Abstract
Aromatic side chains are attractive probes of protein dynamic, since they are often key residues in enzyme active sites and protein binding sites. Dynamic processes on microsecond to millisecond timescales can be studied by relaxation dispersion experiments that attenuate conformational exchange contributions to the transverse relaxation rate by varying the refocusing frequency of applied radio-frequency fields implemented as either CPMG pulse trains or continuous spin-lock periods. Here we present an aromatic 1H R1ρ relaxation dispersion experiment enabling studies of two to three times faster exchange processes than achievable by existing experiments for aromatic side chains. We show that site-specific isotope labeling schemes generating isolated 1H–13C spin pairs with vicinal 2H–12C moieties are necessary to avoid anomalous relaxation dispersion profiles caused by Hartmann–Hahn matching due to the 3JHH couplings and limited chemical shift differences among 1H spins in phenylalanine, tyrosine and the six-ring moiety of tryptophan. This labeling pattern is sufficient in that remote protons do not cause additional complications. We validated the approach by measuring ring-flip kinetics in the small protein GB1. The determined rate constants, kflip, agree well with previous results from 13C R1ρ relaxation dispersion experiments, and yield 1H chemical shift differences between the two sides of the ring in good agreement with values measured under slow-exchange conditions. The aromatic1H R1ρ relaxation dispersion experiment in combination with the site-selective 1H–13C/2H–12C labeling scheme enable measurement of exchange rates up to kex = 2kflip = 80,000 s–1, and serve as a useful complement to previously developed 13C-based methods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Conformational dynamics in proteins on the microsecond to millisecond time scales are often linked to biological function (Mittermaier and Kay 2009). Transiently populated high-energy states play important roles in enzyme catalysis (Boehr et al. 2006; Cole and Loria 2002; Eisenmesser et al. 2002) or ligand binding by conformational selection (Demers and Mittermaier 2009; Malmendal et al. 1999). Such conformational transitions generally lead to a modulation of NMR parameters as the chemical shift (Gutowsky and Saika 1953), residual dipolar coupling (Igumenova et al. 2007; Vallurupalli et al. 2007) or strong scalar coupling (Weininger et al. 2013b). The conformational dynamics can be probed by NMR relaxation dispersion methods (Palmer 2004; Palmer et al. 1991), such as R1ρ (Akke and Palmer 1996; James et al. 1977) or Carr-Purcell-Meiboom-Gill (CPMG) experiments (Carr and Purcell 1954; Loria et al. 1999a, 1999b; Meiboom and Gill 1958), and chemical exchange saturation transfer (CEST) methods (Forsen and Hoffman 1963; Palmer and Koss 2019; Vallurupalli et al. 2012, 2017) in the case of slower time scales.
Phenylalanine, tyrosine, histidine and tryptophan all have aromatic side chains, which make them an interesting subgroup of amino acids that serve multiple functions in proteins. Aromatic side chains are bulky and constitute a significant proportion of the protein hydrophobic core. They typically form pairs or clusters on the basis of specific aromatic-aromatic interactions (Burley and Petsko 1985, 1989). Furthermore, aromatic residues are overrepresented in protein binding interfaces, where they contribute a significant part of the binding free energy (Birtalan et al. 2010; Bogan and Thorn 1998; Lo Conte et al. 1999). Finally, tyrosine and especially histidine play critical roles in enzyme catalysis, where they make up 6% and 18%, respectively, of all catalytic residues (Bartlett et al. 2002). Histidine can exist in three different states, one protonated and two neutral tautomeric forms. Despite their generally tight packing, Phe and Tyr residues undergo intermittent 180° transitions (‘ring flips’) of the χ2 dihedral angle and thereby provide unique information of transient 'breathing' processes of proteins (Dreydoppel et al. 2020; Wagner 1980; Wagner et al. 1976; Weininger et al. 2014b). Each of these properties makes aromatic side chains highly interesting and powerful probes for studying protein dynamics.
Recent developments have enabled straightforward and robust site-selective 13C labeling of aromatic side chains (Kasinath et al. 2013; Lundström et al. 2007; Schörghuber et al. 2018; Teilum et al. 2006; Weininger 2019), which eliminate unwanted relaxation pathways and coherent magnetization transfer via one-bond 13C–13C couplings. These advancements have made possible advanced studies of protein dynamics involving aromatic side chains, including methods to characterize fast (ps–ns) timescale dynamics via 13C relaxation rate constants (Weininger et al. 2012a) that enable studies of order parameters (Boyer and Lee 2008; Kasinath et al. 2013, 2015). Furthermore, slower (µs–ms) timescale dynamics have been probed with 13C relaxation dispersion experiments, either using CPMG (Weininger et al. 2012b) or R1ρ refocusing elements (Weininger et al. 2014a). In particular, these relaxation dispersion methods have been used to measure ring-flip rates (Dreydoppel et al. 2020; Weininger et al. 2014b) and transient histidine tautomerization (Weininger et al. 2017). Complementary 1H CPMG relaxation dispersion experiments are applicable to a subset of aromatic sites, namely His δ2, His ε1 and Trp δ1. Other positions are inaccessible due to sizeable 3JHH couplings and possibly strong 1JCC, where 1 J-couplings are equal or greater than the chemical shift difference, can cause severe artifacts (Raum et al. 2018). Site-selective 1H–13C/2H–12C labeling addresses both issues and allows to obtain artifact-free 1H CPMG relaxation dispersion profiles (Raum et al. 2019).
