Abstract
Tiller number is a key component of wheat plant architecture having a direct impact on grain yield. Because of their viability, biotic resistance, and abiotic stress tolerance, wild relative species are a valuable gene source for increasing wheat genetic diversity, including yield potential. Agropyron glael, a perennial hybrid of Thinopyrum intermedium and Th. ponticum, was created in the 1930s. Recent genome analyses identified five evolutionarily distinct subgenomes (J, Jst, Jvs, Jr, and St), making A. glael an important gene source for transferring useful agronomical traits into wheat. During a bread wheat × A. glael crossing program, a genetically stable translocation line, WT153397, was developed. Sequential in situ hybridizations (McGISH) with J-, St-, and D-genomic DNA probes and pSc119.2, Afa family, pTa71, and (GAA)7 DNA repeats, as well as molecular markers specific for the wheat 6D chromosome, revealed the presence of a 6DS.6Jvs Robertsonian translocation in the genetic line. Field trials in low-input and high-input breeding nurseries over four growing seasons demonstrated the Agropyron chromosome arm’s high compensating ability for the missing 6DL, as spike morphology and fertility of WT153397 did not differ significantly from those of wheat parents, Mv9kr1 and ‘Mv Karizma.’ Moreover, the introgressed 6Jvs chromosome arm significantly increased the number of productive tillers, resulting in a significantly higher grain yield potential compared to the parental wheat cultivars. The translocated chromosome could be highly purified by flow cytometric sorting due to the intense fluorescent labeling of (GAA)7 clusters on the Thinopyrum chromosome arm, providing an opportunity to use chromosome genomics to identify Agropyron gene variant(s) responsible for the tillering capacity. The translocation line WT153397 is an important genetic stock for functional genetic studies of tiller formation and useful breeding material for increasing wheat yield potential. The study also discusses the use of the translocation line in wheat breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hexaploid wheat (Triticum aestivum L., 2n = 6x = 42, BBAADD) is one of the most important grain crops, grown on about 220 million hectares worldwide (FAOSTAT 2020). To feed the world’s growing population, the current wheat yield production of 763.7 million tons (International Grain Council, 2020/2021) should be increased by 60% until 2050 (https://iwyp.org/global-challenge). However, thousands of years of cultivation and domestication of wheat have led to a reduction in genetic diversity (Tiwari et al. 2014), which has hampered the development of high-yielding cultivars and yield stability under biotic and abiotic stresses (Sears 1981; Jiang et al. 1993; Friebe et al. 1996; Oliver et al. 2006).
Improving wheat structural and reproductive traits contributes to increased crop yield potential (YP) (Reynolds et al. 2011), which is the maximum yield per unit land area achieved under optimal environmental conditions. Among the morphological traits, tiller formation correlates linearly with the number of spikes and has a strong effect on grain yield (Darwinkel 1980). In wheat, two types of tillers form the following: productive tillers, which lead to the formation of spikes and thus significantly contribute to the total number of grains per plant and grain yield, and non-productive tillers, which reduce grain yield (Kirby and Jones 1977; Kulshrestha and Chowdhury 1987).
Species related to wheat are a valuable source of genetic variation for expanding the genetic basis of bread wheat (Tester and Langridge 2010; Mujeeb-Kazi et al. 2013) and improving its morphological and reproductive traits (Reynolds et al. 2011). The introgression of the 1RS chromosome arm from rye (Secale cereale, L.) into wheat in the form of 1BL.1RS wheat-rye Robertsonian translocation has been widely used in wheat breeding programs (Rabinovich 1998). This translocation is present in many wheat cultivars worldwide and has improved the yield potential and biotic stress resistance of cultivated wheat (Zeller 1973; Carver and Rayburn 1994; Moreno-Sevilla et al. 1995).
Wheat wild relatives have also long been used in wheat breeding, primarily to improve biotic resistance and abiotic stress tolerance. Pest and disease-resistance genes are involved in over 80% of the beneficial traits introgressed into cultivated species from wild relatives (Hajjar and Hodgkin 2007). Beginning in the late 1970s, CIMMYT produced and characterized interspecific and intergeneric crosses to study genetic relationships and to introgress genetic variation from wild species into wheat (Mujeeb-Kazi and Hettel 1995). Because their homoeologous relationship with wheat is closer than that of other perennial species (King et al. 1993; Mujeeb-Kazi and Hettel 1995; Ceoloni et al. 2014), Thinopyrum species were involved in one of the earliest alien gene transfers from wild species into wheat. Wheatgrasses have been used successfully to improve wheat biotic resistance and abiotic stress tolerance. However, only a few studies have been published on the beneficial effect of wild alien chromatin on wheat yield potential (Monneveux et al. 2003; Kuzmanović et al. 2014; Gao et al. 2021). The 7DL.7Ag translocation from Lophopyrum elongatum (syn: Thinopyrum ponticum), which carries the leaf rust resistance gene Lr19, was found to significantly increase grain yield (Monneveux et al. 2003). New knowledge in the genetics of desired traits, increased availability of wild relatives in gene banks, improved interspecific hybridization techniques, and advanced molecular technologies may lead to an expansion of the scale of agronomic traits introgressed from related wild species into cultivated wheat.
The genus Thinopyrum, which belongs to the wheat tertiary gene pool, has a variable genome structure ranging from diploid to auto- and allodecaploid species (Ceoloni et al. 2014). For example, the exact genomic constitution of the perennial Th. intermedium (Host) Barkworth & D.R. Dewey, a segmental autoallohexaploid (2n = 6x = 42) species, has shown ambiguous features. Liu and Wang (1993) proposed the genome constitution JeJeJeJeSS, with the subgenome Je originating from Th. elongatum (Host) D. Dewey (2n = 2x = 14, EE) and the subgenome S originating from Pseudoroegneria spicata (Pursh) A. Löve (2n = 2x = 14, SS or StSt). Later, it was suggested to rename the S genome with St, since the genomes of the Aegilops species in the section Sitopsis were also assigned the S letter. Using genomic in situ hybridization (GISH), Chen et al. (1998) found that signals of the St-genomic probe were located at the centromeric region on 11 out of the 28 J-genome chromosomes, prompting them to rename the genome symbol to JJJsJsStSt. According to the findings of Deng et al. (2013), Kruppa and Molnár-Láng (2016), and Cseh et al. (2019), who found only 8 to 10 centromeric St GISH signals, the number of Th. intermedium chromosomes with St centromeres varies by genotype. Kishii et al. (2005) suggested that Dasypyrum villosum L. (2n = 2x = 14, VV) was also involved in the evolution of Th. intermedium because the V-genomic probe showed strong signals along the length but lacked signals around centromeric regions in several chromosomes and telomeric regions of all chromosomes. Wang et al. (2015) proposed the most recent change to the genomic designation as JvsJrSt, where Jvs and Jr represent the ancestral genomes of present-day Jb of Th. bessarabicum (Savul. & Rayss) Á.Löve (2n = 2x = 14) and Je of Th. elongatum, respectively, with Jvs being more ancient.
Th. intermedium and Th. ponticum (Podp.) Z.-W. Liu & R.-C. Wang (2n = 10x = 70; JJJJJJJsJs) are valuable gene sources for wheat since they have useful agronomic and quality traits such as high grain protein content, bread-making quality, resistance to rust diseases, and tolerance to low temperature, drought, moisture, and salinity (Dubcovsky et al. 1994; Tang et al. 2000; Colmer et al. 2006; Garg et al. 2014; Tong et al. 2021).
In order to pyramid genes and QTLs encoding useful agronomic traits of Th. intermedium and Th. ponticum in a bridge material that can be used as a gene source for introgression breeding of wheat, N. V. Tsitsin developed an artificial hybrid, Agropyron glael (2n = 8x = 56) in the 1930s by crossing Th. intermedium (syn. A. glaucum (Desf. ex DC.) Roem. & Schult) and Th. ponticum (syn. A. elongatum (Host) P. Beauv.). A detailed molecular cytogenetic analysis of A. glael revealed that it carried 18 J, 28 JSt, and 8 St chromosomes with J-St and JStSt translocations and chromosomes with decreased fluorescence intensity, resulting in unique hybridization patterns (Kruppa and Molnár-Láng 2016). The A. glael clone No. 8 obtained from the Moscow Main Botanical Garden was used in the wheat introgression breeding program at the Centre for Agricultural Research (Kruppa et al. 2016). In 2001, the winter wheat line ‘Martonvásári 9 kr1’ (Mv9kr1) carrying recessive crossability alleles (kr1kr1kr2kr2) (Molnár-Láng et al. 1996) was crossed with A. glael. Backcrossing and selfing the hybrid plants resulted in the selection of several partial amphiploid lines (Kruppa et al. 2016), which were then backcrossed with wheat to select addition, substitution, or translocation lines.
