Abstract
Bloodstream infection (BSI) caused by bacteria is highly pathogenic and lethal, and easily develops whole-body inflammatory state. Immediate identification of disease-causing bacteria can improve patient prognosis. Traditional testing methods are not only time-consuming, but such tests are limited to laboratories. Recombinase polymerase amplification combined with lateral flow dipstick (RPA-LFD) holds great promise for rapid nucleic acid detection, but the uncapping operation after amplification easily contaminates laboratories. Therefore, the establishment of a more effective integrated isothermal amplification system has become an urgent problem to be solved. In this study, we designed and fabricated a hermetically sealed integrated isothermal amplification system. Combining with this system, a set of RPA-LFD assays for detecting S. aureus, K. peneumoniae, P. aeruginosa, and H. influenza in BSI were established and evaluated. The whole process could be completed in less than 15 min and the results can be visualized by the naked eye. The developed RPA-LFD assays displayed a good sensitivity, and no cross-reactivity was observed in seven similar bacterial genera. The results obtained with 60 clinical samples indicated that the developed RPA-LFD assays had high specifcity and sensitivity for identifying S. aureus, K. peneumoniae, P. aeruginosa, and H. influenza in BSI. In conclusion, our results showed that the developed RPA-LFD assay is an alternative to existing PCR-based methods for detection of S. aureus, K. peneumoniae, P. aeruginosa, and H. influenza in BSI in primary hospitals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bloodstream infection (BSI) caused by bacteria can develop clinical symptoms such as chills, high fever, tachycardia and even shock, which can be a serious threat to people’s health (Allerberger and Kern 2020; Kolesnichenko et al. 2021; Shukla et al. 2021). Studies have shown that applying antibiotics within one hour of monitoring a patient for hypotension is beneficial in improving patient survival, otherwise patient mortality increases progressively with the delay in administration (Kumar et al. 2006). Therefore, rapid and accurate identification of pathogenic bacteria is extremely crucial for subsequent medication guidance. For emergency, traditional culture method for diagnosing bacterial infection is not completely satisfactory due to the time-consuming procedures and high-level laboratories, which is unfavorable to timely and rational medication (Palavecino 2020; Peri et al. 2022). Although real-time polymerase chain reaction (real-time PCR) omits culture procedures and shortens turnaround time, real-time PCR still requires sophisticated thermal cycling instruments and is also difficult to be employed without centralized laboratories (Hawkins and Guest 2017; Singh and Roy-Chowdhuri 2016).
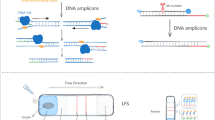
To further simplify operational procedures and improve clinical diagnostic performance, a novel isothermal amplification technique, called recombinase polymerase amplification (RPA), was introduced in this study. It relies on three core enzymes containing DNA polymerase, single strand DNA binding protein and recombinase to aid DNA amplification (Piepenburg et al. 2006). Typically, amplifying nucleic acids can be completed under 37–42℃ condition within 20 min (McQuillan and Wilson 2021; Zhang et al. 2021a; Zheng et al. 2021). Fluorescent-based and lateral flow dipstick (LFD)-based detection has been widely established for various targets among the detection format of RPA amplicons (Behrmann et al. 2020; Shelite et al. 2021; Wang et al. 2022; Xu et al. 2021). Herein, to meet the needs for rapid detection on first aid and emergency treatment, especially for resource-limited settings and poorly equipped laboratories, combining RPA assays and LFD strips (designated as RPA-LFD) is a desirable option.
In this study, Staphylococcus aureus, Pseudomonas aeruginosa, Klebsiella pneumonia and Haemophilus influenzae were selected as detection targets after reviewing literatures and the BSI bacteria profile of Daping Hospital (Cui et al. 2022; Lisowska-Łysiak et al. 2021; Martinez and Wolk 2016; Mendes et al. 2018; Stryjewski and Boucher 2009; Wisplinghoff et al. 2004). Therefore, a panel of RPA-LFD assays targeting the above four common pathogenic bacteria was developed. Additionally, we described a customized integrated isothermal amplification system for the RPA-LFD assays to minimize cross-contamination risk from reopening the lid after amplification and to ensure results’ accuracy. We determined the sensitivity and specificity of RPA-LFD assays before using the newly developed integrated isothermal amplification system to retrospectively detect 60 bacteria-spiked blood samples.
