Abstract
Vitamin C (L-ascorbic acid) is an essential nutrient for human health, and its deficiency has long been known to cause scurvy. Sodium-dependent vitamin C transporters (SVCTs) are responsible for vitamin C uptake and tissue distribution in mammals. Here, we present cryogenic electron microscopy structures of mouse SVCT1 in both the apo and substrate-bound states. Mouse SVCT1 forms a homodimer with each protomer containing a core domain and a gate domain. The tightly packed extracellular interfaces between the core domain and gate domain stabilize the protein in an inward-open conformation for both the apo and substrate-bound structures. Vitamin C binds at the core domain of each subunit, and two potential sodium ions are identified near the binding site. The coordination of sodium ions by vitamin C explains their coupling transport. SVCTs probably deliver substrate through an elevator mechanism in combination with local structural arrangements. Altogether, our results reveal the molecular mechanism by which SVCTs recognize vitamin C and lay a foundation for further mechanistic studies on SVCT substrate transport.
Similar content being viewed by others
Introduction
Vitamin C is essential for human health and participates in numerous physiological processes, including collagen hydroxylation1,2,3, hormone synthesis3,4, the demethylation of DNA and histones5, hypoxia-inducible factor hydroxylase-mediated oxygen sensing6, free radical scavenging7, and immune regulation8. In these processes, vitamin C functions as an enzymatic cofactor or an antioxidant. For example, vitamin C is a cofactor for prolyl and lysyl hydroxylases that hydroxylate collagen1,2, and vitamin C deficiency causes collagen instability and scurvy9, a disease characterized by muscle weakness, tooth loss and impaired wound healing10.
Humans are not able to synthesize ascorbic acid de novo and need to absorb exogenous vitamin C from the diet due to the inactivation mutation of the gene encoding the gulono-gamma-lactone oxidase that catalyzes ascorbic acid synthesis from glucose11. Membrane transporters are thus required for the uptake of vitamin C into the body. While glucose transporters (GLUTs), including GLUT1, GLUT3 and GLUT4, take up dehydroascorbic acid (the oxidized form of vitamin C) by facilitating diffusion12,13,14, sodium-dependent vitamin C transporters (SVCTs) mediate the active uptake of L-ascorbic acid (the reduced form) with high substrate specificity15,16,17,18,19,20,21. Three different isoforms, SVCT1-3, encoded by SLC23A1-3, have been identified in the human SVCT family. SVCT1 and SVCT2 are symporters that cotransport sodium and vitamin C with a 2:1 stoichiometry down electrochemical Na+ gradients across cell membranes17,18. SVCT1 is primarily expressed in epithelial tissues, such as the intestine and kidney, and plays a key role in maintaining the whole-body homeostasis of vitamin C10,15,22. In contrast, the widely expressed SVCT2 distributes vitamin C to various tissues10,15,19, including the brain, retina, placenta, spleen, and prostate. Both Slc23a1 and Slc23a2 knockout mice showed high perinatal mortality23,24. SVCT3 does not transport vitamin C, and its function is still unknown10.
SVCTs belong to the nucleobase-ascorbate transporter (NAT) family25,26, the members of which transport a variety of physiological substrates, including xanthine, uric acid, uracil, and vitamin C. Substrate specificity and transport of NAT proteins were suggested to be related to a signature motif with a conserved sequence [Q/E/P]-N-X-G-X4-T-[R/K/G] (X indicates a nonspecific amino acid)27 (Supplementary Fig. 1). Previous structural studies of the bacterial uracil transporter UraA and the fungal xanthine transporter UapA provided important insights into nucleobase recognition and transport28,29,30. However, mammalian SVCTs specifically catalyze sodium-dependent vitamin C transport.
In this work, we present the cryogenic electron microscopy (cryo-EM) structures of MmSVCT1 in both the apo and substrate-bound states, revealing the structural basis of vitamin C binding and transport.
Results
Overall structure of MmSVCT1
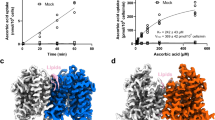
MmSVCT1 was recombinantly expressed in HEK293F cells, and its activity was examined by an in vitro ascorbic acid transport assay (Methods). In this assay, cells were first incubated with ascorbic acid and then washed and lysed to release the imported ascorbic acid, which reduces Fe3+ to Fe2+, resulting in a colorimetric product with a characteristic absorbance at 593 nm. Cells overexpressing wild-type MmSVCT1 displayed robust ascorbic acid uptake compared to control cells (Fig. 1a). Diclofenamic acid, an SVCT inhibitor31, efficiently inhibited transport activity (Fig. 1a), verifying that the uptake of ascorbic acid was mediated by MmSVCT1.
a Ascorbic acid uptake activities of MmSVCT1. WT, wild-type; Diclo, diclofenamic acid. Data points are the mean ± SEM (n = 5 independent experiments). P values from the two-tailed t-test are indicated on the bar chart. Source data are provided as a Source Data file. b, c Cryo-EM density map (b) and cartoon representation (c) of the MmSVCT1 dimer with one subunit colored by individual elements and the other colored gray. Vitamin C is shown as spheres. d Domain topology of the MmSVCT1 subunit. e One MmSVCT1 subunit viewed from the top colored as in b–d. The disulfide bond between Cys78 and Cys136 is marked by S‒S. EL1 and EL2 are colored blue and red, respectively. f Slab view of the surface electrostatic potential of MmSVCT1 showing the inward-open conformation.
