Abstract
Carcinogenesis is a complicated process that involves the deregulation of epigenetics resulting in cellular transformational events, such proliferation, differentiation, and metastasis. Epigenetic machinery changes the accessibility of chromatin to transcriptional regulation through DNA modification. The collaboration of epigenetics and gene transcriptional regulation creates a suitable microenvironment for cancer development, which is proved by the alternation in cell proliferation, differentiation, division, metabolism, DNA repair and movement. Therefore, the reverse of epigenetic dysfunction may provide a possible strategy and new therapeutic targets for cancer treatment. Many dietary components such as sulforaphane and epigallocatechin- 3-gallate have been demonstrated to exert chemopreventive influences, such as reducing tumor growth and enhancing cancer cell death. Anticancer mechanistic studies also indicated that dietary components could display the ability to reverse epigenetic deregulation in assorted tumors via reverting the adverse epigenetic regulation, including alternation of DNA methylation and histone modification, and modulation of microRNA expression. Therefore, dietary components as therapeutic agents on epigenetics becomes an attractive approach for cancer prevention and intervention at the moment. In this review, we summarize the recent discoveries and underlying mechanisms of the most common dietary components for cancer prevention via epigenetic regulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1. Introduction
Epigenetic alternation is referred to the change in gene expression without changing in DNA sequence. Epigenetic modifications include DNA methylation, histone modification (acetylation, methylation, and phosphorylation), and microRNA expression, playing a critical role in many cellular processes [1-4]. The initiation and progression of cancer have been proved as the results of the accumulation of genetic mutations which lead to aberrant cellular functions [3]. Genetic mutations may cause the activation of oncogenes and the inactivation of tumor suppressor genes. In recent decades, dietary components which are widely present in daily dietary, have been proved to exhibit a beneficial effect in cancer prevention and treatment. They reverse epigenetic deregulation by modulation of DNA methylation and histone modification, as well as alternation of microRNA expression [4-7]. This review will introduce some common dietary components possessing anticancer effects and elucidate their mechanisms.
2. Epigenetic modifications
2.1. DNA methylation
In mammalian cells, DNA methylation almost occurs at the 5’ position of cytosine residues within cytosine-phosphate-guanine (CpG) island dinucleotides [8], which is catalyzed by DNA methyltransferases (DNMTs) and S-adenosyl-methionine (SAM) as the methyl donor [9]. So far, five DNMTs have been identified, including DNMT1, DNMT2, DNMT3a, DNMT3b, and DNMT3L, and only three possess catalytic methyltransferase activity (DNMT1, DNMT3a, and DNMT3b) [9, 10]. DNMT1 is responsible for maintaining existed methylation patterns during DNA replication by adding methyl groups to corresponding daughter strands at the hemimethylated CpG sites. DNMT3a and DNMT3b carry out de novo methylation and preferentially target unmethylated CpG sites. DNMT3L lacks intrinsic methyltransferase activity, but can facilitate the methylation of retrotransposons by interaction with DNMT3a and 3b [9]. DNMT3L also recognizes the unmethylated lysine 4 of histone H3 to enhance the de novo methylation of DNA by the recruitment and activation of DNMT3a [11]. In normal cells, CpG islands of most genes are maintained in unmethylated, allowing RNA polymerase II to bind and transcription to proceed. Cancer cells often maintain in global hypomethylation status and hypermethylation of promoter CpG islands. Global hypomethylation can lead to chromosomal instability and mutations and result in the re-activation of oncogenes. DNA hypermethylation of the promoter CpG islands of tumor suppressor genes causes their transcriptional silencing [12]. Hence, aberrant promoter methylation allows the cancer cells to have a potent ability in growth and invasion [2, 3, 9].
2.2. Histone modifications
DNA is tightly compacted by histone proteins [13]. Octamer of histone proteins (two molecules of each H2A, H2B, H3, and H4) are wrapped with 146 bp of DNA to form a nucleosome. Histone proteins regulate the chromatin dynamic via changing chromatin structure by electrostatic charge alternation or providing protein recognition sites for modification [14, 15]. Histone modification occurs at the N-terminal tail, which subsequently affects DNA processes, including transcription, DNA repair, and DNA replication. Histones can be modified post-translationally by acetylation, methylation, phosphorylation, sumoylation, biotinylation, ubiquitination, ADP-ribosylation, deamination, proline isomerization, and propionylation [16-19].
Chromatin can exist in two different states, closed configuration (heterochromatin) and open configuration (euchromatin). The closed chromatin configuration is hard to access for the transcriptional machinery and generally harbor transcriptionally inactive genes. Histones (mainly H3) can be acetylated at lysine residues of N-terminal tails, which regulates chromatin into open or closed form. Acetylation neutralizes the positive charge of histones followed by dis-association from the negatively charged DNA backbone, leading to the open chromatin structure and making accessible to transcriptional machinery. Thereby, histone acetylation is generally associated with transcriptional activation [17-19]. Histone acetylation is carried out by histone acetyltransferases (HATs) and histone deacetylases (HDACs). HATs catalyze histone acetylation, while HDACs catalyze deacetylation by the removal of acetyl group, resulting in compact chromatin configuration and restricting the transcription factor access, and thereby inhibiting gene expression [18, 19]. HATs are classified into five families, Gcn5-related N-acetyltransferase (GNAT), MYST, p300/CBP, nuclear receptor coactivators (SRC), and TAFII250 families [20]. So far, eighteen human HDACs have been identified and divided into four classes, including Class I, II, III, and IV [20].