Materials and methods
Primers and NFO probes of RPA
After a systematic literature search and sequence alignment with DNAMAN software, target gene regions were identified for above four common pathogenic bacteria (Fig. S1). Nuc, CelB, Eta, and Fuck gene of S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae were respectively selected as the target genes (Gadsby et al. 2015; Jiang et al. 2020a; Meyler et al. 2012; Salman et al. 2013; Song et al. 2000; Tian et al. 2019). Primer sets and corresponding NFO probes were designed. All primers and probes of Basic RPA and RPA-LFD assays were synthesized and purified by Beijing Genomics Institute (BGI) Biotechnology Corporation using polyacrylamide gel electrophoresis (PAGE) and high-performance liquid chromatography (HPLC) respectively. The oligonucleotide sequences of primers and probes of Basic RPA and RPA-LFD assays were showed in Tables 1 and 2.
Bacterial strains and DNA preparation
Bacterial DNA for sensitivity and specificity assays were extracted from standard reference strains: S. aureus ATCC29213, K. pneumonia ATCC700603, P. aeruginosa ATCC27853, and H. influenza ATCC49247. DNA of all these bacteria was extracted using TIANamp Bacteria DNA Kit (Tiangen Biotech Co., Ltd., Beijing, China) according to the manufacturer’s instructions (Ahmad et al. 2020). All 60 bacteria-spiked blood samples were collected from Daping Hospital. Bacteria-spiked blood samples were firstly lysed using Red Blood Cell Lysis Buffer (Sansure Biotech Inc., Hunan, China) before bacterial DNA was extracted by TIANamp Bacteria DNA Kit. The extracted DNA was stored at -80℃ until next use.
RPA conditions
A series of Basic RPA assays were carried out to screen out the best primer sets producing the highest analytical sensitivity. The best primers were chosen for subsequent RPA-LFD experiments. Basic RPA and RPA-LFD reactions were both performed in 50 µL volume using DNA Basic kits and LFD kits respectively (Amp-Future Biotech Co., Ltd., Weifang, China) (Sun et al. 2021). Each basic RPA reaction contained 29.4 µL A buffer, 2 µL forward primer (10 µM), 2 µL reverse primer (10 µM), 2 µL sample, 12.1 µL nuclease-free water. Each RPA-LFD reaction included 29.4 µL A buffer, 2 µL forward primer (10 µM), 2 µL reverse primer (10 µM), 0.6 µL NFO probe (10 µM), 5 µL DNA sample, 8.5 µL nuclease-free water. The above mixed reagents were transferred to the reaction tubes containing a dried enzyme pellet provided by the kit, subsequently 2.5 µL B buffer (280 mM) was added to lids. The tubes were closed carefully, vortexed and centrifuged briefly. They were immediately placed in a matched metal heat block and incubated at 38℃ for 10 min. Nuclease-free water was used as the negative control. Finally, the Basic RPA products were purified by phenol-chloroform method (Solarbio, Beijing, China) and analyzed by 1.5% agarose gel electrophoresis. RPA-LFD products were detected by LFD strips (Amp-Future Biotech Co., Ltd., Weifang, China). The amplicons of RPA-LFD were diluted 20-fold in buffer (Milenia Biotec GmbH, Germany). Then LFD strips were placed vertically in tubes containing the diluted RPA-LFD products for 5 min.
Specificity and sensitivity of RPA-LFD assays
Each RPA-LFD assay’s specificity was verified by detecting seven similar bacterial genera. A series of bacteria for detecting possible cross reactions were shown in Table S1. DNA extracted from reference bacterial strains was used as the positive control. To evaluate established RPA-LFD assays’ sensitivity from bacteria-spiked blood samples, bacterial DNA was 10-fold diluted to prepare samples of final serial concentrations ranging from 6 × 105 CFU/mL to 6 CFU/mL. RPA-LFD reactions were prepared according to above RPA-LFD conditions. The experiment was repeated three times for low concentration samples.
Real-time PCR assay
Real-time PCR master mix was prepared, and reaction procedures were set according to the instructions of the real-time PCR kit (Promega, USA). The primers and probes of real-time PCR used in the study are shown in Table S2 (Gadsby et al. 2015; Salman et al. 2013). The reaction was performed on CFX96 real-time PCR detection system (Bio-Rad, USA).
Evaluating bacteria-spiked blood samples for RPA-LFD assays with the integrated isothermal amplification system
In order to keep RPA-LFD assays in a closed environment and reduce the problem of cross-contamination, an integrated isothermal amplification system was designed and applied in the RPA-LFD assays. The system integrates RPA assays, LFD strips detecting amplicons, detection devices, and matched metal heat blocks, where the detection device consists of a top cover and a bottom container including a bottom pad, a reaction chamber and an inspection window (Fig. S2). After amplification, results can be interpreted by visualizing the presence or absence of the corresponding bands on the LFD strips with the naked eye. As shown in Figs. 1, 5 µL of DNA template was added to the premixed RPA reaction buffer. Then, the assay device was incubated in metal heat blocks at 38 °C for 10 min. Thereafter, the device was tilted so that the LFD strip located at the inspection window was immersed in the diluted amplicons and the results were interpreted visually based on the presence or absence of bands in the test and control lines. With this integrated isothermal amplification system, the whole process could be completed in less than 15 min, from the time the DNA sample was added into the device to when the reaction result was interpretated. Furthermore, the whole experiment was carried out step by step in a closed space, without aerosol contamination. To perform clinical sample evaluation experiments, 60 clinical samples were collected to construct bacteria-spiked blood samples to verify the feasibility of this new system. The performance of this novel integrated isothermal amplification system was compared with that of mass spectrometry.
