Abstract
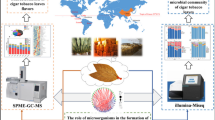
Roasted-rice leachate fermentation, a distinctive local tobacco fermentation method in Sichuan, imparts a mellow flavor and glossy texture to tobacco leaves, along with a roasted rice aroma. In order to find out the impact of roasted-rice leachate on cigar tobacco leaves, the physicochemical properties, volatile flavor profile, and microbial community were investigated. The content of protein significantly decreased after fermentation. The volatile flavor compounds increased following roasted-rice leachate fermentation, including aldehydes, alcohols, acids, and esters. High-throughput sequencing identified Staphylococcus, Pseudomonas, Pantoea, Oceanobacillus, Delftia, Corynebacterium, Sphingomonas, Aspergillus, Weissella, and Debaryomyces as the primary genera. Network and correlation analysis showed Debaryomyces played a crucial role in roasted-rice leachate fermentation, due to its numerous connections with other microbes and positive relationships with linoelaidic acid, aromandendrene, and benzaldehyde. This study is useful for gaining insight into the relationship between flavor compounds and microorganisms and provides references regarding the effect of extra nutrients on traditional fermentation products.
Key points
• Volatile flavor compounds increased following roasted-rice leachate fermentation
• Staphylococcus was the primary genera in fermented cigar
• Debaryomyces may improve the quality of tobacco leaves
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cigar tobacco is a crucial non-food economic crop around the world and has witnessed increased popularity (Delnevo et al. 2015). The cigar production process includes harvesting, curing, fermentation, and rolling. Diverging from the flue-curing process employed in cigarette tobacco, cigar tobacco undergoes air-cured (Andersen et al. 1982; Zhao et al. 2007). The temperature of air-curing is lower, causing uncomplete transformation of the chemicals in cigar. Unfermented cigar tobacco emits a pungent and unpleasant smell, rendering it unsuitable for direct use (Huang et al. 2010). Therefore, all cigar tobaccos need to go through the stage of fermentation (Reid et al. 1937; Frankenburg and Gottscho 1955). Consequently, the judicious selection of appropriate fermentation treatments holds paramount importance in enhancing the quality of cigar tobacco.
Box packing or stacking forms are generally used for the fermentation of tobacco leaves. Roasted-rice leachate fermentation (RLF) is a distinctive local tobacco fermentation method in Sichuan (Stiebritz 2012), aiming to accentuate the aroma characteristics and enhance the quality of tobacco leaves (Shilin et al. 2016). The RLF entails making cigar tobacco leaves soaked with roasted-rice leachate at the beginning of fermentation. The rice is stir-fried, partially cooked, placed in an appropriate amount of boiled water until it turns brown, and then filtered after cooling to obtain the roasted-rice leachate. The roasted-rice is full of saccharides and starch and could improve the total sugar and reduced the sugar content of cigar (Ren et al. 2023). Meanwhile, carbohydrates are key players in tobacco aroma formation and quality determination (Banožić et al. 2020). The purpose of using roasted-rice for fermenting cigar tobacco leaves is to utilize its inherent nutrients to impart a distinctive flavor. Park et al. (2022) found that roasting enhanced the content of resistant starch in rice, which is beneficial to the production of nutritional functional products. Moreover, Shi et al. (2018) found that roasting process of rice could enhance the aroma compounds like furans and pyrazines. The RLF is primarily utilized for filler cigar leaves (Shilin et al. 2016) and usually begins in March and September, lasting for a month. After RLF, cigar tobacco leaves transform into a reddish brown, glossy state with a mellow taste, resulting in a pure fried rice scorched cigar.
Chemical composition, like reducing sugar, total sugar, and nitrogen, played an important role in the flavor formation of cigar and was commonly used to assess the internal quality of tobacco (Ruan et al. 2021). The studies on the impact of roasted rice leachate on the chemical composition of tobacco were limited; previous research (Zhao 2013) suggests a reduction in chemical substances, an acceleration of their transformation, and an improvement in the quality and aroma of tobacco leaves. However, there has been insufficient research on this unique tobacco fermentation technology. Further exploration is warranted to elucidate the microbial and metabolite changes in tobacco fermentation. Microorganisms contribute significantly to the transformation of tobacco flavors during the fermentation process (Wu et al. 2022). Furthermore, roast-rice leachate was shown to suppress the growth of mold and yeast during stacking fermentation (Shilin et al. 2016), but the microbial composition was not investigated. However, the impact of roasted-rice leachate on the metabolic activity of tobacco microorganisms remains unexplained. With an increasing focus on the characteristic tobacco fermentation and the corresponding flavor-forming mechanism, RLF is gaining increasing attention by scholars. Therefore, it is worthy to systematically investigate RLF applied in cigar fermentation.
Accordingly, this work focused on cigar tobacco leaves in different stages of RLF and mainly studied the microbial community structure, succession, and their relationships with flavor compounds. These results may provide a research basis and better target for quality regulation of fermented cigars with roasted-rice leachate tobacco fermentation.
Materials and methods
Sample preparation
The tobacco samples were from the Deyang cigar No. 1 variety, provided by the Sichuan Provincial Branch of China National Tobacco Crop Tobacco Science Institute. Moderately mature cigar samples from the upper and middle sections of tobacco plants were selected for experiment to ensure consistency. Following the air-curing process, a total of 2000 kg of cigar tobacco leaves underwent stacked fermentation with roasted-rice leachate under the condition of 20 °C and 70% humidity. The preparation of roasted-rice leachate was executed as follows: rice was heated at 200 °C for 10 min, added to boiling water in a 1:3 ratio, and boiled for an additional 10 min. Subsequently, dark brown and viscous roasted-rice leachate was obtained through the processes of cooling and filtering. The roasted-rice leachate was evenly sprayed onto the surface of cigar tobacco leaves in a 30% (v/w) ratio. The entire fermentation lasted for 28 days. Whenever the temperature at the center of the tobacco stack dropped to 40 °C, the tobacco stack was turned over. Cigar samples were systematically taken out from tobacco stack at different time points (days 0, 14, 21, and 28). Detailed information about the cigar tobacco is presented in Table 1. Three biological replicates of each group (18 samples in total) were collected, and stored at − 80 °C for chemical analysis and DNA extraction.
