Abstract
Lung adenocarcinoma (LUAD) is a prevalent form of non-small cell lung cancer with a rising incidence in recent years. Understanding the mutation characteristics of LUAD is crucial for effective treatment and prediction of this disease. Among the various mutations observed in LUAD, KRAS mutations are particularly common. Different subtypes of KRAS mutations can activate the Ras signaling pathway to varying degrees, potentially influencing the pathogenesis and prognosis of LUAD. This study aims to investigate the relationship between different KRAS mutation subtypes and the pathogenesis and prognosis of LUAD. A total of 63 clinical samples of LUAD were collected for this study. The samples were analyzed using targeted gene sequencing panels to obtain sequencing data. To complement the dataset, additional clinical and sequencing data were obtained from TCGA and MSK. The analysis revealed significantly higher Ki67 immunohistochemical scores in patients with missense mutations compared to controls. Moreover, the expression level of KRAS was found to be significantly correlated with Ki67 expression. Enrichment analysis indicated that KRAS missense mutations activated the SWEET_LUNG_CANCER_KRAS_DN and CREIGHTON_ENDOCRINE_THERAPY_RESISTANCE_2 pathways. Additionally, patients with KRAS missense mutations and high Ki67 IHC scores exhibited significantly higher tumor mutational burden levels compared to other groups, which suggests they are more likely to be responsive to ICIs. Based on the data from MSK and TCGA, it was observed that patients with KRAS missense mutations had shorter survival compared to controls, and Ki67 expression level could more accurately predict patient prognosis. In conclusion, when utilizing KRAS mutations as biomarkers for the treatment and prediction of LUAD, it is important to consider the specific KRAS mutant subtypes and Ki67 expression levels. These findings contribute to a better understanding of LUAD and have implications for personalized therapeutic approaches in the management of this disease.
Similar content being viewed by others
Introduction
The prognosis of Lung adenocarcinoma (LUAD) has greatly improved with advancements in molecular subtyping and corresponding treatment methods1,2,3. However, despite advancements, LUAD continues to be a leading cause of cancer-related mortality globally4,5,6,7,8, primarily attributable to its high incidence of recurrence and metastasis9,10. The Ki67 index in clinical immunohistochemistry is a measure of the growth and proliferation rate of tumor cells. In the context of tumor biology, it is widely acknowledged that a positive correlation exists between the Ki67 index and tumor growth and proliferation rates. Specifically, tumors with a high Ki67 index are typically associated with a high degree of malignancy, which translates into greater aggressiveness and a higher likelihood of distant metastasis. This relationship has been recognized by the World Health Organization (WHO), which in its 5th edition classification, proposed the term “carcinoid tumors with elevated mitotic counts and/or Ki67 proliferation rates” to describe tumors with high Ki67 indices11. Numerous studies have recommended the utilization of Ki67 immunohistochemistry for diagnostic testing of bronchial or lung biopsies12 making Ki67 a critical indicator for tumor diagnosis and pathological grading. In clinical practice, patients with a high Ki67 index are usually associated with a poor prognosis13,14.
Mutations in various genes have been found to be associated with the initiation, development, and metastasis of diverse types of tumors15,16. Among the identified mutations, the RAS family of genes is the most common, with KRAS being the most representative17. An activating KRAS mutation was discovered in human lung cancer samples in 1984, which were absent in the corresponding normal tissue. Several studies on human cell lines have confirmed this finding. Furthermore, it has been observed that somatic KRAS mutations are frequently present in lung cancers18. KRAS encodes a small GTPase membrane-bound protein that exists in two distinct forms: a GDP-bound form in the inactive state and a GTP-bound form in the activated state. Normally, when KRAS binds to GTP, the last phosphate group of GTP is cleaved, converting it to GDP, which inactivates it. However, when the KRAS gene is mutated, the KRAS protein remains activated by binding to GTP, and it no longer depends on the stimulation of superior signals. This results in hyperactive downstream signaling pathways that initiate and maintain hyperactive signaling in tumor genesis19,20,21. It is worth noting that KRAS-activating mutations are not universally carcinogenic in the absence of other interference. Different KRAS mutations may lead to different activation degrees of the Ras signaling pathway, which may be correlative with pathogenesis and prognosis22,23. However, the relationship between different subtypes of KRAS mutations and the pathogenesis and prognosis of LUAD remains unclear.