Bacteria-spiked blood samples operation procedures using the integrated isothermal amplification system. (A) Placed rubber rings on the detection device and inserted LFD at inspection windows. (B-C) RPA reaction mixtures and buffer were added to left channel and right channel of bottom container respectively. (D) Inserted top cover into the reaction chamber and sealed with sealing films. (E) The assembled detection device was heated in a matched metal heat block at 38 °C for 10 min. (F-G) Bottom pad was pierced by top cover. Then, tilting and gently shaking the detection device. (H) Observed test results
Specificity of RPA-LFD assays. A, B, C, D and E represent the results of S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae RPA-LFD specificity assays, respectively. Results showed that only positive control samples and targeted bacterial samples produced amplification signals, whereas the other pathogen samples and the negative control produced no amplification signals. NC: negative control; C: control line; T: test line
Sensitivity of RPA-LFD assays. A, B, C, D and E represent the results of S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae RPA-LFD sensitivity assays, respectively. Serially diluted DNA concentration of targeted bacteria (6 × 105 CFU/mL, 6 × 104 CFU/mL, 6 × 103 CFU/mL, 6 × 102 CFU/mL, 60 CFU/mL and 6 CFU/mL per reaction) was tested by RPA-LFD assays at 38 °C for 10 min. This experiment was repeated three times for low-concentrated samples (60 CFU/mL-6 CFU/mL). NC: negative control; C: control line; T: test line
Evaluating bacteria-spiked blood samples for RPA-LFD assays with the integrated isothermal amplification system. A: 1, 2, 3, and 4 represent A. baumannii, K. peneumoniae, P. aeruginosa, and H. influenzae; 5 represents negative control; 6–15 represent the clinical isolated S. aureus samples. B: 1, 2, 3, and 4 represent A. baumannii, P. aeruginosa, H. influenzae, and S. aureus; 5 represents negative control; 6–15 represent the clinical isolated K. peneumoniae samples. C: 1, 2, 3, and 4 represent A. baumannii, K. peneumoniae, H. influenzae, and S. aureus; 5 represents negative control; 6–15 represent the clinical isolated P. aeruginosa samples. D: 1, 2, 3, and 4 represent A. baumannii, K. peneumoniae, P. aeruginosa, and S. aureus; 5 represents negative control; 6–15 represent the clinical isolated H. influenzae samples
Results
Primer screening and identification
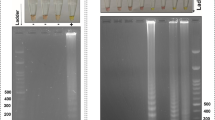
RPA is a multienzyme-assisted isothermal amplification technique where primers play an important role in unwinding templates, and different primer combinations produce different amplification effects. Therefore, a series of primer screening experiments are essential. Several sets of forward and reverse primer screens were conducted according to the manufacturer’s instructions. As shown in Fig. S3, the best primer sets for S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae were identified as R2/F3, R3/F2, R3/F2, and R1/F1 based on the recommendations of RPA and the brightness of the electrophoretic bands. The four selected primer sets were used for subsequent RPA-LFD experiments.
Specificity and sensitivity of RPA-LFD assays
The specificity of RPA-LFD assays for four bacteria species was confirmed by testing seven similar bacterial genera. As can be seen in Fig. 2, only positive control and target bacteria showed test lines, demonstrating that established RPA-LFD reactions have good specificity and no cross reactions occurred. The sensitivity of RPA-LFD assays was determined using a concentration of 6 × 105 CFU/mL to 6 CFU/mL of bacterial DNA extracted from bacteria-spiked blood samples. The results showed that RPA-LFD assays can detect S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae with a sensitivity as low as 600 CFU/mL, 60 CFU/mL, 60 CFU/mL and 6 CFU/mL respectively (Fig. 3). By naked eye, the RPA-LFD assays showed sensitivity ranging from 6 × 102 to 6 CFU/mL. The sensitivity of real-time PCR ranged from 6 × 103 to 60 CFU/mL (Fig. S4). Therefore, the sensitivity of RPA-LFD assays was significantly better than that of real-time PCR method.
