-D mutants) used in this study were derived from our previous study (Chen et al. 2015). All rice seeds in this study were propagated in the paddy field in Guangzhou, China. For laboratory work, rice plants were grown in a greenhouse under a16-h-light/8-h-dark cycle at 30 °C. No significant differences were observed when plants were grown in the greenhouse compared to the paddy field.
Drought response assay
For drought tolerance test, wild type and rel1-D seedlings were grown in soil for 2 months, and then treated by withholding water for 14 days, followed by recovery irrigation for 7 days. Leaves from 1-month-old seedling were selected for dark-induced leaf senescence and ABA treatment. Seeds of wild type and rel1-D were germinated at 37 °C, and then transferred into 1/2 MS medium for 14 days at 30 °C under a16-h-light/8-h-dark cycle. Seedlings from 3rd leaf stage were selected for PEG treatment: 1) plants treated with 0%, 10%, 20%, 30% 40% PEG4000 for 24 h, and the rolling index was measured. The rolling index was measured as previous study (Shi et al. 2007). 2) Plants treated with 20% PEG4000 with 5 time courses, including 0 h, 3 h, 6 h, 12 h and 24 h. The SOD activity was measured as previous study (Zhang et al. 2005).
ABA treatment and chlorophyll content measurement
Leaves were treatment with distillation-distillation water with 0 μM or 20 μM ABA at 28 °C in darkness for 5 days. Chlorophyll content was measured as followed: leaves were milled in 95% ethanol; homogenate solution adjusted to 50 ml final volume; measure the absorbance at 652 nm. Calculate the chlorophyll content according to the following formula: chlorophyll content (mg/g) = (A652 × V)/ (34.5 × m); V was the final volume of homogenate solution, and m was the weight of leaves.
RNA extraction and quantitative real-time PCR
Total RNA was extracted using the RNeasy Plant Mini Kit (Qiagen) according to the manufacturer’s instructions. The first strand of cDNA was synthesized using TransScript First-Strand cDNA Synthesis SuperMix (TransGen Biotech) and quantitative real-time polymerase chain reaction (qRT-PCR) was performed as previously described (Chen et al. 2015). The relative expression level of a target gene was normalized to that of rice UBC. All primers used in qRT-PCR are listed in Additional file 1: Table S12.
Sequence reads mapping
Raw data (raw reads) of fastq format were firstly processed through in-house perl scripts. In this step, the clean data (clean reads) were obtained by removing reads containing adapter, reads containing poly-N and low quality reads from raw data. At the same time, quality parameters of clean data including Q20, Q30, GC-content and sequence duplication level were used for data filtering. All the succeeding analyses were carried out using high quality clean data. Reference genome and gene model annotation files were downloaded from The MSU Rice Genome Annotation Project Database website at http://rice.plantbiology.msu.edu. An index of the reference genome was built using Bowtie2 v2.2.5 (Langmead et al. 2009) and paired-end clean reads were aligned to the reference genome using TopHat v2.0.14 (Trapnell et al. 2010). TopHat was chosen as the mapping tool, because it can generate a database of splice junctions based on the gene model annotation file, and thus give a better mapping result than other non-splice mapping tools.
Quantification and differential expression analysis of transcripts
HTSeq v0.6.1 (http://www-huber.embl.de/users/anders/HTSeq) was used to count the reads numbers mapped to each transcript. The parameter FPKM (expected number of Fragments per kilo-base of transcript sequence per Millions base pairs sequenced) was used to quantify transcripts expression. FPKM was calculated based on the mapped transcript fragments, transcript length and sequencing depth. Currently, this is the most commonly used method for estimating transcript expression (Trapnell et al. 2010). Differential expression analysis of two conditions/groups was performed using the DESeq R package (1.10.1) (Anders and Huber, 2010). DESeq provides statistical routines to determine differential expression in digital gene expression data using a model based on the negative binomial distribution. The resulting q-values (p-adjusted) were adjusted using the Benjamini and Hochberg’s approach for controlling the false discovery rate (Anders and Huber, 2010). Genes with an adjusted |log2 (FC)| ≥ 1 and FDR < 0.05 were assigned as differentially expressed.
iTRAQ assay and data analysis
The iTRAQ assay was performed by the BGI Company. Briefly, about 100 μg protein was subjected to the LC-ESI-MS/MS analysis based on the Triple TOF 5600. The proteins identification was performed by using Mascot search engine (Matrix Science, London, UK; version 2.3.02). For protein quantitation, it was required that a protein contains at least two unique spectra. The quantitative protein ratios were weighted and normalized by the median ratio in Mascot. We used ratios of p-values < 0.05, and fold changes > 1.5 or < 0.67 was considered as significant. Functional annotations of the proteins were conducted using Blast2GO program against the non-redundant protein database (https://www.ncbi.nlm.nih.gov/protein/). The keg database (http://www.genome.jp/kegg/) and the COG database (http://www.ncbi.nlm.nih.gov/COG/) were used to classify and group these identified proteins.