It is expected that 1H R1ρ experiments make it possible to access faster time scales, since the attainable refocusing frequency scales with the gyromagnetic ratio and the RF field amplitude, ω1 = γB1. 1H R1ρ experiments have successfully been applied to the aromatic position H8 of adenine and guanine and in RNA (Steiner et al. 2016), which is not affected by sizeable 3JHH couplings or cross relaxation with remote protons. The situation is more challenging in certain aromatic protein residues, where 3JHH is on the order of 7–8 Hz and the chemical shift difference between vicinal ring protons is often small. Here we demonstrate these problems can be solved by site-selective 1H–13C/2H–12C labeling, which is necessary and sufficient to ensure artifact-free 1H R1ρ relaxation dispersion data for aromatic side chains in proteins.
Materials and methods
Protein samples
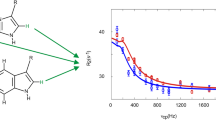
The B1 domain of the bacterial antibody-binding protein G (GB1) containing the mutations T2Q, N8D and N37D (QDD variant) was expressed and purified as described previously (Lindman et al. 2006), with three different labeling patterns: (i) site-selective 13C/12C using 2-13C1 glucose with natural abundance 1H incorporation (Lundström et al. 2007), (ii) site-selective 1H–13C/2H–12C labeling protein (Fig. 1) using site-selectively 1H/2H and 13C labeled -ketoacids as precursors (Lichtenecker 2014; Lichtenecker et al. 2013), 80 mg/L for Phe, 350 mg/L for Tyr, 10 mg/L for Trp, (iii) same as (ii) but with additional 70% background deuteration by expression in D2O (implemented only in the case of Phe ε*). NMR samples contained around 1 mM protein in 20 mM HEPES, 90% H2O/10% D2O with addition of small amounts of NaN3. The pH of the NMR samples was adjusted to 7.0.
NMR spectroscopy
All experiments were acquired on a Bruker Avance III spectrometer at a static magnetic field strength of 14.1 T, equipped with a room temperature probe. 1H R1ρ relaxation dispersion experiments were performed using the pulse sequence shown in Fig. 2, with spectral widths of 14.0 ppm (1H) and 30.0 ppm (13C), by 1024 and 128 points, respectively. Nineteen experiments were performed at temperatures of 15, 20, 25, 30 and 35 °C. R1ρ rate constants were measured in separate experiments conducted on-resonance with the signal of interest resulting in tilt angles of 90° from the z-axis, using a constant-time relaxation period (Mulder et al. 2001) of 20 ms and spin-lock field strengths varying between 1000 and 9000 Hz; the 4-ms adiabatic ramps used to align the magnetization along the effective spin-lock field do not achieve perfect alignment below 1000 Hz. Spectra were processed with NMRPipe (Delaglio et al. 1995) and analyzed with PINT (Ahlner et al. 2013).
Pulse sequence for the 1H R1ρ constant-time relaxation dispersion experiment for measuring conformational exchange of aromatic side chains in specifically 1H–13C/2H–12C labeled proteins. All pulses are applied along the x-axis unless otherwise indicated. Narrow (wide) solid bars indicate rectangular high-power 90° (180°) pulses. The continuous-wave spin-lock relaxation periods T and their flanking 4 ms tan/tanh adiabatic profiles (Mulder et al. 1998) are outlined in gray. The adiabatic sweep is initiated 25 kHz downfield or upfield of the spin-lock frequency. The wide semi-ellipse on 13C represents a REBURP (Geen and Freeman 1991) pulse with a bandwidth of 40 ppm. The pulse sequence can be modified to accommodate non-constant relaxation periods or off-resonance spin-locks by including a pair of 13C 180° pulses to mitigate the effects of dipole–dipole/CSA cross-correlated relaxation (Korzhnev et al. 2002; Massi et al. 2004). The delay τ can be set to 1.6 ms (Phe and Tyr), 1.35 ms (all aromatics) or 1.25 ms (His). The phase cycle is: ϕ1 = (4y, 4 − y), ϕ2 = (x, − x), ϕ3 = (x, x, − x, − x), ϕrec = (x, –x, –x, x, − x, x, x, − x). Pulsed field gradients G1–4 are employed to suppress unwanted coherences and artifacts, while GC and GH are encoding and decoding gradients, respectively, for echo/anti-echo coherence selection, obtained by inverting the signs of GH (Kay et al. 1992). The delay ∆ is equal to GC. For every second t1 increment ϕ2 and the receiver were incremented. Gradient durations (in ms) and relative power levels (in %) are set to (duration, power level) G1 = (1.0, 13), G2 = (0.5, 10), G3 = (1.0, 30), G4 = (1.0, 90), GC = (1.0, 80), GH = (1.0, -20.1)
Data analysis
Relaxation rates in the rotating reference frame were calculated from the signal intensities measured in presence and absence of the spin-lock pulse, I(ω1) and I0, respectively, according to (Mulder et al. 2001)
R1ρ relaxation dispersions were fitted to the general equation for symmetric exchange derived by Miloushev and Palmer (2005) using fixed populations, p1 = p2 = 0.5, and treating ∆δ as a free parameter, which was compared to values measured under slow-exchange conditions at – 5 °C and 200 MPa. Data modeling utilized the Levenberg–Marquardt nonlinear least-squares optimization algorithm (Press et al. 2002) implemented in MATLAB. Errors in the fitted parameters were estimated using 1000 Monte-Carlo simulations per fit; the reported errors correspond to one standard deviation.