Evaluating bacteria-spiked blood samples for RPA-LFD assays with the integrated isothermal amplification system
After the RPA-LFD assays were verified, 10 S. aureus-spiked blood samples, 10 K. peneumoniae-spiked blood samples, 10 P. aeruginosa-spiked blood samples, and 10 H. influenzae-spiked blood samples were used for clinical confirmation. In addition, 5 negative controls bacteria-spiked blood samples were made for each RPA-LFD assay to ensure accuracy. From the Fig. 4, it can be seen that the newly RPA-LFD assay had high specifcity and sensitivity for identifying S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae in BSI, which may become a powerful tool for rapid and reliable diagnosis of BSI caused by these common pathogenic bacteria in primary hospitals.
Discussion
Bacterial infections and secondary serious infections from diseases are important causes of death among the elderly, infirm and disabled. Traditional methods and sequencing-based assays for bacterial identification are time-consuming and instrument-intensive, as well as require trained staff, which are not suitable for low-resource areas (Boolchandani et al. 2019; Li et al. 2018; Otašević et al. 2018).
Isothermal amplification techniques, which require much less sophisticated amplification instruments and possess fast reaction kinetics, have been rapidly developed in recent years (Craw and Balachandran 2012; Li and Macdonald 2015; Zhao et al. 2015). Compared with other isothermal amplification assays, the RPA technique used in this study has the shortest reaction time, the simplest experimental design, and the lowest energy consumption (Crannell et al. 2014; Kong et al. 2019). Some studies have shown that experiments can be done using human body temperature (Natoli et al. 2021; Schuler et al. 2015). In just 16 years since its invention in 2006, RPA has reported extensive research on pathogenic microbes and other aspects (Ivanov et al. 2021; Jiang et al. 2020b; Koo et al. 2016; Wang et al. 2021).
In order to develop a set of experimental platforms capable of detecting common bacterial infections for timely treatment of infections, several pairs of primers and their corresponding probes were designed for four bacteria species (S. aureus, K. peneumoniae, P. aeruginosa, and H. influenzae). The best primer pairs were identified by forward and reverse primer screening experiments using agarose gel electrophoresis and used for subsequent RPA-LFD experiments. The results of specificity assays showed that the four established RPA-LFD assays had no cross reactions with other bacterial species. DNA from bacteria-spiked blood samples was tested for RPA-LFD sensitivity assays. In contrast to previous studies, the matrix of the test samples less involved blood samples, which are an important sample matrix for diagnosing infectious diseases (Chen et al. 2018; Ghosh et al. 2012; Helfrich et al. 2015). Here, to evaluate the practical clinical application of the established RPA-LFD assays, blood samples were used as sample matrix to construct clinical bacteremia samples. Our results showed the sensitivity of established RPA-LFD assays can up to 60 CFU/mL, and even 6 CFU/mL, which was better than that of real-time PCR method. However, the limitation of this study is that the relatively low sensitivity of RPA-LFD in the detection of S. aureus. The possible reasons for the low sensitivity may be related to the colony traits of the bacteria themselves, and corresponding nucleic acid extraction protocols should be optimized according to characteristics of different bacteria (Barbaccia et al. 2020; Bogut and Magryś 2021; Chiarelli et al. 2020; Mizukami et al. 2020).
In order to achieve simple and rapid operation, we designed an integrated isothermal amplification system, which consists of RPA assays, LFD strips detecting amplicons, detection devices, and matched metal heat blocks. This system greatly simplifies the detection process of traditional RPA-LFD technique and effectively avoids the contamination caused by amplification products. At the same time, the system developed by us can preliminarily realize the function of point-of-care testing (Chakravorty et al. 2017; Taki et al. 2021; Zhang et al. 2021b). In short, compared with the conventional techniques, such as real-time quantitative PCR, our detection technology has the advantages of simple equipment, rapid operation, and low cost. With the further decline in the price of detection reagents such as recombinant enzymes in the future, the technology will have a lower cost and produce better benefits. Of course, we will continue to improve this work in the future.
In summary, the RPA-LFD assay is time-saving, more effective and sensitive than conventional identification methods, which has the potential to be applied in primary hospitals (Chen et al. 2021; Yin et al. 2017). Moreover, this novel integrated isothermal amplification system will become a powerful tool for the identification of bacteria or other pathogens, especially suitable for use in low-resource settings (Sadaow et al. 2020).
Data Availability
Data of this study are included in the article and the primary data can be provided from the corresponding author.