Results
Response of rel1-D to drought stress
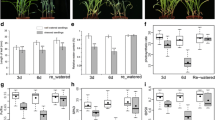
Leaf rolling generally leads to reduction of water loss, and thereby enhances tolerance to multiple stresses (Lang et al. 2004; Zhang et al. 2009; Zou et al. 2011). Therefore, we were specifically interested in investigating the involvement of REL1 in stress response. To address this issue, WT and rel1-D seedlings were undergone the drought assay. Phenotypic analysis showed that almost all of the WT plants displayed severe growth retardation and wilting while the rel1-D plants exhibited less abnormal phenotypes by withholding irrigation for 14 days, and then the rel1-D but not the WT was recovered upon re-watering for 7 days (Fig. 1a and b). To further explore the drought tolerance of rel1-D, leaves from WT and rel1-D were treated by polyethylene glycol (PEG) 4000 at different concentrations. Our results indicated that leaves of rel1-D were much more insensitive to the treatment than those of WT (Fig. 1c). The superoxide dismutase (SOD) activity is generally used as an important indicator for drought tolerance. Therefore, the SOD activity of WT and rel1-D that treated by 20% PEG4000 was measured at 5 time courses. Our results demonstrated that the change pattern of SOD activity was similar between WT and rel1-D, but the corresponding levels were higher in rel1-D than those in WT (Fig. 1d), suggesting that up-regulating REL1 suppressed the boost of reactive oxygen species (ROS) under drought stress. To further gain insight into the REL1-mediated drought resistance, we then evaluated the expression of drought-responsive marker genes. Our results demonstrated that OsDT11, OsSAPK2, OsMYB2, OsDREB1A and OsbHLH148, functioning as positive regulators in drought response (Lou et al. 2017; Li et al. 2017; Chen et al. 2008; Dubouzet et al. 2003; Seo et al. 2011; Yang et al. 2012), were significantly up-regulated in the rel1-D (Fig. 1e-i). Consistently, the similar patterns of above marker genes were also found in the RNA-seq result (Additional file 2: Table S1). These results suggested that enhanced the expression of REL1 leads to the substantial higher expression of drought-responsive genes endogenously, eventually resulting in the tolerance of rel1-D to drought.
Response of rel1-D to drought stress. a, Phenotype of WT and rel1-D with drought treatment. Two-month-old seedlings were used, bars = 10 cm. b, Survival rate of WT and rel1-D were derived from the (A). c, Third leaf stage seedling of WT and rel1-D were treated by different concentration of PEG, bars = 5 mm. d, Measurement of SOD activity in wild type and rel1-D. E-I, Expression of drought-responsive genes in wild type and rel1-D. b and e-i, Data were presented as mean ± S.E., * p-value < 0.05, ** p-value < 0.01, two-tailed, two sample Student’s t test
Taking into account that REL1 is an abaxial leaf rolling regulator, we wondered whether other abaxial leaf rolling associated genes is also involved in the response to drought stress. To address this issue, we examined their expression pattern in WT and rel1-D at different time courses by PEG treatment. Surprisingly, REL1 exhibited normal expression during the treatment in WT, implying it is not related with drought stress. However, the expression level of REL1 was substantial higher in rel1-D than that in WT even it was attenuated at 3 h but gradually increased afterward in rel1-D (Additional file 3: Figure S1A). The ACL1 was not changed by the PEG treatment in both WT and rel1-D at 24 h (Additional file 3: Figure S1B), suggesting this leaf rolling gene is not involved in drought response. The ACL2 was significantly enhanced at 24 h in WT, implying it might participate in drought stress. Notably, ACL2 was retarded from 3 h to 12 h but increased to the normal level at 24 h in rel1-D with the similar level of that in WT (Additional file 3: Figure S1C), suggesting that constitutive expression of REL1 would repress ACL2 during drought stress. The Roc5 was down-regulated at 12 h and increased to the normal level at 24 h in WT, but gradually decreased in rel1-D during treatment (Additional file 3: Figure S1D), demonstrating Roc5 was suppressed by up-regulation of REL1 under drought stress. Taken together, we proposed that rel1-D mediated drought tolerance independent on the three leaf rolling genes, but ACL2 may be also involved in drought stress in a distinct pathway.