The advent of immune checkpoint inhibitors (ICIs) has revolutionized tumor therapy. Tumor cells have the ability to evade the immune system by activating immune checkpoint pathways that suppress anti-tumor immune responses. The principle behind the use of ICI is to block the co-inhibitory signaling pathway, thereby promoting the immune system to clear tumor cells24. Currently, there are no effective targeted therapies for KRAS mutations25. Fortunately, studies have shown that patients with KRAS mutations could benefit from ICI26,27. Therefore, KRAS mutation status may represent a potential biomarker of clinical benefit from ICI in LUAD28. However, it is worth noting that the heterogeneity of KRAS mutation may be a key consideration in the treatment efficacy of ICI in LUAD29. A common molecular lesion is a somatic missense KRAS mutation that introduces amino acid substitutions at positions 12, 13, and 6130. The mutation typically occurs in exon 2 at codon 12, and is rarely found at exon 3 codon 6121,31. Although the development of ICI has changed the treatment prospects of LUAD, but the therapeutic effects are not ideal32. Given that different KRAS mutations vary in both form and function, identifying specific KRAS mutations that are responsive to ICIs is a crucial aim of ongoing research.
The tumor mutation burden (TMB) refers to the overall count of detected somatic gene coding errors, base substitutions, gene insertions, or deletions per one million bases33. Tumor cells with a high frequency of gene mutations often display an abundance of tumor antigens on their surfaces, making them more susceptible to attacks by the body's immune system. As a result, the tumor mutation burden (TMB) has emerged as a promising biomarker for predicting the effectiveness of immune checkpoint inhibitor (ICI) therapies. Previous research has provided evidence that TMB can serve as a predictive factor for the clinical response to ICI treatments34. In addition, we also conducted an analysis to investigate the relationship between the Ki67 immunohistochemical index and TMB.
In this study, we aim to address the critical need for accurate identification and characterization of missense mutations in the KRAS gene in LUAD. While previous research has established the significance of KRAS mutations in lung cancer development and progression, there remains a lack of clarity regarding the specific types and consequences of these mutations. Our study seeks to fill this knowledge gap by comprehensively analyzing missense mutations in the KRAS gene using two targeted next-generation sequencing (NGS) panels. By doing so, we aim to provide a comprehensive understanding of the landscape of missense mutations in KRAS and their potential implications in lung cancer. This study's novelty lies in its characterization of mutations, shedding light on their prevalence, distribution, and potential functional consequences. Ultimately, our findings have the potential to contribute to the development of targeted therapies and personalized treatment approaches for LUAD patients with KRAS missense mutations.
Materials and methods
Data preparation
This study included a cohort of 63 patients who underwent immunohistochemistry (IHC) at Hangzhou Hospital of Traditional Chinese Medicine between November 2020 and January 2022. The mutational status of 9 genes or 1059 genes was determined using next-generation sequencing (NGS). Specifically, two panels were utilized for NGS testing: the 9-gene panel and the 1059-gene panel. The latter includes the genes in the former, and detailed gene information for both panels can be found in Supplementary Table 1. For the purposes of this study, our focus was on the 9 genes that were included in both panels.
The study adhered to the ethical principles outlined in the Declaration of Helsinki (revised in 2013). Approval for this retrospective study was obtained from the medical ethics committee of Hangzhou TCM Hospital (Approval No. 2023KLL050). Given the retrospective design of the study, informed consent was not required by the medical ethics committee of Hangzhou TCM Hospital.
DNA was extracted from the case samples obtained. DNA was extracted from the tumor and the blood. The Rapid DNA FFPE Kit (ZYMO) was used for extraction from formalin-fixed paraffin-embedded (FFPE) sections. Qubit dsDNA HS Analysis Kit (LIFE) and agarose gel electrophoresis were used for DNA quantification. DNA library construction was completed using the ABclonal lone Rapid DNA Library Preparation Kit according to the manufacturer’s protocol. Covaris S220 was used to physically cut 50-200 ng DNA for A-tailing, connector ligation, and PCR amplification. The mixture was hybridized with Boke’s hybridization probe for 16 to 18 h at 65 °C, followed by capture using M270 strand avidin beads for 45 min. Then 15 cycles of post-capture amplification were performed to obtain the capture library. All samples were sequenced using two NGS testing panels that included 1059 cancer-related genes. The panel used for each sample is listed in Supplementary Table 2. Trimmomatic is used to filter low-quality reads and BWA MEM is used to map reads. GATK SortSam, MarkDuplicates, CollectHsMetrics, BaseRecalibrator, ApplyBQSR, AnalyzeCovariates, CollectSequencingArtifactMetrics, Mutect2, GetPileupSummaries, CalculateContamination, FilterMutectCalls, FilterByOrientationBias is used for mutation calling and filtering.