Activation parameters of the ring flips were determined by non-linear regression of the flip rates, kflip = kex/2, on the temperature T, using the Eyring equation. The Eyring equation was parameterized as
where kB and h denote Boltzmann’s and Planck’s constants, respectively, and ∆H‡ and ∆S‡ are the activation enthalpy and activation entropy, respectively.
Hartmann–Hahn transfer calculations
A nucleus affected by a spin-lock radio frequency pulse with field strength \({\omega }_{1}=-\gamma {B}_{1}\) will perceive an effective field strength \({\omega }_{\mathrm{eff}}={({\Omega }^{2}+{{\omega }_{1}}^{2})}^{1/2}\) and will be oriented at an angle \(\theta ={\mathrm{tan}}^{-1}({\omega }_{1}/\Omega )\) from the static magnetic field, with \({\Omega =\omega }_{0}{-\omega }_{c}\) denoting the offset between the nuclear precession frequency ω0 and the carrier frequency ωc of the spin-lock pulse. In the case of two spins I and S, coupled by a scalar-coupling constant J, Hartmann–Hahn matching of the two effective fields causes magnetization transfer between them, according to the coherence transfer function given by (Brath et al. 2006; van de Ven 1995)
where TSL is the duration of the spin-lock period, \(D={({\Delta }^{2}+ {{J}_{\mathrm{eff}}}^{2})}^{1/2}\), \(\Delta ={\omega }_{\mathrm{eff},I}-{\omega }_{\mathrm{eff},S}\), \({J}_{\mathrm{eff}}=\frac{1}{2}J(1+\cos({\theta }_{I}-{\theta }_{S}))\) and AHaHa is the coherence transfer amplitude
Hartmann–Hahn mediated magnetization transfer from the monitored proton of the 1H-13C moiety to vicinal protons in the aromatic ring scales the intensity by a factor of 1 – FHaHa(ω1) for a given spin-lock period TSL. Consequently, the apparent R1ρ rate constant is given by:
where the first term describes the R1ρ rate constant according to Eq. (1), i.e., in the absence of Hartmann–Hahn matching. Similarly, as a first approximation, in the case of n protons coupled to the monitored proton, the apparent rate constant is:
AHaHa and FHaHa were calculated for scalar-coupled protons in the aromatic rings of GB1 using J = 3JHH = 7 Hz, TSL = 20 ms, and the resonance frequencies measured in the spectrum acquired at 25 °C: 0.22 ppm between Y33 ε* and δ*; 0.65 ppm (0.16 ppm) between F52 ε* and δ*(ζ); and 0.98 ppm (0.13 ppm) between W43 ζ3 and ε3 (η2). In case of frequencies averaged by fast ring-flips, i.e. δ* and ε* in Phe and Tyr, it is sufficient to use them directly.
Results and discussion
We performed aromatic 1H R1ρ relaxation dispersion experiments on the small protein domain GB1, 1H–13C labeled in three different ways: (i) natural abundance protonation and site selective 13C/12C labeling; (ii) additional selective deuteration of vicinal hydrogen sites in order to eradicate 3JHH couplings resulting in site-selective 1H–13C/2H–12C labeling (Fig. 1); or (iii) as (ii) with additional non-specific background deuteration at a level of 70%. There are five symmetric aromatic residues (Phe and Tyr) in GB1: Y3, F30, Y33, Y45 and F52. All but Y33 undergo relatively slow ring-flip processes that have been studied by aromatic 13C R1ρ relaxation dispersion experiments (Dreydoppel et al. 2020); in particular Y3δ, Y3ε, F30δ, Y45ε and F52ε have been studied previously and provide valuable points of reference for comparison with our present work. Individual cross peaks from the two symmetric sites of the aromatic rings could be observed under slow exchange conditions (–5 °C, 200 MPa) for Y3δ, Y3ε, F30δ, F30ε and F52ε. Based on these results, the following 13C and 1H chemical shift differences could be extracted: Y3δ (2.11 ppm; 0.40 ppm), Y3ε (1.40 ppm; 0.50 ppm), F30δ (5.39 ppm; 0.84 ppm), F30ε (0.00 ppm; 0.56 ppm) and F52ε (1.76 ppm; 0.00 ppm). F52ε, which has a 1H chemical shift difference of zero, serves as a negative control in the present study, since it should not exhibit any exchange-mediated dependence of the R1ρ rate constant on the spin-lock field strength. In addition, Y33ε, which is solvent exposed and undergoing very fast ring flips, is also expected to show an essentially flat dispersion profile. Furthermore, the two labeled sites in tryptophan, W43δ1 and W43ζ3 also serve as useful controls: W43δ1 does not experience sizeable 3JHH couplings, and neither site undergoes exchange, since this residue does not readily undergo ring flips. In a previous study, site-selective 1H–13C/2H–12C labeling resulted in 99% correct labeling in case of Phe ε* and 75% in case of Trp ζ3, whereas Tyr ε* labeling yielded only 4% incorporation for reasons that are not clear (Raum et al. 2019). Here, we increased the amount of tyrosine precursor fourfold in the protein expression medium, resulting in 17% of the desired labeling for Tyr ε*, while the remaining 83% have natural abundance isotope incorporation. Therefore, 0.9% of the sample is 13C labeled at Tyr ε and protonated at the δ position. Thus, in the present sample the latter isotopomer is diluted by a factor of 20 compared to the desired one, which is sufficient to suppress 3JHH coupling artifacts.