The present study reports on the development of a wheat/A. glael disomic translocation (WT153397) with positive effects on reproductive tiller formation and grain yield when compared to the wheat parents. Further aims were to (1) identify the chromosome composition of the WT153397 introgression line using molecular cytogenetic, flow cytometric, and molecular marker analyses, and (2) evaluate the effects of the added alien chromosome segment on wheat agronomic performance in field trials conducted over four growing seasons. The study also reports on the inclusion of the translocation line in wheat breeding.
Materials and methods
Plant materials
The Mv9kr1 × A. glael BC1F1 partial amphiploid genotype (Kruppa et al. 2016; Kruppa and Molnár-Láng 2016) was backcrossed with the facultative elite wheat cultivar Mv Karizma (Supplementary Figure S1). The wheat/A.glael translocation line WT153397 was selected in the BC3F2 generation using molecular cytogenetic techniques. The WT153397 line, along with the hexaploid wheat crossing partners Mv9kr1, ‘Chinese Spring’ (CS), and ‘Mv Karizma,’ was used for field phenotyping experiments in Martonvásár low-input and high-input nurseries over a 4-year period (2019, 2021, 2022 and 2023). Beginning in 2018, the WT153397 line was involved in a wheat breeding program in Martonvásár. The maternal WT153397 line was crossed with the Martonvásár elite cultivar Mv Nádor, and the offspring were selected based on the plants’ tillering ability and spike length. The phenotype of the F5 generation was assessed in the high-input nursery in 2023 and compared to that of the Mv9kr1 wheat line and the cultivar Mv Nádor.
For molecular marker analyses, genomic DNA was isolated from the line WT153397, the wheat genotypes Mv9kr1 and ‘Mv Karizma,’ the parental A. glael clone No. 8, the Aegilops tauschii (2n = 2x = 14, DD) accession MvGB605, and the durum wheat (T. turgidum ssp. durum) cultivar Langdon (2n = 4x = 28, BBAA).
Genomic DNA isolated from Th. bessarabicum, Pseudoroegneria spicata, Aegilops tauschii, Secale cereale L., Oryza sativa L., T. aestivum, and T. turgidum ssp. durum was used as probes or blocking DNA for GISH, or as templates to amplify specific DNA sequences for fluorescence in situ hybridization (FISH).
Probe preparation and fluorescence in situ hybridization
Mitotic chromosome preparations were made from germinating root tips, as described by Lukaszewski et al. (2004). In order to visualize Thinopyrum (A. glael) chromatin in the wheat genetic background, the following genomic probes were applied: total genomic DNA from Th. bessarabicum was labeled with biotin-11-dUTP (Roche, Mannheim, Germany) and used as J-genomic probe, while DNA from Ps. spicata was labeled with digoxigenin-11-dUTP (Roche) or biotin-11-dUTP using random priming labeling and used as St-genomic probe, as described by Kruppa and Molnár-Láng (2016). Digoxigenin-11-dUTP-labeled DNA isolated from Ae. tauschii was used as a D-genomic probe.
For the FISH experiments, Afa family and pSc119.2 DNA repeats were amplified from the genomic DNA of Ae. tauschii and S. cereale, respectively, and labeled using PCR (Nagaki et al. 1995; Contento et al. 2005) with digoxigenin-11-dUTP (Roche) and biotin-16-dUTP (Roche, Mannheim, Germany), respectively. The 18S unit of the 45S rDNA was amplified using PCR from rice genomic DNA (Chang et al. 2010) and labeled with 50% biotin-16-dUTP and 50% digoxigenin-11-dUTP. Microsatellite (GAA)7 was amplified from T. aestivum genomic DNA and labeled with digoxigenin-11-dUTP (Roche) using PCR (Pedersen and Linde-Laursen 1994). Anti-digoxigenin-rhodamine Fab fragments (Roche) and streptavidine-Alexa Fluor 488 (Molecular Probes, Waltham, USA) were used to detect digoxigenin and biotin, respectively.
Chromosomal composition of the line WT153397 was determined using sequential multicolor genomic in situ hybridization (mcGISH) and fluorescence in situ hybridizations (FISH). GISH reaction mix (15 µL per slide) containing 50% formamide, 2 × SSC, 10% dextran sulfate, 100 ng each of the J- and St-genomic probes, and 3 µg T. aestivum blocking DNA was allowed to hybridize overnight at 42 °C. The hybridization mixes used in the GISH experiments to visualize St- and D-genomic parts of the translocated chromosome contained the same amount of St- and D-genomic probes and durum wheat blocking DNA as described above. GISH signals were documented under a microscope after signal detection and chromosome counterstaining with 2 µg mL−1 DAPI (4′,6-diamidino-2-phenylindole, Amersham, Germany). Following washing (3 × 30 min in 4 × SSC Tween, 2 × 5 min 2 × SSC at 25 °C), the slides were re-hybridized with DNA repeat probes.
The protocol for the FISH experiments was modified as follows. The hybridization temperature was 37 °C for probes pSc119.2, Afa family, and pTa71, and 42 °C for probe (GAA)7 (in a third step hybridization). The hybridization mixture (30 µL per slide) contained 20 ng of pTa71, 70 ng of each of the pSc119.2 and Afa family probes, and salmon sperm as blocking DNA.
The GISH and FISH hybridization sites were documented using an AxioImager M2 fluorescence microscope (Zeiss, Oberkochen, Germany) equipped with an AxioCam MRm CCD camera (Zeiss, Oberkochen, Germany) and filter sets appropriate for DAPI (Zeiss Filterset 49), Alexa Fluor 488 (Zeiss Filterset 38), and Rhodamine (Zeiss Filterset 20). The images were processed with the AxioVision 4.8.2. software (Zeiss, Oberkochen, Germany).
Flow cytometric chromosome analysis and sorting
Suspensions of intact mitotic metaphase chromosomes from the line WT153397 were prepared from synchronized root tips cells of young seedlings following Vrána et al. (2016a; 2016b). Prior to flow cytometric analysis, chromosomes in suspension were fluorescently labeled by FISHIS (Giorgi et al. 2013) with oligonucleotide probe 5′-FITC-(GAA)7-FITC-3′ (Integrated DNA Technologies, USA) and stained with DAPI. Bivariate flow karyotyping and chromosome sorting were carried out using a FACSAria II SORP flow cytometer and sorter (Becton Dickinson Immunocytometry Systems, San José, USA) as described by Molnár et al. (2016) and Said et al. (2019, 2022). The chromosome samples were analyzed at a rate of 900–1400 particles per second, and bivariate flow karyotypes of FITC vs. DAPI fluorescence were acquired. The sorting window was set to the chromosome population of interest on the dot plots, and chromosomes were flow-sorted at a rate of 15–24 per second. Approximately 3000 chromosomes from each chromosome fraction were flow-sorted onto a microscope slide into a 3 µL drop of PRINS buffer supplemented with 2.5% sucrose (Kubaláková et al. 1997). The chromosome content of the flow-sorted fractions was determined using FISH, as described by Kruppa et al. (2016).
Molecular marker analysis
A total of 26 simple sequence repeats (SSR) and two conserved ortholog set (COS) markers previously mapped on chromosome 6D were selected from the GrainGenes database (https://wheat.pw.usda.gov/GG3/) and used to characterize the wheat/A. glael translocation in the line WT153397. Genomic DNA was extracted from fresh young leaves of wheat genotypes Mv9kr1 and ‘Mv Karizma,’ the parental A. glael clone No. 8, the Ae. tauschii accession MvGB605, the durum wheat cultivar Langdon, and the WT153397 translocation line using a Quick Gene-Mini80 device (FujiFilm, City, Japan) and a QuickGene DNA tissue kit (FujiFilm) according to the manufacturer’s instructions. The PCR reactions were carried out in an Eppendorf Mastercycler (Eppendorf, City, Germany). Reaction mixes (15 µL) contained 20 ng of template DNA, 1.5 µL of 10 × key reaction buffer (MgCl2 final concentration of 1.5 mmol/L), 200 µmol/L of each dNTP, 0.2 µmol/L of forward and reverse primers, and 0.375 U of TEMPase Hot Start DNA Polymerase (VWR International, City, Belgium). The PCR profiles and annealing temperatures of the primer pairs used in the reactions are available in the GrainGenes database (https://wheat.pw.usda.gov/GG3/) and are summarized in Supplementary Table S1. The PCR amplicons were separated using a Fragment Analyzer Automated CE System (Advanced Analytical Technologies, City, USA) equipped with a 96-capillary array cartridge (effective length 33 cm) and analyzed with the PROsize v2.0 software (Advanced Analytical Technologies).