Physicochemical properties
The total nitrogen was determined by carbon and nitrogen analyzer (Primacs SNC100, Skalar) based on the combustion method. The protein content of cigar samples was measured by protein extraction kits (G0418W, Suzhou Grace Bio-technology, Suzhou, China) based on the biuret reaction between protein and bicinchoninic acid. The starch content was measured according to GB 5009.9–2016 for starch. The contents of reducing sugars and total sugars were determined based on 3,5-dinitrosalicylic acid assay. For each sample, three independent biological replicates were collected for chemical analysis.
Volatile flavor compounds analysis
The volatile flavor compounds (VFCs) of cigar tobacco leaves were extracted by headspace solid-phase microextraction (HS-SPME) following the method based on our previous study. A total 500 mg of cigar tobacco powder was weighed in a 10 mL glass vial precisely, adding chromatographic pure standards phenethyl acetate as internal standard. Briefly, samples were equilibrated at 70 °C for 20 min, then extracted by a 50/30 µm DVB/CAR/PDMS fiber (Supelco, Bellefonte, PA, USA) at 70 °C for 30 min. Gas chromatography-mass spectrometer (GC–MS) was used to analyze the volatile flavor compounds (VFCs) of tobacco leaves. The instruments and detailed procedure of GC–MS were absolutely according to our previous literature (Liu et al. 2022c).
Microbial community analysis
The E.Z.N.A.® soil DNA Kit (Omega Bio-tek, USA) was utilized to extract microbial genomic DNA from 18 samples based on the manufacturer’s guidelines. Genomic DNA samples underwent purity and concentration assessment based on the value of A260, A230, and A280 with the instrument of NanoDrop 2000 (Thermo Scientific, USA), and integrity evaluation through 1% (w/v) agarose gel electrophoresis. The fungal gene’s ITS1F-ITS2R hypervariable region and bacterial 16S rRNA gene’s V3-V4 hypervariable region were amplified using primer pairs 338F (5'-ACTCCTACGGGAGGCAGCAG-3')/806R (5'-GGACTACHVGGGTWTCTAAT-3') and ITS1F (5'-CTTGGTCATTTAGAGGAAGTAA-3')/ITS2R (5'-GCTGCGTTCTTCATCGATGC-3') via an ABI GeneAmp 9700 PCR thermocycler (ABI, CA, USA) respectively. The PCR components contained 0.8 µL of forward primer (5 µM), 0.8 µL of reverse primer (5 µM), 10 µL of 2 × Pro Taq, 200 µg of template DNA, and 8.4 µL of ddH2O. Electrophoresis with 2% agarose gel was used to extract resulting amplicons. Then amplicons were purified with the AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, USA) according to instructions provided by the manufacturer and quantified using Quantus Fluorometer (Promega, USA). Last, the purified amplicons were pooled in equal amounts and subjected to paired-end sequencing on an Illumina MiSeq PE300 platform (Illumina, San Diego, USA), according to the established instructions by Majorbio Bio-Pharm Technology (Shanghai, China).
The demultiplexing, quality-filtering, and merging of raw sequencing data were respectively performed by fastp v0.20.0 (Chen et al. 2018) and FLASH v1.2.11 (Magoč and Salzberg 2011). Using a 97% similarity threshold (Edgar 2013), Operational taxonomic units (OTUs) were subjected to clustering with the UPARSE v11 software (Stackebrandt and Goebel 1994). This process was followed by the identification and elimination of chimeric sequences to ensure accuracy. The taxonomic classification of each OTU representative sequence was then determined using the RDP Classifier version 2.13 (Wang et al. 2007). This classification was performed by comparing the sequences against the 16S rRNA database for bacterium and ITS databases for fungus, employing a confidence threshold set at 0.7. The original sequencing dates have been made accessible and are archived in the NCBI Sequence Read Archive under the BioProject accession number PRJNA881809.
Statistical analysis
In the conducted experiments, triplicates were performed on each sample to guarantee robust repeatability, and results were depicted as the mean values with their corresponding relative standard deviations. Significant variations between the samples were investigated using one-way ANOVA with the application of Duncan’s multiple-range test. Distinctions of bacterial and fungal communities among cigar tobacco leaves during fermentation were analyzed and mapped by principal co-ordinates analysis (PCoA), based on Bray–Curtis distance matrix. The heatmaps of VFCs among samples were made by TB tools (v2.069). The correlations (Pearson) between physicochemical properties and dominant microbes were determined by R software (v4.1.2), the same as the correlations (spearman) between VFCs and dominant microorganisms and the interaction among dominant microorganisms. A p-value threshold of less than 0.05 (n = 3) was used to determine the relevance of the associations. Gephi (v0.10.1) was applied for visualizing interaction among dominant microbes.
Results
Evolution of physicochemical properties during RLF
The contents of five physicochemical properties including total nitrogen, protein, starch, reduced sugar, and total sugar are shown in Fig. 1. The content of protein decreased from the beginning to the end of RLF. The starch content in upper cigar tobacco leaves decreased after fermentation. The concentrations of both reducing sugars and total sugars generally exhibited a declining pattern. These results indicated that the fermentation with roasted-rice leachate had a significant effect on physicochemical properties of cigar tobacco. Besides, protein content in middle leaves was higher than that in upper leaves.