Methylation is also involved in histone modification. Unlike acetylation, methylation does not change the protein charge [16, 21]. Histone methylation occurs in lysine and arginine residues, which are catalyzed by histone lysine methyltransferase and histone arginine methyltransferase respectively [22]. Until now, twenty-four sites of methylation of histone are recognized, including 17 lysine residues and 7 arginine residues [23]. Lysine residues can be mono-, di-, or trimethylated, whereas arginine residues are mono- or dimethylated. The methylation of lysine is responsible for the alternation of chromatin structure [21]. Monomethylation of histone 3 (H3) at lysine 4 (H3K4me1) and trimethylation of H3 at lysine 4 (H3K4me3) are associated with transcriptional activation, whereas trimethylation of histone H3 at lysine 9 (H3K9me3), lysine 20 (H3K20me3), and lysine 27 (H3K27me3) are correlated with transcriptional inactivation [14]. Dysregulation in polycomb repressive complex 2 -mediated histone H3K27me3 is frequently observed in many types of cancers [14]. Enhancer of zeste homolog 2 (EZH2) is the catalytic component of PRC, which selectively trimethylates H3K27. Evidences suggest that overexpression of EZH2 is highly associated with cancer progression and outcome in different cancers. Therefore, EZH2 is regarded as a therapeutic target for cancer treatment [24]. Histone demethylation is catalyzed by histone demethylases, including LSD and JMJC families [25].
Histone phosphorylation is regulated by kinases and phosphatases via adding or removing phosphate groups from the hydroxyl group of serines, threonines, and tyrosines of histone N-terminal tails. Phosphorylation of histone also alters the charge of histone, resulting in the structure alternation of chromatin [26].
2.3. Non-coding microRNAs
MicroRNAs are small, noncoding regulatory RNAs ranging in size from 17 to 25 nucleotides, which are matured by Dicer/Drosha RNase form hairpin-structured precursors [27]. MicroRNAs post-transcriptionally inhibit gene expression by recognizing complementary target sites in the 3’-untranslatied regions of target mRNAs [28]. MicroRNAs play critical roles in cell proliferation and differentiation, cell cycle control, and cell death, as well as tumor development and metastasis [28, 29]. Aberrant microRNAs expression is observed in diverse cancers, and the expression varies in cancer phenotypes and stages. MiRNAs have been demonstrated their importance in de novo methylation of imprinted loci and act on mRNA of DNMTs and HDACs [30]. Each miRNA is capable of regulating the expression of many genes, allowing them to simultaneously regulate multiple cellular signaling pathways [28, 29]. Hence, miRNAs have the potential to be used as biomarkers for cancer diagnosis and prognosis, as well as therapeutic targets [31, 32].
3. Dietary components in cancer prevention and treatment
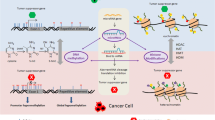
Many bioactive components in diets have been demonstrated to be effective in cancer prevention and intervention through epigenetic alternation [4-8]. In this review, we introduce some common dietary components and their epigenetic targets in cancer. Epigenetic modifications affected by dietary components are presented in Figure 1. Some bioactive components in diet and their epigenetic targets are summarized in Table 1.
3.1. Epigallocatechin-3-gallate (EGCG)
A huge number of studies implicated the anticancer properties of EGCG, a catechin isolated from green tea, with a positive correlation between green tea consumption and the inhibition on cancer [33, 34]. EGCG have a potent inhibitory effect on DNMT activities and DNA methylation in human cancer cells, including esophageal, colon, prostate, and breast cancer cells [35-37]. Treatment of EGCG leads to the demethylation of reactivation of p16, Retinoic acid receptor (RAR)β, MGMT, and human mutL homolog 1 (hMLH1), and glutathione S-transferase p (GSTP) in human oesophageal cancer KYSE 510 cells [35]. The catechol group of catechins plays an important role in which these compounds exert their anticancer functions by inhibiting the methyltransferase activities. Catechol group is an excellent substrate for the methylation mediated by catechol-O-methyltransferase (COMT). COMTmediated methylation leads to the depletion of the methyl donor SAM and promotes the formation of S-adenosyl-L-homocysteine (SAH), inhibiting DNA methylation [38]. Thereby, these catechol- contained compounds inhibit methyltransferase activity by acceleration of COMT-mediated methylation, increasing the levels of SAH. Additionally, the molecular modeling studies indicated that EGCG can directly inhibit the DNMTs through the inhibitory interaction between the gallic acid moiety of EGCG and the catalytic sites of the DNMTs [36]. Furthermore, EGCG promotes the degradation of DNMT3a and HDAC3 [39], as well as the inhibition of HDAC activity [40]. In skin cancer cells, SCC-13 and A431, EGCG reduces the level of polycomb proteins including EZH2, EED, SUZ12, Mel18, and Bmil, which results in the reduction of H3K27me3 and H2AK119ub formation, as well as survival of cancer cells [41]. EGCG also have an effect on miRNAs in human cancer cells. In hepatocellular carcinoma HepG2 cells treated with EGCG, thirteen miRNAs are upregulated and forty-eight miRNAs are down-regulated [42].
Epigenetic modifications regulated by dietary components. The main epigenetic modifications regulated by dietary components include DNA methylation, histone modifications (actylation/deacetylation, methylation/demethylation), and microRNAs. DNA methylation and histone modifications can alter chromatin structure. MicroRNAs can degrade mRNA and modulate translation process.