The physical positions of the markers on chromosome 6D were determined in silico using a sequence similarity approach. The publicly available primer sequences of the markers (Supplementary Table S1) were used as queries in BLASTn searches (Ensembl Plants release 46; https://plants.ensembl.org/Multi/Tools/Blast) against the reference pseudomolecules of A, B, and D genomes of hexaploid wheat (International Wheat Genome Sequencing Consortium (IWGSC) et al. 2018). The start position of each primer sequence was determined by the best hits obtained on the wheat reference sequence using the BLASTn package of the Blast Command Line Application 2.9.0 (ftp:/ftp.ncbi.nlm.nih.gov/) with the following parameters: -task ‘BLASTn’; -evalue 1e-5; -max target seqs 2; -max hsps 1.
Field phenotyping experiments and experimental design
Morphological traits of the line WT153397 and the wheat genotypes Mv9kr1, ‘Chinese Spring,’ and ‘Mv Karizma’ were evaluated in a low-input, pesticide-free location (Tükrös nursery, Martonvásár, Hungary; geographic coordinates: 47°18′40″N 18°46′56″E) in the 2018–2019, 2020–2021, 2021–2022 and 2022–2023 growing seasons, as well as in a high-input nursery (Bulgárföld, Martonvásár, Hungary, geographic coordinates: 47°19′39″N, 18°47′01″E) in the 2021–2022 and 2022–2023 seasons. During the 2022–2023 growing season, the phenotype of the WT153397 × ‘Mv Nádor’ F5 progeny plants was also assessed. In the low-input Tükrös nursery, 40 grains of each genotype were sown (4 × 1 m rows; 10 grains per row) with a 0.15 m row spacing, whereas in the high-input experiments, 250 grains of each genotype were sown in 2 m2 plots (5 × 2 m rows; 50 grains per row) with a row distance of 0.15 m, as previously described by Mikó et al. (2014). The trial plots in the high-input nursery were machine-planted (HEGE-80 plot driller) and hand-harvested. Herbicides, insecticides, and artificial fertilizers were used in the high-input fields when necessary, but fungicides were not. The soil has a clayey chernozem texture and a pH of 7.25. The soil contained 2.8% w/w humus, 210 mg/kg P2O5, and 210 mg/kg K2O. The annual average N intake as NPK combined fertilizer was 120 kg/ha active ingredient (Tóth et al. 2022). Herbicides, fungicides, and insecticides were not used in the low-input nursery. Genotypes were sown close to each other to minimize the confounding effects of soil and microclimatic conditions. Additional growing parameters of the field trial locations are summarized in Supplementary Table S2. The weather conditions varied between the years (Supplementary Table S3) but were typical of the Central-European climate measured over the last 25 years (Vida et al. 2022). The first season had average rainfall but high temperatures, with several heat days. The following two seasons were drier and warmer, especially during the grain-filling period. The last season had the most precipitation and the fewest heat days.
In the high-input nursery, a randomized small-plot complete block design with three replications was used. The plot size was 2 m2. The outside rows were always excluded during the analysis. In the low-input nursery, borders were sown at the beginning and end of the previous and next parcels, respectively. For phenotypic characterization, ten typical plants were selected from each genotype in each location and season. Plant height (cm) and tillering (number of tillers per plant) were measured in the field immediately before harvest. The length of the main spike (cm), the number of spikelets per main spike, the number of seeds per main spike, the number of seeds per spikelet, and the number of seeds per plant were determined after harvest. The total number of spikes and the yield of the plots were also measured in the high-input experiment. Seed length and width (mm) as well as thousand grain weight (g) were measured using a MARVIN 5.0 Seed Analyzer (MARViTECH GmbH, Germany). The heading time of each genotype grown in a low-input nursery between 2019 and 2023 was determined as the time between January 1 and the day when 50% of the inflorescences reached the DEV59 developmental stage (Tottman 1987).
Statistical analysis
Significant differences were calculated from ten biological replicates for each year between the line WT153397 and the wheat genotypes Mv9kr1, ‘Chinese Spring,’ and ‘Mv Karizma,’ as well as between the line WT153397 × ‘Mv Nádor’ F5 generation and wheat control genotypes (‘Mv Nádor’ Mv9kr1) using the Tukey’s post hoc test of the IBM SPSS Statistics 20.0 software (SPSS Inc., Chicago, IL, USA).
Results
Development and molecular cytogenetic analysis of the line WT153397
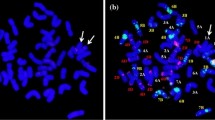
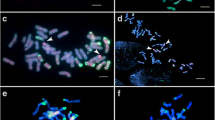
A schematic representation of the crossing program used to develop wheat/A. glael introgression lines is shown in Supplementary Figure S1. Partial amphiploids from the BC1F7 generation were backcrossed with ‘Mv Karizma’ in two cycles to produce BC2F1 and BC3F1 populations. A molecular cytogenetic screening for the presence of Agropyron chromosomes and wheat/Agropyron translocations was performed on the BC3F2 generation. Because A. glael contains genomes J, Js, and St, a combination of digoxigenin-labeled St-genomic probe from Ps. spicata (red) and biotin-labeled J-genomic probe from Th. bessarabicum (green) was used for in situ hybridization. GISH analysis with this probe combination (combination 1) identified a wheat line (WT153397) with 42 chromosomes, including a disomic wheat/A. glael translocation chromosome. A red fluorescent signal indicated that a St-genomic segment had been terminally translocated to a wheat chromosome, whereas the absence of green hybridization signals indicated that no J-genomic fragment was involved in the translocation (Fig. 1a).
Molecular cytogenetic analysis of the wheat/A. glael translocation line WT153397. a GISH with Ps. spicata St-genomic DNA (red) revealed a disomic wheat/A. glael translocation, while biotin-labeled Th. bessarabicum Jb-genomic DNA (green) produced no hybridization signals. The chromosomes were counterstained with DAPI (blue). Inset: the translocated chromosome. b FISH on the same mitotic metaphase cell with the Afa family (red), pSc119.2 (green), and pTa71 (yellow) DNA repeat probes. Inset: The FISH pattern of a normal wheat chromosome 6D (left) and the translocated chromosome (right). c FISH pattern of the (GAA)7 microsatellite probe on the same mitotic cell. Inset: the translocated chromosome with intense GAA clusters on the long arm. d McGISH performed with a combination of digoxigenin-labeled D-genomic- (red) and biotin-labeled St-genomic (green) probes, together with A- and B-genomic blocking DNA from T. turgidum ssp. durum. The red fluorescent signal indicates that only the short arm of the translocation has derived from the wheat D genome. Inset: the translocated chromosome. Scale bars = 10 µm
The slides used for GISH were then subjected to FISH using repetitive DNA probes pSc119.2, Afa family, and pTa71 (Fig. 1b) to identify the wheat chromosome involved in the translocation. The resulting FISH hybridization patterns confirmed the presence of 14 A-genome and 14 B-genome chromosomes. Strong Afa signals also confirmed the presence of one pair for each 1D-5D and 7D chromosome with normal FISH karyotype and the absence of 6D chromosomes. However, a strong Afa signal was detected on the telomeric part of the translocated wheat chromosome arm, which corresponded to the short arm of 6D, suggesting that the A. glael chromosome segment was translocated to the 6D wheat chromosome.
In a third step, using the (GAA)7 microsatellite probe, intense GAA clusters were detected along the recombinant chromosome arm from the near centromeric region to the telomeric part (Fig. 1c), which is not typical for the D genome (Pedersen and Langridge 1997). This result suggests that the rearranged chromosome arm does not contain D-genome fragments.
To further investigate the structure of the translocation and to visualize its D-genomic part, we used a second combination of biotin-labeled St-genomic probe and digoxigenin-labeled D-genomic probe, as well as A- and B-genomic blocking DNA from T. turgidum ssp. durum. The GISH analysis revealed that only the short arm of the translocation originated from the D genome, while the long arm originated from A. glael, with the terminal part belonging to the St genome (Fig. 1d).
