Abstract
There is an urgent need to develop effective antiviral drugs to prevent the viral infection caused by constantly circulating SARS-CoV-2 as well as its variants. The main protease (Mpro) of SARS-CoV-2 is a salient enzyme that plays a vital role in viral replication and serves as a fascinating therapeutic target. PF-07304814 is a covalent inhibitor targeting SARS-CoV-2 Mpro with favorable inhibition potency and drug-like properties, thus making it a promising drug candidate for the treatment of COVID-19. We previously solved the structure of PF-07304814 in complex with SARS-CoV-2 Mpro. However, the binding modes of PF-07304814 with Mpros from evolving SARS-CoV-2 variants is under-determined. In the current study, we expressed six Mpro mutants (G15S, K90R, M49I, S46F, V186F, and Y54C) that have been identified in Omicron variants including the recently emerged XBB.1.16 subvariant and solved the crystal structures of PF-07304814 bound to Mpro mutants. Structural analysis provided insight into the key molecular determinants responsible for the interaction between PF-07304814 and these mutant Mpros. Patterns for PF-07304814 to bind with these investigated Mpro mutants and the wild-type Mpro are generally similar but with some differences as revealed by detailed structural comparison. Structural insights presented in this study will inform the development of novel drugs against SARS-CoV-2 and the possible conformation changes of Mpro mutants when bound to an inhibitor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In late December of 2019, a novel coronavirus was detected in Wuhan city, situated in Hubei Province of central China [1, 2]. In February 2020, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was designated as the name of this newly emerging coronavirus by the International Committee on Taxonomy of Viruses (ICTV). SARS-CoV-2 is known as the pathogenic agent responsible for the coronavirus disease-19 (COVID-19) [3]. Being highly transmissible, SARS-CoV-2 has spread quickly to the entire world and posed a serious threat to global public health [4, 5]. As of April 26, 2023, over 764 million confirmed cases and more than 6.9 million deaths have been reported worldwide, with the numbers still increasing (https://covid19.who.int/). The impressive commitment made by the biomedical research community led to rapid development of several safe and effective vaccines [6,7,8,9], which plays an important role in reducing the rates of infection, hospitalization, and mortality. However, the continuing emergence of SARS-CoV-2 variants highlights that the battle against COVID-19 is far from over. Five variants have been characterized by the World Health Organization (WHO) as variants of concern (VOCs), namely Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Delta (B.1.617.2), and Omicron (B.1.1.529), and several variants have been designated as variants of interest (VOIs), including Lambda (C.37) and Kappa (B.1.617.1). Omicron is currently the most prevalent SARS-CoV-2 variant and has evolved into many sub-variants with high immune evasion ability, including BA.5, BF.7, BQ.1, XBB.1.5, and the newly emerged XBB.1.16 [10,11,12,13,14], which may associate with the reduced effectiveness of available vaccines. Therefore, effective and curative treatment measures are still urgently needed to combat COVID-19.
Development of small molecules that exert antiviral efficacies against SARS-CoV-2 infection by inhibiting its main protease (Mpro), a vital enzyme in the viral life cycle, continues to be a relevant trend in searching for COVID-19 treatments [15]. Mpro is able to cleave the bio-synthesized viral polyprotein 1ab (pp1ab) at 11 specific sites to produce a variety of mature non-structural proteins necessary for viral replication and transcription [16, 17]. In most cases, this enzyme has a recognition sequence of (Leu-Gln)-(Ser-Ala-Gly), with the bond between glutamine and serine being the cleavage site [18, 19]. Such an enzymatic cleavage specificity is absent in humans, which indicates the absence of toxicity of potential Mpro inhibitors [18, 19]. Thus, inhibitors targeting the main protease can effectively impede the infection of SARS-CoV-2 and represent promising antiviral drug candidates.
In the last three years, various Mpro inhibitors have been discovered by using the high-throughput screen or structure-based drug design [20]. Among these, PF-07304814, also named lufotrelvir, is a highly soluble phosphate prodrug of PF-00835231 and a first in class 3CL inhibitor to treat SARS-CoV-2 infection [21]. It is developed by Pfizer and need to be administered by the intravenous route. When administrated, PF-07304814 will be cleaved by ubiquitous human alkaline phosphatase and metabolized into its active form, namely PF-00835231 (Fig. 1), which exhibits potent antiviral efficacy against SARS-CoV-2 as well as favorable absorption, distribution, metabolism, excretion (ADME) profiles [21]. The crystal structure of SARS-CoV-2 Mpro bound to PF-00835231 has been determined [22]. Recently, our group also solved the crystal structure of SARS-CoV-2 Mpro bound to the drug candidate PF-07304814 (PDB ID 7VVP) and illustrated the molecular mechanism for interaction [23]. The carbonyl carbons of hydroxymethyl ketone (HMK) warheads of PF-00835231 and PF-07304814 form irreversible covalent bonds with the sulfur atom of Mpro active-site cysteine (Cys145). These data provide structural basis and support that PF-00835231 as well as PF-07304814 is a valuable scaffold for developing effective drugs to treat COVID-19.