Response of rel1-D to ABA
Since ABA both regulates drought response and leaf senescence (Lou et al. 2017; Li et al. 2017; Chen et al. 2008; Dubouzet et al. 2003; Seo et al. 2011; Yang et al. 2012), we wondered whether REL1 was also involved in the response to ABA. To address this issue, leaves from WT and rel1-D were treated by 0 and 20 μM ABA from 1 to 5 days by the dark-induced leaf senescence assay, respectively. Under the ABA treatment, rel1-D displayed early senescence phenotype and rapid degradation of chlorophyll (Chl) rather than that in WT (Fig. 2a and b), indicating it was hypersensitive to ABA. Notably, REL1 is significantly repressed by the ABA treatments (Fig. 2c). Therefore, we concluded that boosting REL1 accelerates ABA-induced leaf senescence. It was worthy to figure out that rel1-D leaves started to turn yellowish while the wild-type leaves still remained green at the 3 days. Five days after dark treatment, the Chl content in the rel1-D leaves was about 2-fold less than in the wild-type leaves (Fig. 2a and b). To further explore the relationship between REL1 and ABA-induced senescence, we detected the expression of the senescence marker genes Osl85 and SGR, and found that they were induced by dark-treatment assay as previously reported (Lee et al. 2001; Jiang et al. 2011). Notably, their levels were significantly higher in rel1-D as compared to WT, as well as with ABA rather than without ABA treatment (Fig. 2d and e). Therefore, we proposed that overexpression of REL1 also triggers the natural leaf senescence and ABA would accelerate this response.
Response of rel1-D to ABA. a, Response of rel1-D to ABA during dark-induced leaf senescence. Leaves from 1-month-old seedling of WT and rel1-D were incubated with 20 μM ABA for 1 to 5 days, bars = 5 cm. b, Chlorophyll content of WT and rel1-D by ABA treatments. c, Expression of REL1 in response to ABA treatment. d, Expression of Osl85 in response to ABA treatment. e, Expression of SGR in response to ABA treatment. B-E, Data were presented as mean ± S.E.. c-e, Multiple comparisons, Duncan, p-value < 0.01
Transcriptomic profiling of the rel1-D mutant
To further investigate the regulatory mechanism of REL1-mediated leaf rolling and bending, we performed a RNA-seq analysis with the leaves of rel1-D mutants and wild type plants at the tillering stage, since the most obvious leaf rolling and bending phenotypes were occurred at this stage. In total, 487 differentially expressed genes (DEGs) were identified with the stringent criteria (|log2 (FC)| ≥ 1, and FDR < 0.05). To verify these DEGs, 10 randomly selected DEGs were detected by the qRT-PCR, and their trends were similar as RNA-seq (Additional file 4: Figure S2), indicating that the RNA-seq result was qualified for following study. Of these DEGs, 247 and 240 transcripts were up-regulated or down-regulated in rel1-D as compared to wild type, respectively (Additional file 5: Table S2). Even the statistical examination of most BR genes was not significant, there were still two BR signaling genes were induced in rel-D (Additional file 6: Figure S3), suggesting that BR was also involved in regulating the leaf morphology of rel1-D.Notably, the ABA pathway was significantly changed (Additional file 7: Figure S4), further supporting the note that ABA was involved in rel1-D mediated leaf morphology and response. To further characterize these DEGs, we performed the gene ontology (GO) enrichment analysis. In respect to the up-regulated DEGs, they were significantly assigned to certain cellular component GO terms, including cell wall, external encapsulating structure and cell periphery (Fig. 3a; Additional file 8: Table S3). In terms of the molecular function GO term, these DEGs were mainly associated with hydrolase activity, transcription factor activity and catalytic activity (Fig. 3b; Additional file 9: Table S4). In respect to the biological process GO term, these DEGs were significantly involved in multiple stimuli and stresses (Fig. 3c; Additional file 10: Table S5). Investigation of the down-regulated DEGs showed that they were also significantly associated with vacuole and endoplasmic reticulum (Fig. 3d; Additional file 11: Table S6), catalytic activity and transport activity (Fig. 3e; Additional file 12: Table S7), and response to various stimuli and metabolic processes (Fig. 3f; Additional file 13: Table S8). Taken together, our results suggested that boosting REL1 apparently perturbed the homeostasis of stress dynamics in specific organelles, such as cell wall and vacuole, eventually leading to the abnormal leaf morphology.