Flow cytometric analysis and chromosome sorting
The presence of large GAA clusters on the wheat/A. glael translocated chromosome was confirmed by flow cytometric analysis and chromosome sorting, in which the chromosome suspension was labeled by FISHIS using a FITC conjugated (GAA)7 oligonucleotide probe. The bivariate flow karyotype of wheat/A. glael translocation line WT153397 (Supplementary Fig. S2) revealed a chromosome population that is not typical for the standard flow karyotype of hexaploid wheat (Said et al. 2022). The position of the translocated chromosome on the dot plot indicated that its relative DNA content was lower than that of the B-genome chromosomes, but similar to that of the wheat A- and D-genome chromosomes. However, because the Agropyron chromosome segment has prominent (GAA)7 clusters, its FITC fluorescence was much higher than that of the wheat A- and D-genome chromosomes. These differences allowed us to distinguish the 6D/A. glael translocated chromosome from the wheat A-, B-, and D-genome chromosomes (Supplementary Figure S2a). The analysis of 126 chromosomes flow-sorted from the candidate population onto microscopy slides after FISH with pSc119.2, Afa family and pTa71 (Supplementary Figure S2b) confirmed that the translocation could be sorted at high purity (94.4%) with negligible contamination by wheat chromosomes 2D (2.3%), 4A (1.65%) and 4B (1.65%).
The structure of the wheat/Agropyron translocation
The homoeologous relationships between the introgressed A. glael chromatin and wheat were also investigated using molecular markers specific for wheat group 6 chromosomes. Twenty-six SSR and two COS markers previously mapped on chromosome 6D were used, and their physical positions on 6D were determined using BLASTn (Supplementary Table S4).
All 11 markers specific for the short arm of chromosome 6D were detected in the introgression line WT153397, but none of the 17 markers specific for 6DL amplified PCR products on the translocated chromosome, indicating that the long arm of the 6D chromosome was missing from the translocation (Figs. 2 and 3). Five of the 17 markers mapped on the 6DL chromosome arm produced polymorphic fragments between wheat and A. glael. All of these markers amplified Agropyron-specific products in the line WT153397 suggesting that the missing chromosome arm 6DL was replaced by an Agropyron chromosome segment homeologous for wheat group 6 chromosomes (Fig. 3a, b, and c). As a result, the translocated chromosome can be identified as a 6DS.6Ag Robertsonian translocation.
Capillary gel electrophoretogram of 6DL chromosome-specific SSR markers. The markers Xcfd95 and BE518349 amplified specific PCR fragments on both the parental wheat genotypes (Mv9kr1 and ‘Mv Karizma’) and A. glael. The PCR amplicons specific for 6DL in the parental wheat genotypes and Ae. tauschii are indicated by black arrows. The polymorphic fragments (red arrows) amplified on A. glael and the line WT153397 indicate that the wheat 6DL is replaced by a chromosome segment from A. glael in the translocated chromosome. The control durum wheat cultivar Langdon, which lacks D genome, did not produce 6DL-specific PCR fragments. A 35–500 bp DNA ladder was used as a molecular-weight size marker
a The physical locations of twenty-six SSR and two COS markers on the wheat 6D chromosome and the translocated chromosome of the line WT153397. The 6DS-specific markers are present in both wheat and WT153397, but the 6DL-specific markers are absent from the long arm of the translocated chromosome of WT153397, with the exception of five markers (marked with asterisks) that amplified polymorphic PCR fragments. b In situ hybridization with the D-genomic probe (red) confirms the presence of the 6DS chromosome arm in the line WT153397, but the intense (GAA)7 clusters on the long arm of the translocated chromosome exclude the presence of the 6D long arm. Horizontal lines in Figure a and b mean the centromere (C) position
Morphological traits and yield components of the WT153397 line
In order to clarify whether the introgressed Agropyron chromosome segment influences wheat morphological traits, the line WT153397 was compared with the parental wheat genotypes Mv9kr1, ‘Chinese Spring,’ and ‘Mv Karizma’ in four field trials (2019, 2021, 2022, and 2023) under low-input, and two trials (2022, 2023) under high-input conditions (Supplementary Table S5 and Fig. 4). The spike and seed morphology of the WT153397 translocation line and the wheat control genotypes are presented in Fig. 5.
Plant heights (A), number of spikes/plant (tillering) (B), number of grain/plant (C), and thousand-grain weight (D) at wheat control genotypes (Mv9kr1, ‘Mv Karizma,’ and ‘Chinese Spring’) and the WT153397 translocation line in four growing seasons (2018–2019, 2020–2021, 2021–2022, and 2022–2023) in low-input and high-input nurseries, Martonvásár, Hungary. Different letters indicate significant differences in Tukey’s post hoc test at P < 0.05. Bars: ± standard deviation
Spike and seed morphology (A) and flowering time (B) of the parental wheat genotypes Mv9kr1, ‘Chinese Spring’, and ‘Mv Karizma’, and the wheat/A.glael translocation line WT153397 grown in the high-input nursery (spikes were collected in Martonvásár, July 2022) wheat/A. glael translocation line WT153397 and the parental wheat genotypes Mv9kr1, ‘Mv Karizma’, and ‘Chinese Spring’ grown in the high-input nursery (spikes were collected in Martonvásár, July 2022)
The presence of the alien chromosome segment in the new translocation line has no noticeable negative side effects when compared to the morphological and yield components of wheat. The plant height for WT153397 generally did not differ from that of ‘Mv Karizma’ and Mv9kr1, or WT153397 plants were 11–12% shorter in low-input (2021) and high-input (2022) trials (Fig. 4A). On the other hand, Tukey’s post-hoc analysis revealed a high tillering capacity for WT153397, with values ranging from 10.6 to 16.3. This significantly exceeded the number of productive tillers per plant (6.5–12.0) of the wheat genotypes Mv9kr1 and ‘Mv Karizma,’ as well as ‘Chinese Spring’ (6.8–8.7) under both low- and high-input conditions over four growing seasons (Fig. 4B). WT153397 and ‘Mv Karizma’ had a weak tendency for longer spikes and a higher number of seeds per main spike compared to the other wheat genotypes, but differences between WT153397 and the parental wheat genotypes were significant only in the low-input nursery in 2022 (Supplementary Table S5 and Fig. 4). There were no significant differences in the parameters of spikelets per spike and seeds per spikelet between WT153397 and its wheat parents. The increased number of productive tillers and the slightly longer spikes resulted in the highest grain yield (seeds/plant) value of the line WT153397, which was 37–79% higher than the values of ‘Mv Karizma’ under low-input conditions, and 17–46% higher under high-input condition (Supplementary Table S5). This implies that normal sowing density has no effect on the tillering ability and yield potential of the newly developed wheat/A. glael translocation line.
There was a tendency for higher TGW of Mv9kr1 relative to ‘Mv Karizma’ because its grains are wider than those of ‘Mv Karizma’ (Figs. 4 and 5, Supplementary Table S6.). The translocation line had the smallest grains with shorter lengths and width. (It is worth noting that the grain width of the translocation did not differ from that of Mv Karizma.) This grain morphological difference of WT153397 was reflected in the smallest TGW. The superior tillering ability of the line WT153397 over Mv9kr1 and ‘Mv Karizma’ resulted in higher yield weight values (g/plant), which was significantly higher in the 2022 high-input and 2023 low-input experiments (Fig. 4D, Supplementary Table S6). The flowering time of WT153397 exceeded that of ‘Mv Karizma’ by 2–5 days in the 4 years studied, indicating that the introgressed A. glael chromosome segment has no effect on wheat flowering time.
Because the grain size differed between the two wheat parents Mv9kr1 and Mv Karizma, we wanted to know if the positive effect of the A. glael introgression on tillering capacity and yield can be expressed in a different wheat background and whether the small grain size can be increased. To accomplish this, the WT153397 line was crossed with a Hungarian top winter wheat variety ‘Mv Nádor’ as part of a breeding program. The WT153397 × ‘Mv Nádor’ F5 progeny was studied on plants grown in the high-input nursery together with the recurrent parental wheat ‘Mv Nádor’ and Mv9kr1 in the 2023 growing season (Supplementary Table S7). The WT153397 × ‘Mv Nádor’ F5 plants had a significantly higher tillering ability (almost twice as many spikes/plant), number of seeds/main spike, and fertility (number of seeds/spikelet) than both wheat control genotypes. Moreover, there was no difference in the TGW between the F5 plants and ‘Mv Nádor,’ indicating that the smaller TGW observed in WT153397 can be increased in an advanced wheat genetic background.