To date, multiple mutations in the main proteases of emerging SARS-CoV-2 variants are identified [24, 25], which may perturb the interaction network between PF-00835231 and SARS-CoV-2 Mpro, thus affecting its potency. PF-07304814 has the similar molecular structure with its active form. Determining the interacting details between PF-07304814 and Mpro mutants will provide valuable information on drug resistance to PF-00835231 and therapeutic implication for COVID-19. In this study, we solved the crystal structures of PF-07304814 in complex with several Mpro mutants, each carrying a previously reported single amino acid substitution, and revealed the structural basis for their interactions. The results provide structural insights for understanding the possible interaction differences between PF-07304814 and various mutant Mpros, and will add to develope more effective drugs to treat viral infection caused by SARS-CoV-2 as well as its variants.
Results
Structure determination of Mpro mutants-PF-07304814 complexes
A total of six previously reported mutations (G15S, K90R, M49I, S46F, V186F, and Y54C) are included in this investigation. The occurrence rates for G15S, K90R, M49I, S46F, and V186F mutations among 15,476,050 sequenced SARS-CoV-2 Mpro genes in the Global Initiative on Sharing All Influenza Data (GISAID) (accessed on April 27, 2023) are 0.19%, 1.34%, 0.015%, 0.025%, and 0.016%, respectively, while only 13 reports are available regarding the occurrence of Y54C mutation. All these six mutations are identified in the wildly contagious Omicron variants. G15S and K90R mutations, previously dominant in Lambda and Gamma variants, respectively, are also found in the recently emerged XBB.1.16 subvariant. In order to figure out the interaction details between PF-07304814 and these six mutant Mpros, the co-crystallization method was used to determine the structures of Mpro mutants bound to PF-07304814. All Mpro mutants were expressed in E. Coli cells, purified to near homogeneity, and then incubated with excessive PF-07304814 (Fig. 1). The prepared samples were successfully crystallized and the crystal structures of PF-07304814 in complex with these Mpro mutants were solved to 2.29-Å (G15S), 1.95-Å (S46F), 2.05-Å (M49I), 1.75-Å (Y54C), 1.97-Å (K90R), and 1.70-Å (V186F) resolution, respectively. All these complex structures are in space group P1211, which is different with that (P212121) of wild-type Mpro-PF07304814 complex [23]. Data collection and refinement statistics are summarized in Table 1.
Overall structure of Mpro mutants bound to PF-07304814
In these solved complex structures, each mutant Mpro is displayed as a dimer, which is exactly the form with enzymatic activity. Like the wild-type SARS-CoV-2 Mpro, each protomer of the dimeric Mpro mutants contains three major domains (domain I-III). Domain I resides between residues 10 through 99 and forms an anti-parallel β sheet. Domain II is located between residues 100 through 184 and is composed of anti-parallel β sheet structures. Domain III starts at the residue 201 and ends at the residue 303, which is consisting of several ɑ helices and connects to domain II by a long loop region (residues 185–200). All the SARS-CoV-2 Mpro mutants bind with two PF-07304814 molecules. PF-07304814 was found to insert into the active sites of mutant Mpros (Fig. 2), which is situated in the cleft between domains I and II. We then extracted the electron density maps of the ligand PF-07304814, the catalytic residues (including His41 and Cys145), and the mutant residues. Cys145 is a nucleophilic reagent in the hydrolysis process of SARS-CoV-2 Mpro mutant, and the binding of PF-07304814 will inhibit its hydrolysis reaction. As unambiguously shown by the electron density maps, the carbonyl carbon of the hydroxymethylketone (HMK) warhead of PF-07304814 covalently interact with the sulfur atom of Cys145 in SARS-CoV-2 Mpro mutants (Fig. 3). Two of these mutations (G15S and K90R) are distal to the substrate binding site of SARS-CoV-2 Mpro, while the other four mutations (M49I, S46F, V186F, and Y54C) are located in or nearby the substrate binding site (Figs. 2 and 3). However, the electron densities for mutant residues S46F and Y54C could only be traced in one protomer of the Mpro (Fig. 2). The quality of the fitting of two PF-07304814 molecules in the electron density maps has been evaluated by two parameters, namely the real-space R-factor (RSR) and the real-space correlation coefficient (RSCC). Calculated RSR values for the ligand 1 and ligand 2 range from 0.10 to 0.15 and from 0.08 to 0.17, respectively, while RSSC values for the ligand 1 and ligand 2 range from 0.87 to 0.93 and from 0.90 to 0.93, respectively (Table 2).