Evolution of volatile flavor compounds during RLF
SPME–GC–MS was used to extract and determine VFCs of cigar tobacco leaves among different fermentation stages. A total of 28 representative VFCs were selected to analyze their distribution in each group, including 6 ketones, 5 esters, 4 aldehydes, 4 pyrazines, 3 alkenes, 2 acids, 2 alcohols, 1 alkane, and 1 furan (Fig. 2 and Table S1). The composition of VFCs in unfermented cigar tobacco leaves (Raw-up and Raw-mid) were similar, while ethyl nonanoate and 1-nonene in Raw-up were significantly higher than that in Raw-mid. Indeed, the RLF significantly transformed the VFCs profile of cigar tobacco. For example, solanone, phytofuran, and phytol increased after fermentation, while lots of compounds such as benzeneacetaldehyde, hexahydrofarnesol, 2,6,10-trimethyltridecane, and hexahydropseudoionone were decreased after fermentation. Compared to middle cigar after fermentation, upper cigar has lower content of myosmine, ethyl palmitate, ethyl tetradecanoate, and L-nicotine, but higher content of aromandendrene, linoeladidc acid, and benzaldehyde. Notably, day 21 may be a crucial fermentation time points of middle cigar tobacco leaves, because hydroxydehydrostevic acid, citral, octyl formate, nicotine 1-N-oxide, and anabasine occurred in F21-mid and were not detected in F28-mid.
Overview of microbial community
A total of 1,116,718 and 1,661,297 raw reads were obtained, with an average of 60,847 and 62,039 high-quality bacterial and fungal sequences per sample, respectively. The information of sequence data and alpha-diversity indices among samples was summarized in Table 2. With the increase of sequencing times, the sparse curve of microbial community tended to be gentle, and the coverage of all samples exceeded 97.9% (as shown in Figure S1), indicating that the sequencing depth of this work was sufficient. Both the Chao index and Shannon index of microbial community in cigar tobacco leaves decreased after fermentation (Table 2). This trend suggests that the fermentation involving roasted-rice leachate could reduce microbial richness and diversity in cigar tobacco leaves.
Comparison of microbial profiles
The microbial taxonomic composition of tobacco leaves at different fermentation stages was studied at the phylum and genus levels. The microbial community composition is described by the histogram in Fig. 3, which clearly revealed the relative abundance of each species and the highest abundance between the groups (details in Table S2). At the phylum level, the bacterial community of middle and upper cigar tobacco leaves was similar (Fig. 3). Significant differences in dominant phylum before and after fermentation. Proteobacteria was the most dominant phyla in unfermented cigar tobacco leaves with an abundance of more than 95%, which converted into Firmicutes and Actinobacteria after the beginning of fermentation. Ascomycota was the most predominant phylum in the fungal community (Fig. 3), approximately accounted for 45 ~ 85% of fungi phyla.
At the genus level, the top 19 genera of bacterial flora and the top 9 genera of fungi were selected from all samples, and less than 0.01% were labeled as “others” (Fig. 3). Staphylococcus, Pseudomonas, Pantoea, Oceanobacillus, Delftia, Corynebacterium, Sphingomonas, Aspergillus, Weissella, and Debaryomyces were primary genera in RLF cigar, detected in all groups. After fermentation, the changes of microbial community genus level reflected in dominant bacterial genus from Pantoea, Pseudomonas, and Delftia to Staphylococcus, Oceanobacillus, and Corynebacterium; dominant fungal genus from Aspergillus and Weissella to Aspergillus, Weissella, and Debaryomyces. Corynebacterium, Oceanobacillus, Aerococcus, and Lactobacillus were the characteristic bacteria genus after fermentation; Debaryomyces, Cutaneotrichosporon, and Trichosporon were the characteristic fungal genus.
Dimension reduction analysis of microbial community
The distinction of relative abundance and diversity of microbial communities was analyzed by PCoA on a two-dimensional plot, based on the Bray–Curtis distance matrix. The variation of bacterial community among samples was 73.97% (PC1 variance) and 9.06% (PC2 variance), demonstrating a pronounced regional separation (Fig. 4). The distinction in bacterial community PCoA was statistically significant (p = 0.001). The unfermented samples in the middle and upper leaves (Raw-up and Raw-mid) were distinctly separated from each other. The fermented samples (F28-up, F14-mid, F21-mid, and F28-mid) exhibited close clustering, suggesting a similar bacterial composition in the samples after fermentation. In addition, there was a clear separation between unfermented and fermented samples. The variation of fungal community was 63.67% (PC1 variance) and 23.44% (PC2 variance), with a statistically significant difference among groups (p = 0.001) as depicted in Fig. 4. The unfermented samples in the middle and upper leaves were distinctly separated from each other as well. While there was no pronounced clustering found in the fermented samples.
The significant differences in microflora among roasted-rice leachate fermented tobaccos were performed by Linear discriminant analysis Effect Size (LEfSe), with a threshold for linear discriminant analysis score set to a minimum of 4 (Kruskal–Wallis test, p < 0.05). This analysis served to validate the results obtained from PCoA and to discern key groups during various fermentation stages, specifically, the identification of biomarkers. In Fig. 5, a total of 32 branches were observed, including 3 phyla, 4 classes, 8 orders, 9 families, and 8 genera. For the unfermented cigar samples (Raw-up and Raw-mid), distinctive microorganisms at the genus level included Pantoea, Pseudomonadales, Methylobacterium-Methylorubrum, and norank_f_Mitochondria. The biomarkers of day 21 (F21-mid) included Lactobacillales and Aerococcus, while the biomarkers of day 28 (F28-up and F28-mid) were Staphyloccus, Corynebacterium, and Microccaceae. No biomarker was found for the day 14 (F14-mid) of RLF. LEfSe analysis of fungal community (Fig. 5) shows that Altemaria, Aspergillus, Bullera, Epicoccum, and Golubevia were biomarkers of the unfermented upper cigar leaves (Raw-up). As for the fermented cigars, Cutaneotrichosporon, g_unclassified_f_Nectriacea, and Debaryomyces were the biomarkers of day 14 (F14-mid), 21 (F21-mid), and 28 (F28-up), respectively. No biomarkers were found in Raw-mid and F28-mid.
