Abstract
Changing climatic conditions and the emphasis on the cultivation of genetically stable and resilient varieties as well as efficiently managing water and mineral resources require the commencement of appropriate research already at the stage of plant breeding. For this purpose, breeders must have the necessary tools not only in the form of an experimental network, but also statistical tools that enable the correct interpretation of the obtained results. In the presented research, the additive main effects and multiplicative interaction (AMMI) model, supplemented with cluster analysis, was used to assess the stability and yielding level of 26 spring wheat genotypes, in six locations. The main reason for the yield variability in studied genotypes was environmental factor (89%). In spite of differential conditions in the experimental network locations, the studied environments, which had a similar effect on the genotypes, in the growing season of 2021, were grouped. The AMMI stability value (ASV), yield stability index (YSI) and genotype selection index (GSI) coefficients were used to evaluate the studied genotypes. Based on the analyses, the following genotypes were selected for further breeding work: STH 21-03, STH 21-09 and KOH 18279, as stable and widely adapted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common wheat (Triticum aestivum) is one of the key species whose yield-building potential and stability are largely taken into account in the global strategy of feeding the population (Grote et al. 2021). The growth and development of cultivated plants, and in particular the yield potential, are influenced by multiple abiotic and biotic factors, as well as human activities (cultivation and fertilization). These aspects, along with genetic conditions, affect the shape and size of the genotype–environmental interaction (GEI) and thus the yield stability of a variety. This consistency in the response to a complex set of factors, combined with the ability to maintain a relatively high mean for yield, is a prerequisite for broad genotype adaptation (Mądry et al. 2006b). Varieties with higher adaptability are preferred in agricultural practice. It is also a condition for registration by Research Centre for Cultivar Testing (COBORU) in Poland, because the new variety should have a satisfactory economic value, which in practice means an appropriate level of yielding throughout the country.
Extensive climatic changes related to the occurrence of periodic droughts and rising temperatures as well as the lack of constant environmental conditions at the experimental environments require the breeder to create varieties that are prepared for the unprecedented changes. The key aspect is the development of an experimental network enabling a comprehensive study of breeding materials: on the one hand, environmentally diverse, in order to study the interactions taking place, and on the other, allowing for a more complete disclosure of genotypic differences not burdened by the influence of other factors (Annicchiarico 1997; Gauch and Zobel 1997).
An effective statistical tool for the analysis of the abovementioned issues is the additive main effects and multiplicative interaction (AMMI) model (Gauch 2006; Zobel et al. 1988). It enables drawing conclusions about the stability of the studied genotypes and the nature of their adaptation and allows grouping of objects and locations on the basis of the GEI profile (Crossa 1990). According to Gauch et al. (2008), it is the most suitable method for plant breeding applications, and its main benefit is faster progress in yield and other traits.
The aim of the study was to investigate the changes in the yield level of spring wheat breeding lines and reference varieties, in response to various environmental conditions. The scope of the research presented included two aspects: genotypic and environmental. The stability of genotypes in terms of yielding was assessed, their hierarchy was constructed in terms of wide adaptation, and the similarity of the studied genotypes in terms of the GEI profile was analyzed, also in relation to the COBORU reference variety. The environmental profile of the research, in one-year terms, included analysis of the experimental network, grouping of locations similarly affecting the studied genotypes, identification of environments least influencing the yield level and allowing more complete disclosure of genotypic differences.
Materials and methods
The research material was the preliminary spring wheat trial lines, established in 2021. These were 23 genotypes from Plant Breeding Strzelce Ltd. Co., IHAR Group, Poland, and three reference varieties: Harenda, Jarlanka–COBORU standards and KWS Dorium. The experiment was set up in the randomized complete block (RCB) system, in three repetitions and six locations [BOH—Borowo (52° 07′ 12″ N, 16° 47′ 19″ E), KOH—Kończewice (53° 10′ 55″ N, 18° 33′ 20″ E), KBP—Kobierzyce (50° 58′ 17″ N, 16° 55′ 50″ E), MAH—Małyszyn (52° 44′ 19″ N, 15° 10′ 17″ E), SMH—Smolice (51° 42′ 12″ N, 17° 10′ 10″ E), and STH—Strzelce (52° 18′ 41″ N, 19° 24′ 22″ E)] (Fig. S1). Plot size to harvest was 10 m2. The experimental results, grain yield (kg plot−1), were statistically analyzed.