These morphological traits of the line WT153397 indicate that the 6DS.6Ag translocation is a compensatory type and may be a promising gene source for improving wheat morphology and yield components.
Discussion
A. glael is a promising gene variant reservoir for wheat breeding. The present study is the first to report on a genetically stable introgression line with 42 chromosomes, including a homozygous translocated chromosome carrying a chromosome arm from A. glael.
Molecular cytogenetic analyses revealed that wheat chromosome arm 6DS was involved in the translocation. The introgressed Agropyron chromosome arm was identified by a terminal St-genomic signal, missing GISH signals with Th. bessarabicum DNA and massive GAA clusters from the near centromeric region to the telomere. These results are consistent with those of Cseh et al. (2019), who found that all of the Jvs-genome chromosomes of Th. intermedium, the parental species of A. glael, have telomeric hybridization signals with the St-genomic probe, eight of them also have centromeric signals, and the V-genomic probe from D. villosum hybridizes with all of them. Giorgi et al. (2013) also reported that the D. villosum V-genome chromosomes contain a high number of GAA microsatellite repeats. The strong GAA clusters allowed fluorescent labeling of the V chromosomes in FISHIS using FITC-conjugated GAA oligonucleotide probes and discrimination of the V chromosomes on the bivariate (FITC vs. DAPI) flow karyotype (Giorgi et al. 2013). Previous studies also indicated that the Jvs genome preserved repetitive sequences from the D. villosum V genome (Kishii et al. 2005; Mahelka et al. 2011; Cseh et al. 2019). It seems reasonable to conclude that GAA microsatellites are common repetitive DNA fractions in D. villosum and the Th. intermedium Jvs genome. A molecular cytogenetic analysis of diploid ancestors of St- and Jb-genomes, Ps. spicata, and Th. bessarabicum, respectively, revealed that only one St-genome chromosome contains a GAA signal, while the Jb chromosomes do not contain microscopically detectable GAA clusters (Linc et al. 2017).
Based on previous findings and our current results showing the presence of GAA clusters in the introgressed chromosome arm of WT153397, we conclude that the transferred A. glael chromosome arm belongs to the Jvs genome. Our molecular marker analysis confirmed the presence of the 6DS chromosome arm in the line WT153397, as well as the fact that the introgressed chromosome arm is homoeologous with the missing 6DL. The homoeologous relationship between the diploid Thinopyrum genomes and the wheat genome has been established in several publications (Wang et al. 2010; Gaál et al. 2018; Grewal et al. 2018; Baker et al. 2020; Yang et al. 2021). A comparison of Th. bessarabicum and wheat chromosomes revealed that the macrosynteny between the two species is preserved, and the majority of Th. bessarabicum chromosome arms are collinear with the corresponding A-, B-, and D-genome chromosomes of hexaploid wheat (Grewal et al. 2018). Baker et al. (2020) found that the D genome had the most significant BLAST hits for the markers on the diploid Th. elongatum genetic map and that good overall synteny is maintained between wheat and Th. elongatum, proving the close relationship between the two genomes. Based on synteny analyses between the Th. elongatum and wheat genomes, Yang et al. (2021) also confirmed that the 2A and 2E chromosomes, as well as the 5D and 5E chromosomes, appear to have a good collinearity relationship. Their results agreed with those of Gaál et al. (2018), who used EST sequences to study the macrosyntenic relationship between Th. elongatum and wheat genomes. Even in polyploid wheatgrasses, the homeology of their genomes with wheat genomes is well preserved.
Kantarski et al. (2017) applied genotyping-by-sequencing (GBS) technology to develop a consensus genetic map for Th. intermedium. By aligning the 10,029 mapped markers to the barley reference sequence, significant syntenic relationships between the linkage groups of Thinopyrum and barley were found. Later, using an Axiom 35 K Wheat-Relative Genotyping Array to characterize 187 wheat/Th. intermedium introgression lines, 634 chromosome-specific SNPs were validated (Cseh et al. 2019). The authors found a significant macrosyntenic relationship between the 21 chromosomes of Th. intermedium and their homoeologues from the wheat D genome, including the 6Jvs-6D synteny. The close 6Jvs-6D homoeologous relationship is also supported by the present study, as we observed good compensation ability of the introgressed A. glael chromosome arm for the missing 6DL for morphological traits and fertility. After four seasons of field trials, there were no significant differences in the spike morphology (spike length, spikelets per spike) and fertility between the introgression line WT153397 and the wheat genotypes Mv9kr1 and ‘Mv Karizma.’
The quality analyses revealed no significant differences in total protein and dietary fiber content (total arabinoxylan), as well as the glutenin/gliadin (Glu/Gli) ratio and unextractable polymeric protein content (UPP%) between the WT153397 line and the wheat parents (data not shown). Because group 6 chromosomes contain genes that determine wheat glutenin and gliadin proteins (Payne et al. 1984, 1987), the similar quality traits of WT153397 support the good compensation ability of 6JVS chromosome arm for the loss of 6DL. However, additional experiments are needed to obtain precise data on the grain chemical composition and bread-making quality of this translocation.
It is important to note that approximately 93.75% of the genes in Mv9kr1 originate from the wheat cultivar Mv9, which adapted well to the agro-ecological conditions of Central Europe (Molnár-Láng et al. 1996), while ‘Mv Karizma’ is a modern elite wheat cultivar registered in 2009. However, field experiments confirmed the translocation line’s outstanding tillering ability, which significantly exceeded the capacity of productive tiller formation in all wheat control genotypes under both low- and high-input conditions.
Tillering is a complex trait determined by the combined effects of multiple genes. Although four single genes (tin1, tin2, tin3, and ftin) responsible for tiller inhibition have been identified on wheat chromosomes 1A, 2A, and 3A (Spielmeyer and Richards 2004; Kuraparthy et al. 2007), it was found that a majority of underlying variation for tillering was controlled by quantitative trait loci (QTL) primarily distributed on chromosomes 1A, 2A, 2D, 6A, 6D, 7A, and 7D (Li et al. 2010; Ren et al. 2018; Wang et al. 2018). Our results show that replacing the 6DL chromosome arm with the 6Jvs chromosome arm increased the capacity for tiller formation, which is consistent with the findings of Bilgrami et al. (2020). They identified two metaQTLs (MQTL6D-3 and MQTL6D-4) on the terminal region of wheat chromosome 6DL. Our results indicate that QTLs on the 6Jvs chromosome arm influence tiller formation more positively than QTLs on the 6DL.
Because the 6DS.6Jvs translocated chromosome can be purified at high purity using flow cytometric sorting, identifying key genes on 6Jvs responsible for tiller formation using mutant chromosome sequencing (MutChromSeq) will be facilitated. EMS mutagenesis is a suitable method for producing low-tillering wheat mutants, as demonstrated by Dong et al. (2023) for the cloning of a key tiller number regulatory gene, TN1. Combining EMS mutagenesis with flow cytometric chromosome sorting and sequencing by MutChromSeq is a powerful approach for rapid gene isolation and identification (Sánchez-Martín et al. 2016), as demonstrated by the cloning of disease resistance genes (Yu et al. 2022; Holden et al. 2022) or genes related to plant morphology (Borrill et al. 2022). The development of low-tillering knock-out EMS mutants, followed by flow sorting and sequencing of the 6DS.6Jvs translocated chromosomes from the mutant and wild type WT153397, may be an effective strategy in the future to obtain gene content of 6Jvs and identify the candidate gene(s) affecting tillering.