Deuteration of vicinal protons is necessary in order to achieve artifact-free aromatic 1H R1ρ relaxation dispersion profiles
Figure 3 shows the resulting 1H R1ρ relaxation dispersion profiles for the 1H–13C labeled ε sites of residues Y3, Y33, F30, and F52 (a-d), and for the δ1 and ζ3 sites of W43 (e–f). There is a notable variation among the dispersion profiles of the different residues and between the two1H/2H labeling patterns for each residue. The non-deuterated samples of residues Y33ε, F30ε, F52ε, and W43ζ3 (Figs. 3b–d and f; red symbols) all exhibit anomalous, ‘inverted’ relaxation dispersion profiles characteristic of sizeable (7–8 Hz) 3JHH coupling (Raum et al. 2018). In contrast, the dispersion profile of Y3ε (Fig. 3a) does not appear anomalous, because the effect of Hartmann–Hahn matching is in part masked by a substantial chemical exchange contribution. In general, the magnitude of the artifact is more pronounced in the case of Phe (Fig. 3c, d) compared to Tyr (Fig. 3a, b), because the investigated 1Hε spins in Phe couple to two vicinal proton spins (δ and ζ) instead of one (δ) in Tyr. In the case of Trp ζ3, the magnitude is closer to the Tyr case. In the non-deuterated sample, W43δ1 displays a flat relaxation dispersion profile (Fig. 3e), as a result of its small coupling constant, 3JHH < 2 Hz, and absence of exchange. This result is in keeping with the expectation outlined above regarding the role of W43δ1 as a negative control.
Aromatic 1H R1ρ relaxation dispersions recorded on-resonance (tilt angle θ = 90° from the z-axis) at a static magnetic field-strength of 14.1 T and a temperature of 25 °C. Dispersion profiles obtained from uniformly protonated and site-selectively 13C/12C labeled samples are shown in red, from site-selectively 1H–13C/2H–12C labeled samples in blue, and site-selectively 1H–13C/2H–12C labeled samples with additional 70% background deuteration in green. Dispersion profiles are shown for Y3ε (A), Y33ε (B), F30ε (C), F52ε (D), W43δ1 (E) and W43ζ3 (F)
Site-selective deuteration of the vicinal hydrogen sites efficiently removes the effect of 3JHH coupling, resulting in ‘normal’-looking relaxation dispersion profiles (Fig. 3; blue symbols). Y33ε, F52ε, and W43ζ3 (Figs. 3b, d, f) show flat dispersion profiles in the deuterated sample, again in agreement with expectations (very fast exchange, no 1H chemical shift difference between the symmetric sites, and no exchange, respectively; see above). In the case of F30ε, the site-selectively deuterated sample reveals a fast exchange process that is completely hidden by the strong-coupling effect in the protonated sample (Fig. 3c). Finally, for Y3ε where the relaxation dispersion profile shows clear signs of exchange even in the non-deuterated sample, the profile is more pronounced in case of site-selective deuteration (Fig. 3a). This comparison clearly shows that the Y3ε profile in the non-deuterated sample is compromised and cannot be analyzed to yield reliable exchange parameters. Additional, unspecific 70% background deuteration of the protein does slightly shift the 1H R1ρ to lower values, but does not alter the profile shape (Fig. 3c, d, green symbols). A quantitative analysis comparing the results obtained from the samples with or without background deuteration shows that identical exchange parameters are obtained, within the margin of error (see below). This result demonstrates that site-selective deuteration in the aromatic ring alone is sufficient to obtain artifact free relaxation dispersion profiles, and further indicates that long-range ROE effects (Lundström and Akke 2005; Weininger et al. 2013a) do not cause concern in the present case.
Taken together, the present results illustrate the detrimental effects of 3JHH couplings on the relaxation dispersion data obtained on non-deuterated samples, and clearly demonstrate that high-quality 1H R1ρ relaxation dispersion profiles can only be obtained using samples with site-selective deuteration. Our present approach enables 1H R1ρ relaxation dispersion experiments for Phe, Tyr, and Trp ζ3, and thus completes the repertoire of probes to cover all aromatic 1H-13C positions. By contrast, only Trp δ1, His δ2 and His ε1 are accessible also in non-deuterated samples, because these positions do not involve sizeable scalar coupling to vicinal protons.