Gene Ontology (GO) analysis for differentially expressed genes (DEGs). a, Significant cellular component GO terms of the up-regulated DEGs. b, Significant molecular function GO terms of the up-regulated DEGs. c, Significant biological process GO terms of the up-regulated DEGs. d, Significant cellular component GO terms of the down-regulated DEGs. e, Significant molecular function GO terms of the down-regulated DEGs. f, Significant biological process GO terms of the down-regulated DEGs
Proteomics analysis of rel1-D mutant
To further analyze the function of REL1 in leaf morphology at protein level, we then performed the isobaric tags for relative and absolute quantitation (iTRAQ) analysis on the above materials. In total, 3657 peptides were identified (Additional file 14: Table S9). Using the p-value < 0.05 and fold change > 1.5 or < 0.67 as significant cutoff, 20 and 29 differentially expressed proteins (DEPs) were up-regulated or down-regulated, respectively. To verify these DEPs, 10 randomly selected DEPs were detected by the qRT-PCR, and their changing pattern were similar as iTRAQ (Additional file 15: Figure S5), indicating that the iTRAQ result was qualified for following study. Subsequently, GO enrichment analysis of these DEPs revealed that they were enriched for cellular component GO terms related to inter- and intra- cellular organelles, ribosome and plastid envelope (Fig. 4a; Additional file 16: Table S10). In respect to the molecular function GO term, significant enrichments were found in amylase activity, transport activity and ion binding (Fig. 4b; Additional file 16: Table S10). Regarding the biological process, the DEPs were grouped into multiple catabolic processes, response to water transport and stimuli (Fig. 4c; Additional file 16: Table S10). It was worthy to mention that 24 out of 49 DEPs were annotated or predicted to be plastid-localized proteins (Additional file 17: Table S11), while REL1 was previously implicated as plastid protein (Chen et al. 2015). Notably, 4 DEPs have been reported to regulate stress response (Table 1), including OsPIP1;1, OsPIP1;2, SUS2 and OsGLP8–7 (Liu et al. 2013; Mosa et al. 2012; Xiao et al. 2014; Breen and Bellgard 2010), further supporting the notion that rel1-D actives the endogenous stress responses. Collectively, we proposed that REL1 may coordinate the chloroplast DEPs to regulate leaf morphology by altering the metabolic process and stressful dynamics.
Integrative analysis of transcriptome and proteome for rel1-D
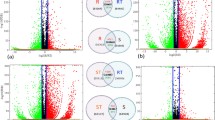
Integrative analysis of transcriptome and proteome may provide new insights into the identification of interest key genes. In total, 3575 co-expressed genes and proteins were identified (Fig. 5a). Then only 2 co-expressed genes/proteins, LOC_Os05g09740 and LOC_Os02g37654, were found between the 487 DEGs and 49 DEPs (Fig. 5b). To broad view the genome-wide change, 246 DEPs were screened by a less stringent criteria (with fold change > 1.5 or < 0.67), and 234 out of these low criteria DEPs (DEPs-low) were identified in the RNA-seq data (Fig. 5c). Integrating the DEGs and the 246 DEPs-low, there were 8 genes/proteins found in each other (Fig. 5d). These 8 genes modulated stresses response and had distinct expression pattern, of which 5 genes showed similar trends in transcription and translation level but 3 genes exhibited opposite trends (Table 2). Taken together, we proposed that the molecular mechanism underlying REL1-mediated leaf phenotype was likely different between transcriptional and post-translation levels.