The transcriptomic data from LUAD patients were obtained from The Cancer Genome Atlas (TCGA) database. Specifically, RNA-seq data was available for 496 patients with LUAD from the TCGA portal (https://portal.gdc.cancer.gov/). In addition, MSK data of NSCLC patients on TMB results and survival data were directly downloaded from the cBioPortal for Cancer Genomics (https://www.cbioportal.org/study/summary?id=msk_impact_2017). These data sources were chosen for their comprehensive and reliable collection of genomic and clinical data of patients with lung cancer, which enabled us to conduct a thorough analysis of the biomarkers of interest in our study.
Local clinical cohorts
Clinical patient information included gender, age, smoking history, pathological stage, and immunohistochemical scores of Ki67, NapsinA, TTF, EGFR, and P53. Detailed clinical data can be found in Supplementary Table 2. Pathological staging was evaluated by experienced pathologists using the TNM staging system. Staging and pathological diagnosis were manually organized based on pathological reports. The ggplot2 package in R is used to draw bar charts and violin charts.
Immunohistochemistry (IHC) analysis of biomarkers
IHC was performed to analyze the expression of specific biomarkers TTF, p53, EGFR, NapsinA, and Ki67 in lung cancer tissue samples. Briefly, tissue sections were deparaffinized, rehydrated, and subjected to antigen retrieval. Endogenous peroxidase activity was blocked with hydrogen peroxide, and non-specific binding was blocked with serum. The tissue sections were then incubated with primary antibodies against the biomarkers of interest, followed by secondary antibodies conjugated with horseradish peroxidase. The IHC staining was visualized with diaminobenzidine (DAB) and counterstained with hematoxylin. Images were captured using a microscope equipped with a digital camera. The binary variables of conventional IHC biomarkers used for statistical analysis were as follows: positive or negative for TTF, p53, EGFR, and NapsinA; proliferation index of Ki67. The Ki67 index was determined by counting at least 200 cells within the area of highest Ki67 expression.
Division of missense and non-missense mutations
We partitioned the samples into two distinct cohorts, distinguishing between those harboring KRAS missense mutations and those with non-missense mutations, with the aim of ascertaining the utility of KRAS missense mutations as potential biomarkers for ICI therapy. A missense mutation is characterized by the alteration of a given codon, such that it results in the substitution of the original codon with a novel one, encoding a different amino acid and consequently leading to a modification in the amino acid sequence. In this study, we considered all mutations that bring about changes in amino acids, ascertained through the Genome Analysis Toolkit (GATK) analysis, as falling under the category of missense mutations. It is noteworthy that KRAS mutations predominantly manifest as single-base missense mutations, with a significant proportion clustering within three well-defined hotspots located at codon 12, 13, and 6135,36. In the NGS panels, probes are designed specifically targeting these KRAS missense mutation sites. In both the TCGA and MSK datasets, the original missense mutation labels obtained from the database are utilized. Non-missense mutations, on the other hand, pertain to the samples that fall outside the category of missense mutations.
mRNA expression profile and enrichment analysis
Correlation analysis of gene expression was conducted using the ggplot2 package in R. Differential gene analyses were performed using the DESeq2 package to associate gene signatures with KRAS missense or the control group. Gene sets from the MsigDB database were obtained using the msigdbr package in R. Enrichment functional pathway maps were generated using the ClusterProfiler package. Heat maps were created using the pheatmap package.
TMB and survival analysis
The local clinical data TMB was defined as the ratio of the number of somatic nonsynonymous variants in the coding regions, including indels and deletions, detected in tumor tissue to the size of the region that was analyzed. TMB results from the TCGA data were calculated using the maftools package. TMB results and survival data from the MSK database were downloaded directly. The IHC score of Ki67 and the expression of Ki67 were classified based on the median value. Survival analysis and visualization were performed using the Survminer package in R.
Statistical analysis
Data analysis was conducted using R version 3.6.0. The Wilcoxon test was employed to compare continuous variables, including gene expression values, IHC Ki67 scores, and TMB. Kaplan-Meier survival curves were generated and the log-rank test was performed using the Survminer package in R. Significance tests in this study were two-sided, with a p-value <0.05 indicating statistical significance. A p-value <0.01 was deemed marginally significant.