Hartmann–Hahn transfer explains artifacts in aromatic 1H R1ρ relaxation dispersion experiments
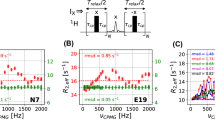
The artifacts described above for the non-deuterated samples (Fig. 3) can be rationalized as increased homonuclear Hartmann–Hahn transfer from the proton of interest (I) to a scalar coupled vicinal proton (S) with increasing spin-lock field strength (ω1). Figure 4 shows the expected values of the amplitude (AHaHa) of the transfer function and the relative loss of magnetization for spin I (1–FHaHa), as a function of the spin-lock field strength and the difference in offset between spins I and S; see Eqs. (3–5). The magnetization loss expected for the case where spin I has two J-coupled vicinal protons, as for Phe ε, is included in Fig. 4c; see Eq. (6). We calculated the amount of magnetization lost from I during the spin-lock period for those residues that do not show exchange contributions to R1ρ (Y33ε, F52ε, and W43ζ3), and estimated their resulting apparent R1ρ rate constant as a function of ω1. To do so, we took the average R1ρ rate constant measured on site-selectively 1H–13C/2H–12C labeled samples to represent the artifact-free first term of Eq. (6), and added the second term involving the calculated value of (1–FHaHa). As shown in Figs. 4d–f, in all three cases the experimental data points acquired on the uniformly protonated sample are reproduced well by the calculated function describing the apparent R1ρ values. Thus, Hartmann–Hahn transfer quantitatively explains the observed artifacts in aromatic 1H R1ρ relaxation dispersion experiments acquired on uniformly protonated samples.
Hartmann–Hahn coherence transfer between spin-locked proton spins. The carrier frequency of the spin-lock is chosen to match the resonance frequency of spin I. A Contour levels of the coherence transfer amplitude, AHaHa = 0.5, as a function of the spin-lock ω1 and the resonance offset from the spin-lock carrier for spin S, Ω. Contour lines are shown for various differences in resonance frequency offsets between the coupled nuclei: 0.2 ppm (red), 0.4 ppm (yellow), 0.6 ppm (green), 0.8 ppm (blue) and 1.0 ppm (magenta). B AHaHa plotted as a function of spin-lock field strength for different resonance frequency offsets as in (A). C Loss of coherence according to the transfer function, 1 – FHaHa, for resonance frequency offsets of 0.2 ppm (red) and 0.6 ppm (green), and the product of both (black). D–F Measured data points from Y33ε (D), F52ε (E), and W43ζ3 (F) as given in Fig. 3, together with predicted apparent R1ρ dispersion profiles (black lines), calculated using Eq. 6 and the artifact-free data measured on deuterated samples (blue points)
Comparison of ring flip rates of 1H and 13C based aromatic R1ρ relaxation dispersion experiments
We acquired relaxation dispersion profile for Y3ε and F30ε in the site-selectively deuterated sample at three different temperatures (Fig. 5). The artifact-free relaxation dispersions could be fitted to the general equation for symmetric exchange, resulting in flip rates at each temperature together with a global chemical shift difference for each residue (Table 1). The derived chemical shift differences are ∆δ = (0.46 ± 0.03) ppm for Y3ε and (0.47 ± 0.02) ppm for F30ε, in good agreement with the values measured in the spectrum under slow exchange conditions (0.50 ppm and 0.56 ppm, respectively). In the case of Y3ε, the resulting flip rates are virtually the same, within the margin of error, as those determined from 13C based experiments (Fig. 6a, Table 1). Consequently, the activation enthalpy (∆H‡) and activation entropy (∆S‡) determined from the temperature dependence of the flip rates are identical within the margin of error to previous estimates (Fig. 6a): ∆H‡ = (82 ± 8) kJ mol−1 for 1H versus (87 ± 14) kJ mol−1 for 13C; and ∆S‡ = (111 ± 26) J mol−1 K−1 versus (126 ± 46) J mol−1 K−1. In case of F30ε, we cannot directly compare flip rates determined by 1H and 13C R1ρ experiments, because its 13Cε chemical shift difference is zero. However, ring flip rates have been measured for F30δ using 13C R1ρ relaxation experiments at two higher temperatures, although the flip rate is far outside the optimal range of the R1ρ experiment in this case and should only be interpreted semi-quantitatively (Fig. 6b). At lower temperatures F30δ becomes broadened beyond detection, because of its large 13Cδ chemical shift difference. The rates derived from the 1H R1ρ experiments seem to be a bit higher (as gauged from the extrapolated line in Fig. 6b), although the two data points from the 13C R1ρ experiments seem to follow the temperature dependence determined from the 1H R1ρ experiments. Background deuteration in the F30ε sample leads to a minor improvement of the fit in the region of the highest refocusing frequencies (Fig. 5 bc), where the ROE effect is the most severe. The derived values of ∆H‡ and ∆S‡ with and without background deuteration are also identical, within the margin of error (Fig. 6b): ∆H‡ = (47 ± 4) kJ mol−1 for 1H with protonated background versus (50 ± 1) kJ mol−1 with deuterated background; and ∆S‡ = (0 ± 13) J mol−1 K−1 versus (10 ± 4) J mol−1 K−1. In summary, aromatic 1H R1ρ experiments allow the determination of correct exchange parameters, provided that the aromatic ring is site-selectively 1H/2H labeled; further deuteration of the background is not needed.