MmSVCT1 structures were determined in the presence and absence of L-ascorbic acid to overall resolutions of 3.2 and 3.5 Å, respectively, using single-particle cryo-EM (Fig. 1b, c, Supplementary Figs. 2–4 and Supplementary Table 1). The structures in the apo and substrate-bound states were highly similar with an RMSD of 0.53 over 894 Cα atoms (Supplementary Fig. 5), so the 3.2-Å vitamin C-bound structure is referred to for most of the description. Similar to other NAT transporters29,30, MmSVCT1 forms a homodimer (Fig. 1b, c), in which each MmSVCT1 subunit consists of 14 transmembrane segments (TM1–TM14) (Fig. 1d, e). TM1–TM7 and TM8–TM14 are inverted structural repeats related by a pseudo twofold axis parallel to the membrane (Supplementary Fig. 6). The transmembrane domain is spatially separated into a gate domain (TM5–TM7 and TM12–TM14) and a core domain (TM1–TM4 and TM8–TM11) (Fig. 1d, e). Transmembrane helices from the gate domain, including TM5, TM6, and TM12 from one subunit and TM13 from the neighboring subunit, form the dimeric interface of MmSVCT1 largely through hydrophobic interactions and van der Waals contacts (Supplementary Fig. 7). Lipid binding was previously shown to be important for UapA dimer formation32, but the lipid binding site is not conserved in MmSVCT1 (Supplementary Fig. 8), and no strong lipid-like density near the UapA lipid binding site was found in MmSVCT1. The core domain contains a vitamin C binding site located between the half helices of TM3 and TM10 (Fig. 1c, e). Above the core domain, there is an elongated extracellular loop (EL2) between TM3 and TM4 (Fig. 1d, e), on which Cys136 forms a disulfide bond with Cys78 on EL1 between TM1 and TM2, contributing to the structural integrity. Both the apo and substrate-bound structures adopt an inward-open conformation, and a solvent-accessible channel sandwiched between the core domain and gate domain can reach the substrate from the cytoplasmic side (Fig. 1f and Supplementary Fig. 5). As in UapA, TM13 of MmSVCT1 contributes to the formation of the inward-facing translocation channel of the neighboring subunit (Supplementary Fig. 9), which may play a role in substrate release30.
Extracellular interfaces
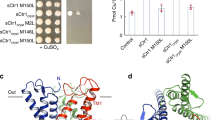
Two tightly packed extracellular interfaces between the core domain and gate domain (interface I and II) comprise the closed extracellular gate of the inward-open structures of MmSVCT1 (Fig. 2a, b). The two interfaces are separated by a central pore with a small radius impermeable to the solvent (Fig. 2a and Supplementary Fig. 10). Localized above the substrate binding site, Interface I is predominantly formed by hydrophobic residues, including Phe119 from TM3 and multiple residues from TM1 and TM12. In addition, Arg213 from TM5 establishes polar interactions with the side chain of Glu75 and the carbonyl group of Phe71 from TM1 on the extracellular side (Fig. 2a), further stabilizing the interface. The alanine substitution of Leu72, Phe119, Arg213 or Ile446 on interface I substantially reduced the transport activity of MmSVCT1 (Fig. 2c), while their overall expression and plasma localization showed no major difference from WT MmSVCT1 (Supplementary Fig. 11). Consistent with our structural and functional analysis of interface I, mutations in TM5 and TM12 of UapA have been previously shown to affect the transport dynamics, substrate affinity, and specificity33. Interface II is formed by residues from TM3, TM8, TM14a, and TM14b (Fig. 2b), in which the side-chain hydroxyl group of Ser127 establishes two hydrogen bonds with the carbonyl group of Val502 and the side-chain hydroxyl group of Thr506. Interface II is important in that several residues on TM3 and TM8 directly participate in the formation of substrate and ion binding sites, as discussed below. The extensive interactions of the extracellular interfaces between the core domain and gate domain maintain MmSVCT1 structures in the inward-facing conformation.
a One subunit of MmSVCT1 viewed from the gate domain with a zoomed-in view of interface I. The TM13 helix was removed for clarity. The central pore calculated by the program Hole is shown as a dotted mesh. b Magnified view of interface II. c Transport activity of MmSVCT1 variants with point mutations at the extracellular interface. Data points are the mean ± SEM (n = 5 independent experiments). P values from the two-tailed t-test are indicated on the bar chart. Source data are provided as a Source Data file.