The data were analyzed using the additive main effects and multiplicative interaction (AMMI) model (Gauch and Zobel 1990), for spring wheat grain yield. The AMMI model first fits the additive effects for the main effects of genotypes (G) and environment (E), followed by multiplicative effects for GEI by principal component analysis (PCA). The results of the AMMI analysis are presented as biplot graphs. The AMMI model (Nowosad et al. 2016) is expressed by the following formula:
where yge is the trait mean of a genotype g in environment e, μ is the grand mean, αg is the mean genotype deviation, βe is the mean environment deviation, N is the number of PCA axes retained in the adjusted model, λn is the eigenvalue of the PCA axis n, γgn is the genotype score for the PCA axis n, δen is the score eigenvector for the PCA axis n and Qge is the residual, including the AMMI noise and pooled experimental error. AMMI analysis was performed in the R Studio (http://www.r-project.org/) using the “AMMI” function from “agricolae” package. The stability of genotypes was assessed using the AMMI stability value (ASV) coefficient (Purchase et al. 2000):
where SSIPCA1 is the sum of squares for IPCA1, SSIPCA2 is the sum of squares for IPCA2, and the IPCA1 and IPCA2 scores are the genotype scores in the AMMI model. The lower ASV value, the greater the stability of the genotype in the studied environments. The genotype selection index (GSI)—the sum of the ASV and yield stability index (YSI) ranking positions (Farshadfar 2008), was calculated for each genotype.
In order to better understand the relationships between the studied genotypes and to compare the GEI profiles of the studied genotypes with the Harenda (a long-term reference variety used in COBORU, with a high and stable yield level over the years), cluster analysis was used as a supporting tool (Hühn and Truberg 2002). Grouping was carried out using the hierarchical Ward method, and the results were visualized as dendrogram. Cluster analysis was performed in the R Studio (http://www.r-project.org/) using package “factoextra.”
Results
The analysis of variance of 26 genotypes, in six environments, showed a significant influence of all the analyzed factors on the yield of spring wheat grain (Table 1). As much as 89.0% of the total sum of the squared deviations for the yield means was explained by the main effects of environments. Such a high value of this parameter indicates a huge differentiation of the environmental averages of the yield. The expression of the trait was significantly modified by the conditions in a given environment, which is evident for the complex polygenic trait, which is the yield. The main effects of the genotypes explained a similar level of observed variability (4.3%) to the interaction effects of GEI (5.2%). This means that 9.5% of the trait variation could be used to detect genotypes with narrow adaptation (Mądry et al. 2006a).
PCA showed three significant interaction components. The first two, explaining a total of 78.2% of the GEI effects variability, were used to construct biplots in the AMMI analysis (Figs. 1, 2).
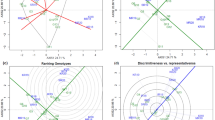
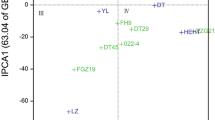
The AMMI method was used to study the structure and yield level of spring wheat for the breeding program purposes. Based on the ASV values (Table 2) and the interpretation of biplot 1 (Fig. 1), the most stable of the studied genotypes were: KOH 18293 (0.14), STH 21-03 (0.17), STH 21-02 (0.21), KOH 19367 (0.25), KOH 19034 (0.28), KOH 18280 (0.28), KOH 19209 (0.32), STH 21-09 (0.33), KOH 19174 (0.38), STH 21-08 (0.41), STH 21-04 (0.42). In the biplot 1 of the first and second main components (Fig. 1), these genotypes are located in the near distance from the origin of the coordinate system.