References
Ahmad T, Ullah S, Moosa A, Liu Y, Chengrong N, Shujian H (2020) First Report of Pre-harvest Soft Rot of Peach Fruit (Prunus persica) caused by Enterobacter mori in China. Plant Dis Aug 18. https://doi.org/10.1094/PDIS-06-20-1285-PDN
Allerberger F, Kern WV (2020) Bacterial bloodstream Infection. Clin Microbiol Infect 26:140–141. https://doi.org/10.1016/j.cmi.2019.10.004
Barbaccia P, Francesca N, Gerlando RD, Busetta G, Moschetti G, Gaglio R, Settanni L (2020) Biodiversity and dairy traits of indigenous milk lactic acid bacteria grown in presence of the main grape polyphenols. FEMS Microbiol Lett 367. https://doi.org/10.1093/femsle/fnaa066
Behrmann O, Bachmann I, Spiegel M, Schramm M, Abd El Wahed A, Dobler G, Dame G, Hufert FT (2020) Rapid Detection of SARS-CoV-2 by low volume real-time single tube reverse transcription recombinase polymerase amplification using an Exo probe with an internally linked Quencher (Exo-IQ). Clin Chem 66:1047–1054. https://doi.org/10.1093/clinchem/hvaa116
Bogut A, Magryś A (2021) The road to success of coagulase-negative staphylococci: clinical significance of small colony variants and their pathogenic role in persistent Infections. Eur J Clin Microbiol Infect Dis 40:2249–2270. https://doi.org/10.1007/s10096-021-04315-1
Boolchandani M, D’Souza AW, Dantas G (2019) Sequencing-based methods and resources to study antimicrobial resistance. Nat Rev Genet 20:356–370. https://doi.org/10.1038/s41576-019-0108-4
Chakravorty S, Simmons AM, Rowneki M, Parmar H, Cao Y, Ryan J, Banada PP, Deshpande S, Shenai S, Gall A, Glass J, Krieswirth B, Schumacher SG, Nabeta P, Tukvadze N, Rodrigues C, Skrahina A, Tagliani E, Cirillo DM, Davidow A, Denkinger CM, Persing D, Kwiatkowski R, Jones M, Alland D (2017) The New Xpert MTB/RIF Ultra: improving detection of Mycobacterium tuberculosis and Resistance to Rifampin in an assay suitable for point-of-care testing. mBio 8. https://doi.org/10.1128/mBio.00812-17
Chen J, Xu Y, Yan H, Zhu Y, Wang L, Zhang Y, Lu Y, Xing W (2018) Sensitive and rapid detection of pathogenic bacteria from urine samples using multiplex recombinase polymerase amplification. Lab Chip 18:2441–2452. https://doi.org/10.1039/c8lc00399h
Chen N, Si Y, Li G, Zong M, Zhang W, Ye Y, Fan L (2021) Development of a loop-mediated isothermal amplification assay for the rapid detection of six common respiratory viruses. Eur J Clin Microbiol Infect Dis 40:2525–2532. https://doi.org/10.1007/s10096-021-04300-8
Chiarelli A, Cabanel N, Rosinski-Chupin I, Zongo PD, Naas T, Bonnin RA, Glaser P (2020) Diversity of mucoid to non-mucoid switch among carbapenemase-producing Klebsiella pneumoniae. BMC Microbiol 20:325. https://doi.org/10.1186/s12866-020-02007-y
Crannell ZA, Rohrman B, Richards-Kortum R (2014) Equipment-free incubation of recombinase polymerase amplification reactions using body heat. PLoS ONE 9:e112146. https://doi.org/10.1371/journal.pone.0112146
Craw P, Balachandran W (2012) Isothermal nucleic acid amplification technologies for point-of-care diagnostics: a critical review. Lab Chip 12:2469–2486. https://doi.org/10.1039/c2lc40100b
Cui J, Li M, Cui J, Wang J, Qiang X, Liang Z (2022) The proportion, species distribution and dynamic trends of bloodstream Infection cases in a tertiary hospital in China, 2010–2019. Infection 50:121–130. https://doi.org/10.1007/s15010-021-01649-y
Gadsby NJ, McHugh MP, Russell CD, Mark H, Conway Morris A, Laurenson IF, Hill AT, Templeton KE (2015) Development of two real-time multiplex PCR assays for the detection and quantification of eight key bacterial pathogens in Lower Respiratory Tract Infections. Clin Microbiol Infect 21:788e1. -788.e13
Ghosh C, Jain I, Shinde CP, Chakraborty BS (2012) Rapid and sensitive liquid chromatography/tandem mass spectrometry method for simultaneous determination of enalapril and its major metabolite enalaprilat, in human plasma: application to a bioequivalence study. Drug Test Anal 4:94–103. https://doi.org/10.1002/dta.241