Correlation between the transcript and protein levels. a, Venn analysis of all expressed mRNA and all expressed proteins. b, Venn analysis of differentially expressed genes (DEGs) and differentially expressed proteins (DEPs) with stringent criteria (p-value < 005). c, Venn analysis of all expressed transcripts and differentially expressed proteins with less stringent criteria. d, Venn analysis of differentially expressed transcripts and differentially expressed proteins with less stringent criteria
Discussion
During plant growth and development, leaf rolling is an adaptive rather than passive response to the abiotic and biotic stresses in plants (Kadioglu et al. 2012; Kadioglu and Terzi, 2007). Our previous study has implicated that REL1 positively regulates leaf rolling through altering the profile of bulliform cells and leaf bending by coordinated expression of BR related genes (Chen et al. 2015). However, the biological function of REL1 and the relevant regulatory mechanism still remains to be further elucidated. Here, we further explored its regulatory role in leaf morphology by transcriptomic and proteomic analyses, as well as co-expression analysis. Our results may provide a new insight into the REL1-mediated leaf morphology in rice.
Advance studies have also been evident that water deficiency is one of the most major reasons for the formation of leaf rolling, which effectively reduces transpiration and thus is potentially useful in drought tolerance (Kadioglu et al. 2012). As expected, up-regulation of drought resistance marker genes and tolerance to drought treatment in rel1-D suggested that REL1 positively participates in drought resistance. In addition, our result also indicated that REL1-mediated drought tolerance might be integrated the ABA pathway. Therefore, we proposed that REL1 might coordinate ABA pathway to regulate drought tolerance. Comparative analysis of REL1 and other abaxial leaf rolling genes demonstrated that they might play opposite roles in response to drought tolerance, suggesting that they might also function independently during the formation of leaf rolling under stresses. Surprisingly, we also found that REL1 is involved in leaf senescence and ABA response, which would be another interesting issue to be addressed regarding the biological function of REL1.
Although REL1 encodes an unknown but species-specific protein, genome-wide profiling may facilitate our understanding on the biological function of REL1 in determining leaf morphology. Transcriptomic and proteomic profiling demonstrated that the change of leaf morphology in rel1-D was highly associated with the metabolic changes and stresses response. However, it is still unclear whether REL1 directly or indirectly catalyzes specific primary and/or secondary metabolism so that generating stressful dynamics endogenously. Taking into account the chloroplast localization of REL1, the chloroplast DEGs may be functionally associated with REL1. Unexpectedly, only 4 DEGs were grouped into chloroplast GO term under the stringent statistical criteria. Differing from the DEGs, almost half of the DEPs were chloroplast localized proteins. Combining the integrative analyses of transcriptom and proteasome, two possibilities were proposed: 1) REL1 regulates leaf morphology at the post-transcriptional level independent on the chloroplast genes; 2) REL1 regulates leaf morphology at the post-translation levels through direct or indirect regulation of chloroplast proteins. These issues would be quite interesting for further study. Expectedly, a large part of the DEGs and DEPs were both enriched into the stress response GO term, further demonstrating that up-regulation of REL1 generates endogenous stresses for the plant. Several genes/proteins identified in both DEGs and DEPs would be the candidates for future study in terms of leaf rolling and bending, particularly the response to multiple stresses.
Interestingly, a recent study reported that another gene REL2, encoding a DUF630 and DUF632 domains containing protein, regulates leaf rolling and bending as well (Yang et al. 2016). It was worthy to figure out that the gene LOC_Os06g44610, significantly down-regulated in rel1-D, also encodes a membrane associated DUF588 domain containing protein, suggesting a possible role of DUF family genes in the leaf development. Although REL1 has a functional relationship with REL2, it is still challenged by: 1) the distinct localization pattern of these two proteins since REL2 is a plasma membrane localized protein while REL1 is a chloroplast protein; 2) REL2 likely regulates bulliform cell through auxin pathway while REL1might be much more related to BR pathway. In addition, much more transcriptomics datasets would benefit the further understanding of the regulatory network between REL1 and other leaf development genes by co-expression analysis. Meanwhile, genetic analysis by constructing double (or multiple) mutant would also benefit our knowledge of the correlation among these leaf morphology genes.
Conclusions
Leaf rolling is generally related to stress response. By the drought assay, we identified the involvement of rel1-D in such biological process. Furthermore, the RNA-seq and iTRAQ-based profiling of WT and rel1-D also indicated that enhanced expression of REL1 resulted in the alteration of stress responsive genes and hormone genes. These genes might be the potential targets for extending our understanding of the REL1-mediated leaf morphology, and would also be valuable resources for further exploring and/or genetic engineering the molecular mechanism of stress tolerance in rice.
One sentence summary
REL1 is involved in drought tolerance, leaf senescence and hormones sensitivity by altering the profiles of relevant response genes in rice.