3.2. Curcumin
Curcumin (diferuloylmethane), a yellow polyphenol from the rhizomes of Curcuma longa, is commonly used as spice and food coloring agent. The major components in the isolated curcuminoid complex are curcumin (approximately 80%), demethoxycurcumin (approximately 17%), and bisdemethoxycurcumin (approximately 3%) [43]. Curcuma longa has been used in traditional medicine in Asia for thousands of years. Recently, curcumin has been identified to exhibit antitumor and apoptosis-induction activity in a variety of human cancer cell lines [44-50]. Moreover, curcumin have also been applied in cancer treatment, including pancreatic cancer, multiple myeloma, and colorectal cancer [51]. Curcumin inhibits DNMT1 activity by covalently blocking the catalytic thiol group of Cys1226 binding site [52]. Prostate LNCaP cells treated with curcumin causes the demethylation of the CpG islands of the NEUROG1 and NRF1 genes. Curcumin also has the effects on the protein expression of HDACs, increasing the expression of HDAC1, 4, 5, and 8 and decreasing HDAC3 [44]. In leukemia K562 and HEL cells, curcumin elevates the expression of SOCS1 and SOCS3 via inhibiting HDAC8 expression to increase the acetylation of histone in the regions of SOCS1 and SOCS3 promoters [45]. Acetylation of the histone protein p300/CBPB and the non-histone protein p53 can be inhibited by curcumin through inhibition on HAT activity [53]. Furthermore, curcumin can enhance the anticancer effect on HDAC inhibitor, trichostatin A, in breast cancer cells SKBR3 and 435eB [46]. In MDA-MB-435 breast cancer cells, curcumin induces the downregulation of EZH2 expression through MAPK pathway [47]. In MCF-7 breast cancer cells and leukemia cells, curcumin upregulates the expression of miR-15 and miR-16, resulting in Bcl-2 downregulation and apoptosis induction [49, 50]. In human pancreatic cells, curcumin increases miR-22 and inhibits miR-199a*. In non-small cell lung cancer cells, the proapoptotic effects of curcumin depend on miR-192-5p/215 induction [48]. MiRNAs and their targeted genes modulated by curcumin also comprise miR-15a, miR-16, miR-21, mir-22, miR-26, miR-34, miR-101, miR-146, miR-200, miR-203, and let-7 [43].
3.3. Resveratrol
Resveratrol (3,5,4’-trihydroxystilbene) is a natural phytoalexin existing in several plants, such as grapes, berries, plums, and peanuts. Its anticancer effects are achieved by inhibiting the growth of cancer cells and inducing apoptosis [54, 55]. Resveratrol is capable of inhibiting the epigenetic silencing of the BRCA1 tumor suppressor protein [56]. Furthermore, resveratrol partially restores H3K9 mono-methylation, DNMTs, and MBD2 at the BRCA1 promoter in MCF-7 cells [56]. Resveratrol shows to have the inhibitory effects on the class III HDACs, such as SIRT1, SIRT2, SIRT3, and p300 [57]. Resveratrol provides an effect in chemoprevention via SIRT1-encoded proteins in an in vivo skin tumor model [58]. In prostate cancer cells, resveratrol enhances p53 acetylation and apoptosis by inhibition of the metastasisassociated protein 1 (MTA)–nucleosome remodeling deacetylation (NuRD) complex [59]. In SW480 colon cancer cells, resveratrol affects the expression of Dicer, PDCD1, and PTEN by downregulation of oncogenic miRNAs, including miR-17, miR-21, miR-25, miR-92a, and miR-196a, while inhibiting the TGFβ by upregulation of miR-663 [60]. In MCF-7 breast cancer cells, resveratrol upregulates miR-663 and miR-773, which inhibits cell proliferation by inhibiting the eukaryotic translation elongation factor 1A2 (eEF1A2) at the mRNA and protein levels [61]. In CLI-5 and A549 lung adenocarcinoma cells, resveratrol downregulates miR-520h and induces miR-520h-mediated signaling pathway, resulting in the inhibition of forkhead box C2 (FOXC2) and the subsequent suppression of tumor metastasis in both in vitro and in vivo model [62]. In resveratrol-induced apoptosis and tumor suppression of neuroblastoma, EZH2 and H3K27me3 repression are mediated by miR-137 [63].
3.4. Flavonoids
Flavonoids (bioflavonoids) are widely in plants, including fruits, vegetables, and beverages (coffee, tea, beer, wine, and juice) [64]. Flavonoids, such as quercetin, fisetin, and myricetin, have been shown to inhibit DNMT activity in different cancer cells [65]. Quercetin (3,5,7,3’,4’-pentahydroxyflavone) is the most abundant flavonoid in nature. In leukemia HL-60 cells, quercetin increases histone H3 acetylation which results in the promotion of the expression of FasL [66]. Quercetin exhibits the potential in the activation of HATs and the inhibition of HDACs, contributing to histone acetylation [66]. Furthermore, quercetin has an inhibitory effect on HAT activity and blocks TNF-α-induced acetylation and phosphorylation of histone H3 at the IP-10 and MIP-2 gene promoter in murine intestinal epithelial cells [67]. Quercetin also exerts an anticancer activity by regulation of miRNAs. The combination of quercetin and hyperoside significantly inhibits the invasion and migration of PC3 prostate cancer cells through downregulation of miR-21 [68]. The combination of quercetin and hyperoside mediates miR-27a inhibition, inducing Sp-repressor ZBTB10 increase which inhibits Sp and survivin, leading to apoptosis of 786-O renal cancer cells [69].
Isoflavones are derived primarily from soybeans, including genistein, daidzein, and glycitein. Genistein inhibits the proliferation, invasion, and metastasis of cancer cells [70]. In addition, genistein has been showed to have a chemopreventive effect against various types of cancer cells, including prostate, esophageal, and colon cancer. Genistein is a phytoestrogen which binds to α and β estrogen receptors and regulates the intracellular signaling pathway to mimic the actions of endogenous estrogen, 17β-estradiol [71]. In esophageal and prostate cancer cells, genistein reverses aberrant DNA methylation which results in reactivation tumor repressor genes, including as p16 INK4A, RARβ, MGMT, phosphatase and tensin homolog (PTEN), cylindromatosis (Turban tumor syndrome, CYLD) by inhibiting DNMT activity [72]. In prostate cancer cells, genistein increases the acetylation of H3K9 at p53 and FOXO3a promoter through inhibition of SIRT1 activity [73]. Genistein also inhibits miR-21 in A-498 renal cancer cells growth and tumor growth in xenograft model through cell cycle arrest and apoptosis induction [74]. In pancreatic cancer cells, genistein inhibits cell growth and induces apoptosis through the up-regulation of miR-34a and Notch-1 signaling pathway [75].