The best yielding genotype KOH 19678 (7.63 kg) was the least stable (ASV = 1.37) and highly adapted to the conditions of Smolice. The most stable genotype KOH 18293 was on the 14th place in the yield ranking (6.81 kg). As evident, stability in itself cannot be the basis for selection in breeding program. The most stable genotype is not always, or even relatively rarely, the highest yielding genotype. The coefficient, taking into account both the stability and the level of yielding, was proposed by Farshadfar (2008). The genotype selection index—GSI, allows ordering and creation of transparent rankings of genotypes breeding value. Genotypes with the lowest GSI coefficient (Table 2), and thus broad adaptation, were: STH 21-03 (6), STH 21-09 (13), KOH 263 (15), KOH 18279 (16), KOH 18280 (16), STH 21-08 (17), KOH 19209 (19), STH 21-04 (19). They were located to the right of the vertical line, indicating the overall mean yield of all objects, on the biplot 2 for the first main component (IPC1) and the mean grain yield (Fig. 2). Particular attention was paid to the objects: STH 21-03, STH 21-09 and KOH 18279, which, apart from the highest values of the GSI index (6, 13 and 16, respectively), also had an average yield above the reference variety Harenda (7.24, 7.22 and 7.29 kg, respectively) (Table 2).
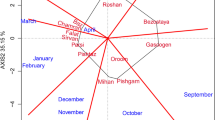
In analyzed growing season, the mean temperatures in individual months of the trial were similar in all analyzed environments. The way in which the locations were grouped reflected the distribution of rainfall, which may suggest a large influence of this factor on the generation of the observed variability (Fig. 3).
Based on the cluster analysis, the analyzed genotypes were divided into four homogeneous groups (Figs. 4, 5). The reference variety Harenda was assigned to cluster no. 1 along with six other genotypes, e.g., STH 21-09 (Table 3). KOH 18279 was the genotype with the most similar GEI profile to Harenda. Cluster No. 1 was characterized by the highest genotypic mean for the yield and average stability, as measured by the ASV coefficient. The genotypes adapted to the KBP conditions were grouped there, while at the same time poorly adapted to the BOH and MAH conditions. The lowest average ASV coefficient (highest stability) was found in cluster no. 2, which included objects adapted to the KBP and KOH conditions, at the same time poorly adapted to the SMH conditions. Cluster No. 3, including STH 21-03, was characterized by good adaptation to SMH conditions, and poor adaptation to KBP conditions. The average ASV coefficient for this cluster was slightly lower than that of cluster no. 4 (higher stability), and the average yield slightly exceeded cluster no. 2. The objects with the least favorable, from the breeding point of view, ASV parameter values and average yield, were collected in cluster no. 4. These were genotypes adapted to the BOH, STH and MAH conditions, and at the same time poorly adapted to the SMH and KOH conditions. Interestingly, it also contained the Jarlanka (COBORU quality standard) and KWS Dorium varieties.
Based on obtained results, the experimental network was analyzed, on an annual basis. The highest mean of yielding for all genotypes was obtained in KBP (9.42 kg) and the lowest in BOH (5.05 kg) (Table 2). The largest difference between the mean yields for genotypes was obtained in SMH (3.78 kg), and the smallest in STH (1.36 kg). The location that brought the lowest variability of the interaction effects was KOH, which was manifested by the shortest vector of environmental parameters among all the investigated locations. The yielding of genotypes in this environment was the best reflection of the yield ordering for the means from the entire experiment, and thus, best represented the genotypic means of the studied trait. Due to the weakest GE interactions in KOH, genotypic differences could be revealed. The location where the strongest GE interactions occurred was SMH. Here, the greatest differences were observed between the mean yields of the studied genotypes and in replications. The environments that group together should reflect the consistency of the influence on the genotypes and thus define the direction of selection (Cooper et al. 1993). A similar ordering of the mean yields was obtained in KOH and KBP as well as in MAH, BOH and STH—the vectors of the abovementioned groups of locations create acute angles (Fig. 1) (Gauch and Zobel 1997). SMH revealed its distinctiveness. The observed correlations between the analyzed locations were confirmed by grouping the environments, based on the genotypes GE interaction profiles (Fig. 5), carried out using the Ward’s method.
Discussion
The literature provides many examples of the AMMI model application in the studies of GEI shape, for various features of high economic importance species, such as maize (Branković-Radojčić et al. 2018; Bocianowski et al. 2019a; Shojaei et al. 2021), wheat (Crossa et al. 1991; Mahmodi et al. 2011; Verma et al. 2015; Verma and Singh 2021; Bocianowski and Prażak 2022), rapeseed (Nowosad et al. 2016; Liersch et al. 2020), barley (Nowosad et al. 2018; Bocianowski et al. 2019b) or sugar beet (Abbasi and Bocianowski 2021). In this study, the AMMI method was used to examine the structure and level of spring wheat yield for the purposes of the breeding program.