Interaction network of microbial communities
The co-occurrence and co-exclusion patterns of microbial (both bacterial and fungal) communities were investigated to clarify the interaction within microbial communities in the characteristic fermentation of the cigar tobacco leaves, using the Spearman rank correlation (| ρ |> 0.5, p < 0.05). The interaction networks of representative bacterial taxa consisted of 28 nodes (genera) and 285 edges (Fig. 6). Most of genera in the phylum of Proteobacteria exhibited positive correlations with the other bacteria. Staphylococcus, belonging to the phylum of Firmicutes, has the highest abundance and generally displayed negative correlations with the other bacteria. Within the phylum of Acinobacteriota, Corynebacterium represented mainly negative correlation with the bacteria, while Curtobacterium represented mainly positive correlation with the bacteria. Additionally, the bacteria were negatively correlated with the fungi within the roasted-rice fermented cigar samples, as shown in Fig. 6. For instance, Debaryomyces had the most connections over bacterial and fungal communities and was negatively correlated with the majority of the bacterial genera.
Co-occurrence and co-exclusion network of microbial communities in tobacco. a Interaction among dominant bacterial communities. b Interaction network of dominant bacterial and fungal communities. Edge colors denote positive (red) or negative (green) correlations, with edge thicknesses scaled to the absolute Spearman correlation coefficients. The nodes were proportionally sized by the amounts of connections, and colored by different phylum groups
Correlation analysis
The correlation between 5 physicochemical properties and top 15 bacteria and top 15 fungi was analyzed by the Pearson correlation analysis and displayed on the heatmap (Fig. 7). The results in Fig. 7 show that Atopostipes was significantly associated with total sugar; Pantoea, Pectobacterium, Atopostipes, and Corynebacterium were significantly associated with protein and reduced sugar; Pseudomonas was significantly associated with total nitrogen. In fungus (Fig. 7), Trichosporon, unclassified_f_Aspergillaceae, and unclassified_o_Saccharomycetales were significantly associated with total nitrogen; Cladosporium, Alternaria, and Stemphylium were significantly associated with reduced sugar.
The correlation between microbial and physicochemical indexes in cigar tobacco leaves (*p < 0.05; **p < 0.01; ***p < 0.001). a Correlation of bacteria and physicochemical factors. b Correlation of fungi and physicochemical factors. TS, total sugar; P, protein; RS, reduce sugar; TN, total nitrogen; S, starch
The Spearman correlation analysis of metabolite absolute abundance and microbial relative abundance delineated distinct metabolite-microbe associations (Fig. 8). Most significantly, myosmine exhibited a significant negative correlation with Methylobacterium-Methylorubrum and Pseudomonas; benzeneacetaldehyde exhibited a significant negative correlation with Delftia; neophytadiene was positively correlated with Sphingomonas (p < 0.001). Ester compounds were mainly associated with Stagonosporopsis and g_norank_f_Mitochondria. Ketones were positively associated with Pseudomonas, Methylobacterium-Methylorubrum, and Sphingomonas. There was no significant correlation (p < 0.05) observed between acid compounds and microorganism.
Discussion
Effect of RLF on cigar substance profile
Roasted-rice leachate can provide nutrients for tobacco fermentation thus accelerate the chemical reaction in the fermentation process to improve the fermentation quality (Onmankhong et al. 2021; Park et al. 2022). Indeed, compared to conventional cigar fermentation, this work has evidenced certain specific transition of physicochemical properties of cigar tobacco leaves during RLF. The RLF process primarily manifests changes in physicochemical properties, notably marked by a reduction in protein, starch, total sugar, and reducing sugar content. Concurrently, the total nitrogen content remains relatively constant. Protein and starch are macromolecular substances in the tobacco (Banožić et al. 2020). High content of protein or starch is negative to the grade and smoking characteristics of tobacco leaves (Vansuyt et al. 2003; Zhang et al. 2005). Properly reducing the content of protein and starch in the fermentation process is conducive to reducing the offensive odor of tobacco leaves and ensuring mellow aroma of tobacco (Dai et al. 2020). Interestingly, protein content in the upper leaves was higher than that in the middle leaves, and remained almost the same after fermentation, suggesting that more aroma could be produced in the upper leaves. Otherwise, the decrease of total sugar and reducing sugar might result from the growth of microorganism and Maillard reaction.
The VFC profile of roasted-rice fermented tobacco leaves exhibited expected distinctions from the unfermented counterpart. Briefly, aldehydes, alcohols, acids, and esters increased following RLF, while hydrocarbon decreased. Studies have documented that aldehydes and ketones, heterocyclic compounds, and esters significantly contributed to the aroma, such as benzaldehyde (Samira Jandoust 2014), furan (Yuan et al. 2023), and pyridine (Deblander et al. 2015). The hydrocarbon content gradually declined during RLF. Alkanes, resulting from the oxidation and degradation of fatty acids, contribute to aroma and sweetness (Gamou and Kawashima 1980; Maggi et al. 2010). However, high odor thresholds limited their impact on the overall flavor. Concurrently, previous works have suggested that hydrocarbons were the important intermediates in the formation of heterocyclic compounds; for instance, alkenes can form aldehydes and ketones under certain conditions (Wang et al. 2021; Chang et al. 2022; Bian et al. 2022). In this work, aldehydes were generated at the middle stage of RLF, suggesting that the reduction in hydrocarbons may be attributed to aldehyde formation. Aldehydes, byproducts of lipid oxidation in the fermentation process (Qin et al. 2013), play a crucial role in enhancing the flavor of cigars (Zheng et al. 2022). They typically impart a pleasant aroma reminiscent of smell similar to grass, malt, fruit, and cheese (Wu et al. 2013; Wang et al. 2018). The formation of pyridine is another consequence of the lipid oxidation pathway, as the aldehydes and ketones required for pyridine formation are generated by lipid oxidation (Zamora et al. 2020). Pyridine was detected on day 21 during RLF, while the content of aldehydes and ketones reached the peak on day 14 and subsequently declined on day 21. The increase of alcohols and acids contributes to the generation of esters. Esters also contribute to the flavor of cigar tobacco. Ethyl palmitate was produced during fermentation, which is a flavor substance with a waxy, fruity, creamy, milky white color, and a light creamy flavor (Tabaszewska et al. 2022). In summary, the flavor of cigar was enhanced after RLF.
