Abstract
Background
Protein phosphatase class 2 C (PP2C) is the largest protein phosphatase family in plants. Members of the PP2C gene family are involved in a variety of physiological pathways in plants, including the abscisic acid signalling pathway, the regulation of plant growth and development, etc., and are capable of responding to a wide range of biotic and abiotic stresses, and play an important role in plant growth, development, and response to stress. Apocynum is a perennial persistent herb, divided into Apocynum venetum and Apocynum hendersonii. It mainly grows in saline soil, deserts and other harsh environments, and is widely used in saline soil improvement, ecological restoration, textiles and medicine. A. hendersonii was found to be more tolerant to adverse conditions. The main purpose of this study was to investigate the PP2C gene family and its expression pattern under salt stress and to identify important candidate genes related to salt tolerance.
Results
In this study, 68 AvPP2C genes and 68 AhPP2C genes were identified from the genomes of A. venetum and A. hendersonii, respectively. They were classified into 13 subgroups based on their phylogenetic relationships and were further analyzed for their subcellular locations, gene structures, conserved structural domains, and cis-acting elements. The results of qRT-PCR analyses of seven AvPP2C genes and seven AhPP2C genes proved that they differed significantly in gene expression under salt stress. It has been observed that the PP2C genes in A. venetum and A. hendersonii exhibit different expression patterns. Specifically, AvPP2C2, 6, 24, 27, 41 and AhPP2C2, 6, 24, 27, 42 have shown significant differences in expression under salt stress. This indicates that these genes may play a crucial role in the salt tolerance mechanism of A. venetum and A. hendersonii.
Conclusions
In this study, we conducted a genome-wide analysis of the AvPP2C and AhPP2C gene families in Apocynum, which provided a reference for further understanding the functional characteristics of these genes.
Similar content being viewed by others
Background
Plants are exposed to different environmental adversities such as drought, salinity, and high temperature during growth and development, which can affect the balance of ions, regulation of hormones, and the reception and transmission of signals in the plant body [1, 2]. These factors ultimately impact the growth and development of the plant and can significantly reduce crop yield and quality [3,4,5,6]. Plants have evolved an array of response mechanisms to thrive in challenging environments. Among these is protein reversible phosphorylation, a complex process that entails two opposing chemical reactions: protein phosphorylation and dephosphorylation. This modification process is facilitated by protein kinases (PKs) and protein phosphatases (PPs), and can promptly adjust the biological properties of proteins based on external stimuli. It plays a crucial role in plant growth and development, signal transduction, and in mitigating the effects of stress due to unfavorable conditions [7]. Phosphorylation occurs at specific sites in proteins, including serine (Ser), threonine (Thr), and tyrosine (Tyr), and is regulated by protein kinases (PKs) and protein phosphatases (PPs). PPs are classified based on their substrate specificity, with Ser/Thr phosphatases (STPs), protein Tyr phosphatases (PTPs), and dual-specificity phosphatases (DSPTPs) being the main categories [8]. PTPs are further divided into two groups, phosphoprotein metal phosphatases (PPM) and phosphoprotein phosphatases (PPP), based on crystal structure, amino acid sequence, and response to inhibitors [9, 10]. The PPP family includes various types of protein phosphatases, such as PP1, PP2A, PP2B, PP4, PP5, PP6, and PP7, while type 2 C protein phosphatase (PP2C) is the largest protein phosphatase family in plants. Although closely related to the PPP family, PP2C has no sequence homology and belongs to the PPM family [9,10,11,12]. PP2C has a conserved catalytic structure at the C-terminus and a functional extension region at the N-terminal that determines its various functions [12]. In Arabidopsis, distinct subgroups of PP2C genes exhibit specific functions. It is worth noting that these genes belong to the PP2C family, which plays a vital role in regulating plant growth and development. Group A comprises genes that participate in abscisic acid (ABA) signaling, such as ABI1 and ABI2. These genes interact with SnRK2, binding and dephosphorylating it, which inhibits the ABA signaling pathway and stress response [13,14,15,16]. Group B is responsible for mitogen-activated protein kinase (MAPK) signaling, while Group C is mainly involved in flower development [17]. Members of Group D may have negative effects on cell elongation, but can also have positive effects on the ABA signaling pathway in response to saline stress [18,19,20]. Group E is responsible for regulating the opening and closing of plant stomata, while Group F induces stress responses in bacteria. The PP2C gene family has been extensively researched in various plants, such as rice [21, 22], maize [23, 24], and wheat [25], among others. Their roles in plant growth, development, and ability to combat negative stressors are increasingly being comprehended.
Apocynum is a perennial, persistent herb that grows mainly in harsh environments such as saline soils and deserts, and is widely used in saline land improvement, ecological restoration, textile and medicine [26,27,28,29,30,31,32]. Apocynum is divided into Apocynum venetum and Apocynum hendersonii, and these two species differ in morphological characteristics and physiological properties. The former is characterized by red stems and flowers, while the latter is more salt tolerant and sports white flowers, making it an ideal choice for soil improvement and restoration [33, 34]. Currently, a comprehensive systematic analysis of the PP2C gene family in A. venetum and A. hendersonii remains elusive, particularly with regards to their expression patterns and underlying functional mechanisms in response to salt stress. Therefore, studying the role and mechanism of the PP2C gene family in Apocynum under salt stress is of significant importance for improving the salt tolerance of plants.
In this study, we performed genome-wide identification and analysis of PP2C gene family members in Apocynum. 68 AvPP2C genes and 68 AhPP2C genes were identified, and the physicochemical properties, gene structure, conserved structural domains, cis-acting elements, chromosomal location distribution, gene duplication, and phylogenetic relationships of PP2C gene family members were analyzed. And the expression patterns of PP2C genes in different tissues of Apocynum were analyzed based on the transcriptome data of our group. The response of the PP2C gene after NaCl stress was analyzed by qRT-PCR study. The results of this study laid the foundation for the study of PP2C gene function in Apocynum and provided a reference for the excavation and genetic improvement of Apocynum resistance genes.
Results
Identification, characteristics and phylogenetic analysis of the PP2C gene family in Apocynum
In this study, we used BLASTp to identify 68 members of the AvPP2C gene family in A. venetum genome and 68 members of the AhPP2C gene family in the A. hendersonii genome, respectively, using the sequences of 80 AtPP2C proteins from Arabidopsis thaliana as a reference. We used TBtools to map the positional distribution of the PP2C genes on chromosomes of A. venetum and A. hendersonii respectively, which were named AvPP2C1-AvPP2C68 and AhPP2C1-AhPP2C68 according to their chromosomal positions (Fig. 1). Except for AhPP2C68 which was not assembled, the remaining 68 AvPP2C genes and 67 AhPP2C genes were located on 11 chromosomes, respectively. There were fewer genes on chromosomes 4, 8 and 10.