3.5. Isothiocyanates
Isothiocyanates (ITCs), metabolites of glucosinolates, are found in cruciferous vegetables, such as broccoli, cabbage, brussels sprouts, watercress, kale, and cauliflower, which provide a remarkable anticancer effect on pancreatic, prostate, ovarian, and breast cancers [76, 77]. The known potent anticancer effects of ITCs are from allyl isothiocyanate (AITC), benzyl isothiocyanate (BITC), phenethyl isothiocyanate (PITC), and sulforaphane. The anticancer activities of ITCs can be divided into chemopreventive and chemotherapeutic effects. ITCs have a potential in inhibition of carcinogenesis via acting on detoxification, inflammation, apoptosis, and cell cycle, as well as epigenetic regulation [77]. ITCs are found to inhibit tumorigenesis through inhibition of HDACs [78, 79]. The inhibition of HDAC activity also contributes to the increase of tumor suppressor gene p21WAF1 and the pro-apoptotic gene Bax [80]. In myeloma cells, PITC can inhibit HDACs and induce DNA demethylation of tumor suppressor p16 gene, which leads to its reactivation [81]. In HL-60 leukemia cells, PITC mediates cell growth arrest which is associated with the reduction of HDAC activity and increase of acetyltransferase p300 and p21WAF1 acetylation [82]. In hyperplastic (BPH1) and prostate cancer cells (LNCaP and PC3), sulforaphane induces cell cycle arrest and apoptosis by inhibition of HDAC activity and class I and II HDAC proteins, as well as increase of H3 acetylation followed by induction of p21 WAF1 expression [83]. The inhibition of HDAC is related to sulforaphane metabolite sulforaphanecysteine that can fit the enzyme pock and form a bidentate ligand through the interaction of α-carboxyl group of the cysteine moiety and buries zinc atom [79]. In vivo data showed that administration of sulforaphane to Apc min mice can inhibit intestinal tumors and increase histone acetylation of global DNA and the promoter regions of p21 WAF1 and Bax [84]. PITC has been proved to exert a lung chemopreventive effect by modulated the cigarette smokeinduced alternations on miRNA expression. Furthermore, the combination of PITC and indole 3-carbinol reverses the effects of cigarettes on these miRNAs [85, 86]. In prostate cancer cells, PITC inhibits androgen receptor by enhancing miR-17 mediated suppression of p300/CBP-associated factor (PCAF), a co-regulator of androgen receptor [87]. Sulforaphane treatment causes the reduction of EZH2 and Bmi-1 expression in SCC-13 skin cancer cells and also reduces H3K27me3, which is associated with the accumulation of cells at G2/M phase, the decrease of cyclin B1, cyclin A, cyclin dependent kinases (CDK) 1/2, and the increase of p21WAF1 expression [88]. Additionally, PITC modulates miR-27a, miR-20a, and miR-17-5p, resulting in the apoptosis of pancreatic cancer cells [89].
4. Bioavailability of dietary components
The anticancer effects of dietary components like EGCG, curcumin, and resveratrol have been reported in a wide variety of cancers [33, 34, 90, 91]. Clinical trial results showed that the prostate cancer risk declined with increasing frequency, duration, and quantity of green tea consumption, and the dose-response relationships were also significant, suggesting that EGCG is effective against prostate cancer [33, 34]. Curcumin and resveratrol, either alone or in combination with other agents, have demonstrated their potential effects against colorectal cancer, pancreatic cancer, breast cancer, prostate cancer, multiple myeloma, lung cancer, oral cancer, and head and neck squamous cell carcinoma [90-91].
Along with laboratory-based results, some clinical trials have demonstrated the chemopreventive and chemotheputic effects of dietary components. However, most of the clinical trials of these components such as EGCG, curcumin, resveratrol, and genisten as anticancer agents are poor bioavailability, raising the debate of the practical application of in vitro results in physiological states [51, 92, 93]. Clinical studies of these promising molecules demonstrated their low bioavailability, which are hindered by poor water solubility and absorption, as well as rapid metabolism and clearance [51, 92, 93]. The formation of sulphate and glucuronide conjugated by intestine and liver also reduced the bioavailability of some phenolic compounds [51, 93]. Thus, continuing research on these dietary components is needed to provide some possible solutions to overcome these problems. To increase the bioavailability, longer circulation, higher permeability, and resistance to metabolic processes of dietary components, a variety of approaches have been developed, including synthesis of analogues, nanoparticles, liposomes, micelles, and phospholipid complexes [94-96]. The combination use of adjuvants also serves as a useful strategy to improve the bioavailability of dietary components. Recent studies showed that the administration of curcumin or resveratrol with piperine significantly improve their bioavailability through the inhibition of glucuronidation, elevating their concentrations in plasma [97, 98].
5. Conclusion and future perspectives
Based on the studies mentioned, it is clear that these dietary components act on different epigenetic targets leading to epigenetic modifications and execute anticancer activities. Current studies also showed that dietary components used in combination with traditional chemotherapy creates potential synergistic effects, including reverting chemotherapy resistance, overcoming the side effects from chemotherapy, and increasing chemotherapy sensitivity, which are all very important for the successful treatment of cancer patients. Insight understanding of the global patterns of epigenetic modifications by dietary components in cancer will necessarily help to develop better strategies to prevent and cure cancer. Therefore, sufficient preclinical data is required for the better understanding of the epigenetic targets and pathways altered by these dietary components to increase the efficacy of their anticancer properties. Additional clinical studies are also needed to analyze the safety profile of dosages, the routes of administration, tissue distribution, as well as bioavailability alone, and in combination with other chemotherapeutic agents in order to obtain the maximum beneficial effects of these dietary components as anticancer agents. Despite these challenges, persistent research on dietary components will offer more epigenetic targets and promising strategies for theprevention and treatment against cancers in the future.