Effect of fermentation on microbe, correlation between microorganisms and volatile substances in RLF
The fermentation of cigar tobacco leaves primarily results from the concerted action of microorganisms and a variety of complex enzyme systems, ultimately leading to transformation of the chemical composition and flavor components (Wang et al. 2011). Roasted-rice leachate provides a distinctive environment conducive to cigar fermentation. Although a substantial amount of organic matter in rice undergoes carbonization at high temperatures during the parched rice-making process, the retained carbon, nitrogen, and inorganic salts provide additional nutrients for microbial growth (Wang et al. 2022b; Yuan et al. 2022). The microbial communities in RLF process exhibited significant variation between unfermented and fermented cigars. The transition manifested in the dominance shift of bacteria, transitioning from Pseudomonas and Pantoea to Staphylococcus and Corynebacterium. Simultaneously, the dominant fungi shifted from Aspergillus and Wallemia to Aspergillus, Wallemia, and Debaryomyces. Staphylococcus, Pseudomonas, Pantoea, Corynebacterium, Aspergillus, and Weissella were widely reported as the dominant microorganisms of fermented tobacco leaves. Generally, the succession of bacterial community was more intensive than that of fungal community during RLF. Despite the decreasing richness of the microbial community during RLF, it is essential to note that high microbial richness does not necessarily lead to better quality (Wang et al. 2022a).
Microorganisms give rise to diverse flavor compounds during fermentation. Corynebacterium, Oceanobacillus, Aerococcus, Lactobacillus, and Debaryomyces were the characteristic genus after RLF, which has been proven to be beneficial bacteria and significantly improved of ureolysis pathway, nitrate reduction pathway, and fermentation pathway (Di Giacomo et al. 2007; Sami et al. 2021; Mardawati et al. 2022). Corynebacterium and Oceanobacillus exhibit alkaline tolerance, potentially accounting for their predominance in the fermentation process (Hirota et al. 2013; Pang et al. 2020). Additionally, Corynebacterium (Mhatre et al. 2022; Gao et al. 2022b; Liu et al. 2022b), Oceanobacillus (Ning et al. 2023), and Staphylococcus (Jia et al. 2021; Dias et al. 2022; Li et al. 2023) have been demonstrated to exert beneficial impacts on the quality of fermented products. However, Pantoea, Defltia, and Pectobacterium in the early stages of RLF may produce a limited contribution to fermentation (Bajpai et al. 2012; Li et al. 2020a) and induce the blight to plant leaves (Gao et al. 2022a; Lao et al. 2022).
Pseudomonas and Sphingomonas have been proven to play crucial roles in tobacco fermentation (Li et al. 2020b). In the RLF, Pseudomonas and Sphingomonas were related with total nitrogen, protein, and reduce sugar, and had positive correlation with nonanal, hexahydropseudoionone, neophytadiene, and megastigmatrienone. Neophytadiene was able to help certain flavor substance transfer into the flue gas and reduce irritation of tobacco (Liu et al. 2022a). Interestingly, the abundance of Pseudomonas was increased and the abundance of Sphingomonas was reduced during the conventional fermentation (Li et al. 2020b), but both of Pseudomonas and Sphingomonas were decreased in the RLF. Acinetobacter, Sphingomonas, and Aspergillus are presumed to be important contributors to tobacco volatiles (Wu et al. 2022). Fungus, such as Aspergillus and Debaryomyces, also contribute to the flavor. As one of the major yeasts in dairy products (Haastrup et al. 2018), Debaryomyces could form pseudohyphae and reproduce by multilateral budding, which is considered to have the potential for aroma enhancement (Shruthi et al. 2022). In this study, the abundance of Debaryomyces increased after the onset of the fermentation and exhibited a positive correlation with Corynebacterium, Staphylococcus, and Weissella (Fig. 6). Additionally, Debaryomyces showed a positive relationship with linoelaidic acid, aromandendrene, and benzaldehyde. Therefore, Debaryomyces played a crucial role in RLF as a fungal genus. The correlation between microorganisms and volatile components obtained in this work also coincides with previous studies, but there are also new findings unique to roasted-rice fermentation. All discoveries of these association pairs can be used for directional enhanced cigar fermentation.
The present study investigated the physicochemical properties, volatile flavor compounds, and microbial community of cigars during RLF. The content of protein, reside sugar and total sugar decreased after fermentation. The flavor compounds, such as aldehydes, alcohols, acids, and esters, increased following RLF. Staphylococcus, Pseudomonas, Pantoea, Oceanobacillus, Delftia, Corynebacterium, Sphingomonas, Aspergillus, Weissella, and Debaryomyces were the primary genera. Debaryomyces played a crucial role in RLF, due to its numerous connections with other microbes and positive relationships with linoelaidic acid, aromandendrene, and benzaldehyde. These results contribute to the understanding of chemical and microbial transition profile and mechanism during RLF, providing valuable insights for the further application of characteristic fermentation technology.
Data availability
The datasets generated for this study can be found in the NCBI Sequence Read Archive (SRA, https://www.ncbi.nlm.nih.gov/sra/PRJNA881809) database under the BioProject accession number PRJNA881809.