Chromosomal distribution and localization of the PP2C gene family in A. venetum and A. hendersonii. The blue rectangular bars represent chromosomes of Apocynum, and chromosome names are shown on the left side of each chromosome. The 0–27 Mb scale bar on the left represents chromosome length. (A) Chromosome distribution and localization of AvPP2Cs. (B) Chromosome distribution and localization of AhPP2Cs
We analyzed the physicochemical properties such as molecular weight (MW), theoretical isoelectric point (pI), and subcellular localization of the 68 AvPP2C proteins and 68 AhPP2C proteins (Table S1). The 68 AvPP2C genes encoded proteins were located at lengths ranging from 278 to 1086, theoretical isoelectric points ranging from 4.38 to 9.78, and molecular weights ranging from 30369.58 to 123024.02 Da. The 68 AhPP2C genes encoded proteins with lengths ranging from 138 to 1092, theoretical isoelectric points ranging from 4.62 to 9.8, and molecular weights of 15540.2-123582.74 Da. The results of subcellular localization show that PP2C genes have the ability to appear in various areas of the cell, including the nucleus, chloroplast, cell wall, cell membrane, cytoplasm, mitochondria, and peroxisome. Some specific PP2C genes, such as AvPP2C9, 10, 16, 19, 20, 21, 29, 31, 35, 38, 47, 53, 62 and AhPP2C9, 10, 21, 29, 31, 35, 37, 38, 39, 48, 53, 54, 58 are predicted to be present in multiple cellular structures.
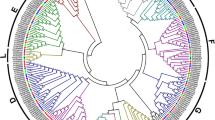
To investigate the phylogenetic relationship between A. venetum, A. hendersonii and Arabidopsis thaliana PP2C genes, a phylogenetic tree was constructed using the neighbor-joining (NJ) method based on 80 AtPP2C genes of Arabidopsis thaliana, 68 AvPP2C genes of A. venetum and 68 AhPP2C genes of A. hendersonii (Fig. 2). Based on phylogenetic analysis, the PP2C genes in Apocynum were categorized into 13 subgroups (A-L). These subgroups contained 9, 4, 4, 8, 7, 6, 4, 6, 3, 2, 2, 7, 2 AvPP2C genes and 9, 4, 4, 9, 7, 6, 3, 6, 3, 2, 2, 7, 2 AhPP2C genes, with similar groupings to those found in Arabidopsis. Notably, PP2C proteins from both Apocynum and Arabidopsis were present in each subfamily, suggesting a common ancestor. Interestingly, AvPP2C3, AvPP2C7, AvPP2C31, AvPP2C61, AhPP2C3, AhPP2C7, AhPP2C31, and AhPP2C60 of Apocynum did not cluster with any other group. The evolutionary tree also showed that genes in A. venetum and A. hendersonii were tightly clustered, indicating high homology and similar gene functions.
Gene structures and conserved motifs analyses of the AvPP2C genes and the AhPP2C genes
Based on their phylogenetic relationships, the structures and conserved domains of the AvPP2C and AhPP2C genes were studied. The gene structures include positional information such as coding sequence (CDS), untranslated regions (UTR), introns and PP2C structural domains (Fig. 3A and C). Genes within the same subgroup share similar gene structures, but there are variations in the length and arrangement of their CDSs and UTRs. For instance, all PP2C genes in group C contained UTR and CDS regions, but their lengths varied. AvPP2C42 had the lengthiest UTR region, while 16 genes in A. venetum had no UTR regions and 14 genes in A. hendersonii had no UTR regions. AvPP2C39 and AhPP2C40 had the highest number of CDS regions, totaling 16. After examining the number of introns, it was found that three AvPP2C genes and four AhPP2C genes in Apocynum had no introns, while the remaining 129 PP2C genes contained introns.
In order to study the structure and function of PP2C proteins and their evolutionary relationships, the conserved domains of AvPP2Cs proteins and AhPP2Cs proteins were analyzed using the online MEME website. The analysis revealed 15 different conserved domains, which were given the names motif 1 to motif 15 (Fig. 3A and B, Table S2). The results demonstrated that PP2C proteins in the same subgroups possess a similar distribution that may be linked to their functions. Motif 1 (except AvPP2C62, AhPP2C67, AhPP2C68), motif 2 (except AvPP2C31, AvPP2C44, AhPP2C45, AhPP2C31), and motif 3 (except AhPP2C68, AhPP2C19, AvPP2C19, AvPP2C48) were present in all subgroups. Motifs 6, 7, 11, 13 and 15 were absent in subgroups C and D, while motif 5 was only present in subgroups C and D, except AhPP2C68. Motif 9 and 10 were present only in subgroup D, and motif 11 was present only in subgroup E, but not in other groups.
Phylogenetic tree, conserved domains and gene structures of 68 AvPP2Cs proteins and 68 AhPP2Cs proteins. (A) The phylogenetic tree was constructed using all AvPP2C proteins and AhPP2C proteins with 1000 replicates on each node. (B) The conserved domains identified by the Batch CD-search Tool of the NCBI website. 15 colored squares represent motif1-motif15 and gray lines represent amino acid lengths. (C) The gene structures of AvPP2C genes and AhPP2C genes. The green rectangles are UTRs, yellow rectangles are CDSs, and pink rectangles are PP2C structural domains, and the gray lines represent introns
Cis-element analysis of the PP2Cs promoter in A. venetum and A. hendersonii
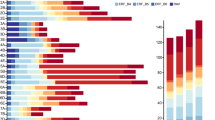
Using the PlantCARE online database, we analyzed and predicted the cis-acting elements in the promoter regions of A. venetum and A. hendersonii, and conducted a visual analysis using TBtools software. A total of 68 cis-acting elements were retrieved. Among them, there were 32 light-responsive elements, 13 phytohormone-responsive elements, 12 stress-responsive elements, and 11 elements related to plant growth and development (Fig. 4, Table S3). It was found that all the promoters of AvPP2Cs and AhPP2Cs contained light-responsive elements, phytohormone-responsive elements, and stress-responsive elements. However, elements associated with plant growth and development were missing in AvPP2C18, AvPP2C33, AvPP2C59, AvPP2C61, AvPP2C65, AhPP2C18, AhPP2C19, AhPP2C32, AhPP2C58, AhPP2C60, and AhPP2C66. Box 4 was identified as the most abundant light-responsive element, with 263 in A. venetum and 290 in A. hendersonii. This suggests that the expression of PP2C genes may be strongly influenced by light conditions. Among the phytohormone response elements, a large number of abscisic acid response elements (ABRE) and AAGAA-motifs were found, with AvPP2Cs having 148 ABREs and 132 AAGAA-motifs, and AhPP2Cs having 141 ABREs and 129 AAGAA-motifs. The presence of these elements suggests that PP2C genes play an important role in ABA-mediated stress resistance pathways. The ethylene response element (ERE) was also found in large numbers, with AvPP2Cs having 147 and AhPP2Cs having 156. This indicates that the ethylene signaling pathway may be involved in the regulation of PP2C gene expression. The stress-responsive elements MYB and MYC were found to be the most abundant with 247 and 235 in A. venetum and 242 and 238 in A. hendersonii, respectively. These elements are involved in drought and ABA-induced responses. Another stress-responsive element, ARE, was found to be essential for anaerobic induction, with 133 in A. venetum and 130 in A. hendersonii. The large number of these elements suggests that most PP2C genes may respond to various stresses and are closely related to plant stress resistance. In A. venetum and A. hendersonii, the number of the same type of cis-acting elements differs, reflecting subtle differences in gene expression regulation between the two plants, which may be related to their ecological adaptability or genetic background.