Vitamin C and sodium binding site
In the cryo-EM density maps of the substrate-bound structure, prominent nonprotein density can be observed in a pocket surrounded by TM1, TM3, TM8, and TM10 (Fig. 3a and Supplementary Fig. 12a, b). A vitamin C molecule can be reasonably placed into the density (Supplementary Fig. 12a, b), although an alternative conformation cannot be completely ruled out with the current local resolution. Previous in silico docking analysis has suggested similar substrate binding in human SVCT234. Interestingly, some extra density can be observed outside the binding pocket between the core and gate domains, which is likely to be detergents or lipids (Supplementary Fig. 13), as seen in the UapA structure30. Most of the residues lining the binding site are polar residues and are highly conserved among SVCTs (Fig. 3a and Supplementary Fig. 1). TM3 and TM10 form a crossing above the substrate. As a result, Ser117 and Phe119 on the connecting loop between the β1 strand and TM3 helix directly interact with the substrate (Fig. 3a), while Ala118 forms hydrogen bonds with the side chain of Asn384 on TM10 (Supplementary Fig. 14a), stabilizing the loop. Below the substrate, residues in the sequence 387Thr-Ser-Ser-Ser, which immediately precede the NAT motif comprising the TM10 helix, establish multiple interactions with vitamin C (Fig. 3a), contributing to substrate specificity determination. Following the 387Thr-Ser-Ser-Ser sequence, Pro391 in MmSVCT1, the initial residue in the NAT motif, interacts with the vitamin C molecule with its bulky side chain through van der Waals contact (Fig. 3a). Consistent with our observation, previous studies have suggested that residues corresponding to Ser389 and Pro391 are key elements that distinguish vitamin C transporters from other nucleobase transporters35,36. In addition, Cys62 and Thr66 on TM1 interact with the substrate from one lateral side, and Glu341 and Asp345 on TM8 stabilize the sodium ions and substrate from the other side (Fig. 3a, b). We performed mutagenesis studies to evaluate the functional significance of these residues around the vitamin C binding site. While the overall expression and plasma localization showed no major difference with WT MmSVCT1 (Supplementary Fig. 11), all alanine substitutions (C62A, S117A, T387A, S389A, or P391A) profoundly reduced vitamin C uptake (Fig. 3c), indicating their critical roles in substrate recognition. The Km of WT MmSVCT1 is close to that of human and rat SVCTs reported previously15,18,20,21 (Supplementary Fig. 15). The C62A and P391A mutants exhibited Km and Vmax values similar to that of WT MmSVCT1, while other mutants showed nearly no activity (Supplementary Fig. 15). Notably, the strong vitamin C density is absent in the apo map despite some unknown densities (Supplementary Fig. 12b, c). Moreover, there is no major structural change in the binding pocket upon substrate binding (Supplementary Fig. 14b), probably because the abundant hydrogen bonds in the unwound regions of TM3 and TM10 stabilize the pocket (Supplementary Fig. 14a). The vitamin C recognition of MmSVCT1 is different from that of the bacterial phosphorylation-coupled vitamin C transporter UlaA37.
a Residues in the proximity of the vitamin C molecule. Sodium ions are shown as blue spheres, and a potential water molecule (Wat) is shown as a red sphere. Interactions are indicated by dashed lines. b Coordinating residues of Na1. c Uptake activity of MmSVCT1 variants with point mutations at vitamin C and sodium ion binding sites. Data points are the mean ± SEM (n = 5 independent experiments). P values from the two-tailed t-test are indicated on the bar chart. Source data are provided as a Source Data file.
The calculated surface electrostatic potential of the binding pocket in the MmSVCT1 protein is rather negative (Fig. 1f). Sodium ion binding, together with the positively charged ends of TM3 and TM10 dipoles, is thus required for the recognition of the negatively charged vitamin C (pKa = 4.70) at physiological pH (Fig. 3a). Two potential Na+ binding sites, designated as Na1 and Na2, were identified around the substrate binding pocket (Fig. 3a). The coordination of Na1 is provided by the main-chain carbonyl oxygen of Ser388 and side-chain oxygens of Glu341, Asp345, Ser390, and vitamin C (Fig. 3b), with coordination distances of 2.2–2.8 Å. Asp345 in MmSVCT1, corresponding to His245 in proton symporter UraA, was suggested to play a key role in sodium symport28, which is consistent with its direct Na+ coordination. Importantly, all mutations of the coordinating residues—E341A, D345A, and S390A—abolished transport activity (Fig. 3c), showing their functional significance. The ion at the tentative Na2 site is less well coordinated and interacts with the substrate and a potential water molecule that is stabilized by Glu341 (Fig. 3a).
Transport mechanism
To understand the transport mechanism of MmSVCT1, we compared our inward-open MmSVCT1 structure with a previously reported occluded UraA structure29. The gate domain of MmSVCT1 and UraA can be aligned exceptionally well with a backbone RMSD of ~1.3 Å (Fig. 4a), rationalizing our analysis based on the comparison of two structures. The largest structural difference between the gate domains of the two structures is from the loop EL7 between TM13 and TM14 (Fig. 4a); two beta strands in the loop of UraA are replaced by elongated TM13 helix and TM14a in MmSVCT1. When superposed by the gate domains, the intracellular segments of the core domain helices undergo outwards movement from the occluded structure to the inward-open structure, resulting in the opening of the intracellular side of the MmSVCT1 structure (Fig. 4b–d). Along with that, the substrate bound in the core domain shows a downward shift of ~5 Å (Fig. 4b), which is consistent with an elevator mechanism first proposed for GltPh38. However, instead of a rigid-body movement, the core domain is also subjected to local structural arrangements: the extracellular segments of the core domain helices of MmSVCT1 undergo an anticlockwise rotation compared to their UraA counterparts (Fig. 4b, c). Thus, MmSVCT1 explores a mechanism combining an elevator-like shift with local structural changes to deliver the substrate (Fig. 4d).