Hawkins S, Guest PC (2017) Multiplex analyses using real-time quantitative PCR. Methods Mol Biol 1546:125–133. https://doi.org/10.1007/978-1-4939-6730-8_8
Helfrich YR, Maier LE, Cui Y, Fisher GJ, Chubb H, Fligiel S, Sachs D, Varani J, Voorhees J (2015) Clinical, Histologic, and Molecular Analysis of Differences between Erythematotelangiectatic Rosacea and Telangiectatic Photoaging. JAMA Dermatol 151:825–836. https://doi.org/10.1001/jamadermatol.2014.4728
Ivanov AV, Safenkova IV, Zherdev AV, Dzantiev BB (2021) Recombinase polymerase amplification assay with and without nuclease-dependent-labeled Oligonucleotide Probe. Int J Mol Sci 22. https://doi.org/10.3390/ijms222111885
Jiang L, Li X, Gu R, Mu D (2020a) Nanoparticles-based Biosensor coupled with Multiplex Loop-mediated isothermal amplification for detection of Staphylococcus aureus and Identification of Methicillin-Resistant S. Aureus. Infect Drug Resist 13:1251–1262. https://doi.org/10.2147/IDR.S243881
Jiang L, Ching P, Chao CC, Dumler JS, Ching WM (2020b) Development of a sensitive and Rapid Recombinase Polymerase Amplification Assay for Detection of Anaplasma phagocytophilum. J Clin Microbiol 58. https://doi.org/10.1128/JCM.01777-19
Kolesnichenko SI, Lavrinenko AV, Akhmaltdinova LL (2021) Bloodstream Infection etiology among children and adults. Int J Microbiol 2021:6657134. https://doi.org/10.1155/2021/6657134
Kong M, Li Z, Wu J, Hu J, Sheng Y, Wu D, Lin Y, Li M, Wang X, Wang S (2019) A wearable microfluidic device for rapid detection of HIV-1 DNA using recombinase polymerase amplification. Talanta 205:120155. https://doi.org/10.1016/j.talanta.2019.120155
Koo KM, Wee EJ, Trau M (2016) Colorimetric TMPRSS2-ERG Gene Fusion detection in Prostate Cancer urinary samples via recombinase polymerase amplification. Theranostics 6:1415–1424. https://doi.org/10.7150/thno.15250
Kumar A, Roberts D, Wood KE, Light B, Parrillo JE, Sharma S, Suppes R, Feinstein D, Zanotti S, Taiberg L, Gurka D, Kumar A, Cheang M (2006) Duration of hypotension before initiation of effective antimicrobial therapy is the critical determinant of survival in human septic shock. Crit Care Med 34:1589–1596. https://doi.org/10.1097/01.CCM.0000217961.75225.E9
Li J, Macdonald J (2015) Advances in isothermal amplification: novel strategies inspired by biological processes. Biosens Bioelectron 64:196–211. https://doi.org/10.1016/j.bios.2014.08.069
Li J, Macdonald J, von Stetten F (2018) Review: a comprehensive summary of a decade development of the recombinase polymerase amplification. Analyst 144:31–67. https://doi.org/10.1039/c8an01621f
Lisowska-Łysiak K, Lauterbach R, Międzobrodzki J, Kosecka-Strojek M (2021) Epidemiology and Pathogenesis of Staphylococcus Bloodstream Infections in humans: a review. Pol J Microbiol 70:13–23. https://doi.org/10.33073/pjm-2021-005
Martinez RM, Wolk DM (2016) Bloodstream Infections. Microbiol Spectr 4. https://doi.org/10.1128/microbiolspec.DMIH2-0031-2016
McQuillan JS, Wilson MW (2021) Recombinase polymerase amplification for fast, selective, DNA-based detection of faecal indicator Escherichia Coli. Lett Appl Microbiol 72:382–389. https://doi.org/10.1111/lam.13427
Mendes RE, Sader HS, Castanheira M, Flamm RK (2018) Distribution of main Gram-positive pathogens causing bloodstream Infections in United States and European hospitals during the SENTRY Antimicrobial Surveillance Program (2010–2016): concomitant analysis of oritavancin in vitro activity. J Chemother 30:280–289. https://doi.org/10.1080/1120009X.2018.1516272
Meyler KL, Meehan M, Bennett D, Cunney R, Cafferkey M (2012) Development of a diagnostic real-time polymerase chain reaction assay for the detection of invasive Haemophilus influenzae in clinical samples. Diagn Microbiol Infect Dis 74:356–362. https://doi.org/10.1016/j.diagmicrobio.2012.08.018
Mizukami K, Iwasaki Y, Kawakami E, Hirata M, Kamatani Y, Matsuda K, Endo M, Sugano K, Yoshida T, Murakami Y, Nakagawa H, Spurdle AB, Momozawa Y (2020) Genetic characterization of Pancreatic cancer patients and prediction of carrier status of germline pathogenic variants in cancer-predisposing genes. EBioMedicine 60:103033. https://doi.org/10.1016/j.ebiom.2020.103033