Analysis of cis-acting elements in the promoter region of AvPP2Cs and AhPP2Cs. The distribution of cis-acting elements in the promoter region is shown on the left. On the right side is the heat map of cis-acting elements for the light-responsive, phytohormone-responsive, and-responsive, and the color concentration of the squares indicates the number of cis-acting elements. (A) Analysis of cis-acting elements in the promoter region of AvPP2Cs. (B) Analysis of cis-acting elements in the promoter region of AhPP2Cs
Collinear analysis and evolutionary analysis of AvPP2Cs proteins and AhPP2Cs proteins
The expansion of gene families in the genome is predominantly caused by tandem and segmental duplications. To gain insight into the PP2C gene family, 68 AvPP2C genes and 68 AhPP2C genes were analyzed for duplication (Fig. 5A and B, Table S4). It was discovered that in A. venetum, one tandem duplication pair was located on chromosome LG06 (39/40), whereas 17 pairs of segmental duplication genes were unevenly distributed on chromosomes other than LG08 and LG10. In A. hendersonii, there were 2 tandem replication gene pairs located on chromosome LG06 (38/39, 40/41), and 17 fragment replication gene pairs were unevenly distributed on 9 chromosomes except for LG08 and LG10. These findings suggest that the evolution of PP2Cs in Apocynum was primarily driven by segmental duplication events, as the number of segmental duplication gene pairs was significantly greater than that of tandem duplications.
Furthermore, the study investigated the homology between A. venetum, A. hendersonii, and Arabidopsis thaliana to understand the evolutionary relationship of PP2C genes (Fig. 5C, Table S5). The results indicated that there were 84 homologous gene pairs between A. venetum and Arabidopsis thaliana, and 82 homologous gene pairs between A. hendersonii and Arabidopsis thaliana. Among these pairs, A. venetum and A. hendersonii had the highest number of homologous gene pairs on chromosome LG06, with 13 pairs each. These findings suggest that A. venetum and A. hendersonii may have a high degree of homology with Arabidopsis.
Collinearity analysis of the AvPP2C genes. (A) Distribution of AvPP2C genes on chromosomes and gene duplications. Red lines represent gene fragment duplications. (B) Distribution of AhPP2C genes on chromosomes and gene duplications. Red lines represent gene fragment duplications. (C) Comparative analysis of PP2C genes between Arabidopsis and A. hendersonii and A. venetum. Blue lines represent homologs of PP2C genes
Tissue-specific expression of AvPP2C genes and AhPP2C genes in different tissues
To gain a better understanding of the function of PP2C genes in the growth and development of A. venetum and A. hendersonii, we analyzed the expression patterns of PP2C genes using RNA-Seq data of roots, stems, and leaves of both species (Fig. 6). Most of the PP2C genes were expressed in all tissues, except for four genes (AvPP2C19, AvPP2C38, AhPP2C39, and AhPP2C68), which were barely expressed. In A. venetum, AvPP2C47 was highly expressed in roots, stems, and leaves. AvPP2C9, AvPP2C24, AvPP2C49, AvPP2C60, and AvPP2C68 were moderately expressed in roots; AvPP2C13 and AvPP2C60 were highly expressed in stems; AvPP2C13 was highly expressed in leaves, and AvPP2C7 and AvPP2C60 were moderately expressed in leaves. In A. hendersonii, AhPP2C29 and AhPP2C67 were highly expressed in roots, while AhPP2C24, AhPP2C13, AhPP2C49, AhPP2C48, and AhPP2C59 were moderately expressed in roots. AhPP2C9, AhPP2C13, AhPP2C59, and AhPP2C48 were highly expressed in stems, and AhPP2C13, AhPP2C48, and AhPP2C59 were highly expressed in leaves.
Relative expression changes of AvPP2C genes and AhPP2C genes under NaCl treatment
In order to study the expression of AvPP2C genes and AhPP2C genes under salt stress conditions and to identify important PP2C genes related to salt tolerance. Based on phylogenetic analysis and homology of known PP2C genes in Arabidopsis thaliana, five genes in subgroup D and two genes in subgroup G were selected in A. hendersonii and their expression patterns in different tissues of A. venetum under salt stress were investigated by qRT-PCR (Fig. 7).
The results showed that the AvPP2C1 gene was down-regulated in roots after NaCl stress, with the lowest expression at 50 mM concentration. It was up-regulated in leaves and phloem, with the highest expression at 200 mM and 100 mM concentrations, respectively, reaching 2.14-fold and 1.40-fold of CK. AhPP2C1 gene was up-regulated in leaves and increased with increasing NaCl concentration; it showed down-regulation in bast and decreased with increasing NaCl concentration. AvPP2C2 gene showed up-regulation under NaCl stress, in which the expression of the gene in leaves varied the most with NaCl concentration, which was 10.44 times that of CK. The expression trend of AvPP2C2 in root, xylem and phloem was the same, and all of them reached the maximum expression at 100 mM concentration. While the AhPP2C2 gene was down-regulated in roots and up-regulated in leaves after NaCl stress, there was little change in the xylem and phloem, but the expression reached the maximum at 50 mM concentration in all. AvPP2C6 gene did not change significantly in roots, but showed up-regulation in leaves and xylem phloem, and it is noteworthy that its expression in leaves increased with the increase of NaCl concentration and reached the maximum at 200 mM, which was 13.55 times that of CK. And AhPP2C6 was different from AvPP2C6 in that it showed down-regulation in roots. The expression of AvPP2C14 was different in roots, leaves, xylem and phloem, with the highest in leaves and the lowest in roots; after NaCl stress, the AvPP2C14 gene showed a tendency of increasing and then de-creasing with the increase of NaCl concentration, and its expression reached the maximum in all tissues at the concentration of 50 mM. While the expression of AhPP2C14 did not change significantly under NaCl treatment, it only showed up-regulation in leaves, and the expression of the gene was the highest at 50 mM concentration, and its expression gradually declined with the increase of concentration. AvPP2C24 gene had the highest expression in leaves and showed up-regulation in all tissues after NaCl stress. AhPP2C24 gene was the same as AvPP2C24 gene in that the expression of AhPP2C24 gene was up-regulated in all tissues after NaCl stress, except in phloem at 50 mM concentration. The difference was that the expression of AhPP2C24 gene changed more significantly in leaves after NaCl treatment. Both AvPP2C27 and AhPP2C27 genes showed up-regulation after NaCl treatment, but the expression of AvPP2C27 gene in roots and leaves increased with increasing NaCl concentration, while the expression of AhPP2C27 gene in leaves reached a maximum at 50 mM concentration and then decreased with increasing concentration. The expression of AvPP2C41 and AhPP2C42 genes in roots increased with increasing NaCl concentration, AvPP2C41 gene showed down-regulation in leaves at 100 mM and up-regulation at 50 mM and 200 mM concentrations. While AhPP2C42 gene showed up-regulation after NaCl stress, the expression was maximum in leaves at 50 mM concentration, which was 8.20 times of CK, and then decreased with the increase of NaCl concentration.