Result
Clinical data showed that age and pathological stage were correlated with the IHC Ki67 score
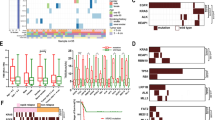
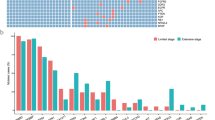
Ki67 is a proliferating cell-associated antigen whose function is closely related to mitosis. It is indispensable in cell proliferation, so the Ki67 proliferation index is an important indicator of immunohistochemistry in pathology. We collected clinical information from 63 NGS samples (including 38 9-gene samples and 25 1059-gene samples) and analyzed these samples. Clinical information included gender, age, smoking history, pathological stage, and immunohistochemical results of the samples (Fig. 1a). The results of clinical information analysis showed that the Ki67 immunohistochemical index of people over 50 years old was significantly higher than that of people under 50 years old (P = 0.0094, Wilcox test) (Fig. 1b). Patients with Tis stage were significantly lower than patients with stage I (P = 0.01, Wilcox test) and patients with stage II-III (P = 0.025, Wilcox test), and patients with stage I were significantly lower than patients with stage II-III (P=0.034, Wilcox test)) (Fig. 1c). There was no significant difference between the sexes in the Ki67 index (P=0.21, Wilcox test) (Fig. 1d).
There were significant differences in Ki67 scores between KRAS missense and non-missense mutations
At the same time, 63 clinical patient samples with 9 shared gene tests and Ki67 IHC indicators were analyzed, and the relationship between samples with different gene mutation types and Ki67 IHC index was analyzed. The results showed that the Ki67 immunohistochemical index of the KRAS gene missense mutation samples was significantly higher than that of the KRAS non-missense mutation samples index (P = 0.047, Wilcox test), while other genes were not statistically different (Fig. 2a). Among the 63 patients with LUAD with Ki67 immunohistochemical index, the proportion of patients with Ki67 immunohistochemical index (≥ median) in the KRAS gene missense mutation group was higher than that in the non-missense mutation group (18/25 vs 18/38) (Fig. 2b). In the TCGA data, the expression level of Ki67 (MKi67) in KRAS gene missense mutation samples was significantly higher than that in KRAS gene non-missense mutation samples (P=0.014, Wilcoxon test) (Fig. 2c).
The relationship between different gene mutation types and Ki67 IHC score. (a) Distribution of Ki67 IHC scores for different mutation types of each gene. (b) Proportion of clinical patients with different KRAS mutation types in the Ki67 IHC score interval. (c) Distribution of Ki67 expression corresponding to different KRAS mutation types in TCGA data.
TCGA data correlation analysis showed that KRAS expression was significantly positively correlated with Ki67 expression. Enrichment analysis showed that both KRAS and Ki67 activated SWEET LUNG CANCER KRAS DN and CREIGHTON ENDOCRINE THERAPY RESISTANCE 2 pathways
We performed a correlation analysis on the expression of KRAS and Ki67 (MKi67) in the TCGA data, and the results showed that they had a statistically significant positive correlation at the RNA level (r = 0.33, P < 0.0001, Spearman) (Fig. 3a). To further explore the difference between KRAS missense and non-missense mutation subtypes, we performed GSEA based on RNA-seq data from TCGA, which revealed a prominent enrichment in KRAS-missense-mutant group of signatures related to SWEET LUNG CANCER KRAS DN (P < 0.0001) and CREIGHTON ENDOCRINE THERAPY RESISTANCE 2 (P< 0.0001) (Fig. 3b–d). We generated heat maps to depict the distribution of RNA levels in the corresponding pathway between the KRAS missense-mutant and non-missense-mutant groups (Fig. 3e,f).
GSEA analysis of mRNA relative expression levels of signal pathways enriched in TCGA dataset. (a) Correlation analysis results of KRAS gene expression and Ki67 gene expression in TCGA dataset. (b) Enrichment analysis showed that SWEET LUNG CANCER KRAS DN and CREIGHTON ENDOCRINE THERAPY RESISTANCE 2 pathways were activated. (c,d) enrichment plot regarding two pathways. (e,f)Heat map of relative mRNA expression of the two pathways.
KRAS missense mutation and Ki67 expression were significantly increased at the TMB level compared with the other groups
Samples with different KRAS mutations and Ki67 expression in clinical data, TCGA data, and MSK data were genotyped. Clinical data showed that the KRAS non-missense mutation and Ki67 IHC low score group were significantly lower than the other groups (P < 0.05, Wilcox test) (Fig. 4a). Also, TCGA data showed that KRAS non-missense mutation and Ki67 IHC low score group were significantly lower than other groups (P < 0.05, Wilcox test). The results also showed that KRAS missense mutation and Ki67 low IHC score group were significantly lower than KRAS missense mutation and Ki67 high IHC score group (P = 0.02, Wilcox test) (Fig. 4b). In MSK data, the TMB of KRAS missense mutation samples was significantly higher than that of KRAS non-missense mutation samples (P < 0.0001, Wilcox test) (Fig. 4c).