Abbreviations
- AITC:
-
allyl isothiocyanate
- BITC:
-
benzyl isothiocyanate
- COMT:
-
catechol-O-methyltransferase
- CpG:
-
cytosinephosphate-guanine
- DNMTs:
-
DNA methyltransferases
- EGCG:
-
epigallocatechin-3-gallate
- EZH2:
-
enhancer of zeste homolog 2
- H3K4me1:
-
Monomethylation of histone H3 at lysine 4
- H3K4me3:
-
trimethylation of histone H3 at lysine 4
- H3K9me3:
-
trimethylation of histone H3 at lysine 9
- H3K20me3:
-
trimethylation of histone H3 at lysine 20
- H3K27me3:
-
trimethylation of histone H3 at lysine 27
- HATs:
-
histone acetyl transferases
- HDACs:
-
histone deacetylases
- ITCs:
-
isothiocyanate
- miRNAs:
-
microRNAs
- PITC:
-
phenethyl isothiocyanate
- SAH:
-
S-adenosyl-L- homocysteine
- SAM:
-
S-adenosyl-methionine
References
Jaenisch R, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Science 2003; 300: 489–92.
Dehan P, Kustermans G, Guenin S, Horion J, Boniver J, Delvenne P. DNA methylation and cancer diagnosis: new methods and applications. Expert Rev Mol Diagn 2009; 9: 651–7.
Hirst M, Marra MA. Epigenetics and human disease. Int J Biochem Cell Biol 2009; 41: 136–46.
Stefanska B, Karlic H, Varga F, Fabianowska-Majewska K, Haslberger A. Epigenetic mechanisms in anti-cancer actions of bioactive food components – the implications in cancer prevention. Brit J Pharmacol 2012; 167: 279–97.
Gonzalez-Vallinas M, Gonzalez-Castejon M, Rodriguez-Casado A, Ramirez de Molina, A. Dietary phytochemicals in cancer prevention and therapy: a complementary approach with promising perspectives. Nutr Rev 2013; 71: 585–99.
Shankar S, Kumar D, Srivastava RK. Epigenetic modifications by dietary phytochemicals: implications for personalized nutrition. Pharmcol Ther 2013; 138: 1–17.
Thakur VS, Deb G, Babcook MA, Gupta S. Plant phytochemicals as epigenetic modulators: role in cancer chemoprevention. AAPS J 2014; 16: 151–63.
Bird AP. CpG-rich islands and the function of DNA methylation. Nature 1986; 321: 209–13.
Issa J, Kantarjian HM. Target DNA methylation. Clin Cancer Res 2009; 15: 3933–46.
Subramaniam D, Thombre R, Dhar A, Anant S. DNA methyltransferases: a novel target for prevention and therapy. Front Oncol 2014; 4: 80.
Ooi SK, Qiu C, Bernstein E, Li K, Jia D, Yang Z, et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 2007; 448: 714–7.
Weber M, Davies JJ, Wittig D, Oakeley EJ, Haase M, Lam WL, et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat Genet 2205; 37: 853–62.
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997; 389: 251–60.
Mills AA. Throwing the cancer switch: reciprocal roles of polycomb and trithorax proteins. Nat Rev Cancer 2012; 10: 669–82.
Suganuma T, Workman JL. Signals and combinatorial functions of histone modifications. Annu Rev Biochem 2011; 80: 473–99.
Shilatifard A. Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu Rev Biochem 2006; 75: 243–69.
Bernstein BE, Meissner A, Lander ES. The mammalian epigenome. Cell 2007, 128: 669–81.
Kouzarides T. Chromatin modifications and their function. Cell 2007; 128: 693–705.
Shahbazian MD, Grunstein M. Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem 2007; 76: 75–100.
Gong F, Miller KM. Mammalian DNA repair: HATs and HDACs make their mark through histone acetylation. Mut Res 2013; 750: 23–30.
Dalvai M, Bystricky K. The role of histone modifications and variants in regulating gene expression in breast cancer. J Mammary Gland Biol Neoplasia 2010; 15: 19–33.
Yamane K, Tateishi K, Klose RJ, Fang J, Fabrizio LA, Erdjument- Bromage H, et al. PLU-1 is an H3K4 demethylase involved in transcriptional repression and breast cancer cell proliferation. Mol Cell 2007; 25: 801–12.
Banister AJ, Kouzarides T. Reversing histone methylation. Nature 2005; 436: 1103–6.
Wee S, Dhanak D, Li H, Armstrong SA, Copeland RA, Sims R, et al. Targeting epigenetic regulators for cancer therapy. Ann NY Acad Sci 2014; 1309: 30–6.
Koooistra SM, Helin K. Molecular mechanisms and potential functions of histone demethylases. Nat Rev Mol Cell Biol 2012; 13: 297–11.
Berger SL. Cell signaling and transcriptional regulation via histone phosphorylation. Cold Spring Harb Symp Quant Bio 2010; 75: 23–26.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 2004; 5: 522–31.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120: 15–7.
Garzon R, Calin GA, Croce CM. MicroRNAs in Cancer. Annu Rev Med 2009; 60: 167–79.
Zhou H, Hu H, Lai M. Non-coding RNAs and their epigenetic regulatory mechanisms. Biol Cell 2010; 102: 645–55.
McDermot AM, Heneghan HM, Miller N, Kerin MJ. The therapeutic potential of microRNAs: disease modulators and drug targets. Pharm Res 2011; 28: 3016–29.