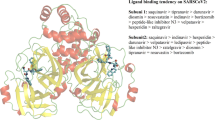
Structural overview of SARS-CoV-2 Mpro mutants in complex with PF-07304814. a SARS-CoV-2 Mpro G15S (green) in complex with PF-07304814 (magentas). b SARS-CoV-2 Mpro K90R (cyan) in complex with PF-07304814 (magentas). c PF-07304814 (magentas) bound to SARS-CoV-2 Mpro M49I (slateblue). d PF-07304814 (magentas) in complex with SARS-CoV-2 Mpro S46F (yellow). e PF-07304814 (magentas) bound to SARS-CoV-2 Mpro V186F (salmon). f PF-07304814 (magentas) bound to SARS-CoV-2 Mpro Y54C (orange). The SARS-CoV-2 Mpro mutants are shown as cartoons, while PF-07304814 molecules and mutated residues are displayed as sticks
The electron density maps of ligands, catalytic dyad residues, and mutant residues in Mpro mutants-PF-07304814 complexes. a-f The 2Fo-Fc density map (1.0σ) of PF-07304814 (magentas) molecules bound to Mpro G15S (a, green), K90R (b, cyan), M49I (c, slateblue), S46F (d, yellow), V186F (e, salmon), and Y54C (f, orange) are shown as a blue mesh. The ligands, catalytic dyad residues, and mutant residues are shown as sticks
The binding energy between PF-07304814 and Mpro mutants
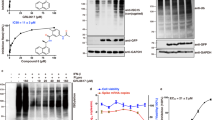
A molecular docking analysis was performed to characterize the binding energy between PF-07304814 and the six SARS-CoV-2 Mpro mutants as well as the wild-type SARS-CoV-2 Mpro. The binding energy values in kcal/mol are described in Table 3. As shown in Table 3, the binding energy between wild-type Mpro and PF-07304814 is -9.2 kcal/mol. In comparison, PF-07304814 shows higher binding energy with the six Mpro mutants. Among these, PF-07304814 shows the least binding energy with V186F Mpro, but shows the highest binding energy with M49I Mpro. The binding energy values between PF-07304814 and these Mpro mutants are − 8.9 (V186F), -7.4 (G15S), -7.4 (K90R), -6.8 (S46F), -6.6 (Y54C), and − 6.3 (M49I) kcal/mol. These interaction energy data may reflect the strength of the enzyme-ligand complex.
Comparison of binding modes of PF-07304814 with different Mpro mutants
To determine whether conformation of different Mpro mutants is changed when bound to PF-07304814, we superposed these six structures with wild-type Mpro-PF-07304814 complex. The root mean square deviation (RMSD) values of equivalent Cα positions range from 0.695 to 0.808 Å. The results showed that the binding patterns of PF-07304814 are not significantly perturbed by these substitutions on SARS-CoV-2 Mpro (Fig. 4). Indeed, two substitutions, namely G15S and K90R, are far from the binding site of PF-07304814 and have little impact on the interaction between PF-07304814 and Mpro. Though four other substitutions, namely S46F, M49I, Y54C, and V186F, situated nearby the binding site of PF-07304814, no significant changes were found in the orientation of PF-07304814.