Introgression breeding is hampered by several factors, including the taxonomic relationship between wheat and its relatives, cross-compatibility, fertility of the hybrids, and their offspring (Zhang et al. 2016). The introgression of large alien chromosome segments is also accompanied by the occurrence of the linkage drag phenomenon, which can have a negative impact on grain yield and quality traits (Johansson et al. 2020). However, by using additional backcrosses, appropriate crossing strategies, and selfings, the introgressed chromosome fragment can be shortened, reducing the effect of linkage drag, which causes poorer agronomic features. The translocation line WT153397 was developed through three backcrosses with the recurrent parents, including two backcrosses with ‘Mv Karizma.’ The agronomic characteristics of the translocation line are most similar to ‘Mv Karizma’ in terms of plant height, as well as spike and seed morphology, though the WT153397 line’s spikes are slightly longer and laxer than those of ‘Mv Karizma.’ Its tillering ability and number of seeds per spike were significantly higher than those of the controls in both the low-input and high-input nurseries, indicating that normal sowing density had no effect on the tillering capacity of the line WT153397. The only morphological trait that differed significantly from the wheat controls was the kernel size (thinner grain) and, as a result, the smaller TGW. The fact that the grain width and TGW increased in the ‘Mv Nádor’ genetic background, as observed in the WT153397 × ‘Mv Nádor’ F5 lines, supports the possibility that the smaller grain width of the line WT153397 originated from ‘Mv Karizma.’ However, larger-scale field trials should be carried out in the future to support the positive contribution of higher tillering capacity of the 6DS.6Jvs translocation to higher wheat yield formation. Mapping and cloning of the Agropyron gene responsible for improved tillering will also aid wheat breeding programs in utilizing this gene source.
Conclusion
Wheat/Thinopyrum introgressions can be used to improve not only stress resistance but also yield components of bread wheat. The translocation line WT153397 is genetically stable, with no evidence of linkage drag. The presence of the A.glael 6Jvs chromosome arm in subsequent generations can be tracked using specific molecular markers as well as molecular cytogenetic methods. According to preliminary results from environments with low and high nutrient supplies, this translocation can even develop reproductive tillers with a 35–45% higher efficiency. The WT153397 × ‘Mv Nádor’ F5 plants demonstrated that the improved tillering caused by the 6D.S6Jvs translocation chromosome can be combined with larger seed size in the advanced wheat genetic background, potentially resulting in higher yield potential. In the future, large-scale field trials and precise breeding programs will be needed to confirm this finding and investigate end-use quality and other agronomic traits. These advantages enable this line to be used in wheat breeding programs as well as functional genomic studies aiming to clarify the genetic control of reproductive tiller formation.
Data availability
All data generated or analyzed during this study are included in this article and its supplementary information file. Seeds of the described translocation line are freely available to any researcher wishing to use them for non-commercial purposes.
References
Baker L, Grewal S, Yang C-Y, Hubbart-Edwards S, Scholefield D, Ashling S, Burridge AJ, Przewieslik-Allen AM, Wilkinson PA, King IP, King J (2020) Exploiting the genome of Thinopyrum elongatum to expand the gene pool of hexaploid wheat. Theor Appl Genet 133:2213–2226. https://doi.org/10.1007/s00122-020-03591-3
Bilgrami SS, Ramandi HD, Shariati V, Razavi K, Tavakol E, Fakheri BA, Mahdi Nezhad N, Ghaderian M (2020) Detection of genomic regions associated with tiller number in Iranian bread wheat under different water regimes using genome-wide association study. Sci Rep 10:14034. https://doi.org/10.1038/s41598-020-69442-9
Borrill P, Mago R, Xu T, Ford B, Williams SJ, Derkx A, Bovill WD, Hyles J, Bhatt D, Xia X, MacMillan C, White R, Buss W, Molnár I, Walkowiak S, Olsen O-A, Doležel J, Pozniak CJ, Spielmeyer W (2022) An autoactive NB-LRR gene causes Rht13 dwarfism in wheat. Proc Nat Acad Sci USA (PNAS) 119:e2209875119. https://doi.org/10.1073/pnas.2209875119
Carver F, Rayburn AL (1994) Comparison of related wheat stocks possessing 1B or 1RS.1BL chromosomes: agronomic performance. Crop Sci 34:1505–1510. https://doi.org/10.2135/cropsci1994.0011183X003400060017x
Ceoloni C, Kuzmanović L, Gennaro A, Forte P, Giorgi D, Rosaria Grossi M, Bitti A (2014) Genomes, chromosomes and genes of the wheatgrass genus Thinopyrum: the value of their transfer into wheat for gains in cytogenomic knowledge and sustainable breeding. Genomics of plant genetic resources: crop productivity, food security and nutritional quality, pp 333–358. https://doi.org/10.1007/978-94-007-7575-6_14
Chang KD, Fang S, Chang FC, Chung MC (2010) Chromosomal conservation and sequence diversity of ribosomal RNA genes of two distant Oryza species. Genomics 96:181–190. https://doi.org/10.1016/j.ygeno.2010.05.005
Chen Q, Conner RL, Laroche A, Thomas JB (1998) Genome analysis of Thinopyrum intermedium and Thinopyrum ponticum using genomic in situ hybridization. Genome 41:580–586. https://doi.org/10.1139/g98-055
Colmer TD, Flowers TJ, Munns R (2006) Use of wild relatives to improve salt tolerance in wheat. J Exp Bot 57:1059–1078. https://doi.org/10.1093/jxb/erj124
Contento A, Heslop-Harrison JS, Schwarzacher T (2005) Diversity of a major repetitive DNA sequence in diploid and polyploid Triticeae. Cytogenet Genome Res 109:34–42. https://doi.org/10.1159/000082379
Cseh A, Yang C, Hubbart-Edwards S, Scholefield D, Ashling SS, Burridge AJ, Wilkinson PA, King IP, King J, Grewal S (2019) Development and validation of an exome-based SNP marker set for identification of the St, Jr and Jvs genomes of Thinopyrym intermedium in a wheat background. Theor Appl Genet 132:1555–1570. https://doi.org/10.1007/s00122-019-03300-9
Darwinkel A (1980) Ear development and formation of grain yield in winter wheat. NJAS 28:156–163. https://doi.org/10.18174/njas.v28i3.17029
Deng C, Bai L, Fu S, Yin W, Zhang Y, Chen Y, Wang RR-C, Zhang X, Han F, Hu Z (2013) Microdissection and chromosome painting of the alien chromosome in an addition line of wheat–Thinopyrum intermedium. PLoS ONE 8:e72564. https://doi.org/10.1371/journal.pone.0072564
Dong C, Zhang L, Zhang Q et al (2023) Tiller Number1 encodes an ankyrin repeat protein that controls tillering in bread wheat. Nat Commun 14:836. https://doi.org/10.1038/s41467-023-36271-z
Dubcovsky J, Galvez AF, Dvořák J (1994) Comparison of the genetic organization of the early salt-stress-response gene system in salt-tolerant Lophopyrum elongatum and salt-sensitive wheat. Theor Appl Genet 87:957–964. https://doi.org/10.1007/BF00225790
FAOSTAT (2020) Food and Agriculture Organization of the United Nations Food and agriculture data. https://www.fao.org/faostat/en/#data/QCL. Accessed 19 Dec 2022
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87. https://doi.org/10.1007/BF00035277
Gaál E, Valárik M, Molnár I, Farkas A, Linc G (2018) Identification of COS markers specific for Thinopyrum elongatum chromosomes preliminary revealed high level of macrosyntenic relationship between the wheat and Th. elongatum genomes. PLoS One 13:e0208840. https://doi.org/10.1371/journal.pone.0208840
Gao L, Koo D-H, Juliana P et al (2021) The Aegilops ventricosa 2NvS segment in bread wheat: cytology, genomics and breeding. Theor Appl Genet 134:529–542. https://doi.org/10.1007/s00122-020-03712-y