The distribution of the studied genotypes in the coordinate system of the IPC1 and IPC2 parameters (Fig. 1) indicates their various reactions in response to the conditions in the studied environments. Mądry et al. (2006a) in their work on the winter wheat yield response to various environmental conditions also obtained a large variety of responses of the studied objects. The presented experiences covered one year of research, which, according to the authors, constitutes a certain limitation of inference possibility. In breeding practice, there is a large rotation of objects in trials, every year different genotypes are tested; therefore, the key aspect is to have an appropriate network of field experiments. Drzazga and Krajewski (2001), in their research on winter wheat, concluded that the shape of GEI in an environment depends on particular season conditions, rather than trial location or genotypes set. They noted the role of a season climatic conditions and their impact on shaping the GEI. In the presented research, also the level of yielding depended mainly on the conditions in the environments, especially rainfall.
The advantage of cluster analysis, based on GEI, has been discussed before (Lin and Binns 1991; Bull et al. 1992; Cooper et al. 1993; Crossa et al. 1993; Baril et al. 1994; Ouyang et al. 1995). By dividing objects and environments into homogeneous groups, interactions within groups are minimized. On the basis of the reaction profile of one of the group’s objects, it is possible to conclude on the behavior of the others, or to select the best representatives of a group—e.g., candidates for model varieties. To add to it, one can limit the set of test environments to the most common of a group. However, according to Crossa et al. (1993), due to the existence of many agglomeration algorithms and distance measures, each of them may give different results. Therefore, the key aspect is appropriate selection of grouping methods and parameters.
An attempt to reduce the number of locations in the experimental network, based on cluster analysis, was made by Baril et al. (1994). The comparison of environments grouping results, made independently in subsequent years, allowed to reduce the number of locations and thus real savings for the breeding program. Lin and Binns (1991) describe a method of regional experiences results analyzing, using jointly cluster analysis and AMMI, for four datasets previously compiled by other authors. The results suggested that this approach provides comprehensive information for breeding purposes. They point out that cluster analysis methods can help plant breeders identify general types of responses among the genotypes tested, but their performance declines with the number of subjects tested.
The utility of the AMMI method, in the conditions of Polish plant breeding company, was confirmed as a tool which enables the selection of stable and widely adaptable genotypes. In addition, the method allows for a more precise prediction of breeding activities effects, in terms of yield, but also many other traits, and thus more efficient progress in breeding. The genotypes of particular interest, in terms of the analyzed issues, turned out to be: STH 21-03, STH 21-09 and KOH 18279. The usefulness of the Ward cluster analysis was confirmed, as a supplement to the AMMI model, that classifies genotypes in a transparent manner, enabling a more complete comparison with reference variety. Particular attention was paid to cluster no. 1, in which the genotypes with the GEI pattern closest to the Harenda’s were grouped. Due to the fact that the presented research covered only one growing season, the possibilities of drawing conclusions about the general relationships between the studied environments were limited, and the experimental network was summarized on an annual basis. The environments that had a similar effect on the studied genotypes in terms of yield shaping, in 2021, were: Kończewice and Kobierzyce, Borowo, Małyszyn and Strzelce. Smolice, which was characterized by the strongest GEI, revealed its distinctiveness.