Structural comparison of SARS-CoV-2 Mpro-PF-07304814 complexes. a Overview of structural superposition of wild-type Mpro-PF-07304814 complex (gray) and Mpro G15S-PF-07304814 complex (green), Mpro K90R-PF-07304814 complex (cyan), Mpro M49I -PF-07304814 complex (slateblue), Mpro S46F-PF-07304814 complex (yellow), Mpro V186F-PF-07304814 complex (salmon), and Mpro Y54C-PF-07304814 complex (orange). The SARS-CoV-2 Mpro and its mutants are shown as cartoons, while PF-07304814 molecules are displayed as sticks. b The representative electronic density map of the ligands. c A enlarged view of structural superpositions
Detailed interaction between PF-07304814 and different Mpro mutants
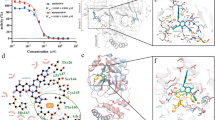
We further analyzed the interaction details between PF-07304814 and different Mpro mutants. PF-07304814 is composed of four moieties, including P1′, P1, P2, and P3 (Fig. 1), and each substituent occupies S1’, S1, S2, and S3 pockets of SARS-CoV-2 Mpro, respectively. Similar with that in wild-type Mpro-PF-07304814 interface [23], a C-S covalent bond is formed between the HMK warhead of PF-07304814 and the active-site cysteine (Cys145) in Mpro mutants and a tetrahedral carbinol complex is generated. In addition, multiple hydrogen-bonding interactions are formed between PF-07304814 and the residues in the active site of Mpro mutants (Fig. 5). A phosphate moiety is presented at the P1′ site of PF-07304814. One of the hydroxyl groups forms a hydrogen bond with the Gly143 backbone NH of mutant Mpros, while the carbinol hydroxyl forms an additional hydrogen-bonding interaction with the backbone NH of Cys145 (Fig. 5). These interactions are also observed in the wild-type Mpro-PF-07304814 interface. However, another hydroxyl group of the phosphate moiety of PF-07304814 forms a water molecule mediated hydrogen bond with Thr26 when bound to wild-type Mpro [20]. Such an interaction can not be seen in the Mpro mutants-PF-07304814 complexes. The S1 pocket of SARS-CoV-2 Mpro has a strong preference for Gln at the P1 position. Upon binding to the wild-type Mpro or mutant Mpros, the lactam ring displayed at the P1 position of PF-07304814 forms hydrogen-bonding interactions with the Nε2 of H163, the backbone oxygen of Phe140, and the side-chain oxygen of Glu166. This moiety of the inhibitor is expected to mimic Gln at the P1 position of the substrate. Moreover, a hydrogen-bonding interaction can be seen between the P1 NH in PF-07304814 and the main-chain carbonyl oxygen of His164 in wild-type and mutant Mpros. A leucine moiety is presented at the P2 position of PF-07304814, while an indole group is displayed at the P3 position. The P2 NH forms a hydrogen-bonding interaction with the side chain of Gln189 in wild-type and mutant Mpros (except Y54C). In Mpro Y54C-PF-07304814 complex, the hydrogen bond between the Gln189 of mutant Mpro and the inhibitor can not be observed. For both wild-type and mutant Mpros, the P3 indole of PF-07304814 forms two another hydrogen-bonding interactions with the backbone carbonyl and NH of Glu166. These structural details established that how lufotrelvir recognizes different SARS-CoV-2 Mpro mutants, which will be helpful in drug resistance monitoring and further drug development.
Structural characterization of PF-07304814 bound to SARS-CoV-2 wild-type Mpro or Mpro mutants. a-f Superposition of the x-ray crystal structures of PF-07304814 bound to wild-type (WT) SARS-CoV-2 Mpro (a-f, gray) and SARS-CoV-2 Mpro G15S (a, green), K90R (b, cyan), M49I (c, slateblue), S46F (d, yellow), V186F (e, salmon), and Y54C (f, orange). PF-07304814 molecules bound to WT Mpro are displayed as gray sticks, while PF-07304814 molecules bound to mutant Mpros are shown as magentas sticks. Hydrogen bonds are shown as black dashed lines between WT Mpro protein and the inhibitor, while hydrogen bonds are shown as red dashed lines in Mpro mutants-PF-07304814 complexes
Discussion
Effective antiviral drugs are urgently required to combat the public health threat caused by SARS-CoV-2 and its variants. Main protease (Mpro) serves as a superior target for anti-SARS-CoV-2 drugs. PF-07304814, also named lufotrelvir, is one of the most advanced Mpro inhibitors that have entered clinical trials and also the phosphate prodrug of PF-00835231 that could increase its bioavailability [21]. In addition, PF-07304814 exhibits good tolerability, ADME, and safety in rat models and preclinical trials [21]. Even that the phase 1 data of PF-07304814 were not released and the phase 2/3 trial of PF-07304814 was also suspended, such a promising Mpro inhibitor provides a valuable scaffold for drug design. In fact, PF-07321332 (nirmatrelvir), the active ingredient in the oral COVID-19 drug PAXLOVID™, was developed by the modification of PF-07304814. Moreover, as a phosphate ester prodrug of PF-00835231, PF-07304814 displays a very similar molecule structure with its active form. This study used structural biology methods to reveal the differences and similarities of PF-07304814 in binding with different Mpro mutants and evaluated the possible conformation change of mutant Mpros when bound to this inhibitor. Information on structural changes of Mpro mutants when bound to PF-07304814 can be useful for understanding how to inhibit mutant Mpros with potential drug resistance and for optimizing the inhibition potency of PF-07321332 as well as PF-00835231. Thus, these data will be informative to develop next generation drugs against wild-type SARS-CoV-2 and its variants.