a Structural comparison of the gate domain between inward-open MmSVCT1 (cyan) and occluded UraA (gray). b, c Movements of the core domain of the inward-open MmSVCT1 (orange or yellow) and occluded UraA (gray) viewed from the gate domain. Vitamin C and uracil are shown as purple and gray spheres, respectively. The directions of movements of the substrate (black arrows), the extracellular segments (green arrows) and intracellular segments (red arrows) of the core domain helices are indicated. d Working model for the vitamin C transport of MmSVCT1. The purple pentagons and blue spheres indicate the substrate and sodium ions, respectively.
Discussion
Previous electrophysiological studies suggested an SVCT binding sequence of Na+, L-ascorbic acid, Na+17,18. We noticed that the substrate binding site is located at a position halfway across the membrane in the core domain, which is surrounded by multiple polar residues and a largely negative electrostatic surface. A sodium ion may first bind near the Na1 site, the stronger ion binding site, partly neutralizing Asp345 and/or Glu341 on TM8, which makes the substrate binding site more suitable for negatively charged vitamin C. Vitamin C binding is then followed by Na+ at the Na2 binding site, which is primarily coordinated by vitamin C. In line with our analysis, a similar transport mechanism of UapA has been proposed previously39, in which the symported proton neutralized the negative charge of D360 and E356 (corresponding to D341 and E345 in MmSVCT1) and then allowed substrate binding and transport. In the inward-facing conformation of MmSVCT1, the bound sodium ions are likely in a prerelease conformation, and their dissipation into the low-sodium cytosol would promote vitamin C release. Thus, the structural analyses of MmSVCT1 presented here rationalize previously reported electrophysiological data and provide critical insights into vitamin C and ion recognition as well as their cotransport.
MmSVCT1 functions as a homodimer with its gate domain forming the dimeric interface. This is similar to the microbial NAT/SLC23 transporters UraA29 and UapA30, anion exchange 1 (SLC4A1)40 and plant borate efflux transporter Bor141, but different from the epithelial anion transporter SLC26A942,43 and plant sulfate transporter AtSULTR4;144, which are dimerized primarily with cytoplasmic domains. Structural comparison between the inward-open MmSVCT1 and the occluded UraA suggests that SVCTs transport vitamin C through an elevator mechanism (Fig. 4b, d). Relative to the gate domain, the substrate-bound core domain undergoes rotation with some local rearrangements to transport the substrate. The gate domain is very similar between the inward-open MmSVCT1 and the occluded UraA (Fig. 4a), which is likely because both structures have closed extracellular gates. We suspect that when the transporter switches to an outward-open conformation, the extracellular interfaces (Interfaces I and II) would be destroyed, and the helices in the gate domain, especially for TM12 in interface I containing a hydrophobic surface (Fig. 2a), would rearrange to adapt to the environment exposed to solvent. This can then lead to structural changes on the dimeric interface side of the gate domain. As the same conformation was observed for both subunits in all known dimeric SLC4/23 structures29,30,40,41 and MmSVCT1 structures in the current study, concurrent rearrangements of the dimeric gate domains may provide a mechanism for coordinating the two neighboring subunits. Indeed, the dimeric interface has been proven to be essential for the substrate transport of UraA29 and UapA30. Future work needs to determine SVCT structures in outward-facing and occluded states to pinpoint the detailed conformational changes during a transport cycle for this family of vitamin C transporters. Together, our structural studies offer important insights into vitamin C recognition and lay a foundation for further mechanistic studies.
Methods
Protein expression, purification and nanodisc reconstitution
MmSVCT1 (SLC23A1, NCBI accession: NP_035527.3) was cloned into a pEZT-BM vector with a C-terminal strep tag and heterologously expressed in HEK293F cells (Thermo Fisher Scientific, R79007) using the BacMam system (Thermo Fisher Scientific). The primers used in this study are provided as a Source Data file. Baculovirus was generated from Sf9 cells (Thermo Fisher Scientific, 11496015) and used to infect HEK293F cells at a volume ratio of 1:40 (virus:cell). HEK293F cells were cultured in suspension in 293F medium (Sino Biological Inc.) at 37 °C and 5% CO2. Sodium butyrate (10 mM) was added to the culture medium to boost protein expression. After 48–72 h, the cells were harvested by centrifugation at 5000 g, resuspended in buffer A (50 mM Tris, pH 8.0, 150 mM NaCl), and homogenized by sonication on ice. A protease inhibitor cocktail (containing 2 μg/ml DNase I, 0.5 μg/ml pepstatin, 2 μg/ml leupeptin, 1 μg/ml aprotinin, and 0.1 mM PMSF) was added to the buffer throughout the purification process. MmSVCT1 was extracted with 1% (w/v) n-dodecyl-β-d-maltopyranoside (DDM, Anatrace) supplemented with 0.2% (w/v) cholesteryl hemisuccinate (Sigma-Aldrich) by gentle agitation for 2 h. After extraction, the supernatant was collected after 60-min centrifugation at 30,000 g and purified with Strep-Tactin XT 4Flow resin (IBA). During resin washing, the buffer was gradually changed to buffer B (50 mM Tris, pH 8.0, 150 mM NaCl, and 0.03% GDN). The protein was then eluted with buffer B supplemented with 50 mM D-biotin, concentrated to a final volume of 500 μl, and further purified by size exclusion chromatography on a Superose 6 10/300 GL column (GE Healthcare) preequilibrated with buffer B.