Garg M, Yanaka M, Tanaka H, Tsujimoto H (2014) Introgression of useful genes from Thinopyrum intermedium to wheat for improvement of bread-making quality. Plant Breed 133:327–334. https://doi.org/10.1111/pbr.12167
Giorgi D, Farina A, Grosso V, Gennaro A, Ceoloni C, Lucretti S (2013) FISHIS: fluorescence in situ hybridization in suspension and chromosome flow sorting made easy. PLoS ONE 8:e57994. https://doi.org/10.1371/journal.pone.0057994
Grewal S, Yang C, Edwards SH, Scholefield D, Ashling S, Burridge AJ, King IP, King J (2018) Characterisation of Thinopyrum bessarabicum chromosomes through genome-wide introgressions into wheat. Theor Appl Genet 131:389–406. https://doi.org/10.1007/s00122-017-3009-y
Vida G, Szunics L, Veisz O, Cséplő M (2022) Breeding for improved gluten strength and yellow pigment content in winter durum wheat. Bulg J Crop Sci 59:16–33
Hajjar R, Hodgkin T (2007) The use of wild relatives in crop improvement: a survey of developments over the last 20 years. Euphytica (2007) 156:1–13. https://doi.org/10.1007/s10681-007-93630
Holden S, Bergum M, Green P, Bettgenhaeuser J, Hernández-Pinzón I, Thind A, Clare S, Russell JM, Hubbard A, Taylor J, Smoker M, Gardiner M, Civolani L, Cosenza F, Rosignoli S, Strugala R, Molnár I, Šimková H, Doležel J, Schaffrath U, Barrett M, Salvi S, Moscou MJ (2022) A lineage-specific Exo70 is required for receptor kinase-mediated immunity in barley. Sci Adv 8:eabn7258. https://doi.org/10.1126/sciadv.abn7258
International Wheat Genome Sequencing Consortium (IWGSC), Appels R, Eversole K et al (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361. https://doi.org/10.1126/science.aar7191
Jiang J, Friebe B, Gill BS (1993) Recent advances in alien gene transfer in wheat. Euphytica 73:199–212. https://doi.org/10.1007/BF00036700
Johansson E, Henriksson T, Prieto-Linde ML, Andersson S, Ashraf R, Rahmatov M (2020) Diverse wheat-alien introgression lines as a basis for durable resistance and quality characteristics in bread wheat. Front Plant Sci 11:1067. https://doi.org/10.3389/fpls.2020.01067
Kantarski T, Larson S, Zhang X, DeHaan L, Borevitz J, Anderson J, Poland J (2017) Development of the first consensus genetic map of intermediate wheatgrass (Thinopyrum intermedium) using genotyping-by-sequencing. Theor Appl Genet 130:137–150. https://doi.org/10.1007/s00122-016-2799-7
King IP, Purdie KA, Orford SE, Reader SM, Miller TE (1993) Detection of homoeologous chiasma formation in Triticum durum x Thinopyrum bessarabicum hybrids using genomic in situ hybridization. Heredity 71:369–372. https://doi.org/10.1038/hdy.1993.151
Kirby EJM, Jones HG (1977) The relations between the main shoot and tillers in barley plants. J Agric Sci 88:381–389. https://doi.org/10.1017/S0021859600034870
Kishii M, Wang RR, Tsujimoto H (2005) GISH analysis revealed new aspect of genomic constitution of Thinopyrum intermedium. Czech J Genet Plant Breed 41:91–95
Kruppa K, Molnár-Láng M (2016) Simultaneous visualization of different genomes (J, JSt and St) in a Thinopyrum intermedium × Thinopyrum ponticum synthetic hybrid (Poaceae) and in its parental species by multicolour genomic in situ hybridization (mcGISH). Comp Cytogenet 10:283–293. https://doi.org/10.3897/COMPCYTOGEN.V10I2.7305
Kruppa K, Türkösi E, Mayer M, Tóth V, Vida G, Szakács É, Molnár-Láng M (2016) McGISH identification and phenotypic description of leaf rust and yellow rust resistant partial amphiploids originating from a wheat × Thinopyrum synthetic hybrid cross. J Appl Genet 57:427–437. https://doi.org/10.1007/s13353-016-0343-8
Kubaláková M, Macas J, Doležel J (1997) Mapping of repeated DNA sequences in plant chromosomes by PRINS and C-PRINS. Theor Appl Genet 94:758–763. https://doi.org/10.1007/s001220050475
Kulshrestha VP, Chowdhury S (1987) A new selection criterion for yield in wheat. Theor Appl Genet 74:275–279. https://doi.org/10.1007/BF00289980
Kuraparthy V, Sood S, Dhaliwal HS, Chhuneja P, Gill BS (2007) Identification and mapping of a tiller inhibition gene (tin3) in wheat. Theor Appl Genet 114:285–294. https://doi.org/10.1007/s00122-006-0431-y
Kuzmanović L, Gennaro A, Benedettelli S, Dodd IC, Quarrie SA, Ceoloni C (2014) Structural-functional dissection and characterization of yield-contributing traits originating from a group 7 chromosome of the wheatgrass species Thinopyrum ponticum after transfer into durum wheat. J Exp Bot 65:509–525. https://doi.org/10.1093/jxb/ert393
Li Z, Peng T, Xie Q, Han S, Tian J (2010) Mapping of QTL for tiller number at different stages of growth in wheat using double haploid and immortalized F2 populations. J Genet 89:409–415. https://doi.org/10.1007/s12041-010-0059-1
Linc G, Gaal E, Molnar I, Icso D, Badaeva E, Molnar-Lang M (2017) Molecular cytogenetic (FISH) and genome analysis of diploid wheatgrasses and their phylogenetic relationship. PLoS ONE 12:1–18. https://doi.org/10.1371/journal.pone.0173623
Liu ZW, Wang RR (1993) Genome analysis of Elytrigia caespitosa, Lophopyrum nodosum, Pseudoroegneria geniculata ssp. scythica, and Thinopyrum intermedium (Triticeae: Gramineae). Genome 36:102–111. https://doi.org/10.1139/g93-014
Lukaszewski AJ, Rybka K, Korzun V, Malyshev SV, Lapinski B, Whitkus R (2004) Genetic and physical mapping of homoeologous recombination points involving wheat chromosome 2B and rye chromosome 2R. Genome 47:36–45. https://doi.org/10.1139/g03-089
Mahelka V, Kopecký D, Patová L (2011) On the genome constitution and evolution of intermediate wheatgrass (Thinopyrum intermedium: Poaceae, Triticeae). BMC Evol Biol 11:1–17. https://doi.org/10.1186/1471-2148-11-127
Mikó P, Löschenberger F, Hiltbrunner J, Aebi R, Megyeri M, Kovács G, Molnár-Láng M, Vida G, Rakszegi M (2014) Comparison of bread wheat varieties with different breeding origin under organic and low-input management. Euphytica 199:69–80. https://doi.org/10.1007/s10681-014-1171-8
Molnár I, Vrána J, Burešová V, Cápal P, Farkas A, Darkó É, Cseh A, Kubaláková M, Molnár-Láng M, Doležel J (2016) Dissecting the U, M, S and C genomes of wild relatives of bread wheat (Aegilops spp.) into chromosomes and exploring their synteny with wheat. Plant J 88:452–467. https://doi.org/10.1111/tpj.13266
Molnár-Láng M, Linc G, Sutka J (1996) Transfer of the recessive crossability allele kr1 from Chinese Spring into the winter wheat variety Martonvásári 9. Euphytica 90:301–305. https://doi.org/10.1007/BF00027480
Monneveux P, Reynolds MP, Aguilar JG, Singh RP, Weber WE (2003) Effects of the 7DL.7Ag translocation from Lophopyrum elongatum on wheat yield and related morphophysiological traits under different environments. Plant Breed 122:379–384. https://doi.org/10.1046/j.1439-0523.2003.00856.x
Moreno-Sevilla B, Baenziger PS, Peterson CJ, Graybosch RA, McVey DV (1995) The 1BL/1RS translocation: agronomic performance of F3-derived lines from a winter wheat cross. Crop Sci 35:1051–1055. https://doi.org/10.2135/cropsci1995.0011183X003500040022x
Mujeeb-Kazi A, Hettel GP (1995) Utilizing wild grass biodiversity in wheat improvement: 15 years of wide cross research at CIMMYT Report No: 2.p. 1–140
Mujeeb-Kazi A, Kazi AG, Dundas I, Rasheed A, Ogbonnaya F, Kishii M, Bonnett D, Wang RR-C, Xu S, Chen P, Mahmood T, Bux H, Farrakh S (2013) Genetic diversity for wheat improvement as a conduit to food security. Elsevier, pp 179–257. https://doi.org/10.1016/B978-0-12-417187-9.00004-8
Nagaki K, Tsujimoto H, Isono K, Sasakuma T (1995) Molecular characterization of a tandem repeat, Afa family, and its distribution among Triticeae. Genome 38:479–486. https://doi.org/10.1139/g95-063
Oliver RE, Xu SS, Stack RW, Friesen TL, Jin Y, Cai X (2006) Molecular cytogenetic characterization of four partial wheat-Thinopyrum ponticum amphiploids and their reactions to Fusarium head blight, tan spot, and Stagonospora nodorum blotch. Theor Appl Genet 112:1473–1479. https://doi.org/10.1007/s00122-006-0250-1