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106
Breen J, Bellgard M (2010) Germin-like proteins (GLPs) in cereal genomes: gene clustering and dynamic roles in plant defence. Funct Integr Genomics 10:463–476
Chen JQ, Meng XP, Zhang Y, Xia M, Wang XP (2008) Over-expression of OsDREB genes lead to enhanced drought tolerance in rice. Biotechnol Lett 30:2191–2198
Chen Q, Xie Q, Gao J, Wang W, Sun B, Liu B, Zhu H, Peng H, Zhao H, Liu C, Wang J, Zhang J, Zhang G, Zhang Z (2015) Characterization of Rolled and Erect Leaf 1 in regulating leave morphology in rice. J Exp Bot 66:6047–6058
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Du H, Huang F, Wu N, Li X, Hu H, Xiong L (2018) Integrative regulation of drought escape through ABA-dependent and -independent pathways in rice. Mol Plant 11:584–597
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763
Fang L, Zhao F, Cong Y, Sang X, Du Q, Wang D, Li Y, Ling Y, Yang Z, He G (2012) Rolling-leaf14 is a 2OG-Fe (II) oxygenase family protein that modulates rice leaf rolling by affecting secondary cell wall formation in leaves. Plant Biotechnol J 10:524–532
Fujino K, Matsuda Y, Ozawa K, Nishimura T, Koshiba T, Fraaije MW, Sekiguchi H (2008) NARROW LEAF 7 controls LEAF shape mediated by auxin in rice. Mol Gen Genomics 279:499–507
Hibara K, Obara M, Hayashida E, Abe M, Ishimaru T, Satoh H, Itoh J, Nagato Y (2009) The ADAXIALIZED LEAF1 gene functions in LEAF and embryonic pattern formation in rice. Dev Biol 334:345–354
Itoh J, Nonomura K, Ikeda K, Yamaki S, Inukai Y, Yamagishi H, Kitano H, Nagato Y (2005) Rice plant development: from zygote to spikelet. Plant Cell Physiol 46:23–47
Jiang H, Chen Y, Li M, Xu X, Wu G (2011) Overexpression of SGR results in oxidative stress and lesion-mimic cell death in rice seedlings. J Integr Plant Biol 53:375–387
Kadioglu A, Terzi R (2007) A dehydration avoidance mechanism: leaf rolling. Bot Rev 73:290–302
Kadioglu A, Terzi R, Saruhan N, Saglam A (2012) Current advances in the investigation of leaf rolling caused by biotic and abiotic stress factors. Plant Sci 182:42–48
Lang Y, Zhang Z, Gu X, Yang J, Zhu Q (2004) Physiological and ecological effects of crimpy leaf character in rice (Oryza sativa L.) II. Photosynthetic character, dry mass production and yield forming. Acta Agron Sin 30:883–887
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Lee RH, Wang CH, Huang LT, Chen SC (2001) Leaf senescence in rice plants: cloning and characterization of senescence up-regulated genes. J Exp Bot 52:1117–1121
Lee Y, Park J, Im K, Kim K, Lee J, Lee K, Park JA, Lee TK, Park DS, Yang JS, Kim D, Lee S (2006) Arabidopsis leaf necrosis caused by simulated acid rain is related to the salicylic acid signaling pathway. Plant Physiol Biochem 44:38–42
Li L, Shi ZY, Li L, Shen GZ, Wang XQ, An LS, Zhang JL (2010) Overexpression of ACL1 (Abaxially Curled Leaf 1) increased bulliform cells and induced abaxial curling of leaf blades in rice. Mol Plant 3:807–817
Li X, Han H, Chen M, Yang W, Liu L, Li N, Ding X, Chu Z (2017) Overexpression of OsDT11, which encodes a novel cysteine-rich peptide, enhances drought tolerance and increases ABA concentration in rice. Plant Mol Biol 93:21–34
Liu C, Fukumoto T, Matsumoto T, Gena P, Frascaria D, Kaneko T, Katsuhara M, Zhong S, Sun X, Zhu Y, Iwasaki I, Ding X, Calamita G, Kitagawa Y (2013) Aquaporin OsPIP1;1 promotes rice salt resistance and seed germination. Plant Physiol Biochem 63:151–158
Lou D, Wang H, Liang G, Yu D (2017) OsSAPK2 confers abscisic acid sensitivity and tolerance to drought stress in rice. Front Plant Sci 8:993