Distribution of TMB results in different data sets with different KRAS mutation types. (a) TMB distribution of different KRAS mutation types in the clinical dataset. (b) TMB distribution of different KRAS mutation types in the TCGA dataset. (c) TMB distribution of different KRAS mutation types in the MSK dataset.
Survival analysis showed that the survival time of KRAS missense mutant samples was longer than that of non-missense mutant samples, and the survival time of Ki67 high-expression samples was longer than that of low-expression samples
Survival analysis can study the influence of different traits on the survival time of patients. Survival analysis is performed on MSK and TCGA data. MSK data showed that the survival time of KRAS missense mutant samples was significantly longer than that of non-missense mutant samples (P = 0.045, Wilcox test) (Fig. 5a). TCGA data showed that the survival time of KRAS missense mutant samples was longer than that of non-missense mutant samples (Fig. 5b), and the survival time of Ki67 high-expression samples was longer than that of low-expression samples (P = 0.0044, Wilcox test) (Fig. 5c).
Discussion
The Ki67 marker, which is related to tumor cell proliferation, has been found to be associated with the progression, metastasis, and prognosis of LUAD37. In our analysis of clinical patient information, it was observed that age and pathological stage were significantly correlated with the Ki67 IHC index, which is consistent with previous studies. However, gender was not found to be correlated with the Ki67 IHC index, possibly because the pathogenesis of LUAD is not solely related to smoking. Further research is necessary to explore this issue by collecting more samples and life information.
The utilization of ICIs have revolutionized cancer treatment by enhancing the immune system to target tumor cells38. The function of ICI is to disrupt the inhibitory signals between tumor cells and immune effector cells, allowing activated T cells to target these tumor cells39. Previous studies have shown that gene mutations in KRAS, FGFR, RET, BRAF, and HER2 have an impact on clinical benefits of ICI in advanced non-small cell lung cancer40. Although previous studies have found that KRAS mutations impact the treatment of ICI, they did not consider the heterogeneity of KRAS41. Therefore, various theories exist regarding the evaluation of KRAS mutations as ICI biomarkers. In this study, the samples were divided into KRAS missense and non-missense mutations. To ensure comprehensive analysis, data sets from three different sources were used. The Ki67 IHC index, an important indicator of immunohistochemistry, also showed a difference between KRAS missense and non-missense mutations in the study.
Among the 63 cases of LUAD patients, a significant difference in the Ki67 immunohistochemical index was observed only between samples with KRAS gene missense mutations and non-missense mutations. Notably, a higher proportion of patients in the missense mutation group exhibited a Ki67 immunohistochemical index equal to or greater than the median value. These findings underscore the distinctive nature of KRAS mutations compared to other mutation types, as well as variations within different forms of KRAS mutations. It is important to note that our study focused solely on investigating differences within the mutant forms of KRAS itself and did not consider the potential interactions between KRAS and other mutations. Further studies are warranted to explore these interactions in greater detail.
Additionally, we conducted an analysis using the TCGA dataset. The results revealed that the expression of Ki67 in samples with KRAS gene missense mutations was significantly higher compared to samples with non-missense mutations. This finding supports the notion that different expression types of the KRAS gene exhibit distinct characteristics, aligning with previous research. Notably, there was a significant positive correlation between KRAS gene expression and Ki67 gene expression. This suggests that elevated expression of both genes may be associated with the pathogenesis of lung adenocarcinoma. Furthermore, enrichment analysis highlighted the co-activation of the SWEET LUNG CANCER KRAS DN pathway by KRAS and Ki67, providing further confirmation of our observations. In the future, it might interesting to predict drugs can target both KRAS mutation and Ki67 using some machine leanring42,43,44 and experimental methods45.
Survival analysis was conducted using the MSK and TCGA data.The results revealed that KRAS missense mutant samples had longer survival times than the control samples, while samples with high expression of Ki67 had longer survival times compared to those with low expression.This suggests that Ki67 can serve as an indicator of KRAS activation, thus enhancing the accuracy of survival time prediction.