References
Akbar K, Samira J, Esmaeil E (2014) Bioinformatics analysis of phenylacetaldehyde synthase (PAAS), a protein involved in flower scent production, in rose. J Phys Chem Biophys 5:1000170. https://doi.org/10.4172/2161-0398.1000170
Andersen RA, Kasperbauer MJ, Burton HR, Hamilton JL, Yoder EE (1982) Changes in chemical composition of homogenized leaf-cured and air-cured burley tobacco stored in controlled environments. J Agric Food Chem 30:663–668. https://doi.org/10.1021/jf00112a010
Bajpai VK, Kang SC, Baek KH (2012) Microbial fermentation of cabbage by a bacterial strain of Pectobacterium atrosepticum for the production of bioactive material against Candida species. J Mycol Méd 22:21–29. https://doi.org/10.1016/j.mycmed.2011.11.004
Banožić M, Jokić S, Ačkar Đ, Blažić M, Šubarić D (2020) Carbohydrates—key players in tobacco aroma formation and quality determination. Molecules 25:1734. https://doi.org/10.3390/molecules25071734
Bian X, Chen J, Fan J, Yang Y, Yu D, Ren L, Ma Z, Wu N, Chen F, Liu X, Wang B, Zhang N (2022) Microbial and genes diversity analysis: relationship between starch conversion and carbohydrate metabolism during Niandoubao fermentation via the glutinous proso millet (GPM) process. Food Control 140:109154. https://doi.org/10.1016/j.foodcont.2022.109154
Chang Y, Xie Y, Zhao C, Ren J, Su W, Zhao W, Wu L, Yu H (2022) Selective oxidation of olefins to aldehydes/ ketones/ carboxylic acids with an efficient and practical polyoxometalate-based iron catalyst. ChemCatChem 14:e202200209. https://doi.org/10.1002/cctc.202200209
Chen S, Zhou Y, Chen Y, Jia G (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Dai J, Dong A, Xiong G, Liu Y, Hossain MdS, Liu S, Gao N, Li S, Wang J, Qiu D (2020) Production of highly active extracellular amylase and cellulase from Bacillus subtilis ZIM3 and a recombinant strain with a potential application in tobacco fermentation. Front Microbiol 11:1539. https://doi.org/10.3389/fmicb.2020.01539
Deblander J, Van Aeken S, Adams A, De Kimpe N, Abbaspour Tehrani K (2015) New short and general synthesis of three key Maillard flavour compounds: 2-Acetyl-1-pyrroline, 6-acetyl-1,2,3,4-tetrahydropyridine and 5-acetyl-2,3-dihydro-4H-1,4-thiazine. Food Chem 168:327–331. https://doi.org/10.1016/j.foodchem.2014.07.088
Delnevo CD, Giovenco DP, Ambrose BK, Corey CG, Conway KP (2015) Preference for flavoured cigar brands among youth, young adults and adults in the USA. Tob Control 24:389–394. https://doi.org/10.1136/tobaccocontrol-2013-051408
Di Giacomo M, Paolino M, Silvestro D, Vigliotta G, Imperi F, Visca P, Alifano P, Parente D (2007) Microbial community structure and dynamics of dark fire-cured tobacco fermentation. Appl Environ Microbiol 73:825–837. https://doi.org/10.1128/AEM.02378-06
Dias I, Laranjo M, Potes ME, Agulheiro-Santos AC, Ricardo-Rodrigues S, Fraqueza MJ, Oliveira M, Elias M (2022) Staphylococcus spp. and Lactobacillus sakei starters with high level of inoculation and an extended fermentation step improve safety of fermented sausages. Fermentation 8:49. https://doi.org/10.3390/fermentation8020049
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996. https://doi.org/10.1038/nmeth.2604
Frankenburg WG, Gottscho AM (1955) The chemistry of tobacco fermentation. I. Conversion of the Alkaloids. B. The Formation of Oxynicotine. J Am Chem Soc 77:5728–5730. https://doi.org/10.1021/ja01626a076
Gamou K, Kawashima N (1980) Studies on leaf surface lipid of tobacco III. Compositional change in alkane during growth and senescence of tobacco leaves. Agric Biol Chem 44:2119–2124. https://doi.org/10.1080/00021369.1980.10864278
Gao L, Li W, Ren B, Hu Y, Ma X, Jia Y, Wang Z (2022a) First report of leaf blight caused by Pantoea agglomerans on wheat in China. Plant Dis. https://doi.org/10.1094/PDIS-07-22-1606-PDN. PDIS-07-22-1606-PDN
Gao R, Sun-Waterhouse D, Xiang H, Cui C, Waterhouse GIN (2022b) The effect of the Corynebacterium glutamicum on the shortening of fermentation time, physicochemical and sensory properties of soy sauce. Int J Food Sci Technol 57:4316–4327. https://doi.org/10.1111/ijfs.15758
Haastrup MK, Johansen P, Malskær AH, Castro-Mejía JL, Kot W, Krych L, Arneborg N, Jespersen L (2018) Cheese brines from Danish dairies reveal a complex microbiota comprising several halotolerant bacteria and yeasts. Int J Food Microbiol 285:173–187. https://doi.org/10.1016/j.ijfoodmicro.2018.08.015
Hirota K, Hanaoka Y, Nodasaka Y, Yumoto I (2013) Oceanobacillus polygoni sp. nov., a facultatively alkaliphile isolated from indigo fermentation fluid. Int J Syst Evol Microbiol 63:3307–3312. https://doi.org/10.1099/ijs.0.048595-0
Huang J, Yang J, Duan Y, Gu W, Gong X, Zhe W, Su C, Zhang K-Q (2010) Bacterial diversities on unaged and aging flue-cured tobacco leaves estimated by 16S rRNA sequence analysis. Appl Microbiol Biotechnol 88:553–562. https://doi.org/10.1007/s00253-010-2763-4
Jia Y, Niu C-T, Xu X, Zheng F-Y, Liu C-F, Wang J-J, Lu Z-M, Xu Z-H, Li Q (2021) Metabolic potential of microbial community and distribution mechanism of Staphylococcus species during broad bean paste fermentation. Food Res Int 148:110533. https://doi.org/10.1016/j.foodres.2021.110533
Lao G, Jin P, Miao W, Liu W (2022) First report of leaf spot on cucumber caused by Pantoea ananatis in Hainan of China. Plant Dis. https://doi.org/10.1094/PDIS-04-22-0819-PDN. PDIS-04-22-0819-PDN