References
Abbasi Z, Bocianowski J (2021) Genotype by environment interaction for physiological traits in sugar beet (Beta vulgaris L.) parents and hybrids using additive main effects and multiplicative interaction model. Eur Food Res Technol 247:3063–3081. https://doi.org/10.1007/s00217-021-03861-4
Annicchiarico P (1997) Additive main effects and multiplicative interaction (AMMI) analysis of genotype–location interaction in variety trials repeated over years. Theor Appl Genet 94:1072–1077. https://doi.org/10.1007/s001220050517
Baril CP, Denis JB, Brabant P (1994) Selection of environments using simultaneous clustering based on genotype–environment interaction. Can J Plant Sci 74:311–317. https://doi.org/10.4141/cjps94-059
Bocianowski J, Prażak R (2022) Genotype by year interaction for selected quantitative traits in hybrid lines of Triticum aestivum L. with Aegilops kotschyi Boiss. and Ae. variabilis Eig. using the additive main effects and multiplicative interaction model. Euphytica 218(2):11. https://doi.org/10.1007/s10681-022-02967-4
Bocianowski J, Nowosad K, Tomkowiak A (2019a) Genotype–environment interaction for seed yield of maize hybrids and lines using the AMMI model. Maydica 64(2):M13
Bocianowski J, Warzecha T, Nowosad K, Bathelt R (2019b) Genotype by environment interaction using AMMI model and estimation of additive and epistasis gene effects for 1000-kernel weight in spring barley (Hordeum vulgare L.). J Appl Genet 60:127–135. https://doi.org/10.1007/s13353-019-00490-2
Branković-Radojčić D, Babić V, Girek Z, Živanović T, Radojčic A, Filipović M, Srdić J (2018) Evaluation of maize grain yield and yield stability by AMMI analysis. Genetika 50(3):1067–1080. https://doi.org/10.2298/GENSR1803067B
Bull JK, Basford KE, De Lacy IH, Cooper M (1992) Classifying genotypic data from plant breeding trials: a preliminary investigation using repeated checks. Theor Appl Genet 85:461–469. https://doi.org/10.1007/BF00222328
Cooper M, Byth DE, De Lacy IH (1993) A procedure to assess the relative merit of classification strategies for grouping environments to assist selection in plant breeding regional evaluation trials. Field Crops Res 35:63–74. https://doi.org/10.1016/0378-4290(93)90137-C
Crossa J (1990) Statistical analysis of multilocation trials. Adv Agron 44:55–85. https://doi.org/10.1016/S0065-2113(08)60818-4
Crossa J, Fox PN, Pfeiffer WH, Rajaram S, Gauch HG (1991) AMMI adjustment for statistical analysis of an international wheat yield trial. Theor Appl Genet 81:27–37. https://doi.org/10.1007/BF00226108
Crossa J, Cornelius PL, Seyedsadr M, Byrne P (1993) A shifted multiplicative model cluster analysis for grouping environments without genotypic rank change. Theor Appl Genet 85:577–586. https://doi.org/10.1007/BF00220916
Drzazga T, Krajewski P (2001) Zróżnicowanie środowisk pod względem stopnia interakcji w seriach doświadczeń z pszenicą ozimą. Biuletyn IHAR 218(219):111–115 (in Polish)
Farshadfar E (2008) Incorporation of AMMI stability value and grain yield in a single non-parametric index (GSI) in bread wheat. Pak J Biol Sci 11(14):1791–1796
Gauch HG (2006) Statistical analysis of yield trials by AMMI and GGE. Crop Sci 46:1448–1500. https://doi.org/10.2135/cropsci2005.07-0193
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761. https://doi.org/10.1007/BF00224240
Gauch HG, Zobel RW (1997) Identifying mega-environments and targeting genotypes. Crop Sci 37:311–326. https://doi.org/10.2135/cropsci1997.0011183X003700020002x
Gauch HG, Piepho HP, Annicchiarico P (2008) Statistical analysis of yield trials by AMMI and GGE: further considerations. Crop Sci 48:866–889. https://doi.org/10.2135/cropsci2007.09.0513
Grote U, Fasse A, Nguyen TT, Erenstein O (2021) Food security and the dynamics of wheat and maize value chains in Africa and Asia. Front Sustain Food Syst 4:617009. https://doi.org/10.3389/fsufs.2020.617009
Hühn M, Truberg B (2002) Contributions to the analysis of genotype × environment interactions: theoretical results of the application and comparison of clustering techniques for the stratification of field test sites. J Agron Crop Sci 188:65–72. https://doi.org/10.1046/j.1439-037X.2002.00549.x