The qRT-PCR results of selected PP2C genes of different tissue samples (root, xylem, phloem, leaf) of A. venetum and A. hendersonii after 14 days of NaCl stress. (A) The relative expression levels of selected AvPP2C genes in A. venetum parts. (B) The relative expression levels of selected AhPP2C genes of A. hendersonii
Discussion
PP2C genes are closely related to abscisic acid ABA signaling pathway, plant adversity stress, and stomatal opening and closing, etc. The PP2C gene families of Arabidopsis thaliana, rice, maize, and wheat have been investigated by previous authors. In this study, we comprehensively analyzed the AvPP2C genes of A. venetum and the AhPP2C genes of A. hendersonii, including the identification of the gene families, phylogenetic relationships, and chromosomal locations, collinear relationship, gene structure, conserved motifs, and expression patterns. A total of 136 PP2C genes, including 68 AvPP2C genes and 68 AhPP2C genes, were identified in Apocynum, and the physicochemical properties and subcellular localization of the genes were predicted, and it was found that the vast majority of PP2C genes were located in the nucleus and chloroplasts, and a few genes might be present in the mitochondria, cell wall, cytoplasm, peroxisomes, and cell membrane. Therefore, it is hypothesized that they may be related to photosynthesis, cell growth and development, and respiration. According to phylogenetic and evolutionary relationship analysis, A. hendersonii and A. venetum are closely related to each other, and a total of 136 PP2C genes in A. hendersonii and A. venetum were divided into 13 subgroups (A-L), which is consistent with the grouping of Arabidopsis thaliana. Previous studies found that genes in the A subgroup of PP2C genes in Arabidopsis mainly inhibit ABA receptor activity and negatively regulate ABA signaling [35]; the B subgroup is involved in the mitogen-activated protein kinase signaling pathway and participates in the process of MAPK phosphorylation, and is capable of dephosphorylating and inactivating MAPK [36]; Subgroup C gene family members are involved in stem cell maintenance and differentiation [37]; Members of subgroup D negatively regulate small auxin up RNA (SAUR)-mediated cell elongation, and PP2C.D1, PP2C.D2, and PP2C.D5 in Arabidopsis act during leaf greening, and PP2C.D1 regulates hook formation by affecting ethylene accumulation [18, 38, 39]. Moreover, members of this group respond to saline and alkaline stress, which may be related to stomatal movement and function, by inhibiting plasma membrane H-ATPase activity. It has been shown that Arabidopsis PP2C.D negatively regulates aluminium resistance by modulating malate secretion [19, 40]; Members of the E subgroup gene family may be associated with inflorescence stem growth, regulation of stomatal signaling, and so on [41,42,43]. Based on the phylogenetic relationship between Apocynum and Arabidopsis, it is hypothesized that the functions of gene family members located in the same subgroup in Apocynum may be similar to those in Arabidopsis. The prediction results of subcellular localization for PP2C gene family members in Apocynum indicate that most PP2C genes in Apocynum are located in the nucleus and chloroplasts. This suggests that PP2C genes may participate in regulating the expression of salt stress-related genes in the nucleus, and may also be involved in regulating the activity of enzymes related to photosynthesis. Additionally, some genes may be located in the cell wall, cytoplasm, cell membrane, mitochondria, and peroxisomes, which may be closely related to reducing water loss, regulating intracellular ion concentration, and scavenging reactive oxygen species in plant cells. These functions can enhance the salt tolerance of plants. The predictions of PP2C gene family in A. venetum and A. hendersonii were partially different. For example, AvPP2C9 was predicted to be located in mitochondria and the nucleus, whereas AhPP2C9 was predicted to be located in chloroplasts, mitochondria, and the nucleus. AvPP2C20 was located in the nucleus and chloroplasts, whereas AhPP2C20 was predicted to be located in chloroplasts only. This may imply differences in the location and function of some PP2C genes in A. hendersonii and A. venetum.
The diversity of gene structures, conserved structural domains, and cis-acting elements is also closely related to the functions of gene families. Genes in the same subgroup are similar in structure, which may imply that they have the same function [44]. The gene structures of A. hendersonii and A. venetum genes from the same evolutionary tree branching were similar, except for the AhPP2C68 gene. However, the gene structures of individual PP2C genes were different again, for example, AvPP2C18 and AhPP2C18 in subgroup L had significantly more CDS regions compared with AvPP2C33 and AhPP2C33, which might be related to the increase of introns during the evolutionary process. The PP2C gene family is evolutionarily conserved, and most PP2Cs have a highly conserved structural domain with a unique non-catalytic N-terminal extension region. The specificity of the function of each sub-group of PP2C genes may be related to an N-terminal substrate docking domain [45]. A total of 15 conserved motifs were identified in the amino acid sequences of AvPP2C and AhPP2C genes, and the distribution of conserved motifs was similar for genes in the same subgroup, which may be closely related to the function of PP2C proteins. The sequence structures of some PP2C genes in A. venetum and A. hendersonii exhibit differences, which may lead to variations in their functions or expression regulation. Cis-acting elements are involved in the regulation of gene expression and are closely related to plant growth and development, hormone responses and responses to various stresses. Among AvPP2C and AhPP2C genes, Box4 elements were the most abundant (AvPP2Cs: 263; AhPP2Cs: 290), followed by MYB elements (AvPP2Cs: 247; AhPP2Cs: 242). MYB elements are closely related to plant abiotic stress response [46]. Among all the response elements, light responsive and stress responsive were the most numerous elements, with a high number of 1,318 stress responsive in A. hendersonii. This may be related to the characteristics of A. hendersonii high-stress tolerance.
In this study, chromosomal position analysis and covariance analysis of PP2C gene family of A. venetum and A. hendersonii were carried out, which is important to understand the amplification mechanism of the genes. It was found that both AvPP2Cs and AhPP2Cs were distributed on 11 chromosomes. Seventeen pairs of segmental duplications and one pair of tandem duplications were identified in A. venetum; 17 pairs of segmental duplications and two pairs of tandem duplications were identified in A. hendersonii. Gene duplication is the main driver of gene family amplification, which mainly includes segmental duplication, tandem duplication, and genomic duplication, and segmental duplication is more conducive to maintaining gene function than tandem duplication [47]. Therefore, we hypothesize that segmental duplication is the main force driving the evolution and expansion of the PP2C gene family, which is similar to the results found for PP2C in Arabidopsis [48], woodland and pineapple strawberry [49], and cucumber [7].