Li H, Liu H, Zeng Q, Xu M, Li Y, Wang W, Zhong Y (2020a) Isolation and appraisal of a non-fermentative bacterium, Delftia tsuruhatensis, as denitrifying phosphate-accumulating organism and optimal growth conditions. J Water Process Eng 36:101296. https://doi.org/10.1016/j.jwpe.2020.101296
Li J, Zhao Y, Qin Y, Shi H (2020b) Influence of microbiota and metabolites on the quality of tobacco during fermentation. BMC Microbiol 20:356. https://doi.org/10.1186/s12866-020-02035-8
Li Y, Cao Z, Yu Z, Zhu Y, Zhao K (2023) Effect of inoculating mixed starter cultures of Lactobacillus and Staphylococcus on bacterial communities and volatile flavor in fermented sausages. Food Sci Hum Wellness 12:200–211. https://doi.org/10.1016/j.fshw.2022.07.010
Liu A, Yuan K, Li Q, Liu S, Li Y, Tao M, Xu H, Tian J, Guan S, Zhu W (2022a) Metabolomics and proteomics revealed the synthesis difference of aroma precursors in tobacco leaves at various growth stages. Plant Physiol Biochem 192:308–319. https://doi.org/10.1016/j.plaphy.2022.10.016
Liu J, Xu J-Z, Rao Z-M, Zhang W-G (2022b) Industrial production of L-lysine in Corynebacterium glutamicum: progress and prospects. Microbiol Res 262:127101. https://doi.org/10.1016/j.micres.2022.127101
Liu T, Guo S, Wu C, Zhang R, Zhong Q, Shi H, Zhou R, Qin Y, Jin Y (2022c) Phyllosphere microbial community of cigar tobacco and its corresponding metabolites. Front Microbiol 13:1025881. https://doi.org/10.3389/fmicb.2022.1025881
Maggi F, Papa F, Cristalli G, Sagratini G, Vittori S (2010) Characterisation of the mushroom-like flavour of Melittis melissophyllum L. subsp. melissophyllum by headspace solid-phase microextraction (HS-SPME) coupled with gas chromatography (GC–FID) and gas chromatography–mass spectrometry (GC–MS). Food Chem 123:983–992. https://doi.org/10.1016/j.foodchem.2010.05.049
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics. https://doi.org/10.1093/bioinformatics/btr507
Mardawati E, Febrianti EA, Fitriana HN, Yuliana T, Putriana NA, Suhartini S, Kasbawati (2022) An integrated process for the xylitol and ethanol production from oil palm empty fruit bunch (OPEFB) using Debaryomyces hansenii and Saccharomyces cerevisiae. Microorganisms 10:2036. https://doi.org/10.3390/microorganisms10102036
Mhatre A, Shinde S, Jha AK, Rodriguez A, Wardak Z, Jansen A, Gladden JM, George A, Davis RW, Varman AM (2022) Corynebacterium glutamicum as an efficient omnivorous microbial host for the bioconversion of lignocellulosic biomass. Front Bioeng Biotechnol 10:827386. https://doi.org/10.3389/fbioe.2022.827386
Ning Y, Zhang L-Y, Mai J, Su J-E, Cai J-Y, Chen Y, Jiang Y-L, Zhu M-J, Hu B-B (2023) Tobacco microbial screening and application in improving the quality of tobacco in different physical states. Bioresour Bioprocess 10:32. https://doi.org/10.1186/s40643-023-00651-6
Onmankhong J, Jongyingcharoen JS, Sirisomboon P (2021) The influence of processing parameters of parboiled rice on its physiochemical and texture properties. J Texture Stud 52:219–227. https://doi.org/10.1111/jtxs.12576
Pang L, He Y, Liu X, Li J, Yang P (2020) The role of a newly isolated strain Corynebacterium pollutisoli SPH6 in waste activated sludge alkaline fermentation. Chemosphere 241:125072. https://doi.org/10.1016/j.chemosphere.2019.125072
Park J, Oh S-K, Chung H-J, Shin DS, Choi I, Park H-J (2022) Effect of steaming and roasting on the quality and resistant starch of brown rice flour with high amylose content. LWT 167:113801. https://doi.org/10.1016/j.lwt.2022.113801
Qin Z, Pang X, Chen D, Cheng H, Hu X, Wu J (2013) Evaluation of Chinese tea by the electronic nose and gas chromatography–mass spectrometry: correlation with sensory properties and classification according to grade level. Food Res Int 53:864–874. https://doi.org/10.1016/j.foodres.2013.02.005
Reid JJ, McKinstry DW, Haley DE (1937) The fermentation of cigar-leaf tobacco. Science 86:404–404. https://doi.org/10.1126/science.86.2235.404
Ren M, Qin Y, Zhang L, Zhao Y, Zhang R, Shi H (2023) Effects of fermentation chamber temperature on microbes and quality of cigar wrapper tobacco leaves. Appl Microbiol Biotechnol 107:6469–6485. https://doi.org/10.1007/s00253-023-12750-7
Ruan Y, Tang Z, Chen Z, Xia T (2021) Effects of combined application of chemical fertilizer and microbial fertilizer on the chemical components contents of flue-cured tobacco leaves. IOP Conf Ser Earth Environ Sci 792:012048. https://doi.org/10.1088/1755-1315/792/1/012048
Sami A, Elimairi I, Patangia D, Watkins C, Ryan CA, Ross RP, Stanton C (2021) The ultra-structural, metabolomic and metagenomic characterisation of the sudanese smokeless tobacco ‘Toombak.’ Toxicol Rep 8:1498–1512. https://doi.org/10.1016/j.toxrep.2021.07.008
Shi Y, Wang L, Fang Y, Wang H, Tao H, Pei F, Li P, Xu B, Hu Q (2018) A comprehensive analysis of aroma compounds and microstructure changes in brown rice during roasting process. LWT 98:613–621. https://doi.org/10.1016/j.lwt.2018.09.018
Shilin LI, Wang Y, Zhang B, Tang Z, Ganrong XU (2016) Effect of roasted-rice leachate on tabacco microorganisms during fermentation process. Chin J Bioprocess Eng 2:41–46. https://doi.org/10.3969/j.issn.1672-3678.2016.02.008
Shruthi B, Deepa N, Somashekaraiah R, Adithi G, Divyashree S, Sreenivasa MY (2022) Exploring biotechnological and functional characteristics of probiotic yeasts: a review. Biotechnol Rep 34:e00716. https://doi.org/10.1016/j.btre.2022.e00716