Nanodisc reconstitution was performed by following a published protocol45. Peak fractions after size exclusion chromatography were pooled and mixed with MSPE3D1 and lipids (POPC: POPG: POPE = 3:1:1) at a molar ratio of 1:4:20 (MmSVCT1: MSPE3D1: lipid). After detergent removal by Bio-Beads SM2 (Bio-Rad), the sample was loaded onto a Superose 6 10/300 GL column preequilibrated with buffer A. Peak fractions of reconstituted MmSVCT1 were collected for cryo-electron microscopy analysis.
Cryo-EM sample preparation and EM data acquisition
For the apo MmSVCT1 sample, cryo-EM grids were prepared by applying 4 μl of MmSVCT1 in GDN (~4.0 mg/ml) to a glow-discharged Quantifoil R2/1 200-mesh copper holey carbon film (Quantifoil, Micro Tools GmbH, Germany). Grids were blotted for 3.0 s under 100% humidity at 4 °C before being plunged into liquid ethane using a Mark IV Vitrobot (Thermo Fisher). For the substrate-bound MmSVCT1 sample, MmSVCT1 in nanodiscs (~3.0 mg/ml) was incubated with 1 mM ascorbic acid for 30 min on ice before grid freezing.
Micrographs were acquired on a Titan Krios microscope (Thermo Fisher) operated at 300 kV with a K3 Summit direct electron detector (Gatan). Images were recorded with EPU software (Thermo Fisher) in counting mode with a pixel size of 0.82 Å. The defocus range was set from −1.5 to −2.7 μm. Each micrograph was dose-fractionated to 30 frames under a dose rate of ~18.7 e-/Å2/s, with a total exposure time of 3 s.
Image processing
Data processing was performed using the software package cyroSPARC46. Micrographs were motion corrected with patch motion correction. The CTF parameters of micrographs were estimated using patch CTF estimation. For the apo MmSVCT1 dataset, using templates selected from the 2D classification of 151 manually picked particles, 2,464,213 particles were automatically picked and extracted from the full dataset of 2789 micrographs. After 2D classification, a small dataset containing 83,323 particles was selected for ab-initio reconstruction to generate the reference maps. Then, a total of 614,477 particles resulting from 2D classification were used for heterogeneous refinement. Four rounds of heterogeneous refinement were performed using two reference maps, one with good secondary structural features and one with an empty micelle. After each heterogeneous refinement, particles assigned to the good class were selected for the subsequent heterogeneous refinement. A final dataset of 213,199 particles was then subjected to non-uniform refinement with C2 symmetry imposed, yielding a reconstruction at a resolution of 3.5 Å.
The dataset for substrate-bound MmSVCT1 in nanodiscs was processed similarly to that of apo MmSVCT1. In brief, 2,370,225 particles were picked and extracted from 4708 micrographs. After 2D classification, a total of 444,100 particles were selected for heterogeneous refinement. Three rounds of heterogeneous refinement were performed with two reference maps generated by ab-initio reconstruction from a small dataset of 88,107 particles. A dataset of 224,682 particles was then subjected to non-uniform refinement with C2 symmetry imposed, yielding a reconstruction at a resolution of 3.2 Å. To improve the resolution, one round of local refinement was performed with a soft mask around the protein density (excluding the belt-like density from the nanodisc), yielding a 2.8 Å resolution cryo-EM map. The 3.2 Å map with better peripheral loop density was used to build the overall structure of substrate-bound MmSVCT1 in nanodiscs, and the 2.8 Å map from local refinement was used to help determine the substrate conformation.
Model building, refinement, and validation
Model building was conducted in Coot47, and an MmSVCT1 structure (AF-Q9Z2J0-F1) from the AlphaFold Protein Structure Database48 was used as an initial reference. Real-space model refinement49 and validation50 were performed in Phenix. For the substrate-bound state, residues 37–239, 258–534, and 550–576 are included in the final structure model; residues 1–36, 240–257, 535–549, and 577–605 are disordered and not modeled. The sugar extended from Asn151 has visible but weak density, so it was not modeled. For the apo state, the final structure model includes residues 43–239, 258–534, and 550–576. The statistics of the model geometries were generated using MolProbity51. The transport pathway of MmSVCT1 was predicted by the program HOLE52. The surface electrostatic potential was calculated by the program APBS in PyMol. Figures were prepared using PyMol53 and ChimeraX54 software. Q-scores characterizing atom resolvability in the map were calculated as previously described55.