Furthermore, the oncogenic KRAS gene mutation not only directly facilitates the proliferation and survival of tumor cells but also exerts an impact on the tumor microenvironment. It has been observed by researchers that the combination of a KRAS G12C inhibitor and an immune checkpoint inhibitor (ICI) can effectively suppress certain types of immunogenic lung cancer46. The efficacy of the combination therapy might be related to the TMB. TMB is indicative of the overall number of mutations present in tumor cells. A higher frequency of gene mutations in tumor cells results in an increased expression of tumor antigens on the cell surface. Consequently, tumor cells become more susceptible to immune system-mediated attack, thereby enhancing the effectiveness of immune checkpoint inhibitors (ICI)47. Different KRAS mutations may have different effects on TMB. We compared the TMB values of clinical data, TCGA data, and MSK data for samples with different KRAS genotypes. The high expression and low expression of Ki67 obtained by KRAS missense mutation and non-missense mutation and median differentiation resulted in four groups of TMB: Kras_missense-ki67_high, KRAS_MISsense-Ki67_low, KRAS_NONMISSENse-Ki67_high, KRAS_NONMISsense-Ki67_low. The results showed that TMB in the KRAS_nonmissense-Ki67_LOW group was significantly lower than that in the other three groups, and there was no statistical difference between the other three groups. The TCGA data were consistent with the clinical data, revealing that the TMB in the KRAS_MISsense-Ki67_low group was significantly lower compared to the KRAS_MISsense-Ki67_HIGH group. In the MSK data, the TMB of samples with KRAS missense mutations was significantly higher than that of samples with KRAS non-missense mutations. In all three datasets, it was consistently observed that the TMB of samples with KRAS missense mutations was significantly higher than those without missense mutations. Additionally, the TMB was found to be higher in samples with high Ki67 expression compared to those with low Ki67 expression. These findings indicate that KRAS gene missense mutations and high Ki67 expression contribute to an increased TMB level, which can potentially serve as a predictive marker for the efficacy of immunotherapy.
Our study demonstrates a significant correlation between KRAS mutant subtypes, Ki67 expression, and increased TMB levels. However, further investigations are necessary to elucidate the underlying mechanisms.Secondly, our study utilized bulk sequencing, disregarding cell heterogeneity48,49. Conducting a single-cell resolution study would be more appropriate for a comprehensive investigation of the mechanisms underlying KRAS mutation, Ki67, and TMB.Additionally, the presence of differences among various databases is inevitable. Nonetheless, the consistent results across different databases lend credibility to our findings. Furthermore, the clinical dataset we used included a small sample size, rendering the data statistically insignificant. Hence, additional data is required for further investigation.
Conclusion
In conclusion, our study revealed significant variations in immunohistochemical Ki67 scores among different KRAS variants. Furthermore, we observed a positive correlation between KRAS expression and immunohistochemical Ki67 scores. Interestingly, Ki67 expression proved to be a more effective prognostic indicator than KRAS mutation status. Additionally, our data demonstrated a link between KRAS missense mutations, higher Ki67 scores, and increased TMB. This finding suggests a potential impact on the efficacy of subsequent immunotherapy. Moreover, integrating Ki67 scores and KRAS mutation type improved the accuracy of survival prediction.
Data availability
TGCA and MSK data are publicly avaiable from https://portal.gdc.cancer.gov/ and https://www.cbioportal.org/study/summary?id=msk_impact_2017. The variant data for this study have been deposited in the Genome Variation Map (GVM) in National Genomics Data Center, Beijing Institute of Genomics, Chinese Academy of Sciences and China National Center for Bioinformation, under accession number GVM000561. (https://bigd.big.ac.cn/gvm/getProjectDetail?Project=GVM000561). The data generated during the current study available from the corresponding author on reasonable request.
References
Guo, L. et al. Constructing molecular classifiers for the accurate prognosis of lung adenocarcinoma. Clin. Cancer Res. 12(11 Pt 1), 3344–3354 (2006).
Chen, X. et al. The Cdc2/Cdk1 inhibitor, purvalanol A, enhances the cytotoxic effects of taxol through Op18/stathmin in non-small cell lung cancer cells in vitro. Int. J. Mol. Med. 40(1), 235–242 (2017).
Yuan, X. et al. Bacterial biomarkers capable of identifying recurrence or metastasis carry disease severity information for lung cancer. Front. Microbiol. 13, 1007831 (2022).
Ushio, R., Murakami, S. & Saito, H. Predictive markers for immune checkpoint inhibitors in non-small cell lung cancer. J. Clin. Med. 11(7), 1855 (2022).
Thandra, K. C., Barsouk, A., Saginala, K., Aluru, J. S. & Barsouk, A. Epidemiology of lung cancer. Contemp. Oncol. (Pozn) 25(1), 45–52 (2021).
Song, Z. et al. Evaluating the potential of T cell receptor repertoires in predicting the prognosis of resectable non-small cell lung cancers. Mol. Ther. Methods Clin. Dev. 18, 73–83 (2020).
Zhao, Y. et al. In vitro neutralization of autocrine IL10 affects Op18/stathmin signaling in nonsmall cell lung cancer cells. Oncol. Rep. 41(1), 501–511 (2019).