Stackebrandt E, Goebel BM (1994) Taxonomic Note: A Place for DNA-DNA Reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Stiebritz C (2012) Going global with Chinese cigars. Tob J Int 2012:96–98
Tabaszewska M, Najgebauer-Lejko D, Zbylut-Górska M (2022) The effect of crataegus fruit pre-treatment and preservation methods on the extractability of aroma compounds during liqueur production. Molecules 27:1516. https://doi.org/10.3390/molecules27051516
Vansuyt G, Souche G, Straczek A, Briat J-F, Jaillard B (2003) Flux of protons released by wild type and ferritin over-expressor tobacco plants : effect of phosphorus and iron nutrition. Plant Physiol Biochem 41:27–33. https://doi.org/10.1016/S0981-9428(02)00005-0
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Nave Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261. https://doi.org/10.1128/AEM.00062-07
Wang J, Qiu Y, Liu J (2011) Study on physical and chemical properties of domestic and imported paper-process reconstituted tobacco. Adv Mater Res 356–360:1894–1899. https://doi.org/10.4028/www.scientific.net/AMR.356-360.1894
Wang C, Liu W, Wei Y, Song H (2018) Dynamic changes of the total content of glycoside aroma components in tobacco leaves in different producing areas during the late growth period. Sci Publ Group. https://doi.org/10.11648/J.JPS.20180605.12
Wang X, Li Y, Li Z (2021) Thiol-initiated photocatalytic oxidative cleavage of the C=C bond in olefins and its extension to direct production of acetals from olefins. Catal Sci Technol 11:1000–1006. https://doi.org/10.1039/D0CY01963A
Wang Q, Wang C, Xiang X, Xu H, Han G (2022a) Analysis of microbial diversity and succession during Xiaoqu Baijiu fermentation using high-throughput sequencing technology. Eng Life Sci 22:495–504. https://doi.org/10.1002/elsc.202200015
Wang Z, Zhang Y, Bo G, Zhang Y, Chen Y, Shen M, Zhang P, Li G, Zhou J, Li Z, Yang J (2022b) Ralstonia solanacearum infection disturbed the microbiome structure throughout the whole tobacco crop niche as well as the nitrogen metabolism in soil. Front Bioeng Biotechnol 10:903555. https://doi.org/10.3389/fbioe.2022.903555
Wu L, He Z, Wu Y, Liu J, Li C, Cao J, Wang B, Min S (2013) Evaluation of aroma components in Chinese Southwest tobacco by headspace gas chromatography-mass spectrometry. Asian J Chem 25:8853–8858. https://doi.org/10.14233/ajchem.2013.14698
Wu X, Cai W, Zhu P, Peng Z, Zheng T, Li D, Li J, Zhou G, Du G, Zhang J (2022) Profiling the role of microorganisms in quality improvement of the aged flue-cured tobacco. BMC Microbiol 22:197. https://doi.org/10.1186/s12866-022-02597-9
Yuan Z, Liu Q, Pang Z, Fallah N, Liu Y, Hu C, Lin W (2022) Sugarcane rhizosphere bacteria community migration correlates with growth stages and soil nutrient. Int J Mol Sci 23:10303. https://doi.org/10.3390/ijms231810303
Yuan X, Zhou J, Zhang B, Shen C, Yu L, Gong C, Xu Y, Tang K (2023) Identification, quantitation and organoleptic contributions of furan compounds in brandy. Food Chem 412:135543. https://doi.org/10.1016/j.foodchem.2023.135543
Zamora R, Lavado-Tena CM, Hidalgo FJ (2020) Oligomerization of reactive carbonyls in the presence of ammonia-producing compounds: a route for the production of pyridines in foods. Food Chem 304:125284. https://doi.org/10.1016/j.foodchem.2019.125284
Zhang C, Lillie R, Cotter J, Vaughan D (2005) Lysozyme purification from tobacco extract by polyelectrolyte precipitation. J Chromatogr A 1069:107–112. https://doi.org/10.1016/j.chroma.2004.10.018
Zhao M, Wang B, Li F, Qiu L, Li F, Wang S, Cui J (2007) Analysis of bacterial communities on aging flue-cured tobacco leaves by 16S rDNA PCR–DGGE technology. Appl Microbiol Biotechnol 73:1435–1440. https://doi.org/10.1007/s00253-006-0625-x
Zhao Y (2013) Study on chemical composition of fermented sun-cured tobacco soaked in extract of cooking rice. Dissertation, Sichuan Agricultural University
Zheng T, Zhang Q, Peng Z, Li D, Wu X, Liu Y, Li P, Zhang J, Du G (2022) Metabolite-based cell sorting workflow for identifying microbes producing carbonyls in tobacco leaves. Appl Microbiol Biotechnol 106:4199–4209. https://doi.org/10.1007/s00253-022-11982-3
Acknowledgements
Thanks to the partner units and Sichuan University for providing tobacco raw materials and platform support; thanks to all authors for the contribution of the manuscript.
Funding
This work was financially supported by the Key Research Project of Sichuan Provincial Branch of China National Tobacco Crop Tobacco Science Institute (SCYC202124).
Author information
Authors and Affiliations
Contributions
XF: investigation, formal analysis, writing—original draft; YQ: conceptualization, supervision; LT: date curation; SG: resources, project administration; CW: investigation, methodology; RZ: resources, investigation; QZ: resources, investigation; KL: resources, investigation HS: project administration, conceptualization; RZ: methodology; SZ: resources, writing—review and editing; YJ: conceptualization, writing—review and editing, supervision; all authors have read the article and agreed to the submitted version.
Corresponding author
Ethics declarations
Ethics approval
The article does not contain any studies with human participants or animals.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fang, X., Qin, Y., Liu, T. et al. Roles of cigar microbes in flavor formation during roasted-rice leachate fermentation. Appl Microbiol Biotechnol 108, 457 (2024). https://doi.org/10.1007/s00253-024-13289-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13289-x