AvPP2C genes and AhPP2C genes showed specific expression in different tissues, and most of the PP2C genes were lowly expressed in roots, stems and leaves. However, among all the PP2C genes, AvPP2C13, AvPP2C47, AvPP2C60, AhPP2C48, and AhPP2C59 showed higher expression in all three tissues: roots, stems, and leaves, whereas AhPP2C29, and AhPP2C67 showed high expression in roots and lower expression in stems and leaves. In cucumber, most of the CsPP2C genes were highly expressed in fertilized ovaries, male, female, and leaf (except CsPP2C11, 41, 5, 33, 50), and low in other tissues [4]. In maize, most of the ZmPP2C genes were expressed in all tissues, with ZmPP2C42 and ZmPP2C47 being the most highly expressed in mature pollen, and ZmPP2C59 being expressed in primary roots and root cortex [50]. This suggests that PP2C genes in different plants may play different roles at different stages of plant growth, and some of these PP2C genes may have special functions in specific tissues.
Studies on Arabidopsis have demonstrated that members of subgroup D respond to saline stress, and AtPP2C.D6 and AtPP2C.D7 interact with SCaBP8 of the SOS pathway, whose phosphatase activity is inhibited under salt stress. AtPP2C.G1 in group G is dependent on ABA to positively regulate salt tolerance. Our study revealed that the expression of certain AvPP2C genes differed significantly from that of AhPP2C genes following NaCl stress. Specifically, the expression of the AvPP2C2 gene was significantly up-regulated after salt stress, and its expression was notably higher in roots, xylem, phloem, and leaves. It indicated that AvPP2C2 gene might play a crucial role in these four tissues when A. venetum resistes salt stress. The expression of AhPP2C2 gene was down-regulated in roots but up-regulated in leaves after salt stress.In contrast, it exhibited a similar expression pattern in xylem and phloem, peaking at a concentration of 50 mM NaCl. AvPP2C2 and AhPP2C2 displayed distinct expression patterns in A. venetum and A. hendersonii under salt stress, which we hypothesize is related to their varying salt tolerance mechanisms. The expression of AvPP2C6 and AhPP2C6 was up-regulated in leaves, xylem and phloem. The change in AvPP2C6 expression was not apparent in roots, whereas AhPP2C6 showed down-regulation in roots. Additionally, AvPP2C24, 27 and 41 in roots displayed similar trends after salt stress. In summary, PP2C genes located within the same subgroup exhibited diverse responses to salt stress, while there may be different expression patterns observed in A. venetum and A. hendersonii, where AvPP2C2, 6, 24, 27, 41 and AhPP2C2, 6, 24, 27, 42 displayed significant differences in expression under salt stress. These fundings suggest that these genes may play significant roles in the salt stress response of A. venetum and A. hendersonii. Under salt stress, the significant expression changes of genes such as AvPP2C2, AvPP2C6, AvPP2C27, AhPP2C27, and AhPP2C42 in leaves may indicate that they play diverse roles in plant salt tolerance mechanisms. These include regulating ion balance, participating in signal transduction, influencing plant growth and development, responding to stress, and activating defense mechanisms. These mechanisms work together to enhance the plant’s ability to adapt to salt stress.
Apocynum, renowned for its remarkable stress tolerance, holds immense potential in deciphering the role of the PP2C gene family under salt stress conditions. By investigating the expression patterns of these genes, we can gain insights into their specific functions in responding to salt stress. In Apocynum, PP2C genes are pivotal in stress response mechanisms, and their precise regulation through advanced gene editing techniques or molecular marker-assisted breeding approaches can facilitate the cultivation of crop varieties with enhanced stress resilience. Additionally, secondary metabolites, particularly flavonoids, in Apocynum exhibit significant medicinal value and play a crucial role in plant growth, development, and defense mechanisms [51]. A comprehensive understanding of the role of PP2C genes in the biosynthesis of these secondary metabolites could serve as a theoretical foundation for future drug research and development. In conclusion, a thorough exploration of the functional characteristics of the PP2C gene family in Apocynum offers novel insights and strategies for stress-resistant breeding, medicinal value exploitation, and other applications.
Conclusions
In this comprehensive study, we have conducted a genome-wide identification and in-depth analysis of PP2C family members in the genomes of A. venetum and A. hendersonii. A total of 68 AvPP2C genes and 68 AhPP2C genes were successfully identified and systematically classified into 13 distinct subgroups. These subgroups were then subjected to a rigorous analysis of subcellular localization, gene structure, conserved structural domains, and cis-acting elements. Upon evaluating the expression patterns of PP2C genes in A. venetum and A. hendersonii under salt stress conditions, we observed significant differences, particularly in the subgroup D and subgroup G genes. These variations in expression might serve as potential indicators of the distinct salt tolerance mechanisms between A. venetum and A. hendersonii. The findings of this study offer valuable insights into the functional roles of PP2C genes and serve as a foundation for future research aimed at elucidating the precise mechanisms underlying salt tolerance in these plant species.
Materials and methods
Identification of PP2C genes in Apocynum
The protein sequences of 80 AtPP2C genes of Arabidopsis were retrieved from Tair (https://www.arabidopsis.org/) [52]. Additionally, we acquired the protein sequences of A. venetum and A. hendersonii from the whole genome data sequenced within our laboratory. To identify the PP2C genes, we employed two distinct methodologies. Initially, we utilized BLASTp to locate potential PP2C proteins within the genomes. Subsequently, we downloaded the PP2C structural domain (PF00481) from Pfam (http://pfam.xfam.org/) and leveraged HMMER 3.0 to search our local protein database [46, 53, 54]. Concurrently, we employed the SMART (https://smart.embl.de/) to predict the structural domain, setting E-value < e− 5 [55, 56]. By intersecting the results from both methods, we extracted the relevant protein sequencesfrom the local protein database using TBtools, and identified the gene IDs through rigorous screening.
Sequence analysis and basic information of the Apocynum PP2C gene family
Physicochemical property analysis of PP2C proteins including number of amino acid, molecular weight, theoretical pI, instability index, aliphatic index, grand average of hydropathicity were analyzed by Protein Paramter Calc in TBtools. Prediction of the subcellular location of PP2C protein by Plant-mPloc (http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/) of Cell-Ploc 2.0.
Analysis of chromosome location and collinearity analysis
Chromosome position information was obtained from the annotation file of the Apocynum genome and the chromosome position distribution was mapped using TBtools. Homology and collinearity within the A. venetum and A. hendersonii genomes and between them and Arabidopsis thaliana were analyzed using the MCScanX software and visualized using the Advanced Circos and Multiple Synteny Plot functions in TBtools [57, 58].