Natoli ME, Chang MM, Kundrod KA, Coole JB, Airewele GE, Tubman VN, Richards-Kortum RR (2021) Allele-specific recombinase polymerase amplification to detect Sickle Cell Disease in Low-Resource settings. Anal Chem 93:4832–4840. https://doi.org/10.1021/acs.analchem.0c04191
Otašević S, Momčilović S, Stojanović NM, Skvarč M, Rajković K, Arsić-Arsenijević V (2018) Non-culture based assays for the detection of fungal pathogens. J Mycol Med 28:236–248. https://doi.org/10.1016/j.mycmed.2018.03.001
Palavecino EL (2020) Rapid Methods for Detection of MRSA in clinical specimens. Methods Mol Biol 2069:29–45. https://doi.org/10.1007/978-1-4939-9849-4_2
Peri AM, Harris P, Paterson DL (2022) Culture-independent detection systems for bloodstream Infection. Clin Microbiol Infect 28:195–201. https://doi.org/10.1016/j.cmi.2021.09.039
Piepenburg O, Williams CH, Stemple DL, Armes NA (2006) DNA detection using recombination proteins. PLoS Biol 4:e204. https://doi.org/10.1371/journal.pbio.0040204
Sadaow L, Sanpool O, Rodpai R, Boonroumkaew P, Maleewong W, Intapan PM (2020) Development of immunochromatographic device as a point-of-care tool for serodiagnosis of human strongyloidiasis cases. Eur J Clin Microbiol Infect Dis 39:465–470. https://doi.org/10.1007/s10096-019-03745-2
Salman M, Ali A, Haque A (2013) A novel multiplex PCR for detection of Pseudomonas aeruginosa: a major cause of wound Infections. Pak J Med Sci 29:957–961. https://doi.org/10.12669/pjms.294.3652
Schuler F, Schwemmer F, Trotter M, Wadle S, Zengerle R, von Stetten F, Paust N (2015) Centrifugal step emulsification applied for absolute quantification of nucleic acids by digital droplet RPA. Lab Chip 15:2759–2766. https://doi.org/10.1039/c5lc00291e
Shelite TR, Bopp NE, Moncayo A, Reynolds ES, Thangamani S, Melby PC, Bloch K, Aguilar PV, Travi BL (2021) Isothermal recombinase polymerase amplification-lateral Flow Point-of-Care Diagnostic Test for Heartland Virus. Vector Borne Zoonotic Dis 21:110–115. https://doi.org/10.1089/vbz.2020.2670
Shukla BS, Warde PR, Knott E, Arenas S, Pronty D, Ramirez R, Rego A, Levy M, Zak M, Parekh DJ, Ferreira T, Gershengorn HB (2021) Bloodstream Infection risk, incidence, and deaths for hospitalized patients during Coronavirus Disease Pandemic. Emerg Infect Dis 27:2588–2594. https://doi.org/10.3201/eid2710.210538
Singh C, Roy-Chowdhuri S (2016) Quantitative real-time PCR: recent advances. Methods Mol Biol 1392:161–176. https://doi.org/10.1007/978-1-4939-3360-0_15
Song KP, Chan TK, Ji ZL, Wong SW (2000) Rapid identification of Pseudomonas aeruginosa from ocular isolates by PCR using exotoxin A-specific primers. Mol Cell Probes 14:199–204. https://doi.org/10.1006/mcpr.2000.0306
Stryjewski ME, Boucher HW (2009) Gram-negative bloodstream Infections. Int J Antimicrob Agents 34 Suppl 4S21–25. https://doi.org/10.1016/S0924-8579(09)70561-8
Sun ML, Lai HY, Chong NY, Liu DF, Zhang ZY, Pang B, Yao J (2021) Simple and feasible detection of Hepatitis B Virus via Combination of Multienzyme Isothermal Rapid Amplification and lateral Flow Dipstick Strip. Front Mol Biosci 8:763079. https://doi.org/10.3389/fmolb.2021.763079
Taki K, Yokota I, Fukumoto T, Iwasaki S, Fujisawa S, Takahashi M, Negishi S, Hayasaka K, Sato K, Oguri S, Nishida M, Sugita J, Konno S, Saito T, Teshima T (2021) SARS-CoV-2 detection by fluorescence loop-mediated isothermal amplification with and without RNA extraction. J Infect Chemother 27:410–412. https://doi.org/10.1016/j.jiac.2020.10.029
Tian Y, Wang L, Zhang J, Han Q, Xia XS, Song Y, Yang G (2019) CelB is a suitable marker for rapid and specific identification of Klebsiella pneumoniae by the loop-mediated isothermal amplification (LAMP) assay. Braz J Microbiol 50:961–967. https://doi.org/10.1007/s42770-019-00144-9