The use of the electronic density to measure the model fit has some limitation in catching all possible problems in the model [26]. A good manner to evaluate how well a subset of atomic coordinates fits the experimental electron density is the Real Space R-factor (RSR) [27]. Another well-established measure of model fit to the experimental data is the real-space correlation coefficient (RSCC) [28]. Moreover, a RSR value less than 0.3 indicates a good agreement between observed and calculated electron densities and a RSCC value greater than 0.8 indicates a good correlation between calculated and observed electron density. In the present study, the RSR and RSSC values for the ligand PF-07304814 range from 0.08 to 0.17 and from 0.87 to 0.93, respectively, thus indicating a good consistency between the model and the density map for the co-crystallized ligands.
As determined in this study, the binding pattern of PF-07304814 is not significantly perturbed by the six single mutations (G15S, K90R, M49I, S46F, V186F, and Y54C), and the ligand largely maintaining the interaction networks observed in the wild-type Mpro with only slight differences. One obvious difference is that a water-mediated hydrogen bond with Thr26 can not be found in the interfaces of mutant Mpro-PF-07304814 complexes. Another obvious difference is that the hydrogen bond between Gln189 from Mpro Y54C and PF-07304814 can not be found. A water bridge between the ligand and Mpro Thr26 plays an important role in ligand binding [29]. Gln189 of SARS-CoV-2 Mpro has also been identified as one of the hot spot residues in the interaction with inhibitors and a hydrogen bonding with Gln189 may enhance the potency of Mpro inhibitor [29, 30]. These differences may impact the binding affinities between Mpro mutants and PF-07304814.
We then calculate the binding energies between PF-07304814 and the targeted protein to have a glimpse of the potential of PF-07304814 as a ligand to the active site of Mpro mutants. Specially, the binding between PF-07304814 and Y54C mutant is much higher than that between PF-07304814 and wild-type Mpro, which may be due to the lost of hydrogen bonding interaction between the Gln189 in Mpro Y54C and the ligand PF-07304814. Other Mpro mutants also show higher binding energy values with PF-07304814 compared to wild-type Mpro, which may be due to the lost of a water meidicated hydrogen bond between Thr26 of Mpros. As evidenced in the literature, the covalent docking may not be able to accurately reveal the real binding strength between the ligand and targeted proteins [31], because covalent docking disregards to explicitly explore the reactivity of the covalent inhibitors. Thus, further biochemical assays are needed in the future to evaluate the inhibition efficacy of PF-07304814 against these Mpro mutants.
The present study investigated the impact of single amino acid substitutions of Mpros on the its interaction with PF-07304814. However, main proteases from the circulating SARS-CoV-2 strains tend to possess multiple mutations. Next steps should also include experimental and structural characterization of the impact of combined mutations on the interactions network between PF-07304814 and SARS-CoV-2 Mpro. As PF-07304814 is a promising drug candidate to treat COVID-19, Mpro mutations that cover a larger variety of SARS-CoV-2 lineages should also be investigated in further study.
Materials and methods
Protein expression and purification
Expression of Mpro mutants of SARS-CoV-2 (including G15S, S46F, M49I, Y54C, K90R, and V186F) was performed according to previous description [23]. Briefly, pET-28a plasmids containing the entire coding sequences were transformed into competent Escherichia coli (E. coli) Rosetta DE3 cells. After grown in LB (Luria-Bertani) broth with the OD600 reaching 0.6–0.8 and induced with 500 µmol/L isopropyl-β-d-thiogalactopyranoside (IPTG), the E. coli cells started to produce mutant Mpro proteins. The cells were then centrifuged at 10,000 × g for 15 min at 4 ℃. The supernatants were discarded, while the cell pellets were collected, resuspended using lysis buffer, and lysed by sonication. The cleared lysate was subjected to immobilized metal affinity chromatography on a HisTrap HP 5 mL column (Cytiva). Imidazole gradient treatment was employed to elute the target proteins. Elution fractions containing mutant Mpro proteins were harvested and incubated with TEV protease to remove the N-terminal His-tag. The digested Mpros were subjected to gel filtration for further purification. High-purity mutant Mpro fractions were collected with centrifuge tubes and concentrated using concentrator tubes with the molecular weight cutoff of 10 kDa (Millipore).