Chen, S. et al. Introduction of exogenous wildtype p53 mediates the regulation of oncoprotein 18/stathmin signaling via nuclear factorkappaB in nonsmall cell lung cancer NCIH1299 cells. Oncol. Rep. 41(3), 2051–2059 (2019).
Liu, H. et al. Evaluating DNA methylation, gene expression, somatic mutation, and their combinations in inferring tumor tissue-of-origin. Front. Cell. Dev. Biol. 9, 619330 (2021).
He, B. et al. TOOme: A novel computational framework to infer cancer tissue-of-origin by integrating both gene mutation and expression. Front. Bioeng. Biotechnol. 8, 394 (2020).
WHO Classification of Tumours Editorial Board. Thoracic Tumours Vol. 5 (International Agency for Research on Cancer, 2021).
La Rosa, S. Diagnostic, prognostic, and predictive role of Ki67 proliferative index in neuroendocrine and endocrine neoplasms: Past, present, and future. Endocr. Pathol. 34(1), 79–97 (2023).
Yao, W. et al. Noninvasive method for predicting the expression of Ki67 and prognosis in non-small-cell lung cancer patients: Radiomics. J. Healthc. Eng. 2022, 7761589 (2022).
Soomro, I. N., Holmes, J. & Whimster, W. F. Predicting prognosis in lung cancer: Use of proliferation marker, Ki67 monoclonal antibody. JPMA J. Pak. Med. Assoc. 48(3), 66–69 (1998).
Shi, X., Young, S., Cai, K., Yang, J. & Morahan, G. Cancer susceptibility genes: Update and systematic perspectives. Innovation (Camb) 3(5), 100277 (2022).
World Health Organization. Regional Office for South-East A: Integrated regional action plan for viral hepatitis, HIV and sexually transmitted infections in South-East Asia; 2022–2026. New Delhi: World Health Organization. Regional Office for South-East Asia (2022).
Westcott, P. & To, M. The genetics and biology of KRAS in lung cancer. Chin. J. Cancer 2, 8 (2013).
Riely, G. J., Marks, J. & Pao, W. KRAS mutations in non-small cell lung cancer. Proc. Am. Thorac. Soc. 6(2), 201–205 (2009).
Karachaliou, N. et al. KRAS mutations in lung cancer. Clin. Lung Cancer 14(3), 205–214 (2013).
Manchado, E. et al. A combinatorial strategy for treating KRAS-mutant lung cancer. Nature 534(7609), 647–651 (2016).
Ghimessy, A. et al. Current therapy of KRAS-mutant lung cancer. Cancer Metastasis Rev. 39(4), 1159–1177 (2020).
Johnson, C., Burkhart, D. L. & Haigis, K. M. Classification of KRAS-activating mutations and the implications for therapeutic intervention. Cancer Discov. 12(4), 913–923 (2022).
Li, S., Balmain, A. & Counter, C. M. A model for RAS mutation patterns in cancers: Finding the sweet spot. Nat. Rev. Cancer https://doi.org/10.1038/s41568-018-0076-6 (2018).
Darvin, P., Toor, S. M., Sasidharan Nair, V. & Elkord, E. Immune checkpoint inhibitors: Recent progress and potential biomarkers. Exp. Mol. Med. 50(12), 1–11 (2018).
Adderley, H., Blackhall, F. H. & Lindsay, C. R. KRAS-mutant non-small cell lung cancer: Converging small molecules and immune checkpoint inhibition. EBioMedicine 41, 711–716 (2019).
Li, Y. et al. Resistance to immune checkpoint inhibitors in KRAS-mutant non-small cell lung cancer. Cancer Drug Resist. 5(1), 129–146 (2022).
Mazieres, J. et al. Immune checkpoint inhibitors for patients with advanced lung cancer and oncogenic driver alterations: Results from the IMMUNOTARGET registry. Ann. Oncol. 30(8), 1321–1328 (2019).
Kim, J. H., Kim, H. S. & Kim, B. J. Prognostic value of KRAS mutation in advanced non-small-cell lung cancer treated with immune checkpoint inhibitors: A meta-analysis and review. Oncotarget 8(29), 48248–48252 (2017).
Uras, I. Z., Moll, H. P. & Casanova, E. Targeting KRAS mutant non-small-cell lung cancer: Past, present and future. Int. J. Mol. Sci. 21(12), 4325 (2020).
Schubbert, S. et al. Biochemical and functional characterization of germ line KRAS mutations. Mol. Cell Biol. 27(22), 7765–7770 (2007).