Ascorbic acid uptake assay
MmSVCT1 transport activity was assayed by measuring ascorbic acid uptake in HEK293F cells. In this assay, the ascorbic acid concentration was determined by using the Ferric Reducing/Antioxidant and Ascorbic Acid kit (Sigma, MAK075). Wild-type MmSVCT1 and variants containing a C-terminal GFP followed by a Strep tag were cloned into a pEZT-BM vector and transiently expressed in HEK293F cells using linear polyethylenimine Max (Polysciences) according to the manufacturer’s protocol. Cells were cultured in suspension in 293F medium (Sino Biological Inc.) at 37 °C and 5% CO2. Twenty-four hours after transfection, the medium was replaced with fresh 293F medium supplemented with 10 mM sodium butyrate. After an additional 24 h of culture, the cells were incubated in fresh HEK293F medium supplemented with 0.1 mM ascorbic acid at 37 °C for 30 min. For the inhibition assay, 0.1 mM diclofenamic acid was added. Approximately 2 × 106 cells were then collected and washed twice with ice-cold PBS and rapidly homogenized in lysis buffer from the kit. After removing insoluble material by centrifuging at 13,000 × g for 10 min at 4 °C, the sample was added to a 96-well plate and incubated with a reaction mix for 2–3 min at room temperature. During this process, Fe3+ was reduced to Fe2+ by ascorbic acid present in the sample, resulting in a colorimetric product. A blank control was set simultaneously for each sample, in which ascorbic oxidase was added to oxidize the ascorbic acid, and its reading was subtracted from the sample reading to obtain the corrected measurement. Meanwhile, a series of ascorbic acid standards with 0 (blank), 2, 4, 6, 8, and 10 nmol/well were made to set up a standard curve. The absorbance at 593 nm was measured for each sample, and the amount of ascorbic acid was determined from the standard curve. All samples were run in 5 repeats.
The Km and Vmax of WT and MmSVCT1 mutants were determined by fitting the data to the Michaelis–Menten equation in Origin software. The initial transport velocities were measured at 1 min over a substrate concentration range of 0–200 μM. All experiments were repeated three times.
Western blot and immunofluorescence imaging
To monitor the expression level and subcellular localization, WT and MmSVCT1 mutants containing a C-terminal GFP followed by a Strep tag were transfected into HEK293T cells (ATCC, #CRL-3216). The cells were maintained in DMEM (Cytiva) with 10% fetal bovine serum (HyClone) and 100 units/ml penicillin plus 100 µg/ml streptomycin. For western blot, cells were harvested 48 h after transfection and solubilized in a buffer containing 50 mM Tris, pH 8.0, 150 mM NaCl, and 1% DDM for 1 h at 4 °C. After centrifugation at 20,000 × g for 30 min, equal amounts of samples were resolved on an SDS-polyacrylamide gel and transblotted onto a PVDF membrane. The membrane was then probed with 1:2000 diluted mouse anti-StrepII-tag monoclonal antibody (ABclonal, AE066, Clone No. AMC0521) and 1:5000 diluted goat HRP-conjugated anti-mouse IgG (BBI Life Sciences, D110087).
Immunofluorescence imaging was performed following the published protocol56. HEK293T cells were grown on coverslips and transfected with MmSVCT1 variants. After 24 h, the cells were fixed with 3.7% formaldehyde in PBS buffer. DNA was stained with 4′,6-diamidino-2-phenylindole (DAPI) from Sigma. Images were collected via the FITC channel (for the GFP signal) or DAPI channel on a DeltaVision deconvolution microscope (GE Healthcare). Data were captured and processed using DeltaVision softWoRx software V6.5.2 (Applied Precision).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support this study are available from the corresponding authors upon request. Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-34094 (Vitamin C-bound MmSVCT1) and EMD-34095 (apo MmSVCT1). Atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 7YTW (Vitamin C-bound MmSVCT1) and 7YTY (apo mMSVCT1). The source data underlying Figs. 1a, 2c and 3c and Supplementary Figs. 11a and 15 are provided as a Source Data file. The sequence of mouse SVCT1 is available through NCBI accession NP_035527.3. Source Data are provided with this paper.
References
Peterkofsky, B. Ascorbate requirement for hydroxylation and secretion of procollagen: relationship to inhibition of collagen synthesis in scurvy. Am. J. Clin. Nutr. 54, 1135S–1140S (1991).
Pihlajaniemi, T., Myllyla, R. & Kivirikko, K. I. Prolyl 4-hydroxylase and its role in collagen synthesis. J. Hepatol. 13, S2–S7 (1991).
Englard, S. & Seifter, S. The biochemical functions of ascorbic acid. Annu. Rev. Nutr. 6, 365–406 (1986).
Eipper, B. A., Stoffers, D. A. & Mains, R. E. The biosynthesis of neuropeptides: peptide alpha-amidation. Annu. Rev. Neurosci. 15, 57–85 (1992).
Young, J. I., Zuchner, S. & Wang, G. Regulation of the epigenome by vitamin C. Annu. Rev. Nutr. 35, 545–564 (2015).