Wang F, Wang L, Chen H, Li N, Wang Y, Li Y, Liang W (2021) Rapid Detection of Bla (KPC), bla (NDM), bla (OXA-48-like) and bla (IMP) carbapenemases in Enterobacterales using recombinase polymerase amplification combined with lateral Flow Strip. Front Cell Infect Microbiol 11:772966. https://doi.org/10.3389/fcimb.2021.772966
Wang H, Dong J, Zhang T, Wang F, Yang R, Zhang Y, Zhao X (2022) A novel rapid detection of Senecavirus A using recombinase polymerase amplification (RPA) coupled with lateral flow (LF) dipstrip. Anal Biochem 646:114627. https://doi.org/10.1016/j.ab.2022.114627
Wisplinghoff H, Bischoff T, Tallent SM, Seifert H, Wenzel RP, Edmond MB (2004) Nosocomial bloodstream Infections in US hospitals: analysis of 24,179 cases from a prospective nationwide surveillance study. Clin Infect Dis 39:309–317. https://doi.org/10.1086/421946
Xu Y, Wu P, Zhang H, Li J (2021) Rapid detection of Mycobacterium tuberculosis based on antigen 85B via real-time recombinase polymerase amplification. Lett Appl Microbiol 72:106–112. https://doi.org/10.1111/lam.13364
Yin F, Liu J, Liu A, Li Y, Luo J, Guan G, Yin H (2017) Rapid diagnosis of Theileria annulata by recombinase polymerase amplification combined with a lateral flow strip (LF-RPA) in epidemic regions. Vet Parasitol 237:125–129. https://doi.org/10.1016/j.vetpar.2017.02.019
Zhang B, Zhu Z, Li F, Xie X, Ding A (2021a) Rapid and sensitive detection of Hepatitis B virus by lateral flow recombinase polymerase amplification assay. J Virol Methods 291:114094. https://doi.org/10.1016/j.jviromet.2021.114094
Zhang Y, Li Q, Guo J, Li D, Wang L, Wang X, Xing G, Deng R, Zhang G (2021b) An Isothermal Molecular Point of Care Testing for African Swine Fever Virus using recombinase-aided amplification and lateral Flow Assay without the need to extract nucleic acids in blood. Front Cell Infect Microbiol 11:633763. https://doi.org/10.3389/fcimb.2021.633763
Zhao Y, Chen F, Li Q, Wang L, Fan C (2015) Isothermal amplification of nucleic acids. Chem Rev 115:12491–12545. https://doi.org/10.1021/acs.chemrev.5b00428
Zheng YZ, Chen JT, Li J, Wu XJ, Wen JZ, Liu XZ, Lin LY, Liang XY, Huang HY, Zha GC, Yang PK, Li LJ, Zhong TY, Liu L, Cheng WJ, Song XN, Lin M (2021) Reverse transcription recombinase-aided amplification assay with lateral Flow Dipstick Assay for Rapid Detection of 2019 Novel Coronavirus. Front Cell Infect Microbiol 11:613304. https://doi.org/10.3389/fcimb.2021.613304
Acknowledgements
We appreciate the generosity of Chongqing Songshan Hospital for providing the standard strain K. pneumonia ATCC700603.
Funding
This work was supported by the Military Medical Frontier Innovation Ability Training Program (No. 2019CXJSC018); Chongqing Medical Scientific Research Project (Joint Project of Chongqing Health Commission and Science and Technology Bureau) (No. 2020MSXM021); Chongqing Medical Scientific Research Project (Joint Project of Chongqing Health Commission and Science and Technology Bureau) (No. 2022QNXM034); Biosafety Construction Project (No. A3702022001); and University Outstanding Talent Support Program.
Author information
Authors and Affiliations
Contributions
JL and MS participated in the research design, performed laboratory work and wrote the manuscript; WL conceptualized the research design and provided of study materials, reagents; QH conceptualized the research design and revised of the manuscript critically. ML and JL are responsible for sample collection. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Ethics Committee of Daping Hospital (Approval date: 16/11/2021, number: 2021 − 223). Informed consent was received from all participating patients in the present study.
Consent for publication
All the authors consented to the publication of this article.
Competing interests
The authors declared that there are no conficts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, J., Shang, MY., Deng, SL. et al. Development of a novel integrated isothermal amplification system for detection of bacteria-spiked blood samples. AMB Expr 13, 135 (2023). https://doi.org/10.1186/s13568-023-01643-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13568-023-01643-7