Liersch A, Bocianowski J, Nowosad K, Mikołajczyk K, Spasibionek S, Wielebski F, Matuszczak M, Szała L, Cegielska-Taras T, Sosnowska K, Bartkowiak-Broda I (2020) Effect of genotype × environment interaction for seed traits in winter oilseed rape (Brassica napus L.). Agriculture 10:607. https://doi.org/10.3390/agriculture10120607
Lin CS, Binns MR (1991) Assessment of a method for cultivar selection based on regional trial data. Theor Appl Genet 82:379–388. https://doi.org/10.1007/BF02190626
Mądry W, Paderewski J, Drzazga T (2006a) Ocena reakcji plonu ziarna rodów hodowlanych pszenicy ozimej na zmienne warunki środowiskowe za pomocą analizy AMMI. Fragm Agron 23(42):130–143 (in Polish)
Mądry W, Talbot M, Ukalski K, Drzazga T, Iwańska M (2006b) Podstawy teoretyczne znaczenia efektów genotypowych i interakcyjnych w hodowli roślin na przykładzie pszenicy ozimej. Biul IHAR 240(241):13–32 (in Polish)
Mahmodi N, Yaghotipoor A, Farshadfar E (2011) AMMI stability value and simultaneous estimation of yield and yield stability in bread wheat (Triticum aestivum L.). Aust J Crop Sci 5(13):1837–1844
Nowosad K, Liersch A, Popławska W, Bocianowski J (2016) Genotype by environment interaction for seed yield in rapeseed (Brassica napus L.) using additive main effects and multiplicative interaction model. Euphytica 208:187–194. https://doi.org/10.1007/s10681-015-1620-z
Nowosad K, Tratwal A, Bocianowski J (2018) Genotype by environment interaction for grain yield in spring barley using additive main effects and multiplicative interaction model. Cereal Res Commun 46(4):729–738. https://doi.org/10.1556/0806.46.2018.046
Ouyang Z, Mowers RP, Jensen A, Wang S, Zheng S (1995) Cluster analysis for genotype–environment interaction with unbalanced data. Crop Sci 35:1300–1305. https://doi.org/10.2135/cropsci1995.0011183X003500050008x
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. S Afr J Plant Soil 17(3):101–107. https://doi.org/10.1080/02571862.2000.10634878
Shojaei SH, Mostafavi K, Omrani A, Omrani S, Mousavi SMN, Illes A, Bojtor C, Nagy J (2021) Yield stability analysis of maize (Zea mays L.) hybrids using parametric and AMMI methods. Hindawi Sci 2021:5576691. https://doi.org/10.1155/2021/5576691
Verma A, Singh GP (2021) AMMI with BLUP analysis for stability assessment of wheat genotypes under multi locations timely sown trials in Central Zone of India. Int J Agric Sci Food Technol 7(1):118–124. https://doi.org/10.17352/2455-815X.000098
Verma A, Chatrath R, Sharma I (2015) AMMI and GGE biplots for G × E analysis of wheat genotypes under rain fed conditions in central zone of India. J Appl Natural Sci 7(2):656–661. https://doi.org/10.31018/jans.v7i2.662
Zobel RW, Wright MJ, Gauch HG (1988) Statistical analysis of a yield trial. Agron J 80:388–393. https://doi.org/10.2134/agronj1988.00021962008000030002x
Funding
The research was carried out as part of the project: “Obtaining a new generation of Polish varieties of rape, cereals and legumes resistant to pests, with better mitigation and adaptation abilities to climate change, with appropriate technological features required by consumers and industry”; Project number: POIR. 01.01.01-00-0782/16-00.
Author information
Authors and Affiliations
Contributions
Conceptualization was contributed by SJ and JB; methodology was contributed by SJ and JB; software was contributed by SJ; validation was contributed by SJ and JB; formal analysis was contributed by SJ and JB; investigation was contributed by SJ, JB and PM; resources were contributed by SJ; data curation was contributed by SJ; writing—original draft preparation, was contributed by SJ and JB; writing—review and editing, was contributed by SJ, JB and PM; visualization was contributed by SJ and JB; supervision was contributed by JB; project administration was contributed by JB; funding acquisition was contributed by PM. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals or humans performed by any of the authors.
Additional information
Communicated by M. A. R. Arif.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jędzura, S., Bocianowski, J. & Matysik, P. The AMMI model application to analyze the genotype–environmental interaction of spring wheat grain yield for the breeding program purposes. CEREAL RESEARCH COMMUNICATIONS 51, 197–205 (2023). https://doi.org/10.1007/s42976-022-00296-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42976-022-00296-9