Schofield, C. J. & Ratcliffe, P. J. Oxygen sensing by HIF hydroxylases. Nat. Rev. Mol. Cell Biol. 5, 343–354 (2004).
Rose, R. C. & Bode, A. M. Biology of free radical scavengers: an evaluation of ascorbate. FASEB J. 7, 1135–1142 (1993).
Ang, A., Pullar, J. M., Currie, M. J. & Vissers, M. C. M. Vitamin C and immune cell function in inflammation and cancer. Biochem. Soc. Trans. 46, 1147–1159 (2018).
Magiorkinis, E., Beloukas, A. & Diamantis, A. Scurvy: past, present and future. Eur. J. Intern. Med. 22, 147–152 (2011).
Burzle, M. et al. The sodium-dependent ascorbic acid transporter family SLC23. Mol. Aspects Med. 34, 436–454 (2013).
Nishikimi, M., Fukuyama, R., Minoshima, S., Shimizu, N. & Yagi, K. Cloning and chromosomal mapping of the human nonfunctional gene for L-gulono-gamma-lactone oxidase, the enzyme for L-ascorbic acid biosynthesis missing in man. J. Biol. Chem. 269, 13685–13688 (1994).
Rumsey, S. C. et al. Dehydroascorbic acid transport by GLUT4 in Xenopus oocytes and isolated rat adipocytes. J. Biol. Chem. 275, 28246–28253 (2000).
Rumsey, S. C. et al. Glucose transporter isoforms GLUT1 and GLUT3 transport dehydroascorbic acid. J. Biol. Chem. 272, 18982–18989 (1997).
Wilson, J. X. Regulation of vitamin C transport. Annu. Rev. Nutr. 25, 105–125 (2005).
Tsukaguchi, H. et al. A family of mammalian Na+-dependent L-ascorbic acid transporters. Nature 399, 70–75 (1999).
Daruwala, R., Song, J., Koh, W. S., Rumsey, S. C. & Levine, M. Cloning and functional characterization of the human sodium-dependent vitamin C transporters hSVCT1 and hSVCT2. FEBS Lett. 460, 480–484 (1999).
Godoy, A. et al. Mechanistic insights and functional determinants of the transport cycle of the ascorbic acid transporter SVCT2. Activation by sodium and absolute dependence on bivalent cations. J. Biol. Chem. 282, 615–624 (2007).
Mackenzie, B., Illing, A. C. & Hediger, M. A. Transport model of the human Na+-coupled L-ascorbic acid (vitamin C) transporter SVCT1. Am. J. Physiol. Cell Physiol. 294, C451–C459 (2008).
Rajan, D. P. et al. Human placental sodium-dependent vitamin C transporter (SVCT2): molecular cloning and transport function. Biochem. Biophys. Res. Commun. 262, 762–768 (1999).
Wang, H. et al. Human Na(+)-dependent vitamin C transporter 1 (hSVCT1): primary structure, functional characteristics and evidence for a non-functional splice variant. Biochim. Biophys. Acta 1461, 1–9 (1999).
Wang, Y. et al. Human vitamin C (L-ascorbic acid) transporter SVCT1. Biochem. Biophys. Res. Commun. 267, 488–494 (2000).
Takanaga, H., Mackenzie, B. & Hediger, M. A. Sodium-dependent ascorbic acid transporter family SLC23. Pflugers Arch. 447, 677–682 (2004).
Corpe, C. P. et al. Vitamin C transporter Slc23a1 links renal reabsorption, vitamin C tissue accumulation, and perinatal survival in mice. J. Clin. Invest. 120, 1069–1083 (2010).
Sotiriou, S. et al. Ascorbic-acid transporter Slc23a1 is essential for vitamin C transport into the brain and for perinatal survival. Nat. Med. 8, 514–517 (2002).
Gournas, C., Papageorgiou, I. & Diallinas, G. The nucleobase-ascorbate transporter (NAT) family: genomics, evolution, structure-function relationships and physiological role. Mol. Biosyst. 4, 404–416 (2008).
Faaland, C. A. et al. Molecular characterization of two novel transporters from human and mouse kidney and from LLC-PK1 cells reveals a novel conserved family that is homologous to bacterial and Aspergillus nucleobase transporters. Biochim. Biophys. Acta 1442, 353–360 (1998).
Koukaki, M. et al. The nucleobase-ascorbate transporter (NAT) signature motif in UapA defines the function of the purine translocation pathway. J. Mol. Biol. 350, 499–513 (2005).
Lu, F. et al. Structure and mechanism of the uracil transporter UraA. Nature 472, 243–246 (2011).
Yu, X. et al. Dimeric structure of the uracil:proton symporter UraA provides mechanistic insights into the SLC4/23/26 transporters. Cell Res. 27, 1020–1033 (2017).
Alguel, Y. et al. Structure of eukaryotic purine/H(+) symporter UapA suggests a role for homodimerization in transport activity. Nat. Commun. 7, 11336 (2016).
Manfredini, S. et al. Design, synthesis and activity of ascorbic acid prodrugs of nipecotic, kynurenic and diclophenamic acids, liable to increase neurotropic activity. J. Med. Chem. 45, 559–562 (2002).