Construction of phylogenetic tree
The phylogenetic tree of the PP2C gene family of Arabidopsis and Apocynum was constructed using MEGA-X with the Neighbor-Joining method, Bootstrap value set to 1000 and other default parameters [59, 60]. Finally, the phylogenetic tree was embellished with iTOL(https://itol.embl.de/) [61].
Analysis of gene structures and protein conserved motifs
The gene structure of Apocynum PP2C was analyzed using the Batch CD-search (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi) on the NCBI website. The MEME website (https://meme-suite.org/meme/tools/meme) was used to predict the conserved structural domains of the PP2C genes in Apocynum, setting the motif number to 15 and the rest to default values [62]. Gene Structure View in TBtools was used to generate the final overall map of the phylogenetic tree, gene structure and conserved structural domains of PP2Cs.
Analysis of cis-acting elements in PP2C gene promoters
The sequences of 2.0 kb upstream of AvPP2C genes and AhPP2C genes were extracted using TBtools and then the results were submitted to PlantCARE website (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/) for cis-acting element prediction [62].
Expression analysis of PP2C genes
From the transcriptome data of roots, stems and leaves of A. venetum and A. hendersonii, the FPKM (Fragments Per Kilobase of exon model per Million mapped fragments) values of AvPP2Cs and AhPP2Cs were extracted and heat maps were generated using TBtools [58].
Plant material treatment methods
Seedlings grown under hydroponic conditions for 6–8 weeks were divided into four groups and treated with different concentrations of NaCl stress (0, 50, 100, and 200 mM). After 14 days of treatment, roots, leaves, xylem and phloem samples were collected and stored in a refrigerator at -80 °C.
RNA isolation and qRT-PCR
Total RNA was extracted using the SteadyPure Plant RNA Extraction Kit (Accurate Biotechnology (Changsha, China) Co., Ltd.) according to previously published protocols [63], and the RNA was reverse transcribed into cDNA using the Evo M-MLV One Step RT-PCR Kit (Accurate Biotechnology (Changsha, China) Co., Ltd.) according to manufacturer’s protocols [64, 65]. Specific primers were designed for qRT-PCR using Primer5 (Table S6). ACT gene was used as an internal reference gene. qRT-PCR was performed on a CFX96 Touch Deep Well Real-Time Quantitative PCR System (Bio-Rad) using SYBR® Green Premix Pro Taq HS qPCR Kit II (Accurate Biotechnology (Changsha, China) Co., Ltd.), and performed according to published protocols [64, 66]. The relative expression of PP2C genes was calculated using the 2−ΔΔCT method and histograms were plotted using GraphPad Prism 8.
Data availability
The sequence information of ArabidopsisPP2C family genes were collected from The Arabidopsis Information Resoure (https://www.arabidopsis.org/). The PP2C family expression data were generated by qRT-PCR. The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- PP2C:
-
Protein phosphatase class 2 C
- qRT:
-
PCR-Quantitative Real-time PCR
- MAPK:
-
Mitogen-activated protein kinase
- MW:
-
Molecular weight
- pI:
-
Isoelectric point
- CDS:
-
Coding sequence
- UTR:
-
Untranslated regions
- CK:
-
Control check
- SAUR:
-
Small auxin up RNA
References
Zhai Z, Ao Q, Yang L, Lu F, Cheng H, Fang Q, Li C, Chen Q, Yan J, Wei Y et al. Rapeseed PP2C37 Interacts with PYR/PYL Abscisic Acid Receptors and Negatively Regulates Drought Tolerance. Journal of agricultural and food chemistry 2024.
Gupta S, Petrov V, Garg V, Mueller-Roeber B, Fernie AR, Nikoloski Z, Gechev T. The genome of Haberlea rhodopensis provides insights into the mechanisms for tolerance to multiple extreme environments. Cell Mol Life Sci 2024, 81(1).
Yang S, Gong Z, Guo Y, Gong J, Zheng S, Lin R, Yang H, Mao L, Qin F, Luo L, et al. Studies on plant responses to environmental change in China:the past and the future. Scientia Sinica Vitae. 2019;49(11):1457–78.
Zhu JK. Abiotic Stress Signaling and responses in plants. Cell. 2016;167(2):313–24.
Waadt R, Seller CA, Hsu PK, Takahashi Y, Munemasa S, Schroeder J. Plant hormone regulation of abiotic stress responses. Nat Rev Mol Cell Biol. 2022;23(10):680–94.
Biswas S, Seal P, Majumder B, Biswas AK. Efficacy of seed priming strategies for enhancing salinity tolerance in plants: an overview of the progress and achievements. Plant Stress 2023, 9.
Zhang G, Zhang Z, Luo S, Li X, Lyu J, Liu Z, Wan Z, Yu J. Genome-wide identification and expression analysis of the cucumber PP2C gene family. BMC Genomics 2022, 23(1).
Cao J, Jiang M, Li P, Chu Z. Genome-wide identification and evolutionary analyses of the PP2C gene family with their expression profiling in response to multiple stresses in Brachypodium distachyon. BMC Genomics 2016, 17.
Luan S. Protein phosphatases in plants. Annu Rev Plant Biol. 2003;54:63–92.
Shazadee H, Khan N, Wang J, Wang C, Zeng J, Huang Z, Wang X. Identification and expression profiling of protein phosphatases (PP2C) Gene Family in Gossypium hirsutum L. Int J Mol Sci 2019, 20(6).
Schweighofer A, Hirt H, Meskiene L. Plant PP2C phosphatases: emerging functions in stress signaling. Trends Plant Sci. 2004;9(5):236–43.
Zhang J, Tao N. Research progress of plant PP2C-type protein phosphatase in ABA signal transduction and adversity stress regulation mechanism. Guangxi Zhiwu / Guihaia. 2015;35(6):935–41.
Peirats-Llobet M, Han SK, Gonzalez-Guzman M, Jeong CW, Rodriguez L, Belda-Palazon B, Wagner D, Rodriguez PL. A direct link between Abscisic Acid Sensing and the chromatin-remodeling ATPase BRAHMA via core ABA signaling Pathway Components. Mol Plant. 2016;9(1):136–47.
Hsu PK, Dubeaux G, Takahashi Y, Schroeder JI. Signaling mechanisms in abscisic acid-mediated stomatal closure. Plant J. 2021;105(2):307–21.
Li ZX, Waadt R, Schroeder JI. Release of GTP Exchange Factor Mediated down-Regulation of Abscisic Acid Signal Transduction through ABA-Induced Rapid Degradation of RopGEFs. PLoS Biol 2016, 14(5).
Shi Y, Liu XN, Zhao SS, Guo Y. The PYR-PP2C-CKL2 module regulates ABA-mediated actin reorganization during stomatal closure. New Phytol. 2022;233(5):2168–84.
Wu P, Wang W, Li Y, Hou X. Divergent evolutionary patterns of the MAPK cascade genes in Brassica rapa and plant phylogenetics. Hortic Res 2017, 4.