Crystallization
The eluted SARS-CoV-2 Mpro mutants were concentrated to 5 mg/mL, and then incubated with PF-07304814 molecules at a molar ratio of 1:3 on ice for 30 min. Crystallization was carried out at 18 °C via the sitting drop vapor diffusion technique. After several days, the crystals of six SARS-CoV-2 Mpro mutants in complex with PF-07304814 were obtained. The final crystallization condition of the Mpro G15S-PF-07304814 complex was 0.15 M Tris pH 8.0, 30% (w/v) PEG 4000. The final crystallization condition of the Mpro S46F-07304814 complex was 0.20 M Na2SO4, 24% (w/v) PEG 3350. The final crystallization condition of the Mpro M49I-07304814 complex was 0.15 M Na2SO4, 20% (w/v) PEG 3350. The final crystallization condition of the Mpro Y54C-07304814 complex was 0.1 M BIS-Tris pH 6.5, 20% (w/v) PEG 3350. The final crystallization condition of the Mpro K90R-07304814 complex was 0.15 M Na2SO4, 22% (w/v) PEG 3350. The final crystallization condition of the Mpro V186F-07304814 complex was 0.16 M Na2SO4, 20% (w/v) PEG 3350.
Data Collection, structure determination, and refinement
Before X-ray diffraction data collection, the crystals of SARS-CoV-2 Mpro mutants in complex with PF-07304814 were soaked in the crystallization buffer consisting of 20% glycerol and then flash cooled in liquid nitrogen. X-ray diffraction data were collected at 100 K at BL10U2 of the Shanghai Synchrotron Radiation Facility (SSRF). All the data sets were processed using HKL2000 software [32]. All the complex structures were determined via the molecular replacement method by using the Phaser program [33]. Coot and Phenix softwares were used for atomic model building and maximum likelihood-based refinement [34, 35]. The stereochemical qualities of the final models were assessed with MolProbity [36]. The data collection and processing statistics and structural refinement statistics for the Mpro mutants-PF-07304814 complexes are shown in Table 1. The values of real-space R-factor (RSR) and real-space correlation coefficient (RSCC) were calculated to measure the electron density fit for PF-07304814 molecules in the complex structures [27, 28].
Molecular docking analysis
The co-crystallized Mpro mutants and the ligands were prepared in advance of docking. For Mpro mutants, the chain B of protein was kept and other atoms were deleted. For the ligand PF-07304814, the residue Cys145 (atoms SG, CA, CB, and C) where Michael addition reaction took place was appended to guide covalent docking. Then, the AutoDockTools-1.5.7 program was employed to add polar hydrogen atoms and assign Kollman charges. The grid box defining the binding pocket was set as the center of the ligand with length of 30 angstrom to all three dimensions. Finally, the ADFR program was applied to generate the affinity maps and perform covalent docking [37].
Availability of data and materials
Coordinates and structure factors for SARS-CoV-2 Mpro G15S, S46F, and M49I in complex with PF-07304814 have been deposited in PDB under accession code 8HVU, 8HVV, and 8HVW, respectively. Coordinates and structure factors for SARS-CoV-2 Mpro Y54C, K90R, and V186F in complex with PF-07304814 have been deposited in PDB under accession code 8HVX, 8HVY, and 8HVZ, respectively.
References
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395(10223):497–506. https://doi.org/10.1016/S0140-6736(20)30183-5.
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A novel coronavirus from patients with Pneumonia in China, 2019. N Engl J Med. 2020;382(8):727–33. https://doi.org/10.1056/NEJMoa2001017.
Coronaviridae Study Group of the International Committee on Taxonomy of V. The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. 2020;5(4):536–44. https://doi.org/10.1038/s41564-020-0695-z.
Wu JT, Leung K, Leung GM. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study. Lancet. 2020;395(10225):689–97. https://doi.org/10.1016/S0140-6736(20)30260-9.
Hui DS, Madani EIA, Ntoumi TA, Kock F, Dar R. The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health - the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis. 2020;91:264–6. https://doi.org/10.1016/j.ijid.2020.01.009.
van Doremalen N, Lambe T, Spencer A, Belij-Rammerstorfer S, Purushotham JN, Port JR, et al. ChAdOx1 nCoV-19 vaccine prevents SARS-CoV-2 pneumonia in rhesus macaques. Nature. 2020;586(7830):578–82. https://doi.org/10.1038/s41586-020-2608-y.
Baden LR, El Sahly HM, Essink B, Kotloff K, Frey S, Novak R, et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N Engl J Med. 2021;384(5):403–16. https://doi.org/10.1056/NEJMoa2035389.
Polack FP, Thomas SJ, Kitchin N, Absalon J, Gurtman A, Lockhart S, et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 vaccine. N Engl J Med. 2020;383(27):2603–15. https://doi.org/10.1056/NEJMoa2034577.