Payne PI, Holt LM, Jackson EA, Law CN (1984) Wheat storage proteins: their genetics and their potential for manipulation by plant breeding. Phil Trans R Soc Lond B 304:359–371. https://doi.org/10.1098/rstb.1984.0031
Payne PI, Nightingale MA, Krattiger AF, Holt LM (1987) The relationship between HMW glutenin subunit composition and the bread-making quality of British-grown wheat varieties. J Sci Food Agric 1987(40):51–65. https://doi.org/10.1002/jsfa.2740400108
Pedersen C, Langridge P (1997) Identification of the entire chromosome complement of bread wheat by two-colour FISH. Genome 40:589–593. https://doi.org/10.1139/g97-077
Rabinovich SV (1998) Importance of wheat-rye translocations for breeding modern cultivar of Triticum aestivum L. Euphytica 100:323–340. https://doi.org/10.1023/A:1018361819215
Ren T, Hu Y, Tang Y, Li C, Yan B, Ren Z, Tan F, Tang Z, Fu S, Li Z (2018) Utilization of a Wheat 55K SNP Array for mapping of major QTL for temporal expression of the tiller number. Front Plant Sci 9:333. https://doi.org/10.3389/fpls.2018.00333
Reynolds M, Bonnett D, Chapman SC, Furbank RT, Manès Y, Mather DE, Parry MAJ (2011) Raising yield potential of wheat. I. Overview of a consortium approach and breeding strategies. J Exp Bot 62:439–452. https://doi.org/10.1093/jxb/erq311
Said M, Cápal P, Farkas A, Gaál E, Ivanizs L, Friebe B, Doležel J, Molnár I (2022) Flow karyotyping of wheat-Aegilops additions facilitate dissecting the genomes of Ae. biuncialis and Ae. geniculata into individual chromosomes. Front Plant Sci 13:1017958. https://doi.org/10.3389/fpls.2022.1017958
Said M, Kubaláková M, Karafiátová M, Molnár I, Doležel J, Vrána J (2019) Dissecting the complex genome of crested wheatgrass by chromosome flow sorting. Plant Genome 12:0. https://doi.org/10.3835/plantgenome2018.12.0096
Sánchez-Martín J, Steuernagel B, Ghosh S, Herren G, Hurni S, Adamski N, Vrána J, Kubaláková M, Krattinger SG, Wicker T, Doležel J, Keller B, Wulff BB (2016) Rapid gene isolation in barley and wheat by mutant chromosome sequencing. Genome Biol 17:1–7. https://doi.org/10.1186/s13059-016-1082-1
Sears ER (1981) Transfer of alien genetic material to wheat. In: Evans LT, Peacock WJ (eds) Wheat science - today and tomorrow. Cambridge University Press, New York., pp 75–89
Spielmeyer W, Richards RA (2004) Comparative mapping of wheat chromosome 1AS which contains the tiller inhibition gene (tin) with rice chromosome 5S. Theor Appl Genet 109:1303–1310. https://doi.org/10.1007/s00122-004-1745-2
Tang S, Li Z, Jia X, Larkin PJ (2000) Genomic in situ hybridization (GISH) analyses of Thinopyrum intermedium, its partial amphiploid Zhong 5, and disease-resistant derivatives in wheat. Theor Appl Genet 100:344–352. https://doi.org/10.1007/s001220050045
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822. https://doi.org/10.1126/science.1183700
Tiwari VK, Wang S, Sehgal S, Vrána J, Friebe B, Kubaláková M, Chhuneja P, Doležel J, Akhunov E, Kalia B, Sabir J, Gill BS (2014) SNP Discovery for mapping alien introgressions in wheat. BMC Genomics 15:273. https://doi.org/10.1186/1471-2164-15-273
Tong K, Wu X, He L, Qiu S, Liu S, Cai L, Rao S, Chen J (2021) Genome-wide identification and expression profile of OSCA gene family members in Triticum aestivum L. Int J Mol Sci 23:469. https://doi.org/10.3390/ijms23010469
Tóth V, Láng L, Vida L, Mikó P, Rakszegi M (2022) Characterization of the protein and carbohydrate related quality traits of a large set of spelt wheat genotypes. Foods 2022(11):2061. https://doi.org/10.3390/foods11142061
Tottman DR (1987) The decimal code for the growth stages of cereals, with illustrations. Ann Appl Biol 110:441–454. https://doi.org/10.1111/j.1744-7348.1987.tb03275.x
Vrána J, Cápal P, Šimková H, Karafiátová M, Čížková J, Doležel J (2016a) Flow analysis and sorting of plant chromosomes. Curr Protoc Cytom 78:5.3.1–5.3.43. https://doi.org/10.1002/cpcy.9
Vrána J, Cápal P, Číhalíková J, Kubaláková M, Doležel J (2016b) Flow sorting plant chromosomes. Methods Mol Biol 1429:119–134. https://doi.org/10.1007/978-1-4939-3622-9_10
Wang RR-C, Larson SR, Jensen KB (2010) Analyses of Thinopyrum bessarabicum, T. elongatum, and T. junceum chromosomes using EST-SSR markers. Genome 53:1083–1089. https://doi.org/10.1139/G10-088
Wang RR-C, Larson SR, Jensen KB, Bushman BS, DeHaan LR, Wang S, Yan X (2015) Genome evolution of intermediate wheatgrass as revealed by EST-SSR markers developed from its three progenitor diploid species. Genome 58:63–70. https://doi.org/10.1139/gen-2014-0186
Wang R, Liu Y, Isham K, Zhao W, Wheeler J, Klassen N, Hu Y, Bonman JM, Chen J (2018) QTL identification and KASP marker development for productive tiller and fertile spikelet numbers in two high-yielding hard white spring wheat cultivars. Mol Breed 38:135. https://doi.org/10.1007/s11032-018-0894-y
Yang G, Boshoff WHP, Li H, Pretorius ZA, Luo Q, Li B, Li Z, Zheng Q (2021) Chromosomal composition analysis and molecular marker development for the novel Ug99-resistant wheat-Thinopyrum ponticum translocation line WTT34. Theor Appl Genet 134:1587–1599. https://doi.org/10.1007/s00122-021-03796-0
Yu G, Matny O, Champouret N et al (2022) Aegilops sharonensis genome-assisted identification of stem rust resistance gene Sr62. Nat Commun 13:1607. https://doi.org/10.1038/s41467-022-29132-8
Zeller FJ (1973) lB/lR wheat-rye chromosome substitutions and translocations. Proc. 4th Intern. Wheat Genet. Symp. 1973, Columbia, Missouri, 209—221
Zhang J, Wu M, Yu JW, Li XL, Pei WF (2016) Breeding potential of introgression lines developed from interspecific crossing between Upland cotton (Gossypium hirsutum) and Gossypium barbadense: heterosis, combining ability and genetic effects. PLoS ONE 11:e0143646. https://doi.org/10.1371/journal.pone.0143646
Acknowledgements
The authors thank Fanni Tóth, Ildikó Lakner-Könyves, Zdeňka Dubská, Romana Šperková, and Jitka Weiserová for their technical assistance.
Funding
Open access funding provided by HUN-REN Centre for Agricultural Research. This work was supported by the ERDF project “Plants as a tool for sustainable global development” (No. CZ.02.1.01/0.0/0.0/16_019/0000827), the Hungarian National Research, Development and Innovation Office (FK145848, K135057, PD145915, TKP2021-NKTA-06 and 2019-2.1.11-TÉT-2019-00074), the Marie Curie Fellowship Grant award “AEGILWHEAT” (H2020-MSCA-IF-2016-746253) and the ELIXIR-CZ project (LM2015047), a component of the international ELIXIR infrastructure that is part of the project “e-Infrastruktura CZ” (LM2018140) within the program Projects of Large Research, Development and Innovations Infrastructures. The research was also funded by the EU Horizon Europe project COUSIN (Nr. 101135314).
Author information
Authors and Affiliations
Contributions
ET, KK, IM, and MML conceived the research. ET, KK, ÉS, LI, and IM wrote the manuscript and prepared the figures and the tables. MML produced the wheat-Agropyron glael hybrids and amphiploids and multiplied the progenies in the field. ET, KK, AF, and EG carried out the FISH and GISH experiments. MS and JD performed the flow cytometric chromosome analysis and sorting. LI made the molecular marker analysis. ÉS and ÉD conducted the statistical analyses. ET, KK, MC, and PM were in charge of the field phenotyping experiments. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Türkösi, E., Szakács, É., Ivanizs, L. et al. A chromosome arm from Thinopyrum intermedium × Thinopyrum ponticum hybrid confers increased tillering and yield potential in wheat. Mol Breeding 44, 7 (2024). https://doi.org/10.1007/s11032-024-01439-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-024-01439-y