Rovira A, Sentandreu M, Nagatani A, Leivar P, Monte E. The sequential action of MIDA9/PP2C.D1, PP2C.D2, and PP2C.D5 is necessary to form and maintain the Hook after Germination in the Dark. Front Plant Sci 2021, 12.
Chen C, Yu Y, Ding X, Liu B, Duanmu H, Zhu D, Sun X, Cao L, Zaib N, Li Q et al. Genome-wide analysis and expression profiling of PP2C clade D under saline and alkali stresses in wild soybean and Arabidopsis. Protoplasma 2018, 255(2):643–654.
Xue TT, Wang D, Zhang SZ, Ehlting J, Ni F, Jakab S, Zheng CC, Zhong Y. Genome-wide and expression analysis of protein phosphatase 2 C in rice and Arabidopsis. BMC Genomics 2008, 9.
Singh A, Jha SK, Bagri J, Pandey GK. ABA Inducible Rice Protein Phosphatase 2 C Confers ABA Insensitivity and Abiotic Stress Tolerance in Arabidopsis. PLoS ONE 2015, 10(4).
Xie W, Liu S, Gao H, Wu J, Liu D, Kinoshita T, Huang C-F. PP2C.D phosphatase SAL1 positively regulates aluminum resistance via restriction of aluminum uptake in rice. Plant Physiol. 2023;192(2):1498–516.
He Z, Wu J, Sun X, Dai M. The Maize Clade A PP2C phosphatases play critical roles in multiple abiotic stress responses. Int J Mol Sci 2019, 20(14).
Guo YZ, Shi YB, Wang YL, Liu F, Li Z, Qi JS, Wang Y, Zhang JB, Yang SH, Wang Y, et al. The clade F PP2C phosphatase ZmPP84 negatively regulates drought tolerance by repressing stomatal closure in maize. New Phytol. 2023;237(5):1728–44.
Wang JY, Li CA, Li L, Gao LF, Hu G, Zhang YF, Reynolds MP, Zhang XY, Jia JZ, Mao XG et al. DIW1 encoding a clade I PP2C phosphatase negatively regulates drought tolerance by de-phosphorylating TaSnRK1.1 in wheat. J Integr Plant Biol 2023.
Guo X, Chai W, Bai J, Ma Z. Cloning and Bioinformatics Analysis of AvFLS Gene from Apocynum Venetum. Mol Plant Breed. 2019;17(15):4978–85.
Huang X, Qiu X, Wang Y, Abubakar AS, Chen P, Chen J, Chen K, Yu C, Wang X, Gao G et al. Genome-wide investigation of the NAC Transcription Factor Family in Apocynum Venetum revealed their synergistic roles in Abiotic Stress Response and Trehalose Metabolism. Int J Mol Sci 2023, 24(5).
Zhang M, Lu X, Ren T, Marowa P, Meng C, Wang J, Yang H, Li C, Zhang L, Xu Z. Heterologous overexpression of Apocynum Venetum flavonoids synthetase genes improves Arabidopsis thaliana salt tolerance by activating the IAA and JA biosynthesis pathways. Front Plant Sci 2023, 14.
Cao C, Lin DF, Zhou YJ, Li N, Wang YW, Gong WB, Zhu ZH, Liu CW, Yan L, Hu ZX et al. Solid-state fermentation of Apocynum Venetum L. by Aspergillus Niger: Effect on phenolic compounds, antioxidant activities and metabolic syndrome-associated enzymes. Front Nutr 2023, 10.
Li X, Li JJ, Su HY, Sun P, Zhang Z, Li MF, Xing H. Physiological and transcriptional responses of Apocynum Venetum to Salt stress at the seed germination stage. Int J Mol Sci 2023, 24(4).
Zhang Y, Liu S, Ma JL, Chen C, Huang P, Ji JH, Wu D, Ren LQ. Apocynum Venetum leaf extract alleviated doxorubicin-induced cardiotoxicity through the AKT/Bcl-2 signaling pathway. Phytomedicine 2022, 94.
Jiang L, Wang L, Tian CY. High lithium tolerance of Apocynum Venetum seeds during germination. Environ Sci Pollut Res. 2018;25(5):5040–6.
Gao G, Abubakar AS, Chen J, Wang Y, Chen P, Chen K, Yu C, Wang X, Qiu X, Huang X et al. Comparative genome and metabolome analyses uncover the evolution and flavonoid biosynthesis between Apocynum Venetum and Apocynum Hendersonii. Iscience 2023, 26(5).
Yuan N, Li MM, Jia CL. De novo transcriptome assembly and population genetic analyses of an important coastal shrub, Apocynum Venetum L. BMC Plant Biol 2020, 20(1).
Ma Y. Regulators of PP2C phosphatase activity function as abscisic acid sensors . Science 2009, 324(5932):1266–1266.
Umbrasaite J, Schweighofer A, Meskiene I. Substrate Analysis of Arabidopsis PP2C-Type Protein Phosphatases. In: Plant Kinases: Methods and Protocols Edited by Dissmeyer N, Schnittger A, vol. 779; 2011: 149–161.
Yu LP, Miller AK, Clark SE. POLTERGEIST encodes a protein phosphatase 2 C that regulates CLAVATA pathways controlling stem cell identity at Arabidopsis shoot and flower meristems. Curr Biol. 2003;13(3):179–88.
Sentandreu M, Martín G, González-Schain N, Leivar P, Soy J, Tepperman JM, Quail PH, Monte E. Functional profiling identifies genes involved in Organ-Specific Branches of the PIF3 Regulatory Network in Arabidopsis. Plant Cell. 2011;23(11):3974–91.
Spartz AK, Ren H, Park MY, Grandt KN, Lee SH, Murphy AS, Sussman MR, Overvoorde PJ, Gray WM. SAUR Inhibition of PP2C-D Phosphatases activates plasma membrane H+-ATPases to promote cell expansion in Arabidopsis. Plant Cell. 2014;26(5):2129–42.
Fu H, Yu X, Jiang Y, Wang Y, Yang Y, Chen S, Chen Q, Guo Y. Salt overly sensitive1 is inhibited by clade D protein phosphatase 2 C D6 and D7 in Arabidopsis thaliana. Plant Cell. 2023;35(1):279–97.
Mishra G, Zhang WH, Deng F, Zhao J, Wang XM. A bifurcating pathway directs abscisic acid effects on stomatal closure and opening in Arabidopsis. Science. 2006;312(5771):264–6.
Sugimoto H, Kondo S, Tanaka T, Imamura C, Muramoto N, Hattori E, Ogawa Ki, Mitsukawa N, Ohto C. Overexpression of a novel Arabidopsis PP2C isoform, AtPP2CF1, enhances plant biomass production by increasing inflorescence stem growth. J Exp Bot. 2014;65(18):5385–400.
Wong JH, Klejchová M, Snipes SA, Nagpal P, Bak G, Wang B, Dunlap S, Park MY, Kunkel EN, Trinidad B, et al. SAUR proteins and PP2C.D phosphatases regulate H+-ATPases and K + channels to control stomatal movements. Plant Physiol. 2021;185(1):256–73.