Piva R, Spandidos D, Gambrai R. From microRNA functions to microRNA therapeutics: Novel targets and novel drugs in breast cancer research and treatment. Int J Oncol 2013; 43: 985–94.
Jian L, Xie LP, Lee AH, Binns CW. Protective effect of green tea against prostate cancer: a case-control study in southeast China. Int J Cancer 2004; 108: 130–5.
Khan N, Adhami VM, Mukhtar H. Green tea polyphenols in chemoprevention of prostate cancer: preclinical and clinincal studies. Nutr Cancer 2009; 61: 836–41.
Fang MZ, Wang Y, Ai N, Hou Z, Sun Y, Lu H, et al. Tea polyphenol (-)-epigallocatechin-3-gallate inhibits DNA methyltransferase and reactivates methylation-silenced genes in cancer cell lines. Cancer Res 2003; 63: 7563–70.
Lee WJ, Shim JY, Zhu BT. Mechanism for the inhibition of DNA methytransferases by the catechins and bioflavonoids. Mol Pharmacol 2005; 68: 1018–30.
Lee WJ, Zhu BT. Inhibition of DNA methylation by caffeic acid and chlorogenic acid, two common catechol-containing coffee polyphenols. Carcinogenesis 2006; 27: 269–77.
Singh B, Shankar S, Srivastava. Green tea catechin, epigallocatechin- 3-gallate (EGCG): mechanisms, perspectives and clinical applications. Biochem Pharmacol 2011; 82: 1807–21.
Moseley VR, Morris J, Knackstedt RW, Wargovich MJ. Green tea polyphenol epigallocatechin 3-gallate, contributes to the degradation of DNMT3A and HDAC3 in HCT 116 human colon cancer cells. Anticancer Res 2013; 33: 5325–33.
Kim SO, Kim MR. (-)-Epigallocatechin 3-gallate inhibits invasion by inducing the expression of Raf kinase inhibitor protein in AsPC-1 human pancreatic adenocarcinoma cells through the modulation of histone deacetylase activity. Int J Oncol 2013; 42: 349–58.
Choudhury SR, Balasubramanian S, Chew YC, Han B, Marquez VE, Eckert RL. (-)-Epigallocatechin-3-gallate and DZNep reduce polycomb protein level via a proteasome-dependent mechanism in skin cancer cells. Carcinogenesis 2011; 32: 1525–32.
Tsang WP, Kwok TT. Epigallocatechin gallate up-regulation of miR-16 and induction of apoptosis in human cancer cells. J Nutr Biochem 2010; 21: 140–6.
Teiten MH, Dicato M, Diederich M. Curcumin as a regulator of epigenetic events. Mol Nurt Food Res 2013; 57: 1619–29.
Shu L, Khor TO, Lee JH, Boyanapalli SS, Huang Y, Wu TY, et al. Epigenetic CpG demethylation of the promoter and reactivation of the expression of Neurog1 by curcumin in prostate LNCaP cells. 2011; AAPS J. 13: 606–14.
Chen CQ, Yu K, Yan QX, Xing CY, Chen Y, Yan Z, et al. Pure curcumin increases the expression of SOCS1 and SOCS3 in myeloproliferative neoplasms through suppressing class I histone deacetylases. Carcinogenesis 2013; 34: 1442–9.
Yan G, Graham K, Lanza-Jacoby S. Curcumin enhances the anticancer effects of trichostatin A in breast cancer cells. Mol Carcinog 2013; 52: 404–11.
Hua WF, Fu YS, Liao YJ, Xia WJ, Chen YC, Zeng YX, et al. Curcumin induces down-regulation of EZH2 expression through the MAPK pathway in MDA-MB-435 human breast cancer cells. Eur J Pharmacol 2012; 637: 16–21.
Ye M, Zhang J, Zhang J, Miao Q, Yao L, Zhang J. Curcumin promotes apoptosis by activating the p53-miR-192-5p/215-XIAP pathway in non-small cell lung cancer. Cancer Lett 2015; 357: 196–205.
Gao SM, Yang JJ, Chen CQ, Chen JJ, Ye LP, Wang LY, et al. Pure curcumin decreases the expression of WT1 by upregulation of miR- 15a and miR-16-1 in leukemic cells. J Exp Clin Cancer Res. 2012; 31: 27.
Yang J, Cao Y, Sun J, Zhang Y. Curcumin reduces the expression of Bcl-2 by upregulating miR-15a and miR-16 in MCF-7 cell. Med Oncol 2010; 27: 1114–8.
Anand P, Kunnumakkara AB, Newman RA, Aggarwal BB. Bioavailability of curcumin: problems and promises. Mol Pharm 2007; 4: 807–18.
Liu Z, Xie Z, Jones W, Pavlovicz RE, Liu S, Yu J, et al. Curcumin is a potent DNA hypomethylation agent. Bioorg Med Chem Lett 2009; 19: 706–9.
Balasubramanyam K, Varier RA, Altaf M, Swaminathan V, Siddappa NB, Ranga U, et al. Curcumin, a novel p300/CREB-binding proteinspecific inhibitor of acetyltransferase, represses the acetylation of histone/nonhistone proteins and histone acetyltransferase-dependent chromatin transcription. J Biol Chem 2004; 279: 51163–71.
Whyte L, Huang YY, Torres K, Mehta RG. Molecular mechanisms of resveratrol action in lung cancer cells using dual protein and microarray analyses. Cancer Res 2007; 67: 12007–17.
Kundu JK, Surh YJ. Cancer chemopreventive and therapeutic potential of resveratrol: mechanistic perspectives. Cancer Lett 2008; 269: 243–61.