Mosa KA, Kumar K, Chhikara S, Mcdermott J, Liu Z, Musante C, White JC, Dhankher OP (2012) Members of rice plasma membrane intrinsic proteins subfamily are involved in arsenite permeability and tolerance in plants. Transgenic Res 21:1265–1277
Seo JS, Joo J, Kim MJ, Kim YK, Nahm BH, Song SI, Cheong JJ, Lee JS, Kim JK, Choi YD (2011) OsbHLH148, a basic helix-loop-helix protein, interacts with OsJAZ proteins in a jasmonate signaling pathway leading to drought tolerance in rice. Plant J 65:907–921
Shi Z, Wang J, Wan X, Shen G, Wang X, Zhang J (2007) Over-expression of rice OsAGO7 gene induces upward curling of the leaf blade that enhanced erect-leaf habit. Planta 226:99–108
Talukdar D, Talukdar T (2014) Leaf rolling and stem fasciation in grass pea (Lathyrus sativus L.) mutant are mediated through glutathione-dependent cellular and metabolic changes and associated with a metabolic diversion through cysteine during phenotypic reversal. Biomed Res Int 2014:479180
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515
Weiner JJ, Peterson FC, Volkman BF, Cutler SR (2010) Structural and functional insights into core ABA signaling. Curr Opin Plant Biol 13:495–502
Wu C, Fu Y, Hu G, Si H, Cheng S, Liu W (2010) Isolation and characterization of a rice mutant with narrow and rolled leaves. Planta 232:313–324
Xiang JJ, Zhang GH, Qian Q, Xue HW (2012) Semi-rolled leaf1 encodes a putative glycosylphosphatidylinositol-anchored protein and modulates rice leaf rolling by regulating the formation of bulliform cells. Plant Physiol 159:1488–1500
Xiao X, Tang C, Fang Y, Yang M, Zhou B, Qi J, Zhang Y (2014) Structure and expression profile of the sucrose synthase gene family in the rubber tree: indicative of roles in stress response and sucrose utilization in the laticifers. FEBS J 281:291–305
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yan S, Yan CJ, Zeng XH, Yang YC, Fang YW, Tian CY, Sun YW, Cheng ZK, Gu MH (2008) ROLLED LEAF 9, encoding a GARP protein, regulates the leaf abaxial cell fate in rice. Plant Mol Biol 68:239–250
Yang A, Dai X, Zhang W (2012) A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63:2541–2556
Yang C, Li D, Liu X, Ji C, Hao L, Zhao X, Li X, Chen C, Cheng Z, Zhu L (2014) OsMYB103L, an R2R3-MYB transcription factor, influences leaf rolling and mechanical strength in rice (Oryza sativa L.). BMC Plant Biol 14:158
Yang SQ, Li WQ, Miao H, Gan PF, Qiao L, Chang YL, Shi CH, Chen KM (2016) REL2, a gene encoding an unknown function protein which contains DUF630 and DUF632 domains controls leaf rolling in rice. Rice 9:37
Zhang GH, Xu Q, Zhu XD, Qian Q, Xue HW (2009) SHALLOT-LIKE1 is a KANADI transcription factor that modulates rice leaf rolling by regulating leaf abaxial cell development. Plant Cell 21:719–735
Zhang Q, Li Y, Pang C, Lu C, Wang B (2005) NaCl enhances thylakoid-bound SOD activity in the leaves of C3 halophyte Suaeda salsa L. Plant Sci 168:423–430
Zou LP, Sun XH, Zhang ZG, Liu P, Wu JX, Tian CJ, Qiu JL, Lu TG (2011) Leaf rolling controlled by the homeodomain leucine zipper class IV gene Roc5 in rice. Plant Physiol 156:1589–1602
Acknowledgements
We are thankful to Dr. Rui Xia (South China Agricultural University) and Dr. Jedrzej Szymanski (Weizmann Institute of Science) for helpful discussions and critical comments on the manuscript. We also thank the National Natural Science Foundation of China and the Science and Technology Program of Guangzhou for supporting this work.
Funding
This work was supported by grants from the National Natural Science Foundation of China (31671645) to Z. Z., and the Science and Technology Program of Guangzhou (201704030021) and the National Natural Science Foundation of China (31741095) to Q. X.