Yang J, Chen R, Hu W, Wu Q, Tong X, Li X. Identification and expression analysis of PP2C gene family in Poncirus trifoliata. J Fruit Sci. 2022;39(4):532–47.
Meskiene I, Baudouin E, Schweighofer A, Liwosz A, Jonak C, Rodriguez PL, Jelinek H, Hirt H. Stress-induced protein phosphatase 2 C is a negative regulator of a mitogen-activated protein kinase. J Biol Chem. 2003;278(21):18945–52.
Hong YQ, Ahmad N, Zhang JY, Lv YX, Zhang XY, Ma XT, Liu XM, Yao N. Genome-wide analysis and transcriptional reprogrammings of MYB superfamily revealed positive insights into abiotic stress responses and anthocyanin accumulation in Carthamus tinctorius L. Mol Genet Genomics. 2022;297(1):125–45.
Lynch M, Conery JS. The evolutionary fate and consequences of duplicate genes. Science. 2000;290(5494):1151–5.
Cannon SB, Mitra A, Baumgarten A, Young ND, May G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004;4:10–10.
Guo L, Lu S, Liu T, Nai G, Ren J, Gou H, Chen B, Mao J. Genome-wide identification and Abiotic Stress Response Analysis of PP2C Gene Family in Woodland and Pineapple strawberries. Int J Mol Sci 2023, 24(4).
Wu H, Zhu L, Cai G, Lv C, Yang H, Ren X, Hu B, Zhou X, Jiang T, Xiang Y et al. Genome-wide identification and characterization of the PP2C family from Zea mays and its role in Long-Distance Signaling. Plants-Basel 2023, 12(17).
Ahmad N, Naeem M, Ali H, Alabbosh KF, Hussain H, Khan I, Siddiqui SA, Khan AA, Iqbal B. From challenges to solutions: the impact of melatonin on abiotic stress synergies in horticultural plants via redox regulation and epigenetic signaling. Sci Hort. 2023;321:12.
He WZ, Liu H, Li YP, Wu ZKY, Xie Y, Yan X, Wang XY, Miao Q, Chen TT, Rahman SU, et al. Genome-wide characterization of B-box gene family in Artemisia annua L. and its potential role in the regulation of artemisinin biosynthesis. Ind Crop Prod. 2023;199:13.
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R. TBtools: an integrative Toolkit developed for interactive analyses of big Biological Data. Mol Plant. 2020;13(8):1194–202.
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, Tosatto SCE, Paladin L, Raj S, Richardson LJ, et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 2021;49(D1):D412–9.
Ren M, Wang Q, Zhang FH, Wang Yl, Wang YY, Li W, Qi KJ Xie ZH, Zhang Sl, Tao ST: genome-wide identification of the GAox gene family and functional characterization of PbGA3ox4 during stone cell formation in Chinese white pear. Sci Hort 2024, 330.
Abubakar AS, Wu YM, Chen FM, Zhu AG, Chen P, Chen KM, Qiu XJ, Huang XY, Zhao HH, Chen JK, et al. Comprehensive Analysis of WUSCEL-Related Homeobox Gene Family in Ramie ( Boehmeria nivea) indicates its potential role in adventitious Root Development. Biology-Basel. 2023;12(12):18.
Wang YP, Tang HB, DeBarry JD, Tan X, Li JP, Wang XY, Lee TH, Jin HZ, Marler B, Guo H et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity</i >. Nucleic Acids Res 2012, 40(7).
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R. TBtools: an integrative Toolkit developed for interactive analyses of big Biological Data. Mol Plant. 2020;13(8):1194–202.
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing platforms. Mol Biol Evol. 2018;35(6):1547–9.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4(4):406–25.
Letunic I, Bork P. Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021;49(W1):W293–6.
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren JY, Li WW, Noble WS. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009;37:W202–8.
Zhu LM, Zeng D, Lei XC, Huang J, Deng YF, Ji YB, Liu J, Dai FF, Li YZ, Shi DD, et al. KLF2 regulates neutrophil migration by modulating CXCR1 and CXCR2 in asthma. Biochim Biophys Acta-Mol Basis Dis. 2020;1866(12):13.
Qiu X, Zhao H, Abubakar AS, Shao D, Chen J, Chen P, Yu C, Wang X, Chen K, Zhu A. Genome-wide analysis of AP2/ERF gene superfamily in Ramie ( Boehmeria nivea L.) revealed their synergistic roles in regulating abiotic stress resistance and Ramet Development. Int J Mol Sci 2022, 23(23).
Zhou XL, Lu JY, Ben W, Zhen G. HOXA11-AS facilitates the proliferation, cell cycle process and migration of keloid fibroblasts through sponging mir-188-5p to regulate VEGFA. J Dermatol Sci. 2022;106(2):111–8.
Yang YY, Shi LX, Li JH, Yao LY, Xiang DX. Piperazine ferulate ameliorates the development of diabetic nephropathy by regulating endothelial nitric oxide synthase. Mol Med Rep. 2019;19(3):2245–53.
Acknowledgements
The authors thank laboratory colleagues for their technical assistance. The authors thank the editors and reviewers for their rigorous evaluation of the manuscript.
Funding
This work was supported by Hunan Provincial Department of Education general project (No.22C0674), ESI Discipline Special Project of Changsha Medical University (No.2022CYY023 and No.2022CYY006), Science Research Project of Hunan Provincial Department of Education (No.23A0662), National Undergraduate Innovation and Entrepreneurship Training Program Project under Grant (No.202210823006), National Natural Science Foundation of China (32372092).
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.W. (Yue Wang); methodology, J.C. (Jiayi Chen) and Y.W. (Yue Wang); software, Y.W. (Yue Wang), X.H. (Xiaoyu Huang) and X.Q. (Xiaojun Qiu); validation, J.C. (Jiayi Chen), Y.W. (Yue Wang) and Y.W. (Yongmei Wu); resources, X.H. (Xiaoyu Huang), X.Q. (Xiaojun Qiu), Q.L. (Qian Lin), and H.Z. (Haohan Zhao); writing-original draft preparation, Y.W. (Yue Wang) and J.C. (Jiayi Chen); writing-review and editing, F.C. (Fengming Chen) and G.G. (Gang Gao). All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The plant material used in this study was A. venetum and A. hendersonii, grown in the laboratory of the Institute of bast fiber crops, Chinese Academy of Agricultural Sciences (CAAS) and no permits were required to collect plant samples. In the present study, all methods were carried out following relevant guidelines and regulations. Ethical approval or consent was not required for this study because no endangered or protected species were involved.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Chen, J., Wang, Y., Wu, Y. et al. Genome-wide identification and expression analysis of the PP2C gene family in Apocynum venetum and Apocynum hendersonii. BMC Plant Biol 24, 652 (2024). https://doi.org/10.1186/s12870-024-05328-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-024-05328-6