Jin L, Li Z, Zhang X, Li J, Zhu F, CoronaVac:. A review of efficacy, safety, and immunogenicity of the inactivated vaccine against SARS-CoV-2. Hum Vaccin Immunother. 2022;18(6):2096970. https://doi.org/10.1080/21645515.2022.2096970.
Kurhade C, Zou J, Xia H, Liu M, Chang HC, Ren P, et al. Low neutralization of SARS-CoV-2 Omicron BA.2.75.2, BQ.1.1 and XBB.1 by parental mRNA vaccine or a BA.5 bivalent booster. Nat Med. 2023;29(2):344–7. https://doi.org/10.1038/s41591-022-02162-x.
Shrestha LB, Foster C, Rawlinson W, Tedla N, Bull RA. Evolution of the SARS-CoV-2 omicron variants BA.1 to BA.5: implications for immune escape and transmission. Rev Med Virol. 2022;32(5):e2381. https://doi.org/10.1002/rmv.2381.
Qu P, Evans JP, Faraone JN, Zheng YM, Carlin C, Anghelina M, et al. Enhanced neutralization resistance of SARS-CoV-2 Omicron subvariants BQ.1, BQ.1.1, BA.4.6, BF.7, and BA.2.75.2. Cell Host Microbe. 2023;31(1):9–17e3. https://doi.org/10.1016/j.chom.2022.11.012.
Zhu A, Wei P, Man M, Liu X, Ji T, Chen J, et al. Antigenic characterization of SARS-CoV-2 Omicron subvariants XBB.1.5, BQ.1, BQ.1.1, BF.7 and BA.2.75.2. Signal Transduct Target Ther. 2023;8(1):125. https://doi.org/10.1038/s41392-023-01391-x.
Zappa M, Verdecchia P, Angeli F. Is the competition between variants the end of severe acute respiratory syndrome coronavirus 2 pandemic? A journey from Wuhan to XBB.1.16. Eur J Intern Med. 2023. https://doi.org/10.1016/j.ejim.2023.04.016.
Kronenberger T, Laufer SA, Pillaiyar T. COVID-19 therapeutics: small-molecule drug development targeting SARS-CoV-2 main protease. Drug Discov Today. 2023;28(6):103579. https://doi.org/10.1016/j.drudis.2023.103579.
Zhao Y, Zhu Y, Liu X, Jin Z, Duan Y, Zhang Q, et al. Structural basis for replicase polyprotein cleavage and substrate specificity of main protease from SARS-CoV-2. Proc Natl Acad Sci U S A. 2022;119(16):e2117142119. https://doi.org/10.1073/pnas.2117142119.
Yadav R, Courouble VV, Dey SK, Harrison J, Timm J, Hopkins JB, et al. Biochemical and structural insights into SARS-CoV-2 polyprotein processing by Mpro. Sci Adv. 2022;8(49):eadd2191. https://doi.org/10.1126/sciadv.add2191.
Lee J, Kenward C, Worrall LJ, Vuckovic M, Gentile F, Ton AT, et al. X-ray crystallographic characterization of the SARS-CoV-2 main protease polyprotein cleavage sites essential for viral processing and maturation. Nat Commun. 2022;13(1):5196. https://doi.org/10.1038/s41467-022-32854-4.
Xiong M, Su H, Zhao W, Xie H, Shao Q, Xu Y. What coronavirus 3 C-like protease tells us: from structure, substrate selectivity, to inhibitor design. Med Res Rev. 2021;41(4):1965–98. https://doi.org/10.1002/med.21783.
Agost-Beltran L, de la Hoz-Rodriguez S, Bou-Iserte L, Rodriguez S, Fernandez-de-la-Pradilla A, Gonzalez FV. Advances in the development of SARS-CoV-2 Mpro inhibitors. Molecules. 2022;27(8). https://doi.org/10.3390/molecules27082523.
Boras B, Jones RM, Anson BJ, Arenson D, Aschenbrenner L, Bakowski MA, et al. Preclinical characterization of an intravenous coronavirus 3CL protease inhibitor for the potential treatment of COVID19. Nat Commun. 2021;12(1):6055. https://doi.org/10.1038/s41467-021-26239-2.
Hoffman RL, Kania RS, Brothers MA, Davies JF, Ferre RA, Gajiwala KS, et al. Discovery of Ketone-Based covalent inhibitors of Coronavirus 3CL Proteases for the potential therapeutic treatment of COVID-19. J Med Chem. 2020;63(21):12725–47. https://doi.org/10.1021/acs.jmedchem.0c01063.