Availability of data and materials
All data supporting the conclusions of this article are provided within the article and its (Additional file 1: Table S12, Additional file 2: Table S1, Additional file 3: Figure S1, Additional file 4: Figure S2, Additional file 5: Table S2, Additional file 6: Figure S3, Additional file 7: Figure S4, Additional file 8: Table S3, Additional file 9: Table S4, Additional file 10: Table S5, Additional file 11: Table S6, Additional file 12: Table S7, Additional file 13: Table S8, Additional file 14: Table S9, Additional file 15: Figure S5, Additional file 16: Table S11, Additional file 17: Table S12, respectively).
Author information
Authors and Affiliations
Contributions
QX and ZZ together designed the experiments. JL performed most of the experiments assisted by SG, BS, QL, XC, and HP. JL and SG performed the drought and ABA response. JL and QL analyzed the RNA-seq and iTRAQ datasets. JL, XC and HP conducted the plant growth in greenhouse and paddy field. JL and QX wrote the manuscript. All authors have discussed the results and contributed to the drafting of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1:
Table S12. Primers used in this study. (XLSX 12 kb)
Additional file 2:
Table S1. Expression of drought-responsive genes in WT and rel1-D. (XLSX 13 kb)
Additional file 3:
Figure S1. Expression of abaxial leaf rolling genes in response to PEG treatment. a Expression of REL1 during different time courses by PEG treatment. b Expression of ACL1 during different time courses by PEG treatment. c Expression of ACL2 during different time courses by PEG treatment. d Expression of Roc5 during different time courses by PEG treatment. a-d Multiple comparisons, Duncan, p-value < 0.01. (TIF 2233 kb)
Additional file 4:
Figure S2. Verification of the RNA-seq. Modes for qPCR were cDNA of WT and the rel1-D leaves in tillering stage, three biological repetitions, error bars were S.E.; the X axis was gene names, the Y axis was log2 (Fold change). (TIF 56 kb)
Additional file 5:
Table S2. Differentially expressed genes in rel1-D mutant. (XLSX 190 kb)
Additional file 6:
Figure S3. Expressions of BR-related genes. A Expressions of 10 BR synthesis-related genes. B Expressions of 31 BR signaling genes. A-B The colors indicated the mean expression level (FPKM), the red color was the highest and the green color was the lowest; red panes highlighted p-value < 0.05. (TIF 87 kb)
Additional file 7:
Figure S4. Expressions of ABA-related genes. A Expressions of 15 ABA synthesis-related genes (positive-related). B Expressions of 3 ABA synthesis-related genes (negative-related). C Expressions of 43 ABA signaling genes. A-C The colors indicated the mean expression level (FPKM), the red color was the highest and the green color was the lowest; red panes highlighted p-value < 0.05. (TIF 141 kb)
Additional file 8:
Table S3. GO enrichment (Cellular Component) of the up-regulated DEGs in rel1-D. (XLSX 121 kb)
Additional file 9:
Table S4. GO enrichment (Molecular Function) of the up-regulated DEGs in rel1-D. (XLSX 121 kb)
Additional file 10:
Table S5. GO enrichment (Biological Process) of the up-regulated DEGs in rel1-D. (XLSX 126 kb)
Additional file 11:
Table S6. GO enrichment (Cellular Component) of the down-regulated DEGs in rel1-D. (XLSX 121 kb)
Additional file 12:
Table S7. GO enrichment (Molecular Function) of the down-regulated DEGs in rel1-D. (XLSX 121 kb)
Additional file 13:
Table S8. GO enrichment (Biological Process) of the down-regulated DEGs in rel1-D. (XLSX 125 kb)
Additional file 14:
Table S9. Peptides identified in iTRAQ. (XLSX 3162 kb)
Additional file 15
Figure S5. Verification of the iTRAQ. Modes for qPCR were cDNA of WT and the rel1-D leaves in tillering stage, three biological repetitions, error bars were S.E.; the X axis was gene names, the Y axis was log2 (Fold change of transcript). (TIF 41 kb)
Additional file 16:
Table S10. Plastid associated differentially expressed proteins in rel1-D. (XLSX 13 kb)
Additional file 17:
Table S11. Primers used in this study. (XLSX 12 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liang, J., Guo, S., Sun, B. et al. Constitutive expression of REL1 confers the rice response to drought stress and abscisic acid. Rice 11, 59 (2018). https://doi.org/10.1186/s12284-018-0251-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12284-018-0251-0