Temperature dependent aromatic1H R1ρ relaxation dispersion data. Experiments were recorded on-resonance (θ = 90° from the z-axis) on site-selectively 1H–13C/2H–12C labeled GB1 at pH 7.0 and a static magnetic field strength of 14.1 T. Data of Y3ε acquired at 25 °C, 30 °C, and 35 °C and F30ε at 15 °C, 20 °C, and 25 °C are represented by light, medium, and dark hues, respectively. A Y3ε (blue), B F30ε (blue), and C F30ε with additionally 70% deuterated background (green). The relaxation dispersions were fitted using a fixed population p1 = p2 = 0.5 and ∆δ as a free parameter with the restrictions: kflip (Thigh) > kflip (Tlow), R2,0 (Thigh) ≤ R2,0 (Tlow). The derived ring flip rate constants (kflip) are given in Table 1
Temperature dependence of ring-flip rates. kflip is plotted as a function of 1/T. A Y3ε and B F30ε flip rates determined from aromatic 1H R1ρ relaxation dispersion measurements are shown in blue (protonated background) and green (deuterated background), and from previous 13C R1ρ relaxation dispersion measurements in grey (Dreydoppel et al. 2020). Solid lines represent non-linear regression of kflip on T according to the Eyring equation, displayed on a logarithmic y-axis to show the expected linearity. Derived activation enthalpies (∆H‡) and entropies (∆S‡) are: Y3 1H (protonated background), (82 ± 8) kJ mol−1 and (111 ± 26) J mol−1 K−1 (A, blue); Y3 13C (87 ± 14) kJ mol−1 and (126 ± 46) J mol−1 K−1 (A, grey); F30 1H (protonated background), (47 ± 4) kJ mol−1 and (0 ± 13) J mol−1 K−1 (B, blue); F30 1H (deuterated background), (50 ± 1) kJ mol−1 and (10 ± 4) J mol−1 K−1 (B, green)
Concluding remarks
There are multiple positions available in aromatic side chains that are suitable for studying exchange dynamics. However, it is not uncommon that certain sites do not experience sufficiently large ∆δ for a given nucleus, or experience exchange rates outside of the optimal relaxation dispersion window for a given nuclide (e.g., 1H or 13C). Thus, it is highly advantageous to be able to measure exchange using different nuclides, so as to probe as many sites as possible in a given protein. For example, our studies of ring flip dynamics in GB1 highlight the advantage of using both 1H and 13C based experiments:Y3 can be probed using all four positions (1Hδ, 13Cδ, 1Hε and 13Cε ), while F30 and F52 can each be probed using only a single probe (1Hε and 13Cε , respectively). Our present approach extends the available toolbox to make all aromatic positions accessible by NMR relaxation dispersion experiments using either CPMG or R1ρ refocusing elements. The higher effective field strength available in 1H R1ρ experiments is clearly of advantage in this context, and allows to reliably study exchange processes up to an exchange rate of about 80,000 s−1 (equal to a flip rate of 40,000 s−1).
Data availability
All data generated or analysed during this study are included in this published article.
References
Ahlner A, Carlsson M, Jonsson BH, Lundström P (2013) PINT: a software for integration of peak volumes and extraction of relaxation rates. J Biomol NMR 56:191–202. https://doi.org/10.1007/s10858-013-9737-7
Akke M, Palmer AG (1996) Monitoring Macromolecular motions on microsecond-millisecond time scales by R1r–R1 constant-relaxation-time NMR spectroscopy. J Am Chem Soc 118:911–912
Bartlett GJ, Porter CT, Borkakoti N, Thornton JM (2002) Analysis of catalytic residues in enzyme active sites. J Mol Biol 324:105–121
Birtalan S, Fisher RD, Sidhu SS (2010) The functional capacity of the natural amino acids for molecular recognition. Mol Biosyst 6:1186–1194. https://doi.org/10.1039/b927393j
Boehr DD, McElheny D, Dyson HJ, Wright PE (2006) The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313:1638–1642
Bogan AA, Thorn KS (1998) Anatomy of hot spots in protein interfaces. J Mol Biol 280:1–9
Boyer JA, Lee AL (2008) Monitoring aromatic picosecond to nanosecond dynamics in proteins via C-13 relaxation: expanding perturbation mapping of the rigidifying core mutation, V54A, in Eglin C. Biochemistry 47:4876–4886
Brath U, Akke M, Yang DW, Kay LE, Mulder FAA (2006) Functional dynamics of human FKBP12 revealed by methyl C-13 rotating frame relaxation dispersion NMR spectroscopy. J Am Chem Soc 128:5718–5727
Burley SK, Petsko GA (1985) Aromatic-aromatic interaction: a mechanism of protein structure stabilization. Science 229:23–28
Burley SK, Petsko GA (1989) Electrostatic interactions in aromatic oligopeptides contribute to protein stability trends. Biotech 7:354–359
Carr HY, Purcell EM (1954) Effects of diffusion on free pecession in nuclear magnetic resonance experiments. Phys Rev 94:630–638
Cole R, Loria JP (2002) Evidence for flexibility in the function of ribonuclease A. Biochemistry 41:6072–6081