Papoutsis AJ, Lamore SD, Wondrak GT, Selmin OI, Romagnolo DF. Resveratrol prevents epigenetic silencing of BRCA-1 by the aromatic hydrocarbon receptor in human breast cancer cells. J Nutr 2010; 140: 1607–14.
Roy SK, Chen Q, Fu J, Shankar S, Srivastava RK. Resveratrol inhibits growth of orthotopic pancreatic tumors through activation of FOXO transcription factors. PLoS ONE2011; 6: e25166.
Boily G, He XH, Pearce B, Jardine K, Mcburney MW. SirT1-null mice develop tumors at normal rates but are poorly protected by resveratrol. Oncogene 2009; 28: 2882–93.
Kai L, Samuel SK, Levenson AS. Resveratrol enhances p53 acetylation and apoptosis in prostate cancer by inhibiting MTA1/NuRD complex. Int J Cancer 2010; 126: 1538–48.
Tili E, Michaille JJ, Alder H, Volinia S, Delmas D, Latruffe N, et al. Resveratrol modulates the levels of microRNAs targeting genes encoding tumorsuppressors and effectors of TGFbeta signaling pathway in SW480 cells. Biochem Pharmacol 2010; 80: 2057–65.
Vislovukh A, Kratassiouk G, Porto E, Gralievska N, Beldiman C, Pinna G, et al. Proto-oncogenic isoform A2 of eukaryotic translation elongation factor eEF1 is a target of miR-663 and miR-744. Brit J Cancer 2013; 108: 2304–11.
Yu YH, Chen HA, Chen PS, Cheng YJ, Hsu WH, Chang YW, et al. MiR-520h-mediated FOXC2 regulation is critical for inhibition of lung cancer progression by resveratrol. Oncogene 2013; 32: 431–43.
Ren X, Bai X, Zhang X, Li Z, Tang L, Zhao X, et al. Quantitative nuclear proteomics identifies that miR-137-mediated EZH2 reduction regulates resveratrol-induced apoptosis of neuroblastoma cells. Mol Cell Proteomics 2014; 14: 316–28.
Bravo L. Polyphenols: chemistry, dietary sources, metabolism, and nutritional significance. Nutr Rev 1998; 56: 317–33.
Ong TP, Moreno FS, Ross SA. Targeting the epigenome with bioactive food components for cancer prevention. J Nutrigenet Nutrigenomics 2012; 4: 275–92.
Lee WJ, Chen YR, Tseng TH. Quercetin induces FasL-related apoptosis, in part, through promotion of histone H3 acetylation in human leukemia HL-60 cells. Onco Rep 2011; 2: 583–91.
Ruiz PA, Braune A, Holzlwimmer G, Quintanilla-Fend L, Haller D. Quercetin inhibits TNF-induced NF-kappaB transcription factor recruitment to proinflammatory gene promoters in murine intestinal epithelial cells. J Nutr 2007; 137: 1208–15.
Yang FQ, Liu M, Li W, Che JP, Wang GC, Zheng JH. Combination of quercetin and hyperoside inhibits prostate cancer cell growth and metastasis via regulation of microRNA-21. Mol Med Rep 2015; 11: 1085–92.
Li W, Liu M, Xu YF, Feng Y, Che JP, Wang GC, et al. Combination of quercetin and hyperoside has anticancer effects on renal cancer cells through inhibition of oncogenic microRNA-27a. Oncol Rep 2014; 31: 117–24.
Banerjee S, Li Y, Wang Z, Sarkar FH. Multi-targeted therapy of cancer by genistein. Cancer Letters 2008; 269: 226–42.
Yoon K, Kwack SJ, Kim HS, Lee BM. Estrogenic endocrinedisrupting chemicals: molecular mechanism of actions on putative human diseases. J Toxicol Environ Health B Crit Rev 2014; 17: 127–74.
Fang MZ, Chen D, Sun Y, Jin Z, Christman JK, Yang CS. Reversal of hypermethylation and reactivation of p16INK4a, RAR beta, and MGMT genes by genistein and other isoflavone form soy. Clin Cancer Res. 2005; 11: 7033–41.
Kikuno N, Shiina H, Urakami S, Kawamoto K, Hirata H, Tanaka Y, et al. Genistein mediated histone acetylation and demethylation activates tumor suppressor genes in prostate cancer cells. Int J Cancer 2008; 123: 552–60.
Zaman MS, Shahryari V, Deng G, Thamminana S, Saini S, Majid S, et al. Up-regulation of microRNA-21 correlates with lower kidney cancer survival. PLoS One 2012; 7: 2.
Xia J, Duan Q, Ahmad A, Bao B, Banerjee S, Shi Y, et al. Genistein inhibits cell growth and induces apoptosis through Up-regulation of miR-34a in pancreatic cancer cells. Curr. Drug Targets 2012; 13: 1750–6.
Razis AFA, Noor NM. Cruciferous vegetables: dietary phytochemicals for cancer prevention. Asian Pacific J Cancer Prev 2013; 1565- 7–0.
Gupta P, Wright SE, Kim SH, Srivastava SK. Phenethyl isothiocyanate: A comprehensive review of anti-cancer mechanisms. Biochim Biophys Acta 2014; 1846: 405–24.
Ho E, Clarke JD, Dashwood RH. Dietary sulforaphane, a histone deacetylase inhibitor for cancer prevention. J Nutr 2009; 139: 2393–69.
Nian H, Delage B, Ho E, Dashwood RH. Modulation of histone deacetylase activity by dietary isothiocyanates and allyl sulfides: studies with sulforaphane and garlic organosulfur compounds. Environ Mol Mutagen 2009; 50: 213–21.
Wang LG, Liu XM, Fang Y, Dai W, Chiao FB, Puccio GM, et al. De-repression of the p21 promoter in prostate cancer cells by an isothiocyanate via inhibition of HDACs and c-Myc. Int J Oncol 2008; 33: 375–80.