Li J, Lin C, Zhou X, Zhong F, Zeng P, McCormick PJ, et al. Structural basis of main proteases of coronavirus bound to drug candidate PF-07304814. J Mol Biol. 2022;434(16):167706. https://doi.org/10.1016/j.jmb.2022.167706.
Motyan JA, Mahdi M, Hoffka G, Tozser J. Potential resistance of SARS-CoV-2 main protease (Mpro) against protease inhibitors: Lessons learned from HIV-1 protease. Int J Mol Sci. 2022;23(7). https://doi.org/10.3390/ijms23073507.
Padhi AK, Tripathi T. Hotspot residues and resistance mutations in the nirmatrelvir-binding site of SARS-CoV-2 main protease: design, identification, and correlation with globally circulating viral genomes. Biochem Biophys Res Commun. 2022;629:54–60. https://doi.org/10.1016/j.bbrc.2022.09.010.
Richardson JS, Richardson DC. Studying and polishing the PDB’s macromolecules. Biopolymers. 2013;99(3):170–82. https://doi.org/10.1002/bip.22108.
Jones TA, Zou JY, Cowan SW, Kjeldgaard M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr A. 1991;47(Pt 2):110–9. https://doi.org/10.1107/s0108767390010224.
Tickle IJ. Statistical quality indicators for electron-density maps. Acta Crystallogr D Biol Crystallogr. 2012;68(Pt 4):454–67. https://doi.org/10.1107/S0907444911035918.
Thakur A, Sharma G, Badavath VN, Jayaprakash V, Merz KM Jr, Blum G, et al. Primer for Designing Main protease (M(pro)) inhibitors of SARS-CoV-2. J Phys Chem Lett. 2022;13(25):5776–86. https://doi.org/10.1021/acs.jpclett.2c01193.
Chan HTH, Moesser MA, Walters RK, Malla TR, Twidale RM, John T, et al. Discovery of SARS-CoV-2 M(pro) peptide inhibitors from modelling substrate and ligand binding. Chem Sci. 2021;12(41):13686–703. https://doi.org/10.1039/d1sc03628a.
Sanches M, Duffy NM, Talukdar M, Thevakumaran N, Chiovitti D, Canny MD, et al. Structure and mechanism of action of the hydroxy-aryl-aldehyde class of IRE1 endoribonuclease inhibitors. Nat Commun. 2014;5:4202. https://doi.org/10.1038/ncomms5202.
Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–26. https://doi.org/10.1016/S0076-6879(97)76066-X.
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ. Phaser crystallographic software. J Appl Crystallogr. 2007;40(Pt 4):658–74. https://doi.org/10.1107/S0021889807021206.
Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr. 2010;66(Pt 2):213–21. https://doi.org/10.1107/S0907444909052925.
Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta Crystallogr D Biol Crystallogr. 2010;66(Pt 4):486–501. https://doi.org/10.1107/S0907444910007493.
Chen VB, Arendall WB 3rd, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr. 2010;66(Pt 1):12–21. https://doi.org/10.1107/S0907444909042073.
Bianco G, Forli S, Goodsell DS, Olson AJ. Covalent docking using autodock: two-point attractor and flexible side chain methods. Protein Sci. 2016;25(1):295–301. https://doi.org/10.1002/pro.2733.
Acknowledgements
H.J. was supported by the Natural Science Foundation of Jiangxi Province (20224BAB216004). J.L. was supported by Jiangxi Natural Science Foundation for Distinguished Young Scholars (20212ACB216001), Jiangxi Key Research and Development Program (20203BBG73063), Jiangxi “Double Thousand Plan (jxsq2019101064)”, and the foundation of Gannan Medical University (QD201910). J.Z. was supported by the National Natural Science Foundation of China (32271260), the CAS “Light of West China” Program (xbzg-zdsys-202005), and the Natural Science Foundation of Jiangxi Province (20224ACB206046).
Author information
Authors and Affiliations
Contributions
JL and JZ designed experiments. HJ, XFZ, and PZ purified and crystallized the protein as well as performed the related assays. PZ, XYZ, XLZ, and JW collected the diffraction data. HJ and XFZ prepared the figures. HJ wrote the manuscript. HJ, JZ, and JL revised the manuscript. The authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare that no conflicts of interest exist.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, H., Zou, X., Zeng, P. et al. Crystal structures of main protease (Mpro) mutants of SARS-CoV-2 variants bound to PF-07304814. Mol Biomed 4, 23 (2023). https://doi.org/10.1186/s43556-023-00134-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s43556-023-00134-2