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe - a multidimensional spectral processing system based on inix pipes. J Biomol NMR 6:277–293
Demers JP, Mittermaier A (2009) Binding mechanism of an SH3 domain studied by NMR and ITC. J Am Chem Soc 131:4355–4367. https://doi.org/10.1021/ja808255d
Dreydoppel M, Raum HN, Weininger U (2020) Slow ring flips in aromatic cluster of GB1 studied by aromatic C-13 relaxation dispersion methods. J Biomol NMR 74:183–191. https://doi.org/10.1007/s10858-020-00303-3
Eisenmesser EZ, Bosco DA, Akke M, Kern D (2002) Enzyme dynamics during catalysis. Science 295:1520–1523
Forsen S, Hoffman RA (1963) Study of moderately rapid chemical exchange reactions by means of nuclear magnetic double resonance. J Chem Phys 39:2892–3000. https://doi.org/10.1063/1.1734121
Geen H, Freeman R (1991) Band-selective radiofrequency pulses. J Magn Reson 93:93–141. https://doi.org/10.1016/0022-2364(91)90034-Q
Gutowsky HS, Saika A (1953) Dissociation, chemical exchange, and the proton magnetic resonance in some aqueous electrolytes. J Chem Phys 21:1688–1694
Igumenova TI, Brath U, Akke M, Palmer AG (2007) Characterization of chemical exchange using residual dipolar coupling. J Am Chem Soc 129:13396
James TL, Matson GB, Kuntz ID, Fisher RW (1977) Rotating frame spin-lattice relaxation in presence of an off-resonance radio-frequency field - investigation of intermediate molecular motions. J Magn Reson 28:417–426. https://doi.org/10.1016/0022-2364(77)90283-9
Kasinath V, Valentine KG, Wand AJ (2013) A C-13 labeling strategy reveals a range of aromatic side chain motion in Calmodulin. J Am Chem Soc 135:9560–9563
Kasinath V, Fu YN, Sharp KA, Wand AJ (2015) A sharp thermal transition of fast aromatic-ring dynamics in ubiquitin. Angew Chem Int Edit 54:102–107
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665. https://doi.org/10.1021/Ja00052a088
Korzhnev DM, Skrynnikov NR, Millet O, Torchia DA, Kay LE (2002) An NMR experiment for the accurate measurement of heteronuclear spin-lock relaxation rates. J Am Chem Soc 124:10743–10753. https://doi.org/10.1021/ja0204776
Lichtenecker RJ (2014) Synthesis of aromatic C-13/H-2-alpha-ketoacid precursors to be used in selective phenylalanine and tyrosine protein labelling. Org Biomol Chem 12:7551–7560. https://doi.org/10.1039/c4ob01129e
Lichtenecker RJ, Weinhaupl K, Schmid W, Konrat R (2013) alpha-Ketoacids as precursors for phenylalanine and tyrosine labelling in cell-based protein overexpression. J Biomol NMR 57:327–331
Lindman S, Xue WF, Szczepankiewicz O, Bauer MC, Nilsson H, Linse S (2006) Salting the charged surface: pH and salt dependence of protein G B1 stability. Biophys J 90:2911–2921. https://doi.org/10.1529/biophysj.105.071050
Lo Conte L, Chothia C, Janin J (1999) The atomic structure of protein-protein recognition sites. J Mol Biol 285:2177–2198
Loria JP, Rance M, Palmer AG (1999a) A relaxation-compensated Carr-Purcell-Meiboom-Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121:2331–2332
Loria JP, Rance M, Palmer AG (1999b) A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J Biomol NMR 15:151–155
Lundström P, Akke M (2005) Off-resonance rotating-frame amide proton spin relaxation experiments measuring microsecond chemical exchange in proteins. J Biomol NMR 32:163–173. https://doi.org/10.1007/s10858-005-5027-3
Lundström P et al (2007) Fractional C-13 enrichment of isolated carbons using [1-C-13]- or [2-C-13]-glucose facilitates the accurate measurement of dynamics at backbone C-alpha and side-chain methyl positions in proteins. J Biomol NMR 38:199–212
Malmendal A, Evenas J, Forsen S, Akke M (1999) Structural dynamics in the C-terminal domain of calmodulin at low calcium levels. J Mol Biol 293:883–899
Massi F, Johnson E, Wang C, Rance M, Palmer AG 3rd (2004) NMR R1 rho rotating-frame relaxation with weak radio frequency fields. J Am Chem Soc 126:2247–2256. https://doi.org/10.1021/ja038721w
Meiboom S, Gill D (1958) Modified spin-echo method for measuring nuclear relaxation times. Rev Sci Instrum 29:688–691
Miloushev VZ, Palmer AG (2005) R(1p) relaxation for two-site chemical exchange: general approximations and some exact solutions. J Magn Reson 177:221–227. https://doi.org/10.1016/j.jmr.2005.07.023
Mittermaier AK, Kay LE (2009) Observing biological dynamics at atomic resolution using NMR. Trends Biochem Sci 34:601–611
Mulder FAA, de Graaf RA, Kaptein R, Boelens R (1998) An off-resonance rotating frame relaxation experiment for the investigation of macromolecular dynamics using adiabatic rotations. J Magn Reson 131:351–357. https://doi.org/10.1006/jmre.1998.1380
Mulder FAA, Skrynnikov NR, Hon B, Dahlquist FW, Kay LE (2001) Measurement of slow (mu s-ms) time scale dynamics in protein side chains by N-15 relaxation dispersion NMR spectroscopy: application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123:967–975