Pyle, E. et al. Structural lipids enable the formation of functional oligomers of the eukaryotic purine symporter UapA. Cell Chem. Biol. 25, 840–848.e844 (2018).
Dimakis, D., Pyrris, Y. & Diallinas, G. Transmembrane helices 5 and 12 control transport dynamics, substrate affinity, and specificity in the elevator-type UapA transporter. Genetics 222, iyac107. https://doi.org/10.1093/genetics/iyac107 (2022).
Kosti, V., Lambrinidis, G., Myrianthopoulos, V., Diallinas, G. & Mikros, E. Identification of the substrate recognition and transport pathway in a eukaryotic member of the nucleobase-ascorbate transporter (NAT) family. PLoS One 7, e41939 (2012).
Kourkoulou, A., Pittis, A. A. & Diallinas, G. Evolution of substrate specificity in the Nucleobase-Ascorbate Transporter (NAT) protein family. Microb. Cell 5, 280–292 (2018).
Kourkoulou, A., Zantza, I., Foti, K., Mikros, E. & Diallinas, G. Context-dependent cryptic roles of specific residues in substrate selectivity of the UapA purine transporter. J. Mol. Biol. 433, 166814 (2021).
Luo, P. et al. Crystal structure of a phosphorylation-coupled vitamin C transporter. Nat. Struct. Mol. Biol. 22, 238–241 (2015).
Boudker, O., Ryan, R. M., Yernool, D., Shimamoto, K. & Gouaux, E. Coupling substrate and ion binding to extracellular gate of a sodium-dependent aspartate transporter. Nature 445, 387–393 (2007).
Diallinas, G. Transporter specificity: a tale of loosened elevator-sliding. Trends Biochem. Sci. 46, 708–717 (2021).
Arakawa, T. et al. Crystal structure of the anion exchanger domain of human erythrocyte band 3. Science 350, 680–684 (2015).
Thurtle-Schmidt, B. H. & Stroud, R. M. Structure of Bor1 supports an elevator transport mechanism for SLC4 anion exchangers. Proc. Natl Acad. Sci. USA. 113, 10542–10546 (2016).
Walter, J. D., Sawicka, M. & Dutzler, R. Cryo-EM structures and functional characterization of murine Slc26a9 reveal mechanism of uncoupled chloride transport. Elife 8, e46986. https://doi.org/10.7554/eLife.46986 (2019).
Chi, X. et al. Structural insights into the gating mechanism of human SLC26A9 mediated by its C-terminal sequence. Cell Discov. 6, 55 (2020).
Wang, L., Chen, K. & Zhou, M. Structure and function of an Arabidopsis thaliana sulfate transporter. Nat. Commun. 12, 4455 (2021).
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D Struct. Biol. 74, 814–840 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Schrodinger, LLC. The PyMOL Molecular Graphics System, Version 1.8 (2015).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Pintilie, G. et al. Measurement of atom resolvability in cryo-EM maps with Q-scores. Nat. Methods 17, 328–334 (2020).
Song, X. et al. Dynamic crotonylation of EB1 by TIP60 ensures accurate spindle positioning in mitosis. Nat. Chem. Biol. 17, 1314–1323 (2021).
Acknowledgements
We thank the Cryo-EM Center at the University of Science and Technology of China for the support of cryo-EM data collection. We thank Dr Yongxiang Gao for assistance with cryo-EM data screening and collection. We thank Dr. Chunlei Cang and Dr. Li He at the University of Science and Technology of China for their help and suggestions. This work was supported by the Ministry of Science and Technology of China (2021YFF0600801 to J.S., 2022YFC2303700 to K.Z. and S.L., and 2022YFA1302700 to K.Z.), the National Natural Science Foundation of China (32071201 to J.S. and 32100967 to M.W.), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0490000 to K.Z.), Center for Advanced Interdisciplinary Science and Biomedicine of IHM (QYPY20220009 to J.S. and QYPY20220019 to K.Z.), start-up funding from the University of Science and Technology of China (KY9100000032, KY9100000056, and KJ2070000080 to K.Z.), and the Fundamental Research Funds for the Central Universities (WK2070000183 to J.S. and WK9100000044 to K.Z.).
Author information
Authors and Affiliations
Contributions
M.W., K.Z., and J.S. conceived the project. M.W., J.H., and Q.C. prepared the samples. M.W., J.H., and K.Z. performed data acquisition. M.W., J.H., S.L., K.Z., and J.S. performed image processing and structure determination. K.Z. and J.S. supervised the project and revised the manuscript. All authors participated in the research design, data analysis and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks George Diallinas and Ming Zhou for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, M., He, J., Li, S. et al. Structural basis of vitamin C recognition and transport by mammalian SVCT1 transporter. Nat Commun 14, 1361 (2023). https://doi.org/10.1038/s41467-023-37037-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-37037-3
- Springer Nature Limited
This article is cited by
-
Structures and mechanisms of the Arabidopsis cytokinin transporter AZG1
Nature Plants (2024)
-
Dimeric transport mechanism of human vitamin C transporter SVCT1
Nature Communications (2024)
-
Interdomain-linkers control conformational transitions in the SLC23 elevator transporter UraA
Nature Communications (2024)