Lu Q, Lin X, Feng J, Zhao X, Gallagher R, Lee MY, et al. Phenylhexyl isothiocyanate has dual function as histone deacetylase inhibitor and hypomethylating agent and can inhibit myeloma cell growth by targeting critical pathways. J Hematol Oncol 2008; 1: 6.
Ma X, Fang Y, Beklemisheva A, Dai W, Feng J, Ahmed T, et al. Phenylhexyl isothiocyanate inhibits histone deacetylases and remodels chromatins to induce growth arrest in human leukemia cells. Int J Oncol 2006; 28: 1287–93.
Clarke JD, Hsu A, Yu Z, Dashwood RH, Ho E. Differential effects of sulforaphane on histone deacetylases, cell cycle arrest and apoptosis in normal prostate cells versus hyperplastic and cancerous prostate cells. Mol. Nutr Food Res 2011; 55: 999–1009.
Myzak MC, Dashwood WM, Orner GA, Ho E, Dashwood RH. Sulforaphane inhibits histone deacetylase in vivo and suppresses tumorigenesis in Apc-minus mice. FASEB J 206; 20: 506–8.
Izzotti A, Calin GA, Steele VE, Cartiglia C, Longobardi M, Croce CM, et al. Chemoprevention of cigarette smoke-induced alterations of microRNA expression in rat lungs. Cancer Prev Res 2010; 3: 62–72.
Izzotti A, Larghero P, Cartiglia C, Longobardi M, Pfeffer U, Steele VE, et al. Modulation of microRNA expression by budesonide, phenethyl isothiocyanate and cigarette smoke in mouse liver and lung. Carcinogenesis 2012; 31: 894–901.
Yu C, Gong AY, Chen D, Solelo Leon D, Young CY, Chen XM. Phenethyl isothiocyanate inhibits androgen receptor-regulated transcriptional activity in prostate cancer cells through suppressing PCAF. Mol Nutr Food Res 57: 2013; 1825–33.
Balasubramanian S, Chew YC, Eckert RL. Sulforaphane suppresses polycomb group protein level via a proteasome-dependent mechanism in skin cancer cells. Mol Pharmacol 2011; 80: 870–8.
Jutooru I, Guthrie AS, Chadalapaka G, Pathi S, Kim K, Burghardt R, et al. Mechanism of action of phenethylisothiocyanate and other reactive oxygen species-inducing anticancer agents. Mol Cell Biol 2014; 34: 2382–95.
Gupta SC, Patchva S, Aggarwal BB. Therapeutic roles of curcumin: lessons learned from clinical trails. AAPS J 2013; 15: 195–218.
Bishayee A. Cancer prevention and treatment resveratrol: from rodent studies to clinical trials. Cancer Prev Res 2009; 2: 409–18.
D’Archivio M, Filesi C, Di Benedetto R, Gargiulo R, Giovannini C, Masella R. Polyphenols: dietary sources and bioavailability. Ann Ist Super Sanita 2007; 43: 348–61.
Walle T, Hsieh F, DeLegge MH, Oatis JE, Jr., Walle UK. High absorption but very low bioavailability of oral resveratrol in humans. Drug Metab Dispos 2004; 32: 1377–82.
Gulseren I, Gorredig M. Storage stability and physical characteristics of tea-polyphenol-bearing nanoliposomes prepared with milk fat globule membrane sphspholipids. J Agric Food Chem 2013; 61: 3242–51.
Mignet N, Seguin J, Chabot GG. Bioabailability of polyphenol liposomes: a challenge ahead. Pharmaceutics 2013; 5: 457–71.
He L, Deng D, Zhou X, Cheng L, Ten Cate JM, Li J, et al. Novel tea polyphenol-modified calcium phosphate nanoparticle and its remineralization potential. J Biomed Mater Res B Appl Biomater 2014 Dec 2. Doi: 10.1002/jbm.b.33333.
Shoba G, Joy D, Joseph T, Majeed M, Rajendran R, Srinivas PS. Influence of piperine on the pharmacokinetics of curcumin in animals and human volunteers. Planta Med 1998; 64: 353–56.
Johnson JJ, Nihal M, Siddiqui IA, Scarlett CO, Bailey HH, Mukhtar H, et al. Enhancing the bioavailability of resveratrol by combining it with piperine. Mol Nutr Food Res 2011; 55: 1169–76.
Acknowledgments
This study was supported by funding from Ministry of Science and Technology, Taiwan (MOST103-2320-B-039-052-MY3; MOST104-2321-B-039-005; MOST104-2314-B-039-034); Ministry of Health and Welfare (MOHW104-TDU-B-212-124-002); National Health Research Institutes, Taiwan (NHRI-EX102- 10245BI) and China Medical University Hospital (DMR-104-097 and DMR-104-108).
Author information
Authors and Affiliations
Corresponding author
Additional information
aChinese Medicinal Research and Development Center, China Medical University Hospital, Taichung 404, Taiwan
bGraduate Institute of Cancer Biology, China Medical University, Taichung 404, Taiwan
cCenter for Molecular Medicine, China Medical University Hospital, Taichung 404, Taiwan
dDepartment of Biotechnology, Asia University, Taichung 413, Taiwan
*Corresponding author. Graduate Institute of Cancer Biology, China Medical University, Taichung 404, Taiwan.
E-mail address: ylyu@mail.cmu.edu.tw (Y.-L. Yu).
Statement of conflicts of interest
No potential conflicts of interest were disclosed.
Open Access This article is distributed under terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided original author(s) and source are credited.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chang, LC., Yu, YL. Dietary components as epigenetic-regulating agents against cancer. BioMed 6, 2 (2016). https://doi.org/10.7603/s40681-016-0002-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.7603/s40681-016-0002-8