Palmer AG (2004) NMR characterization of the dynamics of biomacromolecules. Chem Rev 104:3623–3640
Palmer AG 3rd, Koss H (2019) Chemical exchange. Methods Enzymol 615:177–236. https://doi.org/10.1016/bs.mie.2018.09.028
Palmer AG, Cavanagh J, Wright PE, Rance M (1991) Sensitivity improvement in proton-detected 2-dimensional heteronuclear correlation Nmr-spectroscopy. J Magn Reson 93:151–170. https://doi.org/10.1016/0022-2364(91)90036-S
Press WH, Teukolsky SA, Vetterling WT, Flannery BP (2002) Numerical recipes in C++ : the art of scientific computing, 2nd edn. Cambridge University Press, Cambridge
Raum HN, Dreydoppel M, Weininger U (2018) Conformational exchange of aromatic side chains by (1)H CPMG relaxation dispersion. J Biomol NMR 72:105–114. https://doi.org/10.1007/s10858-018-0210-5
Raum HN, Schorghuber J, Dreydoppel M, Lichtenecker RJ, Weininger U (2019) Site-selective (1)H/(2)H labeling enables artifact-free (1)H CPMG relaxation dispersion experiments in aromatic side chains. J Biomol NMR 73:633–639. https://doi.org/10.1007/s10858-019-00275-z
Schörghuber J et al (2018) Late metabolic precursors for selective aromatic residue labeling. J Biomol NMR 71:129–140. https://doi.org/10.1007/s10858-018-0188-z
Steiner E, Schlagnitweit J, Lundstrom P, Petzold K (2016) Capturing excited states in the fast-intermediate exchange limit in biological systems using (HNMR)-H-1 spectroscopy. Angew Chem Int Edit 55:15869–15872. https://doi.org/10.1002/anie.201609102
Teilum K, Brath U, Lundström P, Akke M (2006) Biosynthetic C-13 labeling of aromatic side chains in proteins for NMR relaxation measurements. J Am Chem Soc 128:2506–2507
Vallurupalli P, Bouvignies G, Kay LE (2012) Studying “Invisible” excited protein states in slow exchange with a major state conformation. J Am Chem Soc 134:8148–8161. https://doi.org/10.1021/ja3001419
Vallurupalli P, Hansen DF, Stollar E, Meirovitch E, Kay LE (2007) Measurement of bond vector orientations in invisible excited states of proteins. Proc Natl Acad Sci USA 104:18473–18477
Vallurupalli P, Sekhar A, Yuwen TR, Kay LE (2017) Probing conformational dynamics in biomolecules via chemical exchange saturation transfer: a primer. J Biomol NMR 67:243–271. https://doi.org/10.1007/s10858-017-0099-4
van de Ven FJM (1995) Multidimensional NMR in liquids: basic principles and experimental methods. Wiley, New York
Wagner G (1980) Activation volumes for the rotational motion of interior aromatic rings in globular-proteins determined by high-resolution H-1-Nmr at variable pressure. FEBS Lett 112:280–284
Wagner G, Demarco A, Wüthrich K (1976) Dynamics of aromatic amino-acid residues in globular conformation of basic pancreatic trypsin-inhibitor (Bpti) 1 H-1 Nmr-studies. Biophys Struct Mech 2:139–158
Weininger U et al (2013) Protein conformational exchange measured by H-1 R-1 rho relaxation dispersion of methyl groups. J Biomol NMR 57:47–55
Weininger U (2019) Optimal isotope labeling of aromatic amino acid side chains for NMR studies of protein dynamics. Methods Enzymol 614:67–86. https://doi.org/10.1016/bs.mie.2018.08.028
Weininger U, Diehl C, Akke M (2012a) C-13 relaxation experiments for aromatic side chains employing longitudinal- and transverse-relaxation optimized NMR spectroscopy. J Biomol NMR 53:181–190
Weininger U, Respondek M, Akke M (2012b) Conformational exchange of aromatic side chains characterized by L-optimized TROSY-selected C-13 CPMG relaxation dispersion. J Biomol NMR 54:9–14
Weininger U, Respondek M, Löw C, Akke M (2013) Slow aromatic ring flips detected despite near-degenerate NMR frequencies of the exchanging nuclei. J Phys Chem B 117:9241–9247
Weininger U, Brath U, Modig K, Teilum K, Akke M (2014a) Off-resonance rotating-frame relaxation dispersion experiment for C-13 in aromatic side chains using L-optimized TROSY-selection. J Biomol NMR 59:23–29
Weininger U, Modig K, Akke M (2014b) Ring flips revisited: C-13 relaxation dispersion measurements of aromatic side chain dynamics and activation barriers in basic pancreatic trypsin inhibitor. Biochemistry 53:4519–4525
Weininger U, Modig K, Geitner AJ, Schmidpeter PAM, Koch JR, Akke M (2017) Dynamics of aromatic side chains in the active site of FKBP12. Biochemistry 56:334–343. https://doi.org/10.1021/acs.biochem.6b01157
Acknowledgements
This research was supported by the Deutsche Forschungsgemeinschaft (WE 5587/1-2 to UW) and the Swedish Research Council (2018-4995 to MA).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dreydoppel, M., Lichtenecker, R.J., Akke, M. et al. 1H R1ρ relaxation dispersion experiments in aromatic side chains. J Biomol NMR 75, 383–392 (2021). https://doi.org/10.1007/s10858-021-00382-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-021-00382-w