Timar, J. The clinical relevance of KRAS gene mutation in non-small-cell lung cancer. Curr. Opin. Oncol. 26(2), 138–144 (2014).
Li, R. et al. Inhibition of granulocytic myeloid-derived suppressor cells overcomes resistance to immune checkpoint inhibition in LKB1-deficient non-small cell lung cancer. Cancer Res. 81(12), 3295–3308 (2021).
Huang, K. et al. Predicting colorectal cancer tumor mutational burden from histopathological images and clinical information using multi-modal deep learning. Bioinformatics https://doi.org/10.1093/bioinformatics/btac641 (2022).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51(2), 202–206 (2019).
Christenson, E. S. et al. Cell-free DNA predicts prolonged response to multi-agent chemotherapy in pancreatic ductal adenocarcinoma. Cancer Res. Commun. 2(11), 1418–1425 (2022).
Prior, I. A., Lewis, P. D. & Mattos, C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 72(10), 2457–2467 (2012).
Gu, W. et al. The Ki-67 proliferation index-related nomogram to predict the response of first-line tyrosine kinase inhibitors or chemotherapy in non-small cell lung cancer patients with epidermal growth factor receptor-mutant status. Front. Med. https://doi.org/10.3389/fmed.2021.728575 (2021).
Yousefi, H., Yuan, J., Keshavarz-Fathi, M., Murphy, J. F. & Rezaei, N. Immunotherapy of cancers comes of age. Expert Rev. Clin. Immunol. 13(10), 1001–1015 (2017).
Palmieri, D. J. & Carlino, M. S. Immune checkpoint inhibitor toxicity. Curr. Oncol. Rep. 20(9), 72 (2018).
Uehara, Y., Watanabe, K., Hakozaki, T., Yomota, M. & Hosomi, Y. Efficacy of first-line immune checkpoint inhibitors in patients with advanced NSCLC with KRAS, MET, FGFR, RET, BRAF, and HER2 alterations. Thorac. Cancer 13(11), 1703–1711 (2022).
Gu, M., Xu, T. & Chang, P. KRAS/LKB1 and KRAS/TP53 co-mutations create divergent immune signatures in lung adenocarcinomas. Ther. Adv. Med. Oncol. 13, 17588359211006950 (2021).
Meng, Y. et al. A weighted bilinear neural collaborative filtering approach for drug repositioning. Brief. Bioinform. 23(2), bbab581 (2022).
Liu, C. et al. An improved anticancer drug-response prediction based on an ensemble method integrating matrix completion and ridge regression. Mol. Ther. Nucleic Acids 21, 676–686 (2020).
Liu, X. et al. A systematic study on drug-response associated genes using baseline gene expressions of the cancer cell line encyclopedia. Sci. Rep. 6, 22811 (2016).
Rubin, E. H. & Gilliland, D. G. Drug development and clinical trials–the path to an approved cancer drug. Nat. Rev. Clin. Oncol. 9(4), 215–222 (2012).
Mugarza, E.A.-O. et al. Therapeutic KRAS(G12C) inhibition drives effective interferon-mediated antitumor immunity in immunogenic lung cancers. Sci. Adv. 8(29), eabm8780 (2022).
Gao, G. et al. KRAS G12D mutation predicts lower TMB and drives immune suppression in lung adenocarcinoma. Lung Cancer 149, 41–45 (2020).
Xu, J. et al. Evaluating the performance of dropout imputation and clustering methods for single-cell RNA sequencing data. Comput. Biol. Med. 146, 105697 (2022).
Xu, J., Cai, L., Liao, B., Zhu, W. & Yang, J. CMF-Impute: An accurate imputation tool for single-cell RNA-seq data. Bioinformatics 36(10), 3139–3147 (2020).
Author information
Authors and Affiliations
Contributions
B.C., P.L. designed the study; J.W., L.D. performed the study, analyzed the data and interpreted data; Z.Z.W., Z.Z. wrote the manuscript; B.X., Y.X., X.L., reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Dong, L., Zheng, Z. et al. Effects of different KRAS mutants and Ki67 expression on diagnosis and prognosis in lung adenocarcinoma. Sci Rep 14, 4085 (2024). https://doi.org/10.1038/s41598-023-48307-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48307-x
- Springer Nature Limited
This article is cited by
-
Effect of 660-nm LED photobiomodulation on the proliferation and chondrogenesis of meniscus-derived stem cells (MeSCs)
Scientific Reports (2024)
-
Exploring the prognosis value, immune correlation, and drug responsiveness prediction of homeobox C6 (HOXC6) in lung adenocarcinoma
Discover Oncology (2024)